Abstract
As an important industrial enzyme, protease is widely used in feed, food and other fields. At present, the insufficient protease activity obtained from microorganisms cannot meet the purpose of industrial production. In this study, Bacillus amyloliquefaciens with high protease production was screened from animal feces by plate transparent circle method. To improve the production of protease, atmospheric room temperature plasma (ARTP) mutagenesis was used in the first round, protease activity reached 315.0 U/mL. Then, to enhance production of protease, 60Co-γ irradiation was used for combined mutagenesis, leading to protease activity of B. amyloliquefaciens FMME ZK003 up to 355.0 U/mL. Furthermore, to realize the efficient production of protease, after optimization of fermentation conditions, protease activity was increased to 456.9 U/mL. Finally, protease activity of B. amyloliquefaciens FMME ZK003 reached 823.0 U/mL in a 5 L fermenter. These results indicate that B. amyloliquefaciens can efficiently produce protease, which provides a good foundation for the industrial production of protease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Protease can catalyze the hydrolysis of protein to generate polypeptides and small amino acids [1, 2], which is widely used in food [3, 4], feed [5, 6], detergent [7, 8], leather [9], and textile [10]. Currently, protease can be produced through two different routes: chemical extraction and microbial fermentation [1, 11, 12]. However, chemical extraction of protease from animals [13, 14] and plants [15] is a complex process with many bottleneck problems, such as high cost, low efficiency and complex purification process [1]. To overcome these problems, protease production by microbial fermentation is regarded as a promising method due to its cheap and renewable raw materials and simplified downstream purification process [16,17,18,19,20]. Naturally occurring producers have been isolated and optimized to produce protease, such as Bacillus subtilis [21], Bacillus sphaericus [22], Bacillus licheniformis [12], Rhodotorula mucilaginosa [23] and Mucor spp. [24]. However, protease activity obtained from these microorganisms is not high enough to be suitable for scale-up production.
Two strategies have been investigated for producing protease in microbial fermentation. Strategy (i) is physical or chemical mutagenesis methods. These methods were used to obtain mutants with high protease production, such as ultraviolet (UV) [25], laser [26], ray [27], ethyl methane sulfonate (EMS) [28], and nitrosoguanidine (NTG) [29]. For example, the activity of protease produced by B. licheniformis was increased by 3.50 times to 357.2 U/mL by combined UV and laser mutagenesis [26]. Strategy (ii) is genetic modification. Genetic engineering was used to improve microbial production of protease [30,31,32,33]. For example, the expression of alkaline protease gene (aprE) was optimized by screening the most efficient expression system, and thus protease activity was improved by 62.2% [32]. On the basis of the above two methods, fermentation optimization can be continued to improve the protease activity. The activity of protease was enhanced by optimizing fermentation conditions [34,35,36]. For example, by optimizing the carbon and nitrogen sources of B. subtilis fermentation medium, protease activity reached 125.0 U/mL [35]. However, there are some problems in these methods, such as low positive mutation rate, heavy metabolic burden and complex fermentation process, so that protease activity is not high enough to be used in industrial production.
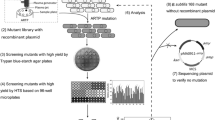
In this study, Bacillus amyloliquefaciens with high protease production was screened by plate transparent circle method. On this basis, atmospheric room temperature plasma (ARTP) and 60Co-γ irradiation were combined to mutagenesis to improve protease production. After optimization of fermentation conditions, protease activity of B. amyloliquefaciens FMME ZK003 reached 823.0 U/mL in a 5 L fermenter. This will lay a good foundation for the efficient production of protease by microbial fermentation.
Materials and methods
Materials
Twenty samples of animal feces (pig, chicken, and cattle) were collected from farms around Wuxi city.
Medium
The casein screening medium was composed of 4 g/L casein dissolved in 20 mL 0.1 mol/L NaOH solution, 20 g/L agar. Medium A was composed of 10 g/L beef extract, 10 g/L tryptone, and 10 g/L NaCl. Luria–Bertani (LB) medium was composed of 10 g/L tryptone, 5 g/L yeast extract, and 10 g/L NaCl. Medium B was composed of 60 g/L glucose, 60 g/L tryptone, 0.5 g/L CaCl2, and 4 g/L Na2HPO4·12H2O. Medium C was composed of 10 g/L yeast extract, 5 g/L tryptone, 5 g/L glucose, and 2 g/L NaCl.
Culture conditions
Seed cultures were prepared in a 250 mL shake–flask containing 50 mL of media. Seed cultures were grown for 16 h at 30℃ with shaking at 200 rpm. The shake flask fermentation conditions are as follows: the seed cultures were transferred as 10% inoculum to a 500 mL shake flask containing 100 mL fermentation medium and then cultivated at 30℃ with shaking at 200 rpm for 1–3 days. The 5 L fermenter fermentation conditions are as follows: fermentation in a 5 L fermenter were conducted with 2.5 L working volumes, containing inoculated at 10%. Airflow was set at 1.5 vvm, and the dissolved oxygen concentration was controlled above 20% saturation by agitation cascade, and pH was maintained at 7.5 by the automatic addition of ammonium hydroxide.
Screening protease-producing strains
During the enrichment process, 1.0 g of the sample was added into a 250 mL shake flask containing sterile water at 30℃ with shaking at 200 rpm for 1 h.
In the preliminary screening, the enrichment solution was diluted at 10 times ratio, and the 10−4, 10−5, 10−6, 10−7, 10−8 folds enrichment solution were, respectively, coated on casein screening plate, which cultured at 30℃ for more than 24 h. Strains that produce protease will appear white transparent circles on the screening plate, while those that do not produce protease will not. Therefore, the potential microorganism range can be determined according to whether there is a white transparent circle.
In the secondary screening, strains with protein degradation ability were isolated and selected by screening medium. The ratio of the diameter of the transparent circle to the diameter of the colony (H/d value) as in Eq. 1 was positively correlated with the protease activity of the strain. The H/d value in the same time was calculated by selecting the isolated strains to the casein screening plate. Strains with the highest ratio were selected for subsequent research.
In the identification of strain species, first, FastPure Bacteria DNA Isolation Mini Kit (Vazyme Biotech Co.,ltd) was used for bacterial genome extraction. The bacterial genome was used as a template, and 27F and 1492R universal primers were used as upstream and downstream primers for PCR amplification. The PCR products were sequenced by GENEWIZ. Then sequence and phylogenetic analysis were performed. BLAST system (http://www.ncbi.nlm.nih.gov/BLAST) was used to analyze the homology of 16S rDNA. Multiple sequence alignment using software DNAMAN, and finally Mega 3.1 software were used to construct phylogenetic tree.
ARTP and 60Co-γ irradiation
The B. amyloliquefaciens FMME ZK001 was cultured at 30 ℃ in LB medium for around 16 h, and then 1 mL of cells was washed three times with phosphate-buffered saline. Finally, the strain was suspended in 1 mL of phosphate-buffered saline and diluted to an OD600 of 3.0 using a gradient.
The prepared 10 μL bacterial suspension was placed in the center of the sterilized metal substrate and placed in an ARTP breeder to induce mutagenesis. The mutagenesis conditions are shown in Table 1. The cultured specimens were placed into a 1.5 mL centrifuge tube containing 990 μL seed medium, and the cells were oscillated with a vortex oscillator for at least 2 min to completely elute the bacteria solution. Then they were transferred into a 500 mL shake flask containing 50 mL seed medium and cultured at 30 ℃ at 200 rpm for 12 h. After proper dilution, 100 μL was spread on the screening solid medium and incubated at 30 ℃ for 12–36 h. The mutant with better protease production ability was selected for further mutagenesis.
For the mutation of 60Co-γ irradiation, we chose irradiation dosage of 0.4, 0.6, 0.8 and 0.9 kGy to irradiate strain suspension, and the other operational parameters were performed as described previously [37]. The irradiated bacterial suspension was diluted appropriately, and 100 μL was applied to solid screening medium, and incubated at 30 ℃ for 12–36 h. First, solid plate screening was carried out, followed by 24 pore plate and shake flask fermentation verification. The protease activity of the obtained fermentation broth was determined, and the strain with the highest enzyme activity was the best mutant screened. The positive mutation rate of mutant was calculated according to the activity of protease produced by fermentation. The mutant with higher protease activity than the starting strain was defined as positive mutant. The positive mutant ratio was defined as Eq. 2.
Protease activity assay
Based on National standards of the People’s Republic of China GB/T 23,527—2009, the activity of protease was assayed by folin reagent method and modified appropriately. The enzyme activity unit was defined as the enzyme amount required to hydrolyze casein to produce 1 μg tyrosine per mL of enzyme solution within 1 min at 40 ℃ and pH 7.5, expressed as U/mL.
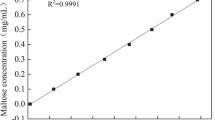
Determination of tyrosine standard curve: absorb 1 mL tyrosine solution (0, 10, 20, 40, 50, 60, 80, 90, 100, 110 μg/mL, respectively), add 5 mL 0.4 mol/L sodium carbonate solution and 1 mL folin reagent, and react in water bath at 40 ℃ for 20 min. The absorbance value at 660 nm was measured by UV–visible spectrophotometer, and the tyrosine standard curve was drawn according to the data.
First, the fermentation broth was pretreated, centrifuged at 4 ℃ at 5000 rpm for 10 min, and the supernatant was filtered by 0.22 μm filter membrane to prepare crude enzyme broth. Then, 1% casein solution were preheated at 40 ℃ for 5 min, and 1 mL casein solution and 2 mL crude enzyme solution were mixed evenly, no enzyme solution was added to the blank tube. The reaction was accurate at 40 ℃ for 30 min. The enzymatic reaction was terminated with 2 mL 10% trichloroacetic acid solution, the reaction solution was left for 5 min, and centrifuged at 5000 rpm at 4 ℃ for 10 min. Take 1 mL centrifuged supernatant, add 5 mL 0.4 mol/L sodium carbonate solution and 1 mL folin reagent in turn, mix well and then take water bath at 40 ℃ for 20 min. After cooling to room temperature, use UV–visible spectrophotometer to measure the absorbance value of 660 nm reaction solution. The protease activity was defined as Eq. 3, A, sample absorption value; N, dilution ratio of enzyme solution, κ the slope of the tyrosine standard curve, V volume of enzyme solution (mL), 5 the sample was diluted 5 times, 30 the reaction time (min).
Protease properties
Effect of temperature on protease activity and stability. The protease activity of crude enzyme solution was measured at 35, 40, 45, 50, 55 and 60 °C. The crude enzyme solution was kept at 35, 45, 55 and 65 °C for 20, 40, 60, 80 and 100 min, respectively, and the residual protease activity was measured at 40 °C.
Effect of pH on protease activity and stability. Different buffer solutions: Na2HPO4–NaH2PO4 and Tris–HCl were used to control pH at 6.0–8.0 and 8.0–9.0, respectively. The crude enzyme solution was reacted with substrates with pH values of 6.0, 6.5, 7.0, 7.5, 8.0, 8.5, 9.0 at 40 °C, and protease activity was measured. The crude enzyme solution was placed in buffer solutions with pH values of 6.5, 7.0, 7.5, 8.0 and 8.5, respectively, and the residual protease activity was measured at 40 °C.
Analytical methods
The OD600 was measured using a spectrophotometer. Glucose was quantified by the M-100 Biosensors Analyzer (Shenzhen Sieman Technology Co., Ltd).
Data analysis
Values are shown as mean ± SD (standard deviation) from three biological replicate experiments.
Results and discussion
Screening protease-producing Bacillus amyloliquefaciens
To obtain protease producing strains, we collected animal feces samples from farms around Wuxi city. A total of 200 protease producing strains were obtained from 908 colonies by plate transparent circle method. To preliminarily evaluate protease production, the ratio of the diameter of the transparent circle to the diameter of the colony (H/d value) was used for preliminary screening and 153 strains were obtained with H/d values ranging from 0.3 to 4.7 (Fig. 1a). To obtain the best strain for protease production, the H/d value was further used for screening. 33 strains with H/d value between 2.0 and 4.8 were obtained, among which the H/d value of strain 153 was 4.7, while the H/d value of other strains was lower than 4.0 (Fig. 1b). These results showed that the protein degradation ability of strain 153 was significantly better than the other strains. An optimal protease producing strain 153 was obtained from animal feces samples using transparent circle preliminary screening and H/d value secondary screening method.
16S rDNA method was used to identify the species of strain 153. With the help of the bacterial genome extraction kit, the genome of strain 153 was extracted and amplified by PCR to obtain 16S rDNA of the strain. Agarose gel electrophoresis detected that the target band size was 1000–2000 bp (Fig. 1c), and the 16S rDNA sequencing result was 1400 bp. According to gene sequencing results, BLAST sequence alignment in NCBI database showed that the 16S rDNA of strain 153 was 100% similar to that of Bacillus amyloliquefaciens. On this basis, the phylogenetic tree of strain 153 was constructed by Mega 3.1 software (Fig. 1d). In summary, strain 153 was identified as B. amyloliquefaciens, named B. amyloliquefaciens FMME ZK001. Previous studies have shown that the high protease producing strain was screened from the soil around maar lake by milk plate screening method [38]. A protease-producing strain was isolated from 30 protease-producing strains, and further identified as B. licheniformis by physiological and biochemical characterization, 16S rDNA gene sequence and phylogenetic analysis [38]. In our study, the sample size was larger and the scope was wider when screening protease-producing strains. The results are more reliable through the combination of plate preliminary screening and H/d value secondary screening method. Similarly, 16S rDNA gene sequence and phylogenetic analysis were also used to identify the strain species.
Improving protease production with ARTP mutagenesis
To obtain high protease producing strain, B. amyloliquefaciens FMME ZK001 was treated with ARTP mutagenesis. First, the treatment time of ARTP mutagenesis needs to be optimized. The optimal mutagenesis time was determined by analyzing the positive mutation rate in each period. Positive mutants mainly appeared in the mutants that were treated with ARTP for 90 s and 120 s, indicating that the treatment time was the optimal condition for ARTP mutagenesis in this study (Fig. 2a). On this basis, B. amyloliquefaciens FMME ZK001 was used as the starting strain for ARTP mutagenesis for 90 s and 120 s. For the preliminary screening stage, according to protease activity at 24 h, 180 mutants growing on the screening plate were screened, and 53 mutants were obtained, among which 33 strains had higher enzyme activity than B. amyloliquefaciens FMME ZK001. The positive mutation rate was 18.3% (Fig. 2a). For the secondary screening stage, protease activity of all 33 positive mutants was higher than that of B. amyloliquefaciens FMME ZK001, and the protease activity of 24 positive mutants increased greatly (Fig. 2b). The protease activity, glucose consumption and OD600 of mutant Z-27 reached 315.0 U/mL, 25.0 g/L and 4.5, respectively, at 24 h (Fig. 2b, c, d). B. amyloliquefaciens FMME ZK001 was mutated by ARTP and a high protease-producing strain Z-27 was obtained, which was named B. amyloliquefaciens FMME ZK002. ARTP mutagenesis has been widely used in Aspergillus oryzae [39, 40] and genus Bacillus [41] to obtain stable and high-producing strains of protease or other products. For example, ARTP mutagenesis was employed to treat spores of Aspergillus oryzae strain 3.042 for selection of high protease producers with an irradiation time of 150 s, the result showed that neutral protease was increased by 17.3% [39]. The protease activity of B. licheniformis TP1 mutated by ARTP was 56.0% higher than that of the original strain [41]. In our study, the mutant B. amyloliquefaciens FMME ZK002 was obtained by mutagenesis through ARTP with an irradiation time of 90 s. The protease activity of B. amyloliquefaciens FMME ZK002 increased from 230 to 315, which was 37.0% higher than that of B. amyloliquefaciens FMME ZK001.
Preliminary and secondary screening results of the first round for mutants with ARTP mutagenesis. a Protease producing mutants obtained by preliminary screening after ARTP mutagenesis under different irradiation time. b Protease activity in secondary screening. c Glucose consumption in secondary screening. d OD600 in secondary screening. The dashed line represents control strain B. amyloliquefaciens FMME ZK001, the asterisk represents optimal mutant
Enhancing protease production by 60Co-γ irradiation
To enhance protease production of B. amyloliquefaciens FMME ZK002, 60Co-γ irradiation was used for compound mutagenesis. First, the irradiation dose of 60Co-γ irradiation was optimized, and the optimal irradiation dose was 0.6 kGy. On this basis, B. amyloliquefaciens FMME ZK002 was used as the starting strain for 60Co-γ irradiation with 0.6 kGy irradiation dose. At preliminary screening, according to protease activity at 24 h, 240 mutants growing on the screening plate were screened, and 120 mutants were obtained. The protease activity of 50 strains was higher than that of B. amyloliquefaciens FMME ZK002, and the positive mutation rate was 20.8% (Fig. 3a). At secondary screening, all positive mutants were screened in shake flask fermentation, and 22 positive mutants had higher protease activity. Among them, the protease activity of mutant E-6 was the most significant increase, and the shake flask level reached 355.0 U/mL (Fig. 3b). Meanwhile, glucose consumption and OD600 reached 29.7 g/L and 5.1, respectively (Fig. 3c, d). The protease production of the strain was enhanced after 60Co-γ irradiation combined mutagenesis, and the optimal mutant E-6 was named B. amyloliquefaciens FMME ZK003. Previous studies have shown that compound mutagenesis combined with 60Co-γ irradiation significantly increases protease production levels [27]. The positive mutation rate calculated by the above two mutagenesis methods shows that the optimization of mutagenesis conditions can improve the positive mutation rate. At the same time, fermentation in shake flasks can be used to screen high protease-producing B. amyloliquefaciens, but the workload is large and the efficiency is not high. To further improve the screening efficiency and positive mutation rate, it is still necessary to establish an efficient screening method [42].
Preliminary and secondary screening results of second round for mutants with 60Co-γ irradiation. a Protease producing mutants obtained by preliminary screening after 60Co-γ irradiation. b Protease activity in secondary screening. c Glucose consumption in secondary screening. d OD600 in secondary screening. The dashed line represents control strain B. amyloliquefaciens FMME ZK002, the asterisk represents optimal mutant
Evaluating protease production of B. amyloliquefaciens
To realize the industrial production of protease, we first analyzed the properties of protease. To evaluate the effect of temperature on protease activity and stability, the relative activity of protease at different temperatures was measured. With the increase of temperature, the relative activity of protease increased first and then decreased, and 45 ℃ was optimal for protease (Fig. 4a). Furthermore, the relative activity of protease decreased with the increase of heat preservation time, and protease had good stability at 35–45℃, keeping more than 50% residual protease activity after heat preservation for 80 min (Fig. 4b). To evaluate the effect of pH on the activity and stability of protease, the relative activity of protease at different pH was measured. With the increase of pH, the relative protease activity increased first and then decreased, and pH 7.5 was the optimal pH for protease (Fig. 4c). The relative activity of protease decreased slowly with the increase of heat preservation time, and protease maintained great stability at pH 7.0–8.0, keeping about 75% residual protease activity after heat preservation for 80 min (Fig. 4d). These results indicated that the optimal temperature and pH of protease were 45℃ and pH 7.5, respectively.
To evaluate the protease production capacity of B. amyloliquefaciens FMME ZK003, fermentation in shake flasks was conducted. During the whole fermentation process, protease activity increased rapidly at 0–12 h and slowly at 12–72 h. The protease activity reached 456.9 U/mL at 72 h, which was 68.9% higher than that of the original strain B. amyloliquefaciens FMME ZK001 (Fig. 5a). Glucose consumption reached 59.8 g/L, and glucose consumption rate reached 0.83 g/L/h, which was 24.6% higher than that of B. amyloliquefaciens FMME ZK001 (Fig. 5b). At 0–36 h, with the consumption of glucose, OD600 increased continuously, maximum OD600 reaching 6.1 at 36 h, increased by 16.6%. However, due to nutrient deficiency and accumulation of toxic metabolites, the cell decays and dies, and the concentration slowly decreases at 36–72 h (Fig. 5c). In addition, protease productivity was increased by 68.9%, and the protease production capacity per unit cell increased by 44.9%. These results proved that B. amyloliquefaciens FMME ZK003 had better protease production ability than B. amyloliquefaciens FMME ZK001 through systematic analysis of fermentation data.
To further verify the protease production capacity of strain B. amyloliquefaciens FMME ZK003, it was characterized by transparent circle and color reaction (Fig. 5d). On the one hand, since H/d value were positively correlated with the protease activity of strain, the protease production capacity could be reflected by continuous photographing of the changes of the transparent circle. With the prolonging of culture time, the protease accumulated continuously, so that casein continued to be consumed, and then the transparent circle gradually became larger. On the other hand, because the folin reagent can be reduced to blue purple by phenolic compounds under alkaline conditions, the protease can react with the substrate casein to generate tyrosine, and the reaction with the folin reagent shows blue purple, so the color depth can reflect the activity of the protease to a certain extent. As shown in Fig. 4d, with the prolongation of fermentation time, the blue–purple gradually became darker, indicating that the accumulation of protease was increased. We evaluated the parameters of B. amyloliquefaciens FMME ZK003 in different ways and proved that it was superior to the original strain B. amyloliquefaciens FMME ZK001. In previous studies, fermentation process of Bacillus were analyzed only by protease activity and substrate consumption to evaluate the capacity of protease production [43]. In our study, a variety of methods to characterize the capacity of protease production were used, such as the detailed parameters in fermentation process, transparent circles and folin reagent color reaction methods, which would make our study more complete and reliable.
Fermentation optimization of protease production
Fermentation optimization plays an indispensable role in improving the production of protease. First, to screen seed medium about protease production of B. amyloliquefaciens FMME ZK003, H/d value was used. The result showed that H/d value of LB medium was significantly higher than that of other three media at 24 h, and bacterial growth was also significantly better than that of other three media (Fig. 6a). Therefore, LB medium was selected as seed medium. Second, to screen the optimal fermentation medium suitable for protease production, H/d value was still used for screening. H/d value of medium B was significantly better than other three media at 48–96 h, and medium B was rich in glucose, peptone and other substances (Fig. 6a). Thus, medium B was selected as fermentation medium. Third, to optimize concentration of carbon source in fermentation medium, protease activity was selected as the screening criterion. The results showed that protease activity increased first and then decreased with the increase of primary glucose concentration of 60, 70, 80, 90, 100, 110, 120 g/L. When initial glucose concentration was 70 g/L, protease activity reached 496.6 U/mL, which was 8.7% higher than initial condition (Fig. 6b). These results indicated that the optimal concentration of glucose was 70 g/L in fermentation medium. To optimize concentration of nitrogen source in fermentation medium, protease activity was also used as evaluation index. The results showed that protease activity increased gradually with the increase of initial nitrogen source concentration from 40 to 90 g/L. When concentration of peptone was 90 g/L, protease activity was the highest, reaching 628.9 U/mL, 37.6% higher than initial condition (Fig. 6c). Thus, adding 90 g/L peptone in the fermentation medium was the best for protease production. Finally, protease fermentation process was further analyzed under the optimal medium conditions. The results showed that with the continuous consumption of glucose, OD600 and protease activity increased gradually. At 72 h, OD600 and protease activity reached 11.0 and 630.7 U/mL, respectively, increased by 79.7 and 38.0% compared with initial condition (Fig. 6d). We got the best protease production conditions by optimizing the production conditions, under which the protease activity was 2.34 times that of original strain B. amyloliquefaciens FMME ZK001. Most of the previous studies also achieved good results when improving protease activity from the aspect of fermentation optimization [44, 45]. In that study, soybean meal, mustard cake, wheat bran, and incubation time have a profound influence on protease production by BSK-1, and the optimization of these significant variables resulted in 2.12-fold (112.0%) enhanced protease yield [44].
Fermentation optimization of protease production. a Effect of medium species on the production of protease. b Effect of glucose concentration on the production of protease. c Effect of peptone concentration on the production of protease. d Protease production of B. amyloliquefaciens FMME ZK003 in shake flask fermentation
Protease production with B. amyloliquefaciens in a 5 L fermenter
To enhance protease production, B. amyloliquefaciens FMME ZK003 was cultured in a 5 L fermenter using the optimized culture conditions. As shown in Fig. 7, glucose consumption reached 84.5 g/L, and glucose consumption rate reached 1.17 g/L/h. OD600 of B. amyloliquefaciens FMME ZK003 increased gradually when it was cultured on the 5 L fermenter, and stopped increasing after it reached the maximum 18.2 at 60 h. During the whole fermentation process, protease activity increased gradually and reached the maximum of 823.0 U/mL at 72 h. In conclusion, B. amyloliquefaciens FMME ZK003 can produce 823.0 U/mL protease in a 5 L fermenter, which was 3.05 times that of original strain B. amyloliquefaciens FMME ZK001.
Conclusions
In this study, a protease producing B. amyloliquefaciens was obtained from animal feces by plate transparent circle method. On this basis, protease production was enhanced by combinatorial mutagenesis of ARTP and 60Co-γ irradiation. After 72 h fermentation in a 5 L fermenter, protease activity of B. amyloliquefaciens FMME ZK003 reached 823.0 U/mL. This provides an opportunity for efficient production of protease in microbial cell factories, and also lays a good foundation for further exploration of the mechanism and property of protease production by B. amyloliquefaciens.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Razzaq A, Shamsi S, Ali A, Ali Q, Sajjad M, Malik A, Ashraf M. Microbial proteases applications. front bioeng. Biotechnol. 2019;7:110.
Contesini FJ, de Melo RR, Sato HH. An overview of Bacillus proteases: from production to application. Crit Rev Biotechnol. 2018;38:321–34.
Wang L, Wang Y-J. Rice starch isolation by neutral protease and high-intensity ultrasound. J Cereal Sci. 2004;39:291–6.
Nalinanon S, Benjakul S, Kishimura H, Shahidi F. Functionalities and antioxidant properties of protein hydrolysates from the muscle of ornate threadfin bream treated with pepsin from skipjack tuna. Food Chem. 2011;124:1354–62.
Hejdysz M, Kaczmarek SA, Kubiś M, Wiśniewska Z, Peris S, Budnik S, Rutkowski A. The effect of protease and Bacillus licheniformis on nutritional value of pea, faba bean, yellow lupin and narrow-leaved lupin in broiler chicken diets. Br Poult Sci. 2020;61:287–93.
Naveed M, Nadeem F, Mehmood T, Bilal M, Anwar Z, Amjad F. Protease—a versatile and ecofriendly biocatalyst with multi-industrial applications: an updated review. Catal Lett. 2021;151:307–23.
Vojcic L, Pitzler C, Körfer G, Jakob F, Ronny M, Maurer K-H, Schwaneberg U. Advances in protease engineering for laundry detergents. New Biotechnol. 2015;32:629–34.
Guleria S, Walia A, Chauhan A, Shirkot CK. Purification and characterization of detergent stable alkaline protease from Bacillus amyloliquefaciens SP1 isolated from apple rhizosphere. J Basic Microbiol. 2016;56:138–52.
Dayanandan A, Kanagaraj J, Sounderraj L, Govindaraju R, Rajkumar GS. Application of an alkaline protease in leather processing: an ecofriendly approach. J Cleaner Prod. 2003;11:533–6.
Rehman R, Ahmed M, Siddique A, Hasan F, Hameed A, Jamal A. Catalytic role of thermostable metalloproteases from bacillus subtilis KT004404 as dehairing and destaining agent. Appl Biochem Biotechnol. 2017;181:434–50.
Gupta R, Beg QK, Lorenz P. Bacterial alkaline proteases: molecular approaches and industrial applications. Appl Microbiol Biotechnol. 2002;59:15–32.
Calik P, Ozdamar TH. Carbon sources affect metabolic capacities of Bacillus species for the production of industrial enzymes: theoretical analyses for serine and neutral proteases and alpha-amylase. Biochem Eng J. 2001;8:61–81.
Ahmed Z, Donkor O, Street WA, Vasiljevic T. Proteolytic activities in fillets of selected underutilized Australian fish species. Food Chem. 2013;140:238–44.
Chalamaiah M, Dineshkumar B, Hemalatha R, Jyothirmayi T. Fish protein hydrolysates: proximate composition, amino acid composition, antioxidant activities and applications: a review. Food Chem. 2012;135:3020–38.
Mala BR, Aparna MT, Mohini SG, Vasanti VD. Molecular and biotechnological aspects of microbial proteases. Microbiol Mol Biol Rev. 1998;62:597–635.
Tufvesson P, Lima-Ramos J, Nordblad M, Woodley JM. Guidelines and cost analysis for catalyst production in biocatalytic processes. Org Process Res Dev. 2011;15:266–74.
Sn N, D J. Optimization of alkaline protease production from Bacillus subtilis NS isolated from sea water. Afr J Biotechnol. 2014;13:1707–13.
Ali N, Ullah N, Qasim M, Rahman H, Khan SN, Sadiq A, Adnan M. Molecular characterization and growth optimization of halo-tolerant protease producing Bacillus Subtilis Strain BLK-1.5 isolated from salt mines of Karak. Pakistan Extremophiles. 2016;20:395–402.
Karray A, Alonazi M, Horchani H, Ben BA. A novel thermostable and alkaline protease produced from Bacillus stearothermophilus isolated from olive oil mill sols suitable to industrial biotechnology. Molecules. 2021. https://doi.org/10.3390/molecules26041139.
Elshaghabee FMF, Rokana N, Gulhane RD, Sharma C, Panwar H. Bacillus as potential probiotics: status, concerns, and future perspectives. Front Microbiol. 2017. https://doi.org/10.3389/fmicb.2017.01490.
Soares VF, Castilho LR, Bon EPS, Freire DMG. High-yield bacillus subtilis protease production by solid-state fermentation. Appl Biochem Biotechnol. 2005;121:311–9.
Elyasi Far B, Yari Khosroushahi A, Dilmaghani A. In silico study of alkaline serine protease and production optimization in Bacillus sp. Khoz1 closed bacillus safensis isolated from honey. Int J Pept Res Ther. 2020;26:2241–51.
Lario LD, Chaud L, Almeida MdG, Converti A, Durães Sette L, Pessoa A. Production, purification, and characterization of an extracellular acid protease from the marine Antarctic yeast Rhodotorula mucilaginosa L7. Fungal Biol. 2015;119:1129–36.
Yegin S, Fernandez-Lahore M, JoseGamaSalgado A, Guvenc U, Goksungur Y, Tari C. Aspartic proteinases from Mucor spp in cheese manufacturing. Appl Microbiol Biotechnol. 2011;89:949–60.
Kalahroudi RJ, Valizadeh V, Atyabi SM, Keramati M, Cohan RA, Aghai A, Norouzian D. Increment in protease activity of Lysobacter enzymogenes strain by ultra violet radiation. Iran J Microbiol. 2020;12:601–6.
Tuly JA, Ma H, Zabed HM, Dong Y, Janet Q, Golly MK, Feng L, Li T, Chen G. Harnessing the keratinolytic activity of bacillus licheniformis through random mutagenesis using ultraviolet and laser irradiations. Appl Biochem Biotechnol. 2022;194:1546–65.
Wang Y, Wang J, Zhang X, Tong Y, Yang R. Genomic and transcriptomic analysis of Bacillus subtilis JNFE1126 with higher nattokinase production through ultraviolet combined 60Co-γ ray mutagenesis. LWT. 2021;147:111652.
Yadav SK, Singh P, Dubey KK, Singh BP. Combined mutagenic improvement of Bacillus licheniformis SK7 for cost-effective protease production. Asian J Bio Sci. 2016;11:91–4.
Zhang X. Applying the mutation of Bacillus subtilis and the optimization of feather fermentation medium to improve Keratinase activity. Adv Biol Chem. 2012;02:64–9.
Zhang JF, Zhu BY, Li XY, Xu XJ, Li DK, Zeng F, Zhou CX, Liu YH, Li Y, Lu FP. Multiple modular engineering of Bacillus Amyloliquefaciens cell factories for enhanced production of alkaline proteases from B. Clausii Front Bioeng Biotechnol. 2022;10:866066.
Zhou CX, Yang GC, Zhang L, Zhang HT, Zhou HY, Lu FP. Construction of an alkaline protease overproducer strain based on Bacillus licheniformis 2709 using an integrative approach. Int J Biol Macromol. 2021;193:1449–56.
Zhou C, Zhou H, Li D, Zhang H, Wang H, Lu F. Optimized expression and enhanced production of alkaline protease by genetically modified Bacillus licheniformis 2709. Microb Cell Fact. 2020;19:45.
Mo QS, Tian Y, Zhang HT, Bu LJ, Lu FP. Using 16S rDNA as target site for homologous recombination to improve the alkaline protease production of Bacillus alcalophilus. Adv Mater Res. 2014;886:349–54.
Suberu Y, Akande I, Samuel T, Lawal A, Olaniran A. Optimization of protease production in indigenous Bacillus species isolated from soil samples in Lagos. Nigeria using response surface methodology: Biocatal Agric Biotechnol; 2019. https://doi.org/10.1016/j.bcab.2019.01.049.
Cai CG, Zheng XD. Medium optimization for keratinase production in hair substrate by a new Bacillus subtilis KD-N2 using response surface methodology. J Ind Microbiol Biotechnol. 2009;36:875–83.
He F, Chao J, Yang D, Zhang X, Yang C, Xu Z, Jiewei T, Yongqiang T. Optimization of fermentation conditions for production of neutral metalloprotease by Bacillus subtilis SCK6 and its application in goatskin-dehairing. Prep Biochem Biotechnol. 2021;52:789–99.
Ding Q, Luo QL, Zhou J, Chen XL, Liu LM. Enhancing l-malate production of Aspergillus oryzae FMME218-37 by improving inorganic nitrogen utilization. Appl Microbiol Biotechnol. 2018;102:8739–51.
Liu H, Zhang Z, Hong P, Zhou C, Yang P, Chen K, Wu J. Isolation of microbial strain producing thermostable protease and characterization of the enzyme. Sci Technol Food Ind. 2017;38:133.
Shu L, Si XG, Yang XD, Ma WY, Sun JL, Zhang J, Xue XL, Wang DP, Gao Q. Enhancement of acid protease activity ofaspergillus oryzaeusing atmospheric and room temperature plasma. Front Microbiol. 2020. https://doi.org/10.3389/fmicb.2020.01418.
Gao X, Liu E, Yin Y, Yang L, Huang Q, Chen S, Ho C-T. Enhancing activities of salt-tolerant proteases secreted by aspergillus oryzae using atmospheric and room-temperature plasma mutagenesis. J Agric Food Chem. 2020;68:2757–64.
Xue G, Chen L, Wu B, He B. Selection of high-yield thermostable protease producing strain by ARTP and the study on its enzymological properties. Sci Technol Food Ind. 2015;36(177–180):206.
Wang S, Luo Q, Liu J, Liu L, Chen X. Mutation and fermentation optimization of Bacillus amyloliquefaciens for acetoin production. Chin J Biotechnol. 2018;34:803–11.
Liu S, Fang Y, Lv M, Wang S, Chen L. Optimization of the production of organic solvent-stable protease by Bacillus sphaericus DS11 with response surface methodology. Bioresour Technol. 2010;101:7924–9.
Singh S, Bajaj BK. Medium optimization for enhanced production of protease with industrially desirable attributes from Bacillus subtilis K-1. Chem Eng Commun. 2015;202:1051–60.
Keshavamurthy M, Vishwanatha T, Kumar SM, Gaddad SM. Enhanced production of alkaline protease from novel bacterium Bacillus cereus GVK21 under submerged fermentation. Biosci Biotechnol Res Commun. 2018;11:416–25.
Funding
This study was supported by the Provincal Outstanding Youth Foundation of Jiangsu Province (BK20211529) and the National Science Fund for Excellent Young Scholars (22122806).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, K., Liu, H., Song, W. et al. Combinatorial mutagenesis of Bacillus amyloliquefaciens for efficient production of protease. Syst Microbiol and Biomanuf 3, 457–468 (2023). https://doi.org/10.1007/s43393-022-00130-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43393-022-00130-7