Abstract
Nowadays, Isomap is one of the most popular nonlinear manifold dimension reductions which have applied to the real-world datasets. However, it has various limitations for the high-dimensional and large-scale dataset. Two main limitations of the Isomap are: it may make incorrect links in the neighborhood graph G and high computational cost. In this paper, we have introduced a novel framework, which we called the FastIsomap. The main purpose of the FastIsomap is to increase the accuracy of the graph by using two state-of-the-art algorithms: a randomized division tree and NN-Descent. The basic idea of FastIsomap is to construct an accurate approximated KNN graph from millions and hundreds of dimensional’ data points and then project the graph into low-dimensional space. We have compared the FastIsomap framework with the existing Isomap algorithm to verify its efficiency and performance, which provided accurate results of the high and large-dimensional datasets.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The high-dimensional data become computationally unmanageable when they have various dimensions and a large number of data points. In numerous research areas, such as machine learning, information visualization, data mining, computer vision [2, 24, 36], and the information visualization community [9, 16, 26], there is an urgent need to invent a low-dimensional data representation of large-scale and high-dimensional data.
The machine learning technique provides dimensionality reduction by the mean of minimal information loss. The critical challenge in the dimension reduction is to protect the structure of the data and perform the transformation with minimal data loss [32]. The dimensional reduction technique can be characterized by linear (supervised) and nonlinear mapping (unsupervised) methods. The linear (supervised) mapping methods are principal component analysis (PCA) [18], multidimensional scaling [36], independent component analysis (ICA), singular value decomposition (SVD), CUR matrix decomposition, compact matrix decomposition (CMD), and non-negative matrix factorization (NMF). The nonlinear (unsupervised) mapping methods are Isomap [35], FastMap, Locally Linear Embedding [30], Laplacian Eigenmaps [2], and Kernel PCA. Hinton and Maaten proposed the high-dimensional local and global structure method, which is known as the t-SNE [34].
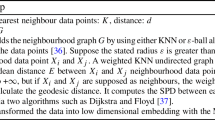
The great importance of studying of manifold learning for nonlinear dimension reduction has attracted much more attention in recent years [40, 43]. We have adopted a nonlinear Isometric feature mapping method (Isomap) [35], which is the most common technique for manifold learning. The key benefits of the Isomap are globally optimal, provable convergence guarantee, and appropriability for the nonlinear manifold. It has been used in many areas, such as image processing [40], robotics [29], computer vision graphics [33], signal processing [16], and pattern recognition [11, 42]. Isomap is viewed as a variant of metric multidimensional scaling (MDS) to model nonlinear data using its geodesic distance. MDS uses approximate geodesic distances between all pairs of data points, rather than Euclidean distances. Given the distance matrix D and data points N, Isomap produces the shortest representation of the eigenvalues of the matrix D. The geodesic distances between neighboring points are approximated by input-space distance. The geodesic distances are calculated as the shortest paths in a neighborhood graph for faraway points [28]. First, build the K nearest neighbor (KNN) graph of the manifold learning based on the distance between the pair of data points in the input space. Second, calculate the geodesic distance matrix between all pairs of data points using the shortest path distance graph algorithm. Further, find a low d-dimensional embedding by executing Eigendecomposition on the geodesic distance dense matrix [32].
In the literature, Choi and Choi [5] proposed the Robust Kernel Isomap method for a lack of topological stability, noises, and outliers problems. They reduced the effects of outliers based on topological structure from the network flow [1, 21]. Choi and Choi [3, 4] also proposed the algorithm for topological stability issues which is called Kernel Isomap. The Kernel Isomap can preserve topological stability while tackling the outliers and noises [21]. Moreover [14], they proposed two variant techniques of Isomap visualization, which are the multi-class multi-manifold-Isomap (MCMM-ISOMAP) and Isomap for classification (ISOMAP-C), respectively. The MCMM-ISOMAP and ISOMAP-C and various other techniques have been proposed to overcome the problem of the Isomap algorithm are that it is very slow.
In our motivation, we have focused on studying the Isomap algorithm problem. The core issue in the Isomap algorithm is lack of topological stability in the neighborhood graph G [1, 28, 37]. The main problem of the Isomap algorithm may make incorrect links in the neighborhood graph G. Isomap method and algorithm are still far from the accurate results and only used for global datasets. We follow the LargeVis method at a limited level in our work [34].
In this paper, we propose a new framework known as FastIsomap to handle the above problems. The framework relies on two components, randomized truncating (KD-tree) trees algorithm (tree building algorithm) and NN-Descent algorithm based on the KNN graph. Our framework uses an effective Randomized Truncated Trees (KD-tree) [7] algorithm to construct an approximated K nearest neighbor graph (KNN) at high accuracy, combine the subgraph data, and refine the graph data with the method known as NN-Descent method. The complete explanation of the techniques and algorithms is given in “Proposed Framework” section. Moreover, the FastIsomap method is much faster than the Isomap. No such type of method has been used in recent years from the Isomap perspective. We have proposed a new algorithm that uses both the local and global datasets.
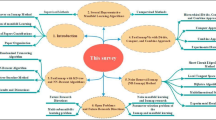
The rest of the paper is organized as follows. Second section provides an overview and background work, which is followed by a brief discussion on manifold learning and Isomap. Details of our proposed framework are discussed in third section, and description of the Randomized KD-tree and NN-Descent. Experiments, details, and results are presented in fourth section. Finally, the last section, the paper is concluded.
Related Work
Manifold Learning
Manifold learning is a technique for nonlinear mapping dimensionality reduction. The manifold learning provides the algorithms for the nonlinear mapping, which is based on the theory that the dimensionality of various information sets is only artificially high. The performance of the nonlinear methods is most reasonable for the global and local frameworks of high-dimensional datasets [35].
The main emphasis in dimensionality reduction is on reducing repetitions and basic pattern findings. It can also be used for data visualization, machine learning, and feature extraction. This technique reduces information dimensionality with minor data loss before proceeding with and executing the analysis. The key challenge in dimension reduction is to protect the structure of the data and to perform the transformation performed with minimal data loss. Moreover, a dimensional reduction technique can be characterized by a linear (supervised) or nonlinear mapping (unsupervised) method. Linear mapping (supervised) methods mean that the information lies in a linear combination subspace, and the actual data variables are exchanged by a smaller set of original data variables. Linear mapping methods can perform the transformation of high-dimensional data into low-dimensional data as a linear arrangement of actual variables. Nonlinear mapping methods can only be applied to originally nonlinear datasets, with high-dimensional and low-dimensional representations of the data points, obtained by calculating the distance between the actual data points [32]. The performance of linear mapping methods is mostly imperfect, as high-dimensional information mainly lies on or near a low-dimensional nonlinear manifold. Although nonlinear techniques, such as local tangent space alignment (LTSA) and Laplacian Eigenmaps (LE), have been proven to be empirically effective on small lab data, they have not been shown to achieve the goal of a global and local framework for high-dimensional data [24, 34].
However, the dimensionality reduction performance can be considerably reduced when using more complex and noisy input data. Because the computation of geodesic distance is sensitive to noisy data [19], which might damage the neighboring local shape or create disjointed edges. Therefore, it cannot be used for linear (classification) tasks or full class information of datasets with labeled data.
Isomap
Isomap is the nonlinear dimension reduction technique that maintains the geodesic distance and creates features during reshaping from high and large-dimensional to low-dimensional metric space. The Isomap is the variant of the metric multidimensional scaling (MDS) to model nonlinear data using its geodesic distance. It is MDS, which uses the geodesic distance between the two data points instead of straight-line Euclidean Distance E for data placed on a nonlinear manifold. The primary purpose of the Isomap is to find out an optimal subspace and maintain the geodesic distance between the data points [22]. The Isomap algorithm describes as follows:
\(\bullet\) Make the local K nearest neighborhood (KNN) graph for all data points
An Isomap method searches the K nearest neighbors of the data points on the manifold. The neighborhood graph data are represented as \(\epsilon\) and K in this section [25]. K is the number of nearest neighbors, and \(\epsilon\) is the max Euclidean search distances E. Construct the neighborhood graph G between the data point’s \(x_{i}\) and \(x_{j}\), if \(x_{i}\) is the \(\textit{K}\) nearest neighbor of \(x_{j}\) or if they are closer than particular distance \(\epsilon\).
\(\bullet\) Estimate the geodesic distance between all data points
After that searching the K nearest neighbors data points in the manifold learning. Isomap calculates the geodesic distance matrix between all pairs of data points \(x_{i}\) and \(x_{j}\) using the shortest path distance graph G and then computes the shortest path distance by using Dijkstra’s and Floyd’s algorithms [15] can be used.
\(\bullet\) Transform the lower-dimensional embedding
In the third step, MDS is applied to the resulting geodesic distance matrix to identify a low d-dimensional embedding by executing Eigendecomposition [6]. Some fundamental features of high-dimensional datasets like handwritten digits, hand gesture images, and face images are detected by Isomap [32].
Moreover, it has a few limitations. However, the geodesic distance is only appropriate for nonlinear datasets, and it can efficiently reproduce the topological structure of datasets. The performance of dimensionality reduction will be greatly reduced when the input data points are noisier and complex because the geodesic distance is sensitive to noise and can be destroyed by the local neighborhood structures [25].
Proposed Framework
In general, given large and high-dimensional datasets \({\mathcal {X}}=\{{x}\in R^d\}\), we aim to denote every dataset \(x_ {i}\)with a low-dimensional vector \(y_{i}\in R^d\), where the value of d is generally 2 or 3. Isomap algorithm possibly fails to transform from large and high-dimensional data into lower-dimensional space. However, our proposed method FastIsomap can efficiently solve this problem. We have used the Euclidean distance in the high-dimensional space instead of geodesic distances because KNN graphs require a metric of distances. Euclidean distance E is used to calculate the similarities of the data points through a square matrix, which provides much more efficient results than geodesic distances.
Our FastIsomap method relies on the two components: randomized truncating (KD-tree) trees algorithm (tree building algorithm) and NN-Descent algorithm based on the KNN graph. We have to use the KD-tree and NN-Descent algorithms for incorrect link problems. These algorithms consist of two steps. First, we have divided all the data points into small datasets and find the K nearest neighbor via a KD-tree algorithm. Also, we have to divide the graph into a subgraph. Second, we have combined the subgraphs data and purified graph data with techniques of the NN-Descent. Further, we have used the KD-tree searcher method for calculating the overall accuracy of our proposed algorithm. We have shown the general framework of our proposed method in Fig. 1.
FastIsomap Building Algorithm-I: Tree Building Randomized Truncating (KD-Tree)
One component of the FastIsomap is based on Randomized Truncating (KD-tree) Trees. The basic idea is to provide better initialization for KD-tree to improve the performance of Isomap significantly. The KD-tree algorithm divides a multidimensional dataset recursively and then constructs the shape of the tree that can be used for fast searching. A KD-tree divides datasets by vector space and makes a binary tree, allowing a logarithmic time complexity for KNN search. In the binary tree, data are split into two subgroups by the size at each level, in which the datasets have the highest variance. Jo et al. [17] proposed the idea of multiple randomized KD-trees for KNN search in higher-dimensional space. The basic idea of randomized KD-trees is that the data are divided by dimension recursively and then randomly picked from a small set of data sizes with the highest variance. Also, KD-tree uses the KD-tree searcher method to find out K nearest neighbor values. Moreover, Muja and Lowe [27] recognized two new algorithms known as randomized KD-trees and hierarchical K-means trees for KNN querying [17].
In this section, we have shown the details of Algorithm I on Building Randomized Truncating (KD-tree) Trees. We have used the KD-tree method for building tree in Algorithm I. In lines 1–2, KD-tree constructs the tree with node and point-set and picks the data d dimensions’ randomly. We have calculated the median over the datasets on data dimension d. Moreover, we have equally divided the tree into two halves, i.e., left-half and right-half, according to the median. After that, we have called the construct-tree function according to (node-left-child, left half) and (node-right-child, right-half) in lines 3–4. Further, in Algorithm I construct-tree function calls recursively and adds node according to the root-node and datasets D (line 5). At the end of Algorithm I, we have calculated the time and accuracy of the KD-tree algorithm by using a KD-tree searcher method (lines 6–8).
KD-tree searcher The KD-tree algorithm uses the KD-tree search model objects method. This method stores the nearest neighborhood search results. In the “Results and Discussion” section, we have included the distance metric, the training datasets, its parameters, and the maximal number of bucket sizes in each leaf node. The KD-tree algorithm splits N by K data points recursively and divides the N data points in K-dimensional space into a binary tree form. When we generated a KD-tree searcher model object, we also found out the stored tree.
Meanwhile, we searched all the nearest neighboring data points to the query data. The KD-tree searcher model objects method uses the Knnsearch and Rangesearch method to find out K nearest neighbor values. We use the Knnsearch method in our work [12] (Table 1).
FastIsomap Efficient Approximate KNN Graph Construction Algorithm-II: NN-Descent
We have used NN-Descent (neighbor exploring) techniques proposed by Dong et al. [8] to refine the resulting graph, which we obtained from the KD-tree method. The NN-Descent techniques are built on the idea that my neighbors’ ”neighbor is likely to be my neighbor.” We have used the randomized division (KD-tree) tree algorithm to construct an approximate KNN graph. The accuracy of the graph may not be so high. That is why we have used the NN-Descent algorithm for the higher accuracy of the graph. First, the execution of the NN-Descent algorithm is started from KNN (K nearest neighbor) graph. Then, NN-Descent continually improved the graph by exploring the neighbors of my neighbors’ corresponding to the specified current graph. We may replicate this iteration several times until the correctness of the graph is improved successfully. Therefore, we have found that the little iteration is enough to get the 100% accurate KNN graph.
In this section, we have shown the details of Algorithm II on an approximate KNN graph and construction of the refinement of KNN graph by the NN-Descent method. Algorithm II calls Algorithm I for refinement of Algorithm I through the NN-Descent Algorithm in line 1. We have initialization of the graph G by randomly creating the test list of KNN for a reverse neighbor of data-point U (line 3). After that, we have checked the different reverse pairs of neighbor U’s (v, w) and RNN (reverse nearest neighbor) list and then calculate the distances (v, w). Further, update the KNN distance list according to w and v (line 4). Moreover, we have calculated the time and accuracy of the refinement graph are the same as that Algorithm I (line 7) (Table 2).
Results and Discussion
We first evaluated our proposed framework to results on the publically available datasets using randomized division tree and NN-Descent algorithms. We analyze the effectiveness and efficiency of the FastIsomap by the general experiments on high-dimensional and large-scale datasets.
Datasets
This section describes the datasets that were used for randomized division tree and NN-Descent algorithms. The experiments were performed on two large-scale and high-dimensional datasets of social networks, Facebook and Twitter, which are available in Leskovec and Krevl [20]. Detailed information on the datasets is listed in Table 3.
Facebook Dataset
This dataset consists of friends circle lists. The Facebook dataset includes profiles node features and circles. This dataset was together from survey members using Facebook APP
Twitter Dataset
This dataset consists of friends circle lists. Twitter dataset includes profiles node features, and circles. Twitter dataset was collected from public sources.
Results and Experiments on KNN Graph Construction
We have presented the distinctive algorithms for K-Nearest-Neighbor (KNN) graphs whose constructions are given below:
-
Randomized division tree (KD-tree)
-
The neighbor exploring technique (NN-Descent) based on KNN graph
FastIsomap: Our new proposed framework has based on the above two algorithms.
KNN Classifier Method
We have used a classification technique for FastIsomap and Isomap. Well-known classification techniques such as KNN classifier [13, 23], SVM [38, 39], BP neural network [31] and decision tree [41]. The KNN classifier method uses the K-fold cross-validation method for calculating accuracy [10]. We used the well-established KNN classifier method for calculating the accuracy of our proposed “FastIsomap” method.
K-fold Cross-Validation-Method—the k-fold cross-validation method is used for the empirical selection method of the parameters. However, for every parameter, several values are verified through a K-fold cross-validation method, and the foremost one is selected [10]. We use the well-known K-fold cross-validation method for calculating the accuracy of both methods, i.e., approximated KNN graph construction algorithm (KD-tree) and refinement of KNN graph construction (NN-Descent). The K-fold cross-validation method randomly divides the accessible data into N partitions of equal-sized datasets. This process is executed N time, which gives N accuracy. For example, given a dataset with N points, and an approximate KNN graph approach should return the N groups of K data points. Each group of data points represents the nearest neighbor and searches the dataset within the respective data points. The accuracy P of the data points N and the number of correct classifications C are defined as:
Then, the accuracy of the output graph is defined as given refine dataset with N points, and an approximate KNN graph approach should output the refinement of N groups of K data points. Each group of data points represents the nearest neighbor and refines the dataset within the respective data point. We used Eq. 1 for the accuracy of the refined graph with the NN-Descent technique.
Tables 4 and 5 show the calculated accuracy of Facebook and Twitter datasets from Eq. (1).
Experiments and Results on KD-Tree and NN-Descent
Although Tables 4 and 5 results are based on the K-fold cross-validation method, Tables 4 and 5 demonstrate the accuracy and time to build the KNN graph for two datasets, such as Facebook and Twitter. To make the graph-based KD-tree and NN-Descent methods very useful, now we require examining how to make the KD-tree graph and refinement of KNN graph construction efficient. Therefore, we have computed the accuracy of the KD-tree and NN-Descent graph with Eq. (1) and compare the performance of the two datasets (Facebook and Twitter) with the refinement of KNN graph construction methods.
Results on KD-Tree of KNN Graph Construction
Figures 2 and 3 show the performance of the KNN graph w.r.t accuracy and time cost. We used the value of \(K=1\) to 5 for calculating the accuracy versus time in the Facebook dataset, which consists of 4039 data points. The computation time for calculating 99% accurate results of Facebook datasets was only 15 minutes. On the Twitter dataset, the value of K is also 1 to 5, which contained the 81,306 data points; it is very difficult to build the KNN graph at 99% the accurate result through the KD-trees technique because Twitter datasets are very large and high. However, the overall performance of the KD-tee algorithm constantly attains the highest accurate results of the dataset at the shortest computation time.
Results on Refinement (NN-Descent) of KNN Graph Construction
Figures 4 and 5 show the performance of the KNN graph w.r.t accuracy and time cost KNN graph. We calculated the highly accurate results of the social network’s datasets from the NN-Descent algorithm. The NN-Descent algorithm first initializes the graph randomly and then refines the graph iteratively with the use of two techniques: sampling and local join [8]. The local join method uses the brute-force searching algorithm for the nearest neighbors. The sampling method is used to verify the small number of points that are involved with the local join. Therefore, the performance of the algorithm is effective. For the Twitter dataset, it is very difficult to build the KNN graph for 100% accurate result through NN-Descent algorithm, because Twitter datasets are so large and higher than Facebook datasets. However, the overall performance of the refinement of the KNN graph construction versus time cost constantly attains the shortest time cost at the highest (100%) accurate results of the datasets. Our proposed FastIsomapVis method for KNN graph structure has produced very effective results for scaling to millions and hundreds of dimensions data points. The FastIsomapVis has conducted only one iteration for the NN-Descent algorithm.
Comparison of Accuracy in the Graph Between the FastIsomap and Isomap
Table 6 demonstrates the overall comparison accuracy of FastIsomap method and Isomap with Facebook and Twitter datasets.
Figure 6 compares the overall accuracy performance of the FastIsomap and Isomap with Facebook and Twitter datasets. We calculated the highly accurate results of the social network’s datasets from the FastIsomap method. The FastIsomap method has provided 100% accuracy as compared to Isomap. The Isomap algorithm is very slow and time-consuming for high-dimensional and large-scale datasets. On the Twitter dataset, it is very difficult to build the KNN graph 72–42% accurate result through the Isomap algorithm. Our proposed FastIsomap method for KNN graph construction has produced very effective results at the scales to millions of datasets with hundreds of dimensions’ data points.
Conclusions
In this paper, we introduced a novel framework, which is called FastIsomap. FastIsomap straightforwardly measures millions of datasets with hundreds of dimensions. We proposed a very efficient and effective algorithm for the construction of an accurately approximated KNN graph and then projected the graph into the low-dimensional space 2D or 3D. Experimental results on two real-world datasets show that the FastIsomap outperforms Isomap for KNN graph construction for both datasets, in terms of both effectiveness and efficiency. In our future work, we design advanced techniques of a hierarchical method for the FastIsomap.
Change history
28 September 2023
A Correction to this paper has been published: https://doi.org/10.1007/s42979-023-02168-3
References
Balasubramanian M, Schwartz EL. The isomap algorithm and topological stability. Science. 2002;295(5552):7.
Belkin M, Niyogi P. Laplacian eigenmaps and spectral techniques for embedding and clustering. In: Advances in neural information processing systems, 2002; p. 585–91.
Choi H, Choi S. Kernel isomap. Electron Lett. 2004;40:1612–3.
Choi H, Choi S. Kernel isomap on noisy manifold. In: Proceedings of the 4th international conference on development and learning, 2005, IEEE, 2005; p. 208–13.
Choi H, Choi S. Robust kernel isomap. Pattern Recogn. 2007;40(3):853–62.
Dadkhahi H, Duarte MF, Marlin B. Isomap out-of-sample extension for noisy time series data. In: 2015 IEEE 25th international workshop on machine learning for signal processing (MLSP), IEEE, 2015; p. 1–6.
Dasgupta S, Freund Y. Random projection trees and low dimensional manifolds. In STOC, Citeseer. 2008; vol. 8, p. 537–46.
Dong W, Moses C, Li K. Efficient k-nearest neighbor graph construction for generic similarity measures. In: Proceedings of the 20th international conference on World wide web, ACM, 2011; p. 577–86.
Fruchterman TM, Reingold EM. Graph drawing by force-directed placement. Softw Pract Exp. 1991;21(11):1129–64.
Geng X, Zhan DC, Zhou ZH. Supervised nonlinear dimensionality reduction for visualization and classification. IEEE Trans Syst Man Cybern Part B (Cybern). 2005;35(6):1098–107.
Gepshtein S, Keller Y. Sensor network localization by augmented dual embedding. IEEE Trans Signal Process. 2015;63(9):2420–31.
Gulraj M, Ahmad N. Mood detection of psychological and mentally disturbed patients using machine learning techniques. IJCSNS. 2016;16(8):63.
Ho TK. Nearest neighbors in random subspaces. In: Joint IAPR international workshops on statistical techniques in pattern recognition (SPR) and structural and syntactic pattern recognition (SSPR), Springer, 1998; p. 640–8.
Hong-Yuan W, Xiu-Jie D, Qi-Cai C, Fu-Hua C. An improved isomap for visualization and classification of multiple manifolds. In: International conference on neural information processing, Springer, 2013; p. 1–12.
Hougardy S. The floyd-warshall algorithm on graphs with negative cycles. Inf Process Lett. 2010;110(8–9):279–81.
Jacomy M, Venturini T, Heymann S, Bastian M. Forceatlas2, a continuous graph layout algorithm for handy network visualization designed for the gephi software. PLoS ONE. 2014;9(6):e98679.
Jo J, Seo J, Fekete JD. A progressive KD tree for approximate k-nearest neighbors. In: 2017 IEEE workshop on data systems for interactive analysis (DSIA), IEEE, 2017; p. 1–5.
Jolliffe I. Principal component analysis. Berlin: Springer; 2011.
Lee JA, Verleysen M. Nonlinear dimensionality reduction of data manifolds with essential loops. Neurocomputing. 2005;67:29–53.
Leskovec J, Krevl A. Snap datasets: Stanford large network dataset collection (2014). http://snap.stanford.edu/data. 2016; p. 49
Li B, Huang DS, Wang C. Improving the robustness of isomap by de-noising. In: 2008 IEEE international joint conference on neural networks (IEEE world congress on computational intelligence), IEEE, 2008; p. 266–270.
Li X, Cai C, He J. Density-based multi-manifold isomap for data classification. In: 2017 Asia-Pacific signal and information processing association annual summit and conference (APSIPA ASC), IEEE, 2017; p. 897–903.
Lowe DG. Similarity metric learning for a variable-kernel classifier. Neural Comput. 1995;7(1):72–85.
Lvd Maaten, Hinton G. Visualizing data using t-SNE. J Mach Learn Res. 2008;9(Nov):2579–605.
Maier M, Luxburg UV, Hein M. Influence of graph construction on graph-based clustering measures. In: Advances in neural information processing systems, 2009; p. 1025–1032.
Martin S, Brown WM, Klavans R, Boyack KW. Openord: an open-source toolbox for large graph layout. In: Visualization and data analysis 2011, international society for optics and photonics, 2011; p. 786806.
Muja M, Lowe DG. Fast approximate nearest neighbors with automatic algorithm configuration. VISAPP (1). 2009;2(331–340):2.
Qu T, Cai Z. An improved isomap method for manifold learning. Int J Intell Comput Cybern. 2017;10(1):30–40.
Ramos FT, Kumar S, Upcroft B, Durrant-Whyte H. A natural feature representation for unstructured environments. IEEE Trans Robot. 2008;24(6):1329–40.
Roweis ST, Saul LK. Nonlinear dimensionality reduction by locally linear embedding. Science. 2000;290(5500):2323–6.
Rumelhart DE, Hinton GE, Williams RJ. Learning representations by back-propagating errors. Nature. 1986;323(6088):533–6.
Sumithra V, Surendran S. A review of various linear and non linear dimensionality reduction techniques. Int J Comput Sci Inf Technol. 2015;6:2354–60.
Takahashi S, Fujishiro I, Okada M. Applying manifold learning to plotting approximate contour trees. IEEE Trans Vis Comput Graphics. 2009;15(6):1185–92.
Tang J, Liu J, Zhang M, Mei Q. Visualizing large-scale and high-dimensional data. In: Proceedings of the 25th international conference on world wide web, international world wide web conferences steering committee, 2016; p. 287–297
Tenenbaum JB, De Silva V, Langford JC. A global geometric framework for nonlinear dimensionality reduction. Science. 2000;290(5500):2319–23.
Torgerson WS. Multidimensional scaling: I. Theory and method. Psychometrika. 1952;17(4):401–19.
Van Der Maaten L, Postma E, Van den Herik J. Dimensionality reduction: a comparative. J Mach Learn Res. 2009;10(66–71):13.
Vapnik V. The nature of statistical learning theory. Berlin: Springer; 2013.
Vapnik V, Vapnik V. Statistical learning theory. New york: Wiley; 1998.
Verma R, Khurd P, Davatzikos C. On analyzing diffusion tensor images by identifying manifold structure using isomaps. IEEE Trans Med Imaging. 2007;26(6):772–8.
Witten IH, Frank E, Hall MA, Pal CJ. Data mining: practical machine learning tools and techniques. Burlington: Morgan Kaufmann; 2016.
Yazdian N, Tie Y, Venetsanopoulos A, Guan L. Automatic ontario license plate recognition using local normalization and intelligent character classification. In: 2014 IEEE 27th Canadian conference on electrical and computer engineering (CCECE), IEEE, 2014; p. 1–6
Zhang B. Multiple features facial image retrieval by spectral regression and fuzzy aggregation approach. Int J Intell Comput Cybern. 2011;4(4):420–41.
Funding
The author(s) received no specific funding grant for this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yousaf, M., Rehman, T.U. & Jing, L. An Extended Isomap Approach for Nonlinear Dimension Reduction. SN COMPUT. SCI. 1, 160 (2020). https://doi.org/10.1007/s42979-020-00179-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42979-020-00179-y