Abstract
Brown planthopper (BPH) is a highly destructive insect pest of rice, causing significant yield loss. Due to its constantly evolving nature, continuous analysis of BPH's protein domain-interacting partners is essential. In the present study, in silico approach was followed to predict 3-D structure of cloned BPH resistant proteins (Bph6, Bph9, Bph14, Bph17, Bph18, Bph26, Bph29 and Bph32) using a comparative modelling approach and their interaction studies. The interactome analysis revealed a key regulator, OsJ_28113, responsible for transducing extracellular signals into intracellular responses, potentially aiding in activating proteins that provide resistance against BPH. The proposed model provides insights into the structure and active sites of these proteins, offering opportunities to develop novel strategies for BPH control in rice plants. The molecular profile analysis revealed that BPH resistance genes containing the CC-NBS-LRR domain have varying lengths of amino acid chains ranging from 1082 for Bph30 to the longest (2024) for Bph6. Bph26 and Bph18 demonstrated high sequence similarity containing NB-ARC and LRR domains. The secondary structure prediction results anticipated that all the proteins, except Bph30, are cytoplasmic and soluble. The in silico findings support the notion that variability in resistance genes is a result of ongoing evolutionary interactions between plants and insect pests. Additionally, the study uncovered higher ligand binding affinities towards jasmonic acid compared to salicylic acid, paving the way for further research on receptor-ligand recognition and signalling mechanisms against rice planthoppers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Being an important staple cereal of Asian countries, rice (Oryza sativa L.) contributes a major portion to human caloric intake and nutrition. Rice productivity is continuously challenged by various abiotic and biotic stresses which account for around 50% of global yield loss (Ishaq and Memon 2017). Among biotic stresses, brown planthopper (BPH, Nilaparvata lugens Stål) is a major devastating insect-pest of rice in South and Southeast Asia. It damages the rice crop by feeding phloem sap using its stylet-type mouthparts (Normile 2008). In recent years, the BPH population has increased significantly due to the rapid adaptation of BPH-sensitive varieties which is further exacerbated by high humidity, optimum temperature, and excessive use of nitrogenous fertilisers beyond the approved doses (Sogawa 2015). Other factors contributing to BPH infestation are higher plant density and indiscriminate application of pesticides during the early development of the host (Rashid et al. 2017; Wang et al. 2008). Both nymphs and adults suck on cell saps of rice leaves, leading to dehydration of leaves, reduction in photosynthetic rate, leaf area, chlorophyll content, nitrogen level of leaf and stem and eventually death of plants resulting in ‘hopper burn’ (Cagampang et al. 1974). Other than the direct harm brought by BPH, it also causes indirect damage by transmitting viruses including rice grassy stunt virus (RGSV) and rice ragged stunt virus (RRSV) (Sogawa 1982; Cha et al. 2008; Cabauatan et al. 2009). There are four different biotypes in which BPH populations have been characterised (Khush et al. 1985). Of these, BPH biotype 4 is the most devastating and predominant in India. While different management strategies to control insect -pest damage, use of a host-plant resistance system is the most durable and environmentally safe approach for managing BPH (Brar et al. 2009; Kumar et al. 2020).
To date, 46 genes/QTLs have been designated from various cultivated and wild relatives of rice and assigned to the different chromosomes of rice. Of these, nine genes namely Bph6, Bph9, Bph14, Bph17, Bph18, Bph26, Bph29, Bph30, and Bph32 (Guo et al. 2018; Zhao et al. 2016; Du et al. 2009; Liu et al. 2015; Ji et al. 2016; Tamura et al. 2014; Wang et al. 2015; Shi et al. 2021; Ren et al. 2016) have been cloned and characterised (Table 1). Among them, Bph9, Bph14, Bph18, Bph26, and Bph30 encode a coiled-coil, nucleotide-binding site and leucine-rich repeat (CC-NBS-LRR) domain proteins. While Bph17, Bph29, and Bph32 encode lectin receptor kinases (LRKs), B3 DNA-binding domain protein, and a unique short consensus repeat (SCR) domain protein, respectively. The CC-NBS-LRR proteins play central roles in perceiving the elicitors/effectors molecules and mounting the appropriate resistance response when infested by insects (Jacob et al. 2013). Wu et al. (2022) proposed that the Bph6 protein enhances rice defense against BPH by regulating the accumulation of cell wall lignin. In another study conducted by Zheng et al. (2021), it was observed that the nymphs and adults of BPH feeding on NIL-Bph6 rice plants exhibited reduced weight gain and growth, indicating suppressed feeding by the BPH. The structure and functional analysis of Bph14 revealed the induction of strong BPH resistance response by activating the salicylic acid (SA) pathway followed by the accumulation of numerous transcription factors (TFs) such as WRKY46 and WRKY72 leading to sieve tube blockage, which reduces the insect’s feeding, growth, and survival (Hu et al. 2017). However, the molecular structure and functions of other BPH resistance genes are poorly known. Therefore, a precise understanding of the structure and function of BPH resistance genes is required to gain deeper insight into the molecular mechanism underlying BPH resistance in rice.
The increasing disparity in connecting DNA sequences with protein structures presents significant challenges in understanding the function of proteins of interest. The current study emphasises on utilising comparative modelling for in silico prediction of 3-D structure of BPH resistant (R)/susceptible (S) proteins. Additionally, an interaction study is conducted to unravel the structural interactions between BPH-specific CC-NBS-LRR genes and other functional genes within the cell. Molecular profiling analysis and secondary structure prediction serve as crucial groundwork for exploring the evolutionary biology of plant- (R) genes. The obtained results contribute to a greater comprehension of the interplay between R protein-elicitor perception and plant defense signalling in response to rice planthoppers. Overall, these findings contribute to a better understanding of BPH resistance mechanisms and offer potential targets for developing effective strategies to combat this destructive rice pest.
Materials and methods
Sequence retrieval
The nucleotide and amino acid sequences of the successfully cloned BPH genes (namely Bph6, Bph9, Bph14, Bph17, Bph18, Bph26, Bph29, Bph30, and Bph32) were downloaded from National Center for Biotechnology Information (NCBI) database (https://www.ncbi.nlm.nih.gov/) and Rice Genome Annotation Project (RGAP) database (http://rice.uga.edu/). The complete coding sequence and protein sequence were retrieved using the gene ids of the cloned BPH genes as given in the reported papers (Supplementary Table 1). The query nucleotide sequences were BLAST (Basic Local Alignment Search Tool) searched to identify the sequences in the other wild and cultivated species genomes submitted in RGAP website (http://rice.uga.edu/analyses_search_blast.shtml) and Ensembl Plants website (http://plants.ensembl.org/index.html). The BLAST search on RGAP and Ensembl revealed very low query coverage among various genomes of wild and cultivated rice. The homology alignment of query sequence was not totally aligned with the subject using default parameters (e-value threshold as1e-5). Similarly, BAC (Bacterial Artificial Chromosome) libraries of the rice genome were used to annotate the target genes. The genomic sequence retrieved from BAC was processed to determine the coding and non-coding regions. The protein sequences were retrieved in FASTA (fast-all) format and processed in Ensembl Plants for identifying target genes with chromosome location. The features of the gene were studied using various options available in the genome browser at the RGAP site. The sequence of methods followed for the present study has been represented in Supplementary Fig. 1.
Physicochemical characterisation of BPH R proteins
Various physical and chemical properties such as molecular weight, theoretical pI (isoelectric point), EI (extinction coefficient), AI (aliphatic index), II (instability index), + R and −R (total number of positive and negative residues) and GRAVY (grand average hydropathy) of cloned BPH R proteins were calculated using ExPASy’s ProtParam web server tool (http://web.expasy.org/protparam/) (Gasteiger et al. 2005).
Phylogenetic analysis and motif identification
The Unrooted phylogenetic tree was constructed using the maximum likelihood method to analyse the evolutionary relationship among the cloned BPH R proteins of rice… All the protein sequences were imported in the MEGA (Molecular Evolutionary Genetic Analysis) software, version 10 (Kumar et al. 2018) (https://www.megasoftware.net/) and the reliability was checked with bootstrap replication value as 1000 and other default parameters remained same. The sequences were then aligned using the MUSCLE algorithm. The conserved motifs were identified using MEME (Multiple Em for Motif Elicitation) software, version 5.1.1 (Bailey et al. 2006).
Domain annotation
Domains from amino acid sequences of cloned BPH genes were extracted through a multi-source domain annotation server, MyCLADE. This server was used to annotate the query dataset with an available set of Pfam domains (http://www.lcqb.upmc.fr/myclade/index.php).
Model preparation
Swiss model
SWISS-MODEL is accessible via a web interface at http://swissmodel.expasy.org, or directly as a link from SWISS-PROT (Boeckmann et al. 2003) entries on the ExPASy server (Appel et al. 1994). The target sequence and template structure were identified and aligned for model building. A local pair-wise alignment of the target BPH sequence to the main template structures was calculated. The integrity of the models was analysed by C-score, giving an estimate of the variability of the template structures at this position in the result files. The parts of the model with no template information were assigned a C-score of 99.
Phyre2
To predict and analyse the protein structure and function, Phyre2 tool was used. Phyre2 uses advanced remote homology detection methods to build 3D models, predict ligand binding sites and analyse the effect of amino-acid variants for the query protein sequences. With Phyre2, function and mutations in the proteins were also predicted. After submitting the query protein sequence, 2° and 3° structures of the models, their domain composition and model quality were interpreted. The 3D structure of the successfully cloned BPH R genes was predicted by submitting the protein sequence of each gene individually (Fig. 1). The analysis included sequence analysis, 2° and disorder prediction, domain analysis and detailed template information. The information to the right of the image showed the structural template on which the top models were based, the confidence and coverage of the model with a link to interact with the 3D model using JSmol within the browser.
Prediction of protein interactions
For an overall understanding of cellular function, knowledge of all functional interactions between the expressed proteins is required, for which STRING (Search Tool for the Retrieval of Interacting Genes/Proteins) database was used (https://string-db.org/). The associations in STRING are included only if direct interactions (physical), as well as indirect interactions (functional) are specific and biologically meaningful (Szklarczyk et al. 2017). STRING showed a linkage of the query proteins with other functional proteins in the cell by collecting and reassessing available experimental data on protein–protein interactions, and importing known pathways and protein complexes from curated databases. For each protein–protein association stored in STRING, a confidence score scaled between zero and one was provided. The confidence score indicated the estimated likelihood that a given interaction is biologically meaningful, specific and reproducible, given the supporting evidence. Hence, the interacting units in STRING are the actual protein-coding gene loci (represented by their main, canonical protein isoform).
Preparation of ligands and protein docking
For docking, the 3D protein structure of BPH R genes and phytochemicals/ligands were downloaded from the NCBI and PubChem Database (https://pubchem.ncbi.nlm.nih.gov/) in.sdf format. The sdf files were converted to.pdb format using Open Babel (O’Boyle et al. 2011). The energy of ligands as well as receptors for docking were minimised using Chimera 1.16 version tool. The ligands selected were jasmonic acid and salicylic acid with previous docking history (Gupta et al. 2019). All of the cloned BPH R proteins were docked individually with both ligands. AutoDock4 version (v4.2.6) (https://autodock.scripps.edu/) was used for docking with the work assisting tool python 3.10.0 (https://www.python.org/) to bind protein and ligand. The protein molecules were processed by adding hydrogen ions, merging non-polar hydrogen atoms, defining AD4 atom types, etc. (Fig. 2). Lamarckian Genetic Algorithm 4.2 was implemented and the receptor was kept rigid all throughout the study. The genetic algorithm was set for 100 runs with other parameters at default (250,000 energy evaluations). The docked models were analysed in UCSF Chimera 1.16 version tool (Pettersen et al. 2004) and were selected based on the appropriate interactive site and docking score. The docked files in pdb format were uploaded in PDBsum to know interacting residues (https://www.ebi.ac.uk/thornton-srv/databases/pdbsum/) of phytochemicals [jasmonic acid and salicylic acid] with cloned BPH R proteins.
Results
Prediction of 3-D structure of BPH R/S proteins
Based on analysis of the targeted proteins, the domain composition of the BPH R and S proteins have indels leading to their different reaction towards BPH infestation. Bph6 has deletion at 821 position in c4ecnA domain in LRR protein, Bph9 and Bph14 have deletions at 451 and 237 position respectively in c7jlvG domain in disease resistance protein roq1, Bph17a, Bph17c and Bph17d have deletions at 655, 623 and 646 positions respectively in c6xr4B domain in LRR serine/threonine-protein kinase 2. The comparison of Bph18 R and S proteins using SWISS-MODEL revealed truncation of protein leading to susceptibility against BPH in rice. Using UCSF Chimera, Bph18 R domains (232–344 and 411–650) and Bph18 S domains (233 to 329) were selected (Fig. 3a and b) and superimposed to visualise the truncation of Bph18 protein in susceptible cultivar (Fig. 3c)where a deletion at 624 position in disease resistance rpp13-like protein4 was found to be associated with the susceptibility of Bph18 (Fig. 3d). Bph26 has deletion at 483 position in c4ecnA domain in LRR protein. These insertion-deletions and their respective positions help to target specific stretches of the gene or editing the DNA at particular locations. The analysis revealed that the majority of the proteins conferring resistance to BPH contained domains such as those found in disease resistance proteins like rpp13-like protein4, roq1, and LRR serine/threonine-protein kinase2. However, in two cases (bph29 and Bph32) no resistance domain was found, still they are providing resistance against BPH (Wang et al. 2015; Zhao et al. 2016). bph29 and Bph32 may act as an enhancer or suppressor for other proteins or may give resistance due to phosphorylation or methylation as they are not directly related to the disease resistance NB-LRR family. The proteins were further explored with the help of STRING software to analyse the Interactome of the BPH R genes.
a 3D structure of Bph18 R domain; b 3D structure of Bph18 S domain; c superimposed 3D model of Bph18 R (represented in grey colour) and Bph18 S (represented in blue colour) to visualise the truncation of Bph18 protein in susceptible gene model., d sequence alignment of Bph18 R and S depicting deletion of amino acid; Proline at 624th position in protein sequence of Bph18 S
Physico-chemical properties
The molecular profile analysis revealed that CC-NBS-LRR domains containing BPH resistance genes have varying lengths of amino acid chains ranging from smallest (1082) for Bph30 to longest (2024) for Bph6. The pI value of cloned BPH resistance genes ranged from 5.83–8.74, where Bph6, Bph14, Bph30 with pI < 7 were found to be acidic and Bph9, Bph18, Bph26 with pI > 7 indicated their basic nature. II of all proteins was above 40 suggesting that these are unstable proteins. The GRAVY index was also measured between − 0.24 and − 0.28 advocated that all proteins are hydrophilic in nature (Table 2). Since the GRAVY index values are negative, the proteins under consideration is considered as hydrophilic.
Phylogenetic and motif analysis
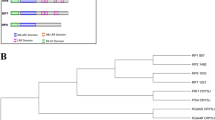
Neighbour-joining tree based on the protein sequence of cloned BPH resistance genes revealed two major groups (Supplementary Fig. 2). The Bph30 (Biotype 1, 2 and 3) and Bph6 (Biotype 1, 2 and 3) are clustered in group I. The most plausible reason is that they are rare atypical BPH resistance genes containing only LRR domains. The Bph26 (Biotype 1 and 2), Bph18 (Korean), Bph9 (Biotype 1, 2 and 3), Bph32 (mixed biotypes from China), and Bph14 (mixed biotypes from China) are clustered in group II. The Bph26 and Bph18 showed high sequence similarity and both contain NB-ARC and LRR domains. The Bph6 and Bph30 contain two LRR domains. Whereas the remaining proteins have NB-ARC and LRR domains. The Bph14 comprises a single NB-ARC domain while Bph9, Bph18, and Bph26 comprise two NB-ARC domains. In addition to that the Bph9, Bph18, Bph26 and Bph32 contain other domains including AAA type ATPase domain. (Supplementary Table 2).
Domain annotation of BPH R genes
Domain annotation of BPH R genes revealed that Bph6 has LRR domain, Bph9 has Rx N-terminal, RTC insert (RNA 3’ terminal phosphate cyclase) domain, NB-ARC (ATPase) domain, and LRR domain, Bph14 has Rx N-terminal, NB-ARC (ATPase) domain and LRR domain and FNIP repeat, Bph17 has β lectin (D-mannose binding lectin), S locus glycoprotein, PAN (PAN-like domain) and Pkinase (Protein kinase domain), Bph18 has Rx N-terminal, NB-ARC (ATPase) domain, LRR domain and FNIP repeat, Bph26 has Rx N-terminal, NB-ARC (ATPase) domain and LRR domain, Bph29 has B3 DNA binding, Bph30 has AAA ATPase domain, LRR domain and FNIP repeat, Bph32 has LEA (Late embryogenesis abundant protein).
Secondary structure prediction
The secondary structure of all the NBS-LRR domain BPH resistance proteins was predicted using SOPMA web servers. The result anticipated that all the proteins were cytoplasmic and soluble except Bph30. The Bph30 is an endomembrane localised protein as it is localised in the endoplasmic reticulum, tonoplast, exocyst but not in the nucleus, Golgi apparatus, peroxisome or plastid. Based on the amino acid sequence, MINNOU server predicted this protein as endomembrane localized protein. The secondary structure prediction results revealed that there were high percentage of alpha helix, extended strands, and random coil accompanied by a small fraction of beta-turn in these proteins. The secondary structure composition of all the proteins is given in Supplementary Table 3. The Bph6 contains alpha helix (34.66%), extended strand (16.41%), beta-turn (2.96%) and random coils (45.97%). The Bph9 comprises a relatively higher alpha helix (52.16%), extended strand (12.35%), beta-turn (4.06%) and random coils (31.43%). Similarly, Bph14 harbours alpha helix (56.99%), extended strand (7.71%), beta-turn (2.65%) and random coils (32.65%). The Bph18 includes alpha helix (52.45%), extended strand (12.81%), beta-turn (3.67%) and random coils (31.08%). The Bph26 encompasses alpha helix (52.71%), extended strand (12.97%), beta-turn (3.28%) and random coils (31.03%). However, the secondary structure composition of the Bph30 was alpha helix (35.77%), extended strand (16.27%), beta-turn (3.60%), and random coils (44.36%). The analysis reveals that the LRR domains of Bph6, Bph9, Bph14, Bph18, Bph26 and Bph30 are mainly composed of Alpha helix conformation and hydrogen-bonded turn (Supplementary Fig. 3).
Interactome analysis of BPH R and S proteins
For each BPH R and S protein, the protein–protein interaction model was made in STRING (Fig. 4). The BPH R and S proteins along with their annotation and identity were explained forming clusters with the cellular functional proteins. In Bph17, G-type lectin s-receptor-like serine/threonine-protein kinase LecRK was found whereas disease related protein1 was found in Bph9, Bph18 and Bph26. The interactome of the R and S genes showed that the BPH R genes interact individually whereas BPH S genes interact with other functional proteins like electron carrier proteins. The Os03g0848700 protein gives resistance against Bph14 in rice and Powdery mildew resistance protein PM3b, putative, expressed in wheat (Table 3). It was observed that protein OsJ_28113 belonging to disease resistance NB-LRR family is encoded by Bph9, Bph18 and Bph26. This protein has one Rx_N domain and two NB_ARC domains. The CC–NB–LRR protein of the NB–LRR family, is an immune receptor type similar to R proteins functioning in disease resistance. OsJ_28113 is encoded by three BPH resistance genes: Bph9, Bph18 and Bph26 derived from different plants, but reveal a similarity in the molecular mechanism for insect resistance. From the interactome, a model can be hypothesised that this protein is the master regulator having g-coupled receptors which transduce the extracellular signals into intracellular responses and activates the protein (Fig. 5). For disease management, we can aim three BPH R genes (Bph9, Bph18 and Bph26) by targeting this particular protein (Fig. 6) to control vulnerability against BPH in rice. No interactions were found for Bph32 with other proteins.
Molecular docking with phytochemicals
To find the best binding mode, Auto Dock was utilised for binding free-energy evaluation. Energy items calculated by Auto Dock comprise intermolecular energy, internal energy, torsional energy and unbound energy. Internal energy is composed of Vander Waals energy, hydrogen bonding energy, desolvation energy and electrostatic energy. The binding energy of Bph6 with jasmonic acid and salicylic acid is 6.26 kcal/mol and − 4.8 kcal/mol respectively. Bph9 has a binding energy of − 7.23 kcal/mol and − 6.49 kcal/mol with jasmonic acid and salicylic acid respectively. Bph14 has a binding energy of − 4.17 kcal/mol and − 4.25 kcal/mol with jasmonic acid and salicylic acid respectively. Bph17a, 17c, 17d have a binding energy of − 5.45 kcal/mol and − 5.06 kcal/mol with jasmonic acid and salicylic acid respectively. Bph18 has a binding energy of − 7.25 kcal/mol and − 6.65 kcal/mol with jasmonic acid and salicylic acid respectively. Bph26 has a binding energy of − 6.9 kcal/mol and − 5.62 kcal/mol with jasmonic acid and salicylic acid respectively. bph29 has a binding energy of − 5.04 kcal/mol with jasmonic acid and − 5.34 kcal/mol with salicylic acid. Bph30 has a binding energy of − 4.72 kcal/mol and − 4.64 kcal/mol with jasmonic acid and salicylic acid, respectively. Bph32 has binding energy of − 5.4 kcal/mol and − 4.75 kcal/mol with jasmonic acid and salicylic acid, respectively (Supplementary Fig. 4). Out of 9 docked complexes, the Auto dock binding energy of LRR domain of 7 complexes was highest with jasmonic acid indicating higher binding affinity of LRR domains towards jasmonic acid compared to salicylic acid (Table 4).
Discussion
Though great advancement has been made in predicting the experimental structure by X-ray crystallography and NMR, there remains a substantial disparity between the number of known proteins and their well-defined structures Therefore, the deployment of computational methods for protein structure prediction is urgently needed to bridge this ‘structure knowledge gap’. In O. sativa, nine BPH resistant genes have been isolated using a map-based cloning approach. Among them, Bph6, Bph9, Bph14, Bph18, Bph26 and Bph30 encode CC-NBS-LRR domain proteins. The other three genes encode lectin receptor kinase, B3 DNA-binding domain, and a SCR domain protein suggesting a unique mechanism of resistance to BPH. Three of the CC-NBS-LRR protein encoding genes Bph9, Bph18, and Bph26 are clustered on chromosome 12, while Bph14 is located on chromosome 3. The CC-NBS-LRR regions are evolutionarily similar and largely found in clusters on plant genomes due to segmental and tandem duplication which generate closely related NBS genes (Meyers et al. 2003; Mchale et al. 2006; Leister 2004). Based on sequence analysis, the co-localization of two BPH resistance genes, Bph18 (Ji et al. 2016) and Bph26 (Tamura et al. 2014) within the genomic region of Bph9, suggested that they are functional allelic forms of Bph9 (Zhao et al. 2016). A number of CC-NBS-LRR genes, despite their sequence similarity, showed amino acid substitutions or deletions in their NBS and LRR regions as documented by Ji et al. (2016). It has been suggested that the LRR domain of plant R proteins plays a crucial role in recognition of specific pathogen effectors and is involved in interaction with specific ligands which, in turn, elicit appropriate defense responses (DeYoung and Innes 2006). This defense strategy of plants, to combat pathogens by creating gene clusters has led to the proliferation of R proteins under divergent selection during the co-evolution of plants and pathogens (Qian et al. 2017; Takken et al. 2006). Out of nine genes, six genes encode NBS-LRR domains. The role of CC-NB-LRR domain proteins is extensively studied in plant immunity. Recently, the R proteins have been reported in insect resistance against wheat hessian fly, etc. Therefore, our study primarily focuses on CC-NBS-LRR domain encoding genes. In this study, superimposition of the secondary structure of CC-NBS-LRR protein from resistant alleles with its susceptible counterpart showed InDels suggesting variations in resistance specificities of CC-NBS-LRR proteins to insect pests over time. Functional characterisation of different domains of CC-NBS-LRR protein has been performed for Bph14 and Bph9 (Hu et al. 2017; Wang et al. 2021). The molecular and functional analysis of CC-NBS1-NBS2-LRR domain of Bph9 showed truncation of the LRR domain in the susceptible allele Bph99311. The lack of LRR domain of susceptible alleles revealed loss of BPH resistance activity (Wang et al. 2021). Comparable outcomes were noted for Bph18, where the truncation of amino acids resulted in susceptibility.
A fully detailed view of all functionally relevant protein interactions of BPH-R and S proteins is essential to locate the molecular functions of individual proteins into their cellular context. The indirect associations such as genetic interactions or shared pathway memberships are equally important as physical interaction for a complete understanding of cellular function, allowing an effective design of experiments, such as site-directed mutagenesis, or the structure-based design of specific inhibitors. The interactome of the R and S genes indicated that in plants, the pathways' functioning relies on a series of reactions, with feedback inhibition playing a significant role. No discernible resistance domain was identified in the case of bph29 and Bph32 as bph29 contains a highly conserved B3 DNA-binding domain (Punta et al. 2012), found exclusively in TFs that interacts with the major groove of DNA (Yamasaki et al. 2004). Bph32 encodes an unknown protein that belongs to the complement control module/SCR domain family, a family of cell adhesion molecules (CAMs) that are considered to be types of lectin, or cell adhesion proteins (Parham 2005). The plant lectins have previously been reported to function as direct defence proteins to inhibit insect feeding (Michiels et al. 2010; Vandenborre et al. 2011) Therefore, it was hypothesised that these genes can be targeted individually by using their interactome and their interaction with other cellular proteins to provide resistance against BPH. Interactome analysis unveiled an additional finding that the genes controlling BPH resistance in rice might also play functional roles in other crops. This so-called ‘interolog’ transfer was based on the observation that orthologs of interacting proteins in one organism often exhibit interaction in another organism leading to the establishment of better orthology relationships (Walhout et al. 2000; Yu et al. 2004). It was observed that protein OsJ_28113 belonging to disease resistance NB-LRR family is encoded by three BPH-R genes: Bph9, Bph18 and Bph26 which can be targeted altogether for disease management. The ability to alter this information lays down the foundation for future functional genomic approaches (CRISPR- Clustered regularly interspaced short palindromic repeats, RNAi- RNA interference, Genetic engineering) to control susceptibility against BPH in rice.
utilising comparative modelling and molecular docking, the binding mechanisms of salicylic acid and jasmonic acid to the domains of BPH-R proteins were explored. The essential cellular processes, like gene regulation and signal transduction, rely on sequence-specific molecular recognition to direct proteins towards preferential interaction with specific nucleic acid or polypeptide ligands. The strength and specificity of such ‘sequence recognition’ play a crucial role in protein–ligand stability (Crocker et al. 2015; Farley et al. 2015; Tanay 2006; Rube et al. 2022). Protein–ligand binding free energy calculations are resultant of sequence-specific molecular interactions, salt bridges and hydrogen bonds interactions in the docking region along with the structural changes during complex unfolding (Fu et al. 2018) Moreover, the calculation of ligand binding affinities revealed that jasmonic acid dependent pathway promotes resistance against BPH with higher affinity than salicylic acid The variation in the hypervariable region of LRR repeats contributes to new ligand binding specificities for broader interaction efficiency. This research aids in advancing our knowledge of the complex mechanisms underlying the interaction of R proteins and ligands, indicating the co-evolutionary arms race between plant resistance and insect adaptation mechanism.
Conclusion
The findings from the in silico study lead to the conclusion that the adaptation of plant species to insect pest elicitors/effector molecules is a result of the ongoing variation in resistance genes as plants and insect pests continue to interact. To gain a deeper insight into the interaction between R genes and elicitors during BPH infection of rice planthoppers, and the activation of signalling cascades during this process, further molecular studies could be pursued. Moreover, a promising avenue for future research involves the development of BPH-resistant rice varieties by combining DNA-encoding BPH-R genes with phytochemicals. Such an approach holds potential for enhancing the plant's defense mechanisms against BPH infestations, paving the way for more effective pest control strategies in rice cultivation.
Availability of data and materials
All the data generated and analysed in this study are available in this article as supplementary material.
Abbreviations
- 3-D:
-
Three-dimensional
- QTL:
-
Quantitative trait loci
- CC-NBS-LRR:
-
Coiled-coil, nucleotide-binding site and leucine-rich repeat
- SCR:
-
Short consensus repeat
- NCBI:
-
National Center for Biotechnology Information
- RGAP:
-
Rice Genome Annotation Project
- BLAST:
-
Basic Local Alignment Search Tool
- BAC:
-
Bacterial artificial chromosome
- FASTA:
-
Fast-all
- STRING:
-
Search Tool for the Retrieval of Interacting Genes/Proteins
- NMR:
-
Nuclear magnetic resonance spectroscopy
- CRISPR:
-
Clustered regularly interspaced short palindromic repeats
- RNAi:
-
RNA interference
References
Appel RD, Bairoch A, Hochstrasser DF (1994) A new generation of information retrieval tools for biologists: the example of the ExPASy WWW server. Trends Biochem Sci 19:258–260
Bailey TL, Williams N, Misleh C, Li WW (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34(suppl_2):W369–W373. https://doi.org/10.1093/nar/gkl198
Boeckmann B, Bairoch A, Apweiler R, Blatter MC, Estreicher A, Gasteiger E, Martin MJ, Michoud K, O’Donovan C, Phan I (2003) The SWISS-PROT protein knowledge base and its supplement TrEMBL. Nucleic Acids Res 31:365–370
Brar DS, Virk PS, Jena KK, Khush GS (2009) Breeding for resistance to planthoppers in rice Planthoppers: new threats to the sustainability of intensive rice production systems in Asia, vol 5. International Rice Research Institute, Los Baños, pp 401–428
Cabauatan PQ, Cabunagan RC, Choi IR (2009) Rice viruses transmitted by the brown planthopper Nilaparvata lugens Stål Planthoppers: new threats to the sustainability of intensive rice production systems in Asia. International Rice Research Institute, Los Baños, pp 357–368
Cagampang GB, Pathak MD, Juliano BO (1974) Metabolic changes in rice plant during infestation by the brown planthopper, Nilaparvata lugens Stål (Hemiptera:Delphacidae). Appl Entomol Zool 19:174–184
Cha YS, Ji H, Yun DW, Ahn BO, Lee MC, Suh SC, Lee CS, Ahn EK, Jeon YH, Jin ID, Sohn JK (2008) Fine mapping of the rice Bph1 gene, which confers resistance to the brown planthopper (Nilaparvata lugens Stål), and development of STS markers for marker-assisted selection. Mol Cells 31:146–151
Crocker J, Abe N, Rinaldi L, McGregor AP, Frankel N, Wang S, Alsawadi A, Valenti P, Plaza S, Payre F, Mann RS (2015) Low affinity binding site clusters confer hox specificity and regulatory robustness. Cell 160(1):191–203
DeYoung BJ, Innes RW (2006) Plant NBS-LRR proteins in pathogen sensing and host defense. Nat Immunol 7:1243–1249
Du B, Zhang W, Liu B, Hu J, Wei Z, Shi Z, He R, Zhu L, Chen R, Han B, He G (2009) Identification and characterization of Bph14, a gene conferring resistance to brown planthopper in rice. Proc Natl Acad Sci USA 106:22163–22168
Farley EK, Olson KM, Zhang W, Brandt AJ, Rokhsar DS, Levine MS (2015) Suboptimization of developmental enhancers. Science 350(6258):325–328
Fu Y, Zhao J, Chen Z (2018) Insights into the molecular mechanisms of protein-ligand interactions by molecular docking and molecular dynamics simulation: a case of oligopeptide binding protein. Comput Math Methods Med 4:3502514. https://doi.org/10.1155/2018/3502514
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) The proteomics protocols handbook. Proteom Protoc Handb. https://doi.org/10.1385/1592598900
Guo J, Xu C, Wu D, Zhao Y, Qiu Y, Wang X, Ouyang Y, Cai B, Liu X, Jing S, Shangguan X, Wang H, Ma Y, Hu L, Wu Y, Shi S, Wang W, Zhu L, Xu X, Chen R, Feng Y, Du B, He G (2018) Bph6 encodes an exocyst-localized protein and confers broad resistance to planthoppers in rice. Nat Genet 50:297–306
Gupta MK, Vadde R, Donde R et al (2019) Insights into the structure–function relationship of brown plant hopper resistance protein, Bph14 of rice plant: a computational structural biology approach. J Biomol Struct Dyn 37:1649–1665. https://doi.org/10.1080/07391102.2018.1462737
Hu L, Wu Y, Wu D et al (2017) The coiled-coil and nucleotide binding domains of BROWN PLANTHOPPER RESISTANCE14 function in signalling and resistance against planthopper in rice. Plant Cell 29:3157–3185. https://doi.org/10.1105/tpc.17.00263
Ishaq W, Memon SQ (2017) Roles of women in agriculture: a case study of rural Lahore, Pakistan. J Rural Dev Agric 1:1–11
Jacob F, Vernaldi S, Maekawa T (2013) Evolution and conservation of plant NLR functions. Front Immunol. https://doi.org/10.3389/fimmu.2013.00297
Ji H, Kim SR, Kim YH, Suh JP, Park HM, Sreenivasulu N, Misra G, Kim SM, Hechanova SL, Kim H, Lee GS, Yoon UH, Kim TH, Lim H, Suh SC, Yang J, An J, Jena KK (2016) Map-based cloning and characterization of the Bph18 gene from wild rice conferring resistance to brown planthopper (BPH) insect pest. Sci Rep 6:343–376
Khush GS, Karim AR, Angeles EG (1985) Genetics of resistance of rice cultivar ARC10550 to Bangladesh brown planthopper biotype. J Genet 64(2):121–125
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547. https://doi.org/10.1093/molbev/msy096
Kumar K, Kaur P, Kishore A et al (2020) Recent advances in genomics-assisted breeding of brown planthopper (Nilaparvata lugens) resistance in rice (Oryza sativa). Plant Breed 139:1052–1066
Leister D (2004) Tandem and segmental gene duplication and recombination in the evolution of plant disease resistance genes. Trends Genet 20:116–122. https://doi.org/10.1016/jtig200401001
Liu YQ, Wu H, Chen H, Liu YL, He J, Kang HY, Sun ZG, Pan G, Wang Q, Hu JL, Zhou F, Zhou KN, Zheng XM, Ren YL, Chen LM, Wang YH, Zhao ZG, Lin QB, Wu FQ, Zhang X, Guo XP, Cheng XN, Jiang L, Wu CY, Wang HY, Wan JM (2015) A gene cluster encoding lectin receptor kinases confers broad spectrum and durable insect resistance in rice. Nat Biotechnol 33:301–305
Mchale L, Tan X, Koehl P, Michelmore RW (2006) Plant NBS-LRR proteins: adaptable guards. Genome Biol 7:212. https://doi.org/10.1186/gb-2006-7-4-212
Meyers BC, Kozik A, Griego A, Kuang H, Michelmore RW (2003) Genome-wide analysis of NBS-LRR—encoding genes in Arabidopsis. Plant Cell 15(4):809–834. https://doi.org/10.1105/tpc009308
Michiels K, Van Damme EJ, Smagghe G (2010) Plant-insect interactions: what can we learn from plant lectins? Arch. Insect Biochem Physiol 73:193–212
Normile D (2008) Reinventing rice to feed the world. Sci 321:330–333
O’Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR (2011) Open Babel: An open chemical toolbox. J Cheminform. https://doi.org/10.1186/1758-2946-3-33
Parham P (2005) The immune system, 2nd edn. Garland Science, New York, pp 244–245
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612. https://doi.org/10.1002/JCC.20084
Punta M, Coggill PC, Eberhardt RY, Mistry J, Tate J, Boursnell C, Pang N, Forslund K, Ceric G, Clements J (2012) The Pfam protein families database. Nucleic Acids Res 40:D290–D301
Qian LH, Zhou GC, Sun XQ et al (2017) Distinct patterns of gene gain and loss: diverse evolutionary modes of NBS-encoding genes in three Solanaceae crop species. G3 Genes Genom Genet 7:1577–1585. https://doi.org/10.1534/g3.117.040485
Rashid MM, Jahan M, Islam KS, Latif MA (2017) Ecological fitness of brown planthopper, Nilaparvata lugens (Stål) to rice nutrient management. Ecol Process 6:1–10
Ren J, Gao F, Wu X, Lu X, Zeng L, Lv J, Su X, Luo H, Ren G (2016) Bph32, a novel gene encoding an unknown SCR domain-containing protein, confers resistance against the brown planthopper in rice. Sci Rep 6:37645
Rube HT, Rastogi C, Feng S, Kribelbauer JF, Li A, Becerra B, Melo LA, Do BV, Li X, Adam HH, Shah NH (2022) Prediction of protein–ligand binding affinity from sequencing data with interpretable machine learning. Nat Biotechnol 40:1520–1527. https://doi.org/10.1038/s41587-022-01307-0
Shi S, Wang H, Nie L et al (2021) Bph30 confers resistance to brown planthopper by fortifying sclerenchyma in rice leaf sheaths. Mol Plant 14:1714–1732. https://doi.org/10.1016/j.molp.2021.07.004
Sogawa K (1982) The rice brown planthopper: feeding physiology and host plant interactions. Annu Rev Entomol 27:49–73
Sogawa K (2015) Planthopper outbreaks in different paddy ecosystems in asia: man-made hopper plagues that threatened the green revolution in rice. In: Heong K, Cheng J, Escalada M (eds) Rice planthoppers. Springer, Dordrecht, pp 33–63
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, Jensen LJ, Mering C (2017) The STRING database in 2017: quality-controlled protein-protein association networks made broadly accessible. Nucleic Acids Res 45:362–368
Takken FL, Albrecht M, Tameling W (2006) Resistance proteins: molecular switches of plant defence. Curr Opin Plant Biol 9:383–390. https://doi.org/10.1016/jpbi200605009
Tamura Y, Hattori M, Yoshioka H, Yoshioka M, Takahashi A, Wu J, Sentoku N, Yasui H (2014) Map-based cloning and characterization of a brown planthopper resistance gene BPH26 from Oryza sativa L. ssp. indica cultivar ADR52. Sci Rep 4:5872
Tanay A (2006) Extensive low-affinity transcriptional interactions in the yeast genome. Genome Res 16(8):962–972
Vandenborre G, Smagghe G, Van Damme EJ (2011) Plant lectins as defense proteins against phytophagous insects. Phytochemistry 72:1538–1550
Walhout AJ, Sordella R, Lu X, Hartley JL, Temple GF, Brasch MA, Thierry-Mieg N, Vidal M (2000) Protein interaction mapping in C. elegans using proteins involved in vulval development. Sci 287:116–122
Wang HY, Yang Y, Su JY, Shen JL, Gao CF, Zhu YC (2008) Assessment of the impact of insecticides on Anagrus nilaparvatae (Hymenoptera: Mymanidae), an egg parasitoid of the rice planthopper, Nilaparvata lugens (Hemiptera: Delphacidae). Crop Prot 27:514–522
Wang Y, Cao LM, Zhang YX (2015) Map-based cloning and characterization of BPH29, a B3 domain-containing recessive gene conferring brown planthopper resistance in rice. J Exp Bot 66:6035–6045
Wang Z, Huang J, Nie L, Hu Y, Zhang N, Guo Q, Guo J, Du B, Zhu L, He G, Chen R (2021) Molecular and functional analysis of a brown planthopper resistance protein with two nucleotide-binding site domains. J Exp Bot 72(7):2657–2671
Wu D, Guo J, Zhang Q, Shi S, Guan W, Zhou C, Chen R, Du B, Zhu L, He G (2022) Necessity of rice resistance to planthoppers for OsEXO70H3 regulating SAMSL excretion and lignin deposition in cell walls. New Phytol 234:1031–1046
Yamasaki K, Kigawa T, Inoue M, Tateno M, Yamasaki T, Yabuki T, Aoki M, Seki E, Matsuda T, Tomo Y, Hayami N (2004) Solution structure of the B3 DNA binding domain of the Arabidopsis cold-responsive transcription factor RAV1. Plant Cell 16(12):3448–3459
Yu H, Luscombe NM, Lu HX, Zhu X, Xia Y, Han JD, Bertin N, Chung S, Vidal M, Gerstein M (2004) Annotation transfer between genomes: protein-protein interologs and protein-DNA regulogs. Genome Res 14:1107–1118
Zhao Y, Huang J, Wang Z, Jing S, Wang Y, Ouyang Y, Cai B, Xin XF, Liu X, Zhang C, Pan Y, Ma R, Li Q, Jiang W, Zeng Y, Shangguan X, Wang H, Du B, Zhu L, Xu X, Feng YQ, He SY, Chen R, Zhang Q, He G (2016) Allelic diversity in an NLR gene BPH9 enables rice to combat planthopper variation. Proc Natl Acad Sci USA 113:12850–12855
Zheng X, Xin Y, Peng Y, Shan J, Zhang N, Wu D, Guo J, Huang J, Guan W, Shi S et al (2021) Lipidomic analyses reveal enhanced lipolysis in planthoppers feeding on resistant host plants. Sci China Life Sci 64:1502–1521
Acknowledgements
The authors gratefully acknowledge the financial support provided by DST/INSPIRE Fellowship. Facilities provided by the School of Agricultural Biotechnology are duly acknowledged.
Funding
This work was supported by DST/INSPIRE Fellowship [IF190371, CRG/2018/001833, and BT/PR31475/ATGC/127/5/2019].
Author information
Authors and Affiliations
Contributions
KN, RP, KK: conceptualisation of the research, proofreading of the manuscript; PK: carried out the study and writing of the original draft; PK, KK, RK: carried out data analysis. All co-authors approved this manuscript before submission.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Human and animal rights
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for publication
Additional informed consent was obtained from all individual participants included in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations
Supplementary Information
Below is the link to the electronic supplementary material.
42485_2023_123_MOESM1_ESM.tif
Supplementary file1 Supplementary Figure 1: Flowchart depicting the sequence of methods followed for the study (TIF 156 KB)
42485_2023_123_MOESM2_ESM.tif
Supplementary file2 Supplementary Figure 2: Neighbour joining tree representing relationship among the map-based cloned BPH resistance genes (TIF 29 KB)
42485_2023_123_MOESM3_ESM.tif
Supplementary file3 Supplementary Figure 3: Graphical representation of amount of different secondary structures in the BPH resistance proteins (TIF 1312 KB)
42485_2023_123_MOESM4_ESM.tif
Supplementary file4 Supplementary Figure 4: Docking of cloned BPH resistant genes with ligand: jasmonic acid and salicylic acid (TIF 3555 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kaur, P., Kumar, P., Kumar, K. et al. In silico analysis of cloned brown planthopper genes unveiled OsJ_28113 as a key regulator in triggering resistance response in rice. J Proteins Proteom 15, 53–66 (2024). https://doi.org/10.1007/s42485-023-00123-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42485-023-00123-7