Abstract
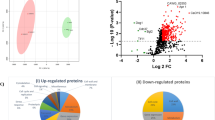
Chlamydospore though considered as a unique and rare morphological form of Candida albicans, regulation and significance of chlamydosporulation is not very clear. SWATH-MS analysis of chlamydosporulation specific proteins revealed that 319 (137-Up regulated and 182-Down regulated) proteins expressed differentially. Functional annotation showed significant modulations in proteins involved in cellular architecture (30), carbohydrate (29), amino acid (17), fatty acid (3), Nucleic acid (14), vitamins (1) metabolism as well as signaling (6), stress response (26), transport (cytoplasmic-21, mitochondrial-6 and nuclear-1), gene expression (transcription-12, RNA processing-6, translation-53, PTM-18), proteolysis (15) etc. Enhanced mannan, β1, 3-glucan and chitin contribute in thickening of cell wall while Hyr1 (218-fold) and Als3 (38.16-fold) dominates the cell surface chemistry of chlamydospores. In addition to ergosterol, enhanced sphingolipids, phospholipids and fatty acids make chlamydospore membrane more sturdy and rigid. Up-regulation of maltase (64-fold) followed by enhanced glycolysis and tricarboxylic acid cycle under nutrient-limiting condition is indicative of chlamydosporulation. Glyoxylate and fermentative pathway reported to facilitate survival of C. albicans under glucose limiting and microaerophilic condition was up-regulated. Enhanced biosynthesis of glutathione, trehalose homeostasis, and inhibition of NAD+ generation ,etc., potentiate oxidative, osmotic and nitrosative stress tolerance. Up regulation of Rsr1 (8.83-fold) and down regulation of Bcy1 (4.20-fold), Tfs1 (negative regulator of RAS) indicates cAMP-PKA pathway activates chlamydosporulation through Efg1 (a morphogenic regulator) in our study. In general, morpho-physiological modulations in C. albicans is a result of different sets of transcriptional programs that facilitate survival under nutrient and oxygen limiting condition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Candida albicans, a polymorphic opportunistic pathogen associated with superficial to life-threatening systemic infections among immunocompromised individuals is included in the list of organisms with potential antibiotic resistance threat, recently (CDC report 2013; Kullberg and Arendrup 2015). Morphogenic plasticity is considered as a survival strategy that enables C. albicans to colonize and invade host tissues by evading host defense mechanisms under a wide range of extreme micro-environments (Brown et al. 2014; Cutler 1991; Ernst 2000; Lim et al. 2012; Ruhnke 2006). Different morphological forms viz. yeast, hyphae, pseudohyphe, chlamydospore, opaque cells and biofilms exhibit differential responses towards host defense mechanisms as well as antifungal agents (Cutler 1991; Lim et al. 2012; Ruhnke 2006; Tyc et al. 2014). Among these, the hyphal form is a prerequisite for tissue invasion and invasive candidiasis and biofilms on indwelling medical devices are considered as difficult-to-treat infections with very high mortality (Mc Manus and Coleman 2014; Mun et al. 2016; Neville et al. 2015; Williams 2011). Considering the significance in virulence, these morphological and growth forms were studied exhaustively in recent years while chlamydospore, considered as a non-virulent form is neglected by scientific community (Bottcher et al. 2016; Citiulo et al. 2009).
Chlamydospores are thick-walled spherical cells (6–10 µm) considered as dormant form induced under unfavorable environmental conditions like oxygen limitation and embedded growth in matrix (Giosa et al. 2017; Nobile et al. 2003; Sonneborn et al.1999). The presence of chlamydospores in clinical specimens is reported but it may be unlikely to play any role in pathogenesis (Citiulo et al. 2009). Chlamydospores are more specialized and relatively rare morphological states formed to survive under harsh conditions (Nobile et al. 2003; Sosinska 2012). It could be an adaptive response towards exposure to reactive oxygen species of host cells or towards co-existing microorganisms (Berman and Sudbery 2002; Douglas et al. 2005). Chlamydospores are metabolically active, can germinate under favorable condition and produce daughter chlamydospores, blastospores, pseudohyphae and true hyphae (Citiulo et al. 2009; Staib and Morschhäuser 2007). Chlamydospores thick wall is providing protection against the adverse micro-environments. However, not much study is available on structure and composition of chlamydospore cell wall (Jansons and Nickerson 1970).
Though various environmental, nutritional and genetic factors are implicated, regulation and significance of chlamydospore formation are not very clear (Bottcher et al. 2016). In present study, we have made an attempt to identify chlamydospore specific proteins using LC–MS/MS analysis. This is the first attempt at identifying chlamydospore specific proteins. Proteomic analysis revealed morphophysiological modulations responsible for altering cellular architecture that enables C. albicans to survive under extreme micro-environments.
Material and methodology
Candida albicans strain and growth condition
Candida albicans ATCC 10231 a quality control strain was procured from Institute of Microbial Technology (IMTECH), Chandigarh, India and maintained on yeast extract peptone dextrose (YPD) agar at 4 °C. Rice extract agar with 1% Tween 80 was used for chlamydospore induction. Yeast Extract Peptone Dextrose (YEPD) broth and Rice extract agar were purchased from Hi-media Laboratories, Pvt. Ltd. Mumbai (India). All the chemicals used in this study were purchased from Sigma-Aldrich Pvt. Ltd., Bangalore (India) and solvents from Qualigens and SD Fine Chemicals Ltd, Mumbai (India).
Induction of chlamydospores
Inoculums were prepared using C. albicans cells grown for 48 h at 30 °C in YEPD broth. Cells were harvested by centrifugation at 448 g for 2 min, washed thrice with sterile distilled water and re-suspended in 1 ml of sterile distilled water (Holmes and Shepherd 1987; Odds 1988).
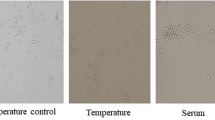
Chlamydosporulation was induced using rice extract agar (Rice extract 1.3 g, Tween 1%, pH 7.1). In brief, sterile rice extract with low melting agar plates were inoculated with 1 × 103 cells per plate (Kelly and Funigiello 1959). Plates were overlaid with sterile, polyethylene sheets of 7.5 cm diameter for anaerobiosis and invasive growth necessary for chlamydospores induction. Plates were wrapped with parafilm and incubated at 30 °C for 14 days (Miller et al. 1974). Chlamydospore formation was monitored by observing the plates under dissecting microscope. Chlamydospores were stained using Lactophenol cotton blue, observed microscopically (OLYMPUS CX21i) and photographed using microscope attached camera (OLYMPUS digital camera E-PL1) (Kim et al. 2002).
Harvesting of chlamydospores
Chlamydospores were harvested from 12-day-old plates grown at 30 °C. In brief, chlamydospores plates were washed with sterile distilled water to remove colonies of yeast phase cells growing on the agar surface. Rice extract agar containing chlamydospores was molten using warm (45 °C) sterile distilled water, centrifugation at 112 g for 1 min. Chlamydospore pellets were washed thrice with sterile distilled water and used for protein extraction. However, chlamydospores as harvested from semisolid agar, traces of hyphae producing chlamydospores were present along with chlamydospores in the sample. Cells from the colonies growing on the rice extract agar surface were harvested by centrifugation, washed with sterile distilled water and used as a control.
Cell surface hydrophobicity
Cell surface hydrophobicity of yeast phase cells and chlamydospores of C. albicans was analyzed using a method developed by Rosenberg, et al. (1980) and Hazen and Hazen (1987). Briefly, yeast cells and chlamydospores were re-suspended in PBS until the OD (at 620 nm) reaches to 0.5. 1.3 ml from each of these suspensions was distributed in three test tubes. 100 μl from each test tube were again added into the wells of 96 well micro titer plates and initial OD (at 620 nm) was recorded using Thermoscan-Ex micro plate reader (Thermo Fisher Scientific Inc., 168 3rd Ave, Waltham, MA 02451, USA). 0.3 ml of octane was added to remaining cell suspension (1.2 ml) separately, mixed vigorously (3 min), and allowed to separate for 15 min. From these suspensions 100 μl were cautiously transferred to the wells of 96-well micro-titer plates and final OD was recorded. Triplicates were used for each samples and experiment was repeated thrice. Percentage CSH was calculated using following formula and compared with control. Results were presented as percentage of CSH ± SD (standard deviation) (Hazen and Hazen 1987; Rosenberg et al. 1980).
Estimation of adhesion
Adhesion of yeast phase cells and chlamydospores of C. albicans was determined using method described by Panagoda et al. (2001) and He et al. (2006). Briefly, 100 μl of cell suspension (1 × 107 cells/chlamydospores/ml) was inoculated in 96 wells micro titer plates and incubated at 30 °C for 90 min with moderate shaking (50 rpm) on orbital shaker for adhesion. After incubation, wells were washed thrice with PBS to remove un-adhered ones and the numbers of adhered cells/chlamydospores in each well were counted, microscopically (Metzer Make inverted Microscope) and compared. The average number of yeast phase cells adhered is considered as 100% adhesion. Experiment was repeated thrice and triplicates were used for each sample and results were shown as adhesion percentage ± SD (standard deviation) (He et al. 2006; Panagoda et al. 2001).
Ergosterol extraction
Ergosterol content of yeast phase cells and chlamydospores of C. albicans was estimated as per Arthington-Skaggs et al. (1999). Briefly, 0.1 g of washed cell/chlamydospore pellets were suspended into 300 μl of ethanolic KOH (25%) and incubated at 85 °C for 1 h and 2 h, respectively. Samples were cooled to room temperature and ergosterol was extracted using n-heptane [75% (v/v)] with vortexing. Layers were allowed to separate and n-heptane layer was transferred to new vial, cautiously. 200 μl of n-heptane layer was diluted to fivefold in ethanol (100%) and spectrum was recorded in the wavelength range of 230–300 nm using a UV–Visible spectrophotometer (Shimadzu Analytical (India) Pvt. Ltd. Mumbai- 400 059, India).
Ergosterol estimation
Ergosterol content was determined using the values of absorbance at 230 nm and 281.5 nm and the formula by Arthington-Skaggs et al. (1999):
where F is the dilution factor in ethanol and 290 and 518 are the E values (percent/centimeter) determined for crystalline ergosterol and 24(28) DHE, respectively. Results were showed as ergosterol percentage ± SD (standard deviation).
Cell lysis and protein extraction
Proteins (chlamydospore and yeast phase cells) were extracted using the protocol optimized by Haar (2007). Briefly, chlamydospores and yeast phase cells (1x108 equivalent cells) were resuspended in 200 µl of freshly prepared lysis buffer (0.1 M NaOH, 0.5 M EDTA, 2% SDS and 2% β-mercaptoethanol) containing (10 µl/ml) PIC (protease inhibitor cocktail) and vortexed. Samples were incubated at 90 °C for 15 min and neutralized using 5 µl of 4 M acetic acid after incubation. Samples were further incubated at 90 °C for 15 min and centrifuged at 2800 g for 5 min. Supernatants were transferred to new vials containing 5 µl of PMSF (Phenyl methane sulphonyl fluoride). Proteins were precipitated using 4 volumes of methanol, 1 volume of chloroform and 3 volumes of sterile distilled water with vortexing. Precipitated proteins were centrifuged at 2800g for 5 min, pellets were washed using 3 volumes of methanol, centrifuged at 2800 g for 5 min and air dried. Air dried pellets were re-suspended in rehydration buffer (6 M urea, 2 M Thiourea, 2% CHAPS, 1% DTT, pH 8.75) (Haar 2007) and protein concentrations were determined as per Bradford method (Bradford 1976).
Sample preparation
Proteins (50 µg) were dissolved in ammonium bicarbonate buffer (50 mM) containing rapigest (0.1%). Proteins were reduced with 3 µl of DTT (100 mM) at 60° C for 15 min, alkylated using 3 µl of Idoacetamide (200 mM) at room temperature for 30 min, and digested at alkaline pH using 2 µg of trypsin per 50 µg of proteins (i.e. 2:50). Digestion was stopped by adding 2 µl of concentrated HCL after 18 h of digestion. Peptides were separated by spinning at high speed (1500 rcf) for 15 min at 4 °C, washed several times with 0.1% TFA, size fractionated (̴̴ 3 kDa) using Zip tip C18 chromatography columns (Millipore; Billerica, MA) and eluted in 100% Acetonitrile. Samples were reconstituted in 15 µl of Acetonitrile (3%) and formic acid (0.1%) with continuous vortexing and used for further analysis (Gillet et al. 2012).
Liquid chromatography and mass spectrometry analysis
Peptides (4 µg) were separated and mass was determined using Micro LC 200 (Eksigent; Dublin, CA) coupled with Triple- TOF 5600 (AB Sciex; Concord, Canada) mass spectrometer in high-sensitivity mode. Equal amounts of samples of chlamydospores and control were spiked to generate the SWATH (sequential window acquisition of all theoretical fragment ion spectra) spectral library of fragment ions and analyzed using Information Dependant acquisition (IDA) (Collins et al. 2013; Gillet et al. 2012; Liu et al. 2006, 2013).
SWATH MS analysis
Swath MS analysis was carried out using the instrument setting as described in Ingle et al. (2017), Collins et al. (2013); Gillet et al. (2012); Liu et al.(2006); Liu et al. (2013). The mass spectral data acquired in triplicates was searched against Candida databases, Uniprot ids were searched using Protein Pilot software and differentially expressed proteins were identified using markerview software. Subsequently SWATH-MS was performed for relative quantification of differentially expressed proteins as mentioned in Ingle et al. (2017).
Statistical analysis
The student t test and probability were performed for statistical analysis. Samples with probability (p) value ≤ 0.05, number of matching peptides ≥ 2 and fold change ≥ 2, were considered for further analysis.
Validation of proteomic data using real-time qPCR analysis of selected genes
Expression of selected genes during chlamydosporulation was evaluated at mRNA level using real-time qPCR analysis. Gene-specific primers were designed using primer3 plus software (Tm 58–60 °C, product size 120–150 bp preferred for primer pairs) (Table 5). Total RNAs were prepared using RNeasy Mini kit (50 reactions) (Cat. No. 74104, Qiagen Pvt. Ltd) by lysing Chlamydospores using lyticase and purified using RNA Sure Mini Kit, Nucleo-pore (Genetix) according to the manufacturer’s instructions. cDNAs were prepared using purified RNA (2 µg) as a template and High-Capacity cDNA Reverse Transcription Kit as per the manufacturer’s instructions (Green et al. 2004). RNA level was measured using KAPA SYBR® FAST qPCR Kit as per manufacturer’s instructions and parameters and CFX96 Touch TM Real-Time PCR Detection System (Biorad Pvt. Ltd). Samples were analyzed in triplicates using biological replicates and data are reported as mean ± SD. Using ANOVA, statistical significance was calculated and p-values less than 0.05 were considered significant. Gene expression was normalized with GAPDH levels and with control cells.
Results
Induction of chlamydosporulation
Embedded and microaerophilic growth on rice extract agar medium at 30 °C induced invasive growth in Candida albicans, initially. The invading hyphae later developed suspensor cells that produced thick-walled chlamydospores, terminally within 4 days of incubation. Further incubation up to 12 days leads to increase in the number of chlamydospores (Fig. 1).
Modulation in cell surface hydrophobicity (CSH), adhesion and ergosterol during chlamydosporulation
In present study, in response to diverse morphology cell surface hydrophobicity, adhesion as well as ergosterol significantly modulated. Cell surface hydrophobicity of C. albicans yeast phase cells and chlamydospores (with filaments) at 30° C were found (37.92 ± 4.98, 31.70 ± 3.64, 27.61 ± 3.12) and (9.12 ± 1.29, 11 ± 3.33, 10 ± 4.05) respectively (Fig. 2a). This showed that CSH of yeast phase cells more as compare to chlamydospores (with filaments). However, adhesion of yeast phase cells and chlamydospore (with filaments) at 30° were found (82 ± 3.61, 79.67 ± 4.51, 77.67 ± 3.79) and (2.33 ± 0.58, 3 ± 2, 2.33 ± 2.08), respectively (Fig. 2b). Yeast pahse cells are more adhesive than chlamydospores (with filaments). In addition to this, ergosterol content was analyzed through spectrophotomer, which showed that chlamydospore (with filaments) contain more ergosterol as compare to yeast phase cells (Fig. 2c). Percent ergosterol content of yeast phase cells and chlamydospores (with filaments) was found (0.0073 ± 0.00076, 0.0087 ± 0.0013, 0.0076 ± 0.001) and (0.124 ± 0.06, 0.0207 ± 0.002, 0.0196 ± 0.0018) respectively (Fig. 2c).
Identification of chlamydosporulation specific proteins using LC–MS/MS analysis
LC–MS/MS analysis identified 1177 proteins out of which, 319 were modulated significantly (Ingle et al. 2017). MS/MS data is submitted to Peptide Atlas and data set is publically available with the data set identifier PASS01061 at http://www.peptideatlas.org/PASS/PASS01061. Differentially expressed proteins were identified; functionally annotated and grouped into different categories according to their functions using databases like CGD, SGD, KEGG and Uniprot etc., using David software (Functional annotation Bioinformatics Microarray). Among these differentially expressed proteins, 137 were up-regulated while 182 were Down-regulated under chlamydospore inducing condition (Ingle et al. 2017). The number of up-regulated and down-regulated proteins in different biological processes viz. metabolism, cell wall and membrane composition, stress response and signaling, gene expression, transport etc. were shown in Tables 1, 2, 3 and 4, Fig. 2a, b.
Modulation of proteins involved in metabolism
A total of 108 proteins involved in metabolism were modulated significantly, out of which 51 were involved in carbohydrate metabolism and energy generation (Table 1). The maximum up regulation of 259.37-fold was observed in case of Pgk1 followed by 64.83 (CAWG_04860/Mal32), 35.99 (Ald4), 17.93 (CAWG_01786/Hxk2), 13.85 (LELG_01826/Icl1) fold while maximum down-regulation of 46.59-fold was observed in case of (CaO19.285/Ctn1) followed by 32.83 (Ams1) and 10.48 (Glo2) fold etc. (Table 1). Five proteins involved in fermentation were up-regulated and maximum up regulation of 12.10-fold was observed in case of Ife2 (Table 1).
Eleven out of the thirteen proteins involved in lipid, sterol and fatty acid biosynthesis were up-regulated and maximum up-regulation of 36.85 was observed in case of Scs7 (Sphingolipid alpha-hydroxylase), while two were down-regulated (Table 1). Seventeen proteins expressed differentially indicating significant modulation in amino acid metabolism, wherein Gcv1 was up-regulated by 10.66-fold while maximum down-regulation of 14.55-fold was observed in case of Gln1 (Table 1). Similarly, differential expression of fourteen proteins indicates significant modulation in nucleotide metabolism i.e. Ysa1 was down-regulated by 18.19-fold while Ura4 was up-regulated by 6.25-fold (Table 1). Heme (Hem13, Hem15) and Vitamin (Snz1) biosynthesis and fatty acid degradation were enhanced under chlamydospore inducing condition (Table 1).
Modulation of proteins involved in cell wall and membrane biosynthesis
A total of eighteen proteins involved in biosynthesis of cell wall components viz. mannan (2), beta 1, 3-glucan (1), beta 1, 6-glucan (2), chitin (1), cell wall proteins (7) and membrane (5) were modulated significantly during chlamydosporulation (Table 2). The cell wall proteins viz. Hyr1, Als3 (hyphae specific) and Ldg1 were up-regulated maximally by 218.24, 38.16 and 38.43-fold respectively while maximum down-regulation of 49.15-fold was observed in case of Kre9 involved in biosynthesis of beta 1,6-glucan (Table 2). Among the proteins involved in membrane structure and biosynthesis like Fmp52, Scs7, Ino1 etc., were up-regulated by 82.65 36.85 and 12.82-fold respectively (Table 2). It indicates that cell wall of chlamydospore is rich in mannan, chitin and beta 1, 3-glucan while membrane is enriched with sphingolipids, phospholipids, sterols etc.
Modulation of proteins involved in stress response and signal transduction
Twenty out of the twenty-six proteins modulated during chlamydospore growth were involved in oxidative stress response while four in heat stress and one each in metal ion and drug-induced stress (Table 2). Among these, ten were up-regulated while sixteen were down-regulated and maximum up-regulation of 242.48-fold was observed in case of Yhb1 (Nitric oxide dioxygenase), and down-regulation in case of Whs11 (176.48-fold) (Table 2).
Among the six proteins involved in signal transduction, maximum up-regulation of 8.83-fold and down-regulation of 16.7-fold was observed in case of Rsr1 (CLUG_ 03767) and Asr2 (CAWG_02167), respectively (Table 2).
Modulation of proteins involved in transport
Twenty-eight proteins involved in transport were significantly modulated (23 down-regulated, 5 up-regulated) during chlamydospore form growth (Table 2). Among the twenty-one proteins involved in cytoplasmic transport, six were up-regulated and fifteen were down-regulated (Table 2). Maximum up-regulation of 35.08-folds was observed in case of Vma5 (Putative vacuolar H (+)-ATPase) and down-regulation of 23.44-fold in Sbe22 (CaO19.10437), respectively (Table 2).
Modulation of proteins involved in regulation of cell cycle, cytoskeleton, genome organization, replication and repair
Among the sixteen proteins modulated, two, six, five and three were involved in cell cycle, cytoskeleton, genome organization and replication and repair, respectively (Table 3). One of the two involved in cell cycle regulation, four out of the six in cytoskeleton, all the five in genome organization and one of the three involved in replication and repair were up-regulated respectively (Table 3). Among these, maximum up- regulation of 8.15-fold and down-regulation of 12.5-fold was observed in case of Arp7 (genome organization) and Ssp2 (cell cycle) respectively (Table 3).
Modulation of proteins involved in genome function (transcription, RNA processing, translation and proteolysis)
Twelve proteins involved in transcription, six in RNA processing, 71 in translation and fifteen in proteolysis were significantly modulated under chlamydospore form growth (Table 3). Among the twelve proteins involved, eight were involved in transcription initiation and four in elongation (Table 3). All the eight involved in transcription initiation were down-regulated while three of the five involved in elongation were unregulated (Table 3). Similarly, six proteins involved in RNA processing were down-regulated (Table 3).
Out of the seventy-one proteins involved, thirty-eight (21 up-regulated, 17 down-regulated) were involved in ribosomal assembly, nine in translation initiation (2 up-regulated, 7 down-regulated), three in elongation (2 up-regulated, 1 down-regulated), fifteen in post translational modification (2 up-regulated, 13 down-regulated) and six were involved in mitochondrial ribosome assembly (3) and mitochondrial PTM (3) (Table 3). Maximum up-regulation of 16.23-fold and down-regulation of 16.35-fold was observed in case of Rbp4 and Rtg1 respectively (Table 3). On the other hand, Snu13 (CAWG_02750) was down-regulated maximally by 7.24-fold amongst the six down-regulated proteins involved in RNA processing (Table 3). Similarly amongst the 71 proteins involved in translation and post-translational modifications, maximum up regulation of 11.15-fold observed in case of Rps23 while Tif3 was down-regulated maximally i.e. 24.28-fold (Table 3). Among the fifteen proteins involved in proteolysis (2 up-regulated and 13 down-regulated), maximum up-regulation of 66.85-fold and down-regulation of 84.5-fold was observed in case of Pre7 (AGOS_AGL324 W) and Pbi2 (CaO19.10285), respectively (Table 3). Thirteen modulated proteins were uncharacterized (Table 4).
Discussion
Chlamydospore, the thick-walled resting spores induced in response to stresses (like the nutrient limitation, microaerophilic and osmotic stress etc.), is considered as a survival strategy of C. albicans (Citiulo et al. 2009; Staib and Morschhäuser 2005; Nobile et al. 2003). C. albicans cells experience most of these stresses (except temperature 37 °C) during growth on rice extract agar containing Tween 80 in our study, citing the biological significance of our data (Berman and Sudbery 2002; Douglas et al. 2005). Though it was found in some of the tissue samples recently, chlamydospore is considered as a non-virulent form of C. albicans and thus neglected by the scientific community (Palige et al. 2013; Citiulo et al. 2009; Lim et al. 2012). However, few recent studies have provided some insights into the regulation of chlamydospore formation and survival strategies (Bottcher et al. 2016; Giosa et al. 2017; Palige et al. 2013; Lim et al. 2012). Mutant analysis by Navarathna et al. (2016) revealed significance of a component of chromatin remodeling complex (ISW2) in chlamydosporulation (Navarathna et al. 2016). Our study showed that nutrient limiting and microaerophilic condition enhances activity of Ras related protein Rsr1, the upstream regulator while down regulate Bcy1, a negative regulator of cAMP-PKA pathway (Table 2) (Biswas et al. 2007). It suggests that chlamydospore formation is induced through cAMP-PKA pathway (Table 2) (Biswas et al. 2007). Earlier studies have shown that nutrient starvation activate cAMP-PKA pathway through Ras1 and induce chlamydospore formation through EFG1 in presence of minimal cAMP i.e. surplus cAMP is inhibitory (Bottcher et al. 2016). It was further supported by down regulation of adenyl cyclases (to avoid surplus cAMP concentration) and Tfs1, a negative regulator of Ras protein in our study (Bottcher et al. 2016). It confirms that involvement of cAMP-PKA pathway is essential for chlamydospore formation and thus osmotic, heat shock and cell wall integrity damage induced stress tolerance (Bottcher et al. 2016; Chautard et al. 2004; Harcus et al. 2004). However, Hog1 mitogen-activated protein (MAP) kinase and quorum sensing molecule farnesol were also implicated in chlamydosporulation (Eisman et al. 2006; Martin et al. 2005). Chlamydosporulation need reprogramming of the transcriptional program through chromatin remodeling complexes (ISW2) that leads to chlamydospore specific transcripts including components of cytoskeleton (actin, tubulin and cofilin) (Cao et al. 2005; Navarathna et al. 2016). Components of cytoskeleton (actin, tubulin and cofilin) and chromatin remodeling complex (Arp7) were up regulated in our study. Similarly, enhanced levels of components (Yke2 and Cct2) involved in proper folding of alpha, beta tubulin and actin indicate increased strain on PTM machinery, as reported earlier in response to GlcNAc (García-Sánchez et al. 2005; Kamthan et al. 2012). Enhanced Phb12 level in our study contributes in stabilizing newly synthesized proteins (CGD 2010; Schleit et al. 2013). Up-regulation of Pre7 (66.85-fold) a proteasome component indicates that ubiquitin-dependent proteasome activity was enhanced during chlamydospore growth (Table 3) (CGD 2010; Liu et al. 2016; Verma et al. 2000). Up-regulation of Tpd3 in our study confirms enhanced cytokinesis) during chlamydosporulation as reported earlier (Liu et al. 2016; Lorenz et al. 2004; Sarkar et al. 2002).
Nutrient limitation, microaerophilic and osmotic stress modulate metabolism during chlamydospore growth that leads to acquire unique morphological and architectural characteristics in addition to survival using nonconventional and complex carbohydrates (viz. maltose, xylitol etc.) (Fig. 3) (Jamai et al. 2007; Bruno et al. 2006). Significant up-regulation of maltase (64-fold) followed by enhanced glycolysis and tricarboxylic acid cycle in our study confirms earlier hypothesis that states, “Enhanced glycolysis and tricarboxylic acid cycle under glucose limiting condition contribute to link the regulation of chlamydospores production in Candida” (Table 1 and Fig. 3) (Brown et al. 2014; Bottcher et al. 2016; CGD 2010; Han et al. 2011). Enhanced glyoxylate cycle reported to facilitate survival of C. albicans under glucose limiting condition is enhanced in our study (Fig. 3) (Barelle et al. 2006; Ene et al. 2012). Microaerophilic condition triggers alcohol production leading to hyphae induction required for chlamydospore formation (Fig. 3) (Chauhan et al. 2011; Smith et al. 2004). In addition to energy generation, glycolysis provides precursors for the biosynthesis of lipids (storage molecules) and trehalose required for stress (oxidative, heat, desiccation, osmotic etc.) tolerance (Fig. 3) (Brown et al. 2014; Pereira et al. 2001; Yoda et al.2000). Inhibition of NAD+ synthesis increases oxidative stress tolerance and extend lifespan as NAD is an essential cofactor for cellular redox reactions (Table 1) (Pereira et al. 2001; Bedalov et al. 2003; Kato and Lin 2014). Enhanced glutathione biosynthesis, up-regulation of Osm1, Yhb1 and down-regulation of Ysa1 could be the compensatory responses to enhanced oxidative, nitrossative and osmotic stresses during chlamydospore growth (Table 2) (Chen et al. 2008; Cottier et al. 2012; Liu et al. 2000; Michán and Pueyo 2009; Nett et al. 2009; Yadav et al. 2011). Enhanced level of Yhb1 was also validated at RNA level using qRT-PCR analysis (Tables 2, 5 and Figs. 4, 5). Chlamydospore inducing condition further potentiates antioxidant machinery through the maturation of Sod1 and trehalose homeostasis (through up-regulation of Lys7 and Hsp21 respectively) (Dong et al. 2013; Gleason et al. 2014; Mayer et al. 2012). Morpho-physiological and cellular architectural modulations seems to affect cellular and mitochondrial membrane functions like transport and osmotic stability as proteins involved in maintaining these functions like Gdi1, Cof1, Por1 were up-regulated (Cederquist et al. 2012; Curwin et al. 2012; Kamthan et al. 2012; Lin et al. 2010; Cao et al. 2005). However vacuolar transport and vacuolar (H)-ATPase activities (viz. endocytosis, tagging of lysosomal enzymes) were modulated in our study (Cabezon et al. 2009; Cabrera et al. 2013; Johnston et al. 2013).
Metabolic modulation enhances biosynthesis of components of cell wall and membrane during chlamydospore growth. The thick cell wall is reported to provide protection against adverse microenvironment, however, structure and composition of cell wall of chlamydospore is not very clear (Hazel and Williams 1990; Citiulo et al. 2009; Jansons and Nickerson 1970). Different environmental stresses/factors reported to modulate cell surface chemistry viz. cell surface hydrophobicity (CSH) and adhesion, the two important virulence factors facilitating survival of C. albicans (Odds and Bernaerts 1994). These morphophysiological modulations are results of modulation in central metabolic pathway that leads to modulations in cellular architecture and thus cell surface properties (Ingle et al. 2017). Cell surface properties are defined by cell surface chemistry, cell surface hydrophobicity (CSH) and adhesion etc. (Brown et al. 2014; Hazen et al. 1986; Hazen 1990; Klotz and Penn 1987). Cell surface properties are extremely important in survival and virulence of C. albicans (Odds and Bernaerts 1994). CSH determines host-Candida cell interaction, i.e. adhesion and colonization followed by tissue invasion at different tissue sites with varied microenvironments (Ener and Douglas 1992). In general, adhesion increases with increase in CSH i.e. CSH and adhesion is directly proportional (Ener and Douglas 1992; Klotz et al. 1985). More numbers of adhesins in C. albicans compared to non-virulent yeast S. cerevisiae cites the importance of cell surface molecules in pathogenicity (Guthrie and Fink 2002). Csh1 (Cell surface hydrophobicity protein 1) was down regulated in our study (Table 5). Down regulation of Csh1 during chlamydosporulation correlates positively with adhesion (Klotz 1990; Samaranayake et al. 2003). Decreased cell surface hydrophobicity is indicative of decreased virulence in chlamydospores (Fig. 2a) (Guthrie and Fink 2002; Samaranayake et al. 2003).
Our result suggests that enhanced mannan, β1, 3-glucan and chitin biosynthesis thicken and strengthen the cell wall of chlamydospore (Table 2) (CGD 2010). Increased components viz. β-glucan (β 1, 3-glucan) and chitin are reported to strengthen cell wall under hypo-osmotic and hypoxic stress (Table 2) (Ene et al. 2012; Hall 2015; Smits et al. 2001). Interestingly, down regulation of Kre9 (at protein and RNA level) and Phr2 indicates lack of β 1,6-glucan leading to abnormal cross-linking, however enhanced chitin content could be a compensatory response to maintain cell wall strength (Tables 2, 5 and Figs. 3, 4, 5) (Aimanianda et al. 2009; Lee et al. 2012; Smits et al. 2001). Over expression of two hyphae specific GPI (glycosylphosphatidylinositol) anchored proteins (Als3, Hyr1) involved in adhesion, indicates that sample exhibit hyphae producing chlamydospores and those were highly adhesive (Table 2, Fig. 5) (Heilmann et al. 2011; Klis et al. 2009; Richard et al. 2002). However, reduced hydrophobicity (down-regulation of Csh1) as well as down regulation of potential virulence factors (Pst3 and Ttr1) known to affect redox state of target proteins of host cells confirms attenuated virulence in chlamydospores (Table 2) (Collinson et al. 2002; Karababa et al. 2004; Seneviratne et al. 2008). Enhanced biosynthesis confirms that lipids (ergosterol, sphingolipid, phospholipids and fatty acids) are the storage molecules of chlamydospores in addition to providing more strength and rigidity to membranes (Table 2) (Fu et al. 2012; Walther et al. 2006; Young et al. 2002).
Our proteomic data confirms that nutrient limiting and microaerophilic microenvironment is sensed by C. albicans through RAS mediated cAMP-PKA pathway that modulate metabolism to use complex carbohydrates like maltose (Fig. 6). Enhanced degradation of complex carbohydrates releases simple sugars and enhances glycolysis and TCA cycle, significantly. In addition to energy, enhanced glycolysis and TCA cycle provides precursors for biosynthesis of lipids and glycogen (energy reserve), sterol and fatty acids (strengthen membrane), chitin, mannan, Beta, 1–3, glucan (strengthen cell wall architecture) and glutathione and trehalose (potentiate stress tolerance) (Fig. 6). In general, we conclude that chlamydosporulation confer tolerance towards hostile microenvironment by strengthening cellular architecture and stress responses through modulations in metabolic pathways and thus survival of C. albicans.
References
Aimanianda V, Clavaud C, Simenel C, Fontaine T, Delepierre M, Latgé JP (2009) Cell wall β-(1,6)-glucan of Saccharomyces cerevisiae structural characterization and in situ synthesis. J Biol Chem 284:13401–13412
Arthington-Skaggs BA, Jradi H, Desai T, Morrison CJ (1999) Quantitation of ergosterol content: novel method for determination of fluconazole susceptibility of Candida albicans. J Clin Microbiol 37:3332–3337
Barelle CJ, Priest CL, MacCallum DM, Gow NA, Odds FC, Brown AJ (2006) Niche-specific regulation of central metabolic pathways in a fungal pathogen. Cell Microb 8:961–971. https://doi.org/10.1111/j.1462-5822.2005.00676.x
Bedalov A, Hirao M, Posakony J, Nelson M, Simon JA (2003) NAD+ -dependent deacetylase Hst1p controls biosynthesis and cellular NAD+ levels in Saccharomyces cerevisiae. Mole Cell Biol 23:7044–7054. https://doi.org/10.1128/MCB.23.19.7044-7054.2003
Berman J, Sudbery PE (2002) Candida albicans: a molecular revolution built on lessons from budding yeast. Nat Rev Genet 3:918–932. https://doi.org/10.1038/nrg948
Biswas S, Van Dijck P, Datta A (2007) Environmental sensing and signal transduction pathways regulating morphopathogenic determinants of Candida albicans. Microbiol Mol Biol Rev 71:348–376. https://doi.org/10.1128/MMBR.00009-06
Bottcher B, Pöllath C, Staib P, Hube B, Brunke S (2016) Candida species rewired hyphae developmental programs for chlamydospore formation. Front Microb 7:1697. https://doi.org/10.3389/fmicb.2016.01697
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Brown AJ, Budge S, Kaloriti D, Tillmann A, Jacobsen MD, Yin Z, Ene IV, Bohovych I, Sandai D, Kastora S, Potrykus J (2014) Stress adaptation in a pathogenic fungus. J Ex Biol 217:144–155. https://doi.org/10.1242/jeb.088930
Bruno VM, Kalachikov S, Subaran R, Nobile CJ, Kyratsous C, Mitchell AP (2006) Control of the C. albicans cell wall damage response by transcriptional regulator Cas5. PLoS Pathog 2:e21. https://doi.org/10.1371/journal.ppat.0020021
Cabezon V, Llama-Palacios A, Nombela C, Monteoliva L, Gil C (2009) Analysis of Candida albicans plasma membrane proteome. Proteomics 9:4770–4786. https://doi.org/10.1002/pmic.200800988
Cabrera M, Arlt H, Epp N, Lachmann J, Griffith J, Perz A, Reggiori F, Ungermann C (2013) Functional separation of endosomal fusion factors and the class C core vacuole/endosome tethering (CORVET) complex in endosome biogenesis. J Biol Chem 288:5166–5175. https://doi.org/10.1074/jbc.M112.431536
Cao YY, Cao YB, Xu Z, Ying K, Li Y, Xie Y, Jiang YY (2005) cDNA microarray analysis of differential gene expression in Candida albicans biofilm exposed to farnesol. Antimicrob Agents Chemother 49:584–589. https://doi.org/10.1128/AAC.49.2.584-589.2005
CDC Report (2013) Antibiotic resistance threats in the United States
Cederquist GY, Luchniak A, Tischfield MA, Peeva M, Song Y, Menezes MP, Chan WM, Andrews C, Chew S, Jamieson RV, Gomes L (2012) An inherited TUBB2B mutation alters a kinesin-binding site and causes polymicrogyria, CFEOM and axon dysinnervation. Hum Mol Genet 21:5484–5499. https://doi.org/10.1093/hmg/dds393
CGD (2010) Description lines for gene products based on orthologs and predicted Gene Ontology (GO)
Chauhan NM, Raut JS, Karuppayil SM (2011) A morphogenetic regulatory role for ethyl alcohol in Candida albicans. Mycoses 54:6. https://doi.org/10.1111/j.1439-0507.2010.02002.x
Chautard H, Jacquet M, Schoentgen F, Bureaud N, Bénédetti H (2004) Tfs1p, a member of the PEBP family, inhibits the Ira2p but not the Ira1p Ras GTPase-activating protein in Saccharomyces cerevisiae. Eukaryot Cell 3:459–470. https://doi.org/10.1128/EC.3.2.459-470.2004
Chen YL, Kauffman S, Reynolds TB (2008) Candida albicans uses multiple mechanisms to acquire the essential metabolite inositol during infection. Infect Immun 76:2793–2801. https://doi.org/10.1128/IAI.01514-07
Citiulo F, Moran GP, Coleman DC, Sullivan DJ (2009) Purification and germination of Candida albicans and Candida dubliniensis chlamydospores cultured in liquid media. FEMS Yeast Res 9:1051–1060. https://doi.org/10.1111/j.1567-1364.2009.00533.x
Collins BC, Gillet LC, Rosenberger G, Röst HL, Vichalkovski A, Gstaiger M, Aebersold R (2013) Quantifying protein interaction dynamics by SWATH mass spectrometry: application to the 14-3-3 system. Nat Methods 10:1246–1253. https://doi.org/10.1038/nmeth.2703
Collinson EJ, Wheeler GL, Garrido EO, Avery AM, Avery SV, Grant CM (2002) The yeast glutaredoxins are active as glutathione peroxidases. J Biol Chem 277:16712–16717. https://doi.org/10.1128/AAC.48.8.3064-3079.2004
Cottier F, Raymond M, Kurzai O, Bolstad M, Leewattanapasuk W, Jiménez-López C, Lorenz MC, Sanglard D, Váchová L, Pavelka N, Palková Z (2012) The bZIP transcription factor Rca1p is a central regulator of a novel CO2 sensing pathway in yeast. PLoS Pathog 8:e1002485. https://doi.org/10.1371/journal.ppat.1002485
Curwin AJ, von Blume J, Malhotra V (2012) Cofilin-mediated sorting and export of specific cargo from the Golgi apparatus in yeast. Mol Biol Cell 23:2327–2338. https://doi.org/10.1091/mbc.e11-09-0826
Cutler JE (1991) Putative virulence factors of Candida albicans. Annu Rev Microbiol 45:187–218
Dong K, Addinall SG, Lydall D, Rutherford JC (2013) The yeast copper response is regulated by DNA damage. Mol Cell Biol 33:4041–4050. https://doi.org/10.1128/MCB.00116-13
Douglas LM, Alvarez FJ, McCreary C, Konopka JB (2005) Septin function in yeast model systems and pathogenic fungi. Eukaryot Cell 4:1503–1512. https://doi.org/10.1128/EC.4.9.1503-1512.2005
Eisman B, Alonso-Monge R, Roman E, Arana D, Nombela C, Pla J (2006) The Cek1 and Hog1 mitogen-activated protein kinases play complementary roles in cell wall biogenesis and chlamydospore formation in the fungal pathogen Candida albicans. Eukaryot Cell 5:347–358. https://doi.org/10.1128/EC.5.2.347-358.2006
Ene IV, Adya AK, Wehmeier S, Brand AC, MacCallum DM, Gow NA, Brown AJ (2012) Host carbon sources modulate cell wall architecture, drug resistance and virulence in a fungal pathogen. Cell Microbiol 14:1319–1335. https://doi.org/10.1111/j.1462-5822.2012.01813.x
Ener B, Douglas LJ (1992) Correlation between cell-surface hydrophobicity of Candida albicans and adhesion to buccal epithelial cells. FEMS Microbiol Lett 99(1):37–42. https://doi.org/10.1111/j.1574-6968.1992.tb05538.x
Ernst JF (2000) Transcription factors in Candida albicans–environmental control of morphogenesis. Microbiology 146:1763–1774. https://doi.org/10.1099/00221287-146-8-1763
Fu MS, De Sordi L, Mühlschlegel FA (2012) Functional characterization of the small heat shock protein Hsp12p from Candida albicans. PLoS ONE 7:e42894. https://doi.org/10.1371/journal.pone.0042894
García-Sánchez S, Mavor AL, Russell CL, Argimon S, Dennison P, Enjalbert B, Brown AJ (2005) Global roles of Ssn6 in Tup1-and Nrg1-dependent gene regulation in the fungal pathogen, Candida albicans. Mol Biol Cell 16:2913–2925. https://doi.org/10.1091/mbc.e05-01-0071
Gillet LC, Navarro P, Tate S, Röst H, Selevsek N, Reiter L, Bonner R, Aebersold R (2012) Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteom 11:O111–016717. https://doi.org/10.1074/mcp.O111.016717
Giosa D, Felice MR, Lawrence TJ, Gulati M, Scordino F, Giuffrè L, Lo Passo C, D’Alessandro E, Criseo G, Ardell DH, Hernday AD (2017) Whole RNA-sequencing and transcriptome assembly of Candida albicans and Candida africana under chlamydospore-inducing conditions. Genome Biol Evol 9:1971–1977. https://doi.org/10.1093/gbe/evx143
Gleason JE, Li CX, Odeh HM, Culotta VC (2014) Species-specific activation of Cu/Zn SOD by its CCS copper chaperone in the pathogenic yeast Candida albicans. J Biol Inorg Chem 19:595–603. https://doi.org/10.1007/s00775-013-1045-x
Green CB, Cheng G, Chandra J, Mukherjee P, Ghannoum MA, Hoyer LL (2004) RT-PCR detection of Candida albicans ALS gene expression in the reconstituted human epithelium (RHE) model of oral candidiasis and in model biofilms. Microbiology 150:267–275. https://doi.org/10.1099/mic.0.26699-0
Guthrie C, Fink GR (eds) (2002) Guide to yeast genetics and molecular and cell biology: part C. Gulf Professional Publishing, Houston
Haar VDT (2007) Optimized protein extraction for quantitative proteomics of yeasts. PLoS ONE 2:e1078. https://doi.org/10.1371/journal.pone.0001078
Hall RA (2015) Dressed to impress: impact of environmental adaptation on the Candida albicans cell wall. Mol Microbiol 97:7–17. https://doi.org/10.1111/mmi.13020
Han TL, Cannon RD, Villas-Bôas SG (2011) The metabolic basis of Candida albicans morphogenesis and quorum sensing. Fungal Genet Biol 48:747–763. https://doi.org/10.1016/j.fgb.2011.04.002
Harcus D, Nantel A, Marcil A, Rigby T, Whiteway M (2004) Transcription profiling of cyclic AMP signaling in Candida albicans. Mol Biol Cell 15:4490–4499. https://doi.org/10.1091/mbc.e04-02-0144
Hazel JR, Williams EE (1990) The role of alterations in membrane lipid composition in enabling physiological adaptation of organisms to their physical environment. Prog Lipid Res 29:167–227. https://doi.org/10.1016/0163-7827(90)90002-3
Hazen KC (1990) Cell surface hydrophobicity of medically important fungi, especially Candida species. In Microbial Cell Surface Hydrophobicity. ASM Press, Washington, DC, USA, pp 249–295
Hazen KC, Hazen BW (1987) A polystyrene microsphere assay for detecting surface hydrophobicity variations within Candida albicans populations. J Microbiol Methods 6:289–299. https://doi.org/10.1016/0167-7012(87)90066-2
Hazen KC, Plotkin BJ, Klimas DM (1986) Influence of growth conditions on cell surface hydrophobicity of Candida albicans and Candida glabrata. Infect Immun 54:269–271
He XY, Meurman JH, Kari K, Rautemaa R, Samaranayake LP (2006) In vitro adhesion of Candida species to denture base materials. Mycoses 49:80–84
Heilmann CJ, Sorgo AG, Siliakus AR, Dekker HL, Brul S, de Koster CG, de Koning LJ, Klis FM (2011) Hyphal induction in the human fungal pathogen Candida albicans reveals a characteristic wall protein profile. Microbiology 157:2297–2307. https://doi.org/10.1099/mic.0.049395-0
Holmes AR, Shepherd MG (1987) Proline-induced germ-tube formation in Candida albicans: role of proline uptake and nitrogen metabolism. Microbiology 133:3219–3228. https://doi.org/10.1099/00221287-133-11-3219
Ingle S, Kodgire S, Shiradhone A, Patil R, Zore G (2017) Chlamydospore specific proteins of Candida albicans. Data 2:26. https://doi.org/10.3390/data2030026
Jamai L, Ettayebi K, El Yamani J, Ettayebi M (2007) Production of ethanol from starch by free and immobilized Candida tropicalis in the presence of α-amylase. Bioresour Technol 98:2765–2770. https://doi.org/10.1016/j.biortech.2006.09.057
Jansons VK, Nickerson WJ (1970) Chemical composition of chlamydospores of Candida albicans. J Bacteriol 104:922–932
Johnston DA, Tapia AL, Eberle KE, Palmer GE (2013) Three prevacuolar compartment Rab GTPases impact Candida albicans hyphal growth. Eukaryot Cell 12:1039–1050. https://doi.org/10.1128/EC.00359-12
Kamthan M, Mukhopadhyay G, Chakraborty N, Chakraborty S, Datta A (2012) Quantitative proteomics and metabolomics approaches to demonstrate N-acetyl-D-glucosamine inducible amino acid deprivation response as morphological switch in Candida albicans. Fungal Genet Biol 49:369–378. https://doi.org/10.1016/j.fgb.2012.02.006
Karababa M, Coste AT, Rognon B, Bille J, Sanglard D (2004) Comparison of gene expression profiles of Candida albicans azole-resistant clinical isolates and laboratory strains exposed to drugs inducing multidrug transporters. Antimicrob Agents Chemother 48:3064–3079. https://doi.org/10.1128/AAC.48.8.3064-3079.2004
Kato M, Lin SJ (2014) YCL047C/POF1 is a novel nicotinamide mononucleotide adenylyltransferase (NMNAT) in Saccharomyces cerevisiae. J Biol Chem 289:15577–15587. https://doi.org/10.1074/jbc.M114.558643
Kelly JP, Funigiello F (1959) Candida albicans: a study of media designed to promote chlamydospore production. J Lab C Med 53:807–809
Kim D, Shin WS, Lee KH, Kim K, Young Park J, Koh CM (2002) Rapid differentiation of Candida albicans from other Candida species using its unique germ tube formation at 39 C. Yeast 19(11):957–962. https://doi.org/10.1002/yea.891
Klis FM, Sosinska GJ, De Groot PW, Brul S (2009) Covalently linked cell wall proteins of Candida albicans and their role in fitness and virulence. FEMS Yeast Res 9:1013–1028. https://doi.org/10.1111/j.1567-1364.2009.00541.x
Klotz SA (1990) Role of hydrophobic interactions in microbial adhesion to plastic used in medical devices. Micro Cell Surf Hydrophobicity 12:107–136
Klotz SA, Penn RL (1987) Multiple mechanisms may contribute to the adherence of Candida yeasts to living cells. Curr Microbiol 16(3):119–122. https://doi.org/10.1007/bf01568389
Klotz SA, Drutz DJ, Zajic JE (1985) Factors governing adherence of Candida species to plastic surfaces. Infect Immun 50(1):97–101
Kullberg BJ, Arendrup MC (2015) Invasive candidiasis. N Engl J Med 373:1445–1456. https://doi.org/10.1056/NEJMra1315399
Lee KK, MacCallum DM, Jacobsen MD, Walker LA, Odds FC, Gow NAR, Munro CA (2012) Elevated cell wall chitin in Candida albicans confers echinocandin resistance in vivo. Antimicrob Agents Chemother 56:208–217. https://doi.org/10.1128/AAC.00683-11
Lim CY, Rosli R, Seow HF, Chong PP (2012) Candida and invasive candidiasis: back to basics. Eur J Clin Microbiol Infect Dis 31:21–31. https://doi.org/10.1007/s10096-011-1273-3
Lin MC, Galletta BJ, Sept D, Cooper JA (2010) Overlapping and distinct functions for cofilin, coronin and Aip1 in actin dynamics in vivo. J Cell Sci 123:1329–1342. https://doi.org/10.1242/jcs.065698
Liu L, Zeng M, Hausladen A, Heitman J, Stamler JS (2000) Protection from nitrosative stress by yeast flavohemoglobin. Proc Natl Acad Sci 97:4672–4676. https://doi.org/10.1073/pnas.090083597
Liu T, Qian WJ, Mottaz HM, Gritsenko MA, Norbeck AD, Moore RJ, Smith RD (2006) Evaluation of multiprotein immunoaffinity subtraction for plasma proteomics and candidate biomarker discovery using mass spectrometry. Mol Cell Proteom 5:2167–2174. https://doi.org/10.1074/mcp.T600039-MCP200
Liu MS, Li HC, Lai YM, Lo HF, Chen LF (2013) Proteomics and transcriptomics of broccoli subjected to exogenously supplied and transgenic senescence-induced cytokinin for amelioration of postharvest yellowing. J Proteom 93:133–144. https://doi.org/10.1016/j.jprot.2013.05.014
Liu Q, Han Q, Wang N, Yao G, Zeng G, Wang Y, Huang Z, Sang J, Wang Y (2016) Tpd3-Pph21 phosphatase plays a direct role in Sep7 dephosphorylation in Candida albicans. Mol Microbiol 101:109–121. https://doi.org/10.1111/mmi.13376
Lorenz MC, Bender JA, Fink GR (2004) Transcriptional response of Candida albicans upon internalization by macrophages. Eukaryot Cell 3:1076–1087. https://doi.org/10.1128/EC.3.5.1076-1087.2004
Martin SW, Douglas LM, Konopka JB (2005) Cell cycle dynamics and quorum sensing in Candida albicans chlamydospores are distinct from budding and hyphal growth. Eukaryot Cell 4:1191–1202. https://doi.org/10.1128/EC.4.7.1191-1202.2005
Mayer FL, Wilson D, Jacobsen ID, Miramón P, Slesiona S, Bohovych IM, Brown AJ, Hube B (2012) Small but crucial: the novel small heat shock protein Hsp21 mediates stress adaptation and virulence in Candida albicans. PLoS ONE 7:e38584. https://doi.org/10.1371/journal.pone.0038584
Mc Manus BA, Coleman DC (2014) Molecular epidemiology, phylogeny and evolution of Candida albicans. Infect Genet Evol 21:166–178. https://doi.org/10.1016/j.meegid.2013.11.008
Michán C, Pueyo C (2009) Growth phase-dependent variations in transcript profiles for thioredoxin-and glutathione-dependent redox systems followed by budding and hyphal Candida albicans cultures. FEMS Yeast Res 9:1078–1090. https://doi.org/10.1111/j.1567-1364.2009.00558.x
Miller SE, Spurlock BO, Michaels GE (1974) Electron microscopy of young Candida albicans chlamydospores. J Bacteriol 119:992–999
Mun MS, Yap T, Alnuaimi AD, Adams GG, McCullough MJ (2016) Oral candidal carriage in asymptomatic patients. Aust Dent J 61:190–195. https://doi.org/10.1111/adj.12335
Navarathna DH, Pathirana RU, Lionakis MS, Nickerson KW, Roberts DD (2016) Candida albicans ISW2 regulates chlamydospore suspensor cell formation and virulence in vivo in a mouse model of disseminated candidiasis. PLoS ONE 11:e0164449. https://doi.org/10.1371/journal.pone.0164449
Nett JE, Lepak AJ, Marchillo K, Andes DR (2009) Time course global gene expression analysis of an in vivo Candida biofilm. J Infect Dis 200:307–313. https://doi.org/10.1086/599838
Neville BA, d’Enfert C, Bougnoux ME (2015) Candida albicans commensalism in the gastrointestinal tract. FEMS Yeast Res. https://doi.org/10.1093/femsyr/fov081
Nobile CJ, Bruno VM, Richard ML, Davis DA, Mitchell AP (2003) Genetic control of chlamydospore formation in Candida albicans. Microbiology 149:3629–3637. https://doi.org/10.1099/mic.0.26640-0
Odds FC (1988) Candida and Candidosis. Baillière Tindall, London
Odds FC, Bernaerts RIA (1994) CHROMagar Candida, a new differential isolation medium for presumptive identification of clinically important Candida species. J Clin Microbiol 32(8):1923–1929
Palige K, Linde J, Martin R, Böttcher B, Citiulo F, Sullivan DJ, Johann W, Claudia S, Staib C, Steffen R, Bernhard H, Morschhäuser J, Peter Staib J (2013) Global transcriptome sequencing identifies chlamydospore specific markers in Candida albicans and Candida dubliniensis. PLoS ONE 8(4):e61940. https://doi.org/10.1371/journal.pone.0061940
Panagoda GJ, Ellepola ANB, Samaranayake LP (2001) Adhesion of Candida parapsilosis to epithelial and acrylic surfaces correlates with cell surface hydrophobicity. Mycoses 44(1–2):29–35
Pereira MD, Eleutherio EC, Panek AD (2001) Acquisition of tolerance against oxidative damage in Saccharomyces cerevisiae. BMC Microbiol 1:11. https://doi.org/10.1186/1471-2180-1-11
Richard M, de Groot P, Courtin O, Poulain D, Klis F, Gaillardin C (2002) GPI7 affects cell-wall protein anchorage in Saccharomyces cerevisiae and Candida albicans. Microbiology 148:2125–2133. https://doi.org/10.1099/00221287-148-7-2125
Rosenberg M, Gutnick D, Rosenberg E (1980) Adherence of bacteria to hydrocarbons: a simple method for measuring cell-surface hydrophobicity. FEMS Microbiol Lett 9(1):29–33
Ruhnke M (2006) Epidemiology of Candida albicans infections and role of non-Candida albicans yeasts. Curr Drug Targets 7:495–504. https://doi.org/10.2174/138945006776359421
Samaranayake YH, Samaranayake LP, Yau JYY, Ellepola ANB, Anil S, Yeung KWS (2003) Adhesion and cell-surface-hydrophobicity of sequentially isolated genetic isotypes of Candida albicans in an HIV-infected Southern Chinese cohort. Mycoses 46(9–10):375–383. https://doi.org/10.1046/j.0933-7407.2003.00919.x
Sarkar P, Florczyk M, McDonough K, Nag D (2002) SSP2, a sporulation-specific gene necessary for outer spore wall assembly in the yeast Saccharomyces cerevisiae. Mol Genet Genom 267:348–358. https://doi.org/10.1007/s00438-002-0666-5
Schleit J, Johnson SC, Bennett CF, Simko M, Trongtham N, Castanza A, Hsieh EJ, Moller RM, Wasko BM, Delaney JR, Sutphin GL (2013) Molecular mechanisms underlying genotype-dependent responses to dietary restriction. Aging Cell 12:1050–1061. https://doi.org/10.1111/acel.12130
Seneviratne CJ, Wang Y, Jin L, Abiko Y, Samaranayake LP (2008) Candida albicans biofilm formation is associated with increased anti-oxidative capacities. Proteomics 14:2936–2947. https://doi.org/10.1002/pmic.200701097
Smith DA, Nicholls S, Morgan BA, Brown AJ, Quinn J (2004) A conserved stress-activated protein kinase regulates a core stress response in the human pathogen Candida albicans. Mol Biol Cell 15:4179–4190. https://doi.org/10.1091/mbc.e04-03-0181
Smits GJ, van den Ende H, Klis FM (2001) Differential regulation of cell wall biogenesis during growth and development in yeast. Microbiology 147:781–794. https://doi.org/10.1099/00221287-147-4-781
Sonneborn A, Bockmühl DP, Ernst JF (1999) Chlamydospore formation in Candida albicans requires the Efg1p morphogenetic regulator. Infect Immun 67:5514–5517
Sosinska GJ (2012) Adaptations in the wall proteome of the clinical fungus Candida albicans in response to infection-related environmental conditions, Ph.D. Thesis, University of Amsterdam, Netherland
Staib P, Morschhäuser J (2005) Liquid growth conditions for abundant chlamydospore formation in Candida dubliniensis. Mycoses 48(1):50–54. https://doi.org/10.1111/j.1439-0507.2004.01085.x
Staib P, Morschhäuser J (2007) Chlamydospore formation in Candida albicans and Candida dubliniensis–an enigmatic developmental programme. Mycoses 50:1–12. https://doi.org/10.1111/j.1439-0507.2006.01308.x
Tyc KM, Kühn C, Wilson D, Klipp E (2014) Assessing the advantage of morphological changes in Candida albicans: a game theoretical study. Front Microbiol 5:41. https://doi.org/10.3389/fmicb.2014.00041
Verma R, Chen S, Feldman R, Schieltz D, Yates J, Dohmen J, Deshaies RJ (2000) Proteasomal proteomics: identification of nucleotide-sensitive proteasome-interacting proteins by mass spectrometric analysis of affinity-purified proteasomes. Mol Biol Cell 11:3425–3439. https://doi.org/10.1091/mbc.11.10.3425
Walther TC, Brickner JH, Aguilar PS, Bernales S, Pantoja C, Walter P (2006) Eisosomes mark static sites of endocytosis. Nature 439:998. https://doi.org/10.1038/nature04472
Williams SM (2011) Investigating the role of Nrg1p and Tup1p during Candida albicans chlamydospore formation. McNair Scholars J 15:11
Yadav AK, Desai PR, Rai MN, Kaur R, Ganesan K, Bachhawat AK (2011) Glutathione biosynthesis in the yeast pathogens Candida glabrata and Candida albicans: essential in C. glabrata, and essential for virulence in C. albicans. Microbiology 157:484–495. https://doi.org/10.1099/mic.0.045054-0
Yoda K, Kawada T, Kaibara C, Fujie A, Abe M, Hashimoto H, Shimizu J, Tomishige N, Noda Y, Yamasaki M (2000) Defect in cell wall integrity of the yeast Saccharomyces cerevisiae caused by a mutation of the GDP-mannose pyrophosphorylase gene VIG9. Biosci Biotechnol Biochem 64:1937–1941. https://doi.org/10.1271/bbb.64.1937
Young ME, Karpova TS, Brügger B, Moschenross DM, Wang GK, Schneiter R, Cooper JA (2002) The Sur7p family defines novel cortical domains in Saccharomyces cerevisiae, affects sphingolipid metabolism, and is involved in sporulation. Mol Cell Biol 22:927–934. https://doi.org/10.1128/MCB.22.3.927-934.2002
Acknowledgements
Authors are thankful to Prof. Udhav V. Bhosle, Honorable Vice Chancellor, SRTM University, Nanded (MS) India for his encouragement and incessant support. Authors are also thankful to SERB, India for financial support under SERB FAST Track Scheme for Young Scientists to GBZ. GBZ acknowledge geneorous financial support of UGC under UGC-SAP-DRS II and DST under DST-FIST I to the School of Life Sciences, SRTM University, Nanded.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ingle, S., Kazi, R., Patil, R. et al. Proteome analysis of Candida albicans cells undergoing chlamydosporulation. J Proteins Proteom 10, 269–290 (2019). https://doi.org/10.1007/s42485-019-00024-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42485-019-00024-8