Abstract
Tomato leaf curl New Delhi virus (ToLCNDV), a bipartite begomovirus, was first reported to infect tomato and has recently spread rapidly as an emerging disease to Cucurbitaceae crops. To date, the virus has been reported to infect more than 11 cucurbit crops, in 16 countries and regions, causing severe yield losses. In autumn 2022, ToLCNDV was first isolated from cucurbit plants in Southeastern coastal areas of China. Phylogenetic analysis established that these isolates belong to the Asian ToLCNDV clade, and shared high nucleotide identity and closest genetic relationship with the DNA-A sequence from the Chinese tomato-infecting ToLCNDV isolate (Accession no. OP356207) and the tomato New Delhi ToLCNDV-Severe isolate (Accession no. HM159454). In this review, we summarize the occurrence and distribution, host range, detection and diagnosis, control strategies, and genetic resistance of ToLCNDV in the Cucurbitaceae. We then summarize pathways that could be undertaken to improve our understanding of this emerging disease, with the objective to develop ToLCNDV-resistant cucurbit cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Begomoviruses are a serious threat to many economically important crops and cause devastating diseases, worldwide (Navas-Castillo et al. 2011). Based on their genome characteristics, host range, and insect vector, the family Geminiviridae is currently divided into fourteen genera, with Begomovirus being the largest genus (Brown et al. 2015; Varsani et al. 2014). Begomoviruses are classified as monopartite (with a single genomic component) or bipartite (with genomic components referred to as DNA A and DNA B). These geminiviruses are transmitted by the whitefly, Bemisia tabaci (Gennadius) (order: Hemiptera, family: Aleyrodidae), in a circulatory persistent manner and infect many dicotyledonous plant species (Brown et al. 2015).
Tomato leaf curl New Delhi virus (ToLCNDV) was first described in 1995 infecting tomato (Solanum lycopersicum) in northern India. This virus is a rapidly spreading bipartite begomovirus, with two circular single-stranded DNA genomic components each of approx. 2.7 kb, designated as DNA-A and DNA-B (Srivastava et al. 1995). ToLCNDV is transmitted by whiteflies but can also be inoculated mechanically (Lopez et al. 2015; Tsai et al. 2011). Typical symptoms include severe leaf curling, yellow mosaic patterns and swelling of the veins of young leaves and dwarfing through the shortening of internodes (Zaidi et al. 2017).
Following its first report 28 years ago, ToLCNDV has been reported to infect a large number of economically important crops, over a wide range of families, including Cucurbitaceae, Solanaceae, Malvaceae, Fabaceae and Euphorbiaceae (Seal et al. 2006). Particularly in the cucurbits, ToLCNDV can cause severe losses in such vegetables as zucchini (Cucurbita pepo), cucumber (Cucumis sativus), and squash (Cucurbita moschata), and in fruits such as melon (Cucumis melo) and watermelon (Citrullus lanatus), as well as several pumpkin species (Bandaranayake et al. 2014; Siskos et al. 2022; Venkataravanappa et al. 2020; Yamamoto et al. 2021). In Europe, ToLCNDV has had severe negative impacts on various crops, especially cucurbits, with losses of up to 20% in open-field melon production in central Spain (Saez et al. 2017).
Genetic resistance has been identified in several Cucurbitaceae crops, including melon, cucumber, pumpkin, and sponge gourd (Luffa cylindrica) (Kaur et al. 2021; Romero-Masegosa et al. 2021; Saez et al. 2017, 2020, 2021). However, to date, no resistance gene has been successfully mapped and characterized.
In the autumn of 2022, ToLCNDV was isolated from different cucurbit crops being grown in Jiangsu Province, Zhejiang Province and Shanghai, located in the Southeastern coastal areas of China, where infection of ~ 650 hectares caused approx. $US15 million in economic losses (Gu et al. 2023; Zeng et al. 2023).
Here, we briefly describe the history of ToLCNDV research in the Cucurbitaceae and then address the distribution, host range, genetic relationship, detection and diagnosis, control strategies, and genetic resistance of ToLCNDV. We then assess the presence of natural sources of resistant germplasm and mapping of resistance genes, which could be employed to advance our knowledge on this emerging disease in the cucurbits and other crops.
Discovery and distribution
Although ToLCNDV was first described in India in 1995, only partial DNA fragments of begomoviruses were detected from cucurbits in 2000, including cucumber, muskmelon (Cucumis melo L.), cantaloupe melon (Cucumis melo var. reliculatus) and wax gourd (Benincase hispida) (Samretwanich et al. 2000a, 2000b, 2000c). Subsequently, several disease reports indicated that ToLCNDV had caused severe symptoms in bitter gourd, in Pakistan (Tahir and Haider 2005), and infected cucumber, bottle gourd and muskmelon, in Thailand (Ito et al. 2008). The distribution of ToLCNDV in the cucurbits has now been reported in the Indian subcontinent, Southeast Asia, and East Asia. Since 2012, reports of ToLCNDV-infected cucurbits have spread westward, including from Algeria, Spain, Morocco, Tunisia, Italy, Greece, and Iran (Juarez et al. 2014; Kheireddine et al. 2019; Mnari-Hattab et al. 2015; Orfanidou et al. 2019; Panno et al. 2016; Sifres et al. 2018; Yazdani-Khameneh et al. 2016). In recent years, additional countries (Malaysia, Cambodia, Laos, Indonesia, and France) have reported that ToLCNDV has caused problems in cucurbit crops (Chen et al. 2021; Desbiez et al. 2021; Neoh et al. 2023).
To date, according to the retrieved data from NCBI, more than 11 infected cucurbit species have been reported from 16 countries and areas in three continents, including Asia, Europe, and Africa (Table 1). These regions are located mainly in tropical and temperate regions, where ToLCNDV has caused tremendous economic losses in both greenhouse and open-field production of cucurbit crops (Lopez et al. 2015).
Host range and disease symptoms in Cucurbits
ToLCNDV infects a wide range of hosts, which in the cucurbits include the following: melon, cucumber, watermelon, zucchini, pumpkin, gourd, sponge gourd, wax gourd, bottle gourd (Lagenaria siceraria), ridge gourd (Luffa acutangula), bitter gourd (Momordica charantia), ivy gourd (Coccinia grandis), ash gourd (Bragard et al. 2020) (Table 1). Symptoms consist of severe yellowing and mosaic discoloration in young leaves and includes leaf curling, vein swelling and short internodes, along with fruit skin roughness and longitudinal cracking (Bragard et al. 2020; Juarez et al. 2019; Lopez et al. 2015).
Short internodes and curling, vein swelling and mosaic in young leaves were observed on the leaves of melon (Fig. 1A, B) and melon fruits have longitudinal cracks (Fig. 1C) (Siskos et al. 2022). Infected bitter gourd shows mosaic mottling with slight curling of leaves (Fig. 1E) (Kiran et al. 2021). The disease leaves symptoms of zucchini were similar to melon (Fig. 1F) (Juarez et al. 2014) and fruits are of lower quality due to skin roughness and reduced size (Fig. 1G). Watermelon and cucumber leaves show yellowing and upward leaf curling (Fig. 1D, H) (Venkataravanappa et al. 2020; Yamamoto et al. 2021).
Disease symptoms caused by Tomato leaf curl New Delhi virus in foliage and fruit of different cucurbit species. Yellowing and curling of melon (Cucumis melo) leaves (A, B), as well as fruit cracking (C). Watermelon (Citrullus lanatus) plants show yellowing and downward curling symptoms (D). Mosaic mottling with slight curling of leaves in bitter gourd (Lagenaria siceraria) (E). Short internodes and curling, vein swelling and mosaic symptoms in young zucchini (Cucurbita pepo) leaves (F). Zucchini fruits with lower marketability due to the roughness of the skin and reduced size (G). Yellowing and upward leaf curling on cucumber (Cucumis sativus) leaves (H). Images in (A), (F), (G), and (H) refer to the European and Mediterranean Plant Protection Organization (EPPO) global database (https://gd.eppo.int/taxon/TOLCND/photos), posted by Dr. Raffaele Giurato. Figures 1B, C were cited from Siskos et al. (2022) without revision; reprint permission: https://springerlink.bibliotecabuap.elogim.com/article/10.1007/s10681-022-03081-1#rightslink; Fig. 1D was cited from Venkataravanappa et al. (2020) without revision, reprint permission: https://s100.copyright.com/CustomerAdmin/PLF.jsp?ref=e2f56b44-78b5-4777-b376-7ab99eb6039e; Fig. 1E was cited from Kiran et al. (2021) without revision; reprint permission: https://s100.copyright.com/CustomerAdmin/PLF.jsp?ref=625141e1-4cce-4fc2-90dc-f95c540cdfe8. Data is retrieved from NCBI
Genome characterization and sequence diversity
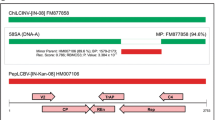
ToLCNDV is a bipartite begomovirus with two circular single-stranded DNA genome components: DNA-A contains the AV1 and AV2 genes, in the virion sense orientation, which encode the coat protein (CP) and pre-coat protein, respectively. In the complementary sense orientation, AC1, AC2, AC3 and AC4 encode the replication-associated protein (Rep), transcriptional activator protein (TrAP), replication enhancer protein (REn) and viral effector, respectively. ToLCNDV DNA-B consists of the BV1 gene, in the virion sense orientation, and the BC1 gene, in the complementary sense orientation, which functions as a nuclear shuttle protein (NSP) and movement protein (MP), respectively. Beta satellites are associated with ToLCNDV (Zaidi et al. 2017). ToLCNDV is unique in that it shares its DNA-B component with several other bipartite begomoviruses, such as Pepper leaf curl Bangladesh virus, Tomato leaf curl Palampur virus, and Bhendi yellow vein mosaic virus (Zaidi et al. 2016).
Globally, some 681 ToLCNDV DNA-A accessions have been deposited in the GenBank database, of which 184 are for the cucurbits (Supplemental Table 1 and 2). A phylogenetic analysis of ToLCNDV isolates from cucurbit plants indicated the presence of two main clades, an Asian and European clade, respectively (Fig. 2). The genetic relationship between these ToLCNDV cucurbit isolates is not associated with the species, but probably relates to the region.
Phylogenetic trees derived from the DNA-A components of Tomato leaf curl New Delhi virus isolates derived from cucurbit plants. The genetic relationships indicate that clade separation was likely not associated with the species, but rather with the various regions. Scale bar representing the genetic distance is shown in the upper left
Detection and diagnosis
ToLCNDV serves as an emerging threat to the global cucurbit industry, and rapid and accurate detection and diagnosis are critical for an effective response to control its spread. Several molecular approaches are available for the detection and identification of ToLCNDV. Polymerase chain reaction (PCR), rolling circle amplification (RCA), sequence analysis and enzyme-linked immunosorbent assay (ELISA) methods can be routinely used for virus detection. The procedures for viral genome sequencing are as follows: Collect diseased samples from cucurbit plants showing symptoms of mosaic, leaf curl; viral DNA is then extracted from approx. 50 mg of dried samples (Gilbertson et al. 1991; Tsai et al. 2011), then specific primers are used for amplification (Chen et al. 2021; Tsai et al. 2011), and the products are sequenced, by next-generation sequencing (NGS), which provides an effective method for virus detection and identification. NGS has been successfully used to sequence the full-length ToLCNDV genome, which has offered a foundation for developing an understanding of evolutionary relationships and the global distribution of ToLCNDV (Chen et al. 2021; Figas et al. 2017).
Commercial ELISA tests (e.g., AGDIA, DSMZ) are available for ToLCNDV detection, and a molecular test, based on loop-mediated isothermal amplification (LAMP)-based molecular test, is available from Enbiotech srl (Palermo, Italy), which affords robust virus detection (Bragard et al. 2020; Panno et al. 2019). The alpha and beta satellites that are associated with ToLCNDV disease can also be detected by molecular means using appropriate primers (quantitative PCR and RCA) (Zaidi et al. 2016).
Emergence of ToLCNDV infecting cucurbits in China
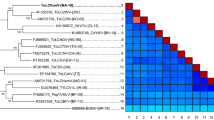
In the autumn of 2022, the occurrence in China of ToLCNDV-infected cucurbit plants was reported in four cities. A phylogenetic tree, constructed with 681 ToLCNDV DNA-A accessions, and 50 isolates, most closely related to the ToLCNDV China isolates from cucurbit plants, were used for an evolutionary analysis. All the DNA-A component of ToLCNDV China cucurbit plants isolates shared high nucleotide identities (99.34–99.85%) with the DNA-A sequence of isolates of ToLCNDV infecting tomato in China (OP356207) (Li et al. 2022) (Fig. 3), and shared 98.36–98.83% with DNA-A genome of ToLCNDV-Severe isolate (Accession no. HM159454) from tomato in New Delhi. This evolutionary analysis suggests that the isolates from cucurbit plants in China were more closely related to the tomato isolates than to other cucurbit isolates from other geographical regions.
Phylogenetic trees of the DNA-A component of ToLCNDV China isolates with other selected ToLCNDV identified from different crops. The ToLCNDV China isolates from cucurbit plants are labeled in pink, and two isolates from tomato plants genetically close to the cucurbits isolates are labeled in blue, two other cucurbit isolates from India and Pakistan are labeled in green. Scale bar representing the genetic distance is shown in the upper left
Disease management
Integrated disease management (IDM) strategies, which combine host resistance, chemical, biological, and cultural control measures, can maximize viral disease control, while also meeting the requirements of environmental and social responsibility (Jones 2004). There is a consensus that effective control of ToLCNDV should be based on such an integrated management approach.
Chemical control
Bemisia tabaci is a very efficient vector of ToLCNDV, and the use of insecticides has long been the main strategy for whitefly management (Legg et al. 2014). However, heavy pesticide use can lead to the emergence of resistant whitefly populations and, long term, can be ecologically harmful. Natural plant bio-stimulants can be used as environmentally friendly alternatives to agricultural chemicals. A recent study reported that such bio-stimulants (e.g., Fullcrhum Alert and BioVeg 500) affected the fitness of zucchini plants against ToCLNDV (Donati et al. 2022). Essential oils (EOs) and hydrosols (HS) derived from natural plant products have also gained attention. For example, EOs and HS from Origanum vulgare (OV), Thymus vulgaris (TV), and Rosmarinum officinalis (RO) were beneficial in treating ToLCNDV-infected greenhouse-grown Cucurbita pepo plants (Taglienti et al. 2022).
Biological control
Biological control of insect vectors would also assist in limiting the spread of ToLCNDV. For example, the predatory mite, Amblyseius swirskii, which feeds on Bemisia tabaci eggs and nymphs (the first instar stage, also called crawlers), is an effective biological control agent. These mites can significantly reduce the secondary transmission of ToLCNDV in zucchini crops (Tellez et al. 2017). Based on these findings, a combination of pesticides compatible with predatory mites could reduce ToLCNDV primary transmission, by eliminating the B. tabaci adults, thus more effectively controlling ToLCNDV transmission (Rodriguez et al. 2019).
Cultural practices
The use of healthy planting material is an important first step in minimizing the spread of ToLCNDV. As seed transmission of ToLCNDV has been reported in India and Italy (Kil et al. 2020; Sangeetha et al. 2018), strict quarantine measures are essential when cucurbit seeds are transported between production areas. Insect and virus-free material is recommended for cucurbits, such as sponge gourd, which are usually vegetatively propagated (Venkataravanappa et al. 2019). In addition, some cultural practices, such as the implementation of a fallow period, early or late planting, physical barriers, intercropping, etc., can also be used to control the whitefly population and associated viral diseases (Hilje et al. 2001). Crop selection is also an important part of disease management. As an example, bitter gourd planted adjacent to tomato plots showed susceptibility to ToLCNDV (Kiran et al. 2021). Weeds can also act as alternative viral hosts; hence, effective and timely application of weed control, is essential. Weeds reported to serve as alternative hosts for ToLCNDV include Ecballium elaterium, Datura stramonium, Sonchus oleraceus, and Solanum nigrum (Juarez et al. 2019).
Host resistance
Resistance to ToLCNDV has been reported in several cucurbit accessions, with five resistant lines, DSG-6, DSG-7, IIHR-137, IIHR-138 and IIHR-Sel-1, identified in sponge gourd (Islam et al. 2010; Kaur et al. 2021). The DSG-6 and DSG-7 resistance to ToLCNDV was shown to be controlled by a single dominant gene (Islam et al. 2010). Three sources of resistance have been identified in squash, including the improved cultivar Large Cheese from the USA (PI 604506), an Indian landrace (PI 381814) and a Japanese accession, BSUAL-252. Here, PI 604506 resistance is conferred by a single recessive gene, located between 799,373 and 986,936 bp on Chromosome 8, whereas the BSUAL-252 resistance was shown to be controlled by a single dominant gene unrelated to this region on Chromosome 8 (Romero-Masegosa et al. 2021; Saez et al. 2016, 2020).
In melon, nine accessions with good resistance to ToLCNDV have been identified: Kharbuja, PI 124112, PI 414723, WM9, WM7, AM 87, IC-274014, PI 282448 and PI 179901 (Lopez et al. 2015; Roman et al. 2019; Saez et al. 2017). Three genomic regions, derived from the wild Indian source, WM7, were identified as resistance-conferring loci, including a major QTL on Chromosome 11 and two additional regions on Chromosomes 12 and 2 (Saez et al. 2017). Further candidate gene validation established that transcript levels of CmARP4 and CmNAC were differentially higher in the inoculated susceptible genotype than in the inoculated resistant genotype (Roman et al. 2019). Studies on the inheritance of resistance in IC-274014 revealed the involvement of one recessive (bgm-1) and two dominant (Bgm-2, Tolcndv) genes (Romay et al. 2019).
Transcriptome analysis has also been employed to study the interactions between ToLCNDV and melon. Here, the structural functionality of differentially expressed genes (DEGs) associated with the main QTLs for ToLCNDV resistance has been reported (Saez et al. 2022). In cucumber, three resistant lines were identified, CGN23089, CGN23423, and CGN23633, in which resistance was controlled by a single recessive gene, and a resistance-related QTL was identified on Chromosome 2 (Saez et al. 2021).
Conclusions and perspective
Cucurbit crops suffer from numerous viral diseases, with ToLCNDV being highly transmitted by whitefly vectors, and it can also be co-infected with multiple other begomoviruses (Zaidi et al. 2016). Here, we outlined the discovery of ToLCNDV in China, its global distribution, host range, detection, and diagnosis. Development of control strategies, including IDM, in combination with breeding for robust genetic resistance, are essential for the detection and control of this emerging disease in cucurbit crops.
Although progress has been accomplished, it is imperative that candidate resistance/susceptibility genes be further studied to develop an understanding of the molecular mechanism(s) by which the ToLCNDV infection process is either prevented or permitted in the host plant. The utility of such validated resistance genes would be in their development as molecular makers for use in breeding ToLCNDV-resistant varieties.
Data availability
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.
References
Bandaranayake WMEK, Wickramarachchi WART, Wickramasinghe HAM, Rajapakshe RGAS, Dissanayake DMKK (2014) Molecular detection and characterization of begomoviruses associated with cucurbitaceae vegetables in Sri Lanka. J Natl Sci Found Sri 42:239–245. https://doi.org/10.4038/jnsfsr.v42i3.7400
Bragard C et al (2020) Pest categorisation of tomato leaf curl New Delhi virus. Efsa J. https://doi.org/10.2903/j.efsa.2020.6179
Brown JK et al (2015) Revision of Begomovirus taxonomy based on pairwise sequence comparisons. Arch Virol 160:1593–1619. https://doi.org/10.1007/s00705-015-2398-y
Chen YJ, Lai HC, Lin CC, Neoh ZY, Tsai WS (2021) Genetic diversity, pathogenicity and pseudorecombination of cucurbit-infecting Begomoviruses in Malaysia. Plants-Basel. https://doi.org/10.3390/plants10112396
Desbiez C, Gentit P, Cousseau-Suhard P, Renaudin I, Verdin E (2021) First report of Tomato leaf curl New Delhi virus infecting courgette in France. New Disease Reports 43:e12006. https://doi.org/10.1002/ndr2.12006
Donati L et al (2022) Effects of organic biostimulants added with Zeolite on zucchini squash plants infected by Tomato leaf curl New Delhi virus. Viruses-Basel. https://doi.org/10.3390/v14030607
Figas MR et al (2017) Inoculation of cucumber, melon and zucchini varieties with Tomato leaf curl New Delhi virus and evaluation of infection using different detection methods. Ann Appl Biol 170:405–414. https://doi.org/10.1111/aab.12344
Gilbertson RL, Rojas MR, Russell DR, Maxwell DP (1991) Use of the asymmetric polymerase chain reaction and DNA sequencing to determine genetic variability of bean golden mosaic Geminivirus in the dominican republic. J Gen Virol 72:2843–2848. https://doi.org/10.1099/0022-1317-72-11-2843
Gu Q et al (2023) First report of tomato leaf curl New Delhi virus infecting several cucurbit plants in China. Plant Dis. https://doi.org/10.1094/PDIS-01-23-0180-PDN
Hilje L, Costa HS, Stansly PA (2001) Cultural practices for managing Bemisia tabaci and associated viral diseases. Crop Prot 20:801–812. https://doi.org/10.1016/S0261-2194(01)00112-0
Islam S, Munshi AD, Mandal B, Kumar R, Behera TK (2010) Genetics of resistance in Luffa cylindrica Roem. against tomato leaf curl New Delhi virus. Euphytica 174:83–89. https://doi.org/10.1007/s10681-010-0138-7
Ito T, Sharma P, Kittipakorn K, Ikegami M (2008) Complete nucleotide sequence of a new isolate of tomato leaf curl New Delhi virus infecting cucumber, bottle gourd and muskmelon in Thailand. Arch Virol 153:611–613. https://doi.org/10.1007/s00705-007-0029-y
Jones RA (2004) Using epidemiological information to develop effective integrated virus disease management strategies. Virus Res 100:5–30. https://doi.org/10.1016/j.virusres.2003.12.011
Juarez M et al (2014) First detection of Tomato leaf curl New Delhi virus infecting zucchini in Spain. Plant Dis 98:857–858. https://doi.org/10.1094/Pdis-10-13-1050-Pdn
Juarez M, Rabadan MP, Martinez LD, Tayahi M, Grande-Perez A, Gomez P (2019) Natural hosts and genetic diversity of the emerging Tomato leaf curl New Delhi virus in Spain. Front Microbiol. https://doi.org/10.3389/fmicb.2019.00140
Kaur M, Varalakshmi B, Pitchaimuthu M, Mahesha B (2021) Screening Luffa germplasm and advanced breeding lines for resistance to Tomato leaf curl New Delhi virus. J Gen Plant Pathol 87:287–294. https://doi.org/10.1007/s10327-021-01010-z
Kheireddine A, Sifres A, Saez C, Pico B, Lopez C (2019) First report of Tomato leaf curl New Delhi virus infecting cucurbit plants in Algeria. Plant Dis 103:3291–3291. https://doi.org/10.1094/Pdis-05-19-1118-Pdn
Kil EJ et al (2020) Seed transmission of Tomato leaf curl New Delhi virus from zucchini squash in Italy. Plants-Basel. https://doi.org/10.3390/plants9050563
Kiran GVNSM, Nagaraju N, Girish TR, Ashwini BN (2021) Molecular investigations reveal bitter gourd crop is more susceptible to tomato leaf curl New Delhi virus infection in diverse crop cultivation practices. 3Biotech. https://doi.org/10.1007/s13205-021-02975-6
Legg JP et al (2014) Biology and management of Bemisia whitefly vectors of cassava virus pandemics in Africa. Pest Manag Sci 70:1446–1453. https://doi.org/10.1002/ps.3793
Lopez C, Ferriol M, Pico MB (2015) Mechanical transmission of Tomato leaf curl New Delhi virus to cucurbit germplasm: selection of tolerance sources in Cucumis melo. Euphytica 204:679–691. https://doi.org/10.1007/s10681-015-1371-x
Mnari-Hattab M, Zammouri S, Belkadhi MS, Doña DB, Nahia EB, Hajlaoui MR (2015) First report of Tomato leaf curl New Delhi virus infecting cucurbits in Tunisia. New Disease Report 31:21–21. https://doi.org/10.5197/j.2044-0588.2015.031.021
Navas-Castillo J, Fiallo-Olive E, Sanchez-Campos S (2011) Emerging virus diseases transmitted by whiteflies. Annu Rev Phytopathol 49:219–248. https://doi.org/10.1146/annurev-phyto-072910-095235
Neoh ZY, Lai H-C, Lin C-C, Suwor P, Tsai W-S (2023) Genetic diversity and geographic distribution of cucurbit-infecting begomoviruses in the Philippines. Plants-Basel 12:272. https://doi.org/10.3390/plants12020272
Orfanidou CG et al (2019) First report of tomato leaf curl New Delhi virus in zucchini crops in Greece. J Plant Pathol 101:799–799. https://doi.org/10.1007/s42161-019-00265-y
Panno S et al (2016) First report of Tomato leaf curl New Delhi virus affecting zucchini squash in an important horticultural area of southern Italy. New Disease Report 33:6–6. https://doi.org/10.5197/j.2044-0588.2016.033.006
Panno S et al (2019) Emergence of tomato leaf curl New Delhi virus in Italy: estimation of incidence and genetic diversity. Plant Pathol 68:601–608. https://doi.org/10.1111/ppa.12978
Rodriguez E, Tellez MM, Janssen D (2019) Whitefly control strategies against Tomato leaf curl New Delhi Virus in greenhouse zucchini. Int J Env Res Pub He. https://doi.org/10.3390/ijerph16152673
Roman B, Gomez P, Pico B, Lopez C, Janssen D (2019) Candidate gene analysis of Tomato leaf curl New Delhi virus resistance in Cucumis melo. Sci Hortic-Amsterdam 243:12–20. https://doi.org/10.1016/j.scienta.2018.07.005
Romay G, Pitrat M, Lecoq H, Wipf-Scheibel C, Millot P, Girardot G, Desbiez C (2019) Resistance against Melon chlorotic mosaic virus and Tomato leaf curl New Delhi virus in melon. Plant Dis 103:2913–2919. https://doi.org/10.1094/Pdis-02-19-0298-Re
Romero-Masegosa J et al (2021) Response of Cucurbitaspp. to tomato leaf curl New Delhi virus inoculation and identification of a dominant source of resistance in Cucurbita moschata. Plant Pathol 70:206–218. https://doi.org/10.1111/ppa.13268
Saez C et al (2016) Resistance to Tomato leaf curl New Delhi virus in Cucurbita spp. Ann Appl Biol 169:91–105. https://doi.org/10.1111/aab.12283
Saez C, Esteras C, Martinez C, Ferriol M, Dhillon NPS, Lopez C, Pico B (2017) Resistance to tomato leaf curl New Delhi virus in melon is controlled by a major QTL located in chromosome 11. Plant Cell Rep 36:1571–1584. https://doi.org/10.1007/s00299-017-2175-3
Saez C et al (2020) A major QTL located in chromosome 8 of Cucurbita moschata is responsible for resistance to Tomato leaf curl New Delhi virus. Front Plant Sci. https://doi.org/10.3389/fpls.2020.00207
Saez C, Ambrosio LGM, Miguel SM, Valcarcel JV, Diez MJ, Pico B, Lopez C (2021) Resistant sources and genetic control of resistance to ToLCNDV in cucumber. Microorganisms. https://doi.org/10.3390/microorganisms9050913
Saez C, Flores-Leon A, Montero-Pau J, Sifres A, Dhillon NPS, Lopez C, Pico B (2022) RNA-Seq transcriptome analysis provides candidate genes for resistance to Tomato leaf curl New Delhi virus in melon. Front Plant Sci. https://doi.org/10.3389/fpls.2021.798858
Samretwanich K, Chiemsombat P, Kittipakorn K, Ikegami M (2000a) Tomato leaf curl geminivirus associated with cantaloupe yellow leaf disease in Thailand. World J Microb Biot 16:401–403. https://doi.org/10.1023/A:1008962402329
Samretwanich K, Chiemsombat P, Kittipakorn K, Ikegami M (2000b) Tomato leaf curl geminivirus associated with cucumber yellow leaf disease in Thailand. J Phytopathol 148:615–617. https://doi.org/10.1023/A:1008962402329
Samretwanich K, Chiemsombat P, Kittipakorn K, Ikegami M (2000c) Yellow leaf disease of cantaloupe and wax gourd from Thailand caused by Tomato leaf curl virus. Plant Dis 84:200. https://doi.org/10.1094/pdis.2000.84.2.200c
Sangeetha B, Malathi VG, Alice D, Suganthy M, Renukadevi P (2018) A distinct seed-transmissible strain of tomato leaf curl New Delhi virus infecting Chayote in India. Virus Res 258:81–91. https://doi.org/10.1016/j.virusres.2018.10.009
Seal SE, Vanden Bosch F, Jeger MJ (2006) Factors influencing begomovirus evolution and their increasing global significance: Implications for sustainable control. Crit Rev Plant Sci 25:23–46. https://doi.org/10.1080/07352680500365257
Sifres A, Saez C, Ferriol M, Selmani KA, Riado J, Pico B, Lopez C (2018) First report of Tomato leaf curl New Delhi virus infecting zucchini in Morocco. Plant Dis 102:1045–1045. https://doi.org/10.1094/Pdis-10-17-1600-Pdn
Siskos L, Cui L, Wang C, Visser RGF, Bai YL, Schouten HJ (2022) A new challenge in melon resistance breeding: the ToLCNDV case. Euphytica. https://doi.org/10.1007/s10681-022-03081-1
Srivastava KM, Hallan V, Raizada RK, Chandra G, Singh BP, Sane PV (1995) Molecular-cloning of Indian Tomato leaf curl virus genome following a simple method of concentrating the supercoiled replicative form of viral-DNA. J Virol Methods 51:297–304. https://doi.org/10.1016/0166-0934(94)00122-W
Taglienti A et al (2022) In vivo antiphytoviral activity of essential oils and hydrosols from origanum vulgare, thymus vulgaris, and rosmarinus officinalis to control Zucchini yellow mosaic virus and Tomato leaf curl New Delhi virus in Cucurbita pepo L. Front Microbiol. https://doi.org/10.3389/fmicb.2022.840893
Tahir M, Haider MS (2005) First report of Tomato leaf curl New Delhi virus infecting bitter gourd in Pakistan. Plant Pathol 54:807–807. https://doi.org/10.1111/j.1365-3059.2005.01215.x
Tellez MD, Simon A, Rodriguez E, Janssen D (2017) Control of Tomato leaf curl New Delhi virus in zucchini using the predatory mite Amblyseius swirskii. Biol Control 114:106–113. https://doi.org/10.1016/j.biocontrol.2017.08.008
Tsai WS, Shih SL, Kenyon L, Green SK, Jan FJ (2011) Temporal distribution and pathogenicity of the predominant tomato-infecting begomoviruses in Taiwan. Plant Pathol 60:787–799. https://doi.org/10.1111/j.1365-3059.2011.02424.x
Varsani A et al (2014) Establishment of three new genera in the family Geminiviridae: Becurtovirus, Eragrovirus and Turncurtovirus. Arch Virol 159:2193–2203. https://doi.org/10.1007/s00705-014-2050-2
Venkataravanappa V, Reddy NL, Shankarappa KS, Reddy MK (2019) Association of Tomato leaf curl New Delhi virus betasatellite and alphasatellite with mosaic disease of spine gourd (Momordica dioica Roxb. Willd) in India. Iran J Biotechnol 17:17–29. https://doi.org/10.2159/ijb.2134
Venkataravanappa V, Ashwathappa KV, Reddy CNL, Shankarappa KS, Reddy MK (2020) Characterization of Tomato leaf curl New Delhi virus associated with leaf curl and yellowing disease of Watermelon and development of LAMP assay for its detection. 3 Biotech. https://doi.org/10.1007/s13205-020-02245-x
Yamamoto H, Wakita Y, Kitaoka T, Fujishiro K, Kesumawati E, Koeda S (2021) Southeast Asian isolate of the Tomato leaf curl New Delhi virus shows higher pathogenicity against tomato and cucurbit crops compared to that of the Mediterranean isolate. Horticult J 90:314–325. https://doi.org/10.2503/hortj.UTD-269
Yazdani-Khameneh S, Aboutorabi S, Shoori M, Aghazadeh A, Jahanshahi P, Golnaraghi A, Maleki M (2016) Natural occurrence of Tomato leaf curl New Delhi virus in Iranian cucurbit crops. Plant Pathology J 32:201–208. https://doi.org/10.5423/Ppj.Oa.10.2015.0210
Zaidi SSEA, Shafiq M, Amin I, Scheffler BE, Scheffler JA, Briddon RW, Mansoor S (2016) Frequent occurrence of Tomato leaf curl New Delhi virus in cotton leaf curl disease affected cotton in Pakistan. PLoS ONE. https://doi.org/10.1371/journal.pone.0155520
Zaidi SSEA, Martin DP, Amin I, Farooq M, Mansoor S (2017) Tomato leaf curl New Delhi virus: a widespread bipartite begomovirus in the territory of monopartite begomoviruses. Mol Plant Pathol 18:901–911. https://doi.org/10.1111/mpp.12481
Zeng R et al (2023) Occurrence of Tomato leaf curl New Delhi virus in cucurbit plants in China. Plant Dis. https://doi.org/10.1094/PDIS-01-23-0059-PDN
Acknowledgements
We thank Prof. Weihong Bao for providing field infectious leaves, and Prof. Robert N. Trigiano for the critical review and revision of this manuscript. This study was partially supported by a grant from the Key Research and Development Program of Hainan Province (ZDYF2021XDNY166), the Key Science and Technology Program for Agricultural (Vegetable) New Variety Breeding of Zhejiang Province (2021C02065), and the Earmarked Fund for Modern Agro-Industry Technology Research System of China (CARS-26-17).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The author declares that no competing interests exist.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cai, L., Mei, Y., Ye, R. et al. Tomato leaf curl New Delhi virus: an emerging plant begomovirus threatening cucurbit production. aBIOTECH 4, 257–266 (2023). https://doi.org/10.1007/s42994-023-00118-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42994-023-00118-4