Abstract
The development of high-throughput techniques with evolving bioinformatics tools has elucidated the adaptation mechanism of plant to certain environmental stress like salinity. The characterization of proteins is critical for plant stress responses and it provides the comprehensive information about the cellular and biochemical pathway involved in various stress mitigation. In the present work, we provide the information about the alteration in wheat proteomic profile in response to plant growth-promoting rhizobacterial (PGPR)-inoculation under abiotic stress like salinity. PGPR facilitate the plant growth and enhance their induced systemic tolerance (ISR) under various stress conditions. The present study was aimed to generate the AcdS− mutant of Enterobacter cloacae SBP-8 which differs in its ability to breakdown the stress ethylene precursor ‘ACC’ (1-aminocyclopropane-1-carboxylic acid). The proteomic profile of wheat (Triticum aestivum L.) plant was investigated under non-saline and high salinity stress (200 mM NaCl) following wild type and its mutant inculcation. A total of four treatments were taken to monitor the differential expression of proteins in the wheat seedlings exposed to high salt stress for 15 days. The major changes concerned the proteins involved in metabolism, ion-transport, photosynthesis, defense and stress responses. The observed changes at the proteomic level in each treatment can be related to effects of salt and bacterial inoculation. The identified proteins were further classified into cellular, biological and molecular function. Bacterial inoculation significantly enhanced the expression of Thioredoxin and Ninja family proteins in addition to Heat shock proteins (Hsp70, Hsp 90), which play a major role in defense against abiotic stress. Taken together, the observed results suggest that bacterial inoculation alleviated the salinity-induced damages by improving the metabolism, photosynthesis and defense-related proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Agricultural crops are often subjected to several stresses which impart an adverse effect or can be lethal also. Severe stress such as heat, salinity, drought, and flooding are the main cause for inhibiting the production of agricultural important crop plants (Suzuki et al. 2014). Among these stresses, salinity is a major abiotic stress factor that limits crop productivity in with increasing negative effects on soil fertility (FAO (2005). Furthermore, the increased salinity stress resulted in oxidative stress which affects the protein functionality and also damage of lipids and nucleic acids (Apel and Hirt 2006; Flagella et al. 2006). In addition, the gradual changing climate of current scenario is also reducing the overall productivity (Singh et al. 2018). Therefore, it is necessary to understand the molecular mechanism of stress protection involved in crop endurance to certain stress.
In response to salt stress, plants perceive stress signals and respond through various adaptive mechanisms to maintain growth and productivity like production of osmolytes, ionic hemeostasis, growth regulation, ROS scavenging activity, etc. These adaptation leads to multiple changes in biochemical, physiological metabolic pathway as well as certain changes in gene expression that might disturb the cellular homeostasis (Munns et al. 2006; Aghaeiet al. 2008). To ameliorate salinity stress, one approach is the use of ACCD-producing plant growth-promoting bacteria which showed the plant growth improvement even at high salt concentrations (Singh et al. 2015). Use of such biocatalyst has been shown to be effective and eco-friendly (Lin et al. 2014; Xu et al. 2014). A variety of mechanism conferred by these microorganism to increase the crop productivity by various mechanisms like production of various phytohormones (Kochar et al. 2011; Singh et al. 2015), providing the nutrients availability (Idriss et al. 2002), and boosting the induced systemic tolerance (IST) (Singh et al., 2015) and induced systemic resistance (ISR) in plants (Niu et al. 2011; Singh and Jha 2016). As a consequence of severe environmental stresses, the stress ethylene level elevated to cause overall inhibitory effects to plant growth (Stearns and Glick, 2003). Under these condition, ACCD bacteria cleaves ACC, the precursor of ethylene to α-ketobutyrate and ammonia (Glick et al. 1998, 1999), thereby lowering the ethylene level to trigger plant protective mechanism (Stearns and Glick 2003). Previous studies (Li and Glick 2001; Sun et al. 2009) have shown that AcdS− mutant strains lacking a functional AcdS gene (encoding ACC deaminase) do not prevent stress ethylene production that severely inhibits the plant growth. The benefits of using ACCD-producing bacteria have certain advantages to exhibit its stress ameliorating activity under diverse conditions and, therefore, confer plant tolerance to a number of different environmental stresses. The effectiveness of ACCD-producing bacteria to counteract the abiotic and biotic stresses has been shown under various conditions like salinity (Singh et al. 2015; Mayak et al. 2004), flooding (Grichko and Glick 2001a, b), metal stress(Belimov et al. 2005), drought (Saradadevi et al. 2017) and pathogens (Wang et al. 2008).
AcdS mutant of Enterobacter cloacae SBP-8 was generated by transposon mutagenesis. Transposon mutagenesis is a technique of creating random mutations and it is easy to create a complete library of mutants. It involves the introduction of the EZ::TN5 < DHFR-1 > TnpTransposome via electroporation. The transposon contains the dihydrofolate reductase (dhfr) gene, which confers trimethoprim resistance (Mates et al. 2007). Most of the proteomic study on wheat has been done by a two-dimensional gel electrophoresis; however, it has certain limitation like poor resolution of proteins having high molecular weight, high pI value, etc. The iTRAQ-based (isobaric tags for relative and absolute quantitation) quantitative analysis of protein expression in combination with LC–MS/MS (liquid chromatography/tandem mass) facilitate the detail analysis of the differential expressed proteins (Zhang et al. 2015). It also allows detection of a large number of proteins without any limitation like stable isotope labelling, poor reproducibility, exceedingly large or small proteins, as well as basic proteins, etc.
Wheat is a major staple food worldwide, and also represents the greatest source of protein in the human diet (Matsuoka et al. 2008). Among the wheat grain proteins, the major ones are prolamines, gliadins, glutenins and non-prolamins including albumins and globulins (Vensel et al. 2005). In response to beneficial bacterial inoculation, many of the plant proteins have been identified as being involved in plant–bacterial interactions, resistance to pathogenic microorganisms, nutrient fixation, etc. The present proteomic study was aimed to dissect the molecular events induced in response to ACCD-producing wild-type (AcdS+) bacteria and its AcdS− mutant under controlled environment of plant growth chamber.
Materials and methods
Bacterial strain and inoculum preparation
In the present study, an ACC deaminase-producing bacterium Enterobacter cloaca SBP-8 was selected based on its plant growth enhancing properties under salt-stress conditions (Singh et al. 2015). The strain was identified as Klebsiella sp. based on partial 16S rRNA gene sequencing, however, later on by full-length sequencing confirmed it as E. cloacae (Singh and Jha 2017). Transposon mutagenesis of selected isolate was performed using EZ::TNTM < DHFR-1 > Transposome TM Kit (Epicentre, USA) following instructions of manufacture. Further detail about the competent cells preparation and electroporation are provided in supplementary File S1. A mixture of competent cells, and transposon was prepared and incubated for 15 min on ice. The mix was transferred into an electroporation cuvette (BioRad, USA), and cells were electroporated at 25 ºC in an electroporator (BioRad, USA) set at 1250 V. After electroporation, the cells were immediately transferred on ice and cultured in 1 ml of SOC medium at 30 °C for 1 h. The 100 µl of cultured cells were spread on Muller-Hinton (M-H) agar plates containing Tm (30 µg/ml) and were incubated at 30 °C for overnight. Bacterial colonies growing on Muller Hinton-Tm agar plates were selected for further confirmation of ACC utilization by growing on the DF-ACC plate. The bacterial colony unable to grow on DF-ACC agar plate was further selected for ACC deaminase enzymatic assay and plant growth test against wild type.
Plant growth and treatments
Wheat plants (Triticum aestivum C-309) was grown under controlled environment of plant growth chamber (Labtech, South Korea) and treated with test isolate (SBP-8) and its AcdS−mutant following standard procedure (Penrose and Glick 2003). The detail about the plant growth and bacterial treatment has been provided in Supplementary File S1. The following sets of treatment were included in the study for protein comparison; T-1 comprised of control plants and AcdS− mutant without salt stress, T-2 control plants and AcdS− mutant with salt stress, T-3 wild type (AcdS+) and AcdS− mutant without salt stress, whereas T-4 included wild type (AcdS+) and AcdS− mutant with salt stress. The plant growth promotion following inoculation of wild type SBP-8 and its AcdS− mutant was evaluated to check the various physiological parameters of wheat (Triticum aestivum) plant.
Protein extraction
Wheat plant (1 g) was ground to powder in liquid nitrogen and further extracted in buffer containing 0.9 M sucrose, 0.5 M Tris–HCl, 5 mM EDTA, 0.1 M KCl, 1% w/v dithiothreitol (DTT) up to 1 ml of volume. The mixture was sonicated for 5 min and mixed with 100% tricarboxylic acid (TCA) and 100% ice-cold acetone in 1:8 ratios. After mixing, the samples were centrifuged at 12,000g for 15 min at 4 °C and obtained pellet was re-suspended in 1 ml ice-cold acetone. Pellet was dried using a Speed-Vac concentrator (Savant Instruments, Hickville, NY, USA) and finally dissolved in 50 mM ammonium bicarbonate with 1%SDS.
Protein enrichment and digestion
Bradford assay was used for estimating the protein concentration and 100 µg protein was further proceed for alkylation with 50 mM iodoacetamide and tryptic digestion. Detail has been provided in supplementary File S1. In brief, the peptides were analyzed on the Waters Synapt G2 Q-TOF instrument and the raw data were processed by MassLynx 4.1 WATERS (version 2.5.1; Matrix Science, London, UK). The individual peptides MS/MS spectra were matched to the database sequence for protein identification on PLGS (Protein Lynx Global Server) software. The peptide ratio > 1.5 were rated as up-regulated, whereas value < 0.75 as down-regulated. Assignment of varying expressed proteins were determined accordingly Gene-ontology (http://www.geneontology.org) for their various functionality like cellular, biological and molecular functions. Similarly, Blast-2 Go algorithm (http://www.blast2go.com) was used for predicting cellular and molecular functionality of identified proteins. The expression profile of 185 differential expressed proteins common to tested treatments (T-1, T-2, T-3, T-4) was constructed through the 2-way hierarchical clustering according to the Permut-Matrix software. Green and red colors were used for demonstrating the high and low expression level of expressed proteins in each treatment.
Statistical analysis
The experiments were performed in triplicates and results were expressed in terms of mean ± standard deviation (SD). Data was analyzed by analysis of variance (ANOVA) and subsequently by Duncan’s multiple range test at p < 0.05.
Results
Effect of wild-type SBP-8 (AcdS +) and its mutant (AcdS −) on plant growth
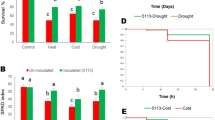
After transposon-mutagenesis, bacterial colonies growing on culture medium containing trimithoprim were selected and replica plated on minimal agar medium possessing 3 mM ACC. Bacterial colonies which did not showed growth on ACC containing selective medium were further subjected to ACC deaminase assay. As compared to wild type, mutant strain did not show the ACC deaminase enzymatic activity. We checked the other PGP traits in AcdS− SBP-8 strain including growth pattern, which were almost found to be unaffected as compared to wild type. After confirmation of mutation, we tested the plant growth promontory effect of mutant and wild-type strain under salt stress. Mutation in the AcdS gene significantly (p < 0.05) affected the various growth parameters of wheat plant when compared to the wild type at 200 mM NaCl stress (Fig. 1a). As seen from Fig. 1a, shoot length and root length was significantly reduced by 28.55% (p < 0.05) and 24.18% (p < 0.05), respectively. Evaluation of biomass in terms of fresh weight/dry weight revealed that mutation in the AcdS gene significantly reduced the fresh weight of 34.24% (p < 0.05) and dry weight of 38.5% (p < 0.05) as compared to wild type (Fig. 1b).
Comparative evaluation of plant growth promotion test following inoculation of a E. cloacae SBP-8 (AcdS+) and SBP-8 (AcdS−) at 200 mM NaCl stress (b; i) plant growth in terms of the shoot and root length (b; ii) biomass measurement in terms of fresh weight and dry weight. Double asterisk (**) represent the significant difference as compared to respective control plants
Annotation and classification of proteins
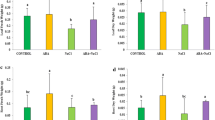
The identified proteins were divided into various functional groups like cell division and chromatid-associated protein, lipid biosynthesis, ion-transport, defense-related protein, proteins involved in metabolic and photosynthesis pathway, protein synthesis, plant growth, seed storage, stress tolerance and other proteins associated with known biological functions. The response of wheat to bacterial inoculation (AcdS− mutant) is the up-regulation of proteins involved in cell division, metabolic pathway, ionic transport, etc. (Fig. 2). Based on the relative score of protein, we calculated the upregulated and downregulated proteins in the percentage (%) form. The highest up-regulation was observed for metabolic pathway protein (226%), followed by stress protein (153%), plant growth and development (131%), lipid biosynthesis (59%), and ionic transport (30%) (Fig. 2a).
The functional group of protein identified in each treatments a treatment T1, b treatment T2, c treatment T3, d treatment T4. Protein showing > 1.5 fold ratio of expression level was considered as positive regulation, whereas value < 0.75 was considered as negative regulation. S1: Cell division; S2: Chromatin associated proteins; S3: Defence and pathogenesis proteins; S4: Ion transport; S5: Lipid Biosynthesis; S6: Metabolic pathway; S7: Photosynthesis; S8: Plant growth and development; S9: Protease inhibitor; S10: Protein synthesis; S11: Seed storage; S12: Stress related proteins; S13: Other proteins
Following salt stress, in the presence of bacterial-inoculation (AcdS− mutant), the up-regulation was observed for defense-responses (173%), photosynthesis (99%), stress tolerance (60%), and ionic transport (21%) (Fig. 2b). Salinity stress leads to down-regulation of proteins belonging to category of seed storage (86%), cell division (41%), lipid biosynthesis (37%), and metabolic pathway (16%). In the other treatment (T-3), protein related to seed storage, protein synthesis, defense responses, and ionic transport were up-regulated by 565%, 428%, 199% and 112%, respectively. The down-regulated proteins were belonging to category of plant growth and development (54%), stress-related (48%), and cell division proteins (41%) (Fig. 2c).Wheat plant treated with AcdS− mutant and salt plus AcdS− mutant attributed to an increase in lipid biosynthesis (319%), seed storage protein (130%), photosynthesis (79%), and ionic transport (54%). The down-regulated proteins were protease inhibitor (55%), cell division (42%), protein synthesis (38%), plant growth and development (36%) and defense-associated proteins (23%) (Fig. 2d).
Based on significant changes of ≥ 1.5 fold or ≤ 0.75, among the 181 proteins, 40 were up-regulated and 141 proteins were down-regulated in T-1 treatment (Fig. 3a). The highest increase in up-regulated protein was observed for metabolic pathway (10), plant growth and development (5), followed by protein synthesis (4) and cell division (3). In response to salinity stress, the proteins were up-regulated for metabolic pathway (11), defense and stress-responses (6), followed by protein synthesis (5) and protease inhibitor (4). The down-regulated proteins were of metabolic pathway (30), photosynthesis (15), chromatin-associated protein (13) and cell-division protein (8) (Fig. 3b). In the T-3 treatment, the expression of 160 protein showed significant changes, of which 48 were up-regulated, and 112 were down-regulated (Fig. 3c). As compared to wild-type AcdS, mutation in the AcdS gene in response to salt stress leads to expression of total 145 protein in the treatment T-4, 36 were up-regulated and 109 were down-regulated (Fig. 3d).
The protein numbers for various functionality in each treatment a treatment T1, b treatment T2, c treatment T3, d treatment T4. S1: Cell division; S2: Chromatin associated proteins; S3: Defence and pathogenesis proteins; S4: Ion transport; S5: Lipid Biosynthesis; S6: Metabolic pathway; S7: Photosynthesis; S8: Plant growth and development; S9: Protease inhibitor; S10: Protein synthesis; S11: Seed storage S12: Stress related proteins; S13: Other proteins
Proteomic analysis
All the identified proteins were classified by gene ontology (GO) software and then classified into three functional groups: biological process, molecular function, and cellular component. The proteins belonging to category of biological process were related to metabolic processes, signalling, stress-response, whereas molecular functional proteins were found related to binding, transporter and receptor activity. List of all the identified protein in selected four treatments has been provided in Supplementary File S2. The cellular proteins were classified into membrane, organelle, vacuolar and nucleus. In the T-1 treatment, a total of 288 were identified, followed by 262, 253 and 237 proteins were recorded in T-2, T-3 and T-4 treatment respectively.
In the T-1 treatment under biological process, 33 proteins were classified for metabolic process, 13 for signalling and 11 for stress responses. Proteins related to cellular responses (10%), protein folding (8%), regulation (9%) and localization (6%) were also identified. Under molecular process, most of the identified proteins were of the binding (22%), transporter (16%) and receptor (15%) activity. Under cellular process, most of the proteins belonging to category of plasma membrane (22%), and organelle (20%) were identified (Fig. 4a). In the T-2 treatment, 31% proteins related to metabolic process, 14% and 11% for stress and cellular process. Among the molecular function, the higher 19% proteins were for binding to DNA, protein and nucleotides and 18% for the transporter activity (Fig. 4b). In the T-3 treatment, the highest 36% proteins were observed for metabolic process, followed by binding (23%) and plasma membrane proteins (20%) (Fig. 4c). Similarly, in the T-4 treatment, the highest number was observed for metabolic process (27%), followed by binding (24%), plasma membrane (22%) and organelle (20%) (Fig. 4d).
Identification and clustering of differentially expressed proteins
For identification of common and unique proteins in each treatment (T-1 to T-4), Venn diagram was constructed. A total of 288, 262, 253 and 237 proteins were observed in Treatment T-1, T-2, T-3 and T-4, respectively. After comparison, it was found that 238 proteins were common between treatment T-1 and T-2, 227 and 201 were found to be common in between T-1/T-3 and T1/T4, respectively. Among the other treatments 229, 218 and 222 were common between treatments T-2/T-3; T-2/T-4 and T-3/T-4, respectively (Fig. 5).
Based on their higher abundance and their presence in all treatments (T-1, T-2, T-3, T-4), a total of 185 differentially expressed proteins were used for hierarchical cluster analysis (Fig. 6). The information about the differential expressed proteins in selected treatments has been provided in Supplementary File S3. In the T-1 treatment, the cluster analysis shows the up-regulation of 31 proteins. The up-regulated proteins were belonging to the category of DNA binding protein EMBP 1, DELLA protein RHT 1, NADP dependent glyceraldehyde 3 phosphate dehydrogenase, fructan 1 exohydrolase w1, ADP ATP carrier protein 2, cold shock protein CS66, granule bound starch synthase 1, etc.
In the T-2 treatment, the cluster contained the 27 up-regulated and 94 down-regulated proteins. The highest up-regulation was observed for histone H3, Ubiquitin activating enzyme E1 2, tubulin beta 3 and beta 4 chain, ATP synthase subunit alpha mitochondrial, whereas down-regulated were belonging to cytochrome f, 50S ribosomal protein L2, casparian strip membrane protein 2, phosphoglycerate kinase, elongation factor 1 beta, protein disulfide isomerise, plasma membrane ATPase, elongation factor 1 alpha, ADP ATP carrier protein 2 mitochondrial, histone H2B 3 and 4, etc. In contrast to AcdS+ strain, AcdS– mutant inoculation showed the higher up-regulation of Transcription factor HBP 1b c38, followed by Inositol 3 phosphate synthase, Tubulin beta 3 chain and glutathione S transferase, whereas the maximum down-regulation was recorded for retinoblastoma related protein 1, S adenosylmethionine synthase, phosphoglycerate kinase cytosolic, elongation factor 1 beta, adenosylhomocysteinase, cold shock protein CS120, plasma membrane ATPase, homogentisate geranyl geranyltransferase, gibberellin 3 beta dioxygenase 2 1, etc.
Discussion
Various aspects of plant growth and development are largely controlled by several environmental stimulus or stress. ACC-deaminase-producing bacteria play a significant role in modulating the plant responses to abiotic stress like salinity. Under the effect of various abiotic environmental stress, plant develop several adaptive mechanism to cope with stresses. These adaptive mechanism could be the up-regulation of proteins belonging to the category of stress-responses, photosynthesis, metabolism, ion-transport and protein synthesis. Therefore, comparative analysis of differentially expressed proteins under various treatments can provide the more insight of plant responses to stress. In the present study, we monitored the proteomic analysis of wheat plant under salt stress following inoculation of Enterobacter cloacae SBP-8 AcdS+ and AcdS– strain. In our study, the major protein belonging to photosynthesis, metabolism, ionic transport and defense-responses were altered in each treatment.
Defense and stress proteins
Under abiotic stress, plants have always suffered from reactive oxygen species (ROS) that is primarily involved in symbiotic interaction between plants and microorganisms (Scheler et al. 2013). To counteract the produced ROS, plants have developed antioxidative machinery to scavenge these molecules. In the present study, catalase and peroxidase were identified and their up-regulation was significantly increased by the inoculation of bacterial strain SBP-8. Regardless of whether they were inoculated with SBP-8 AcdS+ or SBP-8 AcdS−, the expression levels were both up-regulated as compared to control. Catalase is primarily involved in the detoxification of H2O2, senescence and stress response (Almagro et al. 2009). Similarly, 2 Cys-peroxiredoxin acts as an antioxidant enzyme particularly important in the developing shoot and photosynthesizing leaf (Singh and Jha 2017). Previous studies have shown that bacterial inoculation mitigates the effects of abiotic stress-induced oxidative burst by increasing the antioxidative enzyme activity (Israr et al. 2016; Youssef et al. 2016).
Thioredoxins assumed to play a key role in plant protection to oxidative stress by supplying power to reductases, repairing various oxidized proteins or by detoxification of lipid hydroperoxides (Vieira Dos Santos and Rey 2006). It was observed that with respect to salt stress of 200 mM NaCl, the level of few defence proteins such as thioredoxin and Ninja family protein was down-regulated, however, bacterial inoculation increased the level of these proteins under salt stress. Till now, the role of Ninja family proteins in plant defense has not been well known, however, they might act as regulator of abscisic acid (ABA) production during seed germination through the ubiquitin-mediated proteolysis of ABI5/DPBF1 (Singh and Jha 2017). In addition, bacterial inoculation also increased the expression of Hsp 70, Hsp 90 and cold shock protein under salt stress, illustrating that in presence of bacterial inoculum plant was protected from the stress conditions. The up-regulation of these proteins provides the cross-tolerance of wheat plants in response to biotic and abiotic stress. The expression of dehydrin COR410 provides the tolerance to salinity-induced water stress and expressed in roots and leaves during cold acclimation. Xylanase inhibitor protein 1 competitively inhibiting the activity against fungal endo-1,4-beta-d-xylanases belonging to glycoside hydrolase family 10 (GH10) and GH 11, may function in plant defense against secreted fungal pathogen xylanases. It is similar to class III chitinases, however, it does not exhibit the chitinase activity. The Alpha-amylase trypsin inhibitor is involved in defense mechanisms against biotic stress. The expression of pathogenesis-related protein provide tolerance to pathogens, however, their role in salinity stress has been demonstrated in few crop plants (Huang et al. 2012; Wang et al. 2011). Previous study (Ohshima et al. 1990) has shown induction of pathogenesis-related protein by general stress, however, in this study, all the pathogen-responsive proteins induced in the presence and absence of both SBP-8 AcdS+ or SBP-8 AcdS− strain. The defense proteins involved in growth promotion and plant protection were significantly induced in Arabidopsis following inoculation with Paenibacillus polymyxa E681 (Kwon et al. 2016). Wheatwin proteins show antifungal activity against pathogenic fungi and towards the wheat-specific pathogenic fungi F. culmorum and F. graminearum.
Proteins associated with cell wall and cellular structure maintenance
The cell wall of plant is involved in different biological processes like morphogenesis, cell expansion and response tobiotic and abiotic stress (Szymanski and Cosgrove 2009; Hématy et al. 2009). Maintaining the functional integrity of the cell wall during adverse conditions is essential for carrying out optimal activity, osmotic balance and homeostasis. In the present study, bacterial inoculation enhanced the expression of cytoskeletal protein ‘Tubulin’ that maintain the cell wall integrity (Shoji et al. 2006). The increase in the expression of ‘Casparian strip membrane protein’ (CASP) and ‘Xyloglucanendotransglycosylase (XET)’ was observed in bacterial inoculated plants as compared to uninoculated plants. Casparian strip membrane protein recruits the lignin polymerization machinery in the endodermis to prevent the lateral diffusion of molecules and also regulates the cell-wall to membrane junction (Roppolo et al. 2014). CASP form complex with other CASPs and required for a constituent of a plant junctional complex. CASP are shown to establish a plasma membrane and extracelluar diffusion barrier in plants (Roppolo et al. 2011). The net like arrangement of CASPs provide a seal for extracellular space, while allowing the nutrient and water transport across the outer and inner plasma membrane surfaces. The molecular processes involved in regulation the CASPs formation is not properly understood, however, CASP proteins are anchored by an unknown mechanism that might include their interaction with the cytoskeleton or the cell wall (Roppolo and Geldner 2012). Similarly, xyloglucanendotransglycosylase (XET) reconstructs the existing cell wall of xyloglucan by catalyzing the transglycosylation to previous wall-associated xyloglucan molecules (Munoz-Bertomeu et al. 2013). XET enzymes of plants belong to glycoside hydrolase family 16 (GH16). It also modifies the xyloglucan-cellulose framework of plant cell walls, thereby regulating their expansion and strength (Nishitani 1997). In addition for their role in the formation of secondary walled xylem and phloem fibres during secondary wall deposition, the enzyme appears to be of importance in the formation of sieve tubes also (Zhang et al. 2012). The increase in cell wall extensibility in wheat was correlated with enhanced XET activity under abiotic stress conditions (Zhong et al. 2008).
Transporter proteins
Calmodulin (CaM) is a Ca2+-binding regulatory protein widely distributed in plants and animals, and its persistence was observed in transduction of Ca2+ signals. Many of the CaM-binding proteins expressed under stress response in plants, illustrating its role in unfavourable environmental conditions (McCormack et al. 2005). A total of 50 different types of CaM-binding proteins associated to different processes like transport of ions, phytohormone signalling, cytoskeleton function, protein folding, transcriptional regulation, etc. have been identified (Bouche et al. 2005). Plasma membrane-ATPases are major species of membranes ion pumps (Luo et al. 1999) which are integral membrane proteins that use chemical energy to transfer protons to the extracellular environment by a primary active transport (Hasegawa et al. 2000). These plasma membrane bound pump are actively involved in many physiological processes in plants like salt tolerance. Recently, a number of studies have demonstrated that the expression of the plasma membrane-ATPase gene increased in response to salt stress (Kuhlbrandt 2004; Hohm et al. 2014).
Mitochondrial carrier protein ADP-ATP carrier, mediates the export of ATP synthesized within mitochondria and import the phosphate into matrix for oxidative phosphorylation (Klingenberg 2008). These carrier proteins contain the membrane anchoring domains that span the inner mitochondrial membrane and exchange ADP for ATP in a 1:1 ratio (Pedersenet al. 2012). In addition, plant utilize the Na+/H+ antiporters (SOS1 and NHX1) to avoid the cytotoxicity through maintaining the cytosolic ion concentrations. NHX1 is localized in tonoplast to reduce the cytosolic Na+ content by pumping Na+ into the vacuole, while the plasma membrane bound SOS1 extrudes Na+ into apoplast. Similarly, the presence of SOS1 has been associated with antiporters (Na+/H+), responsible for providing the salinity tolerance and also for regulating the Na+transport from root to shoot in plants (Buch-Pedersen et al. 2000). Inoculation of both SBP-8 AcdS+ or SBP-8 AcdS−, enhance the expression of Na+/H+ antiporters, indicating that bacterial-inoculated plants were protected from salinity-induced oxidative stress.
Metabolic proteins
Several metabolic proteins belonging to category NADPH quinine oxidoreductase, adenylsuccinate synthetase, fructan 1 exohydrolase, glutathione gamma glutamylcysteinyl transferase, phosphoglycerate kinase, NADPH dependent glyceraldehydes 3 phosphate dehydrogenase, Serine carboxypeptidase, phyneylalanine ammonia lyase, fructan 1 exohydrolase, obtusifoliol 14 alpha demethylase, Glucose 1 phosphate adenyltransferase, and inositol 3 phosphate synthase were up-regulated following bacterial inoculation both in a wild type and AcdS− mutant inoculation. NADPH quinine oxidoreductase protects the cells against oxidative stress and free radical reactive species under salt stress. It is a key enzyme involved in the metabolism of quinines and performs multiple functions within the cell. This enzyme is considered as a detoxification enzyme based in its ability to detoxify the reactive quinines and its related compounds to less toxic hydroquinones (Ross et al. 2000). Fructan 1 exohydrolase is an enzyme responsible for the hydrolysis of fructans in plants and implicated in the metabolism of inulin (Guo et al. 2005). Inulin has a high biotechnological potential and used for production of ethanol and high-fructose syrup production. Inulin is also known as a prebiotic and is able to stimulate health-promoting bacterial growth in human colon (Macfarlane et al. 2008). Glutathione gamma glutamylcysteinyl transferase are encoded by a large and diverse gene family in plants and perform various catalytic functions including detoxification of herbicides, reduction of organic hydroperoxides formed during oxidative stress and catabolism of tyrosine. The expression of Glutathione gamma glutamylcysteinyl transferase is induced in response to general cellular injury caused by a range of xenobiotics and oxidative stress caused by salinity stress and herbicides (Dixon et al. 2009). It performs a broad level of stress tolerance through a role in cell signalling, involved in primary and secondary metabolism and induction of genes of flavonoid biosynthesis (Dixon et al. 2002). Phosphoglycerate kinase is a major enzyme in glycolysis and is a major ATP generating enzymes in glycolysis. Under abiotic and biotic stress, the induced expression of phosphoglycerate kinase maintains the energy level in the plants (Bharti et al. 2016). Bacterial inoculation also induced the synthesis of Phyneylalanine ammonia lyase (PAL) under salinity stress. Induction of PAL is generally associated with increased synthesis of phenolic compounds such as tannic, gallic, caffeic, chlorogenic and cinnamic acids that are effective against the stress conditions (Basha et al. 2006). In a previous study, it was found that PGPR strains Bacillus pumilus, Bacillus subtilis, Bacillus amyloliquefaciens, and Brevibacillus brevis inoculation also increased the level of PAL and total phenol content in tomato (Girish and Umesha 2005).
Serine carboxypeptidase-like (SCPL) proteins have been associated with synthesis of various metabolites that confer resistance against stress. Many of the functionality of SCPL proteins are not known. However, it estimated that they are involved in provide protection against biotic and/or abiotic stresses (Vivas et al. 2006). The enhanced expression of Fructan-1-exohydrolase following up-regulation of genes in response to bacterial-inoculation signify its role in protection against adverse environmental conditions (Hincha et al. 2002). Obtusifoliol 14 alpha demethylase maintain the membrane fluidity and permeability; modulate activity of membrane-bound proteins and ion channels. In addition, it also serve as precursors for biologically active to regulate growth and development processes. Inositol 3 phosphate synthase participates in the synthesis of phospholipids, maintaining the membrane fluidity and orientation.
Protein synthesis, protease inhibitor and seed storage protein
The differential expression of various synthetical proteins was observed following inoculation of SBP-8 (AcdS+) or SBP-8 (AcdS−) under tested treatments. The expression of various proteins like eukaryotic translation initiation factor 4E 1, eukaryotic translation initiation factor isoform 4G 1, beta amylase, peptidyl prolyl isomerise, glutathione S transferase, protein disulfide isomerase were decreased under high salinity stress (200 mM NaCl) in the absence of bacterial inoculation, whereas various important proteins like eukaryotic translation initiation factor 4G, 30S ribosomal protein S7, eukaryotic translation initiation factor 4B1, eukaryotic translation initiation factor 2, eukaryotic initiation factor 4A, 30S ribosomal protein S8, imidazoleglycerol phosphate dehydratase were up-regulated in the presence of bacterial inoculation under salinity stress. The enhanced expression of these proteins confers that plants were protected from the salinity-induced oxidative damages. The Eukaryotic translation initiation factor 4G is a multi-subunit complex and remains associated with other protein complexes to provide the stability inside the cell under changing environmental conditions. It is a crucial protein required for binding the cellular mRNA to ribosomes and is known to be cleaved as the central part of the mechanism of host translational shutoff under adverse conditions (Bellur and Woodson 2009). The 30S ribosomal protein S7is a primary binding protein that organizes the folding of the 16S rRNA and enables the subsequent binding of other ribosomal proteins like S4. Experimental evidence suggests that 30S ribosomal protein S7 interact with 50S ribosomal subunit and form a stable complex (Robert and Brakier-Gingras 2001). Eukaryotic translation initiation factor 4B1 promotes the eIF4F and eIF4A assembly via RNA-dependent ATP hydrolysis activity. Eukaryotic translation initiation factor 2 is required for initiation of translation and therefore protein synthesis and mediates the binding of tRNAMet to the ribosome in a GTP-dependent manner. This protein is highly conserved among evolutionary remote species and plays a key role in selection of the correct start codon on messenger RNA. Imidazole glycerol phosphate dehydratase catalyzes the complex reaction for biosynthesis of histidine in plants.
The expression of various protease inhibitors like Serine carboxypeptidase 3, Alpha amylase trypsin inhibitor CM1 and CM2, Endogenous alpha amylase subtilisin inhibitor, Serpin Z1A, Z1C, Z2A, Z2B, xylanase inhibitor protein 1 were decreased under tested salinity stress. However, various other important proteins responsible for degradation of damaged/abnormal proteins were up-regulated following bacterial-inoculation under tested salinity stress. The enhanced expressions of Serpins are likely to participate in various biochemical pathways to protect the tissues, cell and organs from oxidative stress (Roberts and Hejgaard 2007). ATP-dependent ClpP protease is a major contributor for mitochondrial protein and removing damaged or mis-folded proteins in mitochondrial matrix. Similarly, Ubiquitin activating enzyme E1 targets the mis-folded or unwanted protein for degradation via proteasome. Aberration of this system leads to the dys-regulation of cellular homeostasis and therefore it is an important way to regulating the protein activity (McGrath et al. 1991).
α-Amylase inhibitors plays a key role in protecting the starch and protein reserves in the endosperm against degradation, particularly under biotic stress (Franco et al. 2002). Starch is the major energy reserve source particularly in the cereal grains (Emes et al. 2003). The pyrophosphorylase (AGPase) reaction is the rate determining reaction in the biosynthesis of stored starch in amyloplasts (Tetlow et al. 2004) and adenosine diphosphate glucose is the key enzyme. The enhanced expression of these enzymes increase the starch content and grain weight under un-favourable conditions.
Photosynthesis and plant growth and development
Salinity stress finally resulted in the decrease of stomatal aperture, which primarily reduce theCO2 availability and thereby minimize the energy for the plant growth. RuBisCO (ribulose-1,5-bisphosphate carboxylase oxygenase), a stroma-localized protein, catalyzes the carboxylation of d-ribulose 1,5-bisphosphate and constitutes up to 50% of all chloroplast proteins (Maroco et al. 2002; Parry et al. 2002). We found a significant increase of the level of RuBisCO large subunit-binding protein at 200 mM NaCl in the presence of bacterial inoculums. The other increased enzymes included photosystem I P700, Photosystem II reaction center protein M, chloroplast envelope membrane protein, photosystem II reaction centre protein L, photosystem II protein D1. Photosystem I (PSI) functions to reduce the carbon dioxide in the reactions of the Calvin cycle. PSI also transfers the electrons to the primary electron acceptor A0 and also generated the powerful reductant P700 (Sage et al. 1990). Similarly, photosystem II reaction center protein M and L, uses the light energy to generate O2 and also play a major role in ATP formation.
Chloroplast envelope membrane proteins are involved in Na+-dependent proton extrusion and are associated with CO2 transport. It indirectly promotes efficient inorganic carbon uptake into chloroplasts. Some of these identified proteins further opens up new areas of investigation that would be helpful to understand the chloroplast metabolism (Sharkey and Seemann 1989). Gibberellin 20 oxidase 1A and 1B catalyze the biosynthesis of gibberellins. The synthesize gibberellins further catalyzes the conversion of GA12 and GA53 to GA9 and GA20, respectively. GA is an essential hormone for developments of plants including stem and root elongation, seed germination, and floral development etc. (Fleet and Sun 2005). The driving signals of DELLA degraded proteins bind to DELLA proteins tagged with ubiquitin for degradation by the 26S proteasome (Griffiths et al. 2006).
Chromatin remodelling and other protein
The decrease in the expression of various chromatin-associated proteins like nuclear ribonuclease Z Fragment, DNA directed RNA polymerase subunit alpha and beta, Maturase K, histone H2B4, H2B 6, H2B 3 and H 3, transcription factor HBP 1b c1 and c38, High mobility group (HMG1 2) like protein, Transcription factor HBP 1b was observed under salinity stress. EmBP-1 belongs to leucine zipper (bZIP) class of transcription factors and has been implicated in the mechanisms of abscisic acid (ABA)-mediated gene regulation and histone gene expression. DNA directed RNA polymerase is responsible for the polymerisation of ribonucleotides into a sequence complementary to the template DNA and catalyzes the transcription of DNA into RNA (Sweetser et al. 1987). The chloroplast maturase K gene (matK) codes for a maturase protein, however, high evolutionary rate of matK has made it usable in phylogenetic reconstructions at high taxonomic levels.
Wheat transcription factors HBP-1a and HBP-1b, binds to the wheat histone gene promoters, illustrating their role in the transcriptional regulation of wheat histone genes (Tabata et al. 1991). HBP-la and HBP-lb exhibit distinct DNA-binding properties: the former binds specifically to the hexamer motif of the H3 promoter, whereas the latter binds to both the H3 hexamer motif (Jakoby et al. 2002). The high mobility group (HMG) proteins are among the most abundant and ubiquitous non-histone proteins in the nucleus (Johns 1982) and subjected to multiple post-translational modifications such as methylation, phosphorylation, acetylation, ADP-ribosylation and glycosylation (Bustin 1999). These HMG proteins are involved in various cellular processes such as replication, transcription or nucleosome assembly (Bustin and Reeves 1996). Among the other proteins, protein RAFTIN 1B required for pollen development is synthesized in the tapetum, and transported at appropriate stages to the microspores.
Conclusion
Proteomic analysis of wheat plant under high salinity stress in the presence and absence of ACC deaminase bacteria has provided a new insight to identify the mechanism of induced systemic tolerance. Present work helped to explore the role of microbes producing ACC deaminase enzyme and their role in alleviation of stress responses. A number of differentially expressed proteins upregulated in the present study have been shown for their role in stress protection to plant against salt stress in previous studies. The majority of these proteins were related to protein metabolism, defense activation, stress protection, maintenance of photosynthesis carbohydrate and energy metabolism. These proteins might work in a synergistically way to enhance tolerance to salt-attack and keep plant growth on a mostly even keel. Some of the other identified protein in this study also could be a promising candidate for to confer stress tolerance. The present study enriched our understanding and paved the way of mechanisms of PGPR-mediated stress tolerance in plants under adverse conditions to promote the agriculture production.
References
Aghaei K, Ehsanpour AA, Komatsu S (2008) Proteome analysis of potato under salt stress. J Proteome Res 7:4858–4868
Almagro L, Ros LG, Belchi-Navarro S, Bru R, Barcelo AR, Pedreño M (2009) Class III peroxidases in plant defence reactions. J Exp Bot 60(2):377–390
Apel K, Hirt H (2006) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Basha SA, Sarma BK, Singh DP et al (2006) Differential methods of inoculation of plant growth-promoting rhizobacteria induce synthesis of phenylalanine ammonia-lyase and phenolic compounds differentially in chickpca. Folia Microbiol 51:463–468
Belimov AA, Hontzeas N, Safronova VI, Demchinskaya SV, Piluzza G, Bullitta S, Glick BR (2005) Cadmium-tolerant plant growth promoting rhizobacteria associated with the roots of Indian mustard (Brassica juncea L. Czern.). Soil Biol Biochem 37:241–250
Bellur DL, Woodson SA (2009) A minimized rRNA-binding site for ribosomal protein S4 and its implications for 30S assembly. Nucleic Acids Res 37:1886–1896
Bharti N, Pandey SS, Barnawal D, Patel VK, Kalra A (2016) Plant growth promoting rhizobacteria Dietzia natronolimnaea modulates the expression of stress responsive genes providing protection of wheat from salinity stress. Sci Rep 6:34768
Bouche N, Yellin A, Snedden WA, Fromm H (2005) Plant-specific calmodulin-binding proteins. Annu Rev Plant Biol 56:435–466
Buch-Pedersen MJ, Venema K, Serrano R, Palmgren MG (2000) Abolishment of proton pumping and accumulation in the E1P conformational state of a plant plasma membrane H+-ATPase by substitution of a conserved aspartyl residue in transmembrane segment 6. J Biol Chem 275(50):39167–39173
Bustin M (1999) Regulation of DNA-dependent activities by the functional motifs of the high-mobility-group chromosomal proteins. Mol Cell Biol 19:5237–5246
Bustin M, Reeves R (1996) High-mobility-group chromosomal proteins: architectural components that facilitate chromatin function. Prog Nucleic Acid Res Mol Biol 54:35–100
Dixon DP, Lapthorn A, Edwards R (2002) Plant glutathione transferases. Genome Biol 3:3004.1–3004.10
Dixon DP, Hawkins T, Hussey PJ, Edwards R (2009) Enzyme activities and subcellular localization of members of the Arabidopsis glutathione transferase superfamily. J Exp Bot 60:1207–1218
Emes MJ, Bowsher CG, Hedley C, Burrell MM, Scrase-Field ES, Tetlow IJ (2003) Starch synthesis and carbon partitioning in developing endosperm. J Exp Bot 54:569–575
FAO (2005) Global network on integrated soil management forsustainable use of salt-affected soils. Land and Plant Nutrition Management Services, Rome. http://www.fao.org/ag/agl/agll/spush
Flagella Z, Trono D, Pompa M, Di Fonzo N, Pastore D (2006) Seawater stress applied at germination affects mitochondrial function in durum wheat (Triticum durum) early seedlings. Funct Plant Biol 33:357–366
Fleet CM, Sun TP (2005) A DELLAcate balance: The role of gibberellin in plant morphogenesis. Curr Opin Plant Biol 8:77–85
Franco OL, Rigden DJ, Melo FR, Grossi-De-Sá MF (2002) Plant alpha-amylases inhibitors and their interaction with insect alpha-amylases. Eur J Biochem 269:397–412
Girish N, Umesha S (2005) Effect of plant growth promoting rhizobacteria on bacterial canker of tomato. Arch Phytopathol Plant Prot 38(3):235–243
Glick BR, Penrose DM, Li J (1998) A model for the lowering of plant ethylene concentrations by plant growth-promoting bacteria. J Theor Biol 190:63–68
Glick BR, Patten CL, Holguin G, Penrose DM (1999) Biochemical and genetic mechanisms used by plant growth promoting bacteria. Imperial College Press, London
Griffiths J, Murase K, Rieu I, Zentella R, Zhang ZL, Powers SJ, Gong F, Phillips AL, Hedden P, Sun TP, Thomas SG (2006) Genetic characterization and functional analysis of the GID1 gibberellin receptors in Arabidopsis. Plant Cell 18:3399–3414
Grichko VP, Glick BR (2001a) Amelioration of flooding stress by ACC deaminase-containing plant growth-promoting bacteria. Plant Physiol Biochem 39:11–17
Grichko VP, Glick BR (2001b) Flooding tolerance of transgenic tomato plants expressing the bacterial enzyme ACC deaminase controlled by the 35S, rolD or PRB-1b promoter. Plant Physiol Biochem 39:19–25
Guo W, Reigan P, Siegel D, Zirrolli J, Gustafson D, Ross D (2005) Formation of 17-allylamino-demethoxygeldanamycin (17-AAG) hydroquinone by NAD(P)H:quinone oxidoreductase 1: role of 17-AAG hydroquinone in heat shock protein 90 inhibition. Cancer Res 65:10006–10015
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Annu Rev Plant Physiol Plant Mol Biol 51:463–499
Hématy K, Cherk C, Somerville S (2009) Host-pathogen warfare at the plant cell wall. Curr Opin Plant Biol 12:406–413
Hincha DK, Zuther E, Hellwege EM, Heyer AG (2002) Specific effects of fructo- and gluco-oligosaccharides in the preservation of liposomes during drying. Glycobiology 12:103–110
Hohm T, Dermarsy E, Quan C, Petrolati LA, Preuten T, Vernoux T et al (2014) Plasma membrane H+-ATPase regulation is required for auxin gradient formation preceding phototropic growth. Mol Syst Boil 10:751
Huang G-T, Ma S-L, Bai L-P, Zhang L, Ma H, Jia P et al (2012) Signal transduction during cold, salt, and drought stresses in plants. Mol Biol Rep 39(2):969–987
Idriss EE, Makarewicz O, Farouk A, Rosner K, Greiner R, Bochow H et al (2002) Extracellular phytase activity of Bacillus amyloliquefaciens FZB45 contributes to its plant-growth-promoting effect. Microbiology 7:2097–2109
Israr D, Mustafa G, Khan KS, Shahzad M, Ahmad N, Masood S (2016) Interactive effects of phosphorus and Pseudomonas putida on chickpea (Cicer arietinum L.) growth, nutrient uptake, antioxidant enzymes and organic acids exudation. Plant Physiol Biochem 108:304–312
Jakoby M, Weisshaar B, Droge-Laser W, Vicente-Carbajosa J, Tiedemann J, Kroj T, Parcy F (2002) bZIP transcription factors in Arabidopsis. Trends Plant Sci 7:106–111
Johns EW (ed) (1982) The HMG Chromosomal Proteins. Academic Press, London
Klingenberg CP (2008) Morphological integration and developmental modularity. Annu Rev Ecol Evol Syst 39:115–132
Kochar M, Upadhyay A, Srivastava S (2011) Indole-3-aceticacid biosynthesis in the biocontrol strain Pseudomonas fluorescens Psd and plant growth regulation by hormone overexpression. Res Microbiol 162:426–435
Kuhlbrandt W (2004) Biology, structure and mechanism of P-type ATPases. Nature 5:282–295
Kwon YS, Lee DY, Rakwal R, Baek SB, Lee JH, Kwak YS et al (2016) Proteomic analyses of the interaction between the plant-growth promoting rhizobacterium Paenibacillus polymyxa E681 and Arabidopsis thaliana. Proteomics 16:122–135
Li J, Glick BR (2001) Transcriptional regulation of the Enterobacter cloacae UW4 1-aminocyclopropane-1-carboxylate (ACC) deaminase gene (acdS). Can J Microbiol 47:359–367
Lin Y, Du D, SiC ZQ, Li Z, Li P (2014) Potential biocontrol Bacillus sp. strains isolated by an improved method from vinegar waste compost exhibit antibiosis against fungal pathogens and promote growth of cucumbers. Biol Control 71:7–15
Luo H, Morsomme P, Boutry M (1999) The two major types of plant plasma membrane H+-ATPases show different enzymatic properties and confer differential pH sensitivity of yeast growth. Plant Physiol 119:627–634
Macfarlane GT, Steed H, Macfarlane S (2008) Bacterial metabolism and health-related effects of galacto- oligosaccharides and other prebiotics. J Appl Microbiol 104:305–344
Maroco JP, Rodrigues ML, Lopes C, Chaves MM (2002) Limitations to leaf photosynthesis in field-grown grapevine under drought—metabolic and modelling approaches. Funct Plant Biol 29:451–459
Mates L, Izsvak Z, Ivics Z (2007) Technology transfer from worms and flies to vertebrates: transposition-based genome manipulations and their future perspectives. Genome Biol 8(Suppl 1):S1
Matsuoka Y, Aghaei MJ, Abbasi MR, Totiaei A, Mozafari J, Ohta S (2008) Durum wheat cultivation associated with Aegilops tauschii in northern Iran. Genet Resour Crop Evol 55:861–868
Mayak S, Tirosh T, Glick BR (2004) Plant growth-promoting bacteria confer resistance in tomato plants to salt stress. Plant Physiol Biochem 42:565–572
McCormack E, Tsai YC, Braam J (2005) Handling calcium signalling: Arabidopsis CaMs and CMLs. Trends Plant Sci 10:383–389
McGrath JP, Jentsch S, Varshavsky A (1991) UBA 1: an essential yeast gene encoding ubiquitin-activating enzyme. Embo J 10:227–236
Munns R, James RA, Läuchli A (2006) Approaches to increasing the salt tolerance of wheat and other cereals. J Exp Bot 57:1025–1043
Munoz-Bertomeu J, Miedes E, Lorences EP (2013) Expression of xyloglucan endotransglucosylase/hydrolase (XTH) genes and XET activity in ethylene treated apple and tomato fruits. J Plant Physiol 170:1194–1201
Nishitani K (1997) The role of endoxyloglucan transferase in the organization of plant cell walls. In: Jeon KW (ed) International review of cytology—a survey of cell biology, vol 173. Elsevier Academic Press Inc, San Diego, pp 157–206
Niu DD, Liu HX, Jiang CH, Wang YP, Wang QY, Jin HL et al (2011) The plant growth-promoting rhizobacterium Bacillus cereus AR156 induces systemic resistance in Arabidopsis thaliana by simultaneously activating salicylate-and jasmonate/ethylene-dependent signalling pathways. Mol Plant Microbe Interact 24:533–542
Ohshima M, Itoh H, Matsuoka M, Murakami T, Ohashi Y (1990) Analysis of stress-induced or salicylic acid-induced expression of the pathogenesis-related 1a protein gene in transgenic tobacco. Plant Cell 2:95–106
Parry MAJ, Andralojc PJ, Khan S, Lea PJ, Keys AJ (2002) Rubisco activity: effects of drought stress. Ann Bot 89:833–839
Pedersen CNS, Axelsen KB, Harper JF, Palmgren MG (2012) Evolution of plant P-type ATPases. Front Plant Sci 3:31
Penrose DM, Glick BR (2003) Methods for isolating and characterizing ACC deaminase-containing plant growth-promoting rhizobacteria. Physiol Plant 118:10–15
Robert F, Brakier-Gingras L (2001) Ribosomal protein S7 from Escherichia coli uses the same determinants to binds 16S ribosomal RNA and its messenger RNA. Nucleic Acids Res 29:677–682
Roberts TH, and Hejgaard J (2007) Serpins in plants and green algae. Funct Integr Genomics 8:1–27
Roppolo D, Geldner N (2012) Membrane and walls: who is master, who is servant? Curr Opin Plant Biol 15:608–617
Roppolo D, Rybel BD, Tendon VD, Pfister A, Alassimone J, Vermeer JEM, Yamazaki M, Stierhof Y-D, Beeckman T, Geldner N (2011) A novel protein family mediates Casparian strip formation in the endodermis. Nature 473:380–383
Roppolo D, Boeckmann B, Pfister A, Boutet E, Rubio MC, DeÂnervaud-Tendon V et al (2014) Functional andevolutionary analysis of the Casparian strip membrane domain protein family. Plan Physiol 165(4):1709–1722
Ross D, Kepa JK, Winski SL, Beall HD, Anwar A, Siegel D (2000) NAD(P)H:quinone oxidoreductase 1 (NQO1): chemoprotection, bioactivation, gene regulation and genetic polymorphisms. Chem Biol Interact 129:77–97
Sage RF, Sharkey TD, Seemann JR (1990) Regulation of Ribulose-1,5-bisphosphate carboxylase activity in response to light intensity and CO2 in the C3 annuals Chenopodium album L. and Phaseolus vulgaris L. Plant Physiol 94:1735–1742
Saradadevi R, Palta JA, Siddique KHM (2017) ABA-mediated stomatal response in regulating water use during the development of terminal drought in wheat. Front Plant Sci 8:1251
Scheler C, Durner J, Astier J (2013) Nitric oxide and reactive oxygen species in plant biotic interactions. Curr Opin Plant Biol 16:534–539
Sharkey TD, Seemann JR (1989) Mild water stress effects on carbon-reduction-cycle intermediates, ribulose bisphosphate carboxylase activity, and spatial homogeneity of photosynthesis in intact leaves. Plant Physiol 89:1060–1065
Shoji T, Suzuki K, Abe T, Kaneko Y, Shi H, Zhu J-K et al (2006) Salt stress affects cortical microtubule organizationand helical growth in Arabidopsis. Plant Cell Physiol 47(8):1158–1168
Singh RP, Jha PN (2016) The multifarious PGPR Serratia marcescens augments induced systemic resistance and enhanced salinity tolerance of wheat (Triticum aestivum). PLoS ONE. https://doi.org/10.1371/journal.pone.0155026
Singh RP, Jha PN (2017) The draft genome sequence of the plant growth promoting rhizospheric bacterium Enterobacter cloacae SBP-8. Genom Data 12:81–83
Singh RP, Jha P, Jha PN (2015) The plant growth promoting bacterium Klebsiella sp. SBP-8 confers induced systemic tolerance in wheat (Triticum aestivum) under salt stress. J Plant Physiol 184:57–67
Singh VK, Singh AK, Singh PP, Kumar A (2018) Interaction of plant growth promoting bacteria with tomato under abiotic stress: a review. Agric Ecosyst Environ 267:129–140
Stearns J, Glick BR (2003) Transgenic plants with altered ethylene biosynthesis or perception. Biotechnol Adv 21:193–210
Sun Y, Cheng Z, Glick BR (2009) The presence of a 1-aminocyclopropane-1-carboxylate (ACC) deaminase deletion mutation alters the physiology of the endophytic plant growth-promoting bacterium Burkholderia phytofirmans. FEMS Microbiol Lett 296:131–136
Suzuki N, Rivero RM, Shulaev V, Blumwald E, Mittler R (2014) Abiotic and biotic stress combinations. New Phytol 203:32–43
Sweetser D, Nonet M, Young RA (1987) Prokaryotic and eukaryotic RNA polymerases have homologous core subunits. Proc Natl Acad Sci 84:1192–1196
Szymanski DB, Cosgrove DJ (2009) Dynamic coordination of cytoskeletal and cell wall systems during plant cell morphogenesis. Curr Biol 19:R800–R811
Tabata T, Nakayama T, Mikami K, Iwabuchi M (1991) HBP-1a and HBP-1b: Leucine zipper-type transcription factors of wheat. EMBO J 10:1459–1467
Tetlow IJ, Morell MK, Emes MJ (2004) Recent developments in understanding the regulation of starch metabolism in higher plants. J Exp Bot 55:2131–2145
Vensel WH, Tanaka CK, Cai N, Wong JH, Buchanan BB, Hurkman WJ (2005) Developmental changes in the metabolic protein profiles of wheat endosperm. Proteomics 5:1594–1611
Vieira Dos Santos C, Rey P (2006) Plant thioredoxins are key actors in oxidative stress response. Trends Plant Sci 11:329–334
Vivas A, Biro B, Nemeth T, Barea JM, Azcón R (2006) Nickel-tolerant Brevibacillus brevis and arbuscular mycorrhizal fungus can reduce metal acquisition and nickel toxicity effects in plant growing in nickel supplemented soil. Soil Biol Biochem 38(9):2694–2704
Wang SL, Wang CY, Huang CY (2008) Microbial reclamation of squid pen for the production of a novel extracellular serine protease by Lactobacillus paracasei subsp. paracasei TKU012. Bioresour Technol 99:3411–3417
Wang Q, Li F, Zhang X, Zhang Y, Hou Y, Zhang S et al (2011) Purification and characterization of a CkTLP protein from Cynan chumkomarovii seeds that confers antifungal activity. PLoS ONE 6(2):e16930
Xu Z, Zhang R, Wang D, Qiu M, Feng H, Zhang N et al (2014) Enhanced control of cucumber wilt diseaseby Bacillus amyloliquefaciens SQR9 by altering the regulation of its degup hosphorylation. Appl Environ Microbiol 80:2941–2950
Youssef SA, Tartoura KA, Abdelraouf GA (2016) Evaluation of Trichoderma harzianum and Serratia proteamaculans effect on disease suppression, stimulation of ROS-scavenging enzymes and improving the tomato grow infected by Rhizoctonia solani. Biol Control 100:79–86
Zhang Z, Fu R, Huber DJ, Rao J, Chang X, Hu M et al (2012) Expression of expansin gene (CDK-Exp3) and its modulation by exogenous gibberellic acid during ripening and softening of persimmon fruit. HortScience 47:378–381
Zhang Q, Sun J, Lu T, Zhang J, Wu C, Li L, He Z, Zhao Y, Liu X (2015) A rapid and sensitive LC-MS/MS method for evaluation of the absolute oral bioavailability of a novel c-Met tyrosine kinase inhibitor QBH-196 in rats. Biomed Chromatogr 29:1650–1656
Zhong YX, Chen JY, Feng HL, Kuang JF, Xiao R, Ou M et al (2008) Expansin and XET genes are differentially expressed during aril breakdown in harvested longan fruit. J Am Soc Hortic Sci 133:462–467
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
42485_2022_83_MOESM1_ESM.docx
Supplementary file1 Supplementary S1. The detail of methodology w.r.t to bacterial inoculation, plant growth and protein extraction (DOCX 17 KB)
42485_2022_83_MOESM2_ESM.xlsx
Supplementary file2 Supplementary S2. List of all the identified protein in treatment T-1, T-2, T-3 and T-4 (XLSX 60 KB)
42485_2022_83_MOESM3_ESM.xlsx
Supplementary file3 Supplementary S3. The differentially expressed proteins with their UNIPROT-ID common to tested four treatments (T-1, T-2, T-3, T-4) were used for hierarchical cluster analysis displaying differential expression. (The customized set of parameters is employed for the analysis, i.e. Low and high expression levels are shown with green and red colors respectively) (XLSX 25 KB)
Rights and permissions
About this article
Cite this article
Singh, R.P., Jha, P.N. Transposon mutagenesis of ACC deamination gene alters the proteomic analysis of wheat plant under non-saline and saline stress. J Proteins Proteom 13, 39–53 (2022). https://doi.org/10.1007/s42485-022-00083-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42485-022-00083-4