Abstract
Klebsiella pneumoniae belongs to Enterobacteriaceae, which is the commonest bacterium causing nosocomial respiratory tract infection. It ranks second in bacteremia and urinary tract infection in gram-negative bacteria. Therefore, the rapid and accurate identification of K. pneumoniae was of great significance for the guide of clinical medication, and timely treatment of patients. The purpose of this study was to establish a rapid and sensitive molecular detection method for K. pneumoniae based on loop-mediated isothermal amplification (LAMP) technology. Firstly, local BLAST and NCBI BLAST were used to analyze the genome of K. pneumoniae. According to the principle of interspecific and intraspecific specificity, CelB (GenBank ID 11847805) was selected as the specific gene. Then, the LAMP and PCR identification systems were established with this target gene. Thirty-six clinical isolates of K. pneumoniae and 50 non-K. pneumoniae were used for the specific evaluation, and both LAMP and PCR could specifically distinguish K. pneumoniae from non-K. pneumoniae. A 10-fold series diluted positive plasmids and simulated infected blood samples were used as the templates in the sensitivity assay, and the results showed that the sensitivity could reach 1 copy/reaction. In summary, a rapid, specific, and sensitive LAMP method was established to detect K. pneumoniae in clinics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
K. pneumoniae was widely distributed in nature, which was one of the normal microorganisms in the intestine [1]. As a conditional pathogen, it belongs to gram-negative bacteria. When the body’s immunity declines, it enters the lung through the respiratory tract and causes the fusion of large lobes or small lobes, and finally caused pneumonia [2, 3]. It also ranks second in bacteremia and urinary tract infection in gram-negative bacteria, which is a great threat to people’s health [4,5,6].
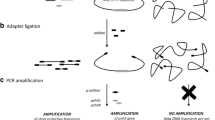
At present, traditional methods were used for bacterial identification and drug sensitivity identification in many hospitals, including microscopy, disk diffusion method, biochemistry, serotype, and antibiotic dilution method (minimal inhibitory concentration method) [7]. Although it has the advantages of easy operation and low cost, it was time-consuming and has low accuracy. With the rapid development of bioinformatics and molecular biology, conserved nucleic acid sequences with interspecies-specific and intraspecies commonality in the pathogens have been discovered one after another [8]. Based on this, an army of detection techniques has been established, such as the polymerase chain amplification (PCR), real-time fluorescent PCR, loop-mediated isothermal amplification (LAMP) [9,10,11]. The specific DNA fragments were amplified in vitro by PCR technology was developed in 1985 by Mullis et al. [12]. In 2000, Notomi et al. developed a novel, constant-temperature, enzyme-based nucleic acid amplification technique, and named it LAMP, which was a viable and cost-effective alternative to molecular diagnostics [13]. This technique was based on a set of specific primers (at least four primers, two internal primers (BIP and FIP) and two external primers (B3 and F3)) to recognize six different regions of the DNA at constant temperature (60 °C to 65 °C) [13]. Complementary strand synthesis was initiated base on the target DNA region, produce the mixture of stem-loop DNA with various fragment sizes and shapes, and the by-product was white pyrophosphate ion precipitated [14, 15]. Thus, the LAMP could be used in the POCT (point-of-care testing) with the advantages of rapidity, specificity, sensitivity, simplicity, and low cost of operation [16,17,18] (Table 1).
In this study, the clinical strains of K. pneumoniae were used as the research object to establish a rapid LAMP detection method. It can make up the deficiency of traditional detection methods by accurately detect patients infected with pathogens in a timely and accurate manner during clinical practice, and guide the use of drugs promptly and accurately.
Materials and methods
Strain culturing and the extraction of genomic DNA
Thirty-six clinical isolates of K. pneumoniae and fifty (5 species) other non-K. pneumoniae clinical isolates including Escherichia coli (15), Pseudomonas aeruginosa (7), Staphylococcus epidermidis (10), Staphylococcus aureus (8), and Acinetobacter baumannii (10) were collected from March 2018 to July 2018 and kindly provided by the laboratory of the First People’s Hospital of Yunnan Province. The above strains were identified by routine isolation culture, gram staining, and mass spectrometry (VITEK®MS) [19, 20]. All the strains were inoculated in LB liquid medium, in 37 °C, 180 rpm for 12 h, and preserved for later use. The bacterial genomic DNA kit (Zomanbio, China) was used to extract the genome DNA from bacteria. The extracted genome DNA was stored at − 20 °C for further use.
Specific gene screening
The specific gene of K. pneumoniae was screened similarity as the previous study [21, 22]. Briefly, the formatted non-redundant nucleic acid database was downloaded from the National Center for Biotechnology Information (NCBI) as the local database. The local BLAST was performed using K. pneumoniae subsp. pneumoniae HS11286 (GenBank No. NC_016845.1) genome as a query sequence against the above-mentioned database. Potential specific genes were further analyzed by NCBI BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PROGRAM=blastn) with the database excluding or including the sequences of K. pneumoniae. The specific gene must be highly conserved among the K. pneumoniae strains and show no significant similarity to other species.
Primer designing
After the specific gene screening, CelB (ID: 11847805) as the target gene was finally obtained. Four oligonucleotide primers (outer and inner primers, F3/B3 and FIP/BIP, respectively) used for the LAMP reaction were designed by Primer Explorer version 5 software (http://primerexplorer.jp/lampv5/index.html). The outer primers (B3/F3) were also used in the PCR assays, all the primers were synthesized by TSINGKE Biological Technology Co., Ltd. (Kunming, China).
Establishment of LAMP and PCR reactions
The genomic DNA of K. pneumoniae as a template was subjected to the LAMP and PCR amplification. The LAMP reaction system consisting of 12.5 μL of 2 × isothermal Mastermix (pH 8.8, with SYBR green I as the fluorescence dye to combine with the minor groove of the DNA double helix), 8 μM FIP and BIP (4 μL), 1 μM B3 and F3 (0.5 μL), 100 ng of genomic DNA, and finally add sterilizing water up to 25 μL. The reaction was carried out in Genie®II with a process of incubation at 65 °C for 30 min and melting curve analysis at 98 °C to 80 °C. The PCR reaction was carried out in a 25 μL system, including 12.5 μL 2 × TSINGKE Master Mix (containing 1 U DNA polymerase, 1.5 mM MgCl2, 200 μM dNTP), 10 μM B3 and F3, 50 ng of genomic DNA. The PCR reaction process was pre-denaturation at 95 °C for 5 min, with 35 cycles including denaturation at 95 °C for 30 s, annealing for 30 s, extension at 72 °C, and a final extension at 72 °C for 5 min. 5 μL of the PCR products were used in the 2% agarose gel electrophoresis at 120 V, 30 min, and the agarose gel was stained by Gel stain (Beijing TRANSGEN BIOTECH Co., Ltd.).
Specificity evaluation of LAMP and PCR reactions
Thirty-six clinical isolates of K. pneumoniae and 5 other non-K. pneumoniae clinical isolates (E. coli, P. aeruginosa, S. epidermidis, S. aureus, and A. baumannii) were used for the specific evaluation of LAMP and PCR reactions. The PCR test was used as the gold standard in preliminary experiments for the specific test prior to the LAMP test. All experiments were repeated twice.
Sensitivity evaluation of LAMP and PCR reactions
The sensitivity of LAMP and PCR reactions were evaluated using two different templates, the serially diluted 10-fold positive plasmids (108–100 copies) (constructed as the previous study [21]) and blood sample mimicking infection. The counted K. pneumoniae strain was serially diluted by 10-fold in PBS and then mixed with blood from the healthy specific-pathogen-free female Kunming mice (Laboratory Animal Center of Kunming Medical University, Kunming, Yunnan; weight, 22–25 g; age, 5 weeks) at 1:1 proportion to mimic infection. The mice were housed in the animal experiment center of Kunming University of Science and Technology at 25 °C with a 12 h/12 h light/dark cycle and access to food and water ad libitum. The blood sample was lysed by a solution containing 125 mM NaOH, 1 mM EDTA, and 0.1% Tween 20, and then a solution containing 125 mM HCl and 10 mM Tris-HCl, and the suspension was used as the template for LAMP and PCR assays. All experiments were repeated 2 times.
Results
Screening the specific gene and primer designing
Through local BLAST, 700 potential specific genes were obtained. Conclusively, 4 potential specific genes were obtained after NCBI BLAST, which met the criterion about the interspecific specificity and intraspecies universality. Then, the primers for LAMP reaction were designed on the Primer Explorer V5 software (http://primerexplorer.jp/lampv5/index.html). In this step, four primers were designed for the six regions of the target gene in the LAMP, and the primers should be confirmed by the primer BLAST and PCR assay. Considering the above harsh selection conditions, the CelB gene (GenBank ID 11847805) was finally identified as the only one that can be used in the detection of K. pneumoniae. The CelB gene was involved in the encoding of cellobiose-specific PTS family enzyme IIC component (YP_005227087.1), which participated in the metabolism of K. pneumoniae, and could be as a new target for the identification of K. pneumoniae.
Specificity of LAMP and PCR reactions
Thirty strains of K. pneumoniae and 50 other non-K. pneumoniae clinical isolates were used for the specificity evaluation. Before the LAMP reaction, PCR test was used for the specific test. All the K. pneumoniae were positive, while all the non-K. pneumoniae were negative, the partial results were shown in Fig. 1A. The LAMP results including the amplification curve, melting curve, and the fluorescence visualization were consistent with the PCR, which was shown in Fig. 2 A–C.
Specificity and sensitivity of the PCR assay for detecting the target gene CelB using the primers B3/F3 (a–c). a Specificity of the PCR assay for detecting the target gene of CelB. Genomic DNA of K. pneumoniae (14A0882, 14A0794, 13A14310, 13A14525, 13A13753, 13A13943) was used as the template for PCR in lane 1 to lane 6, the template in lane 7 to lane 11 were E. coli (1611NY0004), P. aeruginosa (0007280366), S. epidermidis (14B13012), S. aureus (15A14046). Negative control (NC), with the sterile distilled water as the template in lane 12. b Sensitivity of the PCR assay for detecting the target gene of CelB in the positive plasmids. The positive plasmids of K. pneumoniae and they were serially diluted 10-fold as templates for PCR assay. The lane 1 to 9 were 108–100, lane 10 was the negative control. c Sensitivity of the PCR assay for detecting the target gene of CelB in the bacterial solutions. The bacterial solutions were serially diluted 10-fold with the mice blood as a volume ratio of 1:1. These mixtures were lysed and then the suspension was used for PCR assay. The lane 1 to 8 were 107–100, lane 9 was the negative control. M: 2000 marker (Takara Biomedical Technology (Beijing) Co., Ltd). All the experiments were repeated three times
Specificity and sensitivity of the LAMP assay for detecting the target gene CelB using the primers B3/F3, FIP/BIP (a–f). (a–c) Specificity of the LAMP assay for detecting the target gene of CelB by Genie®II. Genomic DNA of K. pneumoniae (14A0882, 14A0794, 13A14310, 13A14525) and four non-K. pneumoniae (E. coli (1611NY0004), P. aeruginosa (0007280366), S. epidermidis (14B13012), S. aureus (15A14046)) were used as the template for LAMP test. a the amplification curve, b melting curve, and the fluorescence visualization c. (d–f) sensitivity of the LAMP assay for detecting the target gene of CelB by Genie®II. Gradient diluted bacterial solution of K. pneumoniae (105–100) was mixed with the mice blood as a volume ratio of 1:1. These mixtures were lysed and then the suspension was used for LAMP assay, positive plasmid (PC) as the positive control, negative control (NC), with the sterile distilled water as the template. d The amplification curve, e melting curve, and the fluorescence visualization f. All the experiments were repeated three times
Sensitivity of LAMP and PCR reactions
After calculation, the initial concentration of the positive plasmid was 2.13 × 108 copies/μL, which was first diluted to 1 × 108 copies/μL, then subjected to 10-fold serial dilution, diluted from 108 to 100. One microliter of each gradient concentration of the recombinant plasmid was used as the template for the sensitivity detection of the PCR reaction. Take a 10-fold gradient of K. pneumoniae with an initial concentration of 1.05 × 1010 CFU/mL, dilute from 108 to 101, 2 μL of each gradient of the bacterial solution to make a blood sample, lysed by heat. Two microliters each gradient concentration (107 to 100 for PCR, 105 to 100 for LAMP, positive plasmid (1 × 106 copies/μL) as the positive control for LAMP) of the blood sample lysate was used as a template for sensitivity detection of the LAMP and PCR reaction. As can be seen from Figs. 1 B and C and 2 D–F, both the positive plasmid and the blood sample were used as templates, the detectability of LAMP and PCR reaction can achieve 1 copy/reaction.
Discussion
In China, the infection rate of K. pneumoniae has become second only to that of Escherichia coli [5]. The World Health Organization reported in 2013 that diarrhea caused by K. pneumoniae was the main reason for the 29% death (Two million children each year) of global children [23]. K. pneumoniae was reported to account for 9.03% of total bacterial infection in hospitals [24, 25]. Faced with such a severe situation, the rapid and accurate diagnostic methods are of great significance for drug use.
The main methods for pathogen detection were including the culture method, routine smear microscopic examination, and immunological assay in clinical testing. With the rapid development of molecular biology technology, PCR technology was considered to be a good choice in addition to the above techniques, and has been more commonly applied to pathogenic microorganism detection. Compared with traditional methods, PCR technology has the advantages of high specificity, sensitivity, and rapidity [26, 27]. However, PCR requires expensive thermal cycling equipment, and finally requires agarose gel electrophoresis to further verify the detection results, which was time-consuming and inapplicable. Since 2000, LAMP technology has been widely used for the detection of pathogenic bacteria, including bacteria, fungi, viruses, and parasites, due to its rapidity, simplicity, and high sensitivity [16, 18, 28]. In this study, LAMP combining PCR technology was established for the fast identification.
After local BLAST and NCBI BLAST, the CelB gene was finally obtained as the target gene for the detection of the K. pneumoniae. After the extensive literature review, we found that this gene had been annotated a few years ago, but it has never been used for the identification of K. pneumoniae. In this study, this gene was used in the identification for the first time. In the results, it has been found that this gene can successfully identify K. pneumoniae.
LAMP provides an alternative to PCR-based assays; due to its isothermal nature, it may be more suitable for the detection of pathogenic bacteria in the front-line or mobile laboratories in developing countries. LAMP technology was at least 10 times more sensitive than PCR technology [29,30,31]. But in this study, the sensitivity of PCR was same to the LAMP by optimizing the PCR reaction system, which suggested that there was still room for improvement in the sensitivity of PCR.
This study established a rapid LAMP detection system for K. pneumoniae. PCR and LAMP technology was used to detect 86 strains of pathogens (the 50 non-K. pneumoniae including the commonest multi-drug-resistant bacteria and the most likely to cause contamination bacteria); the positive rate was 100% and no false-positive results were found, indicating that the detection method with better specificity. The detectability of LAMP can reach 1 copy/reaction, which proved to be highly sensitive. The LAMP detection system established in this study laid the foundation for subsequent research.
References
Khaertynov KS, Anokhin VA, Rizvanov AA, Davidyuk YN, Semyenova DR, Lubin SA, Skvortsova NN (2018) Virulence factors and antibiotic resistance of Klebsiella pneumoniae strains isolated from neonates with sepsis. Front Med (Lausanne) 5:225. https://doi.org/10.3389/fmed.2018.00225
Lin YT, Tseng KY, Yeh YC, Yang FC, Fung CP, Chen NJ (2014) TREM-1 promotes survival during Klebsiella pneumoniae liver abscess in mice. Infect Immun 82(3):1335–1342. https://doi.org/10.1128/IAI.01347-13
Rock C, Thom KA, Masnick M, Johnson JK, Harris AD, Morgan DJ (2014) Frequency of Klebsiella pneumoniae carbapenemase (KPC)-producing and non-KPC-producing Klebsiella species contamination of healthcare workers and the environment. Infect Control Hosp Epidemiol 35(4):426–429. https://doi.org/10.1086/675598
Kobayashi SD, Porter AR, Freedman B, Pandey R, Chen L, Kreiswirth BN, DeLeo FR (2018) Antibody-mediated killing of carbapenem-resistant ST258 Klebsiella pneumoniae by human neutrophils. mBio 9(2). https://doi.org/10.1128/mBio.00297-18
Lin WH, Tseng CC, Wu AB, Yang DC, Cheng SW, Wang MC, Wu JJ (2015) Clinical and microbiological characteristics of peritoneal dialysis-related peritonitis caused by Klebsiella pneumoniae in southern Taiwan. J Microbiol Immunol Infect 48(3):276–283. https://doi.org/10.1016/j.jmii.2013.10.002
Long SW, Olsen RJ, Eagar TN, Beres SB, Zhao P, Davis JJ, Brettin T, Xia F, Musser JM (2017) Population genomic analysis of 1,777 extended-Spectrum Beta-lactamase-producing Klebsiella pneumoniae isolates, Houston, Texas: unexpected abundance of clonal group 307. mBio 8(3). https://doi.org/10.1128/mBio.00489-17
Bachman MA, Breen P, Deornellas V, Mu Q, Zhao L, Wu W, Cavalcoli JD, Mobley HL (2015) Genome-wide identification of Klebsiella pneumoniae fitness genes during lung infection. mBio 6(3):e00775. https://doi.org/10.1128/mBio.00775-15
Barreda-Garcia S, Miranda-Castro R, de-Los-Santos-Alvarez N, Miranda-Ordieres AJ, Lobo-Castanon MJ (2018) Helicase-dependent isothermal amplification: a novel tool in the development of molecular-based analytical systems for rapid pathogen detection. Anal Bioanal Chem 410(3):679–693. https://doi.org/10.1007/s00216-017-0620-3
Mekata T, Kono T, Savan R, Sakai M, Kasornchandra J, Yoshida T, Itami T (2006) Detection of yellow head virus in shrimp by loop-mediated isothermal amplification (LAMP). J Virol Methods 135(2):151–156. https://doi.org/10.1016/j.jviromet.2006.02.012
Meserve D, Wang Z, Zhang DD, Wong PK (2008) A double-stranded molecular probe for homogeneous nucleic acid analysis. Analyst 133(8):1013–1019. https://doi.org/10.1039/b804853c
Zhu G, Ye X, Dong Z, Lu YC, Sun Y, Liu Y, McCormack R, Gu Y, Liu X (2015) Highly sensitive droplet digital PCR method for detection of EGFR-activating mutations in plasma cell-free DNA from patients with advanced non-small cell lung cancer. J Mol Diagn 17(3):265–272. https://doi.org/10.1016/j.jmoldx.2015.01.004
Mullis K, Faloona F, Scharf S, Saiki R, Horn G, Erlich H (1992) Specific enzymatic amplification of DNA in vitro: the polymerase chain reaction. 1986. Biotechnology 24:17–27
Notomi T, Okayama H, Masubuchi H, Yonekawa T, Watanabe K, Amino N, Hase T (2000) Loop-mediated isothermal amplification of DNA. Nucleic Acids Res 28(12):E63
Caipang CM, Haraguchi I, Ohira T, Hirono I, Aoki T (2004) Rapid detection of a fish iridovirus using loop-mediated isothermal amplification (LAMP). J Virol Methods 121(2):155–161. https://doi.org/10.1016/j.jviromet.2004.06.011
Kang SI, Her M, Kim JY, Lee JJ, Lee K, Sung SR, Jung SC (2015) Rapid and specific identification of Brucella abortus using the loop-mediated isothermal amplification (LAMP) assay. Comp Immunol Microbiol Infect Dis 40:1–6. https://doi.org/10.1016/j.cimid.2015.03.001
Ptaszynska AA, Borsuk G, Wozniakowski G, Gnat S, Malek W (2014) Loop-mediated isothermal amplification (LAMP) assays for rapid detection and differentiation of Nosema apis and N. ceranae in honeybees. FEMS Microbiol Lett 357(1):40–48. https://doi.org/10.1111/1574-6968.12521
Rodel J, Bohnert JA, Stoll S, Wassill L, Edel B, Karrasch M, Loffler B, Pfister W (2017) Evaluation of loop-mediated isothermal amplification for the rapid identification of bacteria and resistance determinants in positive blood cultures. Eur J Clin Microbiol Infect Dis 36(6):1033–1040. https://doi.org/10.1007/s10096-016-2888-1
Soliman H, Saleh M, El-Matbouli M (2015) Detection of fish pathogens by loop-mediated isothermal amplification (LAMP) technique. Methods in Molecular Biology (Clifton, NJ) 1247:163–173. https://doi.org/10.1007/978-1-4939-2004-4_12
Martins KB, Ferreira AM, Mondelli AL, Rocchetti TT, Lr d SCM (2018) Evaluation of MALDI-TOF VITEK((R))MS and VITEK((R)) 2 system for the identification of Staphylococcus saprophyticus. Future Microbiol 13:1603–1609. https://doi.org/10.2217/fmb-2018-0195
Sarier M, Sepin N, Duman I, Demir M, Hizel A, Goktas S, Emek M, Kukul E, Soylu A (2018) Microscopy of Gram-stained urethral smear in the diagnosis of urethritis: which threshold value should be selected? Andrologia 50(10):e13143. https://doi.org/10.1111/and.13143
Li C, Shi Y, Yang G, X-s X, Mao X, Fang Y, Zhang A-M, Song Y (2019) Establishment of loop-mediated isothermal amplification for rapid detection of Pseudomonas aeruginosa. Experimental and Therapeutic Medicine 17:6. https://doi.org/10.3892/etm.2018.6910
Li C, Shi YQ, Yang GY, Xia XS, Mao XQ, Fang Y, Zhang AM, Song YZ (2019) Loop-mediated isothermal amplification assay for rapid detection of hypothetical protein gene in Escherichia coli clinical isolates. Clin Lab 65(4). https://doi.org/10.7754/Clin.Lab.2018.180826
Ferkol T, Schraufnagel D (2014) The global burden of respiratory disease. Annals of the American Thoracic Society 11(3):404–406. https://doi.org/10.1513/AnnalsATS.201311-405PS
Miriagou V, Cornaglia G, Edelstein M, Galani I, Giske CG, Gniadkowski M, Malamou-Lada E, Martinez-Martinez L, Navarro F, Nordmann P, Peixe L, Pournaras S, Rossolini GM, Tsakris A, Vatopoulos A, Canton R (2010) Acquired carbapenemases in gram-negative bacterial pathogens: detection and surveillance issues. Clinical Microbiology and Infection: the Official Publication of the European Society of Clinical Microbiology and Infectious Diseases 16(2):112–122. https://doi.org/10.1111/j.1469-0691.2009.03116.x
Queenan AM, Bush K (2007) Carbapenemases: the versatile beta-lactamases. Clin Microbiol Rev 20(3):440–458, table of contents. https://doi.org/10.1128/CMR.00001-07
Bharathi MJ, Murugan N, Rameshkumar G, Ramakrishnan R, Venugopal Reddy YC, Shivkumar C, Ramesh S (2013) Comparative evaluation of uniplex, nested, semi-nested, multiplex and nested multiplex PCR methods in the identification of microbial etiology of clinically suspected infectious endophthalmitis. Curr Eye Res 38(5):550–562. https://doi.org/10.3109/02713683.2013.772205
Teranishi H, Ohzono N, Inamura N, Kato A, Wakabayashi T, Akaike H, Terada K, Ouchi K (2015) Detection of bacteria and fungi in blood of patients with febrile neutropenia by real-time PCR with universal primers and probes. J Infect Chemother 21(3):189–193. https://doi.org/10.1016/j.jiac.2014.11.008
Ohtsuka K, Yanagawa K, Takatori K, Hara-Kudo Y (2005) Detection of Salmonella enterica in naturally contaminated liquid eggs by loop-mediated isothermal amplification, and characterization of Salmonella isolates. Appl Environ Microbiol 71(11):6730–6735. https://doi.org/10.1128/AEM.71.11.6730-6735.2005
Abdoli A, Dalimi A, Soltanghoraee H, Ghaffarifar F (2016) Molecular detection of toxoplasma gondii in house sparrow (Passer domesticus) by LAMP and PCR methods in Tehran, Iran. J Parasit Dis 40(4):1317–1321. https://doi.org/10.1007/s12639-015-0680-2
Dong D, Liu W, Li H, Wang Y, Li X, Zou D, Yang Z, Huang S, Zhou D, Huang L, Yuan J (2015) Survey and rapid detection of Klebsiella pneumoniae in clinical samples targeting the rcsA gene in Beijing, China. Front Microbiol 6:519. https://doi.org/10.3389/fmicb.2015.00519
Kitamura M, Aragane M, Nakamura K, Watanabe K, Sasaki Y (2017) Rapid identification of drug-type strains in Cannabis sativa using loop-mediated isothermal amplification assay. J Nat Med 71(1):86–95. https://doi.org/10.1007/s11418-016-1031-z
Acknowledgments
The authors would like to express their sincere gratitude to Mr. Yaoqiang Shi for his help in the language of the article.
Funding
This work was supported by grants from Yunnan Science and Technology Commission (2015BC001 and 2015DH010).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Responsible Editor: Elizabeth Andrade Marques.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tian, Y., Wang, L., Zhang, J. et al. CelB is a suitable marker for rapid and specific identification of Klebsiella pneumoniae by the loop-mediated isothermal amplification (LAMP) assay. Braz J Microbiol 50, 961–967 (2019). https://doi.org/10.1007/s42770-019-00144-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-019-00144-9