Abstract
This study may enhance our understanding of AvFLS function in plants under salt stress and may provide a new tool for the improvement of plant salt tolerance in the field. We isolated and identified AvFLS from Apocynum venetum. To further characterize the potential role of AvFLS in salt tolerance and to explore the relationship between FLS and salt resistance, we generated transgenic Arabidopsis lines overexpressing AvFLS. Tissue-specific expression analysis and salt-stress experiments identified AvFLS as a salt-inducible gene that is highly expressed in leaves of A. venetum. Subcellular localization analysis showed that AvFLS was located in the cytoplasm, consistent with other plant FLS proteins.The overexpression of AvFLS in Arabidopsis thaliana significantly improved the salt-stress tolerance of the transgenic plants: under salt stress, transgenic Arabidopsis exhibited improved flavonoid accumulation, seed germination rate, plant growth, chlorophyll content, and fresh weight compared to wild-type plants. Comparison of malonaldehyde (MDA), soluble sugar, and proline contents between the transgenic and wild-type plants indicated that the improved salt tolerance associated with AvFLS overexpression was due to decreased membrane damage. AvFLS overexpression also led to the upregulation of endogenous Arabidopsis genes involved in flavonoid biosynthesis. The results of this study demonstrate the potential utility of the AvFLS gene for molecular crop breeding, both to increase the contents of valuable flavonoids and to improve crop productivity in saline fields.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Soil salinity is one of major abiotic stressors of plants, affecting both growth and yield. Plants adapt to changes in habitat by continuously adjusting energy flow and secondary metabolic pathways to maintain growth and development while resisting biotic and abiotic stresses (Goyal et al. 2012; Bita et al., 2019; Thakur et al. 2019; Yadav et al. 2021). Flavonoids, a class of important secondary metabolites, are extensively distributed across the plant kingdom and can be divided into six categories: flavones, isoflavones, flavanones, flavonols, flavanols, and anthocyanins (Agati et al. 2012). Flavonoids play important roles in plant development, growth, and stress resistance (Hartmann 2007; Vickers et al. 2009). Flavonols are important component of flavonoids and are critical for the regulation of plant growth and development, as well as the response to biotic and abiotic stresses (Reginato et al. 2014; Flowers and Muscolo 2015; Shah and Smith 2020). For example, flavonols promote the growth of plant roots under drought stress and alleviate the inhibitory effects of abscisic acid on lateral root growth (Nguyen et al. 2013). Similarly, 5-aminoallylic acid induces the accumulation of flavonols in the guard cells of Arabidopsis thaliana, and flavonols inhibit ABA-induced stomatal closure by scavenging hydrogen peroxide; this process regulates external gas flow under stress conditions (An et al. 2016). Finally, comparisons of the effects of flavonoids among various drought-tolerant tomato varieties demonstrated that, under drought stress, the increase in flavonoid content in drought-tolerant varieties was significantly greater than that in drought-sensitive varieties (Sánchez-Rodríguez et al. 2011).
Flavonoids are synthesized using common key enzymes such as chalcone synthase (CHS), chalcone isomerase (CHI), flavanone 3-hydroxylase (F3H), flavonol synthase (FLS), and dihydroflavonol reductase (DFR). Flavonols and anthocyanins are synthesized via the same pathway before diverging at the point of dihydroflavonol production. At this point, dihydroflavonols are converted to anthocyanins or flavonols by DFR or FLS, respectively (Vu et al. 2015; Park et al. 2019). Flavonol synthase (FLS), a member of the 2-oxoglutarate-dependent dioxygenase (2-ODD) family, is an important rate-limiting enzyme in the plant flavonol metabolic pathway (Muir et al. 2001; Tanaka et al. 2008). FLS not only participates in plant growth and development, but also plays an important role in the plant defense response to biotic and abiotic stresses, such as ultraviolet radiation and pathogen invasion (Xu et al. 2012; Fang et al. 2019; Li et al. 2020; Yu et al. 2020; Dong and Lin 2021). Indeed, FLS activity was first reported in suspension cultures of parsley cells irradiated with ultraviolet (Britsch et al. 1981). Ultraviolet radiation led to a significant increase in plant flavonoid contents, and flavonols, the production of whose is catalyzed by FLS, can protect plants such as corn from ultraviolet damage (Falcone Ferreyra et al. 2010). For example, the Ginkgo biloba FLS (GbFLS) gene is induced in response to ultraviolet light, sodium chloride (NaCl), exogenous abscisic acid, and salicylic acid (Xu et al. 2012), thereby promoting flavonoid accumulation and improving salt tolerance. Two other 2-ODD genes in the flavonoid anabolic pathway, anthocyanin synthase (ANS) and flavanone 3-hydroxylase (F3H), might be somewhat interchangeable or complementary with FLS (Kawai et al. 2014; Guo et al. 2020; Wang et al. 2021a, 2021b, 2021c, 2021d). For example, Arabidopsis FLS1 partially catalyzes F3H, while Arabidopsis ANS shares some of the functions of FLS (Owens et al. 2008; Stracke, et al. 2009; Martens et al. 2010).
The flavonol content of various plants is strictly regulated by the FLS genes (Verhoeyen et al 2002; Ma et al. 2014; Wang et al. 2016; Xu et al. 2020a, b). FLS genes have been well-studied in many plants (Koes et al. 1994; Falcone Ferreyra et al. 2012; Fujino et al. 2018; Yonekura-Sakakibara et al. 2019; Liu et al. 2021). The full-length FLS gene sequence was first cloned from petunia (Froemel et al. 1985) and has subsequently been identified and cloned from a variety of other plants, including Petunia hybrida (Holton et al. 1993), A. thaliana (Owens et al. 2008; Preuß et al. 2009; Falcone Ferreyra et al. 2010), Litchi chinensis (Liu et al. 2018), Fagopyrum tataricum (Li et al. 2012), Scutellaria baicalensis (Kim et al. 2014), Ginkgo biloba (Xu et al. 2012), Camellia sinensis (Jiang et al. 2020), Acacia confusa (Toh et al. 2013), Vaccinium uliginosum (Zhang et al. 2016a, b), and Oryza sativa (Park et al. 2019). FLS gene functions and characteristics may differ both among and within species. For example, all five FLS genes from Vitis vinifera are expressed in the flowers, but only VvFLS4 and VvFLS5 are expressed in fruits (Fujita et al., 2003). Similarly, the functional properties of the six FLS genes found in A. thaliana (AtFLS1–6) differ: the AtFLSl protein has very strong catalytic activity, AtFLS3 has very low enzymatic activity, and the remaining AtFLS proteins have no enzymatic activity (Owens et al. 2008). In addition, the transcriptional abundance of Triticum aestivum FLS (TaFLS1) under salt stress was significantly greater in the salt-tolerant variety SR3 as compared to the salinity-sensitive cultivar Jinan 177 (JN177) (Wang et al. 2014). TaFLS1 improved the salinity tolerance of A. thaliana at the seedling stage but not at the germination stage. (Wang et al. 2016). Finally, the overexpression of AtFLS1 in A. thaliana significantly altered seed coat color and flavonoid accumulation without affecting growth performance or abiotic stress tolerance as compared to the wild type (Nguyen et al. 2016).
Apocynum venetum Linn. (Apocynaceae) is a perennial, halophytic forage plant with high medicinal value and strong stress resistance that grows on saline-alkali lands, sandy wastelands, and floodplains in northern China. A. venetum is rich in flavonoids (Chen et al. 2020; Xu et al. 2021; Yang et al. 2021; Abubakar et al. 2022), and salinity stress was shown to significantly increase flavonol content (Xie et al., 2014; Jiang et al. 2020; Xu et al. 2020a, b; Wang et al. 2021a, b, c, d). However, the physiological and biochemical mechanisms underlying the antioxidant protection conferred by flavonoids in A. venetum plants adapted to salt stress remain unclear. Previously, we cloned the key enzyme gene in the flavonoid biosynthesis pathway of A. venetum, namely AvFLS (Guo et al. 2019). To further characterize the potential role of AvFLS in salt tolerance and to explore the relationship between FLS and salt resistance, we generated transgenic Arabidopsis lines overexpressing AvFLS. This study may enhance our understanding of AvFLS function in plants under salt stress and may provide a new tool for the improvement of plant salt tolerance in the field.
2 Materials and methods
2.1 Cloning and analysis of AvFLS
2.1.1 Plant materials and salt stress treatments
A. venetum seeds were collected from wild plants in Shaya County, Xinjiang Uygur Autonomous Region, China (40˚92´N, 82˚21´E; 957 m). Seedlings were cultured as previously described (Guo et al. 2022). To assess the response of the FLS gene of A. venetum (AvFLS) to salt stress, six-week-old plantlets were subjected to one of two treatments: (1) To determine the tissue-specificity of the AvFLS gene under control conditions and under salt stress, NaCl (0, 50, or 300 mM) was added to the hydroponic growth solution. After two weeks of salt stress, seedling roots, leaves, and stems were collected, frozen in liquid nitrogen, and stored at − 80 °C. (2) To determine the expression patterns of AvFLS under prolonged salt stress, plants were subjeted to 0, 50, 100, 150, 200, or 300 mM NaCl stress treatment for 144 h. Leaves were sampled at 6 and 144 h. At each sampling point, the second pair of unfolded leaves (from the top) were collected, immediately frozen in liquid nitrogen, and stored at − 80°C for RNA extraction.
2.1.2 Gene cloning and bioinformatics analysis
The full-length cDNA of AvFLS was downloaded from the GenBank database (accession no. MK391176.1) (Guo et al. 2019). The coding sequence was amplified using the pEASY-Uni Seamless Cloning and Assembly Kit (TransGen Biotech) with the gene-specific primers AvFLS-F1, AvFLS-F2, AvFLS-R1, and AvFLS-R2 (Table S1). Sequences from other species homologous to the predicted AvFLS protein sequence were identified and aligned using DNAMAN (Ver. 7.0) (Liu et al. 2018; Guo et al. 2019; Li et al. 2021). Phylogenetic analysis was carried out using the neighbor joining method (bootstrap resampling test with 1000 replicates) in MEGA version 10.0 software (https://megasoftware.net/). Eustoma russellianum was used as the outgroup (Wang et al. 2021a, b, c, d; Guo et al. 2022; Wang et al. 2022a, b).
2.1.3 AvFLS expression patterns and subcellular localization
To compare AvFLS expression patterns among tissues and treatment groups, RNA was first isolated from the collected samples and reverse transcribed to cDNA as previously described (Jeong et al., 2018;Wang et al. 2021a, b, c, d ;Zhang et al., 2021). We performed real-time quantitative PCRs (qRT-PCRs) using the cDNA template and the housekeeping gene Actin as an internal control, as described previously (Zhang et al., 2021). PCRs were performed on an ABI Prism7500 (Applied Biosystems, USA). The gene-specific primers (AvFLS-qRT-F/R) and actin primers (Actin-qRT-F/R) used for PCR are shown in Table S1. Three biological replicates and three technical replicates of each sample were performed. The comparative CT method (2−ΔΔCt) (Wang et al. 2021a, b, c, d;Zhang et al., 2021;Wang et al. 2022a, b) was used to calculate the relative expression of each target gene.
Using the gateway method, the full-length coding sequence of AvFLS was fused to the modified pCAMBIA1302-GFP vector under the control of the cauliflower mosaic virus (CaMV) 35S promoter to generate the 35S::AvFLS-GFP fusion construct. The 35S::AvFLS-GFP and 35S::GFP fusion proteins were introduced into different epidermal cells of Nicotiana tabacum using the Agrobacterium-mediated transformation method (Zhang et al. 2016a, b). The transformed N. tabacum leaves were cultivated for 2–6 days and then examined under a confocal laser scanning microscope (Zeiss Lsm 700, Zeiss, Jena, Germany).
Effects of salt stress on transgenic Arabidopsis overexpressing AvFLS.
2.1.4 Generation of transgenic Arabidopsis overexpressing AvFLS
The AvFLS fragment was inserted into the pEarleyGate 100 vector, and the recombinant protein was transformed into wild-type (WT) A. thaliana (Columbia-0) using the Agrobacterium-mediated floral dip method as previously described (Holton, et al. 1993; Muir,et al. 2001; Li et al. 2021). The specific primers used are listed in Supplementary Table S1. Successfully transformed seeds were selected on MS medium containing 50 mM kanamycin as previously described (Nguyen, et al. 2016; Wang, et al. 2021a, b, c, d). The T3 generation transgenic lines (OE1) with the highest levels of AvFLS expression and with 100% resistance to kanamycin were considered homozygous and were thus selected for further analysis.
2.1.5 Salt-stress treatment and effects on biomass
To test the response of AvFLS-transgenic Arabidopsis to salinity stress, WT and OE1 seeds were seeded in 1/2 MS medium containing 0, 25, 50, 75, or 100 mM NaCl. Once cotyledons appeared, the number of germinated seeds was counted every 24 h for seven days. The seed germination percentage was calculated as the total number of seeds germinated over the seven day period/total number of seed planted × 100% (Ellis et al. 1986).
Four-week-old WT and OE1 seedlings were stressed using 100 mM NaCl. Leaves were collected from the treated plants at 0 h, 6 h, 24 h, 72 h, 120 h, and 168 h after salt treatment for the assessment of biomass, photosynthetic pigments, soluble sugar, proline, malondialdehyde (MDA), and total flavonoids. Each parameter was measured in six replicates. Fresh samples were weighed, oven-dried at 60 ± 65°C for 16 h (Delgado-Pertı́ñez et al. 2000), and weighed again to obtain seedling dry and fresh weights.
2.1.6 Measurement of physiochemical parameters
Chlorophyll a and chlorophyll b were extracted from leaves (0.1 g) using a mixture of acetone, ethanol and water in a volume ratio of 4.5:4.5:1. A spectrophotometer (Shimadzu, UV-1800, Suzhou, China) was used to measure absorbance at 663 nm and 645 nm. Chlorophyll concentrations were calculated as follows (Gratani 1992):
where D663 and D645 correspond to absorbance at 663 and 645 nm, respectively; v is the extraction volume (ml); and W is the leaf weight (g).
Soluble sugar content was measured using Anthrone colorimetry (Yemm and Willis 1954). We constructed a standard curve reflecting the linear relationship between the mass concentration of glucose (X) and the absorbance value (Y) as follows: y = 9.095x − 0.0075 (R2 = 0.998) (Supplementary Fig. 1A). This standard curve was reliable for glucose concentrations of 0–0.1 mg/ml. Proline was extracted and quantified using the method of Bates et al. (1973); malondialdehyde content was measured using a thiobarbituric acid reaction following Heath and Packer (1968); and total flavonoid concentration was determined using the aluminum nitrate method (Park et al. 1997) with rutin as the standard.
2.1.7 Expression patterns of genes associated with stress tolerance
To explore the possible molecular mechanisms underlying the participation of FLS in the salt-stress response, we used qRT-PCR to quantify the transcript levels of five genes encoding enzymes in the flavonoid biosynthesis pathway (AtCHI, AtCHS, AtANS, AtF3H, and AtFLS1) in transgenic and WT plants under salt stress. Four-week-old WT and OE1 seedlings were exposed to 100 mM NaCl for 0 h, 6 h, 24 h, 72 h, and 120 h. Samples were then collected for RNA extraction and cDNA synthesis. RNA was isolated from the collected samples and reverse transcribed to cDNA as previously described (Wang et al. 2021a, b, c, d; Zhang et al., 2021). qRT-PCRs were performed as described above using gene-specific primers (Supplementary Material Table 1).
2.1.8 Statistical analysis
All data are presented as means ± standard deviation (SD). Significant differences among treatments were identified using one-way ANOVAs in SPSS 20.0, followed by Duncan’s multiple range tests. We considered p < 0.05 statistically significant.
3 Results
3.1 Analysis of AvFLS in A. venetum
3.1.1 Bioinformatic sequence analysis
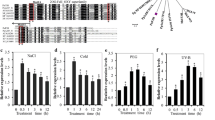
The full-length cDNA sequence of AvFLS had an open reading frame of 1212 bp and was predicted to encode a polypeptide containing 335 amino acids, with a predicted molecular weight of 38.33 kDa and a predicted isoelectric point of 6.11. Multiple sequence alignments showed that the deduced amino acid sequence of AvFLS was highly homologous with known FLS proteins from other plant species (Fig. 1).
Relationship between AvFLS and other known FLS proteins from representative plant species. Multiple sequence alignment. Thick black line indicates the highly conserved N-terminal 2-ketoglutarate domain; fine black line indicates the 2OG-Fe (II) oxygenation enzyme domain. The regions marked H–X-D, H–X, and R-X-S correspond to the regions conserved across genes in the 2-ODD superfamily
A GenBank conserved domain database search and functional analysis revealed that AvFLS possessed putative conserved domains belonging to the 20G-FeII Oxy super family, as well as a highly conserved N-terminal region found in proteins with 2-oxoglutarate/Fe(II)-dependent dioxygenase activity. Importantly, the positions of many of the conserved sequence motifs identified in AvFLS, including the HXD motif for ligating ferrous iron and the RXS motif for binding 2-oxoglutarate (2OG), were similar between the AvFLS sequence and the other FLS sequences (Fig. 1). Phylogenetic analysis recovered AvFLS in a sister relationship with the FLS protein from Coffea arabica and within a larger clade comprised of the FLS proteins from Solanum lycopersicum, Nicotiana tabacum, Gentiana triflora, Eustoma grandiflorum, and Sesamum indicum (Fig. 2).
3.1.2 Expression patterns and subcellular localization
To explore the possible role played by AvFLS in the response of A. venetum to salt stress, we quantified its expression patterns under control and salt-stress conditions using qRT-PCR. AvFLS was ubiquitously expressed in all tested tissues (Fig. 3). AvFLS was significantly upregulated in the leaves as compared to the roots and stems across all treatment groups, and AvFLS expression level increased significantly as salinity increased (Fig. 3). These results suggested that AvFLS may be involved in the stress response of A. venetum, similar to other known FLS genes such as MdFLS1 in Malus domestica (Li et al. 2021) and BnFLS in Brassica napus (Vu et al. 2015).
Gene expression patterns of AvFLS in various tissues of A. venetum in response to salt stress. The values shown are the means ± SDs of three biological replicates. Error bars represent the standard deviation of three replicates. Different letters within a treatment group indicate significant differences based on one-way ANOVAs (P < 0.05)
As salt stress was prolonged, the expression patterns of AvFLS in A. venetum leaves changed substantially. After 6 h, relative AvFLS gene expression was significantly greater in leaves exposed to 200 and 300 mM NaCl as compared to leaves exposed to 100 mM NaCl or less (Fig. 4). However, after 144 h of salt treatment, the AvFLS gene was significantly upregulated in the leaves exposed to 200 and 300 mM NaCl as compared to all other treatment groups (Fig. 4). This suggested that the AvFLS gene was induced by salt stress, and that the response fluctuated over time. This may indicate that AvFLS plays an important role in the resistance of A. venetum to salt stress.
Relative expression level of the AvFLS gene in the leaf tissues of A. venetum exposed to various concentrations of NaCl (0–300 mM). (a) After 6 h of salt treatment. (b) After 144 h of salt treatment. The values shown are the means ± SDs of three biological replicates. Error bars represent the standard deviation of three replicates. Different letters within a treatment group indicate significant differences based on one-way ANOVAs (P < 0.05)
To determine the subcellular location of AvFLS, the recombinant AvFLS-green fluorescent protein (GFP) was transformed into N. tabacum mesophyll protoplasts. The pCAMBIA1302GFP vector was used as a control. The control GFP protein was expressed throughout the cell, while the GFP signal of AvFLS was localized in the cytoplasm (Fig. 5). As the AvFLS does not possess a transmembrane-spanning domain (Guo et al. 2019), we concluded that AvFLS was a cytoplasmic protein.
3.2 Effects of AvFLS overexpression on the salinity tolerance of Arabidopsis
3.2.1 Growth performance was improved in transgenic Arabidopsis under salt stress
Under control conditions (0 mM NaCl), there was no significant difference in seed germination rate between the WT and transgenic plants (Fig. 6a). Exposure to 25–100 mM NaCl for 7 d significantly decreased the germination rate of both the WT and overexpression line OE1, but the germination rate was significantly higher in the transgenic line as compared to the WT across all treatment groups (Fig. 6a).
Growth performance of transgenic Arabidopsis overexpressing AvFLS as compared to WT Arabidopsis under salt stress. a AvFLS expression in AvFLS-over-expressing (OE1) and wild-type (Col-0) Arabidopsis and in Apocynum venetum. b OE1 and Col-0 Arabidopsis seed germination in 1/2 MS medium supplemented with NaCl (0, 25, 50, 75 and 100 mM). c Five-week-old OE1 and Col-0 Arabidopsis seedlings grown under control conditions (without NaCl). d Four-week-old OE1 and Col-0 Arabidopsis seedlings after treatment with 100 mM NaCl for one week. The values shown are the means ± SDs of six biological replicates. Error bars represent the standard deviation of three replicates. Different letters within a treatment group indicate significant differences based on one-way ANOVAs (P < 0.05). Scale bars: 5 cm
After growth at control conditions for five weeks, the transgenic Arabidopsis were somewhat larger than the WT plants (Fig. 6b). After four-week-old WT and transgenic plants were treated with 100 mM NaCl for one week, the growth of both lines was severely curtailed. However, the growth of the transgenic line was noticeably better than that of the WT line (Fig. 6c).
Under control conditions, the biomass of the transgenic plants was greater than that of the WT plants (both wet and dry weights; Fig. 7). Although the biomass of both lines decreased significantly under increased salinity, the biomass of the transgenic line remained significantly higher than that of the WT (Fig. 7): after 7 d of exposure to 100 mM NaCl, the fresh and dry weights of the transgenic plants were 24.95% and 14.08%, respectively, greater than the fresh and dry weights of the WT (Fig. 7).
Dry and fresh weights of transgenic and wild-type Arabidopsis under salt stress. The values shown are the means ± SDs of three biological replicates. Error bars represent the standard deviation of six replicates. Different letters within a treatment group indicate significant differences based on one-way ANOVAs (P < 0.05)
3.2.2 AvFLS overexpression mitigated the negative effects of salinity on the physiochemical parameters of Arabidopsis
Chlorophyll a, chlorophyll b, and total chlorophyll contents generally decreased in both WT and transgenic Arabidopsis seedlings as exposure to 100 mM NaCl was prolonged. However, chlorophyll a content in the transgenic plants was significantly greater than that in the WT plants at all times points measured (P < 0.05): after 0 h, 6 h, 24 h, 72 h, 120 h, and 168 h of exposure to 100 mM NaCl, the chlorophyll a content of the transgenic plants was 39.33%, 7.62%, 28.45%, 59.64%, 32.03%, and 93.08%, respectively, greater than that of the WT plants (Fig. 8A). Chlorophyll b content in the transgenic plants was also significantly greater than that in the WT plants at all times points measured except 6 h (P < 0.05): after 0 h, 6 h, 24 h, 72 h, 120 h, and 168 h of exposure to 100 mM NaCl, the chlorophyll b content of the transgenic plants was 19.50%, 6.8%, 27.40%, 77.72%, 109.72%, and 112.94%, respectively, greater than that of the WT plants (Fig. 8B). Similar to the trend in chlorophyll a content, total chlorophyll levels in the transgenic plants were significantly greater than those in the WT plants at all times points measured (P < 0.05; Fig. 8C).
Chlorophyll content of transgenic and wild-type Arabidopsis under salt stress. a Chlorophyll a content. b Chlorophyll b content. c Total chlorophyll content. The values shown are the means ± SDs of six biological replicates. Error bars represent the standard deviation of three replicates. Different letters within a treatment group indicate significant differences based on one-way ANOVAs (P < 0.05)
Soluble sugar content fluctuated in both transgenic (OE1) and WT A. thaliana as the duration of salt stress (100 mM NaCl) was prolonged (Fig. 9a). However, the soluble sugar content of the transgenic line (OE1) was significantly greater than that of the WT at 0 h, 6 h, 24 h, and 120 h (Fig. 9a). At 168 h, the soluble sugar content of the WT was significantly greater than that of the transgenic line, and at 72 h, there was no significant difference in soluble sugar content between the lines (Fig. 9a and Supplementary Material Fig. 10a).
Physiological indexes in the leaves of transgenic A. thaliana overexpressing AvFLS (OE1) and wild-type A. thaliana (Col-0) under prolonged salt stress (100 mM NaCl). a Soluble sugar content. b Proline content. c MDA content. d Total flavonoid content. The values shown are the means ± SDs of six biological replicates. Error bars represent the standard deviation of three replicates. Different letters within a treatment group indicate significant differences based on one-way ANOVAs (P < 0.05)
In both transgenic and WT Arabidopsis leaves, proline content gradually increased with time under salt stress (100 mM NaCl): after 24 h, 72 h, 120 h, and 168 h of salt stress, proline content in the WT leaves increased 2.29-fold, 6.60-fold, 6.68-fold, and 13.73-fold compared to control conditions (0 h), while proline content in the transgenic leaves increased 6.69-fold, 5.09-fold, 11.44-fold, and 24.30-fold compared to control conditions (Fig. 9b). However, the proline content of the transgenic A. thaliana leaves treated with 100 mM NaCl for 72 h was lower than that of the WT, and this difference was maintained until the end of the experiment (Fig. 9b and Supplementary Material Fig. 10b).
Under control conditions (0 h), MDA content in the WT Arabidopsis leaves was significantly greater than that in the transgenic leaves and, as salt stress was prolonged, MDA content increased significantly in both WT and transgenic lines (Fig. 9c). However, the MDA content of the transgenic leaves was significantly lower than that of the WT leaves at all time points except 72 h and 120 h (Fig. 9c). The MDA content of the transgenic leaves was 30.7%, 35.3%, 16.3%, and 16.7% lower than that of the WT leaves at 0 h, 6 h, 24 h, and 168 h, respectively. This suggested that that the overexpression of AvFLS reduced the degree of salt-induced membrane lipid peroxidation in Arabidopsis, reducing MDA production and the associated tissue damage.
The standard curve of rutin was Y = 12.537x + 0.00184 (R2 = 0.9979), and this standard curve had good linearity for rutin concentrations of 0–0.07 mg/L (Supplementary Material Fig. 10b).
At all time points except 6 h, total flavonoid content was significantly greater in the transgenic leaves than in the WT leaves (Fig. 9d). At 0 h, 24 h, 72 h, 120 h and 168 h, total flavonoid contents in the transgenic leaves were 37.2%, 32.2%, 60.3%, 37.1%, and 54.3%, respectively, greater than total flavonoid contents in the WT leaves (Fig. 9d). This suggested that the overexpression of AvFLS might significantly increase total flavonoid content in A. thaliana.
3.2.3 AvFLS overexpression upregulated the expression of flavonoid pathway genes in Arabidopsis
The heterologous expression of the FLS gene affects the synthesis of total flavonoids and anthocyanins in plants, and may also lead to the downregulation or non-expression of endogenous genes (Ohno et al. 2011; Nguyen, et al. 2016; Jeyaraj et al. 2017; Jiang et al. 2020). Under control conditions, the relative expression of the AtFLS gene did not differ significantly between the transgenic (OE1) and WT plants (Fig. 10a). However, under salt stress, AtFLS was significantly upregulated in the WT plants as compared to the transgenic plants at all time points except 72 h (Fig. 10a). The expression of AvFLS peaked in the OE1 plants after 24 h of salt treatment; at the same time point, the expression of AtFLS was relatively low (Fig. 10b).
Expression levels of genes related to flavonoid synthesis pathway under salt stress. a Expression of the Arabidopsis flavonol synthase gene (AtFLS) in Col-0 and OE1 plants. b Expression of the A. venetum flavonol synthase gene (AvFLS) in OE1 plants. c Expression of the Arabidopsis dihydroflavonol-4-reductase gene (AtDFR) in Col-0 and OE1 plants. d Expression of the Arabidopsis anthocyanin synthase gene (AtANS) in Col-0 and OE1 plants. e Expression of the Arabidopsis flavanone 3-hydroxylase gene (AtF3H) in Col-0 and OE1 plants. f Expression of the Arabidopsis chalcone synthase gene (AtCHS) in Col-0 and OE1 plants. g Expression of the Arabidopsis chalcone isomerase gene (AtCHI) in Col-0 and OE1 plants. The values shown are the means ± SDs of three biological replicates. Error bars represent the standard deviation of three replicates. Different letters within a treatment group indicate significant differences based on one-way ANOVAs (P < 0.05)
AvFLS overexpression also affects the expression patterns of some genes in the flavonoid synthesis pathway of A. thaliana. Two genes in the anthocyanin metabolic pathway, AtDFR and AtANS, were significantly upregulated in the transgenic line as compared to the WT after 24 h, 72 h, and 120 h of salt stress (Fig. 10c, d). Similarly, the genes encoding flavanone 3-hydroxylase (AtF3H; Fig. 10e) and chalcone synthase gene (AtCHS; Fig. 10f) were significantly upregulated in the transgenic lines as compared to the WT lines after 120 h of salt stress. The chalcone isomerase gene (AtCHI) was significantly upregulated in both the transgenic and the WT lines as salt stress was prolonged; there was no significant difference in expression between the two lines (Fig. 10g).
4 Discussion
Flavonol synthase is a key enzyme in the plant flavonoid metabolic pathway and therefore plays an important role in flavonoid biosynthesis. Plant flavonoids, which are regulated by the flavonoid metabolic pathway, determine flower and seed-coat colors (Lou et al. 2014; Zhang et al. 2018). In addition, increases in flavonoid content can increase plant tolerance of biotic and abiotic stresses (Yuan, et al. 2015; Baskar et al. 2018; Li et al. 2018), because flavonoids can act as antioxidants or pro-oxidants to eliminate or reduce the tissue damage associated with the reactive oxygen species produced as a result of adverse environmental conditions (Lee et al. 2004; Tattini et al. 2004; Eghbaliferiz and Iranshahi 2016). Therefore, further systematic physiological and molecular experiments of FLS gene will be helpful to further reveal the mechanism of FLS gene in improving plant stress.
In this study, we successfully isolated and characterized AvFLS cDNA from A. venetum, demonstrated that AvFLS encodes a functional protein, and showed that AvFLS was ubiquitously expressed throughout A. venetum, including leaves, stems, and roots. AvFLS expression levels were higher in the leaves and lower in the stems and roots, these results are consistent with the research results of Wang et al.,(2021a, b, c, d).
To further investigate the flavonol synthase function of the FLS protein from A. venetum, the amino acid sequence of AvFLS was compared with predicted FLS sequences from other species using a phylogenetic tree. Phylogenetic analyses recovered A. venetum in a close relationship with predicted FLS proteins from Capsicum baccatum (Wang et al. 2022a, b), Nicotiana tabacum (Shi et al. 2017), and Solanum lycopersicum (Morimoto and Tao 2016; Gao et al. 2018).
At the subcellular level, AvFLS appeared to be localized in the cytoplasm and not in the nucleus, consistent with the observation that flavonoids are synthesized and localized in the cytoplasm in A. thaliana (Saslowsky et al. 2005; Kuhn et al. 2011) and Dendrobium officinale (Yu et al. 2020). However, there are many flavonoid metabolic enzymes that have been localized in the cytoplasm, including CHS, CHI, and FLS (Kuhn et al. 2011).
In response to stress, plants may synthesize osmotic regulators, such as proline, soluble sugars, and betaine to improve stress resistance (Farooq et al. 2009; Hao et al. 2021). Proline accumulation is triggered by environmental stress (Hare and Cress 1997; Maggio et al., 2021) to protect cells from stress-associated damage (Hong et al. 2000; Kavi and Sreenivasulu et al., 2014). We observed that salt stress significantly increased the proline content of the leaves of both WT Arabidopsis seedlings and Arabidopsis seedlings overexpressing AvFLS. After 168 h of salt stress (100 mM NaCl), the proline content in the leaves of the WT was 522.8 μg· g−1, a 13.7-fold increase compared to proline content under control conditions (0 h). Similarly, the proline content in the transgenic leaves was 419.37 μg·g−1 after 168 h of salt stress, corresponding to a 24.3-fold increase in proline content compared to control conditions (0 h). Under control conditions and during the early stages of salt stress (0 h, 6 h, and 24 h), soluble sugar content was significantly greater in the transgenic plants than in the WT plants. Thus, AvFLS overexpression improved osmoregulation and increased soluble sugar accumulation in Arabidopsis, enhancing resistance to salt stress. Similarly, the overexpression of the FLS gene of Chrysanthemum morifolium (CmFLS) improved the stress resistance of tobacco (Wang et al. 2021a, b, c, d).
Photosynthesis provides the energy required for plant growth and development, and is thus critical for increases in plant biomass and economic crop yield (Richards et al., 2006; Long et al. 2006; Zhu et al. 2008; Zhu et al. 2010). Chlorophyll content reflects the efficiency of plant photosynthesis. Therefore, chlorophyll concentration indicates the physiological state of the plant and can be considered a biochemical marker of stress tolerance (Arnon 1949; Rahneshan et al., 2018). Indeed, chlorophyll concentration has been specifically identified as an important physiological indicator of plant salt tolerance (Singh and Gautam 2013; Bernal-Vicente et al. 2018). Under stress conditions, the chlorophyll content of plants with strong stress resistance is higher than the chlorophyll content of plants with poor stress resistance (Billings and Mooney 1968; Quick et al. 1992; Mukami et al. 2019). After 168 h of salt exposure, the contents of both chlorophyll a and chlorophyll b were significantly greater in the transgenic Arabidopsis seedlings overexpressing AvFLS that in WT plants, indicating that the transgenic plants were more resistant to salt stress. Consistent with this, when the Euphorbia kansui flavonol synthase gene (EkFLS) was overexpressed in A. thaliana, chlorophyll content increased, affecting photosynthesis and promoting stress resistance (Wang et al. 2021a, b, c, d).
Under abiotic stress, plants produce large amounts of peroxides, and the resulting oxidative damage to plant cell membranes produces MDA (Parvanova et al. 2004). Due to their polyphenolic structure, flavonoids act as strong antioxidants in plants, scavenging excess free radicals (Djeridane et al. 2006). Increases in flavonoid content improve the antioxidant capacity of plants, which in turn improves plant stress tolerance (Ashraf 2009; Akula and Ravishanka 2011). Here, MDA contents in the transgenic plants were lower than those of the WT under control and salt-stress conditions, indicating that AvFLS overexpression reduced the lipid peroxidation of cell membranes in Arabidopsis.
Studies have shown that in some cases, when the overexpressed exogenous gene is highly similar to the endogenous gene, both genes in the transgenic plant are inhibited (i.e., co-suppression; Facchini 2001). For example, the overexpression of the rice (Oryza sativa) FLS gene OsFLS in tobacco led to the downregulation of the early and late biosynthetic genes in the flavonoid pathway; the endogenous FLS gene NtFLS was particularly strongly inhibited (Park et al. 2019). Here, the expression levels of AtFLS did not differ significantly between the transgenic and WT A. thaliana lines under control conditions, but AtFLS was significantly downregulated in the transgenic plants as compared to the WT under salt stress. This indicated that overexpression of the AvFLS gene moderately suppressed the expression of the endogenous AtFLS gene in Arabidopsis. However, due to the strong upregulation of the AvFLS gene, total flavonoid content was significantly greater in the AvFLS-overexpression line under both salt-stress and control conditions. Consistent with this, AvFLS-overexpressing transgenic tobacco plants accumulated flavonoids to a greater extent than WT plants, which resulted in a significant positive effect on the recovery of growth under salinity stress (Wang et al. 2021a, b, c, d).
FLS transcript levels affect the contents of various flavonols in plants, and thus participate in the regulation of the plant response to various external stresses (Khare et al. 2018; Sharma et al. 2019; Hou et al. 2020). For example, the contents of flavonols, such as quercetin and kaempferol, increased significantly in Tartary buckwheat seedlings under salt stress, and the FtFLS2 gene was simultaneously significantly upregulated (Li et al. 2018). Similarly, ultraviolet radiation increased flavonol accumulation and strongly upregulated F3H and FLS in soybean (Kim et al. 2008). It also has been reported that EkFLS overexpression is strongly correlated with an increase in flavonoid synthesis and therefore in abiotic stress tolerance in Arabidopsis (Wang et al. 2021a, b, c, d). In addition, FtFLS2 was upregulated by SA and NaCl, affecting the flavonol biosynthetic pathway (Li et al. 2013). Similarly, overexpression of BnFLS increased flavonol accumulation and exhibited enhanced tolerance to multiple abiotic stressors in the transgenic plants (Vu et al. 2015). Treatment with the flavonoid quercetin not only alleviated the adverse effects of mannitol-induced osmotic stress, but also upregulated CHI and FLS in Apocynum seedlings (Yang et al. 2021). The heterologous expression of the FLS gene led to the downregulation or non-expression of endogenous genes in N. tabacum (Jiang et al. 2020), while the overexpression of AvFLS in Arabidopsis downregulated the endogenous FLS gene AtFLS in our research work. In vivo, FLS and DFR genes competitively determine the accumulation of flavonols, anthocyanins, and flavan-3-ols in plants (Mahajan et al. 2011; Zhao et al. 2014; Luo et al. 2016; Akita et al. 2018). However, the overexpression of AvFLS, the FLS gene from the halophyte A. venetum, in Arabidopsis upregulated not only the genes involved in flavonoid biosynthesis but also the genes involved in anthocyanin synthesis. That is, the expression levels of AtDFR and AtANS in the transgenic plants after 24 h, 72 h, and 120 h of salt stress were significantly higher than those in wild-type Arabidopsis at the same time points. Interestingly, studies have shown that, in FLS-overexpressing plants, the early genes in the flavonoid synthesis pathway (CHS and F3H) were not expressed or were expressed at low levels (Park et al. 2019; Li et al. 2021). Our results showed that the expression levels of AtCHS and AtF3H in transgenic plants treated with salt for 6 h and 24 h were significantly lower than those of wild-type plants at the same time points. However, these genes were significantly upregulated in the transgenic plants compared to the wild-type plants after 120 h salt treatment. At the same time, we also found that the falvonoid content of transgenic Arabidopsis overexpressing AvFLS was significantly greater than that of WT Arabidopsis after 120 h of salt treatment. The upregulation of these genes indicated that the activity of the flavonoid synthesis pathway increased in response to salt stress in the transgenic overexpression line, corresponding to the observed significant increase in total flavonoid content.Together, these results indicate that FLS genes in halophytes and glycophytes may play different functions under salt stress, and it is necessary to continue to study the related mechanisms in subsequent studies.
5 Conclusion
Overexpression of the AvFLS gene in transgenic Arabidopsis improved plant growth and development under salt stress compared to the WT. In transgenic Arabidopsis, the expression level of the endogenous AtFLS gene differed significantly from that of wild type. In addition, AvFLS expression in transgenic Arabidopsis increased with prolonged salt stress. This study not only helped to clarify the molecular mechanisms of salt-tolerance genes, but also provided a theoretical basis for the further mining of Apocynum genes, the molecular breeding of plants with improved resistance to salt stress, and the consequent improvement of crop quality.
References
Abubakar AS, Feng X, Gao G, Yu C, Chen J, Chen K et al (2022) Genome wide characterization of R2R3 MYB transcription factor from Apocynum venetum revealed potential stress tolerance and flavonoid biosynthesis genes. Genomics 114:110275. https://doi.org/10.1016/J.YGENO.2022.110275
Agati G, Azzarello E, Pollastri S, Tattini M (2012) Flavonoids as antioxidants in plants: location and functional significance. Plant Sci 196:67–76. https://doi.org/10.1016/j.plantsci.2012.07.014
Akita Y, Kitamura S, Mikami R, Ishizaka H (2018) Identification of functional flavonol synthase genes from fragrant wild cyclamen (Cyclamen purpurascens). J Plant Biochem Biotechnol 27:147–155. https://doi.org/10.1007/s13562-017-0423-9
Akula R, Ravishankar GA (2011) Influence of abiotic stress signals on secondary metabolites in plants. Plant Signaling Behav 6:1720–1731. https://doi.org/10.4161/psb.6.11.17613
An Y, Feng X, Liu L, Xiong L, Wang L (2016) ALA-induced flavonols accumulation in guard cells is involved in scavenging H2O2 and inhibiting stomatal closure in Arabidopsis cotyledons. Front Plant Sci 7:1713. https://doi.org/10.3389/fpls.2016.01713
Arnon DI (1949) Copper enzymes in isolated chloroplasts. Polyphenoloxidase in Beta Vulgaris Plant Physiol 24:1–15. https://doi.org/10.1104/pp.24.1.1
Ashraf M (2009) Biotechnological approach of improving plant salt tolerance using antioxidants as markers. Biotechnol Adv 27:84–93. https://doi.org/10.1016/j.biotechadv.2008.09.003
Baskar, V, Venkatesh, R, Ramalingam, S (2018) Flavonoids (antioxidants systems) in higher plants and their response to stresses. Antioxid Antioxid Enzymes Higher Plants 253–268. https://doi.org/10.1007/978-3-319-75088-0_12
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207. https://doi.org/10.1007/bf00018060
Bernal-Vicente A, Cantabella D, Petri C, Hernández JA, Diaz-Vivancos P (2018) The salt-stress response of the transgenic plum line J8–1 and its interaction with the salicylic acid biosynthetic pathway from mandelonitrile. Int J Mol Sci 19:3519. https://doi.org/10.3390/ijms19113519
Billings WD, Mooney HA (1968) The ecology of arctic and alpine plants. Biol Rev 43:481–529. https://doi.org/10.1111/j.1469-185X.1968.tb00968.x
Bita CE, Gerats T (2013) Plant tolerance to high temperature in a changing environment: scientific fundamentals and production of heat stress-tolerant crops. Front Plant Sci 4:273. https://doi.org/10.3389/fpls.2013.00273
Britsch L, Heller W, Grisebach H (1981) Conversion of flavanone to flavone, dihydroflavonol and flavonol with an enzyme system from cell cultures of parsley. Z. Naturforsch., C. J Biosci 36:742–750. https://doi.org/10.1515/znc-1981-9-1009
Chan YJ, Won JL, Hai AT, Cao ST, Suk-Whan H, Hojoung L (2018) AtMybL-O modulates abscisic acid biosynthesis to optimize plant growth and ABA signaling in response to drought stress. Appl Biol Chem 61:473–477. https://doi.org/10.1007/s13765-018-0376-2
Chen, C, Wang, C, Liu, Z, Cai, Z, Hua, Y, Mei, Y, et al. (2020) iTRAQ-based proteomic technique provides insights into salt stress responsive proteins in Apocyni Veneti Folium (Apocynum venetum L.). Environ Exp Bot 180. https://doi.org/10.1016/j.envexpbot.2020.104247
Davies KM, Schwinn KE, Deroles SC, Manson DG, Lewis DH, Bloor SJ et al (2003) Enhancing anthocyanin production by altering competition for substrate between flavonol synthase and dihydroflavonol 4-reductase. Euphytica 131:259–268. https://doi.org/10.1023/A:1024018729349
Delgado-Pertı́ñez M, Gómez-Cabrera A, Garrido A (2000) Predicting the nutritive value of the olive leaf (Olea europaea): digestibility and chemical composition and in vitro studies. Anim Feed Sci Technol 87:187–201. https://doi.org/10.1016/S0377-8401(00)00195-4
Djeridane A, Yousfi M, Nadjemi B, Boutassouna D, Stocker P, Vidal N (2006) Antioxidant activity of some Algerian medicinal plants extracts containing phenolic compounds. Food Chem 97:654–660. https://doi.org/10.1016/j.foodchem.2005.04.028
Dong NQ, Lin HX (2021) Contribution of phenylpropanoid metabolism to plant development and plant–environment interactions. J Integr Plant Biol 63:180–209. https://doi.org/10.1111/jipb.13054
Eghbaliferiz S, Iranshahi M (2016) Prooxidant activity of polyphenols, flavonoids, anthocyanins and carotenoids: updated review of mechanisms and catalyzing metals. Phytother Res 30:1379–1391. https://doi.org/10.1002/ptr.5643
Ellis RH, Covell S, Roberts EH, Summerfield RJ (1986) The Influence of Temperature on Seed Germination Rate in Grain Legumes II. Intraspecific variation in chickpea (Cicer arietinum L.) At constant temperatures. J Exp Bot 37:1503–1515. https://doi.org/10.1093/jxb/37.10.1503
Facchini PJ (2001) Alkaloid biosynthesis in plants: biochemistry, cell biology, molecular regulation, and metabolic engineering applications. Annu Rev Plant Physiol Plant Mol Biol 52:29–66. https://doi.org/10.1146/annurev.arplant.52.1.29
Falcone Ferreyra ML, Rius S, Emiliani J, Pourcel L, Feller A, Morohashi K et al (2010) Cloning and characterization of a UV-B-inducible maize flavonol synthase. Plant J 62:77–91. https://doi.org/10.1111/j.1365-313X.2010.04133.x
Falcone Ferreyra ML, Rius SP, Casati P (2012) Flavonoids: biosynthesis, biological functions, and biotechnological applications. Front Plant Sci 3:222. https://doi.org/10.3389/fpls.2012.00222
Fang Y, Deng X, Lu X, Zheng J, Jiang H, Rao Y et al (2019) Differential phosphoproteome study of the response to cadmium stress in rice. Ecotoxicol Environ Saf 180:780–788. https://doi.org/10.1016/j.ecoenv.2019.05.068
Farooq M, Wahid A, Kobayashi N, Fujita D, Basra SMA (2009) Plant drought stress: effects, mechanisms and management. Agron Sustainable Dev 29:185–212. https://doi.org/10.1051/agro:2008021
Flowers, TJ, Muscolo, A (2015) Introduction to the special issue: halophytes in a changing world. AoB Plants 7. https://doi.org/10.1093/aobpla/plv020
Froemel S, De Vlaming P, Stotz G, Wiering H, Forkmann G, Schram AW (1985) Genetic and biochemical studies on the conversion of flavanones to dihydroflavonols in flowers of Petunia hybrida. Theor Appl Genet 70:561–568. https://doi.org/10.1007/BF00305991
Fujino N, Tenma N, Waki T, Ito K, Komatsuzaki Y, Sugiyama K et al (2018) Physical interactions among flavonoid enzymes in snapdragon and torenia reveal the diversity in the flavonoid metabolon organization of different plant species. Plant J 94:372–392. https://doi.org/10.1111/tpj.13864
Fujita A, Goto-Yamamoto N, Aramaki I, Hashizume K (2006) Organ-specific transcription of putative flavonol synthase genes of grapevine and effects of plant hormones and shading on flavonol biosynthesis in grape berry skins. Biosci., Biotechnol. Biochem 70:632–638. https://doi.org/10.1271/bbb.70.632
Gao Y, Liu J, Chen Y, Tang H, Wang Y, He Y et al (2018) Tomato SlAN11 regulates flavonoid biosynthesis and seed dormancy by interaction with bHLH proteins but not with MYB proteins. Hortic Res 5:1–18. https://doi.org/10.1038/s41438-018-0032-3
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930. https://doi.org/10.1016/j.plaphy.2010.08.016
Goyal S, Lambert C, Cluzet S, Mérillon JM, Ramawat KG (2012) Secondary metabolites and plant defence. In Plant Defence: Biol Control 12:109–138. https://doi.org/10.1007/978-94-007-1933-0_5
Gratani L (1992) A non-destructive method to determine chlorophyll content of leaves. Photosynthetica 26:469–473. https://doi.org/10.1007/BF00049537
Guo X, Chai W, Bai J, Ma ZR (2019) Cloning and bioinformatics analysis of AvFLS gene from Apocynum venetum. Mol Plant Breed 17:4978–4985. https://doi.org/10.13271/j.mpb.017.004978
Guo Y, Gao C, Wang M, Fu FF, El-Kassaby YA, Wang T et al (2020) Metabolome and transcriptome analyses reveal flavonoids biosynthesis differences in Ginkgo biloba associated with environmental conditions. Ind Crops Prod 158:112963. https://doi.org/10.1016/j.indcrop.2020.112963
Guo X, Wang Z, Cai D, Song L, Bai JL (2022) The chloroplast genome sequence and phylogenetic analysis of Apocynum venetum L. PLoS ONE 17:e0261710. https://doi.org/10.1371/journal.pone.0261710
Hao S, Wang Y, Yan Y, Liu Y, Wang J, Chen S (2021) A review on plant responses to salt stress and their mechanisms of salt resistance. Horticulturae 7:132. https://doi.org/10.3390/HORTICULTURAE7060132
Hare PD, Cress WA (1997) Metabolic implications of stress-induced proline accumulation in plants. Plant Growth Regul 21:79–102. https://doi.org/10.1023/A:1005703923347
Hartmann T (2007) From waste products to ecochemicals: fifty years research of plant secondary metabolism. Phytochemistry 68:2831–2846. https://doi.org/10.1016/j.phytochem.2007.09.017
Heath RL, Packer L (1968) Photoperoxidation in Isolated Chloroplasts: I. Kinetics and Stoichiometry of Fatty Acid Peroxidation. Arch Biochem Biophys 125:189–198. https://doi.org/10.1016/0003-9861(68)90654-1
Holton TA, Cornish EC (1995) Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell 7:1071–1083. https://doi.org/10.1105/tpc.7.7.1071
Holton TA, Brugliera F, Tanaka Y (1993) Cloning and expression of flavonol synthase from Petunia hybrida. Plant J 4:1003–1010. https://doi.org/10.1046/j.1365-313x.1993.04061003.x
Hong Z, Lakkineni K, Zhang Z, Verma DPS (2000) Removal of feedback inhibition of Δ1-pyrroline-5-carboxylate synthetase results in increased proline accumulation and protection of plants from osmotic stress. Plant Physiol 122:1129–1136. https://doi.org/10.1104/pp.122.4.1129
Hou M, Zhang Y, Mu G, Cui S, Yang X, Liu L (2020) Molecular cloning and expression characterization of flavonol synthase genes in peanut (Arachis hypogaea). Sci Rep 10:17717. https://doi.org/10.1038/s41598-020-74763-w
Jeyaraj A, Liu S, Zhang X, Zhang R, Shangguan M, Wei C (2017) Genome-wide identification of microRNAs responsive to Ectropis oblique feeding in tea plant (Camellia sinensis L.). Sci Rep 7:1–16. https://doi.org/10.1038/s41598-017-13692-7
Jiang X, Shi Y, Fu Z, Li WW, Lai S, Wu Y et al (2020) Functional characterization of three flavonol synthase genes from Camellia sinensis: Roles in flavonol accumulation. Plant Sci 300:110632. https://doi.org/10.1016/j.plantsci.2020.110632
Jiang L, Wu X, Zhao Z, Zhang K, Tanveer M, Wang L et al (2021) Luobuma (Apocynum)–Cash crops for saline lands. Ind Crops Prod 173:114146. https://doi.org/10.1016/j.indcrop.2021.114146
Kavi Kishor PB, Sreenivasulu N (2014) Is proline accumulation per se correlated with stress tolerance or is proline homeostasis a more critical issue? Plant, Cell Environ 37:300–311. https://doi.org/10.1111/pce.12157
Kawai Y, Ono E, Mizutani M (2014) Evolution and diversity of the 2–oxoglutarate-dependent dioxygenase superfamily in plants. Plant J 78:328–343. https://doi.org/10.1111/tpj.12479
Khare E, Mishra J, Arora NK (2018) Multifaceted interactions between endophytes and plant: developments and prospects. Front Microbiol 9:2732. https://doi.org/10.3389/fmicb.2018.02732
Kim BG, Kim JH, Kim J, Lee C, Ahn J (2008) Accumulation of flavonols in response to ultraviolet-B irradiation in soybean is related to induction of flavanone 3-beta-hydroxylase and flavonol synthase. Mol Cells 25:247–252. https://doi.org/10.1016/j.febslet.2008.03.057
Kim YB, Kim K, Kim Y, Tuan PA, Kim HH, Cho JW et al (2014) Cloning and characterization of a flavonol synthase gene from Scutellaria baicalensis. Sci World J 2014:980740. https://doi.org/10.1155/2014/980740
Koes RE, Quattrocchio F, Mol JN (1994) The flavonoid biosynthetic pathway in plants: function and evolution. BioEssays 16:123–132. https://doi.org/10.1002/bies.950160209
Kuhn BM, Geisler M, Bigler L, Ringli C (2011) Flavonols accumulate asymmetrically and affect auxin transport in Arabidopsis. Plant Physiol 156:585–595. https://doi.org/10.1104/pp.111.175976
Lee J, Koo N, Min DB (2004) Reactive oxygen species, aging, and antioxidative nutraceuticals. Compr Rev Food Sci Food Saf 3:21–33. https://doi.org/10.1111/j.1541-4337.2004.tb00058.x
Li C, Bai Y, Li S, Chen H, Han X, Zhao H et al (2012) Cloning, characterization, and activity analysis of a flavonol synthase gene FtFLS1 and its association with flavonoid content in tartary buckwheat. J Agric Food Chem 60:5161–5168. https://doi.org/10.1021/jf205192q
Li X, Kim YB, Kim Y, Zhao S, Kim HH, Chung E et al (2013) Differential stress-response expression of two flavonol synthase genes and accumulation of flavonols in tartary buckwheat. J Plant Physiol 170:1630–1636. https://doi.org/10.1016/j.jplph.2013.06.010
Li Q, Yu HM, Meng XF, Lin JS, Li YJ, Hou BK (2018) Ectopic expression of glycosyltransferase UGT 76E11 increases flavonoid accumulation and enhances abiotic stress tolerance in Arabidopsis. Plant Biol 20:10–19. https://doi.org/10.1111/plb.12627
Li P, Xu Y, Zhang Y, Fu J, Yu S, Guo H et al (2020) Metabolite profiling and transcriptome analysis revealed the chemical contributions of tea trichomes to tea flavors and tea plant defenses. J Agric Food Chem 68:11389–11401. https://doi.org/10.1021/acs.jafc.0c04075
Li P, Lei K, Liu L, Zhang G, Ge H, Zheng C et al (2021) Identification and functional characterization of a new flavonoid synthase gene MdFLS1 from apple. Planta 253:1–15. https://doi.org/10.1007/S00425-021-03615-2
Liu W, Xiao Z, Fan C, Jiang N, Meng X, Xiang X (2018) Cloning and characterization of a flavonol synthase gene from Litchi chinensis and its variation among litchi cultivars with different fruit maturation periods. Front Plant Sci 9:567. https://doi.org/10.3389/fpls.2018.00567
Liu W, Feng Y, Yu S, Fan Z, Li X, Li J et al (2021) The flavonoid biosynthesis network in plants. Int J Mol Sci 22:12824. https://doi.org/10.3390/IJMS222312824
Long SP, Zhu XG, Naidu SL, Ort DR (2006) Can improvement in photosynthesis increase crop yields? Plant, Cell Environ 29:315–330. https://doi.org/10.1111/j.1365-3040.2005.01493.x
Lou Q, Liu Y, Qi Y, Jiao S, Tian F, Jiang L et al (2014) Transcriptome sequencing and metabolite analysis reveals the role of delphinidin metabolism in flower color in grape hyacinth. J Exp Bot 65:3157–3164. https://doi.org/10.1093/jxb/eru168
Luo P, Ning G, Wang Z, Shen Y, Jin H, Li P et al (2016) Disequilibrium of flavonol synthase and dihydroflavonol-4-reductase expression associated tightly to white vs. red color flower formation in plants. Front Plant Sci 6:1257. https://doi.org/10.3389/fpls.2015.01257
Ma D, Sun D, Wang C, Li Y, Guo T (2014) Expression of flavonoid biosynthesis genes and accumulation of flavonoid in wheat leaves in response to drought stress. Plant Physiol Biochem 80:60–66. https://doi.org/10.1016/j.plaphy.2014.03.024
Maggio A, Miyazaki S, Veronese P, Fujita T, Ibeas JI, Damsz B et al (2002) Does proline accumulation play an active role in stress-induced growth reduction? Plant J 31:699–712. https://doi.org/10.1046/j.1365-313X.2002.01389.x
Mahajan M, Ahuja PS, Yadav SK (2011) Post-transcriptional silencing of flavonol synthase mRNA in tobacco leads to fruits with arrested seed set. PLoS ONE 6:e28315. https://doi.org/10.1371/journal.pone.0028315
Martens S, Preuß A, Matern U (2010) Multifunctional flavonoid dioxygenases: flavonol and anthocyanin biosynthesis in Arabidopsis thaliana L. Phytochemistry 71:1040–1049. https://doi.org/10.1016/j.phytochem.2010.04.016
Morimoto T, Tao R (2016) Functional characterization of Prunus SLFLs in transgenic Petunia. Acta Hortic 1231:85–90. https://doi.org/10.17660/actahortic.2019.1231.15
Muir SR, Collins GJ, Robinson S, Hughes S, Bovy A, Ric De Vos CH et al (2001) Overexpression of petunia chalcone isomerase in tomato results in fruit containing increased levels of flavonols. Nat Biotechnol 19:470–474. https://doi.org/10.1038/88150
Mukami A, Ngetich A, Mweu C, Oduor RO, Muthangya M, Mbinda WM (2019) Differential characterization of physiological and biochemical responses during drought stress in finger millet varieties. Physiol Mol Biol Plants 25:837–846. https://doi.org/10.1007/s12298-019-00679-z
Nguyen HN, Kim JH, Hyun WY, Nguyen NT, Hong SW, Lee H (2013) TTG1-mediated flavonols biosynthesis alleviates root growth inhibition in response to ABA. Plant Cell Rep 32:503–514. https://doi.org/10.1007/s00299-012-1382-1
Nguyen NH, Kim JH, Kwon J, Jeong CY, Lee W, Lee D et al (2016) Characterization of Arabidopsis thaliana FLAVONOL SYNTHASE 1 (FLS1)-overexpression plants in response to abiotic stress. Plant Physiol Biochem 103:133–142. https://doi.org/10.1016/j.plaphy.2016.03.010
Ohno S, Hosokawa M, Kojima M, Kitamura Y, Hoshino A, Tatsuzawa F et al (2011) Simultaneous post-transcriptional gene silencing of two different chalcone synthase genes resulting in pure white flowers in the octoploid dahlia. Planta 234:945–958. https://doi.org/10.1007/s00425-011-1456-2
Owens DK, Alerding AB, Crosby KC, Bandara AB, Westwood JH, Winkel BS (2008) Functional analysis of a predicted flavonol synthase gene family in Arabidopsis. Plant Physiol 147:1046–1061. https://doi.org/10.1104/pp.108.117457
Park YK, Koo MH, Ikegaki M, Contado JL (1997) Comparison of the flavonoid aglycone contents of Apis mellifera propolis from various regions of Brazil. Arq Biol Tecnol 40:97–106. https://doi.org/10.1007/s002849900274
Park S, Kim DH, Park BR, Lee JY, Lim SH (2019) Molecular and functional characterization of Oryza sativa flavonol synthase (OsFLS), a bifunctional dioxygenase. J Agric Food Chem 67:7399–7409. https://doi.org/10.1021/acs.jafc.9b02142
Parvanova D, Ivanov S, Konstantinova T, Karanov E, Atanassov A, Tsvetkov T et al (2004) Transgenic tobacco plants accumulating osmolytes show reduced oxidative damage under freezing stress. Plant Physiol Biochem 42:57–63. https://doi.org/10.1016/j.plaphy.2003.10.007
Preuß A, Stracke R, Weisshaar B, Hillebrecht A, Matern U, Martens S (2009) Arabidopsis thaliana expresses a second functional flavonol synthase. FEBS Lett 583:1981–1986. https://doi.org/10.1016/j.febslet.2009.05.006
Quick WP, Chaves MM, Wendler R, David M, Rodrigues ML, Passaharinho JA et al (1992) The effect of water stress on photosynthetic carbon metabolism in four species grown under field conditions. Plant, Cell Environ 15:25–35. https://doi.org/10.1111/j.1365-3040.1992.tb01455.x
Reginato MA, Castagna A, Furlán A, Castro S, Ranieri A, Luna V (2014) Physiological responses of a halophytic shrub to salt stress by Na2SO4 and NaCl: oxidative damage and the role of polyphenols in antioxidant protection. AoB Plants 6:plu042. https://doi.org/10.1093/aobpla/plu042
Richards RA (2000) Selectable traits to increase crop photosynthesis and yield of grain crops. J Exp Bot 51:447–458. https://doi.org/10.1093/jexbot/51.suppl_1.447
Sánchez-Rodríguez E, Moreno DA, Ferreres F, del Mar Rubio-Wilhelmi M, Ruiz JM (2011) Differential responses of five cherry tomato varieties to water stress: changes on phenolic metabolites and related enzymes. Phytochemistry 72:723–729. https://doi.org/10.1016/j.phytochem.2011.02.011
Saslowsky DE, Warek U, Winkel BS (2005) Nuclear localization of flavonoid enzymes in Arabidopsis. J Biol Chem 280:23735–23740. https://doi.org/10.1074/jbc.M413506200
Shah A, Smith DL (2020) Flavonoids in agriculture: Chemistry and roles in, biotic and abiotic stress responses, and microbial associations. Agronomy 10:1209. https://doi.org/10.3390/agronomy10081209
Sharma A, Shahzad B, Rehman A, Bhardwaj R, Landi M, Zheng B (2019) Response of phenylpropanoid pathway and the role of polyphenols in plants under abiotic stress. Mol Online 24:2452. https://doi.org/10.3390/molecules24132452
Shi J, Li W, Gao Y, Wang B, Li Y, Song Z (2017) Enhanced rutin accumulation in tobacco leaves by overexpressing the NtFLS2 gene. Biosci Biotechnol Biochem 81:1721–1725. https://doi.org/10.1080/09168451.2017.1353401
Singh PK, Gautam S (2013) Role of salicylic acid on physiological and biochemical mechanism of salinity stress tolerance in plants. Acta Physiol Plant 35:2345–2353. https://doi.org/10.1007/s11738-013-1279-9
Stracke R, De Vos RC, Bartelniewoehner L, Ishihara H, Sagasser M, Martens S et al (2009) Metabolomic and genetic analyses of flavonol synthesis in Arabidopsis thaliana support the in vivo involvement of leucoanthocyanidin dioxygenase. Planta 229:427–445. https://doi.org/10.1007/s00425-008-0841-y
Tanaka Y, Sasaki N, Ohmiya A (2008) Biosynthesis of plant pigments: anthocyanins, betalains and carotenoids. Plant J 54:733–749. https://doi.org/10.1111/j.1365-313X.2008.03447.x
Tattini M, Galardi C, Pinelli P, Massai R, Remorini D, Agati G (2004) Differential accumulation of flavonoids and hydroxycinnamates in leaves of Ligustrum vulgare under excess light and drought stress. New Phytol 163:547–561. https://doi.org/10.1111/J.1469-8137.2004.01126.X
Thakur M, Bhattacharya S, Khosla PK, Puri S (2019) Improving production of plant secondary metabolites through biotic and abiotic elicitation. J Appl Res Med Aromat Plants 12:1–12. https://doi.org/10.1016/j.jarmap.2018.11.004
Toh HC, Wang SY, Chang ST, Chu FH (2013) Molecular cloning and characterization of flavonol synthase in Acacia confusa. Tree Genet Genomes 9:85–92. https://doi.org/10.1007/s11295-012-0536-1
Verhoeyen ME, Bovy A, Collins G, Muir S, Robinson S, De Vos CHR et al (2002) Increasing antioxidant levels in tomatoes through modification of the flavonoid biosynthetic pathway. J Exp Bot 53:2099–2106. https://doi.org/10.1093/jxb/erf044
Vickers CE, Gershenzon J, Lerdau MT, Loreto F (2009) A unified mechanism of action for volatile isoprenoids in plant abiotic stress. Nat Chem Biol 5:283–291. https://doi.org/10.1038/nchembio.158
Vu TT, Jeong CY, Nguyen HN, Lee D, Lee SA, Kim JH et al (2015) Characterization of Brassica napus flavonol synthase involved in flavonol biosynthesis in Brassica napus L. J Agric Food Chem 63(35):7819–7829. https://doi.org/10.1021/acs.jafc.5b02994
Wang M, Qin L, Xie C, Li W, Yuan J, Kong L et al (2014) Induced and constitutive DNA methylation in a salinity-tolerant wheat introgression line. Plant Cell Physiol 55:1354–1365. https://doi.org/10.1093/pcp/pcu059
Wang F, Kong W, Wong G, Fu L, Peng R, Li Z et al (2016) AtMYB12 regulates flavonoids accumulation and abiotic stress tolerance in transgenic Arabidopsis thaliana. Mol Genet Genomics 291:1545–1559. https://doi.org/10.1007/s00438-016-1203-2
Wang M, Ren T, Huang R, Li Y, Zhang C, Xu Z (2021a) Overexpression of an Apocynum venetum flavonols synthetase gene confers salinity stress tolerance to transgenic tobacco plants. Plant Physiol Biochem 162:667–676. https://doi.org/10.1016/j.plaphy.2021.03.034
Wang M, Zhang Y, Zhu C, Yao X, Zheng Z, Tian Z et al (2021b) EkFLS overexpression promotes flavonoid accumulation and abiotic stress tolerance in plant. Physiol Plant 172:1966–1982. https://doi.org/10.1111/ppl.13407
Wang Y, Shi Y, Li K, Yang D, Liu N, Zhang L et al (2021c) Roles of the 2-Oxoglutarate-Dependent Dioxygenase Superfamily in the Flavonoid Pathway: A Review of the Functional Diversity of F3H, FNS I, FLS, and LDOX/ANS. Mol Online 26:6745. https://doi.org/10.3390/molecules26216745
Wang Y, Zhou LJ, Wang Y, Liu S, Geng Z, Song A et al (2021d) Functional identification of a flavone synthase and a flavonol synthase genes affecting flower color formation in Chrysanthemum morifolium. Plant Physiol Biochem 166:1109–1120. https://doi.org/10.1016/j.plaphy.2021.07.019
Wang D, Dong W, Murray J, Wang E (2022a) Innovation and appropriation in mycorrhizal and rhizobial symbioses. Plant Cell 34:1573–1599. https://doi.org/10.1093/plcell/koac039
Wang Z, Liu A, Liu J, Huang X, Xiao F, Tian M et al (2022b) Substrates and Loaded Iron Ions Relative Position Influence the Catalytic Characteristics of the Metalloenzymes Angelica archangelica Flavone Synthase I and Camellia sinensis Flavonol Synthase. Front Pharmacol 13:902672. https://doi.org/10.3389/FPHAR.2022.902672
Xie W, Zhang X, Wang T, Hu J (2012) Botany, traditional uses, phytochemistry and pharmacology of Apocynum venetum L. (Luobuma): A review. J Ethnopharmacol 141:1–8. https://doi.org/10.1016/j.jep.2012.02.003
Xu F, Li L, Zhang W, Cheng H, Sun N, Cheng S et al (2012) Isolation, characterization, and function analysis of a flavonol synthase gene from Ginkgo biloba. Mol Biol Rep 39:2285–2296. https://doi.org/10.1007/s11033-011-0978-9
Xu N, Liu S, Lu Z, Pang S, Wang L, Wang L et al (2020a) Gene expression profiles and flavonoid accumulation during salt stress in Ginkgo biloba seedlings. Plants 9:1162. https://doi.org/10.3390/plants9091162
Xu X, Gong J, Zhang T, Li Z, Zhang J, Wang L et al (2020b) Insights into antibacterial mechanism of Apocynum venetum L. fiber: Evolution of bioactive natural substances in bast during chemical degumming process. Ind Crops Prod 151:112419. https://doi.org/10.1016/j.indcrop.2020.112419
Xu Z, Wang M, Ren T, Li K, Li Y, Marowa P et al (2021) Comparative transcriptome analysis reveals the molecular mechanism of salt tolerance in Apocynum venetum. Plant Physiol Biochem 167:816–830. https://doi.org/10.1016/j.plaphy.2021.08.043
Yadav B, Jogawat A, Rahman MS, Narayan OP (2021) Secondary metabolites in the drought stress tolerance of crop plants: A review. Gene Rep 23:101040. https://doi.org/10.1016/j.genrep.2021.101040
Yang J, Zhang L, Jiang L, Zhan YG, Fan GZ (2021) Quercetin alleviates seed germination and growth inhibition in Apocynum venetum and Apocynum pictum under mannitol-induced osmotic stress. Plant Physiol Biochem 159:268–276. https://doi.org/10.1016/j.plaphy.2020.12.025
Yemm EW, Willis AJ (1954) The estimation of carbohydrates in plant extracts by anthrone. Biochem J 57:508–514. https://doi.org/10.1042/bj0570508
Yonekura-Sakakibara K, Higashi Y, Nakabayashi R (2019) The origin and evolution of plant flavonoid metabolism. Front Plant Sci 10:943. https://doi.org/10.3389/fpls.2019.00943
Yu D, Huang T, Tian B, Zhan J (2020) Advances in biosynthesis and biological functions of proanthocyanidins in horticultural plants. Foods 9:1774. https://doi.org/10.3390/foods9121774
Yu Z, Dong W, Teixeira da Silva JA, He C, Si C, Duan J (2021) Ectopic expression of DoFLS1 from Dendrobium officinale enhances flavonol accumulation and abiotic stress tolerance in Arabidopsis thaliana. Protoplasma 258:803–815. https://doi.org/10.1007/s00709-020-01599-6
Yuan Y, Qi L, Yang J, Wu C, Liu Y, Huang L (2015) A Scutellaria baicalensis R2R3-MYB gene, SbMYB8, regulates flavonoid biosynthesis and improves drought stress tolerance in transgenic tobacco. Plant Cell, Tissue Organ Cult 120:961–972. https://doi.org/10.1007/s11240-014-0650-x
Zahra R, Fatemeh N, Ali AM (2018) Effects of salinity stress on some growth, physiological, biochemical parameters and nutrients in two pistachio ( Pistacia vera L.) rootstocks. J Plant Interact 13:73–82. https://doi.org/10.1080/17429145.2018.1424355
Zhang C, Liu H, Jia C, Liu Y, Wang F, Wang J (2016a) Cloning, characterization and functional analysis of a flavonol synthase from Vaccinium corymbosum. Trees 30:1595–1605. https://doi.org/10.1007/s00468-016-1393-6
Zhang X, Henriques R, Lin S, Niu Q, Chua N (2016b) Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat Protoc 1:641–646. https://doi.org/10.1038/nprot.2006.97
Zhang C, Li X, Zhan Z, Cao L, Zeng A, Chang G et al (2018) Transcriptome sequencing and metabolism analysis reveals the role of cyanidin metabolism in dark-red onion (Allium cepa L.) bulbs. Sci Rep 8:1–10. https://doi.org/10.1038/s41598-018-32472-5
Zhao D, Jiang Y, Ning C, Meng J, Lin S, Ding W et al (2014) Transcriptome sequencing of a chimaera reveals coordinated expression of anthocyanin biosynthetic genes mediating yellow formation in herbaceous peony (Paeonia lactiflora Pall.). BMC Genomics 15:689. https://doi.org/10.1186/1471-2164-15-689
Zhou XW, Fan ZQ, Chen Y, Zhu YL, Li JY, Yin HF (2013) Functional analyses of a flavonol synthase–like gene from Camellia nitidissima reveal its roles in flavonoid metabolism during floral pigmentation. J Biosci 38:593–604. https://doi.org/10.1007/s12038-013-9339-2
Zhu XG, Long SP, Ort DR (2008) What is the maximum efficiency with which photosynthesis can convert solar energy into biomass? Curr Opin Biotechnol 19:153–159. https://doi.org/10.1016/j.copbio.2008.02.004
Zhu XG, Long SP, Ort DR (2010) Improving photosynthetic efficiency for greater yield. Annu Rev Plant Biol 61:235–261. https://doi.org/10.1146/annurev-arplant-042809-112206
Acknowledgements
We thank LetPub for linguistic assistance and pre-submission expert review.
Funding
This research was supported by the Fundamental Research Funds for the Central Universities(31920220138) and the National Natural Science Foundation of China (Grant No. 31760242).
Author information
Authors and Affiliations
Contributions
XG and DC contributed substantially to the experimental design, conceived the study, and approved the final manuscript. XG contributed to funding, interpreted the data, authored or reviewed the draft, and revised the manuscript. JL and DC carried out mainly experiments, finished the draft of the manuscript, and comprehensively analyzed the data from all experimental results. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Conflict of Interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guo, X., Li, J. & Cai, D. Overexpression of a Flavonol Synthase Gene from Apocynum venetum Improves the Salinity Stress Tolerance of Transgenic Arabidopsis thaliana. J Soil Sci Plant Nutr 24, 2317–2333 (2024). https://doi.org/10.1007/s42729-023-01590-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42729-023-01590-z