Abstract
Sound management of coastal resources is based on science-based decisions. Bottlenose dolphins are found around Puerto Rico; however, limited information exists on the ecology, behavior, sex ratio, distribution patterns, and population structure presenting, challenges in managing the bottlenose dolphin as defined in the Marine Mammal Protection Act of 1972. We sequenced the mitochondrial control region (mtDNA-CR) of 27 live and 11 stranded dolphins from Puerto Rico, five stranded dolphins from Guadeloupe and included sequences from the North Atlantic and the Pacific Ocean. Our genetic data from the new samples indicates the presence of distinct genetic lineages (inshore—represented by coastal individuals) and worldwide-distributed form (represented by both coastal and offshore individuals) in Puerto Rico. DNA divergence between inshore/coastal and offshore haplotypes ranged from 4.34 to 6.58%. All haplotypes from Puerto Rico have been previously reported from the Caribbean and North Atlantic. Genetic analysis yielded a complex population structure without a clear geographic signal; an expected result from a highly mobile marine mammal. A clade consisting exclusively of coastal dolphins of the Caribbean and the western North Atlantic was recovered. Offshore haplotypes from the eastern and western North Atlantic were generally clustered with offshore haplotypes of the Caribbean. Coastal and offshore haplotypes from the Pacific differed from those from the Atlantic. When we partitioned the data by form (coastal vs. offshore) and ocean (Atlantic vs. Pacific), we detected significant population differentiation (FST = 0.4089), indicating limited gene flow between forms and across oceans.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The common bottlenose dolphin (Tursiops truncatus truncatus) is considered the most common nearshore cetacean in the Caribbean (Ward et al. 2001). Geographical variations in size, coloration, habitats, and cranial characteristics of bottlenose dolphins across the world’s oceans have led researchers to differentiate two forms of the species (Hersh and Duffield 1990; Mead and Potter 1995). This distinction has been based on mitochondrial DNA (mtDNA), hemoglobin, parasite loads, prey preferences, and distribution (Hersh and Duffield 1990; Mead and Potter 1995; Hoelzel et al. 1998; Mignucci-Giannoni et al. 1998; Colón-Llavina et al. 2009; Caballero et al. 2011). The offshore form is distinguished by a falcated dorsal fin, a short rostrum, a bulky body, a dark cape pattern, and a white saddle patch in the peduncle area behind the dorsal fin (Herzing and Elliser 2016; Ramos et al. 2016; Van Waerebeek et al. 2017) and is found in deep zones near oceanic islands or in the ocean (Hersh and Duffield 1990). The coastal or inshore form, in contrast, is smaller in size, has lighter coloring, and larger flippers (Mead and Potter 1995; Ramos et al. 2016; Ruenes et al. 2023). However, the features of the two types are not consistent worldwide (Curry and Smith 1997). For example, offshore T. truncatus tend to be smaller in the Pacific Ocean than their nearshore counterparts (Curry and Smith 1997; Bearzi et al. 2009). It is commonly assumed that the inshore form of this species primarily inhabits the coastal zone, while the offshore form is typically found in the pelagic zone. However, recent observations challenge this assumption, as individuals from the offshore form have been observed near the shore in certain areas (Wells et al. 1999). Conversely, individuals corresponding to the inshore form have been observed in far-reaching continental shelf regions (Kenney 1990).
Analysis of mtDNA from bottlenose dolphins from the Caribbean revealed the presence of two genetically differentiated forms; one described as inshore and the other as a worldwide distributed form (Tezanos-Pinto et al. 2009; Caballero et al. 2011). The worldwide distributed form was represented by genetically coastal and offshore individuals that could inhabit both coastal and oceanic habitats. This form exhibits a high level of mtDNA diversity, but no discernible phylogeographic distinction was found among them and no corresponding morphological analysis was made in these assessments (e.g. Tezanos-Pinto et al. 2009; Caballero et al. 2011). The status of various forms of T. truncatus globally is unclear. The 2018 workshop on the taxonomy of the genus Tursiops (Natoli et al. 2019) highlighted various factors contributing to the existing taxonomic uncertainty within the genus. These factors include the wide distribution of Tursiops across diverse and variable environments, limited availability of specimens from numerous regions, variations in research methods and designs, and the intricate and protracted nomenclatural history associated with the genus (Natoli et al. 2019). Following the latest taxonomic distinction, we will use the terms coastal and offshore forms throughout the manuscript as neutral descriptors of the species.
In the Caballero et al. (2011) study, 26 of the analyzed samples were from dolphins stranded in Puerto Rico, and based on the genetic analysis, both forms were identified in Puerto Rico (24 offshore and 2 inshore forms). As these samples came from stranded individuals, no data were available on the geographic origin of the dolphins. Ocean currents can move cetacean carcasses far from residence sites (Peltier et al. 2012). Determining population structure based only on carcasses can fail to detect or infer erroneous patterns of population differentiation. Those patterns are crucial for understanding population structure and dynamics and imperative for management decision-making (Bilgmann et al. 2011). The absence of data from living specimens from Puerto Rico that could lead to a better understanding of the population dynamics of dolphins was the main motivation for undertaking this study.
In the Caribbean Sea and adjacent waters, there are few studies of the genetic structure of known populations, but results suggest significant population differentiation (Caballero et al. 2011). In northern Bahamas (Parsons et al. 2006) and Western North Atlantic (Shintaku 2021), a fine-scale population structure was found between three Tursiops populations suggesting the existence of possibly different units for conservation and management. In Bocas del Toro, in the Caribbean side of Panama, low genetic diversity was found within a well-monitored population (Barragán-Barrera et al. 2013, 2017). Similar results have been reported elsewhere (i.e. Australia, Allen et al. 2016; South Pacific, Sanino et al. 2005; Black Sea, Viaud-Martinez et al. 2008) showing genetic differentiation among regional populations and, in some cases, low diversity (e.g. Fruet et al. 2014) in this highly mobile species. However, there are reports of populations that do not show differentiation, as in the case of the bottlenose dolphins off the mid-North Atlantic, which exhibited shared haplotypes between inshore and offshore types (Castilho et al. 2015). Interestingly, a recent study (Duarte-Fajardo et al. 2023) conducted in the western Caribbean utilizing samples collected in Panama and Colombia, found two genetic forms in Colombian waters, as was found previously in Puerto Rico (Tezanos-Pinto et al. 2009; Waring et al. 2011; Caballero et al. 2011). Furthermore, it was discovered that mtDNA bottlenose dolphin haplotypes from Colombia were nested with haplotypes from Puerto Rico and Honduras, which are classified in the same phylogroup as the worldwide distributed form.
Although the presence of both forms has been reported in Puerto Rico (Tezanos-Pinto et al. 2009; Waring et al. 2011; Caballero et al. 2011), no assessment has been done to determine their extent, distribution, and if there are any interactions between the two forms in the free-ranging population. The unclear composition (e.g. numbers, distribution) and the relationship between the two forms present challenges in managing this species as defined in the Marine Mammal Protection Act of 1972 and the mandatory stock assessments for marine mammals in the U.S. Caribbean.
Rodriguez-Ferrer (2001) reported an estimated census population size of 314 individuals for the southwest coast of Puerto Rico with a more coastal distribution. This work was focused on abundance and distribution, but no information was collected about the population structure and the presence/absence of the two forms in that region. Thus, the objectives of this work were: 1) to characterize the genetic variability and structure, sex ratio, and group composition of bottlenose dolphins throughout Puerto Rico by analyzing DNA from live-biopsied individuals from the south and west coast as well as opportunistic, island-wide strandings; and 2) determine the genetic relationships between T. truncatus dolphins from Puerto Rico, the Caribbean, and worldwide based on a portion of the mitochondrial control region.
Materials and methods
Study area: Puerto Rico
Sampling of free raging dolphins was conducted on the waters off the south and west coast of Puerto Rico (18º 12′ 06″ N, 66º 39′ 52.24″ W) (Fig. 1). Puerto Rico is an archipelago of approximately 140 geographic structures, including islands, and islets of various sizes, surrounded mostly by deep waters (Méndez-Méndez and Fernández 2015). Surrounding Puerto Rico is an insular, narrow shelf on the northern and east coasts (Scheneidermann et al. 1976). The western insular shelf is wide and extends from six to 26 km with an average depth of 18–20 m (Schlee et al. 1999; Ballantine et al. 2008). On the south coast, the insular shelf extends east and narrows again along the eastern coastline (Morelock et al. 1994).
Biopsy sampling surveys were conducted from Aguada in the north to the island of Caja de Muertos in the south (Fig. 1) during two periods (18–31 August 2014 and 19–30 October 2015) when dolphin sightings were reported to peak (Rodriguez-Ferrer 2001). The survey was focused on known dolphin distribution areas (Rodriguez-Ferrer et al. 2017). The survey transects were predetermined based on earlier surveys conducted in the regions (Rodriguez-Ferrer 2001; Rodriguez-Ferrer et al. 2017), which confirmed the existence of a resident population and documented the presence of different coastal and offshore forms.
The surveys were conducted in an open 7-m boat, offering a 360° field of view. Sampling was attempted only under favorable weather conditions (Beaufort scale up to 3; equivalent of a wave height 0.91 m or less). Once a group of dolphins was sighted, data on behaviour, group size and composition were recorded before sampling. In addition, visible diagnostic offshore/coastal characteristics described for bottlenose dolphins in the Caribbean were recorded to distinguish between forms. For the offshore form, the characters used were a large size and bulky body, falcated fin, dark coloration, short rostrum in proportion to body size, and/or a white saddle patch (Herzing and Elliser 2016; Ramos et al. 2016; Van Waerebeek et al. 2017). For the coastal form, we searched for light coloration, small body size, no saddle patch, and rostrum in proportion to body size (Mead and Potter 1995; Ramos et al. 2016).
The boat was then positioned parallel to the swimming group. Skin samples of 27 free ranging dolphins were collected using a standard biopsy protocol (Sinclair et al. 2015). Darts and tips especially designed for small cetaceans (F. Larsen, Ceta-Dart, ACC darts, with floats and vanes for crossbow and sampling heads M8/40 mm) were deployed with a crossbow from a trained licensed marksperson. Adult dolphins were biopsied along their flank below the dorsal fin (Gorgone et al. 2008). The individual was photographed during sampling for identification and cataloguing purposes based on dorsal fin morphology and/or any scarring present. Pictures were then compared and included in an existing dorsal fin catalog (Rodriguez-Ferrer et al. 2017). Tissue samples were preserved in liquid nitrogen and stored in an – 80 °C freezer. We conducted fieldwork under permits from the National Marine Fisheries Service, Southeast Fisheries Science Center, Marine Mammal Protection Act (Scientific Permit Number 14450-01) and the Puerto Rico Department of Natural and Environmental Resources (Permit 2015-IC-047).
Data set of this study
Skin samples collected from stranded dolphins around Puerto Rico were included in the data set. Necropsy reports, if available, were reviewed for pictures and/or descriptions of the specimens to infer gender and possible ecotype. The samples included eight stranded dolphins covering the years between 2006 and 2018 from the Puerto Rico Department of Natural and Environmental Resources tissue bank, and three samples from Puerto Rico from the Caribbean Manatee Conservation Center (2001–2016) (Fig. 2). Also, for comparison purposes, five samples from the Guadeloupe Stranding Network (2013–2015) were included in the set. Finally, 334 control region sequences were extracted from GenBank to augment our dataset (Supplementary Material Table S1), bringing the total to 377 sequences. The samples from Guadeloupe, as well as the GenBank records, were included for comparison purposes since the second objective of this research was to place the Puerto Rico dolphins in the context of the wider distribution of the species in the Caribbean, the North, and South Atlantic and the North and South Pacific. We assigned a dolphin to either coastal/inshore (we kept the nomenclature of the source for continuity) or offshore form based on the classification given by the authors of the studies we included (Supplementary Material Table S1). This was not possible in all cases (178 out of 334 dolphins included in this study). For example, the dolphins from New Zealand (Tezanos-Pinto et al. 2009) had no form information and were excluded from the Pacific group in all statistical tests performed in Arlequin (Tables 1, 2, 3). None of these classifications was based on morphometry to distinguish the two forms; rather, they were based on DNA sequence clustering.
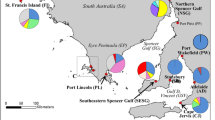
Distribution of samples of Tursiops truncatus from the current study and those of Caballero et al. (2011). Triangles (\({\blacktriangle }\)) represent live animals sampled in this study, circles (●) represent stranded dolphins (years 2001–2018), and squares (\({\blacksquare }\)) represent dolphins stranded in Puerto Rico (1994–2003), as reported in Caballero et al. (2011). Inshore haplotypes are represented by green and purple colors (haplotypes 108 and 124), while offshore haplotypes are colored white, orange, ochre, red, and yellow (haplotypes 9, 12, 46, 72, and 76, respectively)
DNA extraction, PCR and sexing
DNA was extracted from skin samples using the DNeasy kit (Qiagen, Valencia, CA, USA). A 550-bp region of the mitochondrial control region was amplified using the primers tPro-whale (5ʹ-TCACCC AAAGCTGRARTTCTA-3ʹ) and Dlp-5 (5ʹCCATCGWGATGTCTTATTTAAGRGGAA-3ʹ) (Baker et al. 1998) following the same amplification conditions as in Caballero et al. (2011). PCR products were cleaned from excess primers and dNTPs with the ExoSAP-IT™ PCR Product Cleanup Reagent kit (Fisher Scientific, Pittsburgh, PA) and sequenced with Sanger sequencing (Sanger and Coulson 1975). Sex of live animals was determined by a molecular essay where a PCR reaction was performed with the primers TtSRYR (5ʹ-ACCGGCTTCCATTCGTGAACG-3ʹ), PMSRYF (5ʹ-CATTGTGTGGTCTCGTGATC-3ʹ) (Richard et al. 1994), ZFX0582F (5ʹ-ATAGGTCTGCAGA CTCTTCTA-3ʹ) (Bérubé and Palsboll 1996), ZFX0923R (5ʹ-AGAATATGGC GACTTAAGAACG-3ʹ) (Bérubé and Palsboll 1996). We followed the PCR conditions as outlined in Rosel (2003). For stranded dolphins, the sex was determined visually during necropsy.
Data analysis
All successful PCR amplicons were purified from excess primers and unincorporated dNTPs using 4 μL of ExoSAP-IT per 5 μL of PCR product. Samples were plated on 96-well sequencing plates and were processed for Sanger sequencing in both directions using the Big Dye 3.1 Terminator Cycle Sequencing Kit. The ethanol-precipitated products were loaded into an ABI 3130xl 16-capillary Genetic Analyzer at the Sequencing and Genomics Facility of the University of Puerto Rico, Rio Piedras. All DNA sequences have been submitted to GenBank (control region: Accession Numbers PP779129-PP779168).
The DNA traces were visually inspected for quality and accuracy in nucleotide base assignment in Codon Code Aligner v. 8.0.2 (Codon Code Corp.). Sequences were trimmed in Codon Code Aligner and then aligned by the MAFFT Algorithm v. 7 (Bandelt et al. 1999; Katoh and Standley 2013) for further analyses. DnaSP v.6 (Rozas et al. 2017) was used to assign sequences to either coastal/inshore or offshore per location (western North Atlantic inshore, eastern North Atlantic coastal, Caribbean inshore, eastern North Atlantic offshore, western North Atlantic offshore, Caribbean offshore). The best number of phylogroups present in our data, was determined by estimating the highest and significant global FCT (the proportion of genetic variability found among groups that collaterally indicates the best genetic grouping of the data; FCT = 0.173, P = 0.001; Supplementary Material Table S1). The obtained phylogroups were three: Group 1 Atlantic coastal/inshore (western North Atlantic inshore, eastern North Atlantic coastal, Caribbean inshore), Group 2 Atlantic offshore (eastern North Atlantic offshore, western North Atlantic offshore, Caribbean offshore) and Group 3 Pacific (Pacific inshore, Pacific offshore). We then estimated DNA summary statistics (e.g. nucleotide diversity indices and neutrality test statistics Table 1), AMOVA analysis (Table 2), population pairwise FST and ΦST comparisons (Table 3) in Arlequin (Excoffier and Lischer 2010).
The haplotypic data for the population and phylogenetic analysis was constructed as follows: Identical sequences were collapsed to haplotypes in DnaSP with sites with gaps and missing data considered and non-considered. When we included all gaps/missing data we extracted 204 haplotypes and when we excluded them, we extracted 130 haplotypes to be used in the network analysis and phylogenetic reconstruction. We have undertaken the more conservative approach by excluding the sites with missing data. Arlequin files were then generated in DnaSP for downstream analysis.
The female effective population size (Nef) for Puerto Rico populations was estimated using the formula Nef = θ/2 μg, where μ = bp substitution rate per generation and θ = genetic diversity. We used generation time (g = 10 years) as it has been estimated for bottlenose dolphins (Cassens et al. 2005) with a mutation rate of 1.5e−7 (Hoelzel et al. 1991).
Haplotype networks were illustrated with a median-joining network algorithm (Bandelt et al. 1999) using the software PopART v. 1.7.2 (Leigh and Bryant 2015) to depict the geographic distribution of haplotypes and their relatedness visually. Sequence divergences between sequences and inferred phylogroups were estimated in PAUP* (Swofford 2001) using the appropriate model of nucleotide substitution as estimated by the BIC criterion in jModelTest2 (Darriba et al. 2012).
Phylogenetic relationships among dolphin sequences were inferred with the maximum likelihood in RaxML-ng (Kozlov et al. 2019) using 200 bootstrap replicates to assess branch support. We used a control region sequence of an Atlantic spotted dolphin (Stenella attenuata, GQ504130.1) as outgroup. Phylogenetic trees were visualized in iToL (Letunic and Bork 2019) and improved with Adobe Illustrator.
Results
Weather conditions restricted the survey time in the offshore waters, and all sightings were recorded on the nearshore waters (Fig. 1); therefore, the sampling is biased towards the nearshore environment. None of the sighted individuals had notable offshore form characteristics (Herzing and Elliser 2016; Van Waerebeek et al. 2017). A total of 27 biopsy samples were collected from the free ranging individuals, while 33 samples came from stranded animals (Fig. 2).
The sex ratio was 31 females and 29 males. There were four males and one female from Guadeloupe. Genetic analysis showed that 19 of the free-ranging biopsied dolphins belong to the coastal form, and eight biopsied dolphins were of the offshore form. Two of the eight dolphins exhibiting the offshore form have been sighted before in previous surveys, these were identified as males by the molecular essay. The sex ratio for the offshore form was six males and two females. For the coastal form, eight dolphins were re-sighted. Sex ratio of the re-sighted coastal dolphins was seven males and one female. Samples that came from recent strandings included nine offshore and two coastal dolphins. The sex ratio for stranded offshore dolphins was six males and three females, and the coastal form was two females and no males. None of the stranded dolphins could be matched with fins from the fin catalog.
The size range for stranded dolphins in Puerto Rico ranged from 111 to 259 cm in total length. The average length for a stranded offshore dolphin was 231.7 cm while the average size for a coastal dolphin was 226.4 cm. There is no significant difference between the total length of stranded offshore versus the stranded coastal dolphins (One-Way ANOVA, F1,34 = 0.182, P = 0.675). Offshore dolphins are the dominant form in strandings for both sexes and year classes. The north coast of Puerto Rico had the most offshore strandings followed by the south coast (Fig. 2). Strandings of the coastal form were present on all coasts but in a lower frequency (1–3 animals per coast). In total, the free-ranging dolphins included 19 coastal and eight offshore forms, and the stranded dolphins included seven coastal and four offshore forms (Supplementary Material Table 1).
The best fit-model of nucleotide substitution to estimate sequence divergence among coastal and offshore haplotypes from Puerto Rico (Table 4) was selected in jModelTest2, where I = 0.560 and gamma shape (alpha) = 0.359. The offshore haplotypes of Puerto Rico differed from 4.34 to 6.58% from the coastal haplotypes (Table 4). The smallest sequence divergence was observed between the offshore Hap 12 and 14 (0.27%) and the largest between the offshore Hap 12 and the coastal Hap 108 (6.5%). The range of sequence divergence within coastal and offshore dolphin haplotypes was 2.17% and 0.27–3.86%), respectively (Table 4).
The effective population size of the coastal dolphins ranged from 867 to 2400, ((Ne = 0.0026/(2*10*1.5 × 10−7)) and (Ne = 0.0072/(2*10*1.5 × 10−7)), respectively) while for offshore dolphins ranged from 1400 to 3333, ((Ne = 0.0042/ (2*10*1.5 × 10−7)) and (Ne = 0.01/(2* 10*1.5 × 10−7)), respectively).
Haplotype network findings
To reconstruct the haplotype network, the 43 newly generated control region sequences were combined with 334 sequences from GenBank (Fig. 3, Supplementary Material Table S1). Haplotype analysis based on the Median-Joining network (Fig. 3) showed a complex haplotypic structure characterized by the high abundance of singleton sequences (n = 96). The geographic subdivision of haplotypes is mostly visually detected in the Pacific and the eastern Atlantic groups. The most common haplotype of our data set (Hap 124; n = 29) consisted mostly of Caribbean dolphins, while the second most common (Hap 93; n = 26) was exclusive of the Pacific basin. Hap 46 (n = 21) was mostly present in the eastern Atlantic, but several dolphins from the northern Atlantic and Caribbean shared this haplotype. Hap 38 and Hap 41 sequences were shared by two Pacific and western North Atlantic Ocean dolphins, respectively.
Most of the sequences generated from live dolphins of Puerto Rico in the current study belonged to Hap 124 (n = 20), followed by Hap 72 (n = 5) and Hap 46 (n = 2) (Figs. 3 and 4). Hap 124 was shared with previously sampled dolphins from Mexico and Puerto Rico (Caballero et al. 2011), and with the Bahamas (Parsons et al. 2006) (Fig. 3). Hap 46, 76, 72 were shared with dolphins from Costa Rica (Barragán-Barrera et al. 2017), and the Puerto Rico Caballero’s data set (Caballero et al. 2011) all corresponding to the “world distributed form”. Hap 124 (n = 5) and Hap 72 (n = 4) were also present in the stranded dolphins (Fig. 5). Three additional haplotypes were detected from stranded dolphins (Hap 76 (n = 5), Hap 78 (n = 1), and Hap 12 (n = 1)). The dolphin represented by Hap 78 was from Guadeloupe, and Hap 12 represents a female who was stranded on the north coast of Puerto Rico (Fig. 3). Interestingly, Hap 12 is predominantly present (n = 10) in Azores (Quérouil et al. 2007). For Puerto Rico, we identified two haplotypes (108 and 124) as coastal haplotypes and five haplotypes (9, 12, 46, 72 and 76) as offshore haplotypes (Figs. 4, 5). Most live dolphins were coastal form (e.g. Hap 124, n = 20 live, n = 2 stranded from Caballero et al. (2011), n = 1 stranded, current study).
Haplotype sequences based on the mtDNA control region of Tursiops truncatus from Puerto Rico. Live and stranded animals have been included in the current study and those reported in Caballero et al. (2011). The median-joining network algorithm (epsilon = 0) was used as implemented in PopArt
Maximum likelihood tree depicting the phylogenetic relationships of the 130 haplotypes based on the mtDNA control region of Tursiops truncatus from the Atlantic (shown in black text) and Pacific Ocean (shown in magenta text). The closely related dolphin species, Stenella attenuata, is the designated outgroup. Blue circles on the branches indicate bootstrap values above 70%. The eight haplotypes found in Puerto Rico are indicated in red, bold letters. Hap 38 and Hap 41 were present in both the Pacific and Atlantic oceans. Hap2 was represented by a long branch, which has been truncated for better viewing of the tree. The arrow indicates a coastal form exclusive clade
The haplotype network based on control region sequences is not characterized by distinct demarcations of bottlenose dolphins, neither by geography nor by form. A notable exception is the cluster of haplotypes consisting of North-West Atlantic /Caribbean coastal forms on the right side of the network (Fig. 3). The two coastal haplotypes from Puerto Rico (108 and 124) are part of this cluster. Coastal dolphins are present in the Pacific, W. North Atlantic, and the Caribbean, including Central America. The offshore dolphins are more numerous in our data set and are present in all sampled areas, including the eastern North Atlantic (Fig. 3). A few reported coastal and offshore haplotypes mostly from the Pacific and some from the western North Atlantic (e.g. 32–35, 40–44, and 99–102) are genetically similar, oftentimes different by 1 base substitution (Fig. 3).
Population structure and genetic diversity
Even though bottlenose dolphins like other marine mammals are highly mobile, they show strong site fidelity to specific areas mostly driven by group behavior and niche specialization (Louis et al. 2014). Therefore, knowing the standing genetic variation of each region should be regarded as baseline information. The highest number of haplotypes were found in the western Atlantic inshore/coastal dolphins (n = 38, h = 0.99). The second highest number of haplotypes was recorded in eastern Atlantic offshore dolphins (n = 31, h = 0.94), but also this group had the highest number of individuals (N = 120). The lowest haplotypic values were found in the coastal dolphins of Puerto Rico (n = 25, h = 0.08). All phylogroups (Atlantic coastal, Atlantic offshore, and Pacific) exhibited relatively high π and θ values ranging from 0.01 to 0.03. The highest θ value was recorded in the offshore dolpins of Puerto Rico, a subgroup of the Caribbean offshore population (Phylogroup 2, Table 1). The Tajima's D statistic was only significant in the coastal dolphins of Puerto Rico indicating the lack of genetic diversity (2 haplotypes out of 25 mtDNA-CR sequences) in the group or the presence of negative selection. The Fu’s Fs test statistic was significantly different than was expected under neutrality in Atlantic inshore and Pacific offshore (Table 1). Highly negative values of Fu’s Fs statistic are driven by the excess of singletons, suggesting a possible past population expansion event.
The AMOVA test based on the mitochondrial control region indicated that there is a significant population structure (FST = 0.409, P < 0.001) when partitioning the DNA sequences into three groups (Atlantic coastal, Atlantic offshore, and the Pacific) (Table 2). Most genetic variation (∼59%) is allocated within populations (Table 2). High FST values characterize species with distinct populations, with very limited to no gene flow. In the case of the bottlenose dolphins, given our data, the two mitochondrial lineages of the coastal and offshore dolphins in the Atlantic are genetically differentiated (e.g. see DNA divergence values in Table 4 for coastal vs. offshore specimens in Puerto Rico). Pairwise FST and ΦST comparisons based on haplotype frequencies and nucleotide data, respectively, confirmed the presence of significant sequence divergence among populations, including those between the dolphins from the Pacific Ocean and the two dolphin forms found in the Atlantic Ocean (Table 3). The Pacific inshore bottlenose dolphins were the most differentiated group and the western North Atlantic offshore form was the least differentiated group (Table 3).
Phylogenetic analyses
The phylogenetic analysis based on maximum likelihood (ML; Fig. 5) yielded rather similar groups as the haplotype network in Fig. 3. The coastal dolphin group with representatives from the Caribbean, Central America and western North Atlantic was also detected with the ML analysis and supported by > 50 bootstrap value (Fig. 5; clade identified with an arrow). The dolphins from the Pacific marked with the magenta color are distributed on the rest of the tree, intermingled with those of the Atlantic, including the six offshore haplotypes from Puerto Rico (Fig. 5; in red). The eight haplotypes from Puerto Rico were divided into three visible groups as in Fig. 4, however the largest sequence divergence values are observed between coastal and offshore dolphins (Table 4). The group of genetically similar coastal and offshore haplotypes (e.g. 32–35, 40–44, 99–102) was clustered near the coastal clade which consisted of the haplotypes 107–120 and 124–130 (mostly from the Pacific Ocean and a few from the western North Atlantic Ocean).
Discussion
This study confirms the presence of genetically distinct forms in Puerto Rico, not only in stranded dolphins as previously reported, but also in the free-ranging population of the south and west coasts. The two genetically distinguished forms are the inshore, represented by coastal individuals and the worldwide-distributed, represented by both coastal and offshore dolphins. The biased sex ratio of biopsied individuals (17 males:10 females) was due mainly to dolphin behavior as males tend to interact more with the sampling boats (Quérouil et al. 2009); therefore, because of the small number of samples and sex-specific behavior, the reported sex ratio does not represent the true population sex structure. The present study has identified a distinct distribution pattern for the bottlenose dolphin population in Puerto Rico. Analysis of mtDNA data has indicated a prevalence of the coastal form in the collected samples, while strandings primarily involve the offshore individuals (worldwide distributed form). In this study, we assume that stranding dolphins had some level of association with the Puerto Rican waters. The caveat in this scenario is that stranding carcasses can be carried far from their area of origin.
Even though all the 27 biopsied dolphins in this study were sampled in nearshore waters, we identified six dolphins belonging genetically to the offshore form. At the time of sampling, we assumed that these dolphins were of the coastal form because of their smaller size, not bearing any of the cranial or fin diagnostic characteristics of the offshore form, which we could easily distinguish during our field expeditions. Interestingly, two of these six dolphins were males, and were sighted interacting with a group of genetically identified coastal form dolphins. Therefore, we report instances of interaction between forms, which could lead to possible inbreeding opportunities between forms, or perhaps an overlap in distribution with some social interaction but without gene flow (Segura et al. 2006). Alternatively, there is a possibility that these "offshore" individuals, who exhibited coastal characteristics and were biopsied in coastal areas of Puerto Rico, are members of the "worldwide distributed form" previously described in the Caribbean by Caballero et al. (2011). The "worldwide distributed form" of bottlenose dolphins has been proposed to consist of coastal and offshore dolphins. Our genetic data strengthens previous findings reported by Caballero et al. (2011) that the two genetically distinct forms could be found in sympatry in the region. However, we have also observed and biopsied dolphins with clear “offshore” morphological and genetic characteristics away from the insular shelf of Puerto Rico. More field observations are needed to enhance our understanding of the forms, sex ratio and geographic distribution of bottlenose dolphins swimming in the waters of Puerto Rico.
The non-significant difference in total length among the forms for stranded dolphins could suggest that Tursiops in the Caribbean have adapted to warmer conditions; therefore, the size of the offshore form would be similar to that of the coastal form to the naked eye. Our live animal data set included six genetically defined offshore Tursiops that, when sighted in the sea, were identified as coastal due to the size and coloration, strengthening our hypothesis that in some areas of the Caribbean, such as Puerto Rico, the species might have adapted differently than in other regions. Contrary to continental regions, the Caribbean islands have, on average, narrow insular shelves (Hubbard et al. 1981; Smith et al. 1997; Claro and Lindeman 2003; Betancourt et al. 2012); therefore, the coastal form has adapted to the conditions of strong currents, deep waters even near the coast. In other regions the morphological differences are well marked and the morphological distinction between forms is clear. When we compare these areas with the Caribbean, these zones have large shelves, enclosed bays, or estuaries with calmer and shallow waters (Mead and Potter 1995; Segura et al. 2006; Fruet et al. 2017).
No unique dolphin haplotypes have been identified so far in Puerto Rico. All haplotypes in the live animals (this study) and stranded animals (Caballero et al. 2011) from Puerto Rico have been reported elsewhere in the Caribbean. Haplotype 124 is the most common in the Caribbean, indicating that what is common in Puerto Rico is common in the Caribbean. The presence of Hap 12 from a stranded animal from Puerto Rico shared with the Azores could be indicative of a possible migratory population that passes by the north coast of Puerto Rico. Although carcasses drift and may distort the true population range, the fact that a female with the offshore form (large, falcated dorsal fin, short rostrum, offshore mtDNA) stranded in Puerto Rico, may indicate that migratory dolphins could be passing close to the coastal waters of Puerto Rico and increasing the potential of long-range gene flow between the two sides of the Atlantic (Quérouil et al. 2007). There is a need to sample then free-ranging individuals on the north coast of Puerto Rico to help determine if this was an isolated case of a migratory group close to Puerto Rico or that the population shared mtDNA with individuals from the North Atlantic, suggesting recent gene flow among regions (Silva et al. 2008; Castilho et al. 2015). Alternative hypotheses have been suggested to explain the presence of shared haplotypes among distant regions such as the evolutionary interconnection between bottlenose worldwide (Caballero et al. 2011) and possible founder events in the offshore form (Natoli et al. 2005; Tezanos-Pinto et al. 2009; Caballero et al. 2011).
The estimated census population size (Nc = 314) of Puerto Rico’s bottlenose dolphins is based on mark-recapture data (Rodriguez-Ferrer 2001). The census size estimation may be an underrepresentation of the dolphins in Puerto Rico as indicated by the female effective population size of the coastal dolphins (867–2400) and offshore dolphins (1400–3333). Since large areas of Puerto Rico were not surveyed during the Rodriguez-Ferrer (2001) study, the true Nc is likely at least an order of magnitude larger. A great deal of resources (e.g., numerous boats and observers with a lot of time) would be required to survey the waters of Puerto Rico and improve the estimate of the census size; genetic data offers an alternative, cost-effective approach to estimating population statistics.
The presence of three phylogroups in the included mtDNA-CR data set is indicative of the geographic range a wide-roaming species such as bottlenose dolphins can attain but also indicates the presence of behavioral, genetic, and geographic subdivisions within the species. The inclusion of sequences from the wider Caribbean Sea and the North Atlantic showed us that none of the eight haplotypes encountered in Puerto Rico are unique. Rather, they are shared with other dolphins from the Caribbean and the western Atlantic region. At the same time, there are more genetic differences between the coastal and “world-distributed” forms in Puerto Rico than two dolphins of the same form inhabiting different geographic regions (e.g. northeast Caribbean vs. west North Atlantic). The genetic variability present observed in dolphins sampled from Puerto Rico provides useful data to resource managers responsible for implementing conservation strategies for this charismatic mammal. However, the genetic comparisons of locally sampled dolphins against other populations and forms from other geographic locations and different oceans provide insights into the complex composition (e.g., genetic forms, wide distribution) and rudimentary knowledge of the interactions of the two bottlenose dolphin forms throughout the distribution of species.
The phylogenetic analysis generated groups similar to those in the haplotype network, supporting our coastal and offshore classification. This is more evident in the Caribbean and subsequently in Puerto Rico where the two forms can be identified genetically. These forms are considered parapatric populations, and they have the potential to overlap in distribution and they do, as offshore dolphins were observed to interact with coastal dolphins on one occasion in the current study. Yet, they are not interbreeding as far as our samples and markers indicate, evidenced by the absence of shared control region haplotypes between coastal and offshore dolphins, since this marker is maternally inherited. More samples are required to rule out the possibility of introgression between dolphin forms. Depending on the directionality of introgression there could be a decoupling of a form and its mtDNA. For example, mating between a male offshore and a female coastal dolphin would result in progeny carrying the mother’s coastal mtDNA since most animals inherit that genome in a matrilinear fashion. If introgression is widespread, it may explain the presence of coastal mitochondrial haplotypes in offshore forms.
The haplotype diversity found here in Puerto Rico is comparable with other studies of the Caribbean region. Coastal populations are characterized by low haplotype diversity. In the Caribbean, low haplotype diversity has been reported for the Bahamas and Panama (Parsons et al. 2006; Barragán-Barrera et al. 2013). The haplotype diversity for the coastal Caribbean population reported by Caballero et al. (2011) (h = 0.578), is much higher than the one we reported for the coastal population of Puerto Rico (h = 0.080). The existence of low haplotype diversity for the dolphins of Puerto Rico could indicate the small habitat area Puerto Rico provides for dolphins compared to the whole Caribbean. The low genetic diversity could also be explained by the highly philopatric nature of the inshore individuals. The assumption of dolphins' site fidelity should be evaluated by more biological and photo-ID analysis in Puerto Rico. Low genetic diversity could be indicative of reduced population mean fitness and inbreeding among coastal dolphins and should be considered when resource managers in Puerto Rico make conservation plans for the species. Rodriguez-Ferrer et al. (2017) reported a prevalent nearshore distribution of the population; therefore, anthropogenic impacts could be detrimental for a small population with low genetic diversity. In contrast, as expected for the offshore population, we found high haplotype diversity, like Caballero et al. (2011) reported for the region (Puerto Rico h = 0.724 vs. Caribbean h = 0.710). The Puerto Rican population showed a high degree of genetic sequence divergence among the two forms, but when compared to the rest of the region there is no genetic differentiation. Since all haplotypes of Puerto Rico are shared with those of the Caribbean, it indicates that long swimming distances for a strong swimming mammal and lack of obvious natural barriers do not hinder gene flow among Caribbean locations. The ability of long-distance movements of bottlenose dolphins is evident by the number of shared haplotypes between western Atlantic and the Caribbean and across both northern Atlantic coasts. Of the regions analyzed further, the eastern North Atlantic (results not shown) is characterized by high gene diversity, with most dolphins being identified as offshore (Natoli et al. 2005; Quérouil et al. 2007). Offshore dolphins tend to harbor higher genetic diversity than coastal ones even across considerable spatial scales (Quérouil et al. 2007; Tezanos-Pinto et al. 2009; this study in Table 1).
We provided evidence to support the hypothesis that the bottlenose dolphin population in Puerto Rico is part of the Caribbean stock, as has been reported previously (e.g., Barragán-Barrera et al. 2017; Caballero et al. 2011; Duarte-Fajardo et al. 2023; Tezanos-Pinto et al. 2009). The inshore haplotypes of Puerto Rico belong to the phylogroup, which includes the eastern and western North Atlantic coastal/inshore haplotypes, and the offshore Puerto Rico haplotypes belong to the phylogroup, which includes the eastern and western North Atlantic offshore haplotypes. The bottlenose dolphins from the Pacific were distinct genetically and formed another phylogroup. Our results are based on the public sequences we used and the mtDNA-CR, a matrilineal inherited marker. The analysis of matrilinear lineages tells us only part of the story; therefore, a more in-depth analysis of the samples using nuclear markers or next-generation sequencing SNP data could aid in understanding the population shifts and changes. Although our analysis shows that the Puerto Rican population is not distinct from the contiguous Caribbean population, it is important to remember that complex species such as dolphins can adapt to their environment by changing their behaviors, feeding habits, and migratory patterns. Data from photo identification surveys established a resident population, indicating that, while the species appears to be genetically diverse, there is a resident population, with estimates that fall into the small size.
Bottlenose dolphins are commonly sighted in west and southwest Puerto Rico, the most touristic marine region of the island. The coastal communities in the region heavily rely on tourism and commercial fishing, activities that pose possible adverse effects on dolphins by the increased boat traffic, chemical and noise pollution, and reduction of available prey through fishing. These are among the most common threats against marine mammals identified worldwide (Avila et al. 2018). At the same time, marine resource agents are challenged to make management recommendations for bottlenose dolphins as defined in the Marine Mammal Protection Act of 1972 because of uncertainties regarding the distribution and abundance of the two forms in Puerto Rico. Long-term data on dolphin strandings, ecology and distribution of ecotypes, sex-ratio fluctuations, genetic diversity, and population connectivity can assist management decisions in Puerto Rico.
References
Allen SJ, Bryant KA, Kraus RH, Loneragan NR, Kopps AM, Brown AM, Gerber L, Krützen M (2016) Genetic isolation between coastal and fishery-impacted, offshore bottlenose dolphin (Tursiops spp.) populations. Mol Ecol 25(12):2735–2753
Armstrong RA, Singh H (2012) Mesophotic coral reefs of the Puerto Rico Shelf. In: Harris PT, Baker EK (eds) Seafloor geomorphology as benthic habitat: Geohab atlas of seafloor features and benthic habitat. Elsevier, London, pp 365–374
Avila IC, Kaschner K, Dormann CF (2018) Current global risks to marine mammals: taking stock of the threats. Biol Conserv 221:44–58
Baker CS, Flórez-González L, Abernethy B, Rosenbaum HC, Slade RW, Capella J, Bannister JL (1998) Mitochondrial DNA variation and material gene flow among humpback whales of the Southern Hemisphere. Mar Mamm Sci 14(4):721–737
Ballantine DL, Appeldoorn RS, Yoshioka P, Weil E, Armstrong R, Garcia JR, Otero E, Pagan F, Sherman C, Hernandez-Delgado EA, Bruckner A, Lilyestrom C (2008) Biology and ecology of the Puerto Rican coral reefs. In: Riegl BM, Dodge RE (eds) Coral reefs of the USA. Springer Science, Dordrecht, pp 375–406
Bandelt HJ, Foster P, Rohl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Ecol Evol 16(1):37–48
Barragán-Barrera DC, May-Collado LJ, Quiñones-Lebrón SG, Caballero S (2013) Population at risk: low genetic diversity in bottlenose dolphins of Bocas del Toro, Panamá. Final Report. s.l.:IWC SC/65a/SM15
Barragán-Barrera DC, May-Collado LJ, Quiñones-Lebrón SG, Caballero S (2017) High genetic structure and low mitochondrial diversity in bottlenose dolphins of the Archipelago of Bocas del Toro, Panamá: a population at risk? PLoS ONE 12(12):1–22
Bearzi M, Saylan CA, Hwanga A (2009) Ecology and comparison of coastal and offshore bottlenose dolphins (Tursiops truncatus) in California. Mar Freshw Res 60:584–539
Bérubé M, Palsboll P (1996) Identification of sex in cetaceans by multiplexing with three ZFX and ZFY specific primers. Mol Ecol 5(2):283–287
Betancourt, L, Herrera-Moreno A, Beddall K (2012) Spatial distribution of humpback whales (Megaptera novaengliae) in Samana Bay, Dominican Republic. J Cetacean Res Manage SC-64-01:1–10
Bilgmann K, Möller LM, Harcourt RG, Kemper CM, Beheregaray LB (2011) The use of carcasses for the analysis of cetacean population genetic structure: a comparative study in two dolphin species. PLoS ONE 6(5):e20103. https://doi.org/10.1371/journal.pone.0020103
Boisseau O, Leaper R, Moscrop A, Embankment A (2006) Observations of small cetaceans in the Eastern Caribbean, St. Kitts & Nevis: International Whaling Commision Scientific Paper SC/58/SM24
Caballero S, Islas-Villanueva V, Tezanos-Pinto G, Duchene S, Delgado-Estrella A, Sanchez-Okrucky R, Mignucci-Giannoni AA (2011) Phylogeography, genetic diversity and population structure of common bottlenose dolphin in the Wider Caribbean inferred from analyses of mitochondrial DNA control region sequences and microsatellite loci: conservation and management implications. Anim Conserv 15(2):95–112
Castilho CS, Pedone-Valdez F, Bertuol F, Fruet P, Genoves RC, Di Tullio JC, Caon G, Hoffmann LS, Freitas TR (2015) Insights about the genetic diversity and population structure of an offshore group of common bottlenose dolphins (Tursiops truncatus) in the Mid-Atlantic. Genet Mol Res 14(2):3387–3399
Claro R, Lindeman KC (2003) Spawning aggregation sites of snapper and grouper species (Lutjanidae and Serranidae) on the insular shelf of Cuba. Gulf Caribb Res 14(2):91–106
Curry BE, Smith J (1997) Phylogeographic structure of the bottlenose dolphin (Tursiops truncatus): stock identification and implications for management. In: Dizon AE, Chivers SJ, Perrin WF (eds) Molecular genetics of marine mammals. Society for Marine Mammalogy, Kansas, pp 227–247
Darriba D, Taboada GL, Doalla R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9(8):772
Duarte-Fajardo MA, Barragán-Barrera DC, Correa-Cárdenas CA, Pérez-Ortega B, Farías-Curtidor N, Caballero S (2023) Mitochondrial DNA supports the low genetic diversity of Tursiops truncatus (Artiodactyla: Delphinidae) in Bocas del Toro, Panama and exhibits new Caribbean haplotypes. Rev Biol Trop 71(S4):e57291. https://doi.org/10.15517/rev.biol.trop.v71iS4.57291
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 3:564–657
Fahlman A, McHugh K, Allen J, Barleycorn A, Allen A, Sweeney J, Stone R, Faulkner Trainor R, Bedford G, Moore MJ, Jensen FH, Wells RS (2018) Resting metabolic rate and lung function in wild offshore common bottlenose dolphins, Tursiops truncatus, near Bermuda. Front Physiol 9:886. https://doi.org/10.3389/fphys.2018.00886
Fruet PF, Secchi ER, Daura-Jorge F, Vermeulen E, Flores PA, Simoes-Lopes PC, Genoves RC, Laporta P, Di Tullio JC, Freitas TRO, Rosa LD (2014) Remarkably low genetic diversity and strong population structure in common bottlenose dolphins (Tursiops truncatus) from coastal waters of the Southwestern Atlantic Ocean. Conserv Genet 15(4):879–895
Fruet PF, Secchi ER, Di Tullio JC, Simões-Lopes PC, Daura-Jorge F, Costa APB, Vermeulen E, Flores PAC, Genoves RC, Laporta P, Beheregaray LB, Möller LM (2017) Genetic divergence between two phenotypically distinct bottlenose dolphin ecotypes suggests separate evolutionary trajectories. Ecol Evol 7:9131–9143
Gorgone AM, Haase PA, Griffith ES, Hohn AA (2008) Modeling response of target and nontarget dolphins to biopsy darting. J Wildl Manage 72(4):926–932
Hersh SL, Duffield DA (1990) Distinction between northwest Atlantic offshore and coastal bottlenose dolphins based on hemoglobin profile and morphometry. In: Leatherwood S, Reeves RR (eds) The bottlenose dolphin. Academic Press, New York, pp 129–140
Herzing DL, Elliser CR (2016) Opportunistic sightings of cetaceans in nearshore and offshore waters of Southeast Florida. J Northwest Atl Fish Sci 48:21–31
Hoelzel AR, Hancock JM, Dover GA (1991) Evolution of the cetacean mitochondrial D-loop region. Mol Biol and Evol 8:475–493
Hoelzel AR, Potter CW, Best PB (1998) Genetic differentiation between parapatric “nearshore” and “offshore” populations of the bottlenose dolphin. Proc R Soc 265(1402):1177–1183
Hoskin CM, Reed JK, Mook DH (1986) Production and off-bank transport of carbonate sediment, Black Rock, southwest Little Bahama Bank. Mar Geol 73(1–2):125–144
Hubbard DK, Sadd JL, Roberts HH (1981) The role of physical processes in controlling sediment transport patterns on the insular shelf of St. Croix, U.S., Virgin Islands. In: Proceedings of the Fourth International Coral Reef Symposium, University of the Philippines, Manila, 18–22 May 1981, 399–404
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(2):772–780
Kozlov AM, Darriba D, Flouri T, Morel B, Stamatakis A (2019) RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. J Bioinform 35:4453–4455
Leigh JW, Bryant D (2015) PopArt: Full feature software for haplotype network construction. Methods Ecol Evol 6(9):1110–1116
Letunic I, Bork P (2019) Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47(W1):W256–W259. https://doi.org/10.1093/nar/gkz239
Louis M, Viricel A, Lucas T, Peltier H, Alfonsi E, Berrow S, Brownlow A, Covelo P, Dabin W, Deaville R, Stephanis R (2014) Habitat-driven population structure of bottlenose dolphins, Tursiops truncatus, in the North-East Atlantic. Mol Ecol 23(4):857–874
Mead JG, Potter CW (1995) Recognizing two populations of the bottlenose dolphin (Tursiops truncatus) off the Atlantic coast of North America; morphologic and ecologic considerations. IBI Rep 5:31–44
Méndez-Méndez S, Fernández R (2015) Puerto Rico, past and present: an encyclopaedia, 2nd edn. Greenwood, Santa Bárbara
Morelock J, Winget EA, Goenaga C (1994) Geologic maps of the southwestern Puerto Rico Parguera to Guanica insular shelf. Guaynabo: US Geological Survey
Natoli A, Birkun A, Aguilar A, Lopez A, Hoelzel AR (2005) Habitat structure and the dispersal of male and female bottlenose dolphins (Tursiops truncatus). Proc R Soc B Biol Sci 272(1569):1217–1226
Natoli A, Lang A, Archer E, Cipriano F, Krützen Zürich M, Rosel P, Brownell R, Hoezel R, Perrin W (2019) Report of the workshop: Resolving Tursiops taxonomy worldwide. J Cetacean Res Manage 20:523
Parsons KM, Durban JW, Claridge DE, Herzing DL, Balcomb KC, Noble LR (2006) Population genetic structure of coastal bottlenose dolphins (Tursiops truncatus) in the northern Bahamas. Mar Mamm Sci 22(2):276–298. https://doi.org/10.1111/j.1748-7692.2006.00019.x
Peltier H, Dabin W, Daniel P, Van Canneyt O, Dorémus G, Huon M, Ridoux V (2012) The significance of stranding data as indicators of cetacean populations at sea: modelling the drift of cetacean carcasses. Ecol Indic 18:278–290
Quérouil S, Silva MA, Freitas L, Prieto R, Magalhães S, Dinis A, Alves F, Matos JA, Mendonça D, Hammond PS, Santos RS (2007) High gene flow in oceanic bottlenose dolphins (Tursiops truncatus) of the North Atlantic. Conserv Genet 8(6):1405–1419
Quérouil S, Freitas L, Dinis A, Alves F, Cascão I, Prieto R, Silva MA, Magalhães S, Matos J, Santos RS (2009) Sex bias in biopsy samples collected from free-ranging dolphins. Eur J Wildl Res 56(2):151–158
Ramos EA, Castelblanco-Martínez DN, Niño-Torres CA, Jenko K, Gomez NA (2016) A review of the aquatic mammals of Belize. Aquat Mamm 42(4):476–493
Richard KR, McCarrey SW, Wright JM (1994) DNA sequence from the SRY gene of the sperm whale (Physeter macrocephalus) for use in molecular sexing. Can J Sci 72(5):873–877
Rodriguez-Ferrer G (2001) Survey of the bottlenose dolphin (Tursiops truncatus) of the southwest coast of Puerto Rico. Master Thesis, University of Puerto Rico at Mayaguez
Rodriguez-Ferrer G, Appeldoorn RS, Schizas NV (2017) Abundance of the common bottlenose dolphin Tursiops truncatus (Montagu 1821) (Mammalia: Artiodactyla) off the south and west coast of Puerto Rico. LEB 4(4):242–268
Rosel PE (2003) PCR-based sex determination in Odontocete cetaceans. Conserv Genet 4:647–649
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A (2017) DnaSP 6: DNA sequence polymorphism analysis of large datasets. Mol Biol Evol 34(12):3299–3302
Ruenes GF, Lopez R, Laeta M, Cruz D, Sanchez-Campo L, Horst Schulz U, March CG, Hohn AA, Oliviera LR (2023) Life in the Caribbean: age and growth of common bottlenose dolphins Tursiops truncatus in Cuban waters. Mar Mam Sci 39:1019–1030
Sanger F, Coulson AR (1975) A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase. J Mol Biol 94(3):441–448
Sanino GP, Van Waerebeek K, Bressem M-FV, Pastene LA (2005) A preliminary note on population structure in eastern South Pacific common bottlenose dolphins (Tursiops truncatus). J Cetacean Res Manag 7(1):65–70
Scheneidermann N, Pilkey OH, Sounders C (1976) Sedimentation on the Puerto Rico insular shelf. J Sediment Res 46(1):167–173
Schlee JS, Rodriguez RW, Webb RM, Carlo MA (1999) Marine geologic map of the southwestern insular shelf of Puerto Rico, Mayaguez to Cabo Rojo. No 2565, San Juan: Department of Interior
Segura I, Rocha-Olivares C, Flores-Ramirez S, Rojas-Bracho L (2006) Conservation implications of the genetic and ecological distinction of Tursiops truncatus ecotypes in the Gulf of California. Biol Conserv 133(3):336–346
Sinclair C, Sinclair J, Zolman ES, Martinez A, Riishøjgaard LP, Barry KP (2015) Remote biopsy field sampling procedures for cetaceans used during the Natural Resource Damage Assessment of the MSC252 Deepwater Horizon Oil Spill, Pascagoula, Mississippi: NOAA.Technical Memorandum NMFS-SEFSC-670
Smith AF, Rogers CS, Bouchon C (1997) Status of the western Atlantic coral reefs in the Lesser Antilles. In: Balboa, Panama, Proceedings of the 8th Coral Reef Symposium, Smithsonian Tropical Research Institute, June 24–29, 1996:351–356
Shintaku N (2021) Population genomics of bottlenose dolphins (Tursiops truncatus) in the northwest Atlantic. Master's project, Duke University. Retrieved from https://hdl.handle.net/10161/22665
Swofford DL (2001) PAUP: Phylogenetic analysis using parsimony (and other methods) 4.0.b5
Tezanos-Pinto G, Baker CS, Russell K, Martien K, Baird RW, Hutt A, Stone G, Mignucci-Giannoni AA, Caballero S, Endo T, Lavery S, Oremus M, Olavarría C, Garrigue C (2009) A worldwide perspective of the population structure and genetic diversity of the bottlenose dolphins (Tursiops truncatus) in New Zealand. J Hered 100(1):11–24
Van Waerebeek K, Reyes JC, Sanino GP, Félix F, Van Bressem MF, Santillán L, Montes D, García-Godos I, Echegaray M, Venegas-Abad A (2017) Common bottlenose dolphins Tursiops truncatus of Pacific South America, a synoptic review of population identification data. Document SC/67A/SM/10, IWC Scientific Committee Meeting, Bled, Slovenia, May 2017
Waring GT, Josephson E, Maze-Foley K, Rosel PE (2011) U.S. Atlantic and Gulf of Mexico marine mammal stock assessment-2010. NOAA Tech Memo NMFS NE 219(598):02543–1026
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38(6):1358–1370
Acknowledgements
Biopsy sampling was performed by Carrie Sinclair from NMFS and Aaron Barleycorn from Mote Marine Laboratory. We thank the following field assistants: Jennifer Irrizary, Maria Cardona, Nilda Jimenez, DNER Rangers Marine Unit, Jaaziel E. Garcia, Duane Sanabria, Philip Sanchez, Nicholas Hammerman, Jack Olson, Captain Anibal Santiago, and Diana Beltran-Rodriguez. We want to thank Jean Louis Georges and Manolo Rinaldi from the French Caribbean Stranding Network for the Guadeloupe samples. This research was funded by Puerto Rico Sea Grant project# R-101-1-14 to GRF, NVS and RSA. We want to thank the Southwest Fisheries Science Center, Marine Mammal and Sea Turtle Research (MMASTR) Collection for providing samples. This publication was made possible with support from the Sequencing and Genomics Facility of the UPR Río Piedras & MSRC/UPR, funded by NIH/NIGMS-Award Number P20GM103475.
Funding
Puerto Rico Sea Grant College, University of Puerto Rico, # R-101-1-14, Grisel Rodriguez-Ferrer, NIH/NGMS, P20GM103475, Nikolaos V Schizas.
Author information
Authors and Affiliations
Contributions
All authors made contributions to the conception and design of the investigation. The following individuals conducted the material preparation, data collection, analysis and writing of the manuscript: Grisel Rodriguez Ferrer, Nikolaos V. Schizas and Richard S. Appeldoorn. Antonio Mignucci and Renaldo Rinaldi provided samples from stranded individuals for analysis.
Corresponding author
Additional information
Handling editor: Laura Iacolina.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rodriguez-Ferrer, G., Appeldoorn, R.S., Mignucci-Giannoni, A.A. et al. The presence of two distinct mitochondrial lineages in the bottlenose dolphin (Tursiops truncatus) in Puerto Rico and their affinities with previously reported lineages. Mamm Biol (2024). https://doi.org/10.1007/s42991-024-00423-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42991-024-00423-5