Abstract
Twenty five isolates of Acidovorax avenae subsp. avenae causing bacterial leaf streak collected from infected maize crop grown in different geographical regions of Punjab, northern India were characterized for their molecular diversity using twenty five random amplified polymorphic DNA (RAPD) markers. All the primers showed amplification with a total of 1769 amplified fragments. Among RAPD primers, S112 was found to be highly polymorphic with PIC value of 0.95 while OPT-4 was least polymorphic. The size of amplified DNA fragments ranged from 59 bp to 3.9 kbp. Dendrogram based on molecular data generated by 25 RAPD collectively divided all 25 isolates into four groups. From 25 isolates, 20 were used for their pathological characterization and based on the multivar cluster analysis of pathogenicity data, these were divided into three groups, each producing a distinct range of disease score on particular maize inbred/hybrid used in this study. Isolate Aaa2 was found to be the most virulent with an average disease score of 8.6, whereas Aaa17 was found least virulent with average disease score of 5.1. The most virulent isolates of this pathogen were present in central Punjab. This was the first attempt to study the genetic diversity and pathogenic variation in Punjab populations of A. avenae subsp. avenae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Maize (Zea mays L.) crop is grown over diverse landscapes and climatic conditions, and along with wheat and rice, it contributes at least 30% of the food calories to the human population in developing countries (FAOSTAT 2014). It is estimated that maize production needs to be doubled by 2050 to meet the increasing demand of growing human population. One way to meet this demand is through area expansion, which, however, is not a sustainable option. The other option is to reduce the yield losses due to biotic and abiotic stresses. Maize is a major summer season crop across South Asia grown over an area of about 12.1 million hectares and annual productivity of 29.35 quintals per hectare (FAOSTAT 2014). Maize crop is going to get a boost in its production because of its pivotal role in crop diversification program as well as the establishment of CIMMYT-BISA in this region (Kaur et al. 2014). However, biotic stresses play major constraint on its production and cause significant losses. The hot and humid climate particularly during summer season makes the crop more vulnerable to bacterial and fungal pathogens. During the recent years, the incidence of bacterial stalk rot (Kumar 2015; Kumar et al. 2017) and bacterial leaf streak (Dhkal et al. 2016a) has invariably increased on maize.
Bacterial leaf streak (BLS), although considered to be a minor disease, is gaining importance nowadays with variable incidence on different hybrids/inbreds grown in the State. The disease appears as water soaked spots on both upper and lower leaf surface which develop rapidly into long stripes. These lesions/streaks usually have wavy edges in contrast to linear lesion. Sometimes, symptoms start appearing within two days upon infection and whole leaf turns papery within five to six days of inoculation (Dhkal et al. 2016a). BLS can cause upto 30% yield losses under favourable conditions (Techati 2008). The earlier report by Ullasa et al. (1967) from Delhi, India confirmed Xanthomonas rubrilineans as causal agent of BLS. The bacterium also causes red stripe in sugarcane and was described as Pseudomonas rubrilineans by Dange (1972). The species has undergone several changes in its nomenclature (Johnson et al. 1945; Smith and White 1988; Pataky et al. 1997) and finally Willems et al. (1992) reported that maize pathogen Pseudomonas avenae belonged to the genus Acidovorax and reclassified it as A. avenae subsp. avenae. Acidovorax avenae subspecies have been known to be both internally and externally seed-borne (Thind 2012).
Although other subspecies of A. avenae such as citrulli have been extensively studied for their pathogenic and molecular aspects (Shrestha et al. 2013; Silva et al. 2016; Zivanovic and Walcott 2017), little information is available on the extant of diversity of A. avenae subsp. aveane especially from this important maize growing part of India. The present study, therefore aimed to characterize pathogenic and genetic diversity among the Punjab populations of A. avenae subsp. avenae. A better insight into the diversity of A. avenae subsp. avenae will be highly useful in developing strong basis for bacterial disease management in future and hence sustainable maize production system with enhanced yield could be expected.
Materials and methods
Bacterial isolates, their isolation and maintenance
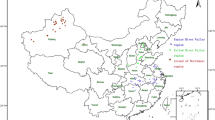
Twenty five isolates of A. avenae subsp. avenae from infected maize plants were collected from farmer’s field in different maize growing regions of Punjab during Kharif 2014. Morphological identification of all the twenty five isolates was done as per procedure described in Bergey’s manual of Determinative Bacteriology (1975). Further, the identity of A. avenae subsp. avenae was confirmed with the help of rDNA based primers (Dhkal et al. 2016a). Each isolate was maintained as pure culture on King’s B agar plates using single bacterial colony.
DNA extraction
Genomic DNA from all the 25 isolates was isolated using Invitrogen Easy DNA® Kit as per manufacture’s protocol. Isolated DNA was purified by adding RNase (Promega Inc) to a final concentration of 10 μg ml-1. The quantity and quality of DNA was checked by using TECAN 2000 Nanoquant Plate reader. The DNA of all the samples was diluted to 25 ng/μl in nuclease free water and stored at -20 °C for further use.
PCR amplification
A set of twenty five RAPD primers (10 base pair oligonucleotides, Operon Technologies Inc.) was used in the present investigations for amplification of 25 isolates of A. avenae subsp. avenae (Table 1). PCR amplification was carried out in final volume of 25 μl, containing 2 mM MgCl2, 1 unit Taq DNA polymerase, 0.1 mM dNTPs, 1X PCR Buffer [Tris HCl (pH 9.0 at 25 °C), KCl and Triton® X-100], 1 μM primer and 50 ng of gDNA. Each reaction for PCR amplification (Saiki et al. 1988) was performed in Eppendorf Master Cycler ProS with initial denaturation of 94 °C for 4 min, 35 cycles of denaturation at 94 °C for 1 min, annealing at 36 °C for 1 min, extension at 72 °C for 2 min and final elongation at 72 °C for 3 min. The amplified products were resolved on 0.8% agarose gels in 0.5 X TBE buffer (Tris base 45 mM, boric acid 45 mM, EDTA 1 mM), stained with ethidium bromide at concentration of 0.5 μg ml−1 of buffer. PCR products were resolved by gel electrophoresis at 5 V cm−1 for 2–3 h. The gel was visualized under UV light and photographed using Gene Snap® software programme (Syngene Inc., UK).
Genetic data analysis
DNA fingerprinting data generated by RAPD based markers were converted into binary matrix form as 1 and 0 representing the presence and absence of band respectively. The size of amplified products by RAPD markers were ascertained by comparing them with the known marker (1 kb ladder, BR Biochem Life Science Pvt. Ltd., India). To find the informativeness of the primer used, the polymorphic information content (PIC) was calculated for each marker by using formula: PIC = 1-n Σ(Pij)2, where Pij is frequency of jth allele in ith primer and summations extend over ‘n” pattern (Botstein et al. 1980). The binary data was further analysed with statistical software DARwin 6.0.10. Dendrogram was generated and the isolates were grouped on the basis of Neighbor Joining using DICE coefficient.
Maize hybrids and inbreds
For pathotypic characterization of different isolates of A. avenae subsp. avenae, nine maize hybrids and inbred lines viz. PMH-1, PMH-2, FR-632, Punjab Sweet Corn-1, CM-600, CM-139, LM-13, CM-140 and CML-25 procured from Maize Section, Department of Plant Breeding and Genetics, Punjab Agricultural University, Ludhiana, India and one hybrid viz. G-5414 from Field Fresh Agriculture Centre of Excellence, Ladhowal, Ludhiana were sown in 12″ earthen pots in the month of May during 2015 and 2016 in the screen house of Department of Plant Pathology, Punjab Agricultural University, Ludhiana, India.
Pathogen inoculation
Out of 25, twenty isolates of A. avenae subsp. avenae representing different genetic groups were used for pathogenicity testing. One-month-old plants of each cultivar were inoculated with all the 20 isolates of A. avenae subsp. avenae individually using whorl prick method described by Dhkal et al. (2016a). Two light pricks with hypodermic needle (0.70 × 32 mm size) were given at the base of the fully expanded leaves. One ml of bacterial suspension (108 cells/ml cfu) was poured into the pricked whorls. Inoculated plants were covered with polybags (12″ × 24″ size) for 48 h and high humidity was maintained by spraying water regularly inside the polybags.
Disease measurement
Disease development on leaves was recorded at 3, 5, 7, 9, 12 and 15 days after inoculation using 0–9 scale given by Pataky et al. (1997). The pathogenic variability of A. avenae subsp. avenae was established on the basis of disease score produced by these isolates on different hybrids and inbreds of maize. The tested isolates were further characterized into different groups. Data was analysed with PAST ver 2.5.1 software using Euclidean coefficient to generate dendrogram.
Results
Molecular characterization
To investigate the degree of genetic diversity among different isolates of A. avenae subsp. avenae, 25 RAPD primers were used. DNA fingerprinting of 25 isolates of A. avenae subsp. avenae collected from different maize growing area of Punjab, NW India showed a distinct banding pattern. All the 25 RAPD primers used in this study showed amplification with a total of 1769 amplified fragments and the number of scorable bands ranged from 35 to 109 across different primers. Among all, S112 showed maximum PIC value of 0.95 while OPT-4 showed minimum PIC value of 0.22 (Table 1). The primer OPT-16 amplified the maximum (109) number of fragments. The fragments thus obtained were amplified within a range of 50 bp to 3.9 kb. Seven isolates of A. avenae subsp. avenae produced one or more unique bands with respect to different primers (Table 2). Isolate Aaa2 could be differentiated by using five random primers whereas Aaa20 and Aaa25 isolates could be differentiated by using eight and nine random primers respectively. Three primers, S1114 (Fig. 1), OPT-1(Fig. 2) and OPT-18 (Fig. 3) produced distinguishable unique bands in Aaa2 (~3.9 kb), Aaa7 (500 bp) and Aaa21 (1250 bp) respectively. In Aaa25, S119 was observed to be highly polymorphic producing at least four repeatable unique bands that helps in differentiating this isolate from other isolates of A. avenae subsp. avenae (Fig. 4).
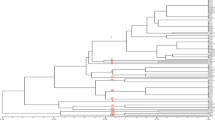
Based on the polymorphic data, these twenty five isolates of A. avenae subsp. avenae were grouped into four distinct genetic groups. Majority of isolates from Ludhiana district were grouped in Group 1 and Group 2 while isolates from Hoshiarpur district were mostly clustered in Group 4. The clustering of molecular data suggested that isolates belonging to the same region tends to group together with each other irrespective of their virulence or pathogenicity (Fig. 5).
Pathotypic characterization
Mean disease score of different isolates were found to be lowest on maize inbred CML-25 (4.5), while it was highest on the hybrid G-5414 (7.5). Among inbreds, highest mean disease score was recorded on CM-139 (Table 3). The isolate Aaa2, isolated from maize fields at PAU Ludhiana was found to be highly virulent as it produced maximum disease score on all maize inbreds and hybrids except CML-25. In contrast, Aaa17 isolated from Makhomajara village of Hoshiarpur district was the least virulent isolate producing mean disease score of 5.1 (Dhkal et al. 2015. Two isolates viz. Aaa7 and Aaa16 failed to produce any symptom on CM-600 and were designated as avirulent. Another isolate, Aaa17, was also found avirulent on CM-139 (Table 3). The present study revealed differential symptom expression of isolates particularly Aaa2 on different maize hybrids and inbreds (Fig. 6).
Based on their response to 20 isolates of A. avenae subsp. avenae, hybrids and inbred lines were divided into three classes viz. highly susceptible, moderately susceptible, and moderately resistant (Fig. 7, Table 4). The data plotted in dendrogram (Fig. 7) clustered moderately resistant lines viz. CM-600 and CML-25 in one group exhibiting mean disease score of 5.1 (Table 4). Six test cultivars viz. CM-139, LM-13, CM-140, FR-632, Punjab sweet corn-1 and PMH-2 showed moderately susceptible reaction to different isolates of A. avenae subsp. avenae with mean disease score of 6.55 which ranged from 6.8 to 6.2. However, two hybrids, G-5414 and PMH-1, were highly susceptible with mean disease score of 7.45 (Fig. 7, Table 4).
Level of susceptibility in test cultivars varied with respect to different isolates of A. avenae subsp. avenae. Inbred line LM-13 was found highly susceptible as 90% isolates were virulent on this line (Fig. 8). It was followed by hybrids G-5414 and PMH-1, which were also highly susceptible to 85% isolates tested. On the other hand, none of isolate was found highly virulent on CML-25 and the disease on this inbred line could not progress beyond a few small streaks.
Discussion
RAPD primers could be highly informative as they scan the whole genome, thus could be used in different studies to observe genetic diversity in population of various bacterial pathogens including Acidovorax species. The current study revealed that isolates of A. avenae subsp. avenae were genetically highly variable. Fontana et al. (2013) also used RAPD primers to study the intra-specific diversity in Acidovorax avenae. Using M13 primer, they differentiated four biotypes among thirty nine isolates. RAPD analysis suggested that the isolates from same geographical region tend to group together irrespective of their virulence spectrum. Similar results have been reported in other plant bacterial pathosystems (Kumar 2015; Lakshmi and Rabindran 2012; Shahrestani et al. 2012; Lore et al. 2011) where authors have used RAPD markers to study the variation in the population of Erwinia chrysanthemi pv. zeae and Xanthomonas oryzae pv. oryzae in different parts of the world. High level of genetic diversity was reported among different pathotypes of Xanthomonas oryzae pv. oryzae collected form Punjab, India using RAPD primers of OPA series (Hunjan 2012). In the present study, RAPD primers differentiated different field isolates of A. avenae subsp. avenae by generating unique amplicons. Similar results were obtained by Mäki-Valkam and Karjalainen (1994) in Erwinia carotovora subsp. atroseptica and Pseudomonas carotovorum.
RAPD primers have also been shown to give clear distinction of Erwinia carotovora subsp. carotovora from pectolytic Pseudomonas species (Parent et al. 1996). Furthermore, Erwinia carotovora subsp. atroseptica and subsp. carotovora could also be distinguished from each other using these markers. In the current study, RAPD primers also produced unique bands in different isolates of A. avenae subsp. avenae. S1114 was also observed to produce unique band in atleast one isolate, while S112 was highly polymorphic. This result is concordant with other study that established pathotypes of X. oryzae pv. oryzae could be differentiated by unique bands produced by different random primers of OPA series Hunjan (2012). He further observed that PBXo-1 pathotype can be differentiated by six different random primers whereas four primers namely S112, S 1114, S1115 and S1120 could differentiate pathotypes PBXo-2, PBXo-4, PBXo-6 and PBXo-179.
The cultivated maize is known to possess genetic resistance against different fungal and bacterial pathognes (Mir et al. 2018; Maschietto et al. 2017; Kumar et al. 2017; Chen et al. 2012; Sharma and Rai 2005; Chen et al. 2002). Although many workers (Kumar 2015; Sinha and Prasad 1975) have reported maize inbred lines resistant to pathogenic bacteria, however limited reports are available with respect to resistance sources against bacterial leaf streak. Dange (1972) identified CM-104, CM-105, CM-106, CM-109, CM-112, CM-201 and CM-300 inbreds possessing resistance to bacterial leaf streak during artificial screening. Our results in this study have confirmed that inbred lines resist A. avenae subsp. avenae more effectively then hybrids. In a parallel study, these inbred lines were also observed to exhibit elevated antioxidant enzyme activities against this pathogen as compared to maize hybrid varieties (Dhkal et al. 2016b).
This is the first report of genetic and pathotypic characterization of A. avenae subsp. avenae isolated from maize crop in India. Finally, the present work concluded that populations of A. avenae subsp. avenae in northern region of India are highly variable. The genetic structure in the pathogen can be further investigated for better understanding of the pathogen adaptation and devise suitable disease resistance breeding programs.
References
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Chen W, Wu J, Yuan H (2002) Identification of resistance on maize germplasm to maize ear rot. J Maize Sci 10:59–60
Chen J, Ding J, Li H, Li Z, Sun X, Li J, Wang R, Dai X, Dong H, Song W, Chen W, Xia Z, Wu J (2012) Detection and verification of quantitative trait loci for resistance to fusarium ear rot in maize. Mol Breed 30:1649–1656
Dange S. R. S., (1972). Studies on bacterial stripe disease of maize caused by Pseudomonas rubrilineans. Ph.D. Thesis. Postgraduate school. Indian Agriculture Research Institute, New Delhi, India
Dhkal M, Hunjan MS, Kaur H (2015) Phenotypic diversity in Acidovorax avenae subsp. avenae causing bacterial leaf streak of maize. Plant Disease Research 30:150–155
Dhkal M, Hunjan MS, Kaur H (2016a) Molecular identification of the pathogen causing bacterial leaf streak disease of maize and standardization of inoculation technique. Plant Disease Research 31:79–85
Dhkal M, Hunjan MS, Kaur H, Kaur R (2016b) Biochemical basis of bacterial leaf streak resistance in maize. Indian Phytopathol 69:373–380
FAOSTAT, (2014). Statistical databases and data-sets of the Food and Agriculture Organization of the United Nations. http://www.fao.org/faostat/en/#data
Fontana DP, Rago AM, Fontana CA, Vignolo GM, Cocconcelli PS, Mariotti JA (2013) Isolation and genetic characterization of Acidovorax avenae from red stripe infected sugarcane in northwestern Argentina. Eur J Plant Pathol 137:525–534
Hunjan M. S., (2012). Pathogenic and molecular characterization of Xanthomonas oryzae pv. oryzae [(Ishiyama) Swings et al.] causing bacterial blight of rice. PhD Thesis, Punjab Agricultural University, India
Johnson AG, Robert AL, Cash L (1945) Further studies of bacterial leaf blight and stalk rot of corn. Phytopathology 35:486 (Abstract)
Kaur H, Mohan C, Vikal Y, Singh M (2014) Pathogenic and molecular characterization of Fusarium moniliforme Sheld, the incitant of fusarium maize stalk rot in the Punjab state of India. Maydica 59:290–297
Kumar A., (2015). Genetic diversity of Erwinia chrysanthemi pv. zeae causing bacterial stalk rot of maize and its management. PhD Thesis, Punjab Agricultural University, India
Kumar A, Hunjan MS, Kaur H, Dhillon HK, Singh PP (2017) Biochemical responses associated with resistance to bacterial stalk rot caused by Dickeya zeae in maize. J Phytopathol 165:822–832
Lakshmi DL, Rabindran R (2012) Molecular variability in Xanthomonas oryzae pv. oryzae isolates of Tamil Nadu. Indian J Plant Protect 40:123–128
Lore JS, Vikal Y, Hunjan MS, Goel RK, Bharaj TS, Raina GL (2011) Genotypic and pathotypic diversity of Xanthomonas oryzae pv. oryzae, the cause of bacterial blight of rice in Punjab state of India. J Phytopathol 159:479–487
Mäki-Valkam T, Karjalainen R (1994) Differentiation of Erwinia carotovora subsp. atroseptica and carotovora by RAPD-PCR. Ann Appl Biol 125:301–309
Maschietto V, Colombi C, Pirona R, Pea G, Strozzi F, Marocco A, Rossini L, Lanubile A (2017) QTL mapping and candidate genes for resistance to fusarium ear rot and fumonisin contamination in maize. BMC Plant Biol 17:20. https://doi.org/10.1186/s12870-017-0970-1
Mir ZR, Singh PK, Zaidi PH, Vinayan MT, Sharma SS, Krishna MK, Vemula AK, Rathore A, Nair SK (2018) Genetic analysis of resistance to post flowering stalk rot in tropical germplasm of maize (Zea mays L.). Crop Prot 106:42–49
Parent JG, Lacroix M, Page D, Vezina L (1996) Identification of Erwinia carotovora from soft rot diseased plants by random amplified polymorphic DNA (RAPD) analysis. Plant Dis 80:494–499
Pataky JK, Toit du LJ, Kerns MR (1997) Bacterial leaf blight on shrunken-2 sweet corn. Plant Dis 81:1293–1298
Saiki RK, Gelfand DH, Stoffel S, Scharf SJ, Higuchi R, Horn GT, Mullis KB, Erlich HA (1988) Primer directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 239:487–491
Shahrestani AT, Kazempour MN, Ebadie AA, Elahinia SA (2012) Genetic diversity of Xanthomonas oryzae pv. oryzae in rice fields of Guilan Province (Iran) using RAPD markers. Agric Trop Subtrop 45:60–65
Sharma RC, Rai SN (2005) Evaluation of maize inbred lines for resistance to maydis leaf blight. Indian Phytopathol 58:339–340
Shrestha RK, Rosenberg T, Makarovsky D, Eckshtain-Levi N, Zelinger E, Kopelowits J, Sikorski J, Burdman S (2013) Phenotypic variation in the plant pathogenic bacterium Acidovorax citrulli. PLoS One 8:e73189. https://doi.org/10.1371/journal.pone.0073189
Silva KMM, Xavier AS, Gama MAS, Lima NB, Lyra MCCP, Mariano RLR, Souza EB (2016) Polyphasic analysis of Acidovorax citrulli strains from northeastern Brazil. Sci Agric 73:252–259
Sinha SK, Prasad M (1975) Varietal screening of maize germplasm against stalk rot pathogen Erwinia carotovora f. Sp. zeae. Labdev J Sci Technol 13:128–133
Smith DR, White DG (1988) Diseases of corn. In: Sprague GF, Dudley JW (eds) Corn and corn improvement. American Society of Agronomy, Madison, pp 687–766
Techati N (2008) Identification and genetic diversity of bacterial leaf streak of corn. Kasetasrt University, Thailand, MSc Thesis
Thind BS (2012) Phytopathogenic Procaryotes and plant diseases. Scientific Publishers, Jodhpur, India
Ullasa BA, Mehta YR, Payak MM (1967) Xanthomonas rubrilineans on Zea mays in India. Indian Phytopathol 20:77–78
Willems A, Goor M, Thielemans S, Gillis M, Kersters K, De Ley J (1992) Transfer of several phytopathogenic Pseudomonas species to Acidovorax as Acidovorax avenae subsp. avenae subsp. nov., Comb.nov., Acidovorax avenae subsp. citrulli, Acidovorax avenae subsp. cattleyae, and Acidovorax konjaci. Int J Syst Bacteriol 42:107–119
Zivanovic M, Walcott RR (2017) Further characterization of genetically distinct groups of Acidovorax citrulli strains. Phytopathology 107:29–35
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dhkal, M., Hunjan, M.S., Kaur, H. et al. Characterization of Acidovorax avenae subsp. avenae causing bacterial leaf streak of maize in Punjab state of India. J Plant Pathol 101, 71–79 (2019). https://doi.org/10.1007/s42161-018-0138-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42161-018-0138-3