Abstract
The main objective of this study is to evaluate the quality of wastewater by molecular identification of enteroviruses, rotaviruses, and adenoviruses in wastewater samples collected from the ElSerw wastewater treatment facility in Damietta Governorate, Egypt. An additional objective is to assess the usefulness of these viruses as markers of viral reduction during wastewater treatment. A treatment facility's inflow and discharge were sampled 48 times. The incidence of enteric viruses was found in 29 wastewater samples (60.4%). 6.25% (3/48), 0% (0/48), 37.5% (18/48), and 20.8% (10/48) of the samples tested positive for enteroviruses (EVs), noroviruses, rotaviruses, and adenoviruses, respectively. Co-infections with two or more viruses were found in 10.4% (5/48) and 2% (1/48) of all cases, respectively. The viral burden in the wastewater treatment plant's discharge effluents dropped non-significantly when compared to intake samples. According to our findings, rotaviruses and adenoviruses have been found in 10 outlet effluent samples. The removal rates for enteroviruses, rotaviruses and adenoviruses were 39%, 61.5% and 33.3%, respectively. As a result of their high frequency and lower removal rates, both rotaviruses and adenoviruses were deemed an appropriate indicator of human enteric viral reduction during the wastewater treatment process.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Egypt's water resources are becoming increasingly scarce in order to meet public drinking and agricultural water needs. Reused wastewater is increasingly being used for agricultural reasons all over the globe as an efficient way to conserve water resources [1, 2]. Pathogens such as enteric viruses and bacteria are common in repurposed wastewater, posing a health risk to subjects, land, animals, and consumers of products irrigated with treated wastewater. Over 150 distinct enteric viruses have been connected to water-borne and food-borne diseases [3]. According to up-to-date World Health Organization (WHO) figures, the annual mortality rate linked with diarrhea among Egyptian infants hit 30 deaths per 100,000. Rotavirus infection was responsible for nearly 3.9% of all documented deaths, while intestinal adenovirus was the third most common viral cause of diarrhea after rotaviruses and noroviruses [4]. There is no single suitable measure that can demonstrate complete reduction of human enteric viruses in wastewater treatment facilities [5, 6]. Conventional bacterial indicators cannot be used to identify the frequency and decline of human enteric viruses during wastewater treatment due to the negligible link between indicators and viruses [7,8,9]. As a consequence, identifying optimal viruses that meet all of the criteria for a viral decline measure is essential [10, 11]. Adenoviruses [12], polyomaviruses [13], F-specific RNA coliphages [14], pepper moderate mottle virus [15], and tobacco mosaic virus [16] have all been proposed as indicators of human enteric viral decline during effluent treatment. However, each of these viruses is insufficient to connect the presence of all human enteric viruses and is unsuitable for evaluating wastewater treatment efficacy [17, 18].
In particular, over 100 species of enteric viruses have been identified as common water pollutants, and the number is expanding due to the emergence of new strains. Indicators are widely used to examine the destiny of pathogenic strains due to the diversity of viruses in the environment. Previously, fecal bacterial indicators (FIB) such as coliform bacteria, Escherichia coli, Enterococcus and Streptococcus spp. were utilized to measure fecal contamination levels in water. Bacteria, on the other hand, have been proven to be significantly less resistant to wastewater treatment and far less environmentally durable than enteric viruses [19,20,21]. As a result, FIB are poor indicators of viral infection risk, meaning that present surveillance programs focused solely on FIB are inadequate. A suitable viral indicator for wastewater contamination evaluation should, ideally, have similar inactivation and retention to the target pathogens and be present in wastewater and wastewater-contaminated environments throughout the year. This would allow for continuous monitoring and would offer information on the level of contamination and the possibility of pathogen presence [22, 23]. Table 1 describes certain enteric viruses found in wastewater that have the potential to be used as indicators; however, not all of these viruses fit these criteria. For example, Respiratory viruses and papillomaviruses have been detected in high concentrations in wastewater but not in polluted environments, which might be due to the rapid destruction of these viruses in water. Furthermore, some enteric viruses, such as hepatitis and Rotaviruses, may be zoonotic, which means that their presence in the environment is caused by agricultural operations rather than human waste. Additionally, enteroviruses, noroviruses, and sapoviruses have substantial seasonality in temperate locations, with peaks in the summer or winter [24]. As a result, these viruses are not found in wastewater or contaminated areas throughout the year [25, 26]. Human adenoviruses (AdVs) and polyomaviruses (PyVs) are often found in polluted environments, and their use as useful fecal indicators has been suggested [27, 28]. As a result, additional research is needed to determine if a single or many suitable markers may be employed as fecal indicators. Thus, based on the results of a one-year monthly monitoring for four human enteric viruses; enteroviruses, noroviruses, rotaviruses, and adenoviruses at a wastewater treatment facility, this study evaluates dependable viruses that can indicate the reduction of human enteric viruses during the wastewater treatment process.
2 Materials and methods
2.1 Samples collection
Wastewater samples were collected at the El-Serw wastewater treatment plant (WWTP) in Damietta, Egypt's influent (raw sewage) and outflow (processed effluent). From January to December 2019, samples were collected every two weeks for a year. One litre of the influent (raw sewage) and outflow (treated effluent) was collected in a clean plastic container and sent in an ice box to Egypt's Environmental virology laboratory, Environment and Climate Change Institute, National Research Centre within 5–6 h. 48 wastewater samples were collected during the study's period.
2.2 Samples processing, nucleic acid extraction, and cDNA synthesis
Using the adsorption–elution method, each sample was filtered individually through a nitrocellulose membrane (0.45 m pore size and 147 mm diameter). According to Katzenelson et al. [29], adsorption viruses have been isolated by 3% beef extract and re-concentrated by organic flocculation. The pellet obtained was dissolved in 1 milliliter of Na2HPO4 (0.14 N, pH 9) and stored at − 70 °C until use. Total viral DNA and RNA were extracted with the QIAamp DNA kit (QIAGEN, Germany) and BioZOL solution (BIOFLUX, Japan), respectively according to the manufacturer’s instruction. DNA and RNA were eluted in 100 µl of elution buffer and stored at −70 °C until use. Nanodrop Spectrophotometer (A260/280 ratio) was used to determine the concentration and quality of the isolated RNA or DNA. The RevertAid RT Reverse Transcription Kit (K1691, ThermoFisher Scientific) and reverse primers specific to viral target were used to create the cDNA.
2.3 Polymerase chain reaction amplification
In a final amount of 50 µl, 100 ng of virus DNA or cDNA, 2 × PCR buffer, 1.5 mM MgCl2, 200 M of dNTPs, 0.5 M of each outer primer, and 5 U of Taq DNA polymerase were used for PCR amplification. The original PCR phase was 95 °C for 10 min, followed by 35 cycles of 94 °C for 30 s, 55 °C for 60 s, and 72 °C for 60 s, followed by 1 cycle of 72 °C for 7 min to extend the reaction using a Bio-Rad PCR platform. Table 2 contains the primer sequences for all specific viruses.
2.4 PCR product sequencing
PCR products were purified using the QIAquick purification kit (Qiagen, Germany) and sequenced directly by used the same primers in the PCR. Consensus sequences were matched to the existing sequences in the NCBI nucleotide collection database library using BLASTN program, http://blast.ncbi.nlm.nih.gov/Blast.cgi.
3 Results and discussion
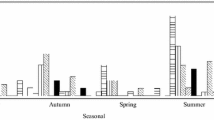
Enteric viruses, such as rotaviruses, enteroviruses, noroviruses and adenoviruses, are significant public health concerns due to their high infectivity and extended survival in the environment. Contaminated water has been identified as one of the primary transmission pathways for diarrheal diseases, highlighting the importance of reducing viral loads prior to effluent release into the environment, especially prior to water reuse [34, 35]. The purpose of this study was to look into the presence of rotaviruses, enteroviruses, noroviruses, and adenoviruses in an Egyptian wastewater treatment facility, as well as to assess the plant's ability to remove viruses. As shown in Table 3 and Fig. 1, rotaviruses were found in wastewater samples with a positive ratio of 37.5% (18/48) followed by adenoviruses with a positive ratio of up to 20.8% (10/48) and enteroviruses with a positive ratio of 6.25% (3/48); however, these ratios were found to be different with those observed in earlier Egyptian studies conducted in environmental or clinical samples [36,37,38,39,40,41,42]. Indeed, the positive ratio for rotaviruses (37.5%) in the current investigation was similar to that observed in previous Egyptian studies on clinical samples (31%), but higher than that observed in environmental samples (8.3–18.75%) [37, 38, 40]. The current study's viral positive rate (20.8%) was higher than in clinical and environmental samples (6.7–8.9%), [39, 41]. Our enterovirus detection rate (6.25%) was lower than the previous study in Egyptian sewage (22%). Variations in incidence rates may be linked to differences in geographical locations, such as rural versus urban areas, population size and society, and other environmental variables. In the current study, considerable levels of positive for enteroviruses, rotaviruses, and adenoviruses were discovered in the majority of intake samples, showing a high prevalence of enteric viruses in raw sewage. Our findings are consistent with previous research from Canada, the United States, and South Africa, where significant numbers of enteric viruses were discovered in raw sewage [18, 43, 44].
Figure 2 depicts the elimination ratios of human enteric viruses throughout the wastewater treatment process, with nearly one-third of the positive adenovirus and enterovirus ratios being adenovirus-negative and enterovirus-negative at final output discharge samples. At final discharge effluent tests, nearly half of the positive rotavirus ratio was rotavirus-negative. Although several methods for dealing with non-detects have been suggested, any of them can result in an overestimation or underestimation of the mean value [45]. The decline rates of the four human enteric viruses studied varied from 33 to 60%, which was lower than previous study findings [46]. Adenovirus removal ratios showed no notable variation, whereas enterovirus and rotavirus removal ratios were considerably higher than adenovirus removal ratios. To demonstrate that they are eliminated more effectively than the indicator virus(es) during the effluent treatment process, a virus(es) with a lower removal ratio than other viruses must be identified. Because of the high frequency of rotaviruses and the high resilience to wastewater treatment processes of adenoviruses, both rotaviruses and adenoviruses are excellent candidates to be a sign of fecal contamination, according to our findings.
4 Conclusion
The viral burden in the wastewater treatment plant's discharge effluents dropped non-significantly when compared to intake samples. As a result of their high frequency and lower removal rates, both rotaviruses and adenoviruses were deemed an appropriate indicator of human enteric viral reduction during the wastewater treatment process. Thus, rotaviruses and adenoviruses were considered suitable markers of human enteric virus removal during the wastewater treatment process.
Data availability
The data generated and/or examined during the present study are not publicly available due to an ongoing research endeavor but are available upon reasonable request from the corresponding author.
References
Al-Saed R (2007) Pathogens assessment in reclaimed effluent used for industrial crops irrigation. Int J Environ Res Public Health 4(1):68–75
Alshammari MS, Derafaa W, Elshaygi EA (2021) Removal of heavy metals from groundwater using silica/activated carbon composite. Desalin Water Treat 238:198–206
Ibrahim C, Cherif N, Hammami S, Pothier P, Hassen A (2016) Quantification and genotyping of rotavirus A within two wastewater treatment processes. Clean: Soil, Air, Water 44(4):393–401
Neuzil KM, Armah GE, Parashar UD, Duncan Steele A (2010) Rotavirus in Africa: shifting the focus to disease prevention. The University of Chicago Press, Chicago
Kitajima M, Iker BC, Pepper IL, Gerba IL (2014) Relative abundance and treatment reduction of viruses during wastewater treatment processes identification of potential viral indicators. Sci Total Environ 488:290–296
Hamza IA, Jurzik L, Uberla K, Wilhelm M (2011) Evaluation of pepper mild mottle virus, human picobirnavirus and torque teno virus as indicators of fecal contamination in river water. Water Res 45:1358–1368
Ottoson J, Hansen A, Björlenius B, Norder H, Stenström TA (2006) Removal of viruses, parasitic protozoa and microbial indicators in conventional and membrane processes in a wastewater pilot plant. Water Res 40:1449–1457
Savichtcheva O, Okabe S (2006) Alternative indicators of fecal pollution: relations with pathogens and conventional indicators, current methodologies for direct pathogen monitoring and future application perspectives. Water Res 40:2463–2476
Hewitt J, Greening GE, Leonard M, Lewis GD (2013) Evaluation of human adenovirus and human polyomavirus as indicators of human sewage contamination in the aquatic environment. Water Res 47:6750–6761
Gerba CP, Betancourt WQ, Kitajima M, Rock CM (2018) Reducing uncertainty in estimating virus reduction by advanced water treatment processes. Water Res 133:282–288
Bibby K, Crank K, Greaves J, Li X, Wu Z, Hamza IA (2019) Metagenomics and the development of viral water quality tools. NPJ Clean Water 2:9
Fong TT, Phanikumar MS, Xagoraraki I, Rose JB (2010) Quantitative detection of human adenoviruses in wastewater and combined sewer overflows influencing a Michigan River. Appl Environ Microbiol 76:715–723
Bofill-Mas S, Albinana-Gimenez N, Clemente-Casares P, Hundesa A, Rodriguez-Manzano J, Allard A et al (2006) Quantification and stability of human adenoviruses and polyomavirus JCPyV in wastewater matrices. Appl Environ Microbiol 72(12):7894–7896
Haramoto E, Fujino S, Otagiri M (2015) Distinct behaviors of infectious F-specific RNA coliphage genogroups at a wastewater treatment plant. Sci Total Environ 520:32–38
Hata A, Kitajima M, Katayama H (2013) Occurrence and reduction of human viruses, F-specific RNA coliphage genogroups and microbial indicators at a full-scale wastewater treatment plant in Japan. J Appl Microbiol 114:545–554
Kitajima M, Sassi HP, Torrey JR (2018) Pepper mild mottle virus as a water quality indicator. NPJ Clean Water 1:19
Shrestha S, Shrestha S, Shindo J, Sherchand JB, Haramoto E (2018) Virological quality of irrigation water sources and pepper mild mottle virus and tobacco mosaic virus as index of pathogenic virus contamination level. Food Environ Virol 10:107–120
Qiu Y, Lee BE, Neumann N, Ashbolt N, Craik S, Maal-Bared R et al (2015) Assessment of human virus removal during municipal wastewater treatment in Edmonton, Canada. J Appl Microbiol 119(6):1729–1739
Lin J, Ganesh A (2013) Water quality indicators: bacteria, coliphages, enteric viruses. Int J Environ Health Res 23(6):484–506. https://doi.org/10.1080/09603123.2013.769201
Prez VE, Gil PI, Temprana CF, Cuadrado PR, Martínez LC, Giordano MO, Masachessi G, Isa MB, Ré VE, Paván JV, Nates SV, Barril PA (2015) Quantification of human infection risk caused by rotavirus in surface waters from Córdoba, Argentina. Sci Total Environ 538:220–229
Sidhu JPS, Ahmed W, Palmer A, Smith K, Hodgers L, Toze S (2017) Optimization of sampling strategy to determine pathogen removal efficacy of activated sludge treatment plant. Environ Sci Pollut Res 24:19001–19010
Scott TM, Rose JB, Jenkins TM, Farrah SR, Lukasik J (2002) Microbial source tracking: current methodology and future directions. Appl Environ Microbiol 68:5796–5809
Xagoraraki I, O’Brien E (2020) Wastewater-based epidemiology for early detection of viral outbreaks. In: O’Bannon DJ (ed) Women in water quality. Michigan State University, UNESCO, East Lansing, pp 75–97
Bosch A, Pinto RM, Guix S (2016) Foodborne viruses. Curr Opin Food Sci 8:110–119
Farkas K, Cooper DM, McDonald JE, Malham SK, de Rougemont A, Jones DL, de Rougemont A, Jones DL, de Rougemont A, Jones DL (2018) Seasonal and spatial dynamics of enteric viruses in wastewater and in riverine and estuarine receiving waters. Sci Total Environ 634:1174–1183
Farkas K, Adriaenssens EM, Walker DI, Mcdonald JE, Malham SK, Jones DL (2019) Critical evaluation of crAssphage as a molecular marker for human-derived wastewater contamination in the aquatic environment. Food Environ Virol 11:113–119
Farkas K, Mannion F, Hillary LS, Malham SK, Walker DI (2020) Emerging technologies for the rapid detection of enteric viruses in the aquatic environment. Curr Opin Environ Sci Health 16:1–6
Fattal B, Vasl RJ, Katzenelson E, Shuval HI (1983) Survival of bacterial indicator organisms and enteric viruses in the Mediterranean coastal waters off Tel-Aviv. Water Res 17:397–402
Katzenelson E, Fattal B, Hostovesky T (1976) Organic flocculation: an efficient second step concentration method for the detection of viruses in tap water. Appl Environ Microbiol 32(4):638
Biscaro V, Piccinelli G, Gargiulo F, Ianiro G, Caruso A, Caccuri F, De Francesco MA (2018) Detection and molecular characterization of enteric viruses in children with acute gastroenteritis in Northern Italy. Infect Genet Evol 60:35–41. https://doi.org/10.1016/j.meegid.2018.02.011
Schreier E, Doring F, Kunkel U (2000) Molecular epidemiology of outbreaks of gastroenteritis associated with small round structured viruses in Germany in 1997/98. Arch Virol 145:443–453
Gray J, Iturriza-Gómara M (2011) Rotaviruses. Methods Mol Biol 665:325–355
Puig M, Jofre J, Lucena F, Allard A, Wadell G, Girones R (1994) Detection of adenoviruses and enteroviruses in polluted waters by nested PCR amplification. Appl Environ Microbiol 60:2963–2970
Cheng H-WA, Lucy FE, Broaders MA, Mastitsky SE, Chen C-H, Murray A (2012) Municipal wastewater treatment plants as pathogen removal systems and as a contamination source of noroviruses and Enterococcus faecalis. J Water Health 10(3):380–389
Ahmed SM, Hall AJ, Robinson AE, Verhoef L, Premkumar P, Parashar UD et al (2014) Global prevalence of norovirus in cases of gastroenteritis: a systematic review and meta-analysis. Lancet Infect Dis 14(8):725–730
Allayeh AK, Al-Daim SA, Ahmed N, El-Gayar M, Mostafa A (2022) Isolation and genotyping of adenoviruses from wastewater and diarrheal samples in Egypt from 2016 to 2020. Viruses 14:2192
El-Gayar MH, Saleh SE, Mohamed AF, Aboulwafa MM, Hassouna NA, Allayeh AK (2022) Isolation, propagation and genotyping of human rotaviruses circulating among children with gastroenteritis in two Egyptian university hospitals. Biology 11:1413
Rizk NM, Allayeh AK (2018) Multiplex semi-nested RT-PCR for genotyping of rotaviruses group A in Giza tap water, Egypt. Asian J Water Environ Pollut 15:217–221
Allayeh AK, ElBaz RM, Saeed NM, ElSayed OM (2018) Detection and genotyping of viral gastroenteritis in hospitalized children below five years old in Cairo, Egypt. Arch Pediatr Infect Dis 6(3):e60288
Rizk NM, Allayeh AK (2020) Genotyping of rotaviruses in River Nile in Giza, Egypt, Iran. Iran J Public Health 49(1):173–180
Gad MA, Allayeh AK, Elmahdy EM, Shaheen MNF, Rizk NM, Al-Herrawy AZ, Saleh FER, Marouf MA (2019) Genotyping and interaction-reality of Acanthamoeba, enteric adenovirus and rotavirus in drinking water, Egypt. Egypt J Aquat Biol Fish 23:65–79
Kamel AH, Ali MA, El-Nady HG, Aho S, Pothier P, Belliot G (2010) Evidence of the co-circulation of enteric viruses in sewage and in the population of Greater Cairo. J Appl Microbiol 108(5):1620–1629
Symonds EM, Griffin DW, Breitbart M (2009) Eukaryotic viruses in wastewater samples from the United States. Appl Environ Microbiol 75(5):1402–1409
Adefisoye MA, Nwodo UU, Green E, Okoh AI (2016) Quantitative PCR detection and characterization of human adenovirus, rotavirus and hepatitis A virus in discharged effluents of two wastewater treatment facilities in the Eastern Cape, South Africa. Food Environ Virol 8(4):262–274
Wood MD, Beresford NA, Copplestone D (2011) Limit of detection values in data analysis: do they matter? Radioprotection 46:S85–S90
Tandukar S, Sherchan SP, Haramoto E (2020) Reduction of human enteric and indicator viruses at a wastewater treatment plant in Southern Louisiana, USA. Food Environ Virol 12:260–263. https://doi.org/10.1007/s12560-020-09433-1
Funding
There was no funding available for the study.
Author information
Authors and Affiliations
Contributions
RS: writing, editing and visualization, reviewing. AA: conceptualization, methodology, writing, editing and visualization.
Corresponding author
Ethics declarations
Conflict of interest
The authors state that they do not have any known competing financial interests or personal ties that could appear to have influenced the work described in this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Soltane, R., Allayeh, A.K. Occurrence of enteroviruses, noroviruses, rotaviruses, and adenoviruses in a wastewater treatment plant. J.Umm Al-Qura Univ. Appll. Sci. 9, 449–454 (2023). https://doi.org/10.1007/s43994-023-00053-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43994-023-00053-6