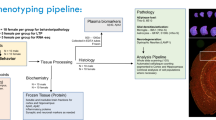
Abstract
Alzheimer’s disease (AD) and age-related cognitive decline represent a growing health burden and involve the hippocampus, a vulnerable brain region implicated in learning and memory. To understand the molecular effects of aging on the hippocampus, this study characterized the gene expression changes associated with aging in rodents using RNA-sequencing (RNA-seq). The glutamate modulator, riluzole, which was recently shown to improve memory performance in aged rats, prevented many of the hippocampal age-related gene expression changes. A comparison of the effects of riluzole in rats against human AD data sets revealed that many of the gene changes in AD are reversed by riluzole. Expression changes identified by RNA-Seq were validated by qRT–PCR open arrays. Riluzole is known to increase the glutamate transporter EAAT2’s ability to scavenge excess glutamate, regulating synaptic transmission. RNA-seq and immunohistochemistry confirmed an increase in EAAT2 expression in hippocampus, identifying a possible mechanism underlying the improved memory function after riluzole treatment.
Similar content being viewed by others
Introduction
Aging is associated with cognitive decline in humans, which impairs quality of life and contributes significantly to health-care costs.1 Similar declines are also observed in rodents and non-human primates.2 Aging is the primary risk factor for the dementia of Alzheimer’s disease (AD); and with significant increases in life expectancy, the prevalence of AD and age-related cognitive disorders is rising.3 The neural circuits affected in aging and AD are similar, involving the glutamatergic connections between cortical areas and with the hippocampal formation, a brain region in the medial temporal lobe that is critical for memory formation.4, 5, 6 The glutamatergic pyramidal neurons of the hippocampus are highly vulnerable to damage in both age-related cognitive decline and in AD.5, 7 However, the effects of glutamatergic modulation on aging remains unknown.
Riluzole is a glutamate modulator approved for treatment of amyotrophic lateral sclerosis.8 Importantly, riluzole treatment for 4 months prevented age-related cognitive decline in rodents through clustering of dendritic spines,9 which form the postsynaptic component of most excitatory synapses.10 Clustering of synaptic inputs is an important neuroplastic mechanism that increases synaptic strength, empowering neural circuits.11, 12 Riluzole’s ability to induce dendritic spines clustering,9 which is dependent on glutamatergic neuronal activity13, 14 and long-term potentiation (LTP),15 suggests that it regulates synaptic glutamatergic activity and prevents glutamate overflow to the extrasynaptic space. Synaptic N-methyl-D-aspartate (NMDA) activity is critical for LTP and memory formation, whereas extrasynaptic NMDA activation is associated with long-term depression (LTD) and excitotoxicity.16, 17, 18
The excitatory amino acid transporter 2 (EAAT2 or GLT-1; Slc1a2) is a high-affinity, Na+-dependent glutamate transporter and the dominant glutamate transporter in the brain.19, 20 Glutamate transporters, including EAAT2, decrease in aging21, 22 and AD,23, 24 and are associated with neurodegeneration.24 They also have a critical role in determining synaptic and extrasynaptic glutamate levels,20, 25 regulating physiological glutamatergic neurotransmission. Riluzole can act to stabilize the inactivated state of the voltage-gated sodium channel and it can increase EAAT2 expression,22, 26, 27, 28 potentiating glutamate uptake.28, 29, 30
Understanding the molecular vulnerabilities of glutamatergic neural circuits can point to novel and more effective treatment targets. In addition, molecular changes resulting from treatments that prevent cognitive decline remain largely unexplored. This study uses the combination of RNA-sequencing (RNA-seq) and open arrays to detect and validate specific molecular pathways that are changed by aging and with riluzole. Importantly, gene expression changes associated with a rescue of cognitive decline under a therapeutic intervention (riluzole) are identified. In addition, genes modulated by riluzole in the rat hippocampus are enriched in many of the same pathways, and in the opposite direction, as those altered in human AD gene expression data sets. These molecular transcriptional profiles in the aging hippocampus and with glutamatergic modulation by riluzole provide mechanistic insights into age-related cognitive decline and provide support for future studies on the role of glutamate transporters as potential therapeutic targets in the aging brain.
Materials and methods
Animals
Young (3-month-old) and aged male Sprague–Dawley rats (retired breeders, 10 months old, Harlan Laboratories (Harlan Sprague Dawley, Indianapolis, IN, USA) were housed at Rockefeller University for the duration of the experiments. All rats were pair-housed in climate-controlled conditions (30–50% humidity, 21±2 °C, 12 h-light/-dark cycle). Separate cohorts of animals were used for RNA extraction and immunohistochemical experiments. For RNA-seq experiments, each group had n=6: 3-month-old (young rats), 10-month-old (middle age rats), 14-month-old riluzole-treated rats and 14-month-old riluzole untreated rats. For immunohistochemistry, 3-month-old rats (n=10), 14-month-old untreated rats (n=9) and 14-month-old riluzole-treated rats (n=10) were used. All procedures were in agreement with the National Institutes of Health and The Rockefeller University Institutional Animal Care and Use Committee guidelines. Sample sizes were chosen to minimize the number of animals used given previously published reports using these methodologies.31, 32, 33
Riluzole treatment
Treated rats had ad libitum access to riluzole solution from 10 months to 14 months of age (17 weeks), and aged-control and young-control rats had ad libitum access to tap water. All rats had ad libitum access to food. The riluzole compound (Sigma-Aldrich, St. Louis, MO, USA) was dissolved in tap water at a concentration of 110 μg ml−1, translating to ∼4.0 mg kg−1 per day per os. To make the solution, riluzole was stirred in room temperature tap water for 6 h. All containers with riluzole were covered with foil to prevent light exposure. Fresh solutions were made every 2–3 days for the duration of treatment.
Tissue processing and immunohistochemistry
One week after the end of riluzole treatment, rats were deeply anesthetized with 100 mg kg−1 sodium pentobarbital and transcardially perfused with 1.0% paraformaldehyde in 0.1 M phosphate buffer (PB; 1 min) followed by 4.0% paraformaldehyde+0.125% glutaraldehyde in 0.1 M PB (12 min). Brains were removed and post-fixed for 6 h in 4.0% paraformaldehyde+0.125% glutaraldehyde in 0.1 M PB (4°C) and transferred to 0.1% sodium azide in PB (4°C) until cutting the following day. Brains were cut on a vibratome (Leica, VT1000S, Leica Biosystems, Buffalo Grove, IL, USA) into 40 μm coronal for immunohistochemistry. Sections were stored in 0.1% sodium azide in PB (4°C). Sections from each animal were washed with phosphate-buffered saline, blocked with 1% bovine serum albumin and incubated in primary antibody for GLT-1a (1:1000 dilution in phosphate-buffered saline; gift from J Rothstein’s Laboratory, Johns Hopkins University) overnight at 4 °C. The tissue was then washed in phosphate-buffered saline, and incubated with fluorescent secondary antibody (AlexaFluor488) for 1 h. The intensity of the labeling was quantified using Nikon Imaging software (Nikon Instruments, Melville, NY, USA) in 50 μm intervals from the cellular layer in each region of hippocampus. Electronic images were coded to blind the rater.
RNA extraction, sequencing and analysis
Wet dissected hippocampus from rapidly decapitated rats were flash frozen on dry ice and stored at −80°C. RNA was extracted using the RNAeasy Lipid Kit (QIAGEN Sciences, Germantown, MD, USA) and Qiacube as per the manufacturer’s instructions. Samples were pooled for sequencing and final RNA integrity was checked using the Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA) prior to library preparation. All samples had RNA Integrity Numbers >8. Sequencing libraries were prepared by the Rockefeller University Genomics Core Facility using the TruSeq RNA Library Preparation Kit v2 (Illumina, San Diego, CA, USA) and bar-coded for multiplexing so that all groups could be run in the same flow cell. Single-stranded reads of 100 bp were collected on a HiSeq 2500 (Illumina) at a sequencing depth of ~60 million reads per sample.
Raw data files were uploaded to Galaxy34, 35 and checked for integrity by FastQC. To remove sequencing artifacts and residual adapter sequences, reads were trimmed by 5–10 bp at the 5′ and 3′ ends and then filtered to remove reads with quality scores <20. Reads were aligned to the rat genome (rn5) using TopHat2 (ref. 36 and then loaded into Strand (Agilent) for quantification of read density by DESeq. Differential expression analyses were conducted in Strand using z-tests that were Benjamin–Hochberg corrected for false discovery rates. Venn diagrams and scatter plots based on significant gene lists were generated using Strand (STRand Analysis Software developed at University of California, Davis' Veterinary Genetics Lab, Davis, CA, USA) and Microsoft Excel (Redmond, WA, USA).
Open-array analysis
cDNA was synthesized using the VILO kit (Life Technologies, Carlsbad, CA, USA) using 2 μg of the same RNA submitted for sequencing for each reaction. Open-array plates were loaded from a 384-well plate as described in the standard Open Array protocol (Life Technologies) and run on a Quantstudio 12 k Flex thermocycler (Thermo Fisher Scientific, Waltham, MA, USA). Counts were exported to Microsoft Excel and used to calculate fold change using the ΔΔCt method.37 All values were normalized to pgk1 expression. Open-array fold change values were plotted against RNA-seq results to generate scatter plots and calculate the r2 values (Prism Software, Irvine, CA, USA).
Pathway analysis
Gene lists from RNA-Seq results were uploaded to the DAVID bioinformatics database (http://david.abcc.ncifcrf.gov/home.jsp). The functional clusters, enrichment scores and gene ontology terms for these categories were obtained from the functional annotation clustering tool. In all of the analysis, enrichment scores above 1.3 were considered significant (P<0.05; http://www.nature.com/nprot/journal/v4/n1/full/nprot.2008.211.html).
The enrichment scores from clusters with similar gene ontology terms were used to compare pathways that were altered in both riluzole and control conditions. Histograms were generated in Microsoft Excel.
Comparisons with previous human AD expression studies
AD expression data were analyzed from the GEO and AMP-AD databases and lists of significantly changed genes (P<0.05) were generated for each study. Some studies examined whole hippocampus,38, 39, 40, 41, 42 whereas others used laser capture microdissection to examine specific subregions of hippocampus, such CA1 and CA3 regions43 or CA1 alone,44 or dentate gyrus and entorhinal cortex.45 Therefore, to control for gene differences that arise from using different regions of the hippocampus and identify the most robust findings that were replicated across studies, only genes identified as significant in at least two studies were used for analysis. There were 2024 genes that were upregulated and 1870 genes that were downregulated with AD and met these criteria. These gene lists were analyzed with the DAVID functional annotation clustering tool as described above. The enrichment scores from these clusters were used to compare genes altered with AD to genes altered with riluzole treatment and the pathways with the highest combined enrichment scores are presented.
Genes significantly changed in aging studies in rats (P<0.05) were also analyzed using the GEO database. One study examined gene expression in the CA1 subregion at different points across the life span.46 Gene lists showing differential expression between 3-month- and 23-month-old rats, and between 3-month- and 12-month-old rats were obtained. A second study compared the genes from the whole hippocampus of 4–6-month-40 and 24–26-month-old rats.47 To control for the differences between regions of the hippocampus, only genes that were significant in both studies were used for analysis. There were 508 genes upregulated and 270 genes that were downregulated in aged rats. These gene lists were analyzed with DAVID functional annotation clustering tool as described above. The enrichment scores from these clusters were compared with the enrichment scores of similar clusters obtained from the 443 genes that were downregulated and the 674 genes that were upregulated with age in our RNA-seq data.
Results
Riluzole rescues age-related expression changes in rats
Hippocampal transcriptional profiles change markedly across the life span. In this study, rats between 3 months (young) and 10 months of age (middle-aged) showed 268 genes increased and 254 genes decreased. There were nearly twice as many changes from the 10-month- to 14-month-old (aged) rats, with 674 genes increased and 443 genes decreased, demonstrating that transcriptional changes are stable across adulthood, but accelerate from middle age onwards (Figure 1a; Supplementary Table 1).
Riluzole treatment of aged rats rescues age-related gene expression changes in the hippocampus. (a) Differential expression analysis revealed gene expression changes across age and with riluzole treatment. Between 10-month-old and 14-month-old rats (blue circle), 674 genes were upregulated and 443 genes downregulated. In all, 1480 genes were changed between age-matched riluzole-treated rats and controls (red circle), with 555 genes increased and 925 genes decreased. (b) Venn diagram illustrating the overlap of 435 genes that were changed by both aging (10–14 months; blue) and by riluzole treatment (red). (c) Scatter plot illustrating the 435 overlapping genes showing fold change by age (10–14 months; x-axis) against fold change with riluzole (y-axis). The upper left quadrant represent 96 genes that had decreased expression with age and increased expression after riluzole treatment. Conversely, the lower right quadrant illustrates 240 genes that were increased with age and decreased by riluzole. (d and e) Histograms illustrating significantly enriched pathways based on genes differentially expressed by either aging or riluzole treatment (enrichment score>1.3 reflects P<0.05). Similar pathways and enrichment scores were observed when comparing genes decreased by aging (yellow bars) and increased by riluzole (orange bars), as well as for genes increased with age (blue bars) and decreased by riluzole (green bars).
Animals treated with the glutamate modulator, riluzole, from 10 until 14 months had 908 genes increased and 927 genes decreased (Figure 1a). Importantly, there is a large overlap of genes (435) that were changed with aging and were also altered by riluzole treatment (Figure 1b). The overlapping genes were plotted to show fold change by age against fold change by riluzole treatment (Figure 1c). The lower right quadrant shows 240 genes that increased with age and were decreased with riluzole treatment. In the upper left quadrant, 96 genes that were decreased with age were increased with riluzole treatment. This profile suggests that riluzole treatment rescues many age-related gene expression changes in the hippocampus.
Differentially expressed genes were organized into functional pathways using the DAVID pathway tools. The pathway classes that were reversed by riluzole are ranked by significance and divided into those that were upregulated with age and downregulated by riluzole (Figure 1d) and pathways that were downregulated with age and upregulated by riluzole (Figure 1e). Many pathways reversed by riluzole treatment were related to synaptic transmission and plasticity. Examples of genes altered by aging that were reversed by riluzole are provided in Table 1. The NMDA receptor subunit NR2b (GRIN2b), a voltage-gated sodium channel subunit (Scn2a1), a calcium/calmodulin protein kinase II alpha (CAMK2A), the microtubule-associated protein 1B (MAP1B), the synaptic scaffolding protein enriched in the postsynaptic density of excitatory synapses SHANK3 and the matrix metalloproteinase 9 (MMP9), each decrease with aging and are increased by riluzole treatment, and have been implicated in learning and neuroplasticity.48, 49, 50, 51, 52, 53, 54 In contrast, isoforms of the GABA receptor (GABRA6) are found to increase with aging and are decreased by riluzole treatment and their blockage may improve memory consolidation.50, 55
Notably, several neuroprotective genes were increased with riluzole treatment, including tropomyosin receptor kinase B (TrkB; NTRK2), which is a receptor for brain-derived neurotrophic factor (Supplementary Table2).56 An example of a gene that was significantly decreased with treatment that is implicated in glutamate signaling is the cysteine/glutamate antiporter (xCT;Slc7a11; Supplementary Table 2). xCT exchanges extracellular cysteine for intracellular glutamate which contributes to the regulation of extrasynaptic glutamate levels57 and could be another potential mechanism for the action of riluzole.
Gene pathways implicated in AD are altered by riluzole
Hippocampal data from nine studies characterizing gene expression changes in post-mortem hippocampal tissue from AD patients,38, 39, 40, 41, 42, 43, 44, 45, 87 were obtained from the GEO (Gene Expression Omnibus, NCBI) and AMP-AD (Accelerating Medicines Partnership-Alzheimer’s Disease) databases. Differentially expressed genes identified in at least two of the nine studies were analyzed in DAVID for comparison with the present pathway results (Figure 2a; Supplementary Tables 3). Many of the pathways altered across studies in AD are also changed with riluzole treatment, including ones related to transmission of nerve impulse and synaptic plasticity (Figure 2b). Examples of genes altered in both AD tissue and by riluzole treatment in rats can be found in Table 2. Importantly, the glutamate transporter EAAT2 is significantly decreased in AD, as well as in aging rats, and was rescued by riluzole treatment. Several genes previously implicated in neural transmission and plasticity are diminished in AD and recovered by riluzole treatment, including: ANK3, an integral membrane protein to the underlying spectrin-actin cytoskeleton that mediates synaptic morphology and transmission;58 CAMK2, a calcium/calmodulin kinase protein and major component of the postsynaptic density that is critically involved in induction of synaptic potentiation and memory,51, 59 and Rab3A, a vesicular trafficking protein that is crucial for synaptic plasticity, learning and memory.60, 61
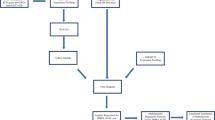
Gene pathways changed by riluzole in aged rats are similar and in the opposite direction to those changed in post-mortem AD brains. (a) Schematic representation of the bioinformatics strategy used to generate pathway lists. Only genes significantly differentially expressed in at least two studies were included. (b) Significantly enriched pathways derived from genes upregulated by riluzole (light green bars) were similar to gene pathways downregulated in AD brains (dark green bars). Gene pathways downregulated by riluzole (light blue bars) also showed similarity to pathways that are increased in AD (dark blue bars). Differentially expressed gene lists from AD brains were derived from publically available data posted in GEO and AMP-AD and subjected to pathway analysis using DAVID (enrichment score>1.3 reflects P<0.05).
Validation of RNA-seq results
Custom open-array technology allows for high-throughput qRT–PCR analysis of gene expression. Fifty-three genes of interest were selected to assay, given their known roles in neuroplasticity, glutamate signaling, learning and memory. Thirty-nine of these 53 genes were significant between at least one RNA-seq condition (Supplementary Table 5). A strong correlation in fold change was observed between genes identified as significant by RNA-seq and the open-array measurements, with three of the comparisons exhibiting r2 values >0.80 (Figure 3a–d). Conversely, no significant differences in gene expression were observed in the open-array analysis that were not already identified as significant by RNA-seq. This concordance, of both the positive and negative data for these genes, demonstrates that our sequencing analysis reflects reliable changes in gene expression.
Expression differences determined by qRT–PCR are highly correlated with RNA-seq results. Scatter plots illustrating fold change levels determined by qRT–PCR open arrays (x-axis) plotted against fold change level calculated from RNA-seq analysis (y-axis) for each comparison group (a) 3m vs 10m, (b) 10m vs 14m, (c) 10m vs 14m riluzole, (d) 14m control vs riluzole. Only genes that reached significance between each condition in the RNA-seq analyses are represented in the scatter plots (Benjamin–Hochberg corrected P<0.05). Several genes of interest and r2 values for each comparison are highlighted.
To further validate the RNA-seq findings, results were compared against previous reports using microarray technology to study gene expression changes with aging in rat hippocampus.46, 47 Differentially expressed gene lists from rat studies were obtained using the GEO database and analyzed for pathway enrichment using DAVID. Many of the same gene pathways were changed in both the RNA-seq data and previously published reports of aged rats (Supplementary Figure 1). This is despite differences in the exact age of the animals, tissue collection techniques and the different array technologies used across these studies. Together, the bioinformatics and open-array results suggest that the RNA-seq data are representative of age- and riluzole-induced gene expression changes.
Riluzole rescues EAAT2 levels after aging
Riluzole is known to increase EAAT2 expression,22, 26, 27, 28 which helps maintain the correct amount of glutamate in the synaptic cleft.20 Failure of EAAT2 leads to glutamate spillover to the extrasynaptic space, which can cause decreased synaptic efficiency, LTD and excitotoxicity.17, 62 EAAT2 is expressed in neurons, axon terminals and glial cells.63, 64, 65, 66, 67 Previous studies have shown that EAAT2 is decreased with aging and AD.21, 24, 23 RNA-seq results from our rodent experiments confirm that EAAT2 is downregulated with age, but importantly, levels of this gene are rescued by riluzole treatment (Figure 4a). Further, we identified increased immunohistochemical labeling for EAAT2 in the distal portion of CA1 (Figures 4b), which confirms the RNA-seq findings, and suggests a potential mechanism by which riluzole may rescue cognitive function to be further validated in future studies. Importantly, this region corresponds to the area in which increased dendritic spine clustering occurred in response to riluzole treatment.9
Riluzole increases EAAT2 expression. (a) Normalized expression values from RNA-seq data for EAAT2 (y-axis) show that gene expression decreases with age (x-axis) but is restored by riluzole treatment. (b) Quantification of fluorescent intensity (y-axis) of CA1 hippocampal sections labeled for EAAT2. Riluzole significantly increased labeling in the region 150–200 μm from the pyramidal cell bodies in aged rats (P<0.05). (c) Example images from 14-month-old controls and treated with riluzole. Red arrows indicate regions of difference.
Discussion
This study reveals gene expression changes that occur with aging and glutamatergic modulation by riluzole in the rat hippocampus. The majority of transcriptional changes identified occurred from middle-aged to aged rats, rather than from young to middle-aged animals, suggesting that a loss of the transcriptional stability during adulthood occurs with aging (Figure 1a). Importantly, riluzole treatment reversed many of the age-related expression changes in the rat hippocampus (Figures 1b), which primarily occurred in pathways associated with synaptic function (Figures 1d). A similar inverse comparison of the pathways changed by riluzole with those identified in the hippocampus of AD patients demonstrated extensive commonality of affected genes (Figure 2 and Table 2), establishing riluzole’s potential as a therapeutic agent for AD. The expression changes identified by sequencing were highly correlated with qRT–PCR results using open-array technology, which validated over 50 genes of interest (Figure 3). Finally, changes in the levels of EAAT2, a gene known to be increased by riluzole treatment,22, 26, 27 were validated by immunohistochemical labeling in hippocampus (Figure 4). This finding raises the possibility that modulation of glutamate transporters is one mechanism by which riluzole can improve cognitive performance in aging. In addition, these high-throughput studies offer an essential library of new targets that warrant further investigation into their role in glutamatergic transmission in the hippocampus and age-related cognitive decline.
The identification of pathways associated with the maintenance of synaptic health as changed by riluzole (Figures 1d; Table 1) is consistent with previous work that demonstrated riluzole prevented age-related cognitive decline through clustering of dendritic spines,9 an important neuroplastic mechanism that has been shown by electrophysiological studies and computational models to allow non-linear summation of synaptic inputs.11, 68 Some examples of genes implicated in learning and plasticity that are reduced in aging and increased by riluzole treatment include: MMP9, which induces structural spine modifications;54 NR2B, which is important for LTP;31 MAP1B, which helps maintainaince of structural plasticity in the adult brain52 and SHANK3, which has an important role in synaptic regulation.53
Many of riluzole’s effects on the aging rat hippocampus were opposite to the changes observed in human hippocampus from AD patients (Figure 2b). This indicates that many of the key pathways altered by glutamate modulation with riluzole are implicated in the development of AD pathology, suggesting riluzole may have therapeutic potential. Notably, as in aging, the top AD pathways identified also involved synaptic transmission and plasticity and were reversed by riluzole. Some examples of genes recovered by riluzole treatment include: ANK3, which mediates synaptic morphology and transmission;58 CAMK2, involved in induction of synaptic potentiation and memory51, 59 and Rab3A, which is crucial for synaptic plasticity, learning and memory.60, 61 Genes that are consistently altered in AD brains and are reversed by riluzole provide potential future therapeutic targets.
Synaptic dysfunction is a critical pathophysiological mechanism in AD that highly correlates with cognitive decline.69, 70 Amyloid-β (Aβ) and phosphorylated tau toxicities, the hallmarks of the neuropathology of AD, are intimately related to glutamatergic dysregulation, whereby oligomers of Aβ disrupt glutamate uptake, inhibiting LTP through excessive activation of extrasynaptic NMDA receptors.71, 72 Oligomers of Aβ also facilitate LTD71, 73 and decrease surface expression of synaptic NMDA receptors.74 In addition, dysregulated glutamate increases release of Aβ75 and tau76, 77 and enhances tau phosphorylation78 and expression,79 forming a vicious cycle of neurotoxicity. Importantly, previous work has shown that EAAT2 haploinsufficiency accelerates cognitive deficits in an AD mouse model (AβPPswe/PS1ΔE9)80 and EAAT2 overexpression improves cognitive and pathological markers in APPSw,Ind AD mouse model.81 These studies support the hypothesis that improved regulation of glutamatergic signaling via enhanced EAAT2 uptake could potentially mitigate toxicities in AD brains.
Finally, increased immunoreactivity for EAAT2 was observed in the same region as increased spine clustering was previously identified in riluzole-treated rats,9 suggesting a potential mechanism by which riluzole can increase cognitive performance. Glutamate transporters have the key role of regulating synaptic transmission, and thereby learning and memory.20, 25 They prevent glutamate spillover to the extrasynaptic space and minimize cross-talk between neighboring synapses.20, 62 Importantly, they also control the time course of synaptic glutamate.82, 83 More recent work suggests that EAAT2 surface trafficking also shapes synaptic transmission.84 Previous studies have suggested that riluzole increases glutamate uptake through both increased EAAT2 expression and stabilization of the inactivated state of voltage-gated sodium channels.22, 29, 30, 26,28 A recent in vivo study using microelectrode arrays coupled with amperometry has shown that riluzole reduces extrasynaptic glutamate levels and enhances cognitive performance which correlated with the increased glutamate uptake measures.27 These mechanisms of riluzole have been hypothesized to facilitate synaptic glutamatergic activity and to increase glutamate–glutamine cycling while preventing glutamate overflow to the extrasynaptic space.26, 85, 86 Activation of extrasynaptic NMDA receptors has been associated with LTD and excitotoxicity, and it is a likely important mechanism in many neurodegenerative diseases, including AD.17, 18
In conclusion, these findings identify molecular pathways implicated in aging that are rescued by administration of a known glutamate modulator, riluzole. Modeling the expression differences in response to riluzole establishes a framework of changes associated with improved learning and memory, against which other treatments can be compared. Further, many of the pathways changed by riluzole have been implicated across multiple studies in the pathophysiology of AD, suggesting glutamate modulators may represent novel treatments for both age-related cognitive decline and AD. Future studies will seek to conclusively demonstrate whether increased expression and activity of glutamate transporters are the essential mechanism underlying riluzole’s ability to improve cognitive performance.
References
Buckner RL . Memory and executive function in aging and AD: multiple factors that cause decline and reserve factors that compensate. Neuron 2004; 44: 195–208.
Burke SN, Barnes CA . Neural plasticity in the ageing brain. Nat Rev Neurosci 2006; 7: 30–40.
Brookmeyer R, Johnson E, Ziegler-Graham K, Arrighi HM . Forecasting the global burden of Alzheimer's disease. Alzheimers Dement 2007; 3: 186–191.
Morrison JH, Hof PR . Life and death of neurons in the aging brain. Science 1997; 278: 412–419.
Morrison JH, Hof PR . Selective vulnerability of corticocortical and hippocampal circuits in aging and Alzheimer's disease. Prog Brain Res 2002; 136: 467–486.
Neves G, Cooke SF, Bliss TV . Synaptic plasticity, memory and the hippocampus: a neural network approach to causality. Nat Rev Neurosci 2008; 9: 65–75.
Braak H, Braak E . Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol 1991; 82: 239–259.
Bensimon G, Lacomblez L, Meininger V . A controlled trial of riluzole in amyotrophic lateral sclerosis. ALS/Riluzole Study Group. N Engl J Med 1994; 330: 585–591.
Pereira AC, Lambert HK, Grossman YS, Dumitriu D, Waldman R, Jannetty SK et al. Glutamatergic regulation prevents hippocampal-dependent age-related cognitive decline through dendritic spine clustering. Proc Natl Acad Sci USA 2014; 111: 18733–18738.
Grutzendler J, Kasthuri N, Gan WB . Long-term dendritic spine stability in the adult cortex. Nature 2002; 420: 812–816.
Larkum ME, Nevian T . Synaptic clustering by dendritic signalling mechanisms. Curr Opin Neurobiol 2008; 18: 321–331.
Polsky A, Mel BW, Schiller J . Computational subunits in thin dendrites of pyramidal cells. Nat Neurosci 2004; 7: 621–627.
Kavalali ET, Klingauf J, Tsien RW . Activity-dependent regulation of synaptic clustering in a hippocampal culture system. Proc Natl Acad Sci USA 1999; 96: 12893–12900.
Kleindienst T, Winnubst J, Roth-Alpermann C, Bonhoeffer T, Lohmann C . Activity-dependent clustering of functional synaptic inputs on developing hippocampal dendrites. Neuron 2011; 72: 1012–1024.
De Roo M, Klauser P, Muller D . LTP promotes a selective long-term stabilization and clustering of dendritic spines. PLoS Biol 2008; 6: e219.
Hardingham GE . Pro-survival signalling from the NMDA receptor. Biochem Soc Trans 2006; 34: 936–938.
Hardingham GE, Bading H . Synaptic versus extrasynaptic NMDA receptor signalling: implications for neurodegenerative disorders. Nat Rev Neurosci 2010; 11: 682–696.
Rusakov DA, Kullmann DM . Extrasynaptic glutamate diffusion in the hippocampus: ultrastructural constraints, uptake, and receptor activation. J Neurosci 1998; 18: 3158–3170.
Furuta A, Rothstein JD, Martin LJ . Glutamate transporter protein subtypes are expressed differentially during rat CNS development. J Neurosci 1997; 17: 8363–8375.
Tzingounis AV, Wadiche JI . Glutamate transporters: confining runaway excitation by shaping synaptic transmission. Nat Rev Neurosci 2007; 8: 935–947.
Potier B, Billard JM, Riviere S, Sinet PM, Denis I, Champeil-Potokar G et al. Reduction in glutamate uptake is associated with extrasynaptic NMDA and metabotropic glutamate receptor activation at the hippocampal CA1 synapse of aged rats. Aging Cell 2010; 9: 722–735.
Brothers HM, Bardou I, Hopp SC, Kaercher RM, Corona AW, Fenn AM et al. Riluzole partially rescues age-associated, but not LPS-induced, loss of glutamate transporters and spatial memory. J Neuroimmune Pharmacol 2013; 8: 1098–1105.
Jacob CP, Koutsilieri E, Bartl J, Neuen-Jacob E, Arzberger T, Zander N et al. Alterations in expression of glutamatergic transporters and receptors in sporadic Alzheimer's disease. J Alzheimers Dis 2007; 11: 97–116.
Masliah E, Alford M, DeTeresa R, Mallory M, Hansen L . Deficient glutamate transport is associated with neurodegeneration in Alzheimer's disease. Ann Neurol 1996; 40: 759–766.
Huang YH, Bergles DE . Glutamate transporters bring competition to the synapse. Curr Opin Neurobiol 2004; 14: 346–352.
Banasr M, Chowdhury GM, Terwilliger R, Newton SS, Duman RS, Behar KL et al. Glial pathology in an animal model of depression: reversal of stress-induced cellular, metabolic and behavioral deficits by the glutamate-modulating drug riluzole. Mol Psychiatry 2010; 15: 501–511.
Hunsberger HC, Weitzner DS, Rudy CC, Hickman JE, Libell EM, Speer RR et al. Riluzole rescues glutamate alterations, cognitive deficits, and tau pathology associated with P301L tau expression. J Neurochem 2015; 135: 381–394.
Gourley SL, Espitia JW, Sanacora G, Taylor JR . Antidepressant-like properties of oral riluzole and utility of incentive disengagement models of depression in mice. Psychopharmacology 2012; 219: 805–814.
Frizzo ME, Dall'Onder LP, Dalcin KB, Souza DO . Riluzole enhances glutamate uptake in rat astrocyte cultures. Cell Mol Neurobiol 2004; 24: 123–128.
Fumagalli E, Funicello M, Rauen T, Gobbi M, Mennini T . Riluzole enhances the activity of glutamate transporters GLAST, GLT1 and EAAC1. Eur J Pharmacol 2008; 578: 171–176.
Faherty SL, Campbell CR, Larsen PA, Yoder AD . Evaluating whole transcriptome amplification for gene profiling experiments using RNA-Seq. BMC Biotechnol 2015; 15: 65.
Maag JL, Panja D, Sporild I, Patil S, Kaczorowski DC, Bramham CR et al. Dynamic expression of long noncoding RNAs and repeat elements in synaptic plasticity. Front Neurosci 2015; 9: 351.
Gray JD, Rubin TG, Hunter RG, McEwen BS . Hippocampal gene expression changes underlying stress sensitization and recovery. Mol Psychiatry 2014; 19: 1171–1178.
Goecks J, Nekrutenko A, Taylor J . Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol 2010; 11: R86.
Blankenberg D, Von Kuster G, Coraor N, Ananda G, Lazarus R, Mangan M et al. Galaxy: a web-based genome analysis tool for experimentalists. Curr Protoc Mol Biol 2010; Chapter 19: 11–21.
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL . TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 2013; 14: R36.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001; 25: 402–408.
Hokama M, Oka S, Leon J, Ninomiya T, Honda H, Sasaki K et al. Altered expression of diabetes-related genes in Alzheimer's disease brains: the Hisayama study. Cereb Cortex 2014; 24: 2476–2488.
Liang WS, Dunckley T, Beach TG, Grover A, Mastroeni D, Walker DG et al. Gene expression profiles in anatomically and functionally distinct regions of the normal aged human brain. Physiol Genomics 2007; 28: 311–322.
Berchtold NC, Coleman PD, Cribbs DH, Rogers J, Gillen DL, Cotman CW . Synaptic genes are extensively downregulated across multiple brain regions in normal human aging and Alzheimer's disease. Neurobiol Aging 2013; 34: 1653–1661.
Silva AR, Grinberg LT, Farfel JM, Diniz BS, Lima LA, Silva PJ et al. Transcriptional alterations related to neuropathology and clinical manifestation of Alzheimer's disease. PLoS One 2012; 7: e48751.
Xu PT, Li YJ, Qin XJ, Kroner C, Green-Odlum A, Xu H et al. A SAGE study of apolipoprotein E3/3, E3/4 and E4/4 allele-specific gene expression in hippocampus in Alzheimer disease. Mol Cell Neurosci 2007; 36: 313–331.
Miller JA, Woltjer RL, Goodenbour JM, Horvath S, Geschwind DH . Genes and pathways underlying regional and cell type changes in Alzheimer's disease. Genome Med 2013; 5: 48.
Blalock EM, Buechel HM, Popovic J, Geddes JW, Landfield PW . Microarray analyses of laser-captured hippocampus reveal distinct gray and white matter signatures associated with incipient Alzheimer's disease. J Chem Neuroanat 2011; 42: 118–126.
Small SA, Kent K, Pierce A, Leung C, Kang MS, Okada H et al. Model-guided microarray implicates the retromer complex in Alzheimer's disease. Ann Neurol 2005; 58: 909–919.
Kadish I, Thibault O, Blalock EM, Chen KC, Gant JC, Porter NM et al. Hippocampal and cognitive aging across the lifespan: a bioenergetic shift precedes and increased cholesterol trafficking parallels memory impairment. J Neurosci 2009; 29: 1805–1816.
Rowe WB, Blalock EM, Chen KC, Kadish I, Wang D, Barrett JE et al. Hippocampal expression analyses reveal selective association of immediate-early, neuroenergetic, and myelinogenic pathways with cognitive impairment in aged rats. J Neurosci 2007; 27: 3098–3110.
Cao X, Cui Z, Feng R, Tang YP, Qin Z, Mei B et al. Maintenance of superior learning and memory function in NR2B transgenic mice during ageing. Eur J Neurosci 2007; 25: 1815–1822.
Nayak TK, Sikdar SK . Time-dependent molecular memory in single voltage-gated sodium channel. J Membr Biol 2007; 219: 19–36.
Shen K, Teruel MN, Connor JH, Shenolikar S, Meyer T . Molecular memory by reversible translocation of calcium/calmodulin-dependent protein kinase II. Nat Neurosci 2000; 3: 881–886.
Glazewski S, Giese KP, Silva A, Fox K . The role of alpha-CaMKII autophosphorylation in neocortical experience-dependent plasticity. Nat Neurosci 2000; 3: 911–918.
Nothias F, Fischer I, Murray M, Mirman S, Vincent JD . Expression of a phosphorylated isoform of MAP1B is maintained in adult central nervous system areas that retain capacity for structural plasticity. J Comp Neurol 1996; 368: 317–334.
Wang X, McCoy PA, Rodriguiz RM, Pan Y, Je HS, Roberts AC et al. Synaptic dysfunction and abnormal behaviors in mice lacking major isoforms of Shank3. Hum Mol Genet 2011; 20: 3093–3108.
Dziembowska M, Wlodarczyk J . MMP9: a novel function in synaptic plasticity. Int J Biochem Cell Biol 2012; 44: 709–713.
Kim DH, Kim JM, Park SJ, Cai M, Liu X, Lee S et al. GABA(A) receptor blockade enhances memory consolidation by increasing hippocampal BDNF levels. Neuropsychopharmacology 2012; 37: 422–433.
Klein R, Nanduri V, Jing SA, Lamballe F, Tapley P, Bryant S et al. The trkB tyrosine protein kinase is a receptor for brain-derived neurotrophic factor and neurotrophin-3. Cell 1991; 66: 395–403.
Massie A, Boillee S, Hewett S, Knackstedt L, Lewerenz J . Main path and byways: non-vesicular glutamate release by system x as an important modifier of glutamatergic neurotransmission. J Neurochem 2015; 135: 1062–1079.
Smith KR, Kopeikina KJ, Fawcett-Patel JM, Leaderbrand K, Gao R, Schurmann B et al. Psychiatric risk factor ANK3/ankyrin-G nanodomains regulate the structure and function of glutamatergic synapses. Neuron 2014; 84: 399–415.
Lisman J, Schulman H, Cline H . The molecular basis of CaMKII function in synaptic and behavioural memory. Nat Rev Neurosci 2002; 3: 175–190.
Thakker-Varia S, Alder J, Crozier RA, Plummer MR, Black IB . Rab3A is required for brain-derived neurotrophic factor-induced synaptic plasticity: transcriptional analysis at the population and single-cell levels. J Neurosci 2001; 21: 6782–6790.
Castillo PE, Janz R, Sudhof TC, Tzounopoulos T, Malenka RC, Nicoll RA . Rab3A is essential for mossy fibre long-term potentiation in the hippocampus. Nature 1997; 388: 590–593.
Asztely F, Erdemli G, Kullmann DM . Extrasynaptic glutamate spillover in the hippocampus: dependence on temperature and the role of active glutamate uptake. Neuron 1997; 18: 281–293.
Furness DN, Dehnes Y, Akhtar AQ, Rossi DJ, Hamann M, Grutle NJ et al. A quantitative assessment of glutamate uptake into hippocampal synaptic terminals and astrocytes: new insights into a neuronal role for excitatory amino acid transporter 2 (EAAT2). Neuroscience 2008; 157: 80–94.
Petr GT, Sun Y, Frederick NM, Zhou Y, Dhamne SC, Hameed MQ et al. Conditional deletion of the glutamate transporter GLT-1 reveals that astrocytic GLT-1 protects against fatal epilepsy while neuronal GLT-1 contributes significantly to glutamate uptake into synaptosomes. J Neurosci 2015; 35: 5187–5201.
Chen W, Mahadomrongkul V, Berger UV, Bassan M, DeSilva T, Tanaka K et al. The glutamate transporter GLT1a is expressed in excitatory axon terminals of mature hippocampal neurons. J Neurosci 2004; 24: 1136–1148.
Danbolt NC, Storm-Mathisen J, Kanner BI . An [Na++K+]coupled L-glutamate transporter purified from rat brain is located in glial cell processes. Neuroscience 1992; 51: 295–310.
Lehre KP, Levy LM, Ottersen OP, Storm-Mathisen J, Danbolt NC . Differential expression of two glial glutamate transporters in the rat brain: quantitative and immunocytochemical observations. J Neurosci 1995; 15: 1835–1853.
Govindarajan A, Kelleher RJ, Tonegawa S . A clustered plasticity model of long-term memory engrams. Nat Rev Neurosci 2006; 7: 575–583.
Selkoe DJ . Alzheimer's disease is a synaptic failure. Science 2002; 298: 789–791.
DeKosky ST, Scheff SW . Synapse loss in frontal cortex biopsies in Alzheimer's disease: correlation with cognitive severity. Ann Neurol 1990; 27: 457–464.
Li S, Hong S, Shepardson NE, Walsh DM, Shankar GM, Selkoe D . Soluble oligomers of amyloid Beta protein facilitate hippocampal long-term depression by disrupting neuronal glutamate uptake. Neuron 2009; 62: 788–801.
Li S, Jin M, Koeglsperger T, Shepardson NE, Shankar GM, Selkoe DJ . Soluble Abeta oligomers inhibit long-term potentiation through a mechanism involving excessive activation of extrasynaptic NR2B-containing NMDA receptors. J Neurosci 2011; 31: 6627–6638.
Cheng L, Yin WJ, Zhang JF, Qi JS . Amyloid beta-protein fragments 25-35 and 31-35 potentiate long-term depression in hippocampal CA1 region of rats in vivo. Synapse 2009; 63: 206–214.
Snyder EM, Nong Y, Almeida CG, Paul S, Moran T, Choi EY et al. Regulation of NMDA receptor trafficking by amyloid-beta. Nat Neurosci 2005; 8: 1051–1058.
Kamenetz F, Tomita T, Hsieh H, Seabrook G, Borchelt D, Iwatsubo T et al. APP processing and synaptic function. Neuron 2003; 37: 925–937.
Yamada K, Holth JK, Liao F, Stewart FR, Mahan TE, Jiang H et al. Neuronal activity regulates extracellular tau in vivo. J Exp Med 2014; 211: 387–393.
Pooler AM, Phillips EC, Lau DH, Noble W, Hanger DP . Physiological release of endogenous tau is stimulated by neuronal activity. EMBO Rep 2013; 14: 389–394.
Sindou P, Lesort M, Couratier P, Yardin C, Esclaire F, Hugon J . Glutamate increases tau phosphorylation in primary neuronal cultures from fetal rat cerebral cortex. Brain Res 1994; 646: 124–128.
Esclaire F, Lesort M, Blanchard C, Hugon J . Glutamate toxicity enhances tau gene expression in neuronal cultures. J Neurosci Res 1997; 49: 309–318.
Mookherjee P, Green PS, Watson GS, Marques MA, Tanaka K, Meeker KD et al. GLT-1 loss accelerates cognitive deficit onset in an Alzheimer's disease animal model. J Alzheimers Dis 2011; 26: 447–455.
Takahashi K, Kong Q, Lin Y, Stouffer N, Schulte DA, Lai L et al. Restored glial glutamate transporter EAAT2 function as a potential therapeutic approach for Alzheimer's disease. J Exp Med 2015; 212: 319–332.
Mennerick S, Zorumski CF . Glial contributions to excitatory neurotransmission in cultured hippocampal cells. Nature 1994; 368: 59–62.
Tong G, Jahr CE . Block of glutamate transporters potentiates postsynaptic excitation. Neuron 1994; 13: 1195–1203.
Murphy-Royal C, Dupuis JP, Varela JA, Panatier A, Pinson B, Baufreton J et al. Surface diffusion of astrocytic glutamate transporters shapes synaptic transmission. Nat Neurosci 2015; 18: 219–226.
Chowdhury GM, Banasr M, de Graaf RA, Rothman DL, Behar KL, Sanacora G . Chronic riluzole treatment increases glucose metabolism in rat prefrontal cortex and hippocampus. J Cereb Blood Flow Metab 2008; 28: 1892–1897.
Brennan BP, Hudson JI, Jensen JE, McCarthy J, Roberts JL, Prescot AP et al. Rapid enhancement of glutamatergic neurotransmission in bipolar depression following treatment with riluzole. Neuropsychopharmacology 2010; 35: 834–846.
Data obtained from the Accelerating Medicines Partnership for Alzheimer's Disease (AMP-AD) Target Discovery Consortium data portal and can be accessed at HYPERLINK. http://dx.doi.org/doi:10.7303/syn2580853; doi:10.7303/syn2580853.
Acknowledgements
This work was supported by DANA Foundation, the Rockefeller University Women & Science Initiative and Alzheimer’s Drug Discovery Foundation to ACP, NIH grant F32 MH102065 to JDG, NIA grant R37 AG06647 to JHM and partial support by grant # UL1 TR000043 from the National Center for Research Resources and the National Center for Advancing Translational Sciences (NCATS).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Rights and permissions
About this article
Cite this article
Pereira, A., Gray, J., Kogan, J. et al. Age and Alzheimer’s disease gene expression profiles reversed by the glutamate modulator riluzole. Mol Psychiatry 22, 296–305 (2017). https://doi.org/10.1038/mp.2016.33
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2016.33
- Springer Nature Limited
This article is cited by
-
Formulating treatment of major psychiatric disorders: algorithm targets the dominantly affected brain cell-types
Discover Mental Health (2023)
-
High-fat diet induces depression-like phenotype via astrocyte-mediated hyperactivation of ventral hippocampal glutamatergic afferents to the nucleus accumbens
Molecular Psychiatry (2022)
-
The stressed synapse 2.0: pathophysiological mechanisms in stress-related neuropsychiatric disorders
Nature Reviews Neuroscience (2022)
-
WNT/β-catenin Pathway: a Possible Link Between Hypertension and Alzheimer’s Disease
Current Hypertension Reports (2022)
-
Cell type-specific potential pathogenic genes and functional pathways in Alzheimer’s Disease
BMC Neurology (2021)