Abstract
China has experienced a notorious increase in nitrogen (N) deposition as a result of anthropogenic activities, particularly in temperate areas. While aboveground biodiversity has been extensively studied, the impact of long-term N deposition on the diversity, composition, and function of the soil microbiome remains largely unexplored. In this study, we evaluated alterations in the diversity, composition, and function of soil bacterial and fungal communities in response to varying levels of N deposition (LN = low N addition, 40 kg N ha−1 yr−1; HN = high N addition, 80 kg N ha−1 yr−1) using Illumina MiSeq sequencing technology in a temperate natural wetland. N deposition had no discernible impact on bacterial α diversity, whereas fungal α diversity exhibited a significant decrease in response to high N addition only. Additionally, N deposition led to a notable increase in the relative abundance of the bacterial phylum Patescibacteria but a decrease in Latescibacteria. The relative abundance of Epsilonbacteraeota was highest in the unamended plots and lowest in the low N addition plots. Furthermore, N addition significantly increased the relative abundance of Ascomycota while decreasing that of Mortierellomycota, with no significant effect observed on Basidiomycota. Structural equation modeling (SEM) indicated that soil organic carbon (SOC), and total and available N were the two primary drivers shaping bacterial and fungal communities. Our study demonstrated that bacterial communities were less responsive to N addition compared to fungal communities, emphasizing the significance of simultaneously evaluating the soil microbiome in response to global changes.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Atmospheric nitrogen (N) deposition primarily results from the emission of excessive reactive N compounds and the subsequent atmospheric transport processes. The increase in N deposition has been significantly driven by industrial and agricultural activities (Gruber and Galloway 2008; Song et al. 2014; Wang et al. 2018). This increase in N can potentially enhance plant primary productivity by increasing N content (LeBauer and Treseder 2008). However, N may also have adverse effects on ecosystems by altering vegetation composition and soil nutrient cycling (Roth et al. 2013; Zhu et al. 2015). N deposition can influence the N cycle at regional and global scales through both dry and wet deposition, thereby impacting the global ecosystem’s biogeochemical cycles (Holland et al. 2005; Fowler et al. 2013). Therefore, atmospheric N deposition has garnered increasing attention in the context of global change ecology. Soil microorganisms are pivotal components within soil ecosystems (Frey et al. 2020) and play essential roles in energy flow, biogeochemical cycling, litter decomposition, and various climate-related processes (Lakshmi et al. 2020; Basu et al. 2021). Alterations in soil microbial composition and function in response to N deposition can lead to critical challenges for ecosystem function and services. Nevertheless, although several studies have characterized the responses of the soil microbiome to N deposition in grassland and forest ecosystems (Yan et al. 2017; Frey et al. 2020, 2023; He et al. 2021; Wang et al. 2021), research on wetland ecosystems subject to long-term N deposition, especially in temperate freshwater wetlands, remains relatively limited.
N deposition can directly or indirectly influence the structure and function of the soil microbiome by altering various edaphic parameters (Nie et al. 2018; Sui et al. 2022). Microorganisms reliant on inorganic N as an energy source may proliferate with increasing N deposition, potentially leading to an increase in microbial diversity (Wang et al. 2021). However, the abundance of some oligotrophic microorganisms may decrease due to increased N availability (Frey et al. 2023), resulting in decreased microbial diversity. Moreover, N deposition can indirectly impact soil microbial functions, compositions, and diversities by altering above-ground vegetation composition and soil properties (Treseder 2008). Lu et al. (2021) observed changes in the α-diversity and composition of soil microbial communities with increasing N deposition in a temperate coastal wetland of China. In contrast, Hu et al. (2022) reported that N deposition did not alter soil microbial community diversity but significantly changed the structure of the soil bacterial community in a temperate swamp meadow wetland of China. These two examples illustrate the persisting contradictions regarding the impact of N deposition on the structure, function, and diversity of soil microbial communities in temperate wetlands. Furthermore, N deposition affects not only the soil microbial composition and abundance but also elicits varying responses in bacterial and fungal composition and abundance (Li et al. 2019). Previous studies have reported that N deposition influences the community composition of ectomycorrhizal fungi in temperate beech forests and subtropical pine plantations (De Witte et al. 2017; Ning et al. 2018). Another study reported that N deposition increased the relative abundance of Basidiomycetes and decreased that of Ascomycetes in a northern hardwood forest soil (Edwards et al. 2011). In temperate steppe and tropical forest soils, N deposition affected bacterial community composition by enhancing the relative abundance of Proteobacteria, Actinobacteria, and Firmicutes while reducing the relative abundance of Acidobacteria and Verrucomicrobia (Zeng et al. 2016; Nie et al. 2018). In contrast, DeForest et al. (2004) reported that elevated N deposition had no observable influence of on the abundance and composition of bacterial or fungal communities in a hardwood forest. Weng et al. (2022) reported that N deposition significantly impacted the metabolic functional potential of bacterial and fungal communities in wetland soils, promoting various metabolic processes of the microbial community, particularly carbohydrate and amino acid-related metabolic processes and archaea-mediated ammonia oxidation. However, N deposition reduced N2 fixation and signal transduction. Therefore, uncertainties persist regarding the effects of N deposition on the composition, abundance, function, and diversity of soil microbial communities.
Wetlands are crucial amphibious ecosystems and are widely recognized as biodiversity hotspots (Sui et al. 2021). They serve a multitude of essential ecological functions, including regulating biogeochemical cycles, preserving biodiversity, and mitigating pollution (Junk et al. 2013; Cai et al. 2021; Sui et al. 2021). The Sanjiang Plain, located in northeastern China, is one of the most important freshwater wetland ecosystems (Weng et al. 2022). It is also China’s most pristine and well-preserved wetland, serving as a significant grain production region (Zhang et al. 2021). Moreover, the Sanjiang Plain plays a crucial role in ensuring national food security and participating in critical ecological processes (Li et al. 2020a, b). However, it is also severely affected by the high N deposition rates occurring in China. For example, Weng et al. (2022) observed a significant decline in soil microbial carbon utilization with increasing N deposition in the Sanjiang Plain. Zhang et al. (2018a, b, c, d) reported that N deposition notably constrained greenhouse gas emissions in Deyeuxia angustifolia wetlands in the Sanjiang Plain. Dai et al. (2018) demonstrated that high N addition significantly reduced plant diversity, whereas low N enhanced it. However, the influence of N addition on the soil microbial diversity, community, and function in temperate wetlands remains unclear.
In May 2010, we initiated a long-term experiment to simulate the increasing N deposition in a typical wetland located in the Sanjiang Plain, northeastern China. Following four consecutive years of elevated N deposition, we investigated the alterations in the diversity, structure, and function of soil microbial communities in response to N addition. Particularly, our objectives were to 1) elucidate the changes in diversity, composition, and function of soil bacterial and fungal communities within the wetland ecosystem; 2) identify which phyla or genera of soil microorganisms exhibited differential responses, such as mycorrhizal or saprophytic fungi; and 3) determine the critical soil physicochemical parameters that influence the composition and diversity of soil bacteria and fungi. Therefore, our study provides a valuable theoretical framework to support the ecological restoration of wetland ecosystems in the Sanjiang Plain facing elevated N deposition.
2 Materials and Methods
2.1 Research area description
This study was conducted at a field experimental station belonging to the Heilongjiang Academy of Sciences, situated within the Honghe National Nature Reserve (47°42′18″–47°52′07″N, 133°34′38″–133°46′29″E), Heilongjiang Province, China (Fig. 1). Since 2010, this site has been used for simulating N (NH4+-N and NO3−-N) deposition. The wetland area covers approximately 1.1 × 104 ha and is primarily dominated by Deyeuxia angustifolia wetlands (Zhang et al. 2018a, b, c, d). The study site exhibits a typical temperate humid/semi humid monsoon climate and exhibits an average annual temperature of 1.9 °C, with an average annual precipitation of 585 mm and evaporation of 1166 mm. The soil in this region is classified as typical bleached stagnant soil and fibrous organic soil according to the USDA soil taxonomy. The dominant vegetation species include Deyeuxia angustifolia, Glyceria spiculose, Carex lasiocarpa, and Carex pseudocuraica.
2.2 Experimental design
Three N deposition treatments were established at the field experimental station, namely, ambient [control (CK), 0 kg N ha−1 year−1], low N addition [(LN), 40 kg N ha−1 year−1], and high N addition [(HN), 80 kg N ha−1 year−1]. Each treatment was replicated in triplicate and randomly allocated with an area of 10 m × 10 m per treatment. The N application rates were selected to mimic a typical fertilization regimen used in local agricultural practices (Song et al. 2016). Each year in May, NH4NO3 was uniformly sprayed in each of the N deposition treatments, whereas the control treatment received an equivalent amount of pure water.
2.3 Soil sample collection and analysis
Following four years of simulated N deposition, soil samples from the top layer (approximately 0–20 cm depth) were collected from each N deposition treatment in 2014. Soil samples were collected using a 5-cm inner diameter auger, and 5–10 soil cores were randomly obtained from each plot after removing the litter layer. The soil samples from each plot were combined, placed in a Ziplock bag, and kept at 4 °C before being promptly transported to the laboratory. The soil samples were then homogenized using a 2-mm mesh size sieve and divided into two parts. One part was stored at − 80 °C for microbial community analysis, whereas the other part was air-dried for the determination of soil physicochemical properties.
2.4 Determine the soil physicochemical properties
Soil physicochemical properties were characterized as described in our previous study (Sui et al. 2022). Briefly, soil pH was determined by preparing a soil–water (deionized water) suspension at a 1:2.5 ratio (w v−1) and shaking it for 30 min. The pH was subsequently measured using a pH meter (Thermo Scientific Orion 3-Star Benchtop, Cambridge, United Kingdom). Soil organic carbon and total N were quantified using an elemental analyzer (Elementar, Langenselbold, Germany). Nitrate nitrogen (NO3−-N) and ammonium nitrogen (NH4+-N) levels were analyzed using a continuous flow analysis system (SAN + + , Skalar Analytical, Netherlands). Total phosphorus was determined using a spectrophotometer. Available phosphorus content was measured through a colorimetric method following extraction with 0.5 M NaHCO3. Soil moisture content (SMC) was assessed by comparing the fresh wet weight with the dry weight obtained after drying at 120 °C for 24 h.
2.5 DNA extraction and PCR amplification
Soil genomic DNA was extracted using the MOBIO soil DNA Isolation Kit (Catalog Number: 12888) from MOBIO (Carlsbad, CA, USA). For each sample, 0.5 g of fresh soil was used to extract total DNA according to the manufacturer’s instructions. The purity and concentration of the extracted DNA were determined prior to PCR amplification using a NanoDrop Spectrophotometer (NanoDrop Technologies Inc., DE, USA). All DNA concentrations were adjusted to 50 (ng μL−1) for subsequent PCR amplification. The amplifications were conducted using the 338F/806R primer pair for bacterial 16S rRNA and the ITS1F/ITS2R primer pair for fungal ITS1 rRNA. The PCR reaction mixture volume was 25 μL and consisted of 12.5 μL PCR Master Mix (TAKARA, Japan), 2.0 μL forward and reverse primers (5 μmol L−1), 2.0 μL DNA, and 6.5 μL of H2O. The PCR amplification program consisted of an initial denaturation step at 95 °C for 45 s, followed by 30 cycles at 95 °C for 45 s, 52 °C (for bacteria) for 45 s or 55 °C (for fungi) for 50 s, and 72 °C for 60 s, and a final extension step at 72 °C for 10 min. Each PCR was performed in triplicate. The PCR products were visualized using 2% agarose gel electrophoresis and then purified using the AXYPrep DNA purification Kit from Axygen Biosciences (CA, United States). Subsequently, the three replicate PCR products for each sample were pooled together and sequenced using the Illumina MiSeq platform (2 × 300 bp) at Majorbio Bio-pharm Technology Co., Ltd. (Shanghai, China). All raw sequence data were deposited into the NCBI Sequence Read Archive database under the following BioProject accession numbers: PRJNA960933.
2.6 Bioinformatic analysis
The bioinformatic analyses of the raw fasta sequences from bacteria and fungi was conducted using the online platform provided by Majorbio Cloud Platform (www.majorbio.com) (Ren et al. 2022). Briefly, the raw PE reads were merged using the Flash software (version 1.2.11). The barcode primers were trimmed from the merged PE reads, and dereplication was performed using QIIME (version 1.9.1). Chimeric sequences were identified and removed from the dataset using Vsearch (Rognes et al. 2016). Operational taxonomic units (OTUs) were detected at a 97% similarity threshold using the Uparse (Edgar 2010) software. Taxonomic classification was carried out using the SILVA (version 138.1) and UNITE (version 8.0) databases for bacteria and fungi, respectively. A threshold value of 0.8 was used for classification in QIIME1 (Caporaso et al. 2010). Prior to estimating α-diversity, the dataset was rarefied to the lowest number of reads present in all soil samples.
2.7 Statistical analysis
The data analysis was also conducted on the Majorbio Cloud Platform (www.majorbio.com). Briefly, soil bacterial and fungal alpha diversity indices (Chao1, Shannon, and Simpson) were computed using QIIME1 (Caporaso et al. 2010). Principal coordinates analysis (PCoA) and permutational multivariate analysis of variance (PERMANOVA) were conducted based on Bray–Curtis dissimilarity at the OTU level on the online platform. This analysis enables the visualization and assessment of differences in community composition between samples. Venn diagrams representing shared and unique OTUs were generated at the OTU level on the online platform. Cladogram diagrams illustrating taxonomic relationships from phylum to species were generated on the online platform. Heatmaps displaying the 30 most abundant bacterial and fungal genera were generated using online tools (https://www.bioincloud.tech/). Correlation heatmaps showing relationships between soil physicochemical parameters and the 30 most abundant bacterial and fungal genera were also generated using online tools (https://www.bioincloud.tech/). Structural equation modeling (SEM), a multivariate statistical method for testing complex path-relation networks, was utilized to examine the direct and indirect effects of N deposition on soil bacterial and fungal community diversity and composition in the R software. An a priori model was constructed based on a literature review and our knowledge of how these predictors are related. Diversity was assessed using the Shannon diversity index, whereas composition was assessed via non-metric multidimensional scaling (NMDS). Model adequacy was determined by χ2 tests (P > 0.05), the goodness-of-fit index (CFI, > 0.9), and Akaike information criteria (AIC). Functional prediction of bacterial communities at the OTU level using FAPROTAX was conducted as described by Louca et al. (2016). Fungal guild predictions at the OTU level were performed using FUNGuild (Nguyen et al. 2016). Both FAPROTAX and FUNGuild analyses were conducted using online tools (https://cloud.majorbio.com/). Statistical significance was determined using Duncan’s multiple comparison method via one-way analysis of variance (ANOVA) with a significance level of 0.05. All relevant statistical analyses were conducted using the SPSS 19.0 software.
3 Results
3.1 Effects of simulated N deposition on soil physicochemical properties
N deposition significantly changed the soil physicochemical properties except TP (Table 1). Specifically, pH, AP, and SMC decreased in response to N, whereas TN, NO3−-N and NH4+-N increased with increased N deposition (Table 1). Furthermore, LN did not have any observable effects on SOC, but HN significantly increased the SOC levels compared to CK (Table 1).
3.2 Variations in the α- and β-diversities of soil bacterial and fungal communities under simulated N deposition
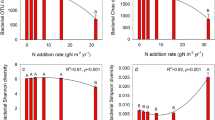
The bacterial α-indices, including Shannon, Chao1, and number of observed OTUs (Sobs), were not significantly different between the N levels (Table 2). However, the Chao1, ACE, and Sobs of fungal diversity were affected by N addition (Table 2). The fungal Chao1, ACE, and Sobs decreased significantly with increasing N addition (P < 0.05), whereas the fungal Shannon index did not change significantly (P > 0.05).
PCoA was conducted to visualize the β diversities of soil microbial communities (Fig. 2). Our findings demonstrated that N addition had a significant impact on the β diversity of fungal communities (Fig. 2b; PERMANOVA: P < 0.05). In contrast, the β diversity of bacterial communities (Fig. 2a) did not exhibit significant changes among the different levels of N addition (Fig. 2a, PERMANOVA: P > 0.05).
3.3 Effects of simulated N deposition on the soil bacterial and fungal compositions
The most dominant bacterial phyla in the LN and HN treatments did not exhibit significant changes compared to the CK (Fig. 3a). However, the relative abundance of Patescibacteria, Firmicutes, Latescibacteria, and Epsilonbacteraeota in the LN and HN treatments did show significant alterations in comparison to the CK (Fig. 3a). Specifically, the relative abundance of Patescibacteria increased significantly, whereas Firmicutes and Latescibacteria decreased significantly with N deposition (Fig. 3a). Notably, the relative abundance of Epsilonbacteraeota was highest in the CK and lowest in the LN treatment (Fig. 3a). Among the bacterial genera, Candidatus_Solibacter, Candidatus_Udaeobacter, Bryobacter, HSB_of53-F07, Candidatus_Koribacter, Bradyrhizobium, Acidotherium, Geobacter, Euin6067, and Halliangium were the most dominant genera across all N deposition levels (Fig. S3 a, relative abundance of top 10 genera). Interestingly, the relative abundance of Acidothermus, Anaeromyxobacter, Gaiella, Bacillus, and Opitutus in the LN and HN treatments exhibited significant differences compared to the CK (Fig. 4a). Specifically, the relative abundance of Acidothermus, Anaeromyxobacter, and Gaiella decreased significantly, whereas that of Opitutus increased with N addition (Fig. 4a).
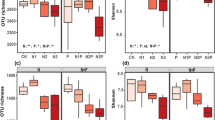
Differences in the relative abundance of bacterial (a) and fungal (b) phyla under different N deposition treatments. The asterisks indicate significant differences between three N depositions at the 0.05 level. Control: (CK), 0 kg N ha−1 year−1; low N addition: (LN), 40 kg N ha−1 year−1; high N addition (HN), 80 kg N ha−1 year−1
Differences in the relative abundance of bacterial (a) and fungal (b) genera under N deposition treatment. The asterisks indicate significant differences between three N depositions at the 0.05 level. Control: (CK), 0 kg N ha−1 year−1; low N addition: (LN), 40 kg N ha−1 year−1; high N addition (HN), 80 kg N ha−1 year−1
Regarding the fungal communities, the relative abundance of Ascomycota and Motierellomycota in the LN and HN treatments exhibited significant changes compared to the CK (Fig. 3b). Notably, N addition significantly increased the abundance of Ascomycota, while decreasing that of Mortierellomycota (Fig. 3b). Mortierella, Archaeorhizomyces, Clavaria, Lachnum, Penicillium, Pseudeurotium, Trichoderma, Acephala, Sordaria, and Arnium were the most dominant genera across all N deposition levels (Fig. S3b, relative abundance of top 10 genera). The relative abundance of Mortierella, Trichoderma, Sordaria, Leohumicola, Paraphaeosphaeria, Coniochaeta, Dimorphospora, Fayodia, Entoloma, and Metarhizium in the LN and HN treatments were significantly different compared to the CK group (Fig. 4b). Particularly, the relative abundances of Mortierella, Sordaria, Leohumicola, Paraphaeosphaeria, Coniochaeta, and Entoloma were higher in the CK group than those in the LN and HN treatments, whereas the relative abundances of Trichoderma, Fayodia, and Dimorphospora were lower in the CK group compared to those in the LN and HN treatments (Fig. 4b).
3.4 Effects of simulated N deposition on soil bacterial and fungal indicator species
The LEfSe analysis identified a total of 7 bacterial and 14 fungal taxa that displayed significant differences in relative abundance (as indicated by LDA effect size scores exceeding 3) among the three N addition treatments (Fig. 5).
3.5 Effects of different soil physicochemical parameters on soil bacterial and fungal communities
SEM was employed to evaluate both the direct and indirect effects of N deposition on the diversity and composition of soil bacterial and fungal communities (Fig. 6). The final model for soil bacteria met our significance criteria (χ2 = 14.65, df = 8, P = 0.07, CFI = 0.94, AIC = − 10.32). Notably, N deposition did not exhibit any significant direct effects on the diversity and composition of the soil bacterial community. However, it did show significant indirect effects through SOC (λ = 0.87, p < 0.05), NH4+ (λ = − 3.67, p < 0.05), NO3− (λ = 3.19, p < 0.05), and TN (λ = − 3.36, p < 0.05) on diversity and composition of soil bacterial community (Fig. 6a). Similarly, the final fungi model met our significance criteria (χ2 = 13.56, df = 9, P = 0.13, CFI = 0.96, AIC = 4.63). Moreover, N deposition exhibited significant indirect effects through SOC (λ = − 0.62, p < 0.05), NH4+ (λ = 2.62, p < 0.05), NO3− (λ = − 2.71, p < 0.05), and TN (λ = 1.79, p < 0.05) on the diversity and composition of the soil fungal community (Fig. 6b).
Path diagrams of the structural equation modeling (SEM) for the relationship of N deposition on the diversity (Shannon index) and composition (NMDS) of soil bacteria (a) and fungi (b). SOC, soil organic carbon; TN, total nitrogen; NH4+, ammonium nitrogen; NO3−, nitrate nitrogen. The values above the line represent the path coefficients. Solid lines indicate positive path coefficients, whereas dashed lines indicate negative path coefficients. * P < 0.05. Control: (CK), 0 kg N ha−1 year−1; low N addition: (LN), 40 kg N ha−1 year−1; high N addition (HN), 80 kg N ha−1 year−1
3.6 Impact of simulated N deposition on the predictions of potential functions
A total of 50 functional potential functions of soil bacterial communities were annotated using FAPROTAX (Fig. 7a), which were primarily associated with carbon (C) and nitrogen (N) processes. Among these functions, only the abundance of anoxygenic_photoautotrophy_H2_oxidizing bacteria within the soil bacterial communities exhibited significant changes across different N additions (Fig. 7a). Conversely, the other potential functions of soil bacterial communities did not show significant alterations (Fig. 7a). Notably, the abundance of anoxygenic_photoautotrophy_H2_oxidizing bacteria tended to decrease with increasing N addition (Fig. 7a).
Impact of N addition on the predictions of potential functions based FAPROTAX analysis for bacteria (a) and FUNGuild analysis for fungi (b). The color gradient (blue = low, red = high) represents the relative abundance of the soil bacterial and fungal functions among the different N addition treatments. The asterisks indicate the differences in different N addition treatments, according to one-way ANOVA at a 0.05 significance level. Control: (CK), 0 kg N ha−1 year−1; low N addition: (LN), 40 kg N ha−1 year−1; high N addition (HN), 80 kg N ha−1 year−1
Regarding the soil fungal communities, a total of 19 potential guilds were annotated using the FUNGuild database (Fig. 7b), primarily falling into the “Saprotrophs,” “Pathogens,” “Endophytes,” “Endomycorrhizal,” and “Arbuscular Mycorrhizal” categories. However, only the “Dung Saprotrophs” and “Litter Saprotrophs” categories exhibited significant changes among the different N additions (Fig. 7b, P < 0.05), whereas the other guilds did not exhibit significant differences among the three N additions (Fig. 7b). Interestingly, both the abundance of “Dung Saprotrophs” and “Litter Saprotrophs” decreased significantly from the CK to the HN treatment (Fig. 7b).
4 Discussion
In contrast to our initial hypothesis, our results demonstrated that elevated N deposition did not significantly alter soil bacterial α and β diversities, but it did lead to a significant reduction in fungal α diversity under high N (HN) deposition, as well as changes in fungal β diversities across the three N deposition levels. These results are inconsistent with previous studies that demonstrated a significant decrease in both soil bacterial and fungal α diversities under N deposition concentrations in coastal wetland ecosystems (Lu et al. 2021; Sun et al. 2022) and wetland forests (Li et al. 2019). However, there have been other conflicting findings, such as those reported by Bledsoe et al. (2020), who observed an increase in soil bacterial α diversity in oligotroph-dominated wetlands; Sun et al. (2022), who reported an increase in soil microbial α diversity in coastal wetlands; and Lee et al. (2015) and Hu et al. (2022), who reported no change in soil microbial α diversity under N deposition in marsh wetland ecosystems and swamp meadow wetland ecosystems. These variations suggest that the responses of α diversity in bacteria and fungi to N deposition can vary among different wetland ecosystems. Firstly, the Sanjiang plain wetland, a typical freshwater wetland, possesses distinct characteristics in terms of vegetation species and soil properties compared to other wetland ecosystems. This variation could lead to differences in the sensitivity of soil microorganisms to external N input (Hu et al. 2022). In nutrient-limited conditions, plants often rely on soil microorganisms to facilitate nutrient acquisition, and in return, plants provide carbon resources to support these soil-associated microorganisms (Bledsoe et al. 2020). Consequently, soil microbial α diversity could increase under N deposition in coastal wetlands due to their poor soil nutrient conditions. However, the Sanjiang plain wetland is not nutrient-limited (Chen et al. 2023), and exogenous N input may primarily be absorbed by above-ground vegetation, with limited facilitation by soil microorganisms. Secondly, the duration of N deposition can have a noticeable impact on soil microbial community diversity. For instance, soil bacterial community diversity may remain stable under short-term N deposition (McHugh et al. 2017) but decrease with long-term N deposition (Zhang et al. 2022). In our case, a four-year N deposition period may not have been sufficient to induce changes in soil bacterial community diversity. Thirdly, the rate of N deposition may also influence soil microbial community diversity. For example, Liu et al. (2020) observed that soil bacterial community diversity remained unchanged when N deposition levels were below 160 g⋅N⋅m–2⋅a–1. In our study, the N deposition levels (i.e. 40 g and 80 g N⋅m–2⋅a–1) were somewhat lower, suggesting that these amounts of N application had no significant effect on bacterial diversity. This could be attributed to the greater plasticity, higher population density, and greater adaptability of microbial communities. Lastly, prior research has established that soil pH plays a critical role in soil bacterial community diversity (Sui et al. 2022), and specific microbial strains exhibit optimal pH ranges (Bledsoe et al. 2020). However, our study demonstrated that pH did not change significantly, remaining within a range of 4.7 to 5.9. Consequently, this lack of significant pH variation may explain why N deposition did not impact soil bacterial community diversity. Conversely, soil fungal diversity appeared to be more sensitive to N deposition, likely due to soil fungi's higher carbon and nutrient assimilation efficiency compared to soil bacteria (Yang et al. 2020). Furthermore, the level of N addition directly affected soil fungal diversity. As indicated by Freitag et al. (2005), N addition effects on bacterial diversity can be either positive or negative, contingent upon the concentration and type of N added. Our study revealed that wetland soil fungal communities responded nonlinearly to N addition; specifically, they remained unaffected by N input below 40 g N⋅m–2⋅a–1, but decreased significantly when N input reached 80 g N⋅m–2⋅a–1. This phenomenon suggests that although wetland fungal communities can withstand low-level N inputs, further increases in N input may surpass a threshold, leading to a shift toward lower diversity within the fungal community.
As anticipated in our second hypothesis, the prevalence of the bacterial phyla Acidobacteria and Proteobacteria, consistent across all N levels, aligns with previous findings in various wetland ecosystems (Hartmann et al. 2017; Zhang et al. 2020a, b; Lu et al. 2021; Zhang et al. 2022). This reaffirms the notion that Acidobacteria and Proteobacteria exhibit a definitive adaptability to different rates of N deposition (Huang et al. 2020). These bacterial phyla play crucial roles in energy metabolism, nutrient cycling, and other ecological functions (Deng et al. 2019; Zhang et al. 2020a, b; Sui et al. 2021; Guo et al. 2022). Our previous study has also highlighted the importance of these bacterial phyla in maintaining soil bacterial communities in wetland ecosystems (Sui et al. 2022) and their dominance in wetland soils (Wang et al. 2019; Sui et al. 2019). However, the abundance of these predominant bacterial phyla did not exhibit significant changes. Only Patescibacteria increased, whereas Firmicutes and Latescibacteria decreased significantly with elevated N deposition.
Patescibacteria, consisting of many unknown bacteria, are often found in anoxic environments and are involved in hydrogen production and sulfur cycling (Frey et al. 2016). Studies by Ren et al. (2020) and Ma et al. (2021) have reported an increase in Patescibacteria with N addition, positively correlating with soil N concentration. This could explain the increase in the abundance of Patescibacteria with elevated N deposition. Firmicutes, traditionally considered copiotrophic bacterial phyla (Hartmann et al. 2017; Perez-Mon et al. 2022), have shown varying responses to N addition in previous research, with some studies reporting an increase (Freedman et al. 2013; Zhang et al. 2014a, b) and others reporting the opposite trend (Zhang et al. 2018a, b, c, d). This study, however, observed a decrease in Firmicutes with N addition. Therefore, these findings suggest that Firmicutes possess characteristics of both copiotrophs and oligotrophs. Firmicutes appear to be opportunistic bacteria capable of adjusting their trophic strategy according to varying nutritional concentrations across a range of ecosystems (Frey et al. 2016; Arcand et al. 2017). These opportunistic Firmicutes possibly have low competitiveness in utilizing N resources, and instead, their survival depends on various substrates, regardless of their availability.
Contrary to our second hypothesis, this study found that the relative abundance of Ascomycota significantly increased with N addition, while Basidiomycota remained unchanged. Some previous studies have reported negative effects of N deposition on the abundance of Ascomycota and positive effects on Basidiomycota in wetland ecosystems (Hu et al. 2022). Conversely, another study found that N deposition had no effect on Ascomycota but decreased the abundance of Basidiomycota in permafrost wetlands (Ma et al. 2021). These discrepancies are not entirely surprising, given the substantial differences in climate, vegetation, and soil properties across the studied ecosystems. The Sanjiang wetland, in particular, differs from coastal and permafrost wetlands not only in climate but also in characteristics such as being phosphorus-limited rather than N-limited (Song et al. 2016; Zhang et al. 2018a, b, c, d) and having lower pH and base cation concentrations compared to the other two wetlands (Li et al. 2020a, b).
In this study, the FAPROTAX method revealed that only anoxygenic_photoautotrophy_H2_oxidizing significantly decreased with N deposition, while other functions did not exhibit significant changes. Simultaneously, the FUNGuild method indicated significant differences in Saprotrophs among the N deposition levels. These findings suggest that alterations in bacterial and fungal compositions can influence soil carbon cycling functions and fungal activity (Frey et al. 2016). However, many studies have reported that N deposition can affect numerous soil microbial functions (Zheng and He 2020; Ma et al. 2021; Zhang et al. 2023). There are several potential explanations for these observations. Firstly, the high soil moisture content in wetland ecosystems, along with lower microbial activity compared to forest and grassland ecosystems, could contribute to these results. The high soil moisture content leads to low soil oxygen levels, which in turn promotes the predominance of saprotrophs among soil fungi. Secondly, the methods based on the FAPROTAX and FUNGuild databases for functional analysis may not capture the functions of certain microorganisms with unique roles, which could explain the inability to detect significant changes in various functions.
In summary, N deposition primarily affects the abundance of soil microorganisms by altering the life cycle strategies of relevant microorganisms, either directly or indirectly, through changes in soil nutrient concentrations (Lu et al. 2021; Ma et al. 2021). For instance, in the case of the Sanjiang plain, which faces a high risk of N saturation and aggravated phosphorus (P) limitation, substantial N addition may directly impact the abundance of copiotrophic fungi such as Ascomycota while inhibiting oligotrophic fungi such as Basidiomycota. Alternatively, N deposition may also alter the decomposition rate of litter, indirectly influencing the composition of the soil fungal community. Therefore, in the context of N addition in temperate wetlands, changes in the composition and diversity of soil microbial communities reflect shifts in the structure and function of wetland ecosystems.
Consistent with our third hypothesis, the soil physicochemical parameters that influence the composition of soil bacterial and fungal communities varied across different N deposition levels. Soil TN and NH4+-N significantly influenced the structure of the soil bacterial community, whereas soil SOC, NH4+-N, NO3−-N, and TN significantly affected the soil bacterial diversity (Fig. 6a). In contrast, the soil fungal community was significantly influenced by NH4+-N, NO3−-N, SOC, and TN (Fig. 6b). These results indicate that both soil bacteria and fungi exhibit close correlations with soil SOC, TN, NH4+-N, and NO3−-N. Additionally, the effects of soil physicochemical parameters on the abundance of dominant soil bacterial and fungal genera were also clearly different. The impact of N deposition on soil microbial community composition remains uncertain. N addition can directly or indirectly alter soil properties by influencing soil microbial communities (Shi et al. 2018). On one hand, soil acidification resulting from N deposition can negatively affect soil microbial community composition and reduce the overall biomass of soil bacteria and fungi (Treseder 2008; Zhang et al. 2018a, b, c, d). On the other hand, N deposition can stimulate plant growth, leading to increased carbon input into the soil (from litter and roots) and providing abundant substrates for microbial growth and reproduction (Wu et al. 2013), thereby promoting microbial proliferation.
In this study, N addition led to increased soil SOC concentrations, indicating a response in active organic carbon in the soil, which is a crucial energy source for microbial growth. These changes are likely due to the stimulatory effects of N addition on microbial growth and the alteration of carbon use efficiency through the increase in active carbon substrates (Song et al. 2017). This finding aligns with Wang et al. (2021), who reported that N deposition significantly influenced bacterial and fungal communities in N deposition conditions by increasing SOC concentrations. Additionally, previous research on soil microbial characteristics in the Sanjiang Plain wetland has indicated that soil pH, NO3−-N, and NH4+-N were also key factors affecting soil microbial community abundance and composition (Weng et al. 2022; Zhang et al. 2022), which aligns with our results. This suggests that soil pH and available N concentration are important factors influencing soil bacterial and fungal communities. The shift in soil pH and alteration of the soil C:N:P stoichiometric ratio, likely induced by N deposition, negatively affects both above- and below-ground communities, altering soil microbial community structure. The mechanism by which soil acidification changes soil bacterial communities includes ecological selection of the bacterial community in response to acidification and changes in the bacterial community composition within the acidified environment (Chen et al. 2015). However, the soil fungal community is also influenced by SOC, TP, and AP. Compared to bacteria, fungi possess a stronger capacity to absorb N and phosphorus (Wu 2011), and they are closely associated with soil nutrients. Ding et al. (2007) found that soil fungal community structure is closely related to available phosphorus, and Xia et al. (2020) and Ma et al. (2021) also reported significant associations between soil fungal community structure and available phosphorus and total phosphorus. These findings align with our results, indicating that the soil physicochemical parameters influencing the fungal community differ from those influencing bacteria.
5 Conclusion
As expected, N deposition led to changes in the structural composition of soil bacterial and fungal communities at both the phyla and genera levels. Long-term and high levels of N deposition had the effect of reducing fungal diversity (as indicated by Shannon and Chao1 indices) and altering fungal community structure, while bacterial diversity remained unaffected in a temperate wetland ecosystem. N deposition exerted significant and dominant effects on the composition and abundance of the soil microbiome, with fungal communities showing greater sensitivity compared to soil bacteria. Exogenous N input significantly increased soil N (total nitrogen = TN, Nitrate nitrogen = NO3−-N, and ammonium nitrogen = NH4+-N) contents and soil organic carbon (SOC), with nitrogen and carbon being the key environmental factors influencing the structure of the soil microbial community. These findings underscore the crucial role of microbes in regulating soil ecological processes within temperate wetland ecosystems under N addition. However, it is important to note that this study focused solely on the effects of N addition on the belowground microbiome, and therefore further research is needed to gain a more comprehensive understanding of the mechanisms through which soil microbiomes respond to global N deposition in wetland ecosystems.
Data Availability
All raw sequence data were deposited into the NCBI Sequence Read Archive database under accession number PRJNA960933.
References
Arcand MM, Levy-Booth DJ, Helgason BL (2017) Resource Legacies of organic and conventional management differentiate soil microbial carbon use. Front Microbiol 8:2293. https://doi.org/10.3389/fmicb.2017.02293
Basu S, Kumar G, Chhabra S, Prasad R (2021) Role of soil microbes in biogeochemical cycle for enhancing soil fertility. In: New and future developments in microbial biotechnology and bioengineering. Elsevier, Amsterdam, pp 149-157
Bledsoe RB, Goodwillie C, Peralta AL (2020) Long-term nutrient enrichment of an oligotroph-dominated wetland increases bacterial diversity in bulk soils and plant rhizospheres. Msphere 5:e00035. https://doi.org/10.1128/mSphere.00035-20
Cai Y, Liang J, Zhang P, Wang Q, Wu Y, Ding Y, Sun J (2021) Review on strategies of close-to-natural wetland restoration and a brief case plan for a typical wetland in northern China. Chemosphere 285:131534. https://doi.org/10.1016/j.chemosphere.2021.131534
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Chen D, Lan Z, Hu S, Bai Y (2015) Effects of nitrogen enrichment on belowground communities in grassland: Relative role of soil nitrogen availability vs. soil acidification. Soil Biol Biochem 89:99–108. https://doi.org/10.1016/j.soilbio.2015.06.028
Chen ZB, Zhang C, Liu ZH, Song CC, Xin S (2023) Effects of long-term (17 Years) Nitrogen input on soil bacterial community in Sanjiang Plain: the largest marsh wetland in China. Microorganisms 11:1552. https://doi.org/10.3390/microorganisms11061552
Dai HY, Sui X, Fu XY, Zhang RT (2018) Effect of community diversity of Deyeuxia angustifolia and CO2 emission flux under nitrogen deposition conditions in Sanjiang Plain. North Horticult 8:123–128. https://doi.org/10.11937/bfyy.20180259
Deng JJ, Zhang Y, Yin Y, Zhu X, Zhu WX, Zhou YB (2019) Comparison of soil bacterial community and functional characteristics following afforestation in the semi-arid areas. PeerJ 7:e7141. https://doi.org/10.7717/peerj.7141
DeForest JL, Zak DR, Pregitzer KS, Burton AJ (2004) Atmospheric nitrate deposition, microbial community composition, and enzyme activity in northern hardwood forests. Soil Sci Soc Am J 68:132–138. https://doi.org/10.1007/s00248-008-9440-5
De Witte LC, Rosenstock NP, Van Der Linde S, Braun S (2017) Nitrogen deposition changes ectomycorrhizal communities in Swiss beech forests. Sci Total Environ 605:1083–1096. https://doi.org/10.1016/j.scitotenv.2017.06.142
Ding JL, Jiang X, Ma MC, Guan DW, Zhao BS, Wei D, Cao FM, Li L, Li J (2007) Structure of soil fungal communities under long-term inorganic and organic fertilization in black soil of Northeast China. J Plant Nut Fertil 23:914–923. https://doi.org/10.11674/zwyf.16357
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Edwards IP, Zak DR, Kellner H, Eisenlord SD, Pregitzer KS (2011) Simulated atmospheric N deposition alters fungal community composition and suppresses ligninolytic gene expression in a northern hardwood forest. PLoS One 6:e20421. https://doi.org/10.1371/journal.pone.0020421
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364. https://doi.org/10.1890/05-1839
Fowler D, Coyle M, Skiba U, Sutton MA, Cape JN, Reis S, Sheppard LJ, Jenkins A, Grizzetti B, Galloway JN, Vitousek P, Leach A, Bouwman AF, Butterbach-Bahl K, Dentener F, Stevenson D, Amann M, Voss M (2013) The global nitrogen cycle in the twenty-first century. Philosophical Transactions of the Royal Society B-Biological Sciences 368:20130164. https://doi.org/10.1098/rstb.2013.0164
Freedman Z, Eisenlord SD, Zak DR, Xue K, He Z, Zhou J (2013) Towards a molecular understanding of N cycling in northern hardwood forests under future rates of N deposition. Soil Biol Biochem 66:130–138. https://doi.org/10.1016/j.soilbio.2013.07.010
Freitag TE, Chang L, Clegg CD, Prosser JI (2005) Influence of inorganic nitrogen management regime on the diversity of nitrite-oxidizing bacteria in agricultural grassland soils. Appl Environ Microbiol 71:8323–8334. https://doi.org/10.1128/AEM.71.12.8323-8334.2005
Frey B, Carnol M, Dharmarajah A, Brunner I, Schleppi P (2020) Only minor changes in the soil microbiome of a sub-alpine forest after 20 years of moderately increased nitrogen loads. Front for Glob Change 3:77. https://doi.org/10.3389/ffgc.2020.00077
Frey B, Moser B, Tytgat B, Zimmermann S, Alberti J, Biederman LA, Risch AC (2023) Long-term N-addition alters the community structure of functionally important N-cycling soil microorganisms across global grasslands. Soil Biol Biochem 176:108887. https://doi.org/10.1016/j.soilbio.2022.108887
Frey B, Rime T, Phillips M, Stierli B, Hajdas I, Widmer F, Hartmann M (2016) Microbial diversity in European alpine permafrost and active layers. FEMS Microbiol Ecol 92:fiw018. https://doi.org/10.1093/femsec/fiw018
Gruber N, Galloway JN (2008) An Earth-system perspective of the global nitrogen cycle. Nature 451:293–296. https://doi.org/10.1038/nature06592
Guo W, Zhang J, Sui X, Hu X, Lei G, Zhou YM, Qi LH (2022) Compartment niche and bamboo variety influence the diversity, composition, network and potential keystone taxa functions of rhizobacterial communities. Rhizosphere 24:100593. https://doi.org/10.1016/j.rhisph.2022.100593
Hartmann M, Brunner I, Hagedorn F, Bardgett RD, Stierli B, Herzog C, Frey B (2017) A decade of irrigation transforms the soil microbiome of a semi-arid pine forest. Mol Ecol 26:1190–1206. https://doi.org/10.1111/mec.13995
He WY, Zhang MM, Jin GZ, Sui X, Zhang T, Song FQ (2021) Effects of nitrogen deposition on nitrogen-mineralizing enzyme activity and soil microbial community structure in a Korean pine plantation. Microb Ecol 81:410–424. https://doi.org/10.1007/s00248-020-01595-6
Holland EA, Braswell BH, Sulzman J, Lamarque JF (2005) Nitrogen deposition onto the United States and Western Europe: synthesis of observations and models. Ecol Appl 15:38–57. https://doi.org/10.1890/03-5162
Hu Y, Chen M, Yang Z, Cong M, Zhu X, Jia H (2022) Soil microbial community response to nitrogen application on a swamp meadow in the arid region of central Asia. Front Microbiol 12:4126. https://doi.org/10.3389/fmicb.2021.797306
Huang L, Zhang G, Bai J, Xia Z, Wang W, Jia J, Wang X, Liu X, Cui B (2020) Desalinization via freshwater restoration highly improved microbial diversity, cooccurrence patterns and functions in coastal wetland soils. Sci Total Environ 15:765. https://doi.org/10.1016/j.scitotenv.2020.142769
Junk WJ, An S, Finlayson CM, Gopal B, Květ J, Mitchell SA, Robarts RD (2013) Current state of knowledge regarding the world’s wetlands and their future under global climate change:a synthesis. Aquat Sci 75:151–167. https://doi.org/10.1007/s00027-012-0278-z
Lakshmi G, Okafor BN, Visconti D (2020) Soil microarthropods and nutrient cycling. In: Environment, climate, plant and vegetation growth. Springer, Berlin, pp 453-472
LeBauer DS, Treseder KK (2008) Nitrogen limitation of net primary productivity in terrestrial ecosystems is globally distributed. Ecology 89:371–379. https://doi.org/10.1890/06-2057.1
Lee SH, Kim SY, Ding W, Kang H (2015) Impact of elevated CO2 and N addition on bacteria, fungi, and archaea in a marsh ecosystem with various types of plants. Appl Microbiol Biotechnol 99:5295–5305. https://doi.org/10.1007/s00253-015-6385-8
Lu G, Xie B, Cagle GA, Wang X, Han G, Wang X, Guan B (2021) Effects of simulated nitrogen deposition on soil microbial community diversity in coastal wetland of the Yellow River Delta. Sci Total Environ 757:143825. https://doi.org/10.1016/j.scitotenv.2020.143825
Li F, Zang SY, Liu YN, Li L, Ni HW (2020a) Effect of freezing–thawing cycle on soil active organic carbon fractions and enzyme activities in the wetland of Sanjiang Plain, northeast China. Wetlands 40:167–177. https://doi.org/10.1007/s13157-019-01164-9
Li L, Chen J, Han X, Zhang W, Shao C (2020) Natural Conditions Influencing Chinese Grassland Ecosystems. In: Grassland Ecosystems of China. Springer, Berlin, pp 49-79
Liu W, Jiang L, Yang S, Wang Z, Tian R, Peng Z, Liu L (2020) Critical transition of soil bacterial diversity and composition triggered by nitrogen enrichment. Ecology 101:e03053. https://doi.org/10.1002/ecy.3053
Li W, Sheng H, Ekawati D, Jiang Y, Yang H (2019) Variations in the compositions of soil bacterial and fungal communities due to microhabitat effects induced by simulated nitrogen deposition of a bamboo forest in wetland. Forests 10:1098. https://doi.org/10.3390/f10121098
Li Y (2018) Carbon input manipulation affects soil bacterial community composition in a temporal steppe under nitrogen deposition. Geoderma 355:113914. https://doi.org/10.1016/j.geoderma.2019.113914
Li Z, Peng Q, Dong Y, Guo Y (2022) The influence of increased precipitation and nitrogen deposition on the litter decomposition and soil microbial community structure in a semiarid grassland. Sci Total Environ 844:157115. https://doi.org/10.1016/j.scitotenv.2022.157115
Ma X, Song Y, Song C, Wang X, Wang N, Gao S, Du Y (2021) Effect of nitrogen addition on soil microbial functional gene abundance and community diversity in Permafrost Peatland. Microorganisms 9:2498. https://doi.org/10.3390/microorganisms9122498
McHugh TA, Morrissey EM, Mueller RC, Gallegos-Graves LV, Kuske CR, Reed SC (2017) Bacterial, fungal, and plant communities exhibit no biomass or compositional response to two years of simulated nitrogen deposition in a semiarid grassland. Environ Microbiol 19:1600–1611. https://doi.org/10.1111/1462-2920.13678
Nguyen NH, Song Z, Bates ST, Branco S, Tedersoo L, Menke J, Schilling JS, Kennedy PG (2016) FUNGuild:an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol 20:241–248. https://doi.org/10.1016/j.funeco.2015.06.006
Nie Y, Wang M, Zhang W, Ni Z, Hashidoko Y, Shen W (2018) Ammonium nitrogen content is a dominant predictor of bacterial community composition in an acidic forest soil with exogenous nitrogen enrichment. Sci Total Environ 624:407–415. https://doi.org/10.1016/j.scitotenv.2017.12.142
Ning C, Mueller GM, Egerton-Warburton LM, Wilson AW, Yan WD, Xiang WH (2018) Diversity and enzyme activity of ectomycorrhizal fungal communities following nitrogen fertilization in an urban-adjacent pine plantation. Forests 9:99. https://doi.org/10.3390/f9030099
Perez-Mon C, Stierli B, Plötze M, Frey B (2022) Fast and persistent responses of alpine permafrost microbial communities to in situ warming. Sci Total Environ 807:150720. https://doi.org/10.1016/j.scitotenv.2021.150720
Pepe-Ranney C, Campbell AN, Koechli CN, Berthrong S, Buckley DH (2016) Unearthing the ecology of soil microorganisms using a high resolution DNA-SIP approach to explore cellulose and xylose metabolism in soil. Front Microbiol 7:703. https://doi.org/10.3389/fmicb.2016.00703
Ren N, Wang Y, Ye Y, Zhao Y, Huang Y, Fu W, Chu X (2020) Effects of continuous nitrogen fertilizer application on the diversity and composition of rhizosphere soil bacteria. Front Microbiol 11:1948. https://doi.org/10.3389/fmicb.2020.01948
Ren Y, Yu G, Shi C, Liu L, Guo Q, Han C, Huang H (2022) Majorbio Cloud: A one-stop, comprehensive bioinformatic platform for multiomics analyses. Imeta 1:e12. https://doi.org/10.1002/imt2.12
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH:a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Roth T, Kohli L, Rihm B, Achermann B (2013) Nitrogen deposition is negatively related to species richness and species composition of vascular plants and bryophytes in Swiss mountain grassland. Agr Ecosyst Environ 178:121–126. https://doi.org/10.1016/j.agee.2013.07.002
Salimi S, Almuktar SA, Scholz M (2021) Impact of climate change on wetland ecosystems: A critical review of experimental wetlands. J Environ Manage 286:112160. https://doi.org/10.1016/j.jenvman.2021.112160
Shi XZ, Hu HW, Wang JQ, He JZ, Zheng CY, Wan XH, Huang ZQ (2018) Niche separation of comammox Nitrospira and canonical ammonia oxidizers in an acidic subtropical forest soil under long-term nitrogen deposition. Soil Biol Biochem 126:114–122. https://doi.org/10.1016/j.soilbio.2018.09.004
Shi Y, Li Y, Yang T, Chu HY (2021) Threshold effects of soil pH on microbial co-occurrence structure in acidic and alkaline arable lands. Sci Total Environ 800:149592. https://doi.org/10.1016/j.scitotenv.2021.149592
Song YY, Song CC, Meng HN, Swarzenski CM, Wang XM, Tan WW (2017) Nitrogen additions affect litter quality and soil biochemical properties in a peatland of Northeast China. Ecol Eng 100:175–185. https://doi.org/10.1016/j.ecoleng.2016.12.025
Song B, Li Y, Yang L, Shi H, Li L, Bai W, Zhao Y (2022) Soil Acidification Under Long-Term N Addition Decreases the Diversity of Soil Bacteria and Fungi and Changes Their Community Composition in a Semiarid Grassland. Microb Ecol 85:1–11. https://doi.org/10.1007/s00248-021-01954-x
Song H, Jiang C, Yu W (2014) Basic features and monitoring methodologies of atmospheric nitrogen deposition. Chin J Appl Ecol 25:599–610
Sui X, Zhang RT, Frey B, Yang LB, Liu MH, Ni HW (2019) Land use change effects on diversity of soil bacterial, Acidobacterial and fungal communities in wetlands of the Sanjiang Plain, northeastern China. Sci Rep 9:1–14. https://doi.org/10.1038/s41598-019-55063-4
Sui X, Zhang RT, Frey B, Yang LB, Liu YN, Ni HW, Li MH (2021) Soil physicochemical properties drive the variation in soil microbial communities along a forest successional series in a degraded wetland in northeastern China. Ecol Evol 11:2194–2208. https://doi.org/10.1002/ece3.7184
Sui X, Frey B, Yang LB, Liu YN, Zhang RT, Ni HW, Li MH (2022) Soil Acidobacterial community composition changes sensitively with wetland degradation in northeastern of China. Front Microbiol 13:1052161. https://doi.org/10.3389/fmicb.2022.1052161
Sun M, Li M, Zhou Y, Liu J, Shi W, Wu X, Gao Z (2022) Nitrogen deposition enhances the deterministic process of the prokaryotic community and increases the complexity of the microbial co-network in coastal wetlands. Sci Total Environ 15:158939. https://doi.org/10.1016/j.scitotenv.2022.158939
Song LH, Liu J, Yan XM, Chang L, Wu DH (2016) Euedaphic and hemiedaphic Collembola suffer larger damages than epedaphic species to nitrogen input. Environ Pollut 208:413–415. https://doi.org/10.1016/j.envpol.2015.10.008
Treseder KK (2008) Nitrogen additions and microbial biomass: A meta-analysis of ecosystem studies. Ecol Lett 11:1111–1120. https://doi.org/10.1111/j.1461-0248.2008.01230.x
Verastegui Y, Cheng J, Engel K, Kolczynski D, Mortimer S, Lavigne J, Neufeld JD (2014) Multisubstrate isotope labeling and metagenomic analysis of active soil bacterial communities. Mbio 5:e01157-e1214. https://doi.org/10.1128/mBio.01157-14
Wang C, Liu D, Bai E (2018) Decreasing soil microbial diversity is associated with decreasing microbial biomass under nitrogen addition. Soil Biol Biochem 120:126–133. https://doi.org/10.1016/j.soilbio.2018.02.003
Wang JQ, Shi XZ, Zheng CY, Suter H, Huang ZQ (2021) Different responses of soil bacterial and fungal communities to nitrogen deposition in a subtropical forest. Sci Total Environ 755:142449. https://doi.org/10.1016/j.scitotenv.2020.142449
Wang N, Gao J, Wei J, Liu Y, Zhuang XL, Zhuang GQ (2019) Effects of wetland reclamation on soil microbial community structure in the Sanjiang Plain. Environ Sci 40:2375–2381. https://doi.org/10.13227/j.hjkx.201809204
Weng XH, Sui X, Liu YN, Yang LB, Zhang RT (2022) Effect of nitrogen addition on the carbon metabolism of soil microorganisms in a Calamagrostis angustifolia wetland of the Sanjiang Plain, northeastern China. Ann Microbiol 72:1–14. https://doi.org/10.1186/s13213-022-01674-8
Wu J, Liu W, Fan H, Huang G, Wan S, Yuan Y, Ji C (2013) Asynchronous responses of soil microbial community and understory plant community to simulated nitrogen deposition in a subtropical forest. Ecol Evol 3:3895–3905. https://doi.org/10.1002/ece3.750
Wu T (2011) Can ectomycorrhizal fungi circumvent the nitrogen mineralization for plant nutrition in temperate forest ecosystems? Soil Biol Biochem 43:1109–1117. https://doi.org/10.1016/j.soilbio.2011.02.003
Xia Q, Rufty T, Shi W (2020) Soil microbial diversity and composition: Links to soil texture and associated properties. Soil Biol Biochem 149:107953
Yan GY, Xing YJ, Xu LJ, Wang JY, Dong XD, Shan WJ, Guo L, Wang QG (2017) Effects of different nitrogen additions on soil microbial communities in different seasons in a boreal forest. Ecosphere 8:e01879. https://doi.org/10.1002/ecs2.1879
Yang S, Xu ZW, Wang RZ, Zhang YY, Yao F, Zhang YG, Ronald FT, Jiang Y, Zou HT, Li H (2017) Variations in soil microbial community composition and enzymatic activities in response to increased N deposition and precipitation in Inner Mongolian grassland. Appl Soil Ecol 119:275–285. https://doi.org/10.1016/j.apsoil.2017.06.041
Yang Y, Cheng H, Gao H, An S (2020) Response and driving factors of soil microbial diversity related to global nitrogen addition. Land Degrad Dev 31:190–204. https://doi.org/10.1002/ldr.3439
Zeng J, Liu XJ, Song L, Lin XG, Zhang HY, Shen CC, Chu HY (2016) Nitrogen fertilization directly affects soil bacterial diversity and indirectly affects bacterial community composition. Soil Biol Biochem 92:41–49. https://doi.org/10.1016/j.soilbio.2015.09.018
Zhang G, Bai J, Zhao Q, Jia J, Wang W, Wang X (2020a) Bacterial succession in salt marsh soils along a short-term invasion chronosequence of Spartina alterniflora in the Yellow River Estuary, China. Microb Ecol 79:644–661. https://doi.org/10.1007/s00248-019-01430-7
Zhang HF, Liu HM, Zhao JN, Li G, Lai X, Li J (2018a) Effects of simulated nitrogen deposition and precipitation change on soil bacterial community structure in a Stipa baicalensis steppe. Acta Ecol Sin 38:244–253
Zhang J, Jin K, Luo Y, Du L, Tian R, Wang S, Xu Z (2023) Responses of Soil Enzyme Activity to Long-Term Nitrogen Enrichment and Water Addition in a Typical Steppe. Agronomy 13:1920. https://doi.org/10.3390/agronomy13071920
Zhang J, Kobert K, Flouri T, Stamatakis A (2014a) PEAR:a fast and accurate Illumina Paired-End mergeR. Bioinformatics 30:614–620. https://doi.org/10.1093/bioinformatics/btt593
Zhang RT, Liu YN, Zhong HX, Chen XW, Sui X (2022) Effects of simulated nitrogen deposition on the soil microbial community diversity of a Deyeuxia angustifolia wetland in the Sanjiang Plain, Northeastern China. Ann Microbiol 72:1–13. https://doi.org/10.1186/s13213-022-01666-8
Zhang RT, Sui X, Xu N, Zhong HX, Fu XY, Ni HW (2018) Responses of greenhouse gas emission to simulated nitrogen deposition in Calamagrostis angustifolia wetlands of Sanjiang Plain, China. J Appl Ecol 29:3191–3198. https://doi.org/10.13287/j.1001-9332.201810.001
Zhang T, Xu F, Huai BD, Yang X, Sui WZ (2020) Effects of land use changes on soil bacterial community diversity in the riparian wetland along the downstream of Songhua River. Environ Sci 41:4273–4283. https://doi.org/10.13227/j.hjkx.202003088
Zhang TA, Chen HY, Ruan H (2018c) Global negative effects of nitrogen deposition on soil microbes. ISME J 12:1817–1825. https://doi.org/10.1038/s41396-018-0096-y
Zhang ZQ, Yao Q, Xu QH, Jiang M, Zhu TC (2021) Hydrological and palynological evidence of wetland evolution on the Sanjiang Plain (NE China) in response to the Holocene East Asia summer monsoon. CATENA 203:105332. https://doi.org/10.1016/j.catena.2021.105332
Zhang X, Wei H, Chen Q, Han X (2014b) The counteractive effects of nitrogen addition and watering on soil bacterial communities in a steppe ecosystem. Soil Biol Biochem 72:26–34. https://doi.org/10.1016/j.soilbio.2014.01.034
Zheng Y, He JZ (2020) Responses of forest soil microbial communities to drought and nitrogen deposition:A review. J Appl Ecol 31:2464–2472. https://doi.org/10.13287/j.1001-9332.202007.035
Zhang XL, Zhai PH, Huang JH, Zhao X, Dong KH (2018d) Responses of ecosystem water use efficiency to spring snow and summer water addition with or without nitrogen addition in a temperate steppe. PLoS ONE 13:e0194198. https://doi.org/10.1371/journal.pone.0194198
Zhu X, Zhang W, Chen H, Mo J (2015) Impacts of nitrogen deposition on soil nitrogen cycle in forest ecosystems: A review. Acta Ecol Sin 35:35–43. https://doi.org/10.1016/j.chnaes.2015.04.004
Acknowledgements
This work was funded by the National Key Research and Development Program of China (2022YFF130090503); the Natural Sciences Foundation of Heilongjiang Province (LH2020C088); the Heilongjiang Province Postdoctoral Research Start-up Fund Project (LBH-Q21167); the Outstanding Youth Foundation of Heilongjiang University (JCL202006); Basic Scientific Research project of Higher Education Institutions of Heilongjiang Province and Heilongjiang Provincial Ecological Environmental Protection Research Project (HST2022ST008), as well as sponsored in part by Program of China Hebei education department (grant No.ZC2023041).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of Interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xin, S., Wang, M., Frey, B. et al. Differential Responses of Soil Bacterial and Fungal Communities to Simulated Nitrogen Deposition in a Temperate Wetland of Northeastern China. J Soil Sci Plant Nutr 24, 467–482 (2024). https://doi.org/10.1007/s42729-023-01557-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42729-023-01557-0