Abstract
Chuanmai104 (CM104), an elite wheat (Triticum aestivum L.) variety that currently produces the highest yield per unit area in southwestern China, plays a critical role in wheat production. The high quality and stability of grain traits are important factors that ensure the high, stable yields of CM104 in different production areas. In this study, six grain traits of CM104 sampled from 19 environments in five provinces of China during 2018–2022 were evaluated. The traits comprised thousand-kernel weight, grain length, grain width, grain length–width ratio, grain circumference, and grain surface area. Fifteen quantitative trait loci (QTLs) associated with the grain traits were identified based on a recombinant inbred lines (F9–10) population derived from the cross between CM104 and the landrace Baimaomai (BMM), nine and six QTLs derived from CM104 and BMM, respectively. Three mainly pleiotropic QTLs derived from CM104, namely QTL10 (grain circumference, grain surface area), QTL11 (grain length, grain circumference), and QTL12 (grain length, grain circumference), were expressed significantly and stably in multiple environments, and explained 3.34–5.06%, 5.32–6.50%, and 6.00–12.13% of the phenotypic variation, respectively. The pyramiding of multiple genes is hypothesized to have contributed to the stability of the CM104 grain traits in different environments. The results provide a basis for future improvement of yield and its stability in wheat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat is an economically important crop that is a major energy and nutrient source for humans. Against the background of a growing global population and climate change, human food demands are increasing rapidly. Wheat breeders are currently focused on developing varieties with high and stable yields to meet the increase in food demand (Eltaher et al. 2021). However, it is difficult to increase wheat yields quickly, which jeopardizes future food security (Li et al. 2016).
Previous studies have indicated that wheat yield is mainly determined by three traits: spike number per area, kernels per spike, and thousand-kernel weight (TKW) (Yang et al. 2016). Thousand-kernel weight is mainly controlled by grain morphology and grain-filling degree (Okamoto et al. 2013). Generally, grain morphology mainly affects TKW through grain length (GL), grain width (GW), grain length–width ratio (GR), grain circumference (GC), and grain surface area (GSA), and ultimately impacts on wheat yield (Wang et al. 2012; Gao et al. 2021; Cristina et al. 2016; Liu et al. 2017). Therefore, research on grain morphology has important applications for improvement of the grain weight in wheat. Identification of the key genes associated with grain morphology is crucial for further improvement of wheat yield. Many quantitative trait loci (QTLs) that contribute to grain morphology or grain weight traits have been identified in durum wheat (Varshney et al. 2000) and in hexaploid wheat distributed on all 21 chromosomes Huang et al. 2003 and 2006; Li et al. 2022; Williams et al., 2014; Okamoto et al. 2013; Patil et al. 2013; Simmonds et al. 2014; Zhang et al. 2015; Kumar et al. 2019; Cabral et al. 2018; Liu et al. 2017). Some of the genes that influence grain morphology have been cloned. For instance, TaGW2 negatively regulates wheat GW and TKW (Su et al. 2011; Wang et al. 2018). The gene DA1-RELATED 1 (TaDA1) is a negative regulator of grain size and can interact with TaGW2 to affect grain weight in wheat (Liu et al. 2020). TaPGS1 is a transcription factor that regulates wheat grain size and grain weight. Overexpression of TaPGS1 significantly increases grain weight, GL, and GW. Further investigation showed that TaPGS1 binding to the FI3 promoter to knockdown FI3 expression significantly reduces grain weight, GL, and GW (Guo et al. 2022). In addition, many genes, such as TaCKX, TaGS5, TaAGP-L, TaSDIR1, TaCYP78A3, TaGS3, TaGW7, and TaGW8, have been confirmed to be involved in the regulation of wheat grain morphology or grain weight (Jabłoński et al. 2020; Wang et al. 2015, 2020; Zhang et al. 2019; Ma et al. 2015, 2017; Zhang et al. 2019; Yan et al. 2019).
Chuanmai104 (CM104), an elite synthetic hexaploid wheat-derived variety bred by the Crop Research Institute, Sichuan Academy of Agricultural Sciences, exhibits numerous desirable traits, including high yield, good grain quality, broad adaptability, and resistance to stripe rust, powdery mildew, and pre-harvest sprouting under the unique ecological conditions of Sichuan Province in China (Li et al. 2014). From 2010 to 2020, CM104 consistently maintained the highest yield increase recorded in the southwestern China wheat production region, and set a record yield of 10,947 kg per hectare for southwestern China in 2020. Recently, CM104 has been widely used as an elite breeding material in the wheat breeding program in southwestern China, and a total of 21 varieties are derived from CM104 in this growing region (Liu et al. 2021). However, the genetic mechanism for the high and stable yield of CM104 remains uncertain.
In this study, CM104 grains were sampled in different years and from different regions to analyze the genetic basis for the high yield and its stability. The grain morphology and grain weight phenotypes were recorded for a population of 251 recombination inbred lines (RILs; F9–10) derived from the cross of CM104 and the wheat landrace Baimaomai (BMM) grown in three environments. The results provide a foundation for analysis of the genetic mechanism for the high yield and stability of CM104, and provide valuable information for pyramiding of multiple genes associated with grain traits for future improvement of wheat varieties.
Materials and methods
Plant materials and mapping population
Nineteen samples of CM104 grains were collected from five provinces in China during 2018–2022, and three BMM samples were collected from Sichuan in 2021–2022 (Fig. 1, Table 1). Recombination inbred lines were derived from the cross between CM104 and BMM (Fig. 2), comprising 251 RILs-F9 (2021) and RILs-F10 (2022) lines. The RILs were grown at Guanghan in 2021 (2021GH) and 2022 (2022GH), and at Shifang in 2022 (2022SF). Approximately 30 seeds of each line were uniformly sown in a single 1-m-long row with 25 cm spacing between rows. Standard field management practices were applied.
Evaluation of grain traits and statistical analysis
Thousand-kernel weight, GL, GW, grain length–width ratio (GR), GC, and GSA of each accession from different environments were recorded using an automatic seed test analyzer (Mini1600, Jie Lai Mei Technology Co., Ltd., Chengdu, China). Regression analysis of the CM104 grain traits was performed with Excel software (Microsoft, Redmond, WA, USA). The correlations among the grain traits were analyzed using the two-tailed Pearson method implemented in IBM SPSS Statistics 25 (IBM Corporation, Armonk, NY, USA). Analysis of variance (ANOVA) was conducted, and then, the accessions were ranked with Duncan’s test and plotted using GraphPad Prism V8.0.2.263. R software (i386 4.1.0) and the lme4 package were used to calculate the broad-sense heritability (H2; Smith et al. 1998) of the wheat grain traits. High-quality images of wheat grains were captured with a Leica M205 FA stereomicroscope (Leica, Germany).
QTL mapping
Genotyping of the CM104 × BMM RILs was performed using the wheat 50 K SNP (single nucleotide polymorphism) genotyping array developed by CapitalBio Corporation (Beijing, China) and synthesized by Affymetrix (California, America). In a previous study, we constructed genetic linkage maps with 3779 polymorphic SNP markers (Liu et al. 2017). The QTL analysis was conducted with inclusive composite interval mapping using GAHP software (Zhang et al. 2022). The QTLs were mapped at a logarithm of odds threshold of 4 based on 1000 permutations and a walking speed of 1.0 cM, with P = 0.001 in a stepwise regression. The QTL effects were estimated as the proportion of phenotypic variance explained by the QTL. The QTLs for a trait with identical, overlapping, or adjacent marker intervals in a linkage group were treated as identical and given a common name. Only QTLs identified in two or three environments were considered. The physical distance between the marker interval of a QTL was analyzed using the International Wheat Genome Sequencing Consortium (http://www.wheatgenome.org/) and EnsemblPlants (http://plants.ensembl.org/info/website/ftp/index.html) databases based on the flanking sequences.
Results
Phenotypic variation and diversity analysis
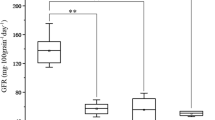
Nineteen samples of CM104 grains from different environments in China were collected to evaluate phenotypic variation in grain morphology and grain weight (Table 2). Of these samples, 14 samples were harvested in Sichuan Province during 2018–2022 (SC group) and five samples were harvested outside Sichuan Province in 2022 (SW group) (Fig. 3). Differences in the means for the grain morphology and grain weight traits between the SC and SW groups were analyzed by performing an ANOVA. The TKW, GW, GC, and GSA traits in the SC group were significantly higher than those in the SW group (P < 0.05; Fig. 3a), suggesting that Sichuan is the most suitable production zone for CM104. Furthermore, ANOVA and a regression analysis were used to evaluate the stability of the grain morphology and grain weight traits. Two CM104 samples (2018CZ and 2021GH) were compared with 12 CM104 samples collected in 2022 from different environments in Sichuan Province, but no significant differences were observed between the two groups (Fig. 3b).
Regression analysis showed that the grain morphology and grain weight traits of CM104 grown in Sichuan Province were highly stable. A highly significant correlation was observed between grain morphology and grain weight traits for all 14 environments (R2 = 0.88–0.95; Fig. 4).
QTL mapping of grain traits
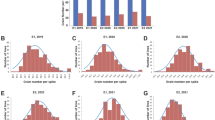
Among the three environments for the RILs, significant differences in TKW, GL, GC, and GSA between CM104 and BMM were observed (P < 0.05; Fig. 5a). The frequency distributions of the investigated traits revealed continuous variation in the RILs population, suggesting that the phenotypic data for the four grain traits were normally distributed and these traits were controlled by multiple QTLs (Fig. 5b).
The estimated H2 values for TKW, GL, GC, and GSA were greater than 0.5 (range 0.52–0.84) (Table 3). Thus, it was assumed that the traits were influenced by genetic factors. Pearson correlation coefficients for the four grain morphologies and grain weight traits in the three environments ranged from 0.58 to 0.96 (Fig. 6).
Pearson’s correlation coefficients (r) between thousand-kernel weight (TKW), grain length (GL), grain width (GW), grain length–width ratio (GR), grain circumference (GC), and grain surface area (GSA). a Grain phenotypic correlation analysis of CM104 was performed in nineteen environments; b grain phenotypic correlation analysis of CM104 × BMM RILs population in three environments
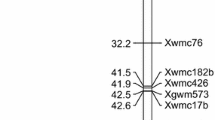
In the three environments, 15 QTLs were identified for GC, GL, GSA, and TKW. Three detected QTLs for TKW explained 6.08–6.84% of the phenotypic variation, 11 QTLs for grain morphology explained 3.35–12.13%, and one QTL for GSA and TKW explained 4.33–5.35%. Three stable putative QTLs (QTL10, QTL11, and QTL12) for grain morphology associated with GC, GL, and GSA were detected in this study. These QTLs were located on chromosomes 5B and 5D, and explained 3.35–12.13% of the phenotypic variation (Table 4, Fig. 7). The additive effect of the individual QTLs indicated that CM104-positive alleles contributed to the three traits.
Discussion
Grain traits of CM104 showed high stability in different environments
Thousand-kernel weight is positively correlated with grain morphology and is an important trait owing to its phenotypic stability and high heritability (Kuchel et al. 2007). Selection for a higher TKW has been conducted during domestication, and therefore, it has been argued that the grain morphology of cereals is a component of the domestication syndrome (Brown et al. 2009). Larger grains generally favor TKW, seedling vigor, and flour yield characteristics (Chastain et al. 1995). Stable wheat production is crucial to global food security (Macholdt and Honermeier 2017). Many factors, such as the phenotypic stability of wheat varieties and genotype × environment interactions, affect wheat yield to different extents (Nielsen and Vigil 2018). Regression analysis and ANOVA are widely used to evaluate the relationship between wheat yield and environment (Chen et al. 2018; Pepó and Győri 2005; Ayalneh et al. 2014; van Frank et al. 2020; Ji et al. 2023). In the present study, CM104 samples were collected from different environments; grain traits in the SC group were significantly higher than those in the SW group. The highest TKW was recorded in the 2022_JY sample (59.62 g/1000 grains), which indicated that JY was the most suitable production location for CM104. As the highest yielding wheat variety grown in Sichuan, CM104 exhibits broad adaptability and stable high yield. However, its adaptability is generally poor outside of Sichuan Province, with the exception of the 2022_XX sample (51.14 g/1000 grains) in Henan Province, which suggested that CM104 can maintain favorable grain traits in specific environments. Correlation analysis showed that the TKW of CM104 was positively correlated with GL, GW, GC, and GSA, but was negatively correlated with GR (Fig. 6). This result indicated that modern breeders prefer rounded seeds, consistent with previous findings (Cheng et al. 2020; Zhang et al. 2022).
Stable QTLs associated with grain morphology and grain weight traits
Grain morphology and grain weight are complex quantitative traits, and are influenced by many main and micro-effect QTLs (Slafer et al. 2014). In previous studies, more than 100 QTLs for TKW, GL, GW, GC, and GSA in wheat have been identified and are distributed on all 21 chromosomes. In addition, wheat TKW is significantly positively correlated with grain morphology, and many QTLs associated with grain morphology have been co-localized with TKW QTLs (Börner et al., 2002; Huang et al. 2003; Huang et al. 2006; Li et al. 2022; Williams et al., 2014; Okamoto et al. 2013; Simmonds et al. 2014; Zhang et al. 2015; Kumar et al. 2019; Cabral et al. 2018; Liu et al. 2017). However, many QTLs for grain morphology are not detectable in different environments and explain only a small proportion of the phenotypic variation for grain weight. Taken together, these factors mean that such QTLs are unsuitable for use in marker-assisted selection for wheat breeding. However, such QTLs are strongly affected by the environment and not all major QTLs are repeatedly detected in all environments (Akram et al., 2020; Li et al. 2019; Suliman et al. 2021; Alemu et al. 2020; White et al. 2022; Gao et al. 2021). Therefore, molecular marker-assisted breeding for wheat grain traits is challenging. The present RIL populations were used for QTL analysis in multiple environments, which is a beneficial approach for breeding to select loci that can be detected in different environments. The present study detected 15 QTLs for grain morphology. With reference to previous studies, four QTLs for TKW identified in previous reports were detected on chromosomes 1A (1), 2A (2), and 2D (1). The co-localized GL and TKW locus QTL1 (537.54–544.74 Mb) is located at a similar position to that of the TKW QTL on chromosome 1A reported by Yang et al. (2020), and three QTLs located on chromosomes 2A and 2D have been reported in multiple studies. Interestingly, all of the favorable alleles associated with TKW were inherited from CM104 (Table 4). Among the 15 QTLs associated with grain morphology and grain weight identified in the present study, 14 QTLs have been located in previous studies except for QTL15. However, the physical location of QTL15 is near that of QTL14, and thus, they may represent a single QTL (Akram et al. 2021; Li et al. 2019; Suliman et al. 2021; Alemu et al. 2020; White et al. 2022; Gao et al. 2021; Okamoto et al. 2013; Tyagi et al. 2015; Gahlaut et al. 2021; Yang et al. 2020; Chidzanga et al. 2022; Sun et al. 2009; Azadi et al. 2015). In the current study, three QTLs associated with grain morphology were identified in multiple environments on chromosomes 5B and 5D, namely QTL10, QTL11, and QTL12, and their favorable alleles were all derived from CM104. Among these three stable QTLs, QTL10 and QTL11 explained 3.34–5.06% and 5.32–6.50% of the phenotypic variation, respectively, whereas QTL12 explained 6.00–12.13% of the phenotypic variation, which was significantly higher than that for QTL10 and QTL11. These results indicated that QTL12 was an important QTL for the grain morphology of CM104 (Table 4). These stable QTLs (QTL10, QTL11, and QTL12) detected in multiple environments provide a foundation for fine-mapping of TKW and grain morphology-related genes and improvement of the yield traits of CM104 by molecular marker-assisted selection. Additionally, the major and minor QTLs presented the excellent genetic basis to ensure the stable phenotype of grain traits of CM104 in multiple environments. It was also an effective strategy to cultivate high and stable yield wheat varieties with widely adaptation by pyramiding major and minor QTLs and utilizing the synergistic effect of major and minor QTLs.
Conclusion
The high-yielding and phenotypically stable wheat variety CM104 harbored favorable alleles for TKW and grain morphology. Eight previously reported QTLs were identified in CM104, but only three stable QTLs (QTL10, QTL11, and QTL12) were detected in multiple environments. Therefore, these three QTLs may be the genetic basis for the phenotypic stability of the TKW and grain morphology of CM104 in multiple environments. Pyramiding major and minor QTLs will be also useful to improve wheat varieties and contribute to the yield stability in multiple environments in future.
References
Akram S, Arif MAR, Hameed A (2021) A GBS-based GWAS analysis of adaptability and yield traits in bread wheat (Triticum aestivum L.). J Appl Genet 62(1):27–41
Alemu A, Feyissa T, Tuberosa R, Maccaferri M, Sciara G, Letta T, Abeyo B (2020) Genome-wide association mapping for grain shape and color traits in Ethiopian durum wheat (Triticum turgidum ssp. durum). Crop J 8(5):757–768
Ayalneh T, Letta T, Abinasa M (2014) Assessment of stability, adaptability and yield performance of bread wheat (Triticum aestivum L.) cultivars in south estern Ethiopia. Plant Breed Seed Sci 67(1):3
Azadi A, Mardi M, Hervan EM, Mohammadi SA, Moradi F, Tabatabaee MT, Pirseyedi SM, Ebrahimi M, Fayaz F, Kazemi M, Ashkani S, Nakhoda B, Mohammadi-Nejad G (2015) QTL mapping of yield and yield components under normal and salt-stress conditions in bread wheat (Triticum aestivum L.). Plant Mol Biol Rep 33:102–120
Brown TA, Jones MK, Powell W, Allaby RG (2009) The complex origins of domesticated crops in the Fertile Crescent. Trends Ecol Evol 24(2):103–109
Cabral AL, Jordan MC, Larson G, Somers DJ, Humphreys DG, McCartney CA (2018) Relationship between QTL for grain shape, grain weight, test weight, milling yield, and plant height in the spring wheat cross RL4452/’AC Domain’. PLoS ONE 13(1):e0190681
Chastain TG, Ward KJ, Wysocki DJ (1995) Stand establishment response of soft white winter wheat to seedbed residue and seed size. Crop Sci 35(1):213–218
Chen H, Deng A, Zhang W, Li W, Qiao Y, Yang T, Zheng C, Cao C, Chen F (2018) Long-term inorganic plus organic fertilization increases yield and yield stability of winter wheat. Crop J 6(6):589–599
Cheng X, Xin M, Xu R, Chen Z, Cai W, Chai L, Xu H, Jia L, Feng Z, Wang Z, Peng H, Yao Y, Hu Z, Guo W, Ni Z, Sun Q (2020) A single amino acid substitution in STKc_GSK3 kinase conferring semispherical grains and its implications for the origin of Triticum sphaerococcum. Plant Cell 32(4):923–934
Chidzanga C, Mullan D, Roy S, Baumann U, Garcia M (2022) Nested association mapping-based GWAS for grain yield and related traits in wheat grown under diverse Australian environments. Theor Appl Genet 135:4437–4456
Cristina D, Ciuca M, Cornea PC (2016) Genetic control of grain size and weight in wheat-where are we now? Sci Bull Ser F Biotechnol 20:27–34
Eltaher S, Baenziger PS, Belamkar V, Emara HA, Nower AA, Salem KF, Alqudah AM, Sallam A (2021) GWAS revealed effect of genotype × environment interactions for grain yield of Nebraska winter wheat. BMC Genomics 22(1):1–14
Gahlaut V, Jaiswal V, Balyan HS, Joshi AK, Gupta PK (2021). Multi-locus GWAS for grain weight-related traits under rain-fed conditions in common wheat (Triticum aestivum L.). Front Plant Sci 12:758631
Gao L, Yang J, Song SJ, Xu K, Liu HD, Zhang SH, Yang XJ, Zhao Y (2021) Genome-wide association study of grain morphology in wheat. Euphytica 217(8):1–12
Guo X, Fu Y, Lee YRJ, Chern M, Li M, Cheng M, Dong H, Yuan Z, Gui L, Yin J, Qing H, Zhang C, Pu Z, Liu Y, Li W, Li W, Qi P, Chen G, Jiang Q, Ma J, Chen X, Wei Y, Zheng Y, Wu R, Liu B, Wang J (2022) The PGS1 basic heli-loop-helix protein regulates Fl3 to impact seed growth and grain yield in cereals. Plant Biotechnol J 20(7):1311–1326
Huang XQ, Cöster H, Ganal MW, Röder MS (2003) Advanced backcross QTL analysis for the identification of quantitative trait loci alleles from wild relatives of wheat (Triticum aestivum L.). Theor Appl Genet 106(8):1379–1389
Huang XQ, Cloutier S, Lycar L, Radovanovic N, Humphreys DG, Noll JS, Somers DJ, Brown PD (2006) Molecular detection of QTLs for agronomic and quality traits in a doubled haploid population derived from two Canadian wheats (Triticum aestivum L.). Theor Appl Genet 113:753–766
Jabłoński B, Ogonowska H, Szala K, Bajguz A, Orczyk W, Nadolska-Orczyk A (2020) Silencing of TaCKX1 mediates expression of other TaCKX genes to increase yield parameters in wheat. Int J Mol Sci 21(13):4809
Ji G, Xu Z, Fan X, Zhou Q, Chen L., Yu Q, Liao S, Jiang C, Feng B, Wang T (2023) Identification and validation of major QTL for grain size and weight in bread wheat (Triticum aestivum L.). Crop J 11(2):564–572
Kuchel H, Williams KJ, Langridge P, Eagles HA, Jefferies SP (2007) Genetic dissection of grain yield in bread wheat. I QTL Analysis Theor Appl Genet 115:1029–1041
Kumar A, Mantovani EE, Simsek S, Jain S, Elias EM, Mergoum M (2019) Genome wide genetic dissection of wheat quality and yield related traits and their relationship with grain shape and size traits in an elite × non-adapted bread wheat cross. PLoS ONE 14(9):e0221826
Li F, Wen W, Liu J, Zhang Y, Cao S, He Z, Rasheed A, Jin H, Zhang C, Yan J, Zhang P, Wan Y, Xia X (2019) Genetic architecture of grain yield in bread wheat based on genome-wide association studies. BMC Plant Biol 19:168
Li T, Deng G, Su Y, Yang Z, Tang Y, Wang J, Zhang J, Qiu X, Pu X, Yan W, Li J, Liu Z, Zhang H, Liang J, Yu M, Wei Y, Long H (2022) Genetic dissection of quantitative trait loci for grain size and weight by high-resolution genetic mapping in bread wheat (Triticum aestivum L.). Theor Appl Genet 135:257–271
Li J, Wan H, Yang W, Wang Q, Zhu X, Hu X, Wei H, Tang R, Li C, Peng Z, Zhou Y (2014) Dissection of genetic components in the new high-yielding wheat cultivar Chuanmai 104. Sci Agric Sin 47(12):2281–2291
Li X, Liu N, You L, Ke X, Liu H, Huang M, Waddington SR (2016) Patterns of cereal yield growth across China from 1980 to 2010 and their implications for food production and food security. PLoS ONE 11(7):e0159061
Liu H, Li H, Hao C, Wang K, Wang Y, Qin L, An D, Li T, Zhang X (2020) TaDA1, a conserved negative regulator of kernel size, has an additive effect with TaGW2 in common wheat (Triticum aestivum L.). Plant Biotechnol J 18(5):1330–1342
Liu Y, Lin Y, Gao S, Li Z, Ma J, Deng M, Chen G, Wei Y, Zheng Y (2017) A genome-wide association study of 23 agronomic traits in Chinese wheat landraces. Plant J 91(5):861–873
Liu Z, Wang Q, Wan H, Yang F, Wei H, Xu Z, Ji H, Li J, Yang W (2021) QTL mapping for adult-plant resistance to powdery mildew in Chinese elite common wheat Chuanmai104. Cereal Res Commun 49:99–108
Ma J, Ding P, Qin P, Liu YX, Xie Q, Chen G, Li W, Jiang Q, Chen G, Lan XJ, Wei YM, Liu C, Zheng YL (2017) Structure and expression of the TaGW7 in bread wheat (Triticum aestivum L.). Plant Growth Regul 82:281–291
Ma M, Wang Q, Li Z, Cheng H, Li Z, Liu X, Song W, Apples R, Zhao H (2015) Expression of TaCYP78A3, a gene encoding cytochrome P450 CYP78A3 protein in wheat (Triticum aestivum L.), affects seed size. Plant J 83(2):312–325
Macholdt J, Honermeier B (2017) Yield stability in winter wheat production: a survey on German farmers’ and advisors’ views. Agronomy 7(3):45
Nielsen DC, Vigil MF (2018) Wheat yield and yield stability of eight dryland crop rotations. Agron J 110(2):594–601
Okamoto Y, Nguyen AT, Yoshioka M, Iehisa JC, Takumi S (2013) Identification of quantitative trait loci controlling grain size and shape in the D genome of synthetic hexaploid wheat lines. Breed Sci 63(4):423–429
Patil RM, Tamhankar SA, Oak MD, Raut AL, Honrao BK, Rao VS, Misra SC (2013) Mapping of QTL for agronomic traits and kernel characters in durum wheat (Triticum durum Desf.). Euphytica 190:117–129
Pepó P, Győri Z (2005) A study of the yield stability of winter wheat varieties. Cereal Res Commun 33:769–776
Simmonds J, Scott P, Leverington-Waite M, Turner AS, Brinton J, Korzun V, Snape J, Uauy C (2014) Identification and independent validation of a stable yield and thousand grain weight QTL on chromosome 6A of hexaploid wheat (Triticum aestivum L.). Bmc Plant Biol 14:1–13
Slafer GA, Savin R, Sadras VO (2014) Coarse and fine regulation of wheat yield components in response to genotype and environment. Field Crop Res 157:71–83
Smith SE, Kuehl RO, Ray IM, Hui R, Soleri D (1998) Evaluation of simple methods for estimating broad-sense heritability in stands of randomly planted genotypes. Crop Sci 38(5):1125–1129
Su Z, Hao C, Wang L, Dong Y, Zhang X (2011) Identification and development of a functional marker of TaGW2 associated with grain weight in bread wheat (Triticum aestivum L.). Theor Appl Genet 122:211–223
Suliman S, Alemu A, Abdelmula AA, Badawi GH, Al-Abdallat A, Tadesse W (2021) Genome-wide association analysis uncovers stable QTLs for yield and quality traits of spring bread wheat (Triticum aestivum) across contrasting environments. Plant Gene 25:100269
Sun XY, Wu K, Zhao Y, Kong FM, Han GZ, Jiang HM, Huang XJ, Li RJ, Wang HG, Li SS (2009) QTL analysis of kernel shape and weight using recombinant inbred lines in wheat. Euphytica 165:615–624
Tyagi S, Mir RR, Balyan HS, Gupta PK (2015) Interval mapping and meta-QTL analysis of grain traits in common wheat (Triticum aestivum L.). Euphytica 201:367–380
Varshney RK, Prasad M, Roy JK, Kumar N, Dhaliwal HS, Balyan HS, Gupta PK (2000) Identification of eight chromosomes and a microsatellite marker on 1AS associated with QTL for grain weight in bread wheat. Theor Appl Genet 100:1290–1294
van Frank G, Rivière P, Pin S, Baltassat R, Berthellot JF, Caizergues F, Dalmasso C, Gascuel JS, Hyacinthe A, Mercier F, Montaz H, Ronot B, Goldringer I (2020) Genetic diversity and stability of performance of wheat population varieties developed by participatory breeding. Sustainability 12(1):384
Wang J, Wang R, Mao X, Zhang J, Liu Y, Xie Q, Yang X, Chang X, Li C, Zhang X, Jing R (2020) RING finger ubiquitin E3 ligase gene TaSDIR1-4A contributes to determination of grain size in common wheat. J Exp Bot 71(18):5377–5388
Wang L, Ge H, Hao C, Dong Y, Zhang X (2012) Identifying loci influencing 1,000-kernel weight in wheat by microsatellite screening for evidence of selection during breeding. PLoS ONE 7(2):e29432
Wang S, Zhang X, Chen F, Cui D (2015) A single-nucleotide polymorphism of TaGS5 gene revealed its association with kernel weight in Chinese bread wheat. Front Plant Sci 6:1166
Wang W, Simmonds J, Pan Q, Davidson D, He F, Battal A, Akhunova A, Trick HN, Uauy C, Akhunov E (2018) Gene editing and mutagenesis reveal inter-cultivar differences and additivity in the contribution of TaGW2 homoeologues to grain size and weight in wheat. Theor Appl Genet 131:2463–2475
White J, Sharma R, Balding D, Cockram J, Mackay IJ (2022) Genome-wide association mapping of Hagberg falling number, protein content, test weight, and grain yield in UK wheat. Crop Sci 62(3):965–981
Williams K, Sorrells ME (2014) Three‐dimensional seed size and shape QTL in hexaploid wheat (Triticum aestivum L.) populations. Crop Sci 54(1):98–110
Xin F (2019) “Mapping of Major QTLs for Kernel Shape in Wheat” in Mater dissertation, Northwest A&F University. Northwest A&F University Library Collection, Shaanxi, China
Yan X, Zhao L, Ren Y, Dong Z, Cui D, Chen F (2019) Genome-wide association study revealed that the TaGW8 gene was associated with kernel size in Chinese bread wheat. Sci Rep 9:2702
Yang C, Li X, Zhang D, Wang H, Shao Y, Fang B, Yue J, Ma F, Qin F (2016) Grey relational analysis and path analysis on wheat yield and its three key factors. J Henan Agric Sci 45(10):19–23
Yang Y, Chai Y, Zhang X, Lu S, Zhao Z, Wei D, Chen L, Hu YG (2020) Multi-locus GWAS of quality traits in bread wheat: mining more candidate genes and possible regulatory network. Front Plant Sci 11:1091
Zhang G, Wang Y, Guo Y, Zhao Y, Kong F, Li S (2015) Characterization and mapping of QTLs on chromosome 2D for grain size and yield traits using a mutant line induced by EMS in wheat. Crop J 3(2):135–144
Zhang S, Guo H, Irshad A, Xie Y, Zhao L, Xiong H, Gu J, Zhao S, Ding Y, Liu, L. (2019) The synergistic effects of TaAGP.L-B1 and TaSSIVb-D mutations in wheat lead to alterations of gene expression patterns and starch content in grain development. Plos One 14(10):e0223783
Zhang Y, Miao H, Wang C, Zhang J, Zhang X, Shi X, Xie S, Li T, Deng P, Wang C, Chen C, Zhang H, Ji W (2022) Genetic identification of the pleiotropic gene Tasg-D1/2 affecting wheat grain shape by regulating brassinolide metabolism. Plant Sci 323:111392
Acknowledgements
This work was supported by National Key Research and Development Plan (2021YFD120060), Sichuan Provincial Finance Department Project (1+3 ZYGG001), Sichuan Province Science and Technology Department Project (2023NSFSC1925, 2021YFYZ0020, 2022NSFSC0161), and the Program of Chinese Agriculture Research System (CARS-03), Strategic Scientist Studio, Sichuan Academy of Agricultural Sciences. We thank Robert McKenzie, PhD, from Liwen Bianji (Edanz) (www.liwenbianji.cn) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declared that they have no conflict of interest to this work. We declare that we do not have any commercial or associative interest that represents a conflict of interest in connection with the work submitted.
Additional information
Communicated by Ankica Kondic-Spika.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, Z., Tang, H., Lin, Y. et al. Identification and stability of QTLs for grain traits in the elite wheat variety ‘Chuanmai104’. CEREAL RESEARCH COMMUNICATIONS (2024). https://doi.org/10.1007/s42976-024-00577-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42976-024-00577-5