Abstract
The disposal of industrial effluents strongly influences low-order streams, which makes them fragile ecosystems that can be impacted by contamination. In central Brazil, the Extrema River spring targets the dumping of pharmaceutical products from the surrounding industries. So, this work aimed to investigate the presence of antibiotics in Extrema River spring samples and the isolation of Staphylococcus aureus, a potential multidrug-resistant bacteria, verifying the antimicrobial resistance profile of these isolates. Three campaigns were carried out in different locals (P1–P3) between October and December 2021, in the dry and rainy seasons. The high-performance liquid chromatography-tandem mass spectrometry (LCMS) approach indicated the presence of sulfamethoxazole (≥ 1 ng/L), metronidazole (< 0.5 ng/L), and chloramphenicol (< 5 ng/L) in the water samples in November (rainy season). S. aureus was isolated in P1 (n = 128), P2 (n = 168), and P3 (n = 36), with greater resistance to trimethoprim-sulfamethoxazole (90%), clindamycin (70%), and gentamicin (60%). The presence of antibiotics in the Extrema River spring may cause S. aureus antibiotic resistance development. The presence of antibiotics and the high percentage of isolated multidrug-resistant S. aureus in the Extrema River spring cause concern and indicate the clandestine dumping of effluents from nearby pharmaceutical industries. Since preserving the springs of low-order streams is important for the environment and public health, we encourage monitoring the wastewater from Extrema River’s nearby pharmaceutical industries and preserving the spring of this river.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The Cerrado biome is the second most extensive vegetation formation on the South American continent. Its area represents one-fourth of Brazil’s entire land surface [1]. It has been considered one of the most important savannas in the world due to the richness of plant and animal species and water resources [2]. The Cerrado’s biodiversity has suffered with the expansion of Brazilian agriculture in the last 50 years, which has caused changes in its soil areas and water bodies [3, 4]. In this anthropization process, industrial activity intensified and guaranteed market in different sectors [5]. As a result, industries have discharged effluents directly into aquatic ecosystems, which alters the runoff volume and water temperature, impairing its integrity and future sustainability [6]. Water pollution has been a global concern, caused by industry and agriculture due to the significant growth of emerging pollutants predicted to increase not less than double until 2035 [7].
The city of Anápolis has 396,526 inhabitants and is located in central Brazil (Goiás State). The city’s economy revolves around the pharmaceutical, transformation, and automobile industries concentrated in the Agroindustrial District of Anápolis (DAIA). DAIA was created in 1976 and produces various types of waste [8]. Part of this industrial waste is disposed of in the Extrema River, which belongs to the Antas River watershed, and its spring is close to the DAIA. Values above the allowed for Fe and Cr and the genotoxic potential of water samples from the Extrema River have already been demonstrated [9]. Low-order rivers, like the Extrema River, are vulnerable to deforestation, agriculture, and urbanization, leading to eutrophication and hypoxia of ecosystems downstream [10, 11]. Springs are ecotones between groundwater, surface water, and the adjacent soil ecosystem, fed by a continuous flow of groundwater with a thermally stable habitat with diverse biota [12]. Springs and their terrestrial surroundings are considered hotspots for the biodiversity of the regions [13]. They are threatened by multiple anthropogenic stressors, such as capture, habitat degradation, and aquatic contaminants [14].
Antibiotics are part of persistent contaminants in aquatic environments through industrial, domestic, or veterinary disposal, whose most prolonged half-life reaches 1800 days [15, 16]. Antibiotic classes have already been recorded in rivers in China [17, 18], Canada [19], the USA [20], and Brazil [21, 22]. The indiscriminate and excessive use of antibiotics has caused an increase in multidrug-resistant bacteria [23]. The relationship between bacterial resistance and the occurrence of antibiotics in surface waters has been reported as harmful to public health, compromising the effectiveness of antimicrobial therapy, as pathogenic microorganisms are becoming resistant to most antibiotics [24]. The spread of antibiotic-resistant bacteria has been classified as one of the three threats to public health in the twenty-first century by the World Health Organization (WHO) [25].
Staphylococcus aureus is a highly resistant Gram-positive coccus to broad-spectrum antimicrobials. S. aureus has been reported to be resistant to methicillin, penicillins, cephalosporins, and carbapenems and tends to develop resistance to quinolones, aminoglycosides, and macrolides [26, 27]. S. aureus is an opportunistic pathogen that causes infections and intoxications in humans and animals, mainly in nosocomial environments [28]. However, staphylococcal infections could evolve to serious systemic infections and death, especially when associated with antimicrobial resistance (AMR) [29]. Initially, outbreaks of antimicrobial-resistant S. aureus infections were associated mainly with compromised patients exposed to hospital environments. However, since the late 90s, with the emergence of new more aggressive community-associated isolates, these infections are no longer limited to these settings. Nowadays, it is common to observe outbreaks of S. aureus infections among young healthy individuals with close contact and sharing common facilities [30].
S. aureus waterborne transmission routes have not been traditionally associated with infections. However, the potential for S. aureus to be spread via aquatic environments is continually evaluated. This microorganism has been identified in freshwater [31, 32], seawater [30, 33,34,35,36], sub-catchment water system [37], drinking water [38, 39], and wastewater [40,41,42,43,44,45]. Additionally, it has been demonstrated that people shed their colonizing organisms into seawater and, therefore, can be sources of potentially pathogenic S. aureus in recreational marine waters [30]. These observations suggest the persistence of S. aureus in the aquatic environment and the water as a possible route for S. aureus transmission.
In addition to the risk of transmission of S. aureus by water, the ESKAPE pathogens (Enterococcus faecium, S. aureus, Klebsiella pneumoniae, Acinetobacter bau-mannii, Pseudomonas aeruginosa, and Enterobacter), that cause AMR crisis faced by hospitals globally, can accumulate AMR genes primarily due to horizontal gene transfer (HGT) aided by plasmids and mobile genetic elements [46, 47]. The origins of the AMR genes are environmental bacteria, mainly soil ones, which have co-evolved with antimicrobial-producing organisms for millennia [48]. There is a temporal lag between the clinical use of a drug and the arrival of relevant mobile AMR genes in human pathogen populations [49]. So, identifying environmental reservoirs of multidrug-resistant bacteria alert to the urgent need for environmental monitoring policies [47]. Thus, the objective of this study was to investigate the presence of antibiotics in the Extrema River’s spring, due to its close proximity to many pharmaceutical industries, and to isolate S. aureus, verifying the AMR profile of these isolates.
Methods
Sampling points
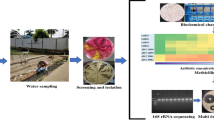
The samples were collected in surface water from the Extrema River spring in Anápolis, central Brazil (Goiás State). Three collection points near the spring were sampled (P1, P2, and P3), as follows: (P1) 16°23'8”S 48°56'38”W, (P2) 16°23'8”S 48°56'37”W, and (P3) 16°23'8”S 48°56'36”W (Fig. 1). The collection area has gallery forest phytophysiognomy, forest vegetation accompanying streams, and small river courses forming closed corridors over water courses, typical of the Brazilian Cerrado. In this region, two other phytophysiognomies are predominant, savanna stricto sensu, and dry forest [50]. Many pharmaceutical industries near the spring area (~ 5 km) have potential clandestine wastewater runoff. The region’s climate is Cwb (tropical altitude, with dry winters and hot, humid summers), according to the Köppen classification [51]. The samples were collected between October (dry season), November, and December (rainy season) in 2021. A monthly collection was carried out. The water quality parameters of the Extrema River are shown in Supplementary Information (Supp. Table S1). The water (2 liters at each point) was collected in sterile amber glass bottles, sealed with a stopper and lid to transport the material at 4 °C. In the laboratory, samples were immediately filtered for S. aureus isolation. The samples intended for high-performance liquid chromatography coupled with mass spectrometry (LCMS) were immediately stored in an ultra-freezer at −80 °C (ColdLab®, Piracicaba, SP, Brazil).
Sample preparation for antibiotic identification
Before analyzing the samples by LCMS, the samples were pretreated as follows: samples were filtered through Millex® 0.45 μm polyvinylidene fluoride (PVDF) membrane filters (Merck KGaA, Darmstadt, Germany). A total of 10 mL of each sample were first lyophilized. The powder residue was resolubilized in methanol three times at 0.5 mL with homogenization, totaling 1.5 mL. This suspension was transferred to a clean microtube and vacuum dried in the SpeedVac. After drying, it was resuspended in 100 μL of mobile phase, followed by vortexing for 5 min, centrifuging at 14000 rpm for 10 min at 8 °C. Finally, 60 μL of the supernatant was transferred to the insert and taken to the chromatographic system for injection.
High-performance liquid chromatography-tandem mass spectrometry (LCMS) conditions
Chromatographic separations were carried out using an ExionLCTM chromatographic system (Sciex, Singapore) composed of a pump with a quaternary solvent manager (AC Pump), an automatic injector (AC Autosampler), and a chromatographic column oven (AC column oven). A Kinetex C8 (150 × 4.6 mm, 5 μm particle size) column with phenomenex guard cartridges (4 × 3 mm) at 35 °C was utilized as a stationary phase. The mobile phase was a mixture of acetonitrile, methanol, and ammonium formate 10 mM containing formic acid 0.2% (5:15:80, v/v/v) on isocratic elution mode at a flow rate of 0.8 mL.min−1, being 20 min of the total chromatographic run. In the injection, the samples were kept at 15 °C, and 20 μL were injected. The tandem mass analysis was performed by a 4500 QTRAP hybrid triple quadrupole linear ion trap mass spectrometer (Sciex, Singapore) equipped with a turbo Ion Spray source. The temperature of the electrospray source was 650 °C. Mass spectrometry (MS) analysis was performed in positive and negative ion modes, 4500 V or −4500 V, respectively. Curtain gas (CUR) was 10 psi, and ion source gas pressures (GS1 and GS2) were 50 psi. MS parameters dependent on compounds include ionization mode, declustering potential (DP), collision energy (CE), and monitored ions (Table 1). These parameters were optimized by infusion of individual standard solutions of each compound at concentrations of 500 μg/L. The software used to control the system functions was Analyst® 1.7.2 (Sciex, Singapore). Only semi-quantitative identification of antibiotics was carried out in this work, that is, the presence above the limit of detection (LoD, Supp. Fig. S1). The LoD was obtained through the lowest analyte concentration that could be determined. To reach this parameter, injections of increasingly lower antibiotic concentrations were performed until a signal/noise ratio of at least 3:1 was obtained. Considering that the samples were concentrated at 100× before injection in the LCMS, the concentration of the antibiotics in the river would be 100× lower than the LoD.
Isolation of Staphylococcus aureus from surface water
Membrane filtration was used to recover S. aureus from surface water samples from the Extrema River, following the methodology of Goldstein et al. (2012) [44] with modifications. Briefly, 10 mL of each triplicate sample was vacuum filtered through a cellulose nitrate filter membrane, with a pore size of 0.45 μm and a diameter of 47 mm (GVS North America, Sanford, USA). The membranes were placed on Baird Parker agar base (KASVI®, São José dos Pinhais, PR, Brazil) and incubated at 37 °C for 24 h. After this period, the amounts of black colonies with presumptive halos of S. aureus were counted. Then, confirmatory catalase, coagulase, and gram stain tests were performed (phenotypic confirmation).
Phenotypic confirmation of Staphylococcus aureus isolates
S. aureus isolates were phenotypically confirmed by catalase- [52] and coagulase-positive (Staphclin latex kit, Laborclin®, Pinhais, PR, Brazil) [53] reactions and Gram-positive staining [54].
Selection of multidrug-resistant Staphyloccocus aureus isolates and antibiotic resistance profile
Colonies of S. aureus were selected from each sampling point and streaked on MRSA Chromogenic agar (Laborclin®, Pinhais, PR, Brazil) and incubated at 37 °C for 24 h [55]. Blue-green colonies were presumptive of MRSA (n = 20 from the three sampled points). These colonies were stored in brain heart infusion (BHI) broth (KASVI®, São José dos Pinhais, PR, Brazil) and added with 15% glycerol at −80 °C in an ultra-freezer (ColdLab®, Piracicaba, SP, Brazil). S. aureus ATCC 25923 and phosphate-buffered saline were used as positive and negative controls, respectively, for quality control and isolation process assurance. Antibiotic susceptibility tests were performed by the disk diffusion method, according to Tsai et al. (2020) [56] and CLSI M100 [57], testing nine different antibiotics (Laborclin®, Pinhais, PR, Brazil) (Supp. Table S2). Multidrug resistance was defined by an S. aureus isolate that grew in the presence of at least three different classes of antibiotics [58]. Cultures of these isolates were deposited in the Elisa F. L. C. Bailão (EFLCB) working collection (collection of microorganism cultures of the State University of Goiás [CCM-UEG], Central Campus) at the Biotechnology Laboratory (LaBiotec).
Genotypic confirmation of Staphylococcus aureus isolates by polymerase chain reaction (PCR)
DNA extraction was performed according to the manufacturer’s protocol (Zymo, CA, USA). The extracted nucleic acid was stored at −80 °C until further analysis. S. aureus nuc gene was used as a specific target using the following primers: forward 5’-GCGATTGATGGTGATACGGTT-3’ and reverse 5’-AGCCAAGCCTTGACGAACTAAAGC-3’ (amplicon = 279 bp) [59]. The following cycle conditions were used: 1× cycle for the initial denaturation step at 94 °C for 5 min; 40× cycles for the denaturation step at 94 °C for 1 min; annealing step at 54 °C for 30 s; extension step at 72 °C for 30 s; 1× cycle for final extension step at 72 °C for 10 min. The nuc amplicon was visualized in a 1.5% agarose gel. A 100 bp ladder (invitrogen by ThermoFisher Scientific, Waltham, Massachusetts, EUA) was added to compare the fragment amplified in the gel.
Statistical analysis
For the antibiotic resistance parameters of S. aureus, the mean ± standard deviation was calculated for each group using BioEstat 5.3 software [60]. For Pearson correlations between dissolved oxygen (DO) and the number of S. aureus isolates, pH and the number of S. aureus isolates, and DO and pH were used in past software 4.06b version [61].
Results
Antibiotic detection in Extrema River spring
It was possible to confirm the presence of sulfamethoxazole (≥ 1 ng/L) at P3 in the November campaign (Supp. Fig. S2). Some antibiotics are suggested to be present also in P3 in the November campaign but appear below the established LoD, metronidazole (< 0.5 ng/L), and chloramphenicol (< 5 ng/L) (Supp. Fig. S3).
Multidrug-resistant Staphylococcus aureus detection in Extrema River spring
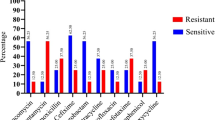
From P1, P2, and P3 samples, 128, 168, and 36 S. aureus colonies were isolated, respectively (Fig. 2). The smaller number of S. aureus isolates in P3 in the three campaigns could be correlated with the lower dissolved oxygen (DO) quantified in P3 (r = 0.870, p = 0.002) in the three campaigns. Moreover, DO and pH were positively correlated in this study (r = 0.780, p = 0.013). The isolates EFLCB 010-029 were genotypically confirmed as S. aureus since the nuc gene was amplified in all samples (Supp. Fig. S4). Of the S. aureus colonies isolated, there was a large number of multidrug-resistance to antibiotics (P1 n = 88.8%, P2 n = 33.3%, P3 = 66.6%) (Fig. 2). In the evaluation of the AMR profile, 90% of S. aureus isolates showed resistance to trimethoprim-sulfamethoxazole, 70% to clindamycin, and 60% to gentamicin. The isolates from P1 showed a high AMR profile (Table 2).
The number of Staphylococcus aureus colonies isolated by sampling point from the Extrema River spring, central Brazil. The number above the bars considers the percentage of multidrug-resistant S. aureus in each collection point. Multidrug resistance was defined by S. aureus isolate that grew in the presence of at least three different classes of antibiotics: chloramphenicol 30 μg, ciprofloxacin 5 μg, clindamycin 2 μg, erythromycin 15 μg, gentamicin 10 μg, rifampicin 5 μg, trimethoprim-sulfamethoxazole 25 μg, tetracycline 30 μg, and cefoxitin 30 μg
Discussion
Low-order streams (1st to 3rd order) dominate a riverside landscape for the function and health of the river network, being fragile ecosystems that are impacted by the discharge of effluents [62]. Improper disposal of effluents brings negative impacts to One Health. Brazil is the only country in South America with specific legislation on the ecotoxicological assessment of effluents, resolutions n° 357/2005 and n° 430/2011 of the National Council for the Environment (CONAMA) [63]. However, many Brazilian states, such as Goiás, do not have their own legislation that would complement federal legislation. In this sense, much still needs to be done to control and monitor the emission of effluents in Brazilian water resources. In this work, we first investigated the presence of antibiotics in the Extrema River spring, a low-order stream close to pharmaceutical industries (DAIA). DAIA has a sewage treatment plant inaugurated in 2003 to treat the industrial effluents from 78 industries. However, nowadays, there are more than 150 industries, and not all pre-treat effluents, which overload the system with a large volume of effluents and cause recurrent leaks [8]. In the November campaign, antibiotics in the Extrema River spring at P3 were observed, namely sulfamethoxazole (≥ 1 ng/L), metronidazole (< 0.5 ng/L), and chloramphenicol (< 5 ng/L). It is important to highlight that P3 is the nearest point from DAIA. In Cerrado, November is considered the month with more volume of rain after a 5-month drought. Maybe the pharmaceutical industries clandestinely discharge more effluents in November, relying on the dilutional property of rainfall. Additionally, the presence of antibiotics in the streams indicates the persistence of degradation, continuous discharge, input in large volumes, and incomplete removal during septic treatment [64, 65]. The half-life of sulfamethoxazole and chloramphenicol in environmental water is 3.1–30 days and 60 days, respectively [66, 67].
Pharmaceutical contaminants have already been reported in low-order streams in China [68], India [69, 70], Japan [71], France [72], the UK [73], and the USA [74]. In international studies, the most detected antibiotic is sulfamethoxazole, appearing in 96% of the analyzes. The antibiotic with the highest concentration is also sulfamethoxazole (> 20,000 ng/l), while the lowest concentration is metronidazole (1800 ng/l). Additionally, 53% of the studies use wastewater as a source, while only 18% of the papers use river water as a source. Therefore, increasing the water analysis in rivers and in other aquatic matrices is very interesting for the scientific community [75]. In Brazil, antibiotics were detected in surface water in the states of Amazonas, Maranhão (including sulfamethoxazole–22–120 ng/L) [76], Minas Gerais [77, 78], Rio Grande do Sul (including sulfamethoxazole > 1.0 ng/L [79], < 300 ng/L [22], < 200 ng/L [80]) [22, 79,80,81], and São Paulo (including sulfamethoxazole < 5 ng/L [82], 0.78–106 ng/L [21]) [21, 82, 83]. To the best of our knowledge, it is the first time that the presence of antibiotics in central Brazil freshwater has been described. However, genes encoding resistance to β-lactams, macrolides, quinolones, fluoroquinolones, tetracyclines, and sulfonamides were recently reported in another central Brazil stream (João Leite River) [84].
The most recent study by Wilkinson et al. (2022) [85], reviewed by Bouzas-Monroy et al. (2022) [86], pointed out that of 1052 river sites monitored in 104 countries, 43.5% (461 sites) contained concentrations of active pharmaceutical ingredients of concern, with approximately 34.1% of the 137 sampling campaigns having at least one site where concentrations were of ecotoxicological concern. Thus, for the conservation of freshwater springs, the Arthington document (2021) [87] points out that a more strategic, integrated, and collaborative global effort is needed between countries through the following solutions: inventory, assessment, and research of contaminated areas; ecosystem restoration and rehabilitation; design and management of protected areas; science and sociological governance. The restoration of springs and biodiversity protection depends on a reliable scientific database that evaluates the ecological results of natural processes and management actions.
The occurrence of antibiotics in the aquatic environment poses a global threat to One Health. First of all, antibiotics can cause acute and chronic toxicity toward aquatic organisms and affect the evolution of the bacterial community structure, which plays a significant role in the ecosystem [65]. Additionally, the presence of antibiotics in freshwater is of special concern because increasing concentrations of discarded antibiotics can force the appearance of some extremely multidrug-resistant strains, not just as the consequence of mutations, but also by HGT, which poses a serious public health problem [88]. S. aureus was isolated from the three sampled points (P1, P2, and P3) in the Extrema River spring. The smaller number of S. aureus isolates in P3 in the three campaigns could be correlated with the lower DO quantified in P3 in the three campaigns. It has been demonstrated that S. aureus is affected by a decrease in dissolved oxygen equal to or below 7 mg/L, because it utilizes a lot of oxygen to survive [89]. Moreover, DO and pH were positively correlated in this study. As pH decreases, hydrogen ions and oxygen react with water, which results in a decrease in the DO [90]. A total of 65% of the isolates presented resistance to at least three classes of antibiotics. Additionally, all the S. aureus isolates are resistant to trimethoprim-sulfamethoxazole in P2 (November) and P3 (December), a drug produced in DAIA and identified in the Extrema River spring, suggesting that the clandestine discharge of antibiotics from industrial effluents in the region may cause S. aureus AMR development. This data is worrying, as S. aureus is a pathogen of significant concern for hospitals and the community. Treating S. aureus infections is becoming challenging, especially considering the emergence of antibiotic-resistant strains [91].
The lack of estimates of human exposure to environmental bacteria, generally antibiotic-resistant bacteria, is a gap that makes it difficult to assess the effects of exposure. Streams and rivers can be important routes for disseminating these antibiotic-resistant bacteria, mainly because people usually use freshwater for recreational activities, ingestion, and irrigation. People have been observed using the Extrema River water for recreational purposes, which could be a source of acquiring and disseminating multidrug-resistant S. aureus strains. Some studies have pointed to the presence of AMR microorganisms and AMR genes in aquatic ecosystems impacted by human activities [25, 92,93,94]. Thus, surveillance and study of microorganisms in impacted freshwater environments can provide information on how microbial communities change because of human activities.
Conclusions
Our data indicate that the Extrema River’s spring is receiving clandestine effluents from pharmaceutical industries. So, it needs restoration and intense environmental surveillance for biodiversity and public health protection. Microbial communities contribute significantly to the quality of surface waters. Moreover, the impacts of land use and industrial effluent disposal on bacteria promote the development of multidrug-resistant strains. Thus, antibiotics in this ecosystem disrupt environmental and public health. This study encourages policies to monitor emerging pollutants in the Extrema River’s spring, which can be helpful since the safe amounts of disposal of these compounds are not yet legally regulated.
Data Availability
Data and materials not included in the manuscript are available under request.
References
IBGE. Instituto Brasileiro de Geografia e Estatística (2019) Biomas e sistema costeiro-marinho do Brasil: compatível com a escala 1:250.000. Rio de Janeiro, Coordenação de Recursos Naturais e Estudos Ambientais. (Relatórios metodológicos, v. 45). 168 p. https://biblioteca.ibge.gov.br/visualizacao/livros/liv101676.pdf
Roa F, Telles MPC (2017) The Cerrado (Brazil) plant cytogenetics database. Comparative. Cytogenetics 11(2):285–297. https://doi.org/10.3897/CompCytogen.v11i2.11395
Hunke P, Mueller EM, Schröder B, Zeilhofer P (2015) The Brazilian Cerrado: assessment of water and soil degradation in catchments under intensive agricultural use. Ecohydrology 8(6):1154–1180. https://doi.org/10.1002/eco.1573
Lemes L, Andrade AFA, Loyola R (2020) Spatial priorities for agricultural development in the Brazilian Cerrado: may economy and conservation coexist? Biodivers Conserv 29(5):1683–1700. https://doi.org/10.1007/s10531-019-01719-6
Latrubesse EM, Arima E, Ferreira ME, Nogueira SH, Wittmann F, Dias MS, Dagosta FCP, Bayer M (2019) Fostering water resource governance and conservation in the Brazilian Cerrado biome. Conserv Sci Pract 1(9):e77. https://doi.org/10.1111/csp2.77
Althoff D, Rodrigues LN, Silva DD (2021) Assessment of water availability vulnerability in the Cerrado. Appl Water Sci 11:176. https://doi.org/10.1007/s13201-021-01521-2
Zadorozhnaya O, Kirsanov D, Buzhinky I, Tsarev F, Abramova N, Bratov A, Muñoz FJ, Ribó J, Bori J, Riva MC, Legin A (2015) Water pollution monitoring by an artificial sensory system performing in terms of Vibrio fischeri bacteria. Sens Actuators B Chem 207(B):1069–1075. https://doi.org/10.1016/j.snb.2014.08.056
Fernandes CE, Barbosa Neto E, Oliveira VC, Fernandes LIFA (2020) Environmental sanitation: the challenges of the DAIA sewage treatment plant in Anápolis (GO). Brazilian Journal of Development 6(7):42426–42436. https://doi.org/10.34117/bjdv6n7-018
Bailão EFLC, Santos LAC, Almeida SS, D’Abadia PL, Morais RJ, Matos TN, Caramori SS, Araújo CST, Silva Neto CM, Almeida LM (2020) Effect of land-use pattern on the physicochemical and genotoxic properties of water in a low-order stream in Central Brazil. Revista Ambiente & Água 15(3):e2486. https://doi.org/10.4136/ambi-agua.2486
Freeman MC, Pringle CM, Jackson CR (2007) Hydrologic connectivity and the contribution of stream headwaters to ecological integrity at regional scales. JAWRA Journal of the American Water Resources Association 43(1):5–14. https://doi.org/10.1111/j.1752-1688.2007.00002.x
Ding J, Jiang Y, Liu Q, Hou Z, Liao J, Fu L, Peng Q (2016) Influences of the land use pattern on water quality in low-order streams of the Dongjiang River basin, China: a multi-scale analysis. Sci Total Environ 551-552:205–216. https://doi.org/10.1016/j.scitotenv.2016.01.162
Lehosmaa K, Muotka T, Pirttilä AN, Jaakola I, Rossi PK, Jyväsjärvi J (2021) Bacterial communities at a groundwater-surface water ecotone: gradual change or abrupt transition points along a contamination gradient? Environ Microbiol 23(11):6694–6706. https://doi.org/10.1111/1462-2920.15708
Cantonati M, Füreder L, Gerecke R, Jüttner I, Cox E (2012) Crenic habitats, hotpots for freshwater biodiversity conservation: toward an understanding of their ecology. Freshw Sci 31(2):463–480. https://doi.org/10.1899/11-111.1
Cantonati M, Fensham RJ, Stevens LE, Gerecke R, Glazier DS, Goldscheider N, Kinight N, Richardson RL, Springer JS, Tockner AE, Klement T (2021) Urgent plea for global protection of springs. Conserv Biol 35(1):378–382. https://doi.org/10.1111/COBI.13576
Ge LK, Chen JW, Wei XX, Zhang SY, Qiao XL, Cai XY, Xie Q (2010) Aquatic photochemistry of fluoroquinolone antibiotics: kinetics, pathways, and multivariate effects of main water constituents. Environ Sci Technol 44(7):2400–2405. https://doi.org/10.1021/es902852v
Guo X, Xiaojun L, Zhang A, Yan Z, Chen S, Wang N (2020) Antibiotic contamination in a typical water-rich city in southeast China: a concern for drinking water resource safety. J Environ Sci Health B 55(3):193–209. https://doi.org/10.1080/03601234.2019.1679563
Xue BM, Zhang RJ, Wang YG, Liu X, Li J, Zhang G (2013) Antibiotic contamination in a typical developing city in South China: occurrence and ecological risks in the Yongjiang River impacted by tributary discharge and anthropogenic activities. Ecotoxicol Environ Saf 92:229–236. https://doi.org/10.1016/j.ecoenv.2013.02.009
Zhang RL, Zhang RJ, Zou SC, Yang Y, Li J, Wang YG, Yu KF, Zhang G (2017) Occurrence, distribution and ecological risks of fluoroquinolone antibiotics in the Dongjiang River and the Beijiang River, Pearl River Delta, South China. Bull Environ Contam Toxicol 99:46–53. https://doi.org/10.1007/s00128-017-2107-5
Kleywegt S, Pileggi V, Yang P, Hao C, Zhao X, Rocks C, Thach S, Cheung P, Whitehea B (2011) Pharmaceuticals, hormones and bisphenol a in untreated source and finished drinking water in Ontario, Canada-occurrence and treatment efficiency. Sci Total Environ 409:1481–1488. https://doi.org/10.1016/j.scitotenv.2011.01.010
Benotti MJ, Trenholm RA, Vanderford BJ, Holady JC, Stanford BD, Snyder SA (2009) Pharmaceuticals and endocrine disrupting compounds in U.S. drinking water. Environ Sci Technol 43(3):597–603. https://doi.org/10.1021/es801845a
Locatelli MAF, Sodré FF, Jardim WF (2011) Determination of antibiotics in Brazilian surface waters using liquid chromatography–electrospray tandem mass spectrometry. Arch Environ Contam Toxicol 60(3):385–393. https://doi.org/10.1007/s00244-010-9550-1
Arsand JB, Hoff RB, Jank L, Bussamara R, Dellegraver A, Bento FM, Kmetzsch L, Falcão DA, Peralba MCR, Gomes AA, Pizzolato TM (2020) Presence of antibiotic resistance genes and its association with antibiotic occurrence in Dilúvio River in southern Brazil. Sci Total Environ 738:139781. https://doi.org/10.1016/j.scitotenv.2020.139781
Porrero MC, Harrison E, Fernández-Garayzábal JF, Paterson GK, Dízer-Guerrier A, Holmes MA, Domínguez L (2014) Detection of mecC-Methicillin-resistant Staphylococcus aureus isolates in river water: a potential role for water in the environmental dissemination. Environ Microbiol Rep 6(6):705–708. https://doi.org/10.1111/1758-2229.12191
Su M, Satola SW, Read TD (2019) Genome-based prediction of bacterial antibiotic resistance. J Clin Microbiol 57(3):e01405–e01418. https://doi.org/10.1128/jcm.01405-18
Rodriguez-Mozaz S, Chamorro S, Marti E, Huerta B, Gros M, Sànchez-Melsió A, Borrego CM, Barceló D, Balcázar L (2015) Occurrence of antibiotics and antibiotic resistance genes in hospital and urban wastewaters and their impact on the receiving river. Water Res 69:234–242. https://doi.org/10.1016/j.watres.2014.11.021
Arunkumar V, Prabagaravarthanan R, Bhaskar M (2017) Prevalence of Methicillin-resistant Staphylococcus aureus (MRSA) infections among patients admitted in critical care units in a tertiary care hospital. Int J Res Med Sci 5:2362–2366. https://doi.org/10.18203/2320-6012.ijrms20172085
Miklasińska-Majdanik M (2021) Mechanisms of resistance to macrolide antibiotics among Staphylococcus aureus. Antibiotics 10(11):1406. https://doi.org/10.3390/antibiotics10111406
Basak S, Singh P, Rajurkar M (2016) Multidrug resistant and extensively drug resistant bacteria: a study. J Pathog 2016:4065603. https://doi.org/10.1155/2016/4065603
Klevens RM, Morrison MA, Nadle J, Petit S, Gershman K, Ray S, Harrison LH, Lynfield R, Dumyati G, Townes JM, Craig AS, Zell ER, Fosheim GE, McDougal LK, Carey RB, Fridkin SK. Active bacterial core surveillance (ABCs) MRSA investigators (2007) Invasive methicillin-resistant Staphylococcus aureus infections in the United States. JAMA, 298 (15):1763-1771. https://doi.org/10.1001/jama.298.15.1763
Plano LRW, Garza AC, Shibata T, Elmir SM, Kish J, Sinigalliano CD, Gidley ML, Miller G, Withum K, Fleming LE, Solo-Gabriele HM (2011) Shedding of Staphylococcus aureus and methicillin-resistant Staphylococcus aureus from adult and pediatric bathers in marine waters. BMC Microbiol 11:5. https://doi.org/10.1186/1471-2180-11-5
Harakeh S, Yassine H, Hajjar S, El-Fadel M (2006) Isolates of Staphylococcus aureus and saprophyticus resistant to antimicrobials isolated from the Lebanese aquatic environment. Mar Pollut Bull 52(8):912–919. https://doi.org/10.1016/j.marpolbul.2005.12.008
Seyedmonir E, Yilmaz F, Icgen B (2015) mecA gene dissemination among staphylococcal and non-staphylococcal isolates shed in surface waters. Bull Environ Contam Toxicol 95(1):131–138. https://doi.org/10.1007/s00128-015-1510-z
Charoenca N, Fujioka RS (1993) Assessment of Staphylococcus bacteria in Hawaii’s marine recreational waters. Water Sci Technol 27(3-4):283–289. https://doi.org/10.2166/wst.1993.0361
Tice AD, Pombo D, Hui J, Kurano M, Bankowski MJ, Seifried SE (2010) Quantitation of Staphylococcus aureus in seawater using CHROMagar™ SA. Hawaii Med J 69(1):8–12. http://www.ncbi.nlm.nih.gov/pmc/articles/pmc3104624/
Soge OO, Meschke JS, No DB, Roberts MC (2009) Characterization of methicillin-resistant Staphylococcus aureus and methicillin-resistant coagulase-negative Staphylococcus spp. isolated from US West Coast public marine beaches. J Antimicrob Chemother 64(6):1148–1155. https://doi.org/10.1093/jac/dkp368
Thapaliya D, Hellwing EJ, Kadariya J, Grenier D, Jefferson AJ, Dalman M, Kennedy K, DiPerna M, Orihill A, Taha M, Smith TC (2017) Prevalence and characterization of Staphylococcus aureus and methicillin-resistant Staphylococcus aureus on public recreational beaches in northeast ohio. GeoHealth 1(10):320–332. https://doi.org/10.1002/2017gh000106
Azuma T, Murakami M, Sonoda Y, Ozaki A, Hayashi T (2022) Occurrence and quantitative microbial risk assessment of methicillin-resistant Staphylococcus aureus (MRSA) in a sub-catchment of the Yodo River Basin Japan. Antibiotics 11(10):1355. https://doi.org/10.3390/antibiotics11101355
Santos AV, Couto CF, Lebron YAR, Moreira VR, Foureaux AFS, Reis EO, Santos LVS, Andrade LH, Amaral MCS, Lange LC (2020) Occurrence and risk assessment of pharmaceutically active compounds in water supply systems in Brazil. Sci Total Environ 746:141011. https://doi.org/10.1016/j.scitotenv.2020.141011
Armstrong JL, Shigeno DS, Calomiris JJ, Seidler RJ (1981) Antibiotic-resistant bacteria in drinking water. Appl Environ Microbiol 42(2):277–283. https://doi.org/10.1128/aem.42.2.277-283.1981
Zieliński W, Korzeniewska E, Harnisz M, Hubeny J, Buta M, Rolbiecki D (2020) The prevalence of drug-resistant and virulent Staphylococcus spp. in a municipal wastewater treatment plant and their spread in the environment. Environ Int 143:105914. https://doi.org/10.1016/j.envint.2020.105914
Galler H, Feieler G, Petternel C, Reinthaler F, Haas D, Habib J, Kittinger C, Luxner J, Zarfel G (2018) Multiresistant bacteria isolated from activated sludge in Austria. Int J Environ Res Public Health 15(3):479. https://doi.org/10.3390/ijerph15030479
Said MB, Abbassi MS, Gómez P, Ruiz-Ripa L, Sghaier S, Ibrahim C, Torres C, Hassen A (2017) Staphylococcus aureus isolated from wastewater treatment plants in Tunisia: occurrence of human and animal associated lineages. J Water Health 15(4):638–643. https://doi.org/10.2166/wh.2017.258
Boopathy R (2017) Presence of methicillin resistant Staphylococcus aureus (MRSA) in sewage treatment plant. Bioresour Technol 240:144–148. https://doi.org/10.1016/j.biortech.2017.02.093
Goldstein RER, Micallef SA, Gibbs SG, Davis JA, He X, George A, Kleinfelter LM, Schreiber NA, Mukherjee S, Sapkota A, Joseph SW, Sapkota AR (2012) Methicillin-resistant Staphylococcus aureus (MRSA) detected at four U.S. wastewater treatment plants. Environ Health Pers 120(11):1551–1558. https://doi.org/10.1289/ehp.1205436
Börjesson S, Matussek A, Melin S, Löfgren S, Lindgren PE (2010) Methicillin-resistant Staphylococcus aureus (MRSA) in municipal wastewater: an uncharted threat? J Appl Microbiol 108(4):1244–1251. https://doi.org/10.1111/j.1365-2672.2009.04515.x
Pendleton JN, Gorman SP, Gilmore BF (2013) Clinical relevance of the ESKAPE pathogens. Expert Rev Anti Infect Ther 11(3):297–308. https://doi.org/10.1586/eri.13.12
Wyres KL, Holt KE (2018) Klebsiella pneumoniae as a key trafficker of drug resistance genes from environmental to clinically important bacteria. Curr Opin Microbiol 45:131–113. https://doi.org/10.1016/j.mib.2018.04.004
D’Costa VM, King CE, Kalan L, Morar M, Sung WWL, Schwarz C, Froese D, Zazula G, Calmels F, Debruyne R, Golding GB, Poinar HN, Wright GD (2011) Antibiotic resistance is ancient. Nature 477(7365):457–461. https://doi.org/10.1038/nature10388
Lewis K (2013) Platforms for antibiotic discovery. Nat Rev Drug Discov 12(5):371–387. https://doi.org/10.1038/nrd3975
Barreto Junior NM, Angelini R. (2003) Mapeamento Topográfico e Delimitação Fitofisionômica Da Área Natural Do Campus Da UEG (Anápolis). Mostra de Iniciação Científica da Universidade Estadual de Goiás; UEG: Anápolis, Brazil.
Cardoso MRD, Marcuzzo FFN (2014) Barros JR (2014) Classificação Climática de Köppen-Geiger Para o Estado de Goiás e o Distrito Federal|Cardoso|ACTA GEOGRÁFICA. Acta Geográfica 8:40–55. https://rigeo.cprm.gov.br/handle/doc/15047
Reiner K (2010) Catalase teste protocol. American Society for Microbiology, Washington, DC 20036. pp. 2–3
Laborclin® (2018) Staphclin latex kit. Pinhais, Paraná, Brazil. https://cdn.media.interlabdist.com.br/uploads/2021/01/570100-STAPHCLIN-LATEX-ESTAFILO-R23mL-KIT-50T-2019.pdf
Shaw K, Mazumder S (2020) Recent prevalence of clinical multidrug resistant Staphylococcus aureus in West Bengal. IOSR Journal of Dental and Medical Sciences, 19:39-44. https://www.iosrjournals.org/iosr-jdms/papers/Vol19-issue1/Series-5/H1901053944.pdf
Goodwin KD, Pobuda M (2009) Performance of CHROMagar™ Staph aureus and CHROMagar™ MRSA for detection of Staphylococcus aureus in seawater and beach sand–Comparison of culture, agglutination, and molecular analyses. Water Res 43(19):4802–4811. https://doi.org/10.1016/j.watres.2009.06.025
Tsai H-C, Tao C-W, Hsu B-M, Yang Y-Y, Tseng Y-C, Huang T-Y, Huang S-W, Kuo Y-J, Chen J-S (2020) Multidrug-resistance in methicillin-resistant Staphylococcus aureus (MRSA) isolated from a subtropical river contaminated by nearby livestock industries. Ecotoxicol Environ Saf 200:110724. https://doi.org/10.1016/j.ecoenv.2020.110724
CLSI M100. Clinical and Laboratory Standards Institute (2020) Performance standards for antimicrobial susceptibility testing, 30th ed. CLSI supplement M100. Clinical and Laboratory Standards Institute, Wayne, PA. https://clsi.org/standards/products/microbiology/documents/m100/
Magiorakos A-P, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, Giske CG, Harbarth S, Hindler JF, Imeter GK, Olsson-liljequist B, Paterson DL, Rice LB, Stelling J, Struelens MJ, Vatopoulos A, Weber JT, Monnet DL (2012) Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect 18(3):268–281. https://doi.org/10.1111/j.1469-0691.2011.03570.x
Brakstad OG, Aasbakk K, Maeland JA (1992) Detection of Staphylococcus aureus by polymerase chain reaction amplification of the nuc gene. J Clin Microbiol 30(7):1654–1660. https://doi.org/10.1128/jcm.30.7.1654-1660.1992
BioEstat. Version 5.3 (2012) Free computer program for data survey - descriptive statistical techniques. 2012. https://www2.assis.unesp.br/ffrei/bioestat.html
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological Statistics software package for education and data analysis. Paleontologia Electronica, 4(1):9pp. https://palaeo-electronica.org/2001_1/past/past.pdf
Ding S, Zhang Y, Liu B, Kong W, Meng W (2013) Effects of riparian land use on water quality and fish communities in the headwater stream of the Taizi River in China. Front Environ Sci Eng 7(5):699–708. https://doi.org/10.1007/s11783-013-0528-x
Brazil, Conselho Nacional do Meio Ambiente - CONAMA (2005) Resolução n° 357 de 17 de março de 2005. Dispõe sobre a classificação dos corpos de água e diretrizes ambientais para o seu enquadramento, bem como estabelece as condições e padrões de lançamento de efluentes, e dá outras providências. Diário Oficial [da] União: seção 1, Brasília, DF, 053:58-63. https://www.siam.mg.gov.br/sla/download.pdf?idNorma=2747
Schenck K, Rosenblum L, Ramakrishnan B Jr, Carson J, Macke D, Nietch C (2015) Correlation of trace contaminants to wastewater management practices in small watersheds. Environ Sci Process Impacts 17(5):956–964. https://doi.org/10.1039/C4EM00583J
Felis E, Kalka J, Sochaki A, Kowalska K, Bajkacz S, Harnisz M, Korzeniewska E (2020) Antimicrobial pharmaceuticals in the aquatic environment-occurrence and environmental implications. Eur J Pharmacol 866:172813. https://doi.org/10.1016/j.ejphar.2019.172813
Ma M, Dillon P, Zheng Y (2019) Determination of sulfamethoxazole degradation rate by an in situ experiment in a reducing alluvial aquifer of the North China plain. Environmental Science & Technology 53(18):10620–10628. https://doi.org/10.1021/acs.est.9b00832
Choi K, Kim Y, Jung J, Kim MH, Kim CS, Kim NH, Park J (2008) Occurrences and ecological risks of roxithromycin, trimethoprim, and chloramphenicol in the Han River, Korea. Environ Toxicol Chem: Int J 27(3):711–719. https://doi.org/10.1897/07-143.1
Liu X, Lv K, Deng C, Yu Z, Shi J, Johnson AC (2019) Persistence and migration of tetracycline, sulfonamide, fluoroquinolone, and macrolide antibiotics in streams using a simulated hydrodynamic system. Environ Pollut 252:1532–1538. https://doi.org/10.1016/j.envpol.2019.06.095
Kumar M, Ram B, Hilanda R, Poopipattana C, Canh VD, Chaminda T, Furumai H (2019) Concurrence of antibiotic resistant bacteria (ARB), viruses, pharmaceuticals and personal care products (PPCPs) in ambient waters of Guwahati, India: Urban vulnerability and resilience perspective. Sci Total Environ 693:133640. https://doi.org/10.1016/j.scitotenv.2019.133640
Gopal CM, Chat K, Ramaswamy BR, Kumar V, Singhal RK, Basu H, Udayashankar HN, Vasantharaju SG, Kumarreddy YP, Lino SY, Balakrishna K (2021) Seasonal occurrence and risk assessment of pharmaceutical and personal care products in Bengaluru rivers and lakes India. J Environ Chem Eng 9(4):105610. https://doi.org/10.1016/j.jece.2021.105610
Hanamoto S, Yamamoto-Ikemoto R (2022) In-stream sorption of azithromycin and levofloxacin in a river receiving sewage treatment plant effluent. Environ Pollut 307:119568. https://doi.org/10.1016/j.envpol.2022.119568
Feitosa-Felizzola J, Chiron S (2009) Occurrence and distribution of selected antibiotics in a small Mediterranean stream (Arc River, Southern France). J Hydrol 364(1-2):50–57. https://doi.org/10.1016/j.jhydrol.2008.10.006
Ashton D, Hilton M, Thomas KV (2004) Investigating the environmental transport of human pharmaceuticals to streams in the United Kingdom. Sci Total Environ 333(1-3):167–184. https://doi.org/10.1016/j.scitotenv.2004.04.062
Bradley PM, Journey CA, Button DT, Carlisle DM, Huffman BJ, et al (2020) Multi-region assessment of pharmaceutical exposures and predicted effects in USA wadeable urbangradient streams. PLOS ONE 15(1):e0228214. https://doi.org/10.1371/journal.pone.0228214
Ilurdoz MS, Sadhwani JJ, Reboso JV (2022) Antibiotic removal processes from water & wastewater for the protection of the aquatic environment - a review. J Water Process Eng 45:102474. https://doi.org/10.1016/j.jwpe.2021.102474
Chaves MJS, Barbosa SC, Malinowski MM, Volpato D, Castro ÍB, Franco TCRS, Primel EG (2020) Pharmaceuticals and personal care products in a Brazilian wetland of international importance: occurrence and environmental risk assessment. Sci Total Environ 734:139374. https://doi.org/10.1016/j.scitotenv.2020.139374
Reis EO, Foureaux AFS, Rodrigues JS, Moreira VR, Lebron YAR, Santos LVS, Amaral MCS, Lange LC (2019) Occurrence, removal and seasonal variation of pharmaceuticals in Brasilian drinking water treatment plants. Environ Pollut 250:773–781. https://doi.org/10.1016/j.envpol.2019.04.102
Barros ALC, Schmidt FF, Aquino SF, Afonso RJCF (2018) Determination of nine pharmaceutical active compounds in surface waters from Paraopeba River Basin in Brazil by LTPE-HPLC-ESI-MS/MS. Environ Sci Pollut Res Int 25:19962–19974. https://doi.org/10.1007/s11356-018-2123-y
Arsand JB, Hoff RB, Jank L, Dallegrave A, Galeazzi C, Barreto F, Pizzolato TM (2018) Wide-scope determination of pharmaceuticals and pesticides in water samples: qualitative and confirmatory screening method using LC-qTOF-MS. Water, Air, & Soil Pollut 229:399. https://doi.org/10.1007/s11270-018-4036-2
Perin M, Dallegrave A, Barnet LS, Meneghini LZ, Gomes AA, Pizzolato TM (2021) Pharmaceuticals, pesticides and metals/metalloids in Lake Guaíba in Southern Brazil: Spatial and temporal evaluation and a chemometrics approach. Sci Total Environ 793:148561. https://doi.org/10.1016/j.scitotenv.2021.148561
Caldas SS, Rombaldi C, Oliveira Arias JL, Marube LC, Primel EG (2016) Multiresidue method for determination of 58 pesticides, pharmaceuticals and personal care products in water using solvent demulsification dispersive liquid-liquid microextraction combined with liquid chromatography-tandem mass spectrometry. Talanta 146:676–688. https://doi.org/10.1016/j.talanta.2015.06.047
Montagner CC, Sodré FF, Acayaba RD, Vidal C, Campestrini I, Locatelli MA, Pescara IC, Albuquerque AF, Umbuzeiro GA, Jardim WF (2019) Ten years-snapshot of the occurrence of emerging contaminants in drinking, surface and ground waters and wastewaters from São Paulo State, Brazil. J Braz Chem Soc 30:614–632. https://doi.org/10.21577/0103-5053.20180232
Pivetta RC, Rodrigues-Silva C, Ribeiro AR, Rath S (2020) Tracking the occurrence of psychotropic pharmaceuticals in Brazilian wastewater treatment plants and surface water, with assessment of environmental risks. Sci. Total Environ 727:138661. https://doi.org/10.1016/j.scitotenv.2020.138661
Gomes RP, Oliveira TR, Gama AR, Vieira JDG, Rocha TL, Carneiro LC (2022) Gene resistance profile and multidrug-resistant bacteria isolated from a stream in midwestern Brazil. Environ Nanotechnol Monit Manag 18:100688. https://doi.org/10.1016/j.enmm.2022.100688
Wilkinson JL, Boxall ABA, Kolpin DW, Leung KMY, Lai RWS, Galbán-Malagón C, Adell AD, Mondon J, Metian M, Marchant RA, Bouzas-Monroy A, Cuni-Sanchez A, Coors A, Carriquiriborde P, Rojo M, Gordon C, Cara M, Moermond M, Luarte T, Teta C (2022) Pharmaceutical pollution of the world’s rivers. Proc Natl Acad Sci U S A 119(8):e2113947119. https://doi.org/10.1073/pnas.2113947119
Bouzas-Monroy A, Wilkinson JL, Melling M, Boxall ABA (2022) Assessment of the potential ecotoxicological effects of pharmaceuticals in the world’s rivers. Environ Toxicol Chem 1-13. https://doi.org/10.1002/etc.5355
Arthington AH (2021) Grand challenges to support the freshwater biodiversity emergency recovery plan. Front Environ Sci 9:664313. https://doi.org/10.3389/fenvs.2021.664313
Jendrzejewska N, Karwowska E (2018) The influence of antibiotics on wastewater treatment processes and the development of antibiotic-resistant bacteria. Water Sci Technol 77(9):2320–2326. https://doi.org/10.2166/wst.2018.153
Mabonga H (2021) Isolation of Escherichia coli and Staphylococcus aureus in surface water sources in Katabi Subcounty, Wakiso District. Student's J of Health Res Africa 2(3):9. https://doi.org/10.51168/sjhrafrica.v2i3.20
Zang C, Huang S, Wu M, Du S, Scholz M, Gao F, Lin C, Guo Y, Dong Y (2011) Comparison of relationships between pH, dissolved oxygen and chlorophyll a for aquaculture and non-aquaculture waters. Water, Air, & Soil Pollut 219:157–174. https://doi.org/10.1007/s11270-010-0695-3
Bravo-Santano N, Behrends V, Letek M (2019) Host-targeted therapeutics against multidrug resistant intracellular Staphylococcus aureus. Antibiotics 8(4):241. https://doi.org/10.3390/antibiotics8040241
Roberto AA, Gray JBV, Engohang-Ndong J, Leff LG (2019) Distribution and co-occurrence of antibiotic and metal resistance genes in biofilms of an anthropogenically impacted stream. Sci Total Environ 688:437–449. https://doi.org/10.1016/j.scitotenv.2019.06.053
Yi X, Lin C, Ong EJL, Wang M, Li B, Zhou Z (2019) Expression of resistance genes instead of gene abundance are correlated with trace levels of antibiotics in urban surface waters. Environ Pollut 250:437–446. https://doi.org/10.1016/j.envpol.2019.04.035
Zhang H, He H, Chen S, Huang T, Lu K, Zhang Z, Wang R, Zhang X, Li H (2019) Abundance of antibiotic resistance genes and their association with bacterial communities in activated sludge of wastewater treatment plants: geographical distribution and network analysis. J Environ Sci 82:24–38. https://doi.org/10.1016/j.jes.2019.02.023
Acknowledgements
We would like to thank Dr. Mirelle Garcia Silva Bailão and Dr. Leonardo Luiz Borges for supporting this research.
Funding
This research was funded by Programa Pesquisa para o SUS: gestão compartilhada em saúde—PPSUS (grant number PPSUS proc. 202110267000295), namely Ministério da Saúde (MS), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Fundação de Amparo à Pesquisa do Estado de Goiás (FAPEG), and Secretaria de Estado da Saúde (SES-GO); and by Universidade Estadual de Goiás (UEG; Pró-Projetos, grant number 202100020012836). IRS and INMS were supported by scholarships from UEG.
Author information
Authors and Affiliations
Contributions
Material preparation, data collection, and analysis were performed by Igor Romeiro dos Santos, Isabela Náthaly Machado da Silva, Jerônimo Raimundo de Oliveira Neto, Naiara Raica Lopes de Oliveira, Adriano Roberto Vieira de Sousa, Anielly Monteiro de Melo, Joelma Abadia Marciano de Paula, Luiz Carlos da Cunha, and Elisa Flávia Luiz Cardoso Bailão. The study conception and design were performed by Igor Romeiro dos Santos, Joelma Abadia Marciano de Paula, Cátia Lira do Amaral, Elisângela de Paula Silveira-Lacerda, Luiz Carlos da Cunha, and Elisa Flávia Luiz Cardoso Bailão. The first draft of the manuscript was written by Igor Romeiro dos Santos and Elisa Flávia Luiz Cardoso Bailão, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
The author hereby transfers to the publisher the copyright of the work.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Lucy Seldin
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
dos Santos, I.R., da Silva, I.N.M., de Oliveira Neto, J.R. et al. The presence of antibiotics and multidrug-resistant Staphylococcus aureus reservoir in a low-order stream spring in central Brazil. Braz J Microbiol 54, 997–1007 (2023). https://doi.org/10.1007/s42770-023-00973-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-00973-9