Abstract
Sleep staging is one of the important methods for the diagnosis of the different types of sleep-related diseases. Manual inspection of sleep scoring is a very time-consuming process, labor-intensive, and requires more human interpretations, which may produce biased results. Therefore, in this paper, we propose an efficient automated sleep staging system to improve sleep staging accuracy. In this work, we extracted both linear and non-linear properties from the input signal. Next to that, a set of optimal features was selected from the extracted feature vector by using a feature reduction technique based on the ReliefF weight algorithm. Finally, the selected features were classified through four machine learning techniques like support vector machine, K-nearest neighbor, decision tree, and random forest. The proposed methodology performed using dual-channel EEG signals from the ISRUC-Sleep dataset under the AASM sleep scoring rules. The performance of the proposed methodology compared with the existing similar methods. In this work, we considered the 10-Fold cross validation strategy; our proposed methods reported the highest classification accuracy of 91.67% with the C4-A1 channel, and 93.8% with the O2-A1 channel using the Random forest classification model. The result of the proposed methodology outperformed the earlier contribution for two-class sleep states classification. The proposed dual-channel sleep staging method can be helpful for the clinicians during the sleep scoring and treatment for the different sleep-related diseases.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Motivation
Maintaining proper health and mental stableness is critical for overall health and well-being. Despite several relevant studies, sleep quality continues as a critical public challenge. Nowadays, people of all age groups are affected by improper sleep quality. Consequently, this scenario can later lead to neurological disorder diseases [1, 2]. Sleep disorders spread over with all categories of the population independently of different genders. This public challenge affects the quality of life in physical and mental health. Multiple insomnia, parasomnias, sleep-related breathing, hypersomnia, bruxism, narcolepsy, circadian rhythms are relevant examples of sleep-related disorders. Some of these disorders can be treated with proper analysis of early symptoms where ensure adequate sleep quality is essential for the patient’s recovery. Moreover, numerous sleep disorders can be nowadays clinically diagnosed through computer-aided technologies [3]. Sleep monitoring is one of the most significant activities in the assessment of sleep-related disturbances and other neural problems. Sleep is a dynamic process and includes different sleep states such as the wake stage, the non-rapid eye movement (NREM), and the rapid eye movement (REM) sleep. Furthermore, the NREM sleep states are divided into four stages, namely NREM stage 1 (N1), stage 2 (N2), stage 3 (N3), stage 4 (N4) [4]. The wake-sleep stage is the awakening period before sleep. The NREM sleep stages are sequentially indicative of light to deep sleep. Stage N1 is light sleep where the eyes move slowly, and the muscle movements are slow. The sleep starts from stage N2, where the eye movement stops, and brain activities decrease. The N3 and N4 stages are treated as deep sleep, and there are no movements in the eyes and muscle. Finally, in the REM stage, the eye movements happened rapidly, and the breathing order increase. The night sleep cycle covers 75% of NREM sleep stages and 25% of a regular sleep night [5].
The sleep assessment can be supported using a sleep test with polysomnographic (PSG) recordings. The PSG recordings are the physiological signals which are col- selected from subjects during sleep. The PSG is a collection of multivariate signal recordings such as electroencephalogram (EEG), electrocardiogram (ECG), electrooculogram (EOG), and electromyogram (EMG) [6]. The EEG signal recordings are used during sleep staging scoring. These signals represent brain activities, and therefore, are suitable for sleep abnormalities evaluation. After data collection, the sleep staging score is performed. The recorded EEG signals are extracted through multiple fixed electrodes located in different places. The process of electrode placement is done according to the international 10/20 placement system [7]. The entire process is carried out by sleep experts who analyze the different patterns of sleep states. The evaluation is made through visual inspection using the recorded data for a specific time window. Consequently, the sleep score is determined through multiple criteria. The criteria for the sleep scoring process are based on the guidelines proposed in Rechtschaffen Kales rules [8]. According to Rechtschaffen Kales guidelines, a sleep stage can be classified as wake (W), non-rapid eye movements (N1, N2, N3, and N4), and rapid-eye movements (REM). Furthermore, the proposed guidelines also include minor changes introduced by the American Academy of Sleep Medicine (AASM) [9]. The AASM manuals have represented the N3, and N4 stages into a single stage (N3) denominated slow-wave-sleep (SWS). The manual and visual evaluation of sleep scoring is complicated, costly, and time-oriented. This manual approach overloads sleep experts who have to monitor and record their patients continuously. Sometimes all these drawbacks may produce biased sleep scoring results. Therefore, these limitations demand to develop an automated sleep staging system [10, 11]. It has been found that most of the existing automated sleep staging system carried basically into two phases: (1) eliminates the irrelevant signal compositions and extracted the features (2) the extracted features fed into the classifiers for classifying the sleep stages [12, 13]. The major contributions in the sleep staging system are based on the EEG signals, some of the contributions based on the EMG signal (or) combinations of EEG, EMG, and EOG signals [14,15,16]. It has been found that most of the sleep staging based on the single-channel EEG signal [17,18,19,20]. However, this process leads to the best accuracy of the research of sleep disturbances [21]. Consequently, the development of automated detection and recognition applications to assist sleep experts in the diagnosis of sleep disorders is critical for enhanced public health.
Contribution
This work focuses on the implementation of an automated sleep stage classification system to identify irregularities that occurred during sleep from distinct medical conditioned subjects. Furthermore, analyses of the current state of the artwork in the field of sleep staging state several challenges that need to be addressed. These challenges are associated with channel selection, feature extraction, and application of the better performance classification method for enhanced accuracy. Therefore, the main aim of the proposed approach is to present the development and implementation of novel automated sleep stage classification techniques based on a single channel of EEG signal for the classification of wake and sleep phases.
Organization of the Paper
Section 2 describes a detailed literature study on sleep staging. In Sect. 3 proposed methodology including experimental data preparation, data preprocessing, feature extraction, feature screening, classification algorithms, and performance metrics used in this paper for sleep staging evaluation. In Sect. 4, we briefly discuss our proposed methodology results and make a result analysis with the state-of-the-art method. Section 5 ends with concluding remarks with future work description.
Literature Study
The different recent studies with subject to sleep staging conducted by the different authors with different methodologies and techniques. The common steps for all the sleep study that (1) data preprocessing, (2) feature extraction, (3) feature selection, and (4) classification. Some of the works it has found that they have obtained multiple channels for sleep recordings and some studies are based on single channels. Some of the contributions are based on time-domain features [13, 22,23,24,25], frequency domain features [26,27,28,29], and non-linear features [30,31,32] for characterizing the sleep stages. Some of the sleep studies are based on sleep staging different classification techniques. It has reported that maximum authors have used support vector machine (SVM) techniques [32,33,34,35,36,37], DT [38], k-nearest neighbor (KNN) [39, 40], RUSBoost [41, 42], Adaboost [43, 44], Bagging [44], Random forest [45], ANN [40].Some of the recent contributions research works with subject to sleep staging are described below here.
Berthier et al. [46] proposed five-state sleep stages classification with the input of EEG signals through fuzzy-logic iterative methods and the accuracy was reported as 82.9%.
Radha et al. [47] considered multiple EEG signals and extracted features from different domains like time and frequency. The author in this work used SVM and Random Forests (RF) were considered for the classification of different sleep stages. The outcome of the research work showed that the best performance occurred by considered frontal EEG signals.
In [48] the author introduced the wavelet tree features and neighboring component analysis and obtained two classification techniques as SVM and ANN. The data used in this research work collected from Sleep-EDF polysomnography records. The overall accuracy achieved for SVM and ANN is 90.30% and 89.93%.
Peker [20] proposed sleep staging analysis using two sleep standards R&K and AASM rules, and extracted complex non-linear features. The obtained features fed into the complex structure of the Neural Network. The accuracy results reported with R&K rules is 91.57% and with AASM is 93.84%.
Ronzhina et al. [49] conducted the sleep study with consideration of spectral features and ANN deployed for six-state classification and the accuracy reported as 82.9%.
Jo et al.[50] proposed four-state sleep stage classification and obtained a genetic-fuzzy classifier for classification and finally, the model achieved 84.6% overall classification accuracy.
Hsu et al. [19] proposed a sleep stage classification with the input of a sleep EEG record, where the authors extracted energy features and fed them into the neural network. The accuracy result reported for five-state sleep stage classification as 87.2%.
Fraiwan et al. [51] extracted time–frequency entropy features to represent the sleep records and used a linear discriminate analysis algorithm for classifying the sleep stages and the overall classification accuracy achieved for six-state sleep stages was 84%.
Eduardo et al. [52] presented an efficient and effective sleep staging scoring system and extracted the frequency behavior from the input channel. The extracted eight statistical features are forwarded to the classifier. The random forest classification model achieved an overall accuracy of 90.9%, 91.8%, 92.4%, 94.3% and 97.1% respectively.
In [53] the sleep study has been performed with EEG signals of the Sleep-EDF dataset, the input signal segmented into different signal sub-bands through butter-worth bandpass filter techniques. The SVM classification techniques deployed for two-state sleep stage classification and achieved a success rate of 92%.
Sousa et al. [54] focused extraction of entropy features from respective channels of EEG along with temporal features and extracted features are classified with SVM and it has reported an overall accuracy of 86.75%.
Heyat et al. [55] proposed a sleep study with the input of EEG signal and extracted power spectral density features and obtained features are forwarded into decision tree classifier. The reported accuracy from the proposed model is 81.25%.
Basha [56] introduced a fuzzy kernel SVM classifier and recurrent neural network for automated sleep staging, obtained statical features from five different frequency bands such as delta, theta, alpha, gamma, and beta. The model reported accuracy with FKSVM as 90.2% and with RNN as 90%.
Sharma et al. [57] introduced two-band energy localized filter techniques for analysis of the sleep characteristics of the subjects with obtaining the time–frequency features. The obtained features are forwarded to the different machine learning classifiers. The highest accuracy reported with Gaussian SVM classifier as 97.4%, 93.3%, 91.3%,89.8% and 88.7% for two-class, three-class, four-class and five- class sleep states classification.
Tian et al. [58] extracted multi-scale entropy properties from EEG signal for characterizing the signal in multiple temporal scales manner. The study was implemented with total epochs of 18,248 of 30 s length from 10 sleep disordered and 10 healthy subjects.
Alickovic et al. [59] used multi-scale principal component analysis and the informative features are extracted from signal sub-bands using discrete wavelet transform techniques. Twenty subjects participated in this ensembling sleep staging analysis.
Methodology
This paper we propose an automated sleep scoring system, where we obtained dual channels of EEG signal for discriminating the sleep behaviour of the subjects. The proposed research work was conducted with four basic steps such as (1) preprocessing the signal, (2) feature extraction, (3) feature screening and finally (4) classification. The complete flow of this proposed methodology is presented in Fig. 1. In this sleep study, we propose two sleep states classification using a dual-channel of EEG signal. The entire sleep scoring was conducted according to AASM sleep standards. Each epoch’s duration of the input signal is 30 s.
Dataset Preparation
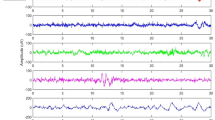
In this proposed study, one subgroup of EEG recordings in this sleep study is obtained from ISRUC-Sleep database, which prepared by the sleep experts in the sleep centre at the Hospital of Coimbra University during 2009–2013 [61]. The whole database is divided into three sections as ISRUC-Sleep subgroup-I, subgroup-II, and subgroup-III. In subgroup-I, 100 subjects one-session sleep recordings were available, all were affected with different types of sleep-related disorder and the average age of subjects in this section is 51 years, similarly, in the subgroup-II, 8 subjects sleep recordings were contained, among them 6 were male and 2 were female gender. In this section, two different recordings were collected from subjects on two different dates; the maximum subjects in this section were affected with sleep apnea events. It has been found that the average age of participated subjects for this section was 47. Finally, the subgroup-III section collected 10 subject’s sleep records, which were completely healthy, controlled, and no prior symptoms with any type of sleep problems in earlier periods. All these sleep recordings are done by the set of sleep exports at the center of sleep in the Hospital of Coimbra University. Each sleep recordings were a collection of whole-night PSG recording containing 6 EEG channel, 2 EOG channel, 3 EMG channel, 1 ECG channel, airflow, abdominal efforts, pulse oximetry, and body position information (Fig. 2, 3).
In the present work, we retrieved sleep records from two channels of EEG signals. The recorded signals are sampled with a sampling frequency of 200 Hz and the length of each epoch of 30 s. Each epoch is labeled with its class names according to its sleep stages Wake (W), NREM (N1), NREM (N2), NREM (N3), and REM(R) and its annotations are for W-0, N1-1, N2-2, N3-3, and REM-5 as per the AASM rules. In this proposed study we have considered the NREM and REM stages to be sleep stages. Both C4-A1 and O2-A1 recorded 30-s epoch’s sleep stages. The details on the dataset are presented in Table 1.
Preprocessing
Generally, it has been seen that the recorded signals are contaminated with the different irrelevant signal compositions such as muscle movement information, eye blinks information, and surrounding noises. All these signal artifacts may lead to wrong interpretations of the sleep characteristics, which ultimately produce the sleep staging results. For eliminating these artifacts from the recorded signals, we used a 10th order Butterworth band-pass filter to remove the artifacts and irrelevant noises that occurred during sleep from sleep EEG signal recordings at the frequency ranges of 0.1–35 Hz. Further to rescale the data, we applied the z-score normalization techniques. After that, the whole recorded signals are segmented into 30 s epochs and each epoch is labeled into a particular sleep stage.
Feature Extraction
For automated analysis of sleep behaviour of subjects, feature-based analysis has been most effective for identifying the different sleep characteristics of the subjects. It has been observed from further sleep studies that proper analysis of features may easier during sleep staging. As we know that brain EEG signals are highly random and non-stationary, for that reason, feature extraction takes an important role concerning sleep stages classification. The features can be classified into two broad categories: linear and non-linear methods.
Time-domain feature analysis: It is one of the direct methods for analyzing sleep EEG records. It helps to analyze the EEG signal patterns with consideration of certain wave patterns such as wave amplitude, duration, and mean value. Sometimes through time-domain analysis, we also often analyze the characteristics of signal correlation, peak detection, and variance analysis. Though the sleep records are continuously in changes form and high instability, we used to analyze the signals skewness, kurtosis, variance, mean, and median and standard deviation parameters. Generally sometimes for analysis of the EEG signal, we also used Hjorth parameters(activity, mobility, and complexity)The activity parameters help to analyze the variance concerning time function, mobility analysis the mean frequency of the signal, and complexity analyzes the changes in frequency level of the signals.
Frequency domain feature analysis: EEG signal has a strong background of frequency-domain characteristics and it plays an important role during sleep staging to characterize the different sleep EEG rhythmic waveforms. Generally, spectral power, spectral entropy, and power ratios are used to analyze the sleep behaviors with different frequency ranges. All these parameters value estimation from power spectral density. Generally frequency domain characteristics were computed from five frequency bands: delta (δ) 0.5–4.5 Hz, theta (θ) 4.5–8.5 Hz, alpha (α) 8.5–11.5 Hz, sigma (σ) 11.5–15.5 Hz, beta (β) 15.5–32.5 Hz.
Non-linear Analysis. It is also an essential part of analyzing the non-linearity properties of the signals. Specifically, EEG signals are typical of non-linear signals. It helps to understand the complex physiological behaviors of the subjects during sleep such as changes in temperature and blood pressure. Though EEG signals are contained highly subjective information, it is necessary to analyze this information’s through different non-linear parameters. The non-linear methods are more effectively analyze and handles the non-linearity presents in the recorded EEG signals. The non-linear analysis is used for the sleep stages scoring method through different entropy features such as spectral entropy, permutation entropy, Renyi entropy, fuzzy entropy, Petrosian fractal dimension, zero-crossing rate, mean Teager energy, etc.
In this proposed research work, we have considered both linear and non-linear analysis of the recorded EEG signals. In total, we extracted 34 features and the extracted features are described in Table 2.
Feature Selection
The main goal of this step is to select the most relevant features, which help dis-criminate the sleep stages features between the five sleep stages categories. Sometimes it has been found that all the extracted features may not be suitable for the classification model, and it may be one of the causes for degradation of the classification results. In this research work, we have used the ReliefF feature selection algorithm for identifying suitable features. It is one of the supervised feature weighting algorithms which evaluate the relevance of the features concerning its class labels [61]. The essential concept behind this algorithm is to select highly commendable features that help to discriminate the sleep behaviour of the subject. As an output, this algorithm assigned a weight to individual input features according to their relevance. It determines how far the features are most discriminate to different instances amounts to different sleep stages. It generates a weight for each feature and the larger the weight of the feature, the higher the association between the features and sleep stages. The main advantage of this algorithm is well managed with noisy and unknown data.
Classification
To distinguish the different characteristics of sleep stages, we employ three machine learning classification algorithms as support vector machine (SVM), decision tree (DT), KNN, and random forest (RF).
(1) SVM It is a more popular classification approach among machine learning techniques and its concept is based on statistical learning theory [62]. It supports both classification and regression methods which easily handle the classification with a huge amount of records. These classification techniques dealt with both the linear and non-linear classification problems. SVM is strongly mathematically approached and it is closely associated with some well-established theories in statistics concepts, for that reason SVM is capable to manage non-linear separation problems by introducing the hyper-plane idea and kernel function. In our experimental part, the radial basis function (RBF) was used as a kernel function [63].
(2) Decision Tree Classifier (DT): It is one of the structured and comprehends classification techniques incomparable to another classification algorithm. Majorly DT used by different type’s classification tasks [64] and the major cause behind this is its simplicity and ease of understanding the rules regarding tree structures. A decision tree is constructed from a considered training dataset and each sample of the dataset is contained feature values and its class labels. Generally, DT is working like inductive inference. The major advantage of DT is, it can deal with noisy data and missing data in the dataset. It is also used for multiple stages and consecutive approaches during the classification procedure. During the first step of classification, the tree is generated, after that one by one data is applied to the classification process. Each node in the decision tree is represented as testing features belong to the training set and the generated branch from this node is the proper value of the feature. There are so many algorithms are designed in connection to DT, but some algorithms are more accepted with subject to different types of classification applications. Some of the most used algorithms in the literature are C4.5, ID3, and C5.
(3) K-nearest neighbor Classifier (KNN): It is one of the mature and simplest theoretical models incomparable to other machine learning classifiers [65]. The main important working style of KNN is to find a similar characteristic between the samples by measuring the distance. It is most acceptable in the case of multi-modal distribution data. It is difficult to decide the boundaries for the different classifiers when the same samples of a certain class are scattered modularly in the different locations of the feature vector. But it can manage with the KNN algorithm; it assigns a label to each input data, and this process is managed by computing the majority of vote of its k-nearest sample points [66]. The major measurement indicator of KNN is to compute the distance between objects located in the feature space, to measure the distance in general two mathematical formula used named the Euclidean distance and Manhattan distance.
(4) Random Forest (RF): This algorithm is proposed by Breimant and this algorithm is one of the popular classification techniques that use multiple tree structures for training the data and predict the samples [67]. Each tree requires randomly sampled data values and separate classifiers. The major difference between RF and other classification techniques is that the input is selected random manner using bootstrap selection methods. This whole method continues till the noisy and outlier samples are not desensitized and at last, the output is computed by voting approaches.
Performance Evaluation Metrics
The proposed study provides an in-depth analysis based on a comparative analysis of multiple different subjects with different session recordings obtained for sleep stage scoring analysis. For that reason, the authors have used multiple evaluation metrics to analyze the performance of the proposed sleep stage classification method.
This study considers four criteria such as the classification accuracy, recall, specificity, and confusion matrix. The confusion matrix is used to evaluate the results obtained from classification algorithms. The authors have analyzed the information about the actual and predict score achieved by the algorithms used. The classification accuracy is to provide information about the correctly classified decisions divided by the total number of cases [68].
The recall is a statistical measure that is also used to evaluate the classification performance by measuring the total number of correctly classified positive examples divided by the total number of positive cases [69].
Specificity is used to decide the performance of a classification test by calculating the number of correctly classified negative examples divided by the total number of negatives [19].
Precision analyses how many of the positively classified were relevant. To calculate the value of precision, we need to divide the correctly classified positive examples by the total number of predicted positives [70].
where: TP is the true positives, FN is the false negatives TN is the true negatives and FP is the false positives.
The F1-score is one of the statistical measures calculated with recall (sensitivity) and precision. Through F1-score, we can compute the harmonic mean, and which can be used to measure the rate of classification [71].
Experimental Results and Discussion
The proposed model focused on sleep scoring for the classification of the two-sleep stages based on dual-channel of EEG signal through machine learning techniques. The two different experiments were executed for two individual channels with sleep recordings of four subjects, who were affected with different types of sleep-related disorders. First of all, we applied preprocessing techniques for reducing the noises and some motion artifacts, which are appeared during the recordings of the signal. We have used the 10th order Butterworth bandpass filter technique for eliminating those irrelevant signal components for better analysis. Each subject having 750 epochs with 6000 sample points. The length of each epoch is 30 s. In this work, we have considered the sleep recordings of 750 epochs with 6000 sample points for each subject. After the data preparation, we have extracted the time domain, frequency domain, and non-linear features for analyzing the sleep behaviour of the subjects. Total of 34 features extracted. To find the suitable features, we employed feature screening techniques as ReliefF weight techniques. For both C4-A1 and O2-A1 channels, the correlation matrix is shown in Figs. 4 and 5. Finally, the selected features are forwarded to the different machine learning techniques such as support vector machine (SVM), k-nearest neighbor (KNN), decision tree (DT), and random forest (RF) for two-sleep states classification. In this study, we used some performance metrics such as classification accuracy, sensitivity, specificity, precision, and F1score for validating the proposed methodology suitability with subject to sleep staging analysis. The proposed method has been implemented using the MATLAB R2017a software for signal preprocessing, feature extraction, feature screening, and classification on a personal laptop with an Intel Core™ i3-4005U CPU 1.70 GHz, 2 core(s), 4 logical processors, 4 GB RAM, and Windows 10 operating system. The confusion matrix results for both channels C4-A1 and O2-A1 channels are presented in Tables 3 and 4 respectively. Similarly, the performance evaluation results for both the channels C4-A1 and O2-A1 are presented in Tables 5 and 6. The graph results of performance metrics are presented in Figs. 6 and 7.
Our analysis is completely documented guidelines of AASM standards, according to the documentation here we have also detect the sleep problem during different stages of sleep through sleep stages classification methods. The whole experiment was applied upon four subjects who were symptoms of sleep problems. To characterize the sleep behavior of each individual subject, we extracted as whole 34 features, through which we are analysis, the changes that happened during sleep in different time intervals and with different frequency ranges. As per our work layout of research work, in the final step, we have obtained supervised machine learning classification techniques like SVM, KNN, DT, and Random forest.
It has been observed from results that with the O2-A1 channel, we reported the best classification accuracy with help of random forest classification techniques with an average of 93.8% to differentiate between wake stages and sleep stages. It has been found that our proposed model achieved good classification accuracy to sleep staging. From this discussion, we concluded that instead of manual sleep stages analysis, automated sleep stages screening is more effective for diagnosis any sort of disorders that occurred due to sleep. The main effect of this proposed work is to consider both male and female subject sleep recordings and obtaining the feature selection techniques for screening the most suitable features for classification tasks. The other important part of this research work is considered four machine learning classifiers for sleep staging. Even though we reported excellent sleep staging accuracy for two-stage classification, but apart from that we have observed some limitations like class imbalance problem and mis-predicting the sleep stages in between N1 and REM sleep stages. It has been seen from the Table that the sleep epochs are quite imbalanced among the sleep stages, which sometimes may produce biased results. The other issue is mispredicting between the N1 and REM sleep stages because of their maximum similarity in their wave patterns, it also affects the sleep staging performances. These above-mentioned issues we will address in our future research work by obtaining the data augmentation techniques and deep learning techniques. To make analysis the performance of the proposed research work, we were made a comparison between our proposed research classification results with those of four other reported methods. Table 6 shows that the performance of the proposed research work is more accurate than other existing contribution work.
Performance Comparisons with the Existing Contributed Classification Sleep Stages Methods
The results achieved in the current research work are compared in between different contributed state-of-art literature, in which the obtained input channel is EEG, two stages classification, statistical features, datasets are addressed. Table 7 presents the performance comparison results of the proposed research methods and five similar existing reported contributions [53, 72,73,74,75,76,77,78] that work also used EEG recordings from the same datasets using a single channel of EEG.
Khalighi et al. [72] used the maximum overlap discrete wavelet transform and obtained both linear and non-linear properties and obtained an mRMR feature selection algorithm for screening the suitable features. The system reported an overall accuracy of 95% for the classification of the wake-sleep stages using SVM classification techniques.
Simoes et al. [73] used the R-square Pearson correlation coefficient and selected relevant features were applied into the Bayesian classifier and achieved an overall classification accuracy of 83%.
Khalighi et al. [74] used three categories of subjects records from the ISRUC-Sleep repository and extracted both temporal and spectral features extracted from the obtained input channel and applied SSM4S classification method, achieved overall classification accuracy with consideration of ISRUC-Sleep Subgroup-I, Subgroup-II, and Subgroup-III as 94.10, 92.40%, and 95.39%, respectively.
Sousa et al. [53] proposed a two-step classifier based on EEG signal, obtained an SVM classifier for distinguishing epochs suspected misclassification, and obtained both times, and frequency domain features and classified features that were forwarded into SVM classifier and reported classification accuracy as 86.75%.
Khalighi et al. [75] designed subject-independent improved automated sleep stage classification with application wake-sleep classification and classified through the SVM classifier and achieved 81.74% overall classification accuracy.
Tzimourta et al. [76] proposed a methodology for human sleep stage classification by considering brain EEG signals and extracted energy features, forwarded into random forest classifier, results with subject to classification accuracy reached 75.29%.
Najdi et al. [77] proposed a sleep study based on the two-layer stacked sparse auto-encoder and obtained frequency, time–frequency, time-domain features were extracted from EEG signals. The resulted classification accuracy was reported as 82.2%.
Finally, Kalbkhani [78] introduced Stockwell transform for signal decomposition and the decomposed features processed through SVM and KNN classifier. The average accuracy reported for SVM is 82.33% and for KNN as 81.00.
Conclusion
In this research work, we proposed an automated sleep staging system based on single-channel EEG signals using machine learning techniques. For a better analysis of the sleep characteristics, we used 10th order Butterworth bandpass filter for eliminating the contaminated signal compositions. One of the most important things of the proposed research work is obtaining the feature screening techniques for identifying the suitable features from the pool of extracted features, which alternatively helps to discriminate the changes in sleep characteristics. The main effectiveness of the proposed method was obtained by four different machine learning classifiers for classifying the sleep stages. It has been noticed from the results that the imbalance sleep stages epochs may create negative impacts on the sleep staging accuracy. In our future directions of research work, we will address the class imbalance issue. Besides we will also include more EEG data to evaluate the performance of the proposed method. We will also apply the deep learning techniques for proper discriminating features without using hand-engineered features. The performance of the proposed automated sleep staging system provides better sleep staging accuracy in comparison to the existing similar methods, which ultimately helpful for the sleep experts during analysis and classification of the sleep patterns.
Data availability statement
All EEG files are available from the ISRUC-SLEEP database (https://sleeptight.isr.uc.pt/ISRUC_Sleep/).
References
Panossian LA, Avidan AY. Review of sleep disorders. Med Clin N Am. 2009;93:407–25. https://doi.org/10.1016/j.mcna.2008.09.001.
Smaldone A, Honig JC, Byrne MW. Sleepless in America: inadequate sleep and relationships to health and well-being of our nation’s children. Pediatrics. 2007;119:29–37.
Hassan AR, Hassan Bhuiyan MI. Automatic sleep scoring using statistical features in the EMD domain and ensemble methods. Biocybern Biomed Eng. 2016. https://doi.org/10.1016/j.bbe.2015.11.001.
Aboalayon KA, Ocbagabir HT, Faezipour M (2014) Efficient sleep stage classification based on EEG signals. In: IEEE long island systems, applications and technology conference, LISAT 2014. https://doi.org/10.1109/LISAT.2014.6845193
Obayya M, AbouChadi F. Automatic classification of sleep stages using EEG records based on Fuzzy c-means (FCM) algorithm. In: Radio science conference (NRSC); 2014. pp. 265–272.
Alickovic E, Subasi A. Ensemble SVM method for automatic sleep stage classification. IEEE Trans Instrum Meas. 2018. https://doi.org/10.1109/TIM.2018.2799059.
Abeyratne UR, Swarnkar V, Rathnayake SI, Hukins C. Sleep-stage and event de- pendency of brain asynchrony as manifested through surface EEG. In: Proceedings of the 29th IEEE annual international conference of the engineering in medicine and biology society 2007, 709–712.
Rechtschaffen A, Kales AA. Manual of standardized terminology, techniques and scoring systems for sleep stages of human Subjects. Washington, DC: U.G.P. Office, Public Health Service; 1968.
Iber C, Ancoli-Israel S, Chesson AL, Quan SF. The AASM manual for the scoring of sleep and associated events: rules, terminology and technical specification. Darien: American Academy of Sleep Medicine; 2007.
Ll C, Zhao Y, Zhang J, Zou JZ. Automatic detection of alertness/drowsiness from physiological signals using wavelet-based nonlinear features and machine learning. Expert Syst Appl. 2015;42:7344–55.
da Silveira TL, Kozakevicius AJ, Rodrigues CR. Single-channel EEG sleep stage classification based on a streamlined set of statistical features in wavelet domain. Med Biol Eng Comput. 2016;55(2):1–10.
Amin HU, Malik AS, Kamel N, Hussain M. A novel approach based on data redundancy for feature extraction of EEG signals. Brain Topogr. 2016;29:207–17.
Diykh M, Li Y, Wen P. EEG sleep stages classification based on time domain features and structural graph similarity. IEEE Trans Neural Syst Rehabil Eng. 2016;24:1159–68.
Acharya UR, Sree SV, Swapna G, Martis RJ, Suri JS. Automated EEG analysis of epilepsy: a review. Knowl-Based Syst. 2013;45:147–65.
Aydın S, Tunga MA, Yetkin S. Mutual information analysis of sleep EEG in detecting psycho-physiological insomnia. J Med Syst. 2015;39:1–10.
Rahman MM, Bhuiyan MIH, Hassan AR. Sleep stage classification using single-channel EOG. Comput Biol Med. 2018;102:211–20.
Diykh M, Li Y, Wen P, Li T. Complex networks approach for depth of anesthesia assessment. Measurement. 2018;119:178–89.
Ebrahimi F, Mikaeili M, Estrada E, Nazeran H. Automatic sleep stage classification based on EEG signals by using neural networks and wavelet packet coefficients. In: 2008 30th annual international conference of the IEEE engineering in medicine and biology society. IEEE; 2008, pp. 1151–54.
Hsu YL, Yang YT, Wang JS, Hsu CY. Automatic sleep stage recurrent neural classifier using energy features of EEG signals. Neurocomputing. 2013;104:105–14.
Peker M. An efficient sleep scoring system based on EEG signal using complex-valued machine learning algorithms. Neurocomputing. 2016;207:165–77.
Bianchi MT. Sleep devices: wearables and nearables, informational and interventional. Consum Clin Metabol Clin Exp. 2017;84:99–108.
Mendez MO, Matteucci S, Cerutti et al. Sleep staging classification based on HRV: time-variant analysis. In: Int. Conf. IEEE Eng. Med. Biol. Soc.; 2009. pp. 9–12.
Radha M, Garcia-Molina G, Poel M, et al. Comparison of feature and classifier algorithms for online automatic sleep staging based on a single EEG signal. Eng Med Biol Soc. 2014;1:1876–80.
Stochholm A, Mikkelsen K, Kidmose P. Automatic sleep stage classification using ear-EEG. Eng Med Biol Soc 2016.
Chriskos P, Kaitalidou DS, Karakasis G, et al. Automatic sleep stage classification applying machine learning algorithms on EEG recordings. In: 2017 IEEE 30th international symposium on computer-based medical systems; 2017. pp. 435–439.
Imtiaz SA, Villegas ER. A low computational cost algorithm for REM sleep detection using single channel EEG. Ann Biomed Eng. 2014;42(11):2344–59.
Imtiaz SA, Jiang ZE, Villegas ER. An ultralow power system on chip for automatic sleep staging. IEEE J Solidst Circ. 2017;52(3):822–33.
Chen C, Liu X, Ugon A. Symbolic fusion: a novel decision support algorithm for sleep staging application. Mobihealth. 2015;16(8):19–22.
Sen B, Peker M, Cavusoglu A, et al. A comparative study on classification of sleep stage based on EEG signals using feature selection and classification algorithms. J Med Syst. 2014;38:18–687.
Liu H, Zhang H, Zhao P, et al. Study on sleep staging based on EEG signals. Chin J Biomed Eng. 2015;34(6):693–700.
Chriskos P, Kaitalidou DS, Karakasis G. et al. Automatic sleep stage classification applying machine learning algorithms on EEG recordings. In: 2017 IEEE 30th international symposium on computer-based medical systems; 2017, pp. 435–39.
Huang CS, Lin CL, Ko LW, et al. A hierarchical classification system for sleep stage scoring via forehead EEG signals. In: 2013 IEEE symposium on computational intelligence, cognitive algorithms, mind, and brain (CCMB), IEEE; 2013. pp. 1–5.
Zhu G, Yan L, Peng W. Analysis and classification of sleep stages based on difference visibility graphs from a single-channel EEG signal. IEEE J Biomed Health Inform. 2014;18(6):1813–21.
See AR, CK Liang. A study on sleep EEG using sample entropy and power spectrum analysis. In: Defense science research conference and expo (DSR), 2011, IEEE; 2011, pp. 1–4.
Yu S, Li P, Lin H. et al. Support vector machine based detection of drowsiness using minimum EEG features. In: International conference on social computing (SocialCom), 2013. IEEE; 2013. pp. 827–35.
Nguyen HD, Wilkins BA, Cheng Q, et al. An online sleep apnea detection method based on recurrence quantification analysis. IEEE J Biomed Health Inform. 2014;18(4):1285–93.
Bandarabadi M, Teixeira CA, Rasekhi J, et al. Epileptic seizure prediction using relative spectral power features. Clin Neurophysiol. 2015;126(2):237–48.
Liang SF, Kuo CE, Hu YH, et al. A rule-based automatic sleep staging method. J Neurosci Methods. 2012;205(1):169–76.
Phan H, Do Q, Vu DL. Metric learning for automatic sleep stage classification. In: 35th annual international conference of the IEEE engineering in medicine and biology society (EMBC), 2013, IEEE; 2013. pp. 5025–28.
Ronzhina M, Janoušek O, Kolárová J, et al. Sleep scoring using artificial neural networks. Sleep Med Rev. 2012;16(3):251–63.
Hassan AR, Bhuiyan MIH. Automated identification of sleep states from EEG signals by means of ensemble empirical mode decomposition and random under sampling boosting. Comput Methods Progr Biomed. 2017;140:201–10.
Hassan AR. A comparative study of various classifiers for automated sleep apnea screening based on single-lead electrocardiogram. In: 2015 international conference on electrical & electronic engineering (ICEEE). IEEE; 2015. pp. 45–8.
Hassan AR. Computer-aided obstructive sleep apnea detection using normal inverse Gaussian parameters and adaptive boosting. Biomed Signal Process Control. 2016;29:22–30.
Hassan AR. A comparative study of various classifiers for automated sleep apnea screening based on single-lead electrocardiogram. In: 2015 international conference on electrical & electronic engineering (ICEEE), IEEE; 2015. pp. 45–8.
Roebuck A, Monasterio V, Gederi E, Osipov M, Behar J, Malhotra A, Clifford GD. A review of signals used in sleep analysis. Physiol Meas. 2013;35(1):R1–57. https://doi.org/10.1088/0967-3334/35/1/r1.
Radha M, Garcia-Molina G, Poel M, Tononi G. Comparison of feature and classifier algorithms for online automatic sleep staging based on a single EEG signal. In: Proceedings of the 36th IEEE annual international conference of engineering in medicine and biology society, Chicago, IL, USA, 26–30 August; 2014. pp. 1876–80.
Savareh A, et al. Performance comparison of machine learning techniques in sleep scoring based on wavelet features and neighboring component analysis. PeerJ. 2018;6:e5247. https://doi.org/10.7717/peerj.5247.
Ronzhina M, Janoušek O, Kolářová J, Nováková M, Honzík P, Provazník I. Sleep scoring using artificial neural networks. Sleep Med Rev. 2012;16(3):251–63. https://doi.org/10.1016/j.smrv.2011.06.003.
Jo HG, Park JY, Lee CK, An SK, Yoo SK. Genetic fuzzy classifier for sleep stage identification. Comput Biol Med. 2010;40:629–34.
Fraiwan L, Lweesy K, Khasawneh N, Wenz H, Dickhaus H. Automated sleep stage identification system based on time-frequency analysis of a single EEG channel and random forest classifier. Comput Methods Programs Biomed. 2012;108(1):10–9. https://doi.org/10.1016/j.cmpb.2011.11.005.
Braun ET, Kozakevicius ADJ, Da Silveira TLT, Rodrigues CR, Baratto G. Sleep stages classification using spectral based statistical moments as features. Rev Inform Teór E Appl. 2018;25(1):11. https://doi.org/10.22456/2175-2745.74030.
Aboalayon KAI, Ocbagabir HT, Faezipour M. Efficient sleep stage classification based on EEG signals. In: IEEE long island systems, applications and technology (LISAT) conference 2014, Farmingdale, NY, USA, May 2014. pp. 1–6. https://doi.org/10.1109/LISAT.2014.6845193.
Sousa T, Cruz A, Khalighi S, Pires G, Nunes U. A two-step automatic sleep stage classification method with dubious range detection. Comput Biol Med. 2015;59:42–53.
Heyat MBB, Lai D, Zhang FIKY. Sleep bruxism detection using decision tree method by the combination of C4-P4 and C4-A1 channels of Scalp EEG’. IEEE Access; 2019. vol. 1–1.
Basha AJ, Balaji BS, Poornima S, Prathilothamai M, Venkatachalam K. Support vector machine and simple recurrent network based automatic sleep stage classification of fuzzy kernel. J Ambient Intell Humaniz Comput. 2020. https://doi.org/10.1007/s12652-020-02188-4.
Sharma M, Patel S, Choudhary S, Acharya UR. Automated detection of sleep stages using energy-localized orthogonal wavelet filter banks. Arab J Sci Eng. 2019. https://doi.org/10.1007/s13369-019-04197-8.
Tian P, Hu J, Qi J, Ye X, Che D, Ding Y, Peng YA. Hierarchical classification method for automatic sleep scoring using multiscale entropy features and proportion information of sleep architecture. Biocybern Biomed Eng. 2017;37(2):263–71. https://doi.org/10.1016/j.bbe.2017.01.005.
Alickovic E, Subasi A. Ensemble SVM method for automatic sleep stage classification. IEEE Trans Instrum Meas. 2018;67(6):1258–65. https://doi.org/10.1109/TIM.2018.2799059.
Khalighi S, Sousa T, Santos JM, Nunes U. ISRUC-Sleep: a comprehensive public dataset for sleep researchers. In: Computer methods and programs in biomedicine; 2016. vol. 124, pp. 180–192.
Kononenko I. Estimating attributes: analysis and extensions of RELIEF. ECML’94; 1994. pp. 171–182.
Cortes C, Vapnik V. Support vector networks. Mach Learn. 1995;20(3):273–97.
Kavzoglu T, Colkesen I. Classification of satellite images using decision trees: Kocaeli case. Electron J Map Technol. 2010;2(1):36–45.
Ocak H. A medical decision support system based on support vector machines and the genetic algorithm for the evaluation of fetal wellbeing. J Med Syst. 2013;37(2):1–9.
Ilhan HO, Bilgin G. Sleep stage classification via ensemble and conventional machine learning methods using single channel EEG signals. Int J Intell Syst Appl Eng. 2017;5(4):174–84.
Shang S, Li X, Zong M, et al. Learning k, forkNN classification. ACM Trans Intell Syst Technol. 2017;8(3):1–19.
Fraley C, Raftery AE. Model-based clustering, discriminant analysis, and density estimation. J Am Stat Assoc. 2002;97(458):611–31.
Shabani F, Kumar L, Solhjouy-Fard S. Variances in the projections, resulting from CLIMEX, boosted regression trees and random forests techniques. Theor Appl Climatol. 2017;97(458):1–14.
Sanders TH, McCurry M, Clements MA. Sleep stage classification with cross frequency coupling. In: 36th Annu. Int. Conf. IEEE Eng. Med. Biol. (EMBC); 2014, pp. 4579–82.
Bajaj V, Pachori R. Automatic classification of sleep stages based on the time-frequency image of EEG signals. Comput Methods Programs Biomed. 2013;112(3):320–8.
Zibrandtsen I, Kidmose P, Otto M, Ibsen J, Kjaer TW. Case comparison of sleep features from ear-EEG and scalp-EEG. Sleep Sci. 2016;9(2):69–72.
Berry RB, Brooks R, Gamaldo CE, Hardsim SM, Lloyd RM, Marcus CL, Vaughn BV. The AASM manual for the scoring of sleep and associated events: rules, terminology and technical specifications. Darien: American Academy of Sleep Medicine; 2014.
Khalighi S, Sousa T, Oliveira D, Pires G, Nunes U. Efficient feature selection for sleep staging based on maximal overlap discrete wavelet transform and SVM. In: Annual international conference of the IEEE engineering in medicine and biology society; 2011.
Simões H, Pires G, Nunes U, Silva V. Feature extraction and selection for automatic sleep staging using EEG. In: Proceedings of the 7th international conference on informatics in control, automation and robotics, vol. 3, pp. 128–33.
Khalighi S, Sousa T, Santos JM, Nunes U. ISRUC-Sleep: a comprehensive public dataset for sleep researchers. Comput Methods Programs Biomed. 2016;124:180–92.
Khalighi S, Sousa T, Pires G, Nunes U. Automatic sleep staging: a computer assisted approach for optimal combination of features and polysomnographic channels. Expert Syst Appl. 2013;40(17):7046–59.
Tzimourta KD, Tsilimbaris AK, Tzioukalia AT, Tzallas MG, Tsipouras LG. EEG-based automatic sleep stage classification. Biomed J Sci Tech Res. 2018;7(4):6032–6.
Najdi S, Gharbali AA, Fonseca JM. Feature Transformation based on stacked sparse auto encoders for sleep stage classification’. In: Technological innovation for smart systems, vol 499; 2017. p. 191–200
Kalbkhani H, Ghasemzadeh PG, Shayesteh M. Sleep stages classification from EEG signal based on stockwell transform. IET Signal Process. 2018;13:242–52.
Hassan AR, Subasi A. A decision support system for automated identification of sleep stages from single-channel EEG signals. Knowl Based Syst. 2017;128:115–24. https://doi.org/10.1016/j.knosys.2017.05.005.
Tripathy RK, Rajendra AU. Use of features from RR-time series and EEG signals for automated classification of sleep stages in deep neural network framework. Biocybern Biomed Eng. 2018. https://doi.org/10.1016/j.bbe.2018.05.005.
Funding
This study is not funded by any organization.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors in the paper have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the topical collection “Data Science and Communication” guest-edited by Kamesh Namudri, Naveen Chilamkurti, Sushma S J and S. Padmashree.
Rights and permissions
About this article
Cite this article
Satapathy, S.K., Loganathan, D. A Study of Human Sleep Stage Classification Based on Dual Channels of EEG Signal Using Machine Learning Techniques. SN COMPUT. SCI. 2, 157 (2021). https://doi.org/10.1007/s42979-021-00528-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42979-021-00528-5