Abstract
Leptospira spp. are bacteria responsible for leptospirosis, a zoonotic disease with considerable impacts on the economy, animal health, and public health. This disease has a global distribution and is particularly prevalent in Brazil. Both rural and urban environments are habitats for Leptospira spp., which are primarily transmitted through contact with the urine of infected animals. Consequently, domestic and wild species can harbor these prokaryotes and serve as infection sources for other hosts. In the context of wild animals, there is a dearth of molecular studies elucidating the roles of various animal and bacterial species in the epidemiology of leptospirosis. Therefore, this study aimed to evaluate the presence of Leptospira spp. DNA in different species of free-living and captive wild animals and to assess the phylogenetic relationships of the identified microorganisms in Rio Grande do Sul, Brazil. The samples were evaluated for the presence of the gene lipL32 by polymerase chain reaction (PCR) and sequencing of the amplified fragment after which phylogenetic analyzes were carried out. DNA from Leptospira spp. was extracted from kidney tissue from wild animals (Mammalia class). Pathogenic Leptospira spp. DNA was detected in 9.6% (11/114) of the samples, originating from nine species of wild animals, including the white-eared opossum (Didelphis albiventris), skunk (Conepatus chinga), geoffroy’s cat (Leopardus geoffroyi), margay (Leopardus wiedii), pampas fox (Lycalopex gymnocercus), capybara (Hydrochoerus hydrochaeris), common marmoset (Callithrix jacchus), neotropical river otter (Lontra longicaudis), and european hare (Lepus europaeus). Phylogenetic analysis revealed the presence of Leptospira borgpetersenii and Leptospira interrogans in these animals. This research is the first study contributing to the epidemiology of leptospirosis by identifying L. borgpetersenii and L. interrogans in free-living and captive wild animals in Rio Grande do Sul, Brazil, potentially acting as bacterial reservoirs. Additionally, our findings can inform sanitary measures for controlling and preventing the disease, thereby safeguarding public health.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacteria of the genus Leptospira serve as the etiological agents of leptospirosis, a disease that significantly impacts the economy, public health, and animal [1, 2]. Classified as a zoonosis, this disease has a global distribution and is particularly prevalent in Brazil [3, 4]. In endemic regions, the persistence of leptospirosis outbreaks is often linked to reservoir hosts capable of harboring Leptospira spp. for extended periods. These hosts may or may not exhibit clinical signs but contribute to the spread of the infectious agent in both rural and urban areas [1, 5, 6].
Environments contaminated with urine from infected animals—such as soil, mud, or water—act as transmission sources for the microorganism to animals and humans, primarily through mucous membranes or skin [7, 8]. While rodents are the main reservoirs of the etiological agent [1, 9], various animal species, including wild animals, can action as hosts and reservoirs for Leptospira spp. in specific regions [5, 6, 10,11,12]. Therefore, investigations into Leptospira spp. in wild animals are important to provide information about the epidemiology of this relevant zoonotic infection, since these animals often coexist with humans and domestic animals [11, 13,14,15].
Given that wild animals often share habitats with humans and domestic animals, studying Leptospira spp. in these species is crucial for understanding the epidemiology of this significant zoonotic [11, 13,14,15]. Understanding the animal host range and geographic distribution of Leptospira species is essential for identifying strains in local animal hosts that can infect people and other animals [16,17,18,19,20]. Tropical countries such as Brazil, which boast extensive biodiversity, provide numerous animal species that warrant investigation as potential Leptospira spp. reservoirs [21], as demonstrated in several Brazilian studies that directly or indirectly detected Leptospira spp. in wild mammals [3, 22, 23]. Therefore, this study aims to assess the presence of Leptospira spp. DNA in various species of free-living and captive wild animals in Rio Grande do Sul, Brazil and to analyze the phylogenetic relationships among Leptospira spp. identified.
Materials and methods
This study examined kidney tissue samples from 114 wild animals, comprising 75 free-living and 39 captive-bred specimens. All animals belonged to Mammalia class and died in Rio Grande do Sul State, in South of Brazil, between 2021 and 2023. They were sent for necropsy without suspicion of leptospirosis to the Laboratório de Patologia Veterinária (Veterinary Pathology Laboratory) at Universidade Federal de Santa Maria (Federal University of Santa Maria, UFSM) (Table 1). During necropsy, a single kidney from each animal was individually harvested and stored at -20°C until molecular analysis. Taxonomic identification was conducted according to the family, genus, and species, as described by Cubas et al. [24] and Hickman et al. [25].
Kidney tissue samples were sent to the Laboratório de Diagnóstico e Pesquisa em Leptospirose (Leptospirosis Diagnostic and Research Laboratory) at UFSM, where were homogenized, and an aliquot (~20 mg) was placed in polypropylene microtubes for total DNA extraction, following a protocol adapted for tissue samples [26]. Tissue fragments were lysed in a buffer containing 2-β-mercaptoethanol, 2% sodium dodecyl sulfate, 10% cetyltrimethylammonium bromide, and 5N sodium chloride. DNA was extracted using a phenol-chloroform method and reconstituted in 40 µL of sterile Tris-EDTA buffer. DNA concentrations were quantified via spectrophotometry.
A polymerase chain reaction (PCR) was performed to amplify a 242-base pair fragment of the lipL32 gene, which encodes for outer membrane proteins exclusively found in pathogenic Leptospira spp. [12]. The sensitivity of the PCR reaction was verified through the detection threshold of the positive control, which detected up to 1.5 × 103 bacteria/mL. The PCR sample was prepared to a final volume of 12.5 µL containing 1 x buffer (Ludwig Biotec, Brazil), 1.5 mM MgCl2 (Ludwig Biotec, Brazil), 0.2 mM dNTPs (Ludwig Biotec, Brazil), 2.5 U of Taq DNA polymerase (Ludwig Biotec, Brazil), 50 nM of each primer (Invitrogen, Brazil) lipL32-45F (5′-AAG CAT TACCGC TTG TGG TG-3′) and lipL32-286R (5′-GAA CTC CCA TTT CAG CGA TT-3′), and 2.5 µL (330 ng/µL) of the extracted DNA sample. The amplification was carried out in a PCR thermal cycler (K960, TION96, Brazil) using a specific set of cycling conditions, consisting of an initial denaturation of 94 °C for 2 min, 35 cycles of 94 °C for 30 s, 53 °C for 30 s, 72 °C for 1 min, followed by a final extension at 72 °C for 5 min. The PCR products were subsequently analyzed through horizontal electrophoresis on a 1% agarose gel, which was stained with non-mutagenic Safer dye (Kasvi, Brazil), observed under ultraviolet light, and photodocumented.
The samples amplified in the PCR were purified using a PCR purification kit (Ludwig Biotec, Brazil) according to the manufacturer’s instructions and sent for DNA sequencing (ACTGene Análises Moleculares, Brazil). The resulting sequences were aligned using the MEGA X software [27], compared among themselves, and with reference sequences available in the GenBank (MN906895, MK328874, MK568983, MK568984). A phylogenetic tree was constructed using Bayesian analysis [28], and the bootstrap resampling method was employed as a phylogeny test with 500 replications [29].
Results
LipL32 was detected in 9.2% (11/114) of the samples examined. Among these amplified samples, ten were identified in at least one distinct species of wild animal evaluated, as listed in Table 1.
Of the positive samples, nine species of wild animals were identified, including the white-eared opossum (D. albiventris) at 18.2% (2/11), skunk (C. chinga) at 18.2% (2/11), geoffroy’s cat (L. geoffroyi) at 9.1% (1/11), margay (L. wiedii) at 9.1% (1/11), pampas fox (L. gymnocercus) at 9.1% (1/11), capybara (H. hydrochaeris) at 9.1% (1/11), common marmoset (C. jacchus) at 9.1% (1/11), neotropical river otter (L. longicaudis) at 9.1% (1/11), and european hare (L. europaeus) at 9.1% (1/11), all from Rio Grande do Sul, Brazil. Regarding the sex of the animals that tested positive, 81.8% (9/11) were males and 18.2% (2/11) were females. In terms of age distribution, 90.9% (10/11) of the animals were adults.
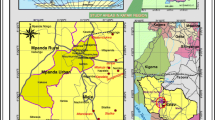
The evaluated samples from free-living animals came from Santa Maria (67/114), Palmeira das Missões (5/114), Lagoa Vermelha (1/114), and Cruz Alta (1/114) municipalities. Captive-bred animal samples were collected from Cachoeira do Sul (28/114) and Santa Maria (12/114) cities. Among the animals that tested positive for pathogenic Leptospira spp. (11/114), 63.6% (7/11) were free-living— 71.4% (5/7) of which were from Santa Maria and 28.6% (2/7) from Palmeira das Missões. The remaining 36.4% (4/11) were captive-bred, with 50% (2/4) from Cachoeira do Sul and 50% (2/4) from Santa Maria.
In the phylogenetic analysis (Figure 1), eleven fragments of the gene lipL32 of Leptospira spp. were sequenced from nine different species of wild animals in Rio Grande do Sul State showed a grouping with sequences belonging to the pathogenic species L. interrogans (OR578518, OR578519, OR578521, OR578522, OR795078, OR795076, OR795077, OR795075) and L. borgptersenii (OR513921, OR513922, OR513923).
Phylogenetic analysis of gene lipL32 sequences from Leptospira interrogans and Leptospira borgptersenii obtained from wild animals in Rio Grande do Sul State, in South of Brazil. The analysis was carried out using the Bayesian method, with 500 bootstraps, in MEGA X software. L. interrogans and L. borgptersenii sequences from the samples analyzed are highlighted in bold
Discussion
The presence of pathogenic Leptospira spp. DNA was predominantly detected in mammals of Carnivora order. Phylogenetic analysis revealed that species L. interrogans and L. borgptersenii are present in wild mammals in Rio Grande do Sul State, Brazil, a critical international transit region for both humans and animals moving between Brazil, Uruguay, and Argentina [30]. In Brazil, various studies have reported the presence of Leptospira spp. DNA in different biomes [3, 30].
Our study revealed the presence of pathogenic Leptospira spp. DNA in diverse wild animals that live in the southernmost state of Brazil, in areas of the Pampa biome, the Mata Atlantica biome and transition zones between these two biomes. This suggests the involvement of wild animals in the epidemiological chain of leptospirosis, highlighting a variety of wild hosts that can act as reservoirs for this pathogen [6]. Among studies employing molecular detection techniques for leptospirosis diagnosis, Carnivora order has been the most extensively studied [3, 11]. In our study, pathogenic Leptospira spp. DNA was detected in 54.55% (6/11) of carnivore samples (Table 1). This higher occurrence in carnivores could be attributed to their extensive terrestrial movements, including through flooded areas, primarily in search of food and preying on other potentially infected animal species [31].
In our study, the presence of L. interrogans (1/23) and L. borgpetersenii (1/23) DNA was detected in kidney tissue of white-eared opossum. These findings might be linked to the omnivorous diet of these animals [22], as well as their extensive habitat range, which includes forests, shrublands, grasslands, and both rural and urban areas [32,33,34], thereby increasing their exposure to Leptospira spp.
Rodents, particularly of the Rodentia order, have been extensively studied in various regions [29, 35,36,37]. However, in this study, L. borgpetersenii DNA was found in one kidney tissue sample from a capybara (1/11). This is, likely, because these large rodents inhabit flood-prone pastures [38], a significant environmental factor for Leptospira spp. transmission [39]. Thus, capybaras are considered important reservoirs for this pathogen, and given their proximity to farm animals and semi-urban areas, they represent a risk to both animal and public health [40].
Epidemiological studies in leptospirosis involving Lagomorpha and Primate orders are relatively scarce. Notably, this study detected L. borgpetersenii DNA in a captive European hare (L. europaeus) (1/3) and L. interrogans DNA in a captive common marmoset (C. jacchus) (1/2). For Artiodactyla order, no positive samples were found in this study, contrasting with findings from other regions. For example, in New Caledonia, deer species tested positive for L. interrogans and L. borgpetersenii DNA [41, 42]. Similarly, pampas deer (O. bezoarticus) from Brazil’s Pantanal biome showed a 3% positivity rate in blood PCR tests [43].
In this study, both L. interrogans and L. borgptersenii were detected. L. interrogans is considered the most widely distributed species globally and has been described in various hosts, including wild animals [30, 43, 44], synanthropic animals [45], domestic animals [46], humans [47, 48], and even environmental samples [49]. L. borgptersenii considered a bacterium that has already been found in rodents [50, 51] and in cattle [52], but is not expected its detection in different wild animal species. However, in this study it was possible to observe that this bacterial species is found circulating in species of wild mammals, such as white-eared opossums, capybara and neotropical river otters, probably due to the proximity of these animals to herds of cattle, as well as rodents possibly infected with Leptospira borgptersenii [51].
Due to the limited number of studies that address the epidemiological aspects of leptospirosis in different regions in Brazil, the importance of this investigation is owing to the detection of important pathogenic Leptospira species in wild animals from Rio Grande do Sul. Factors such as rainfall, water availability, and elevated temperatures significantly influence the survival of Leptospira spp. in the environment [53, 54]. Therefore, the high proportion of molecular detection of Leptospira spp. in free-living wild animals (7/11) from the cities of Santa Maria (5/7) and Palmeira das Missões (2/7) can be attributed to favorable ecological conditions. These include climatic elements that present four distinct seasons, with summer characterized by abundant solar radiation and higher temperatures and winter marked by lower average temperatures [55, 56]. The year and the resulting intense vegetation growth create favorable conditions for the survival of several species of wild animals and the maintenance of Leptospira spp. [56].
Beyond the ecological considerations, free-living animals present a health risk to other animals and humans in the evaluated areas. They also pose occupational risks to environmental police officers, veterinarians, biologists, and other professionals who may come into contact with these animals [57]. Likewise, captive animals constitute an occupational risk for those who work directly with them in settings such as breeding facilities and zoos [58]. In this study, pathogenic Leptospira spp. DNA was detected in samples from captive animals, including common marmosets, capybaras, margays, and European hares. This may be attributable to the stress and behavioral changes experienced by animals in captivity, leading to compromised health [59]. Moreover, these captive settings may be located in urban areas where synanthropic animals serve as important reservoirs for Leptospira spp. [60,61,62,63].
This study is the first to report the molecular detection of pathogenic Leptospira spp., including L. interrogans and L. borgpetersenii, in kidney tissue samples from free-living and captive wild animals in Rio Grande do Sul, Brazil.
Conclusion
This study demonstrates the presence of L. interrogans and L. borgpetersenii DNA in kidney tissue samples from free-living and captive wild animals, predominantly from Mammalia class, in Rio Grande do Sul, Brazil. Therefore, it can be inferred that these animals can act as reservoirs in the epidemiology of leptospirosis. Thus, this research also highlights the need for continuous epidemiological surveillance of leptospirosis in wild mammal populations to mitigate the risks of transmission of the etiological agent to humans and other species of domestic and wild animals. In addition, it is suggested that wild animals be included in the monitoring of the epidemiology of this important zoonotic disease with the aim of guiding leptospirosis control and prevention measures, especially in endemic regions.
Reference
Guernier V, Lagadec E, Cordonin C, Le Minter G, Gomard Y, Pages F, Jaffar- Bandjee MC, Michault A, Tortosa P, Dellagi K (2016) Human leptospirosis on Reunion Island, Indian Ocean: are rodents the (only) ones to blame? PLoS Negl Trop Dis 10:4733. https://doi.org/10.1371/journal.pntd.0004733
Torres FD, Borges ALDSB, Kolesnikovas C, Domit C, Barbosa CB, Carvalho-Costa FA, Di Azevedo MIN, Lilenbaum W (2023) Pinnipeds carriers of pathogenic Leptospira: New data based on molecular characterization. Res Vet Sci 155:62–68. https://doi.org/10.1016/j.rvsc.2022.12.012
Vieira AS, Narduche L, Martins G, Schabib Péres IA, Zimmermann NP, Juliano RS, Pellegrin AO, Lilenbaum W (2016) Detection of wild animals as carriers of Leptospira by PCR in the Pantanal biome. Brazil Acta Trop 163:87–9. https://doi.org/10.1016/j.actatropica.2016.08.001
Marteli AN, Guasselli LA, Diament D, Wink GO, Vasconcelos VV (2022) Spatio-temporal analysis of leptospirosis in Brazil and its relationship with flooding. Geospat Health 17. https://doi.org/10.4081/gh.2022.1128.
Adler B, de la Peña Moctezuma A (2010) Leptospira and leptospirosis. Vet Microbiol 140:287–296. https://doi.org/10.1016/j.vetmic.2009.03.012
Cilia G, Bertelloni F, Fratini F (2020) Leptospira Infections in Domestic and Wild Animals. Pathog 9:573. https://doi.org/10.3390/pathogens9070573
Bharti AR (2003) Peru-United States Leptospirosis C. Leptospirosis: a zoonotic disease of global importance. Lancet Infect Dis 3:757–771. https://doi.org/10.1016/S1473-3099(03)00830-2
Mateus J, Gómez N, Herrera-Sepúlveda MT, Hidalgo M, Pérez-Torres J, Cuervo C (2019) Bats are a potential reservoir of pathogenic Leptospira species in Colombia. J Infect Dev Ctries 13:278–283. https://doi.org/10.3855/jidc.10642
Boey K, Shiokawa K, Rajeev S (2019) Leptospira infection in rats: a literature review of global prevalence and distribution. PLoS Negl Trop Dis 13:e0007499. https://doi.org/10.1371/journal.pntd.0007499
Petrakovsky J, Bianchi A, Fisun H, Nájera-Aguilar P, Pereira MM (2014) Animal leptospirosis in Latin America and the Caribbean countries: reported outbreaks and literature review (2002–2014). Int J Environ Res Public Health 11:10770–10789. https://doi.org/10.3390/ijerph111010770
Vieira AS, Pinto PS, Lilenbaum W (2018) A systematic review of leptospirosis on wild animals in Latin America. Trop Anim Health Prod 50:229–238. https://doi.org/10.1007/s11250-017-1429-y
Ulsenheimer BC, von Laer AE, Tonin AA, Campos AAS, Dos Santos HF, Sangioni LA, Botton SA (2022) Leptospira interrogans in bats in Rio Grande do Sul State, Brazil: epidemiologic aspects and phylogeny. Braz J Microbiol 53:2233–2240. https://doi.org/10.1007/s42770-022-00838-7
Plowright RK, Foley P, Field HE, Dobson AP, Foley JE, Eby P, Daszak P (2011) Urban habituation, ecological connectivity and epidemic dampening: the emergence of Hendra virus from flying foxes (Pteropus spp.). Proc Biol Sci 278:3703–3712. https://doi.org/10.1098/rspb.2011.0522
Lei BR, Olival KJ (2014) Contrasting patterns in mammal–bacteria coevolution: Bartonella and Leptospira in bats and rodents. PLoS Negl Trop Dis 8:e2738. https://doi.org/10.1371/journal.pntd.0002738
Ramírez-Chaves HE, Suárez-Castro AF, González-Maya JF (2016) Recent changes to the list of mammals in Colombia. Ma No 1:1–9. https://doi.org/10.47603/manovol3n1.1-9.
Cilia G, Bertelloni F, Albini S, Fratini F (2021) Insight into the epidemiology of leptospirosis: A review of Leptospira isolations from “unconventional” hosts. Animals 11:191. https://doi.org/10.3390/ani11010191
Hagedoorn NN, Maze MJ, Carugati M, Cash-Goldwasser S, Allan KJ, Chen K, Cossic B, Demeter E, Gallagher S, German R, Galloway RL, Habuš J, Rubach MP, Shiokawa K, Sulikhan N, Crump JA (2024) Global distribution of Leptospira serovar isolations and detections from animal host species: A systematic review and online database. Trop Med Int Health. https://doi.org/10.1101/2023.10.03.23296503
Mazzotta E, Bellinati L, Bertasio C, Boniotti MB, Lucchese L, Ceglie L, Martignago F, Leopardi S, Natale A (2023) Synanthropic and wild animals as sentinels of zoonotic agents: a Study of Leptospira genotypes circulating in Northeastern Italy. Int J Environ Res Public Health 20(5):3783. https://doi.org/10.3390/ijerph20053783
Vieira AS, Pinto PS, Lilenbaum W (2018) A systematic review of leptospirosis on wild animals in Latin America. Trop Anim Health Prod 50:229–238. https://doi.org/10.1007/s11250-017-1429-y
Fratini F, Turchi B, Ebani VV, Bertelloni F, Galiero A, Cerri D (2015) The presence of Leptospira in coypus (Myocastor coypus) and rats (Rattus norvegicus) living in a protected wetland in Tuscany (Italy). Veterinarski Arhiv 85:407–414
Rocha BR, Martins G, Lilenbaum W (2020) An historical view of the experimental leptospiral infection in ruminants. Comp Immunol Microbiol Infect Dis 73:101532. https://doi.org/10.1016/j.cimid.2020.101532
Silva TF, de Quadros APN, do Rêgo GMS, de Oliveira J, de Medeiros JT, Dos Reis LFM, Ribeiro TMP, Carvalho MV, de Mattos PSR, Mathias LA, Paludo GR (2023) Leptospira spp. in Free-Ranging Capybaras (Hydrochoerus hydrochaeris) from Midwestern Brazil. Vector Borne Zoonotic Dis 23:106–112. https://doi.org/10.1089/vbz.2022.0034
Paz LN, Hamond C, Pinna MH (2022) Detection of Leptospira interrogans in Wild Sambar Deer (Rusa unicolor), Brazil. EcoHealth 19:15–21. https://doi.org/10.1007/s10393-022-01577-9
Cubas ZS, Silva JCR, Catão-Dias JL (2014) Tratado de Animais Selvagens-Medicina Veterinária - 2 Vol. São Paulo: Grupo GEN. Available in: https://repositorio.usp.br/item/002652819. Acessed in: 28/08/2023.
Hickman CP, Keen SL, Eisenhour DJ, Larson A, I'Anson H (2022) Princípios Integrados de Zoologia. 18. ed. – Rio de Janeiro: Guanabara Koogan, 2022. Available in: https://www.grupogen.com.br/e-book-principios-integrados-de-zoologia-cleveland-p-hickman-jr-susan-keen-david-j-einsenhour-allan-larson-e-helen-i-anson-guanabara-koogan-9788527738644. Acessed in: 28/08/2023.
Botton SA, Pereira DI, Costa MM, Azevedo MI, Argenta JS, Jesus FP, Alves SH, Santurio JM (2011) Identification of Pythium insidiosum by Nested PCR in cutaneous lesions of Brazilian horses and rabbits. Curr Microbiol 62:1225–1229. https://doi.org/10.1007/s00284-010-9781-4
Fennestad KL, Borg-Petersen C (1972) Leptospirosis in Danish wild mammals. J Wildl Dis 8:343–351. https://doi.org/10.7589/0090-3558-8.4.343
Kobayashi Y, Ogawa A, Sato G, Sato T, Itou T, Samara SI, Carvalho AA, Nociti DP, Ito FH, Sakai T (2006) Geographical distribution of vampire bat-related cattle rabies in Brazil. J Vet Med Sci 68:1097–1100. https://doi.org/10.1292/jvms.68.1097
Souza PG, Amaral BMPM, Gitti CB (2014) Raiva animal na cidade do Rio de Janeiro: emergência da doença em morcegos e novos desafios para o controle. Rev Inst Adolfo Lutz 73:119–124. https://doi.org/10.18241/0073-98552014731596
Ellwanger JH, Ziliotto M, Chies JAB (2022) Protect Brazil’s overlooked Pampa biome. Sci 377:720. https://doi.org/10.1126/science.ade1838
Jorge RS, Ferreira F, Ferreira Neto JS, Vasconcellos SA, Lima ES, Morais ZM, Souza GO (2011) Exposure of free-ranging wild carnivores, horses and domestic dogs to Leptospira spp. in the northern Pantanal. Brazil Mem Inst Oswaldo Cruz 106:441–444. https://doi.org/10.1590/S0074-02762011000400009
Jorge S, Hartleben CP, Seixas FK, Coimbra MA, Stark CB, Larrondo AG, Amaral MG, Albano AP, Minello LF, Dellagostin OA, Brod CS (2012) Leptospira borgpetersenii from free-living white-eared opossum (Didelphis albiventris): first isolation in Brazil. Acta Trop 124:147–51. https://doi.org/10.1016/j.actatropica.2012.07.009
Fernandes JJ, de Lima Peixoto A, de Farias ASS, Junior Pinheiro T, da Costa DF, Silva MLCR, Júnior JPA, Malossi CD, Ullmann LS, de Azevedo SS, Alves CJ, Santos Higino SS (2020) Didelphis albiventris as a carrier of Leptospira sp. in the central nervous tissue in the semiarid region of Northeast. Brazil Comp Immunol Microbiol Infect Dis 73:101560. https://doi.org/10.1016/j.cimid.2020.101560
Weber MM, Roman C, Cáceres NC (2013) Mamíferos do Rio Grande do Sul. Editora UFSM. Available in: https://editoraufsm.com.br/mamiferos-do-rio-grande-do-sul.html. Acessed in: 20/09/2023.
Xu G, Qiu H, Liu W, Jiang X, Chang YF, Wang J, Li Z, Zhu Y, Zhang C, Xiao F (2022) Serological and molecular characteristics of pathogenic Leptospira in rodent populations in Fujian Province, China, 2018–2020. BMC Microbiol 22:151. https://doi.org/10.1186/s12866-022-02566-2
Zhang C, Xu J, Zhang T, Qiu H, Li Z, Zhang E, Li S, Chang YF, Guo X, Jiang X, Zhu Y (2019) Genetic characteristics of pathogenic Leptospira in wild small animals and livestock in Jiangxi Province, China, 2002–2015. PLoS Negl Trop Dis 13:e0007513. https://doi.org/10.1371/journal.pntd.0007513
Schmidt E, Obiegala A, Imholt C, Drewes S, Saathoff M, Freise J, Runge M, Jacob J, Mayer-Scholl A, Ulrich RG, Pfeffer M (2021) Influence of Season, Population and Individual Characteristics on the Prevalence of Leptospira spp. in Bank Voles in North-West Germany. Biology (Basel) 10:933. https://doi.org/10.3390/biology10090933
Cueto GR, Allekotte R, Kravetz FO (2000) Scurvy in capybaras bred in captivity in Argentine. J Zoo Wildl Med 36:97–101. https://doi.org/10.7589/0090-3558-36.1.97
Moreno LZ, Miraglia F, Marvulo MF, Silva JC, Paula CD, Costa BL, Morais ZM, Ferreira F, Neto JS, Dellagostin OA, Hartskeerl RA, Vasconcellos SA, Moreno AM (2016) Characterization of Leptospira santarosai Serogroup Grippotyphosa Serovar Bananal Isolated from Capybara (Hydrochaeris hydrochaeris) in Brazil. J Wildl Dis 52:688–93. https://doi.org/10.7589/2015-09-245
Gonçalves DD, Lopes KFC, Chiderolli RT, Sampieri BR, Rocha VJ, Pachaly JR, Santos IC, Barbosa LN, Mota EA, Pereira UP (2020) Leptospirosis in free-living capybaras (Hydrochaeris hydrochaeris) from a university campus in the city of Araras in São Paulo. Brazil. Semina: Ciênc Agrár 41:159–166. https://doi.org/10.5433/1679-0359.2020v41n1p159
Perez J, Goarant C (2010) Rapid Leptospira identification by direct sequencing of the diagnostic PCR products in New Caledonia. BMC Microbiol 10:325. https://doi.org/10.1186/1471-2180-10-325
Gay N, Soupe MEG, Goarant C (2014) Though not reservoirs, dogs might transmit leptospira in New Caledonia. Int J Environ Res Public Health 11:4316–4325. https://doi.org/10.3390/ijerph110404316
Vieira AS, Rosinha GMS, de Oliveira CE, Vasconcellos SA, Lima-Borges PA, Tomas WM, Mourão GM, Lacerda ACR, Soares CO, de Araújo FR, Piovezan U, Zucco CA, Pellegrin AO (2011) Survey of Leptospira spp in pampas deer (Ozotoceros bezoarticus) in the Pantanal wetlands of the state of Mato Grosso do Sul, Brazil by serology and polymerase chain reaction. Mem Inst Oswaldo Cruz 106:763–768. https://doi.org/10.1590/S0074-02762011000600019
Paz LN, Hamond C, Dias CS, Curvelo VP, Medeiros MA, Oria AP, Pinna MH (2019) Detection of Leptospira spp. in Captive Broad-Snouted Caiman (Caiman latirostris). Ecohealth 16:694–700. https://doi.org/10.1007/s10393-019-01452-0
Rodamilans GM, Fonseca MS, Paz LN, Fernandez CC, Biondi I, Lira-da-Silva RM, Meyer R, Pinna MH, Portela RD (2020) Leptospira interrogans in wild Boa constrictor snakes from Northeast Brazil peri-urban rainforest fragments. Acta Trop 209:105572. https://doi.org/10.1016/j.actatropica.2020.105572
Faria MT, Calderwood MS, Athanazio DA, McBride AJA, Hartskeerl RA, Pereira MM, Ko AI, Reis MG (2008) Carriage of Leptospira interrogans among domestic rats from an urban setting highly endemic for leptospirosis in Brazil. Acta Trop 108:1–5. https://doi.org/10.1016/j.actatropica.2008.07.005
Almeida DS, Paz LN, de Oliveira DS, Silva DN, Ristow P, Hamond C, Costa F, Portela RW, Estrela-Lima A, Pinna MH (2019) Investigation of chronic infection by Leptospira spp. in asymptomatic sheep slaughtered in slaughterhouse. PLoS One 14:e0217391. https://doi.org/10.1371/journal.pone.0217391
Nascimento ALTO et al (2004) Comparative Genomics of Two Leptospira interrogans Serovars Reveals Novel Insights into Physiology and Pathogenesis. J Bacteriol 186:2164–2172. https://doi.org/10.1128/JB.186.7.2164-2172.2004
Hagan JE et al (2016) Spatiotemporal Determinants of Urban Leptospirosis Transmission: Four-Year Prospective Cohort Study of Slum Residents in Brazil. PLoS Negl Trop Dis 10:e0004275. https://doi.org/10.1371/journal.pntd.0004275
Schneider AG, Casanovas-Massana A, Hacker KP, Wunder EA, Begon M, Reis MG, Childs JE, Costa F, Lindow JC, Ko AI (2018) Quantification of pathogenic Leptospira in the soils of a Brazilian urban slum. PLoS Negl Trop Dis 12:1–15. https://doi.org/10.1371/journal.pntd.0006415
Colombo VC, Gamietea I, Loffler SG, Brihuega BF, Beldomenico PM (2018) New host species for Leptospira borgpetersenii and Leptospira interrogans serovar Copenhageni. Vet Microbiol 215:90–92. https://doi.org/10.1016/j.vetmic.2018.01.007
Moinet M, Wilkinson DA, Aberdein D, Russell JC, Vallée E, Collins-Emerson JM, Heuer C, Benschop J (2021) Of Mice, Cattle, and Men: A Review of the Eco-Epidemiology of Leptospira borgpetersenii Serovar Ballum. Trop Med Infect Dis 6:189. https://doi.org/10.3390/tropicalmed6040189
Hamond C, Dirsmith KL, LeCount K, Soltero FV, Rivera-Garcia S, Camp P, Anderson T, Hicks JA, Galloway R, Sutherland G, Schafer IJ, Goris MGA, van der Linden H, Stuber T, Bayles DO, Schlater LK, Nally JE (2022) Leptospira borgpetersenii serovar Hardjo and Leptospira santarosai serogroup Pyrogenes isolated from bovine dairy herds in Puerto Rico. Front Vet Sci 9:1025282. https://doi.org/10.3389/fvets.2022.1025282
Andre-Fontaine G, Aviat F, Thorin C (2015) Waterborne leptospirosis: Survival and preservation of the virulence of pathogenic Leptospira spp. in fresh water. Curr Microbiol 71:136–42. https://doi.org/10.1007/s00284-015-0836-4
Correia L, Loureiro AP, Lilenbaum V (2017) Effects of rainfall on incidental and host-maintained leptospiral infections in cattle in a tropical region. Vet J 220:63–64. https://doi.org/10.1016/j.tvjl.2016.12.016
Heringer I, Jacques AVA (2002) Acumulação de forragem e material morto em pastagem nativa sob distintas alternativas de manejo em relação às queimadas. R Bras Zootec 31:599–604. https://doi.org/10.1590/S1516-35982002000300009
Trentin CB, Trentin AB, Moreira A, Righi E (2021) Características da Vegetação dos Biomas Pampa e Cerrado Monitorados por NDVI. Revista Geoaraguaia 11:69–84
Mergulhão FV (2019) Leptospirose em mamíferos recebidos pelo centro de triagem de animais silvestres do Distrito Federal. Dissertação de mestrado em saúde animal. Brasília/DF. Available in: https://www.repositorio.unb.br/bitstream/10482/36887/1/2019_FernandaVianaMergulh%C3%A3o.pdf. Acessed in: 21/09/2023.
Corrêa SHR, Vasconcellos SA, Morais Z, Teixeira AA, Dias RA, Guimarães MABV, Ferreira F, Ferreira Neto JS (2004) Epidemiologia da Leptospirose em animais silvestres na Fundação Parque Zoológico de São Paulo. BJVRAS 41:189–193. https://doi.org/10.1590/S1413-95962004000300007
Pearson BL, Reeder DM, Judge PG (2015) Crowding increases salivary cortisol but not self-directed behavior in captivity baboons. Am J Primatol 77:462–467. https://doi.org/10.1002/ajp.22363
Ullmann LS, Hoffmann JL, Moraes W, Cubas ZS, Santos LC, Silva RC, Moreira N, Guimaraes AMS, Camossi LG, Langoni H, Biondo AW (2012) Serologic survey for Leptospira SPP. in captive neotropical felids in Foz do Iguaçu, Paraná. Brazil J Zoo Wildl Med 43:223–228. https://doi.org/10.1638/2010-0091.1
Moreno-Beas E, Abalos P, Hidalgo-Hermoso E (2015) Seroprevalence of nine Leptospira interrogans serovars in wild carnivores, ungulates, and primates from a zoo population in a Metropolitan region of Chile. J Zoo Wildl Med 46:774–778. https://doi.org/10.1638/2014-0139.1
Vieira AS, Lilenbaum W (2017) Leptospirosis on captive wild animals in Latin America. Res Vet Sci 115:496–500. https://doi.org/10.1016/j.rvsc.2017.08.001
Acknowledgments
The authors would like to thank the Brazilian development agencies: Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) (financial code 001) and Fundação de Amparo à Pesquisa do Estado do Rio Grande do Sul (FAPERGS).
Funding
Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) (financial code 001), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and Fundação de Amparo à Pesquisa do Estado do Rio Grande do Sul (FAPERGS).
Author information
Authors and Affiliations
Contributions
Bruna Carolina Ulsenheimer: Conceptualization, Methodology, Research, Writing – original draft, Visualization. Helton Fernandes dos Santos, Luís Antônio Sangioni, Rafael Fighera, Matheus dos Santos: Methodology, Writing – review and editing. Sônia Botton, Ana Eucares von Laer, Daniela Brayer Pereira and Alexandre Alberto Tonin: Methodology, Writing – review and editing, Supervision, Funding acquisition.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that there are no conflicts of interest.
Additional information
Responsible Editor: Maria Aparecida Scatamburlo Moreira
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
L. borgpetersenii and L. interrogans in free-living and captive-bred animals;
Pathogenic Leptospira spp. DNA was detected in kidney tissue;
Detection of two pathogenic species of Leptospira spp. in wild mammals;
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ulsenheimer, B.C., Tonin, A.A., von Laer, A.E. et al. Leptospira borgptersenii and Leptospira interrogans identified in wild mammals in Rio Grande do Sul, Brazil. Braz J Microbiol 55, 1941–1948 (2024). https://doi.org/10.1007/s42770-024-01348-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-024-01348-4