Abstract
Septoria tritici blotch (STB) is caused by Zymoseptoria tritici and is one of the most important wheat diseases worldwide. Molecular mechanisms underlying the resistance or susceptibility of wheat to Z. tritici are not fully known. MicroRNAs (miRNAs) are endogenous, small, non-coding RNAs known to play roles during biotic and abiotic stresses in most eukaryotes. To understand their biological function during STB infection, we selected a group of candidate miRNAs, and using RT-qPCR, assessed their expression in two resistant (Arina) and susceptible (Taichung 29) wheat cultivars at 0, 1, 3, 7, and 10 days post-inoculation. Results showed that tae-miR408 and tae-miR444 were downregulated in susceptible plants, while they were upregulated in resistant cultivars. In contrast, tae-miR171 was upregulated in both susceptible and resistant cultivars at the late stages of infection, indicating that this miRNA may not be involved in the resistant host. On the other hand, tae-miR156, tae-miR159, tae-miR167, tae-miR166b, tae-miR169, and tae-miR393 were differentially expressed in resistant and susceptible cultivars at different time points. Differential expression of these miRNAs in susceptible and resistant cultivars suggests their potential role in response to pathogen attacks, however, their exact functions and their gene targets remain unknown. Conclusively, our results provided more insights regarding the role of miRNAs (particularly tae-miR408 and tae-miR444) in STB pathosystem.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat (Triticum aestivum L.) is one of the most important crops in the world and provides about 55% of human carbohydrates (Gill et al. 2004). Wheat plants are constantly exposed to various abiotic and biotic stresses, which cause considerable crop loss and economic damage all over the world (Bray 2000; Ponomarenko et al. 2011; Danon and Eyal 1990). Wheat Septoria tritici blotch (STB) caused by Zymoseptoria tritici is one of these serious diseases and estimated to end with 60% crop loss in favorable conditions (Somasco et al. 1996). In recent years, non-coding RNAs (ncRNAs) have received significant attention since functional RNAs affect many cellular processes (Bentwich 2005; Birney et al. 2007; Washietl et al. 2007). Among these small ncRNAs, regulatory RNAs, owing to their role in regulating the expression of protein-encoding genes, have been increasingly studied (Khraiwesh et al. 2012. In plants, based on the biogenesis and function, small regulatory RNAs can be divided into two main groups: I) microRNAs (miRNAs) and II) Small interfering RNAs (siRNAs). Both types of RNA molecule function as negative regulators of gene expression (Voinnet 2009). It is well documented that miRNAs play a crucial role in regulating the expression of protein-encoding genes via transcriptional target gene silencing (TGS) such as miRNA-directed DNA methylation and/or posttranscriptional gene silencing (PTGS) (Khraiwesh et al. 2012) and exert their effect through mRNA translation disruption or mRNA degradation (Baulcombe 2004; Hannon 2002; Jones-Rhoades et al. 2006; Axtell and Bowman 2008; Axtell et al. 2011; Bartel 2004; Brodersen et al. 2008; Khraiwesh et al. 2010; Xiaoyun and Guiliang 2011). Studies proved the functions of miRNAs in response to biotic and abiotic stresses, modifying and moderating the gene expression and their functions in the plant’s defense system (Khraiwesh et al. 2012; Zhao et al. 2012; Gupta et al. 2012). The miR393 is the first reported miRNA that plays a significant role in regulating Arabidopsis genes in response to pathogens (Navarro et al. 2006). Later, more studies showed the role of miRNAs in various diseases and pathosystems, including powdery mildew infection (Xin et al. 2010) and sheath blight disease in rice (Talesh Sasani et al. 2020).
The discovery and functional characterization of the two main classes of regulatory RNA molecules including miRNAs and siRNAs in plants have changed our understanding of the mechanisms of gene regulation. In plants, miRNAs and siRNAs consist of 21 to 24 nucleotides, and both types function as negative regulators of gene expression (Voinnet 2009). The major difference between these two classes related to their genomic origin and biogenesis. The siRNAs are derived from long double-stranded RNA molecules while miRNAs are single-stranded RNAs and come from endogenous non-coding RNA that is mostly found within the introns of larger RNA molecules. Increasing research has demonstrated that miRNAs play critical roles in developmental and physiological processes as well as biotic and abiotic stress responses (Allen and Weiss 2010; Budak et al. 2015; Lu et al. 2008).
To our knowledge, some research has been performed regarding the interaction of wheat and Zymoseptoria tritici (Ma et al. 2020). However, comparative study in susceptible and resistant cultivars and their predicted target genes remind to be determined. In this paper, we selected nine plant conserved miRNAs (miR156, miR159, miR167, miR171, miR393, miR166b, miR169, miR408, miR444) that were previously unmasked to play an essential role in response to biotic and abiotic stresses (Zhao et al. 2012; Gupta et al. 2012; Navarro et al. 2006; Xin et al. 2010) and characterized the expression variations of the selected miRNAs and their possible target genes in resistant (Ariana cv.) and susceptible (Taichung 29) wheat cultivars. We hypothesized that the expression of selected miRNAs and consequently the expression of their target genes may differ in resistant and susceptible cultivars during the Z. tritici pathogenesis. We inoculated these cultivars with Z. tritici and monitored the expression of miRNAs and their target genes at various time points. Data confirmed a considerable fluctuation in expression of miRNAs.
Materials and Methods
Strains, fungal culture, and inoculation preparation
Seeds of wheat cultivars, Arina (STB resistant cv.) and Taichung 29 (STB sensitive cv), were kindly provided from Wageningen University. The Z. tritici isolate RM155 (IPO2166) was prepared as described previously (Mehrabi et al. 2015). The yeast-like spores were transferred from -80 °C freezer and were subsequently transferred onto Potato Dextrose Agar (PDA,potato 200 g/L, dextrose 20 g/L, agar 15 g/L) plates using a sterile fine needle. The plates were kept at 18 °C for a few days (Somasco et al. 1996; Bentwich 2005; Birney et al. 2007) to allow fungal growth. The Z. tritici isolate then was propagated in YGM (yeast extract 10 g/L, glucose 20 g/L) using a rotary shaker (150 rpm at 18 °C for 120 h). The obtained yeast-like spores supplemented with 0.15% Tween 20 were collected, washed with sterile water, and the concentration was adjusted to 102 spores/mL. Spraying was done until the whole leaf was wet. Finally, the prepared inoculum was kept at -80 °C for long-term storage or inoculation assays.
Inoculation and sample preparation
The wheat seeds were sown into 15 cm diameter pots (Axtell et al. 2011) containing sterile soil and incubated in a greenhouse (16 h light /8 h dark cycle, 20 ± 2 °C). Ten-day-old wheat seedlings (Arina and Taichung 29) were inoculated (using the device, spores were sprayed on the leaves) with Z. tritici spores and were kept at 100% relative humidity for 48 h in black plastic bags. Subsequently, the seedlings were transferred to a greenhouse (85 > % relative humidity, 16 h light/8 h dark regime, 20 ± 2 °C). The negative controls were inoculated by sterile distilled water without spore. The sampling was done on day 0 (before infection), day 1, day 3, day 7, and day 10 after inoculation for treated and control plants. The prepared specimens were stored at -80 for RNA extraction.
Total RNA extraction and cDNA synthesis
According to the manufacturer’s protocol, total RNA was extracted from 0.1 g of the ground samples using Trizol reagent (Ambion, USA). The extracted RNAs were treated by DNase I (Fermentas, USA) to eliminate any remining DNA. Then, concentration and the quality of RNA were evaluated by UV–visible spectroscopy (CECIL-CE2301, England) and on 1% agarose gel, respectively. Synthesis of the mRNAs was done by addition of 1 µl of the simple oligo-dT (Table 1) and 0.5 µl of the random hexamer primers to the 1000 ng of the extracted RNAs in 0.5 ml microtubes. Mixture was incubated at 65 °C for 5 min and immediately transferred into an ice-cold solution. Subsequently, the cDNA synthesis mixture including 1 µl dNTP (40 mM), 4 µl RT buffer (5X) (Fermentas), 0.1 µl RNase inhibitor (Fermentas), 0.5 µl reverse transcriptase (RT) (Fermentas) and DEPC- treated water was added up to 20 µl. Mixtures were incubated in a thermocycler for 60 min at 42 °C and then for 10 min at 70 °C. Following the manufacture’s protocol, to provide cDNA from miRNAs, RNA samples were polyadenylated using a poly-A polymerase. Next, the first-strand cDNA was synthesized using PrimeScriptTM 1st strand cDNA synthesis kit (Takara, Japan) and the addition of anchored Oligo-dT, in a total reaction volume of 25 µl. All of the synthesized cDNAs were monitored by 2% agarose gel.
In silico miRNA target gene prediction and primer designing
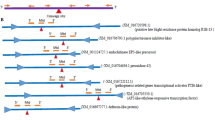
Sequences of selected miRNAs were extracted from the miRBase (www.mirbase.org) database, and their target genes were estimated by employing TarBase (http://diana.cslab.ece.entua.gr/tarbase), (http://plantgrn.noble.org/psRNATarget), (http://www.leonxie.com/targetAlign.php), (http://bioinformatics.psb.ugent.be/webtools/tapir/) online servers (Table 1) (The alignments were shown in supplementary information 2). The miRNAs sequences were chosen as forward primers (with slight modifications at 3' and 5' ends) together with a universal reverse primer (Table 2). To design primer pairs for target genes, Oligo7 and Primer3 web (http://primer3.ut.ee, version 4.1.10) tools were used and further analyzed by IDT-Oligo-analyzer online software (http://euidtdna.com/calc/analyser). The specificity of designed primers was evaluated by NCBI Primer-BLAST tool (http://www.ncbi.nlm.nih.gov/tools/primer-blast/). Confirmed sequences were ordered to synthesize at IDT Co, Belgium.
Quantification of miRNAs expression by Real-time PCR
In order to study the expression of selected miRNAs in response to wheat STB diseases, RT-qPCR was performed by using snRNA-U6 as a house-keeping reference gene. In this regard, ABI Real-Time system (Applied Biosystems StepOne) was set up as follows: 95 °C for 5 min (DNA denaturation, and SYBR Green binding), 45 cycles of 95 °C for 15 s, 53 °C for 20 s, 72 °C for 30 s and a final extension at 72 °C for 8 min. The experiment was done at two replicates, and the Non-template (NTC) and the Non-reverse transcriptase (NRC) samples were used as negative controls to check master mix and cDNAs, respectively. Polyacrylamide gel electrophoresis (PAGE) was carried out to verify qRT-PCR amplification products followed by cloning of the products in TA-pTG19 cloning vector, transferring into competent Escherichia coli (DH5α) cells, screening, and plasmid isolation using the plasmid extraction kit (GeneAll, Korea). The isolated plasmids were subsequently sequenced to confirm cloning accuracy.
Real-time PCR validation of target genes expression
To evaluate the target genes’ expression pattern, Real-time PCR was performed using an ABI (Applied Biosystems StepOne) system with SYBR Green master mix (BioFact, South Korea) and employing β-Actin as housekeeping gene. The following thermal cycles were exerted: 94 °C for 15 min, 40 cycles of 94 °C for 30 s, 60 °C for 30 s, 72 °C for 20 s, and a final extension at 72 ℃ for 8 min. Every sample was repeated in duplicate, and results were monitored by 2% agarose gel.
Statistical analysis of real-time PCR data
Data obtained from the treated plants were compared to control cultivars at the same time points by employing Graphpad Prism-5 software using the 2–ΔΔCT method and the expression fold changes. Correlation between miRNAs and target genes was analyzed based on Pearson's product-moment method by Microsoft Excel-2007 software.
Results
Pathogencity assay indicated difference in disease severity between cultivars
For biological assay, seeds of resistant cultivars (Arina) and susceptible cultivar (Taichung 29) to STB was sown in three biological replicates. Seedling were grown in healthy condition for ten days. Ten days seedlings were inoculated and treated at favorable condition for disease incidence up to 20 days after inoculation. Based on assay, Arina did not show any disease symptom, while Taichung 29 was severely diseased with chlorosis and complete necrosis at 15 dpi. Thus, it was no longer possible to extract RNA at 15 dpi (Fig. 1).
PCR analysis confirmed the quality of extracted RNA and synthesized cDNA
All extracted RNAs and synthesized cDNAs were evaluated qualitatively by using an agarose gel. Results indicated a proper RNA extraction with clear ribosomal bands. Additionally, PCR amplification of the prepared cDNAs using U6 and β-Actin forward primers along with a universal reverse primer confirmed the quality of cDNA synthesis as well as poly-A sequence related to cDNA samples(Fig. 2).
Extracted RNA and synthesized cDNA from miRNAs and their target genes. A Qualities evaluation of the whole extracted RNAs. B Quality of synthesized cDNAs is tested for β-Actin housekeeping gene. C Quality of synthesized cDNAs is tested for U6 housekeeping gene. NRC and NTC stand for no RNA control and no templet control, respectively
Real time analysis indicated tae-miRNA expression variations during plant-pathogen interactions
Expression data obtained from RT-qPCR analysis indicated that candidate miRNAs have different expression pattern at various time points of pathogenesis process (Figs. 3, 4 and 5). Based on expression pattern in both susceptible and resistant cultivars, RT-qPCR analysis proposes three main subgroups of miRNAs the early-responsive group of miRNAs, including tae-miR444 and tae-miR408, which their expression were suppressed in susceptible cultivar, but provoked in resistant one immediately after Z. tritici infection (Fig. 3A, B). The expression pattern in the second group includes tae-miR156, tae-miR159, tae-miR166b, tae-miR167, tae-miR169, and tae-miR393 was somewhat similar to the pattern of the first group of (early-responsive) miRNAs. However, they showed a delayed response compared to the first group (Fig. 4A-F). Tae-miR171, as the only member of the third group, showed equally high expression level at the late stage of infection in both susceptible and resistant cultivars, (Fig. 5).
Expression patterns for the first group of miRNAs during pathogenesis. A tae-miR408. B tae-miR444. If a p-value is less than 0.05, it is showed with one star (*). If a p-value is less than 0.01, it is showed with 2 stars (**). If a p-value is less than 0.001, it is showed with three stars (***). If a p-value is less than 0.0001, it is showed with three stars (****)
MiRNAs affect the expression of their target genes during pathogenesis
To study the effect of miRNAs molecules on their proposed targets, we selected the SPL-Ta.3711 gene due to its described role in wheat powdery mildew pathosystem as a transcription factor and also Ta.138051 (GAMYB1) gene due to its effective function in other diseases (Gupta et al. 2012; Xin et al. 2010). These two genes are targets for miR156 and miR159, respectively (Supplementary information 2). Co-expression study of the miR156 and SPL gene in resistant cultivar indicated a negative correlation between their gene expression. Indeed, a significant negative correlation coefficient of -0.38 (p-value < 0.05) was calculated in which the gene expression level of SPL-Ta.3711 was decreased upon the increase of tae-miR156 expression and conversely, is decreased by increasing the expression of SPL-Ta.3711 gene (Fig. 6A, B). Thanks to the constant expression of tae-miR159 in resistant cultivar, we evaluated its expression correlation with its target gene in susceptible cultivar during pathogenesis. This assessment illustrated a negative correlation between miRNA 159 expression and its target GAMYB3 gene, with a correlation coefficient of -0.18. The results are shown in Fig. 6C, D.
Correlation analysis of the expression for miRNAs and their targets. A Reverse correlation between the expression of tae-miR156 and SPL-Ta.3711. B Negative correlation coefficient of miR156 and SPL-Ta.3711 transcripts in resistant cv. C Converse correlation between the expression of tae-miR159 and GAMYB3. D Negative correlation coefficient of miR159 and GAMYB3 transcripts in the susceptible cultivar
Discussion
Expression of plant genes may differ when plant is growing in a normal condition from when it is treated with biotic or abiotic stresses. The difference in gene expression enable plant to adapt to the imposed stress conditions. Several regulatory processes are involved in global gene expression. MicroRNAs are a group of small non-coding RNAs and are known as regulatory elements of expression for genes that play important roles in response to stress (Sunkar et al. 2007; Phillips et al. 2007). Study on several miRNAs showed variation of expression profiles among plant species and even among tissues of a same species (Tang et al. 2012). Based on this evidence, it is valuable to know the role and relation of miRNAs molecules in disease development.
In this study, the expression patterns of the nine miRNAs including tae-miR156, tae-miR159, tae-miR167, tae-miR171, tae-miR393, tae-miR166b, tae-miR169, tae-miR408, tae-miR444 were assessed in two wheat cultivars Ariana (resistant) and Taichung29 (susceptible) inoculated with Z. tritici. Data revealed that the expression miRNAs were changed when plants were inoculated with fungus. The role of these miRNAs have been previously reported in response to wheat stem rust (Gupta et al. 2012), wheat powdery mildew (Xin et al. 2010), and other fungal and bacterial pathogenes (Zhao et al. 2012; Navarro et al. 2006). However, this study is regarding the expression pattern of miRNAs in the wheat STB pathosystem.
Due to their expression profile, these miRNAs were classified into three main groups. The first group include tae-miR444 and tae-miR408 that their expression increased in resistant cultivars in response to Z. tritici (Fig. 4). Researches have shown that tae-miR408 can regulate the transcript level of gene active in response to biotic and abiotic stresses such as Plantacyanin-like protein, the chemocyanin-like protein (TaCLP1), and the transcripts of several laccase genes involved in cell-to-cell signaling and lignin formation (Gupta et al. 2014; Feng et al. 2013). Notably, such expression changes for genes related to lignin synthesis have also been reported during the interaction of sensitive and resistant cultivars in wheat stem rust pathosystem (Puccinia graminis f. sp. tritici) (Gupta et al. 2012). Studies have also indicated that miR444 does affect the plant growth, stress response, and flowering time of the plant, as well as the hormonal pathways of gibberellic acid and abscisic acid, by regulating gene transcripts such as MADS-box transcription factors and C3HC4 zinc-binding domain protein family (Gupta et al. 2012; Sun 2012; Khong et al. 2015). The decreased expression of MADS-box 26 caused increased resistance of rice to two important fungal (Magnaporthe oryzae) and bacterial (Xanthomonas oryzae) pathogens, as well as increasing of rice tolerance to water deficiency. However, it did not show any detectable effect on growth indexes (Khong et al. 2015). Such cases are in agreement with our findings regarding the increasing expression of tae-miR444 in the resistant cultivar, which probably reduced MADS-box expression and thus caused plant resistance. In wheat stem rust pathosystem, hormonal imbalances also lead to the response and accumulation of miR444 at the beginning of infection for two days post-infection (dpi), and retention to its normal level on day ten post-infection (Gupta et al. 2012). Increase of tae-miR444 expression in the wheat Powdery mildew pathosystem (Erysiphe graminis f. sp. tritici) has been observed after 12 h post-infection (hpi) in susceptible cultivar (Xin et al. 2010), which is consistent with the early hours of infection in the present study.
Expression of the second group of miRNA genes follow a similar patterns to the first group, however with some delay. Their regulation in resistant plant infected with pathogen indicates their possible roles in response to the pathogen. In this group, tae-miR156 is the most important member. In the present study, the inverse correlation between tae-miR156 expression and transcription factor suggest it as one of the targets of Tae-miR156 and agrees with the study indicating its possible role to stress resistance (Fig. 5A) (Wang and Wang 2015). For tae-miR159, although significant increase in expression level was observed in beginning of fungal infection on susceptible cultivar, its expression was suppressed after three dpi (Fig. 5B) (Somasco et al. 1996). In contrast to susceptible ones, the expression level of tae-tae-miR159 in resistant cultivars was induced from seven dpi onwards (Fig. 5B). The expression of miR159 at late time points may decrease expression level of its target genes required for full development and sporlation of fungus. Previous studies have introduced the MYB transcription factor family genes as the tae-miR159 target genes (Wang et al. 2012). This family of transcription factors plays a vital role in the growth and hormonal signaling pathways, secondary metabolites, disease resistance, cell shape, and plant development (Du et al. 2009; Eulgem 2005). Investigation of the barley-Blumeria graminis f. sp. hordei interaction showed that the expression level of tae-miR159 in susceptible cultivar is increased at 12 hpi; however, the expression level of MYB is decreased. (Dağdaş 2009). Nevertheless, the precise role of MYB transcription factor in resistance to fungal diseases needs to be studied.
The other four members of group 2 are tae-miR166b, tae-miR167, tae-miR169 and tae-tae-miR393. The expression of tae-miR167, tae-miR169 and tae-miR393 in resistant cultivar follow similar patterns to tae-miR156 and tae-miR159 inducing about 7dpi, however the expression of tae-miR169 was decreased after 7 dpi at the necrotrophic stages of the pathogenesis (Fig. 5C-F). Expression of tae-miR166b followed similar pattern to tae-miR169, but at very low level. The expression pattern of tae-tae-miR167 in the biotrophic stage and its increase in the fungal necrotrophic stage agrees with the research of Yang et al. (Yang et al. 2006). Auxin Response Factor (ARFs) transcription factors have been identified as tae-miR167 target genes (Yang et al. 2006; Rhoades et al. 2002; Ljung et al. 2002). This function is through the cooperation of tae-miR167 with tae-miR393 and miR160 via precise adjustment of auxin assessment and its signaling (Soltani et al. 2006; Sunkar et al. 2012). It is illustrated that tae-miR393 affects the auxin signaling pathway by regulation of auxin receptors Transport Inhibitor Response 1 (TIR1) and AFB family members (Auxin signaling F-Box proteins) (AFB1, AFB2, AFB3) (Navarro et al. 2006; Xin et al. 2010). These results are consistent with previous studies regarding the role of auxin hormone involvement in plant susceptibility to disease (Navarro et al. 2006). Accumulation of auxin hormone in susceptible cultivar of barley and its absence in resistant cultivar (Dağdaş 2009) during interaction with Blumeria graminis f. sp. hordei indicates the role of TIR1 and ARFs as target genes of miRNAs, specially tae-miR393 in regulation of plant immunity (Navarro et al. 2006). Expression level of tae-miR169 in resistant cultivar to barley powdery mildew is higher than susceptible cultivar, and it shows the negative expression correlation with its target gene in the resistant cultivar (Dağdaş 2009). It has been reported that in rice and Arabidopsis, the expression of the CCAAT-box transcription factor is controlled by miR169 (Combier et al. 2006; Abdel-Ghany and Pilon 2008). Low expression level of tae-miR169 in the susceptible cultivar and expression in biotrophy stage of fungal on resistant cultivar suggest its role in the biotrophic stage of pathogenicity.
The remaining molecule is tae-miR171, the only member of the third group, which showed a similar growing expression pattern in both susceptible and resistant cultivars at the late stages of infection (Fig. 6). It seems that tae-miR171 may not play a pivotal role in resistance or susceptibility in the studied pathosystem. Studies on wheat stem rust disease have shown that the expression level of tae-miR171 is reduced in both resistant and susceptible strains (Gupta et al. 2012). This difference may originates from the type of pathosystem, level of stress, experiment conditions and type of studied plant tissue (Wang et al. 2011).
Conclusion
Collectively, based on the expression patterns of the studied miRNAs, it seems that tae-miR156, tae-tae-miR159, tae-miR167, tae-miR393, tae-miR166b, tae-miR169, tae-miR408, and tae-miR444 are possibly involved in the wheat septoria pathosystem. In adddition, based on their induction time and their expression pattern, they are classified into three main groups comprising the early-responsive group, the delayed responsive group, and the non-responsive group. Fungus Z. tritici is a hemibiotrophic fungus and has a biotrophic stage (up to the 7dpi) in which no disease symptom is visible and a necrotrophic stage in which leaf chlorosis and necrosis occurs on susceptible cultivar (Ponomarenko et al. 2011). The expression pattern of plant miRNAs at the switching stage of biotrophy to necrotrophy has changed. This suggest that for each type of life stage a different group of transcription factors and their downstream genes are responsive to infection. Finding molecular markers in asymptomatic situations of wheat STB disease is of great importance, which could pave the way for the early detection of the disease. Although the studied miRNAs predominantly showed specific and disease-related responses in both susceptible and resistant cultivars. Rapid response and high expression levels of tae-miR408 and tae-miR444 in both cultivars introduces them as good biomarkers for early detection of disease. However, their expression patterns in different wheat cultivars with various degrees of resistance to Z. tritici needs to be assessed carefully.
Data availability
All supporting data would be available upon request.
References
Abdel-Ghany SE, Pilon M (2008) MicroRNA-mediated systemic down-regulation of copper protein expression in response to low copper availability in Arabidopsis. J Biol Chem 283(23):15932–15945
Allen KE, Weiss GJ (2010) Resistance may not be futile: microRNA biomarkers for chemoresistance and potential therapeutics. Mol Cancer Ther 9(12):3126–3136
Axtell MJ, Bowman JL (2008) Evolution of plant microRNAs and their targets. Trends Plant Sci 13(7):343–349
Axtell MJ, Westholm JO, Lai EC (2011) Vive la différence: biogenesis and evolution of microRNAs in plants and animals. Genome Biol 12(4):1–13
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297
Baulcombe D (2004) RNA silencing in plants. Nature 431(7006):356–363
Bentwich I (2005) Prediction and validation of microRNAs and their targets. FEBS Lett 579(26):5904–5910
Birney E, Stamatoyannopoulos JA, Dutta A, Guigó R, Gingeras TR, Margulies EH et al (2007) Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447(7146):799–816
Bray EA (2000) Response to abiotic stress. Biochemistry and molecular biology of plants. 1158–203
Brodersen P, Sakvarelidze-Achard L, Bruun-Rasmussen M, Dunoyer P, Yamamoto YY, Sieburth L et al (2008) Widespread translational inhibition by plant miRNAs and siRNAs. Science 320(5880):1185–1190
Budak H, Kantar M, Bulut R, Akpinar BA (2015) Stress responsive miRNAs and isomiRs in cereals. Plant Sci 235:1–13
Combier J-P, Frugier F, de Billy F, Boualem A, El-Yahyaoui F, Moreau S et al (2006) MtHAP2-1 is a key transcriptional regulator of symbiotic nodule development regulated by microRNA169 in Medicago truncatula. Genes Dev 20(22):3084–3088
Dağdaş YF (2009) Investigating the roles of micrornas in biotic stress responses and functional characterization of a novel ztl-type f-box protein via virus induced gene silencing
Danon T, Eyal Z (1990) Inheritance of resistance to two Septoria tritici isolates in spring and winter bread wheat cultivars. Euphytica 47(3):203–214
Du H, Zhang L, Liu L, Tang X-F, Yang W-J, Wu Y-M et al (2009) Biochemical and molecular characterization of plant MYB transcription factor family. Biochem Mosc 74(1):1–11
Eulgem T (2005) Regulation of the Arabidopsis defense transcriptome. Trends Plant Sci 10(2):71–78
Feng H, Zhang Q, Wang Q, Wang X, Liu J, Li M et al (2013) Target of tae-miR408, a chemocyanin-like protein gene (TaCLP1), plays positive roles in wheat response to high-salinity, heavy cupric stress and stripe rust. Plant Mol Biol 83(4–5):433–443
Gill BS, Appels R, Botha-Oberholster A-M, Buell CR, Bennetzen JL, Chalhoub B et al (2004) A workshop report on wheat genome sequencing: International Genome Research on Wheat Consortium. Genetics 168(2):1087–1096
Gupta OP, Permar V, Koundal V, Singh UD, Praveen S (2012) MicroRNA regulated defense responses in Triticum aestivum L. during Puccinia graminis f. sp. tritici infection. Mol Biol Rep 39(2):817–824
Gupta OP, Sharma P, Gupta RK, Sharma I (2014) Current status on role of miRNAs during plant–fungus interaction. Physiol Mol Plant Pathol 85:1–7
Hannon GJ (2002) RNA interference. Nature 418(6894):244–251
Jones-Rhoades MW, Bartel DP, Bartel B (2006) MicroRNAs and their regulatory roles in plants. Annu Rev Plant Biol 57:19–53
Khong GN, Pati PK, Richaud F, Parizot B, Bidzinski P, Mai CD et al (2015) OsMADS26 negatively regulates resistance to pathogens and drought tolerance in rice. Plant Physiol 169(4):2935–2949
Khraiwesh B, Arif MA, Seumel GI, Ossowski S, Weigel D, Reski R et al (2010) Transcriptional control of gene expression by microRNAs. Cell 140(1):111–122
Khraiwesh B, Zhu J-K, Zhu J (2012) Role of miRNAs and siRNAs in biotic and abiotic stress responses of plants. Biochim Biophys Acta (BBA)-Gene Regulator Mech 1819(2):137–148
Ljung K, Hull AK, Kowalczyk M, Marchant A, Celenza J, Cohen JD et al (2002) Biosynthesis, conjugation, catabolism and homeostasis of indole-3-acetic acid in Arabidopsis thaliana. Springer, Auxin Molecular Biology, pp 249–272
Lu M, Zhang Q, Deng M, Miao J, Guo Y, Gao W et al (2008) An analysis of human microRNA and disease associations. PLoS ONE 3(10):e3420
Ma X, Wiedmer J, Palma-Guerrero J (2020) Small RNA bidirectional crosstalk during the interaction between wheat and Zymoseptoria tritici. Front Plant Sci 10:1669
Mehrabi R, Makhdoomi A, Jafar-Aghaie M (2015) Identification of New Sources of Resistance to Septoria Tritici Blotch Caused by Z ymoseptoria tritici. J Phytopathol 163(2):84–90
Navarro L, Dunoyer P, Jay F, Arnold B, Dharmasiri N, Estelle M et al (2006) A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 312(5772):436–439
Phillips JR, Dalmay T, Bartels D (2007) The role of small RNAs in abiotic stress. FEBS Lett 581(19):3592–3597
Ponomarenko A, Goodwin SB, Kema GH (2011) Septoria tritici blotch (STB) of wheat. Plant Health Instr, Septoria tritici blotch (STB) of wheat. https://doi.org/10.1094/PHI-I-2011-0407-01
Rhoades MW, Reinhart BJ, Lim LP, Burge CB, Bartel B, Bartel DP (2002) Prediction of Plant microRNA Targets Cell 110(4):513–520
Soltani BM, Ehlting J, Hamberger B, Douglas CJ (2006) Multiple cis-regulatory elements regulate distinct and complex patterns of developmental and wound-induced expression of Arabidopsis thaliana 4CL gene family members. Planta 224(5):1226–1238
Somasco O, Qualset C, Gilchrist D (1996) Single-gene resistance to Septoria tritici blotch in the spring wheat cultivar ‘Tadinia.’ Plant Breeding 115(4):261–267
Sun G (2012) MicroRNAs and their diverse functions in plants. Plant Mol Biol 80(1):17–36
Sunkar R, Chinnusamy V, Zhu J, Zhu J-K (2007) Small RNAs as big players in plant abiotic stress responses and nutrient deprivation. Trends Plant Sci 12(7):301–309
Sunkar R, Li Y-F, Jagadeeswaran G (2012) Functions of microRNAs in plant stress responses. Trends Plant Sci 17(4):196–203
Talesh Sasani S, M Soltani B, Mehrabi R, Samavatian H, Padasht-Dehkaei F (2020) Expression Alteration of Candidate Rice miRNAs in Response to Sheath Blight Disease. Iran J Biotechnol
Tang Z, Zhang L, Xu C, Yuan S, Zhang F, Zheng Y et al (2012) Uncovering small RNA-mediated responses to cold stress in a wheat thermosensitive genic male-sterile line by deep sequencing. Plant Physiol 159(2):721–738
Voinnet O (2009) Origin, biogenesis, and activity of plant microRNAs. Cell 136(4):669–687
Wang H, Wang H (2015) The miR156/SPL module, a regulatory hub and versatile toolbox, gears up crops for enhanced agronomic traits. Mol Plant 8(5):677–688
Wang L, Hua D, He J, Duan Y, Chen Z, Hong X et al (2011) Auxin Response Factor2 (ARF2) and its regulated homeodomain gene HB33 mediate abscisic acid response in Arabidopsis. PLoS Genet 7(7):e1002172
Wang Y, Sun F, Cao H, Peng H, Ni Z, Sun Q et al (2012) TamiR159 directed wheat TaGAMYB cleavage and its involvement in anther development and heat response. PLoS ONE 7(11):e48445
Washietl S, Pedersen JS, Korbel JO, Stocsits C, Gruber AR, Hackermüller J et al (2007) Structured RNAs in the ENCODE selected regions of the human genome. Genome Res 17(6):852–864
Xiaoyun JIAJY, Guiliang TANG (2011) MicroRNA-mediated DNA methylation in plants. Front Biol 6(2):133–139
Xin M, Wang Y, Yao Y, Xie C, Peng H, Ni Z et al (2010) Diverse set of microRNAs are responsive to powdery mildew infection and heat stress in wheat (Triticum aestivum L.). BMC Plant Biol 10(1):1–11
Yang JH, Han SJ, Yoon EK, Lee WS (2006) Evidence of an auxin signal pathway, microRNA167-ARF8-GH3, and its response to exogenous auxin in cultured rice cells. Nucleic Acids Res 34(6):1892–1899
Zhao J-P, Jiang X-L, Zhang B-Y, Su X-H (2012) Involvement of microRNA-mediated gene expression regulation in the pathological development of stem canker disease in Populus trichocarpa. PLoS ONE 7(9):e44968
Acknowledgements
This study was supported by the Iranian National Scientific Foundation (Project No. 92043155 provided to RM) as well as foundation provided by Tarbiat Modares University.
Author information
Authors and Affiliations
Contributions
H.S., B.M.S., and R.M. and MKJ conceived, and designed the study and wrote the paper. H.S. and R.M., performed experiments. H.S., B.M.S., and R.M., analyzed the results. All authors reviewed the results and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethical approval
This article is not submitted for publication elsewhere and complies with the Ethical Rules applicable for J. of plant pathology. Also, this article does not contain any studies with human participants or animals.
Conflict of interest
The authors declare no conflicts of interest with any financial organization regarding the material discussed in the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Samavatian, H., Soltani, B.M., Yousefi, F. et al. Susceptible and resistant wheat cultivars show different miRNAs expression patterns in response to Zymoseptoria tritici. J Plant Pathol 105, 437–447 (2023). https://doi.org/10.1007/s42161-022-01278-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42161-022-01278-w