Abstract
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has caused a worldwide pandemic since 2019, spreading rapidly and posing a significant threat to human health and life. With over 6 billion confirmed cases of the virus, the need for effective therapeutic drugs has become more urgent than ever before. RNA-dependent RNA polymerase (RdRp) is crucial in viral replication and transcription, catalysing viral RNA synthesis and serving as a promising therapeutic target for developing antiviral drugs. In this article, we explore the inhibition of RdRp as a potential treatment for viral diseases, analysing the structural information of RdRp in virus proliferation and summarizing the reported inhibitors’ pharmacophore features and structure–activity relationship profiles. We hope that the information provided by this review will aid in structure-based drug design and aid in the global fight against SARS-CoV-2 infection.
Graphical Abstract

Highlights
-
Exploring the potential clinical targets for attenuating coronavirus disease 2019 (COVID-19) by structure-based drug designing of RdRp inhibitors.
-
RdRp catalytic site and druggable cavities predictions of SARS-CoV-2.
-
Pharmacophoric features and structure–activity relationship analysis of different repurposed therapeutic drugs for COVID-19 against RdRp of SARS-CoV-2.
-
Current treatments are logistically challenging, increasing the need for safe and effective oral therapies.
-
Ongoing global efforts to prevent the spread of COVID-19 disease and the current status of SARS-CoV-2.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
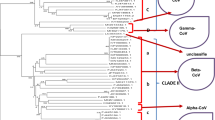
Coronaviruses (CoVs) are enveloped positive-sense RNA viruses that can cause respiratory infections ranging from mild to severe in humans and various animals. In December 2019, a few cases of atypical pneumonia, later identified as coronavirus disease 2019 (COVID-19), were first reported in Wuhan, caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). It is believed that the virus initially jumped from an infected animal to humans during the first week [1, 2]. Upon further examination, the etiological agent was identified as an RNA virus with high genetic homology to both the severe acute respiratory (SARS-CoV) and Middle East respiratory syndrome (MERS-CoV) viruses [3, 4] (Fig. 1).
The ongoing COVID-19 pandemic has issued a grave public health warning, placing millions of people at risk in multiple countries [5]. As of February 19, 2023, the WHO has reported more than 756,581,850 confirmed cases and 6,844,267 deaths [6]. Despite this, there is currently no explicit medication for COVID-19, and the selection of therapeutic drugs is primarily based on experiences treating SARS or other influenza viruses. As such, there is an urgent need to develop safe and effective antiviral medications.
It is worth noting that current medications are explicitly designed for a virus's genome, and the physical structure of SARS-CoV-2 may change over time, potentially limiting their in vivo efficacy. However, some non-structural or accessory proteins may serve as promising molecular targets for developing therapeutic drugs. Rather than directly targeting viral replication, some therapeutic strategies can modify host immune responses to combat the virus or inhibit the inflammatory response generated during the viral invasion, ultimately reducing the physiological response to sickness. The replication of the SARS-CoV-2 genome and its gene transcription is primarily controlled by the viral RdRp, which has been identified as a promising target for designing novel antiviral strategies. Remdesivir, a nucleoside analog, has shown antiviral activity in cell culture and animal models against the RdRp of multiple coronaviruses [7, 8], showing antiviral activity in cell culture and animal models [9]. It is covalently incorporated into the primer strand at the first replicated base pair when a partial double-stranded RNA template is inserted into the central channel of the RdRp and can terminate its chain elongation [10]. Other potential medications with evident anti-SARS-CoV-2 activity in either cellular studies or clinical trials include EIDD-2801, Sofosbuvir, Galidesivir, AT-527, IDX-184, Favipiravir, and Ribavirin [11].
2 Structural Organization of the SARS-CoV-2 Genome
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2; betacoronavirus), a member of the Coronaviridae family, has a genome of 30 kb of single-stranded positive-sense RNA [12, 13], and its pharmacological treatments target both the genome and structure of the virus (Fig. 2). Upon invading the host cell, the viral genome, comprising 14 open reading frames (ORFs), is liberated into the cytoplasm for replication and transcription. ORFs 1a and 1b encode two replicase polyproteins, PP1a and PP1ab, further subdivided into non-structural proteins [14].
The SARS-CoV-2 genome structural organization: it encodes two large ORF1a and ORF1b genes, which encode non-structural proteins that form a replication–transcription complex. The non-structural protein 3 (nsp3) encodes for papain-like protease (PLP), and non-structural protein 5 (nsp5) encodes for 3C-like (3CL) protease
The SARS-CoV-2 virus and its various strains, including the AY.4.2, Delta and Omicron variants, have caused significant disruptions worldwide. Mutations on the spike (S)-gene, which codes for the spike (S) protein, appear to increase transmissibility by binding the S protein to the angiotensin-converting enzyme 2 (ACE2) receptor [15,16,17]. Unfortunately, treatment options for SARS-CoV-2 are currently limited and cannot keep up with the virus’s mutation rate. However, drug design has identified seven primary targets during the search, including the membrane protein, envelope protein, nucleocapsid protein, spike protein, protease, helicase, hemagglutinin esterase and non-structural proteins 16NSPs [18,19,20]. Of these targets, RdRp, also known as nsp12, is a critical protease in the virus’s life cycle with conserved catalytic sites [21]. RdRp forms a replicase complex with other non-structural proteins, such as nsp7 and nsp8, to facilitate viral RNA genome replication and transcription [22]. Notably, the RdRp of SARS-CoV-2 shares 96% sequence similarity with SARS-CoV, indicating the conserved nature of RdRp in coronaviruses [23, 24].
3 Currently Available Therapeutics and Treatment Options for SARS-CoV-2
Current SARS-CoV-2 outbreaks have prompted investment in ongoing studies to exploit various viral target proteins for therapy through five typical mechanisms as follows, but strategies aimed at blocking the viral proteins, as in drug and vaccine development, have primarily failed [25]. It has been shown that SARS-CoV-2 undergoes rapid recombination to generate new strains of altered virulence to escape the host antiviral defensive system, which lead to severe cytokine storms.
-
1.
Enhance a pharmacological immune response (enhance immune function).
-
2.
Destroy the virus, and act on the pathogen itself.
-
3.
Restrict virus entry in the cell (block host cell-binding proteins or virus-binding proteins).
-
4.
Impede virus replication.
-
5.
Treat symptoms.
The first choice is considered the best for the patient since it is potentially long-lasting and "friendly," and may also have preventive properties against subsequent viral attacks. Antiviral medications such as ritonavir, lopinavir, oseltamivir, ribavirin and ganciclovir were tested in several investigations to decrease infection rate and the risk of respiratory problems [26,27,28]. In general, every enzyme and protein involved in the replication of the SARS-CoV-2 virus and the regulation of host cellular machinery is a promising druggable target in the search for treatment options [29]. In this perspective, we summarized the primary host-based and virus-based targets from a structural point of view and information about reported therapeutic drugs with efficacy against various SARS-CoV-2 target proteins (Fig. 3). However, randomized controlled studies are needed to establish the effectiveness of these antiviral medicines against SARS-CoV-2. Therefore, in review, we had the more recent information available from the literatures on biocidal agents, immunomodulatory drugs of SARS-CoV-2 and the clinical characteristics of SARS-CoV-2. We concluded with a possible therapeutic plan against SARS-CoV-2.
4 The Active Cavities of RNA-Dependent RNA Polymerase in Coronavirus
RdRp is a crucial therapeutic target since it is essential for RNA genome replication, and the host does not have a functional protein that can perform the same function. RdRp is considered a promising target in the development and discovery of drugs because of its lack of an analogue in mammalian cells, and its blockage is not anticipated to have target-related adverse effects. The 3-D crystal structure of RdRp of SARS-CoV-2 (PDB ID: 7BTF) was obtained from the protein database with a resolution of 2.9 Å [30, 31]. The catalytic sites of RdRp were predicted using the CASTp server [32] containing four potential drug-binding cavities (Fig. 4). The CASTp web server provides a complete and detailed analysis of surface pockets and protein cavities, essential in judging docking models’ reliability.
A deep and wide groove domain in the core structure of SARS-CoV-2 RdRp depicts a cupped right hand interlinked by “thumb”, “finger” and “palm” subdomains [33]. The thumb subdomain (residues 819–920) forms a tight circle with the finger subdomain (residues 397–581 and 62–679) in the nsp12 [34, 35]. The palm domain, the most conserved domain, contains five of the seven conventional RdRp catalytic motifs (A–E), whereas the finger domains include the remaining two (F and G). The RdRp core, which is structurally conserved, and associated motifs are crucial for viral RdRp catalytic function and could therefore be targeted for therapeutic intervention [36]. The nsp12 (RdRp) is a multi-subunit with a molecular weight of 106 kDa, much larger than other (+) RNA viruses. The nsp12, nsp7 and nsp8 form a 160 kDa replicase complex responsible for the replication and transcription of the viral RNA genome [37]. Since the RdRp enzyme plays a role in replicating the new coronavirus by synthesizing RNA, RdRp has become an attractive therapeutic target for drug research.
5 The Potential Antiviral Drugs Inhibit Viral Replication by Targeting RdRp
This review provides an overview of the potential use of existing antiviral drugs as enzyme inhibitors to repurpose them as effective candidates for inhibiting SARS-CoV-2 polymerase. Notably, coronavirus RdRp screening methods are more complex than those for other viral target proteins, such as proteases [62]. Despite increasing interest in current knowledge, only a few studies have reported on RdRp screening for coronaviruses. This study describes the docking analysis and crystal structures of reported RdRp inhibitors binding with RdRp to elaborate on the structure–activity relationship of the existing drug. Several antiviral compounds have been screened to determine how well-repurposed nucleotide analogues interact with the SARS-CoV-2 RdRp active sites. In vitro research and in silico analysis indicated that some broad-spectrum antiviral medications could potentially be therapeutic platforms against SARS-CoV-2. Depending on their structure and binding affinity, nucleotide analogues mimic the natural substrates of the SARS-CoV-2 RdRp and cause either rapid or gradual chain termination [66, 67]. Moreover, several nucleotide analogues are prodrugs that must undergo phosphorylation inside the cell to have an antiviral effect. In various circumstances, numerous host enzymes carry out intracellular phosphorylation, converting the prodrug into the bioactive triphosphate forms of these drugs [68, 69].
5.1 Favipiravir
Favipiravir (1, Table 1) is an antiviral agent serving as an inhibitor for RdRp in intracellular phosphoribosylation active form, i.e., favipiravir ribofuranosyl-5′-triphosphate (favipiravir-RTP) (Fig. 5). The 6-fluoro-3-hydroxypyrazine-2-carboxamide [favipiravir (FPV)] is a purine nucleotide analogue produced by Toyama Chemicals in Japan to treat influenza and viral infections of the nose and throat [70]. FPV has been shown to have antiviral properties in vitro and in vivo against the influenza viruses (A, B and C) and Lassa, Ebola and other viruses. Although FPV has not yet been approved by the US Food and Drug Administration (FDA), it has recently been examined and found to be a promising therapeutic approach for coronavirus disease 2019 (COVID-19) [40]. The investigation of FPV as a therapeutic for COVID-19 revealed that, although it did not significantly decrease RNA production in vitro, it may still be effectively integrated into RNA products to reduce the precision of RNA synthesis. Like other common nucleotide triphosphate (NTP) substrates, FPV effectively enters RNA products. These findings imply that this drug may have broad-spectrum antiviral efficiency by similar mechanisms against a panel of several RNA viruses. In cell-based experiments, administration of FPV was demonstrated to cause mutations in the progeny virion of SARS-CoV-2, indicating that this drug can evade the coronaviruses’ proofreading mechanism to exhibit antiviral activity through many rounds of RNA synthesis [71].
The reported docking analysis of FPV demonstrated significant binding interactions with crucial amino acid residues of the target receptors [72]. The literature revealed that FPV prodrug showed less critical interaction with the predicted active site of target receptor RdRp (PDB ID: 7BV2) of SARS-CoV-2 (−5.336 kcal/mol) than its active triphosphate metabolite (− 8.951 kcal/mol). Interestingly, docked results showed that FPV prodrug does not form the salt bridge of Mg2+ with the first ribofuranose-linked phosphate group. Reported structure–activity relationship (SAR) analysis of FPV revealed that, for RdRp inhibition, it must involved salt bridge formation of the first ribofuranose-linked phosphate group with the Mg2+ and also show interaction with crucial amino acid residues HIS439, LYS545, ILE548, SER549, ARG553, ARG555, VAL557, LYS621, CYS622, SER682, ASN691, and ASP760. Furthermore, hydrogen bond formation with uracil (U20) in the is crucial for inhibiting viral replication (Fig. 6) [73, 74].
Naydenova et al. create a cryo-electron microscopy (EM) crystal structure of antiviral drug FPV with SARS-CoV-2 RdRp (PDB ID: 7AAP at a resolution of 2.50 Å). The active triphosphate metabolite of FPV-RTP was used by Naydenova et al. to analyse the structure of RdRp of SARS-CoV-2 co-crystallized with FPV using cryo-EM techniques [75]. The cryo-EM structure shows that FPV-RTP terminates chain extension by pairing with the template strand to + 1 nucleotide through noncovalent contacts. FPV-RTP also interacts with ASN691, uracil (U20), LYS545, cytosine (C10) and SER814 through conventional hydrogen bonds (Fig. 7).
FPV showed protective effects against various RNA viral infections in animal models, although it had a low in vitro selectivity against SARS-CoV-2 [half maximal effective concentration (EC50) 61.88 µM], indicating that further in vivo research on this medication against SARS-CoV-2 may be beneficial. Clinical trial data from phases I, II and III showed that FPV had good overall efficacy. It causes chain termination and viral mutagenesis by inhibiting the error-prone viral RdRp [76]. Most patients infected with SARS-CoV-2 (80–85%) have mild to severe sickness and continue to disseminate the virus. FPV used to treat mild to moderate cases can potentially lessen the impact of the continuing epidemic. FPV’s most important side effect was teratogenicity, followed by abnormal liver enzymes, mental symptoms, intestinal symptoms and elevation in serum uric acid in a randomized test of FPV in 241 patients with SARS-CoV-2 [77].
5.2 Ribavirin
Ribavirin (2, Table 1) is a guanosine (purine) analogue that can inhibit viral RNA synthesis. It was first registered in the 1980s and clinically used with interferon for hepatitis B and C. It is also used for viral diseases such as haemorrhagic fever and respiratory syncytial viral pneumonia [78, 79]. The adenosine kinase catalyses the phosphorylation of ribavirin to produce ribavirin monophosphate (RMP), and then nucleoside mono- and di-phosphate kinases catalyse RMP into ribavirin triphosphate (RTP) [80]. Unal et al. reported that the docking analysis of ribavirin against RdRp (PDB ID: 6M71) formed four conventional hydrogen bonds with TRP617, ASP761, SER814 and HIS810 with a binding energy of −6.2 kcal/mol (Fig. 8) [81]. The docking analysis of RMP predicts that it forms a hydrogen bond with crucial amino acid residues ASP544, ASN612, THR601 and ARG476 of RdRp of SARS-CoV-2. Unexpectedly, RMP does not bind the complementary strand nucleotide as effectively and tends to act through a distinct mechanism than other known medications [82, 83]. Elfiky investigated that ribavirin strongly inhibits RdRp (PDB ID: 6NUR) activity with binding affinity −7.8 kcal/mol by interacting with TRP508, TYR510, LYS512, CYS513, ASP514, ASN582 and ASP651 residues of the receptor [84]. Another study showed the binding analysis of RTP with RdRp (PDB ID: 7BV2) of SARS-CoV-2. The analysts predict that RTP is a more potent inhibitor than its prodrug, with a binding affinity of −9.280 kcal/mol (Fig. 8) [73].
Triphosphate moiety is essential for its inhibitory activity, as it involves a nucleotide analogue, RTP, which may hinder the activity of RdRp in three different ways: (1) by competing with the enzyme’s natural substrate without itself serving as a substrate, (2) by acting as an alternative substrate for the polymerase and resulting chain termination or (3) by acting as an alternative substrate for the polymerase without doing chain termination. If RTP just interacted with RNA polymerase as in the previous two situations, viral replication would be instantly inhibited, leading to the breakup of the viral RNA chain [85]. Li et al. create the cryo-EM structure of the drug with PDB ID: 7DFH at a resolution of 2.97 Å [30]. The cryo-EM structure shows that ribavirin-RTP terminates chain extension by pairing with RNA template strand to +1 nucleotide through covalent interaction with guanine (G20) and interacting with two metal ions (Mg2+1005 and Mg2+1004) through attractive charges (Fig. 9).
Ribavirin was tested against SARS-CoV in 2003 and then used therapeutically in conjunction with interferon and corticosteroids; nevertheless, the results of the antiviral effect were inadequate [86]. Commercially, ribavirin is marketed as an oral solution, an oral dosage form capsule and an inhalation formulation. Ribavirin has not been tested in the inhaled dosage when used in the treatment against SARS-CoV, MERS-CoV and SARS-CoV-2. Moreover, the injectable dose is not marketed in the USA [87, 88]. Additionally, in two minor investigations for SARS, oral ribavirin was administered at a 4.0 g dosage followed by 1.2 g for 8 h. Information on the treatment of COVID-19 is currently restricted to studies that employ a combined therapy that includes 400 mg taken twice daily for 14 days [89]. The longer half-life of ribavirin is about 40 days. Increasing doses might make the better condition more quickly achievable. The most frequent side effects of ribavirin treatment for SARS are hypocalcemia and haemolytic anaemia. The probability of adverse side effects is more significant in patients with poor renal function and older age [90, 91].
Furthermore, patients suffering from prior cardiovascular problems should also be considered because haemoglobin lowering raises the risk of exacerbations [92, 93]. Since the metabolism and excretion of ribavirin do not depend on the CYPP450 metabolism pathway, it does not interact with other medications. However, owing to the increasing side effects and toxicity caused by ribavirin, some other treatments should be used with caution [94].
5.3 4′-Cytosine Nucleoside Analogues
A series of 4′-cytosine nucleoside analogues investigated based on their pharmacological and pharmacokinetic characteristics by Janssen, including 4′-chloromethyl-2′-deoxy-2′-fluorocytidine (ALS8112, 3, Table 1); 3′,5′-di-O-isobutyryl (ALS-8176, 4, Table 1); and 4′-azido-2′-deoxy-2′-C-methylcytidine (5, Table 1), was shown to have the most promising inhibiting activity against RSV polymerase and NS5B polymerase, respectively [43,44,45].
ALS-8112 specifically inhibited rhabdoviruses and paramyxoviruses, and ALS-8176 suppressed RSV multiplication in non-human primates, while ALS-8112 also inhibited all RSV strains in vitro. The active 5′-triphosphate metabolite of ALS-8112 formed inside the cell and inhibited the viral RNA polymerase. The literature showed that anti-RSV of 4′-cytosine nucleoside analogues impede the replication of SARS-CoV-2 up to 20 micromolar concentrations in Vero cells [95]. The docking analysis of 4′-cytosine nucleoside analogues against RdRp (PDB ID: 7AAP) of SARS-CoV-2 showed that ALS8112 has less inhibitory potential with a binding affinity of −6.11 kcal/mol (Fig. 10) [96] than its active metabolite ALS8176. The in silico screening of ALS8176 against RdRp (PDB ID: 6M71) of SARS-CoV-2 exhibits a binding affinity of −6.9 kcal/mol (Fig. 10) [97]. Another study investigated the structure–activity relationship of lumicitabine (ALS8176), an oral version of the parent drug ALS8112. In ALS8176, the 3′ and 5′ ester groups are essential for inhibitory activity, forming a hydrogen bond with ASP711 and exhibiting hydrophobic interaction with THR710. The 4-chloromethyl moiety of ALS8176 is a crucial substituent for polymerase specificity and is efficiently recognized by RdRp resulting in chain termination. The docking analysis showed that 4-chloromethyl moiety is involved in attractive charge interaction with LYS47 of receptor protein. The literature study showed that the incorporation of the ALS8176 into the extended RNA primer caused immediate chain termination due to the inability to incorporate subsequent nucleotide into the growing chain [98].
ALS-8112 chose the RNA polymerase-encoding domain of the L gene of RSV for mutations associated with resistance. In biochemical experiments, the recombinant RSV polymerase complex detected the 5′-triphosphate metabolite with high specificity, terminating the process of RNA production. While structurally comparable compounds showed dual RSV/HCV suppression, 5′-triphosphate derivatives did not affect host or viral polymerases, including those from unrelated viruses like hepatitis C. A comparative analysis of RdRp inhibitors by Dweipayan showed that the binding affinity of 4′-azido-2′-deoxy-2′-C-methylcytidine is −4.88 kcal/mol (Fig. 10) [96]. The substantial decrease in binding affinity is due to substituting the azido group instead of halide. The potent nucleoside inhibitor effectively inhibits the NS5B polymerase, i.e., 4′-azido-2′-deoxy-2′-C-methylcytidine, which has a 1.2 mM EC50 value and a high to moderate in vivo efficacy in rats [99].
5.4 2′-C-Methylcytidine (NM 107) and Derivative (Valopicitabine)
The 2′-C-methylcytidine (NM 107, 6, Table 1) is a potent inhibitor of the HCV RNA-dependent RNA polymerase [100]. Several other RNA viruses have been inhibited by it, including pestivirus bovine virus, HCV and flaviviruses, notably West Nile virus, yellow fever virus and dengue-2 virus. The SAR studies show that 2′-C-methylcytidine effectively prevents hepatitis E virus (HEV) replication, making it a promising option for anti-HEV medication development [101, 102]. The research on pyrimidine nucleotide inhibitors as prospective antiviral medications showed that certain nucleotide substituents at the nucleoside’s C2′ and C4′ positions demonstrate blatant therapeutic potential [103, 104]. As virus RNA strands develop, the insertion of 2′-C-modified monophosphates onto their 3′ terminus of the RNA strand encourages the termination of RNA strands’ elongation due to steric restriction between the synthetic 2′-C-group of unnatural and entering natural nucleotide inhibitor [105]. In 2020, several antiviral drugs and 2′-C-methylcytidine under clinical trials for other viral RdRp were computationally investigated against RdRp of SARS-CoV-2 by Elfiky. He used eight different conformations of RdRp (PDB: 7BTF) of SARS-CoV-2 as targets for small antiviral drug molecules. The docking analysis showed that 2′-C-methylcytidine (NM 107) is a potent inhibitor of RdRp of SARS-CoV-2 with a binding affinity of −7.31 kcal/mol [106]. Moreover, the comparative docking analysis of RdRp inhibitors of SARS-CoV-2 showed that 2′-C-methylcytidine phosphorylated to form 2′-C-methylcytidine triphosphate. The 2′-C-methylcytidine triphosphate prevents the virus’s RNA chain elongation and inhibits the RdRp (PDB ID: 7AAP) activity with a binding affinity of −7.19 kcal/mol. Another study showed that 2′-C-methylcytidine inhibits the activity of RdRp (PDB ID: 7BV2) by forming a hydrogen bond with THR556, ASP760, ASP623 and TYR619 of the predicted target site of the receptor (Fig. 11) [107].
Moreover, the structure–activity relationship analysis of valopicitabine, the 3′-O-l-valinyl ester of NM-107, inhibits the activity of RdRp (PDB ID: 7BV2) by forming a hydrogen bond with ARG553, THR556, TYR619 and ASP623 residues. It also shows hydrophobic interaction with LYS714 [107]. According to pharmacokinetic investigations, this novel therapeutic agent has 2′-C-methylcytidine and low oral bioavailability. The 3′-O-l-valinyl ester derivative valopicitabine was created to get over this restriction. Valopicitabine (7, Table 1), the 3′-O-valinyl ester of NM-107, was designed to generate a substance with higher absorption and bioavailability than its parent molecule. Valopicitabine is an acid-stable prodrug with outstanding physicochemical and toxicokinetic properties [108, 109]. Recently a phase III clinical trial for the prodrug valopicitabine (NM283) has been started. It has an EC50 value of 0.67 µM, which might prevent the viral chain elongation and inhibit the function of viral RdRp [110]. Valopicitabine is a more potent inhibitor of RdRp with improved physicochemical and pharmacokinetic profile compared with its parent compound, 2′-C-methylcytidine. The toxicokinetic analysis has shown that valopicitabine (NM283) is an acid-stable prodrug of NM107 with excellent toxicokinetic profiles [111].
5.5 NHC (EIDD-1931) and Molnupiravir (EIDD-2801)
The cytidine deaminase can convert cytidine analogues into cytidine and uridine triphosphates. NHC, EIDD-1931 (8, Table 1) is a β-D-N4-hydroxycytidine orally active ribonucleoside derivative with vast therapeutic potential against a variety of RNA viruses, notably MERS-CoV (EC50 0.56 mM) and SARS-CoV-2 (IC50 0.30 mM). NHC functions as a weak substitute substrate for cytidine triphosphate (CTP) to significantly influence viral replication processes such as translation, replication and encapsulation that depend on these structures [112].
EIDD-1931 is a ribonucleoside analogue that induces RNA viral mutations and has recently been proposed as a COVID-19 treatment option. As the little cytotoxic effect of this drug was seen, NHC displayed a positive effectiveness and safety profile [113, 114]. NHC is a potential candidate in phase II clinical studies to treat symptomatic adolescent outpatients and freshly hospitalized patients with COVID-19. Additionally, preclinical research is being done to assess its potential as a therapy for MERS-CoV infection. Furthermore, EIDD-2801 improved respiratory performance and decreased viral intensity in mice infected with MERS-CoV and SARS-CoV [115]. Considering its physicochemical profile, oral or parenteral option, and veridical, this active ingredient may treat COVID-19 [116, 117].
Molnupiravir, β-D-N4-deoxycytidine, rapidly cleaves to EIDD-1931 in plasma and is then distributed to various organs, where it is converted to 5′-triphosphate (Fig. 12). The virally encoded RdRp uses EIDD-1931 5′-triphosphate as a substrate, and when it integrates into the growing RNA chain, it results in a catastrophic replication error [118, 119]. In the plasma. the host’s esterase rapidly breaks down molnupiravir into EIDD-19311, which is afterwards changed into molnupiravir triphosphate by the host’s kinase once it enters the intracellular space. Molnupiravir triphosphate serves as an active antiviral compound and strongly inhibits RdRp.
The 5′-isopropyl ester of NHC, molnupiravir (9, Table 1), has a wide range of anti-influenza and anti-multiple coronavirus properties. The docking analysis showed that EIDD-1931 formed hydrogen bonds with ASN781 and LYS545, whereas the 5′-isopropyl ester derivative (EIDD-2801) of EIDD-1931 showed strong RdRp (PDB ID: 7BV2) inhibitory activity by forming an H-bond with LYS545, SER759 and uracil (U10) with the binding energy of −6.49 kcal/mol [115, 120]. The structure–activity relationship of molnupiravir triphosphate exhibits strong interaction with RdRp (PDB ID: 7BV2) of SARS-CoV-2 by forming conventional hydrogen bonds with ARG555, U10 and U20. It shows a binding affinity of −8.39 kcal/mol by forming an attractive charge interaction with ARG555, unfavourable attraction with Mg2+1004, hydrophobic interaction with Mg2+1005 and pi–pi stacked interaction with uracil (U20). Moreover, molnupiravir triphosphate also interacts with RdRp of the Delta AY.4 subvariant of SARS-CoV-2 through various types of interaction with crucial amino acid residues (CYS622, TYR622, LYS62, TRP617, ASP761 and U20) with a binding energy of −10.28 kcal/mol [121] (Fig. 13).
Molnupiravir was investigated for mutagenicity in two rodent models in vivo. Based on available genotoxicity evidence and a 72-h treatment period, the FDA suggested that molnupiravir had a low risk of cytotoxicity. Furthermore, treatment with molnupiravir entirely prevented virus transmission to untreated rodents, indicating that early treatment molnupiravir may be able to stop secondary SARS-CoV-2 spread [122, 123].
5.6 Galidesivir
Galidesivir (10, Table 1) is a wide-ranging antiviral drug that has the potential to cure COVID-19 and is well tolerated and effective in phase I investigations in healthy volunteers. Galidesivir is an adenosine nucleoside precursor that prevents viral RNA polymerase. Galidesivir phosphorylated into a triphosphate can resemble the intracellular kinases ATP to treat HCV [124, 125]. The drug’s monophosphate nucleotide is incorporated by viral RNA polymerases into the developing RNA sequence, leading to faster chain termination and inhibiting the function of viral RdRp. It is undergoing a phase II human trial for treating coronavirus in Brazil and other countries [126, 127].
Galidesivir has shown broad-spectrum action in vitro with EC50 ranging from 3 to 68 μM against approximately 20 RNA viruses in nine distinct categories, including flaviviruses, paramyxoviruses, bunyaviruses, arenaviruses and coronaviruses [15, 128]. The docking studies suggest that, compared with ATP (adenosine triphosphate), the active metabolite galidesivir triphosphate demonstrated a greater affinity for SARS-CoV-2 RdRp. The metal ions in the nsp12 structure also consist of two pieces of Zn2+ and Mg2+. ATP forms a salt bridge with Mg2+ to bind with it and activate RNA polymerase [129]. The prodrug galidesivir, according to molecular docking results, creates a salt bridge with Mg2+. It has been found that the interaction of Mg2+ with RdRp inhibitors is essential to connect with the RNA strand. The predicted binding energy for galidesivir is −6.187 kcal/mol, while galidesivir triphosphate showed a binding affinity of −8.994 kcal/mol. Another study showed the inhibitory potential of galidesivir against SARS-CoV-2 RdRp (PDB ID: 6M71) with interaction affinity of −6.81 kcal/mol [130] by forming hydrogen bonds with ASP36, ASP221 and THR206, and various other interactions with crucial amino acids residue of target binding sites (Fig. 14). Moreover, the structure–activity analysis showed that galidesivir triphosphate exhibits more inhibitor activity against RdRp (PDB ID: 7BV2) by forming an attractive charge interaction with Mg2+1005, U20 and ARG553. It also forms a hydrogen bond with ARG555 (Fig. 14) [73]. More clinical and in vitro research on galidesivir is needed [131].
5.7 Remdesivir
Remdesivir (11, Table 1) is a promising small-molecule antiviral medication that has proved effective for RNA viruses from various groups, such as Coronaviridae (MERS-CoV, SARS-CoV and others). Remdesivir, a wide-ranging antiviral drug initially developed to combat Ebola, was produced by Gilead Sciences. It can suppress viral replication by inhibiting the virus’s ability to proliferate by stopping RNA transcription prematurely [132]. However, clinical studies of remdesivir showed that it could significantly reduce SARS-CoV-2 infection in Vero cells during an in vitro trial [133]. Early injection of remdesivir reduced the pulmonary infiltrates in a rhesus macaque model of COVID-19. The replication of SARS-CoV-2 in human nasal and bronchial airway epithelial cells was also reported to be significantly inhibited by remdesivir [134]. These results encouraged its usage in SARS-CoV-2 infection in the absence of other therapeutic therapies [135]. Remdesivir is an adenosine analogue prodrug that effectively inhibits viral RNA polymerases when intracellularly metabolized to an ATP analogue [136]. We can see from the drug’s clinical trials that the drug’s effect on the target RdRp is relatively stable [137].
The structure–activity analysis showed that remdesivir is a prodrug transformed inside cells into triphosphate (RTP, active form) (Fig. 15). Gilead Sciences created monophosphorylated remdesivir to have greater cellular absorption in vivo. Exoribonuclease (ExoN) proofreading efficiency is interfered with, and RdRp polymerization activity is inhibited by the GS-5734 1′-CN group, which is crucial in the termination of RNA replication [138].
The docking analysis revealed that nucleotide analogues block the RdRp activity in viruses by working with the chain termination mechanism. Remdesivir forms a hydrogen bond with ASP761 residue of RdRp (PDB ID: 7BTF) via its −NH2 group on the triazine ring, resulting in a binding affinity of −7.9 kcal/mol. Moreover, through polar interactions, remdesivir interacts with receptor residues TYR619, LYS621, CYS622, ARG624 and ARG553 [139].
Since viral proteases are essential for viral replication, many research groups concentrate on developing inhibitors of these enzymes. The FDA approved remdesivir, an RdRp inhibitor, as the first and sole medication to treat COVID-19 in 2020. The SARS-CoV-2 RdRp complex with the antiviral medication remdesivir has a cryo-EM crystal structure (PDB ID: 7BV2). The co-crystallized structures of remdesivir help us understand the viral RNA replication mechanism [140].
Moreover, the active metabolite (RdRp-RTP) was used by Xu et al. to create the cryo-EM structure of the drug with PDB ID: 7BV2 at a resolution of 2.5 Å. According to the crystal structure, Remdesivir monophosphate (RMP) attaches at the centre of the active site, and pyrophosphate is at the access point of the nucleotide entry channel to the enzyme active site. It impedes the entrance of nucleotide triphosphate to the active site. RMP also interacts with the LYS545 and ARG555 residues in the binding site. ASP623, SER682 and ASN691 are three residues with strong hydrogen bonds in the active site. ASN691 also interacts with a 2′-OH group of the sugar moiety [141]. A noteworthy feature is that the implicit and allosteric binding sites offer a flexible design framework to develop high-potency antagonists of RdRp of SARS-CoV-2 with kinetic selectivity [10] (Fig. 16).
The findings of the ongoing clinical studies in the USA and China will be essential to determine whether remdesivir is a feasible COVID-19 therapeutic option. In addition, the FDA in the USA has approved one clinical test. On 19 January 2020, a 35-year-old Washington resident received remdesivir and did not have to worry about COVID-19. However, further investigation is required into the drug’s therapeutic potential in infected patients. If the trial treatment works, it will be vital to ensure that the medication is commercially available and can meet the challenges of the current epidemic and future outbreaks [142, 143].
We discovered that the medicine could only significantly reduce hospital stay length while not influencing fatality rates. The drug is still being tested, and researchers are using the test results to study mortality reduction. Second, remdesivir therapy was found to cause 66% of adverse responses in clinical trials [144]. As a result, we may conclude that, while this medicine is a novel concept with a well-chosen research direction, it still has several flaws that need to be addressed [145]. Remdesivir inhibits the function of the virus RdRp and prevents the virus ExoN from performing proofreading, inhibiting replication of the virus [146]. Even in non-human primates with diverse viral diseases, such as the rodent SARS-CoV model and the rhesus monkey Ebola virus model, the in-vivo and in-vitro experiments have produced positive results by decreasing viral levels [147, 148].
Initial studies from in vitro test investigations suggested remdesivir’s therapeutic potential for treating human infections carried by novel emerging SARS-CoV-2, for which multiple research organizations launched numerous scientific trials [149].
These investigations have provided a new option for study into designing and altering remdesivir to efficiently generate a large variety of therapeutically active molecules to combat infectious diseases. Nucleotides are biologically active compounds, essential in almost every biochemical function, such as intestinal development, cellular immune system functioning and nutritional digestion. They interact with RNA through base chemisorption, stacking, hydrogen bonds, van der Waal interactions and binding to the phosphate backbone of RNA [150]. The choice of artificial synthetic nucleotide compounds is regarded as novel chemicals concerning antiviral activities. The peptide nucleic acid (PNA) lead molecule has strong specific sequence identification ability and hybridization stability and is not degraded by protease and nuclease [151].
In recent research, Bichismita Sahu and coworkers developed scaffolds formed from PNA and systematically performed in-silico analysis [152]. The designed compounds have good RdRp-binding affinity. They examined the impact of oligonucleotides on the binding effect of the derivatives using different PNA backbones, including 2-pyrrolidinyl amide, 2-aminoethyl prolyl and γ-PNA. The noticeable reduction of binding affinity caused by unaltered PNA’s backbone compared with the cyclic substituent can be mitigated by adding steric hindrance. As a result, investigators used the backbone of γ-PNA instead of the original aeg-PNA. Generally, these alterations give PNA oligomers steric strain, which causes the structure to pre-organize. The research data showed one of the 2-pyrrolidinyl amide and γ-PNA derivatives (Compounds A and B, as shown in Fig. 17) demonstrated better binding capacity than remdesivir. The PNA scaffold with a 2-pyrrolidinyl amide backbone with cyclic moiety showed inhibitory activity (−7.9 kcal/mol) against RdRp (PDB ID: 7BTF) by forming a hydrogen bond with TYR38 and ASP40 and also showed strong ionic interaction with ASP221 (Fig. 17A). However, the inhibitory activity of the PNA scaffold with an open chain backbone is −7.8 kcal/mol (Fig. 17B) [152].
5.8 AT-527
AT-527 (12, Table 1) is a guanosine nucleoside dual prodrug orally administered to patients. It has shown potent efficacy in vitro against clinical strains of HCV and efficacy in vivo with the 1-week monotherapy treatment of patients infected with HCV by explicitly inhibiting the viral RdRp [60, 153]. Moreover, AT-527 was recently reported to be effective and well tolerated at regular dosages of 550 mg for 3 months and showed a high rate of effectiveness in phase II clinical investigations with HCV-infected patients [154]. The free basic form of AT-527, AT-511 (Fig. 18), was investigated in vitro against SARS-CoV-2, and the results of this investigation demonstrate its solid antiviral efficacy. Following oral administration, the bioactive metabolite’s expected concentration in lung tissue increases the prospect that AT-527 could be a successful COVID-19 therapeutic option.
The host cell must first undergo metabolism to produce a biologically active derivative AT-9010 (Fig. 18) of AT-527. More than 40 HCV-infected individuals have received oral treatment with AT-527 in combination with daclatasvir, proving human safety and acceptability [155, 156]. The outstanding pharmacokinetics in combination with a dosage of 550 mg twice a day should result in lung exposures to the active metabolite that are notably higher than the medication’s in vitro EC90 against SARS-CoV-2 multiplication and could, therefore, result in successful antiviral therapy. A phase II clinical investigation is now being conducted to investigate the efficacy of AT-527 for SARS-CoV-2-infected patients [157].
The cellular enzymes metabolize AT-527, a prodrug of a guanosine nucleotide analogue, into the active triphosphate form (AT-9010) by masking both the base and phosphate group. Further, it inhibits the replication of SARS-CoV-2 by targeting the RdRp and the NiRAN (Nidovirus RdRp-associated nucleotidyltransferase) [158].
The docking analysis of AT-527 against RdRp (PDB ID: 7BV2) shows that ribose moiety forms hydrogen bonds with SER814 residue and that the 5′-phosphate interacted with CYS813. The 2′-fluoro group of the ribose ring shows interaction with THR206 (see Fig. 19) [159].
Shannon and co-workers proposed the dual mechanism of action of AT-527 and predicted the cryo-electron microscopic crystal structure of SARS-CoV-2 RdRp in complex with the active form of AT-527 (PDB ID: 7ED5) with the resolution of 2.98 Å [60, 119, 160]. The crystal structure showed that the active form AT-9010 had been incorporated into the RdRp active site at the 3′ end of the RNA product strand. To facilitate the incorporation of AT-9010 into the RNA product strand, the visible 5′ end of the RNA templates comprises four consecutive cytidine bases. At the +1 position of the RNA product, AT-9010 is incorporated and paired with cytosine (C27) on the template strand (Fig. 20). The RdRp–RNA complex appears to be in a post-translocation state, with the second AT-9010 occupying the −1 site and ready to incorporate, preventing other NTPs from being entered. The C27 of the template RNA strand is canonically base-paired with the guanine nucleobase of the incorporated AT-9010. Second, the AT-9010 ribose’s hydrophobic 2′-methyl group increases the inhibitory action by forming a hydrophobic barrier that hinders the incoming NTP’s ribose from being positioned correctly. As a result, the addition of AT-9010 stops the extension of the RNA strand, and the subsequent disruption of the complex’s catalytic site by AT-9010 renders it resistant to ExoN removal. AT-9010 functions as a chain terminator since it precludes further elongation of the product strand. AT-9010 also interacts with crucial amino acid residues of RdRp (LYS545, SER682, C26, ASN691 and CYS622) through conventional hydrogen bonds (Fig. 20) [161].
5.9 2-Methylguanosine Glucose-1-Phosphate (IDX-184)
A prodrug of 2-methylguanosine glucose-1-phosphate named IDX-184 (13, Table 1) targets the viral protein. The native nucleotide guanosine-5′-triphosphate (GTP) competes with it for the virus polymerase catalytic site, preventing polymerization [70, 162].
The 2-C-methylguanosine triphosphate is the active form of this drug. It inhibits HCV in vitro with potency and selectivity. Additionally, it is very well tolerated due to the active structure formed inside the hepatocytes. The toxicity of IDX-184 was decreased due to the active drug’s lower systemic concentration. While in clinical studies to treat HCV, MERS, SARS and Zika virus RdRps, IDX-184 performed better than other medications [163,164,165]. Additionally, four novel IDX-184 derivatives (2-hydroxyphenyl-oxidanyl, 13a), (3,5-dihydroxy phenyl-oxidanyl, 13b), (3-hydroxyphenyl-oxidanyl, compound 13c) and (3-sulfanylphenyl-oxidanyl, 13d, Fig. 21) exhibit promising results in attaching to the SARS-CoV-2 RdRp compared with parent IDX-184 [166]. Currently, drugs that specifically bind and inhibit SARS-CoV-2 proteins are urgently needed. The treatment of COVID-19 may be achieved efficiently by the treatment with novel derivatives 13b and 13c of IDX-184 with the improved binding affinity to the SARS-CoV-2 RdRp. Therefore, following validation of the binding assays, in vitro investigations and in vivo studies, these two derivatives of IDX-184 may be employed to target SARS-CoV-2 RdRp successfully [167, 168].
Docking studies demonstrated the in-depth binding of IDX-184 to the SARS-CoV-2 RdRp (PDB ID: 6NUR). Researchers proposed that continued refinement of this chemical would lead to a more potent molecule useful against SARS-CoV-2 [169]. A guanosine derivative IDX-184 (−9.0 kcal/mol) competes with GTP (−8.7 kcal/mol) for binding. IDX-184 binding is a bit better than GTP. More investigation of the complexes is necessary to fully understand how the docking complexes bind to the SARS-CoV-2 RdRp. The computational analysis of IDX-184 showed that 10 H-bonds were formed with receptor’s residues ARG444, LYS512 (2), ASP651, ASP652, ALA653 (3), TRP691 and GLU702. On the other hand, just one salt bridge was created with ASP514, while two metal contacts were created between IDX-184 and the RdRp’s active site residue ASP652 (Fig. 22) [84].
5.10 2′-Deoxy-2′-Fluoro-β-C-Methyluridine-5′-Monophosphate
The research shows that uracil analogue RdRp inhibitors are the best antiviral drugs with direct action [170]. The uracil derivative PSI-7672 (Fig. 23) can significantly inhibit HCV replication. The −CH3 and −F moiety at the C-2′ position maintain the drug’s potency and safety [171]. The negatively charged nature of substances containing phosphoric acid groups prevents the body from readily absorbing these substances [172]. The first prodrug, PSI-7672, was created when the prodrug concept was eventually adopted. The structure–activity relationship analysis showed that the D-alanine derivative of 2′-deoxy-2′-a-fluoro-β-C-methyluridine-5′-monophosphate is inactive, and only the natural l-amino acid can be active. The amino acid’s isomeric form is essential. At the R1 position of 2′-deoxy-2′-fluoro-β-C-methyluridine-5′-monophosphate (Fig. 23), a –CH3 and CH3–CH2-group can be substituted, which increases the drug’s efficacy. However, replacing a larger alkyl group at the R1 position results in a noticeable loss of molecule activity. The methyl group has the highest potency compared with other alkyl groups. The substitution at the R2 position offers the necessary micromolar inhibitory activity. The inhibitory potential of the drug molecule (2′-deoxy-2′-fluoro-β-C-methyluridine-5′-monophosphate) increases in the short alkyl and branched alkyl groups if the amino acid is alanine and the R3 position is phenyl substituted. However, 2-butyl, n-butyl and n-pentyl esters were cytotoxic.
Halogenated alkyl functional group and phenyl moiety at the R2 position does not sufficiently increase the potency of the drug molecule. According to the phosphoramidate ester group analysis at the R3 position, a phenyl-substituted derivative showed good potency and was not harmful. Finally, it was found that PSI-7851 (Fig. 23) has a favourable pharmacokinetic profile and is an excellent direct-acting antiviral (DAA) for suppressing HCV. PSI-7851 is a combination of two racemic, S-isomer PSI-7976 and R-isomer PSI-7977 (Fig. 23) [173]. Sofosbuvir (PSI-7977) (14, Table 1) has been considered a possible helpful medication for preventing SARS-CoV-2 infection [174]. For treating COVID-19 patients, the administration of PSI-7977 and routine medical care led to higher 2-week recovery efficiency and decreased hospital stays. Moreover, Pinar Mesci et al. revealed that PSI-7977 might treat cognitive problems caused by SARS-CoV-2 [175,176,177]. Sofosbuvir (PSI-7977) treated HCV safely and effectively without interferon. This drug was also effective against the hepatitis A virus, zika virus (ZIKV) infection and yellow fever. Sofosbuvir drug’s high in vitro activity (14,110 nM) and lack of apparent toxicity encourage future in vivo research. Because the RdRp replication process of HCV and SARS-CoV-2 are similar, it was hypothesized that the replication of SARS-CoV-2 was probably suppressed by PSI-7977 (sofosbuvir) [178]. In silico, sofosbuvir can create two hydrophobic interactions with TYR510 and ASP651 and seven hydrogen bonds with TRP508 (3), LYS512 (2), ALA653 and TRP691 of the SARS-CoV-2 RdRp (PDB ID: 6NUR) with a binding energy of −7.5 kcal/mol [179] (Fig. 24).
Furthermore, Chien and co-workers investigate that SARS-CoV-2 RdRp is irreversibly blocked by sofosbuvir triphosphate, preventing polymerase-mediated RNA extension. Sofosbuvir activity was not verified in a human cell-line model, despite modelling and docking studies showing a potential role for the drug in inhibiting SARS-CoV-2 RdRp activity [180]. However, nine randomized clinical studies were undertaken on 66 persons with mild to severe COVID-19 to examine the effectiveness of sofosbuvir in combination with other medications [181]. Clinical recovery within 14 days of treatment was the main consequence. The data from cellular models and in vivo research will help elucidate this drug’s potential against SARS-CoV-2, given its excellent safety profile and oral accessibility [182].
6 Pharmacophore Modelling of RdRp Inhibitors
A pattern of functional groups determines the bioactivity of a molecule called a pharmacophore. By defining the molecular functional characteristics required for a molecule to bind to a specific receptor and then directing the virtual screening of vast datasets of compounds for selecting the most suitable candidates, pharmacophore approaches represent one of the most intriguing tools ever created [183]. Pharmacophoric modelling is based on the idea that identical chemical functionalities and a common spatial arrangement cause biological activity on the same target. The pharmacophoric model depicts the chemical properties of a molecule that can interact with its ligand as geometric elements such as spheres, planes and vectors. Hydrophobic regions (H), hydrogen-bond acceptors (HBAs), hydrogen-bond donors (HBDs), negatively and positively ionizable groups, aromatic groups and metal coordinating sites are the most significant pharmacophoric feature types [184]. To reveal the shape and size of the binding pocket, further size limitations in the form of exclusion volumes in forbidden areas can be included [185, 186]. The RdRp–Remdesivir complex model’s pharmacophore structure was created in recent research [187]. Below are the pharmacophore properties and their x, y and z coordinates and radius in Table 2.
These pharmacophore properties, including one hydrogen-bond donor, six hydrogen-bond acceptors and two aromatic rings, are present in the remdesivir when coupled with SARS-CoV-2 RdRp. Remdesivir interacts with the active site of the RdRp receptor enzyme by hydrogen with GLN773 (3.33 Å) and by cation-pi bonding with LYS47 (3.40 Å) [188, 189]. The pharmacophoric map of RdRp inhibitors revealed that hydrophobic groups, which can take the form of aromatic rings or other structures, are crucial to the compounds’ ability to bind to the active site of the receptor protein [190]. Furthermore, the presence of HBA and HBD groups improves the interaction between inhibitors and the receptor’s active site. Unexpectedly, HBA/HBD groups, like RdRp inhibitors, can enhance their interactions and should be considered vital components in the enzyme inhibition process [10, 191] (Table 3).
7 The Current Status of SARS-CoV-2
The mutation rate of SARS-CoV-2 is exceptionally high since it is a positive-sense single-stranded RNA-encapsulated virus. There are almost 100 mutant strains in existence right now. However, the spike protein mutation accounts for many of these mutant strains [192]. Furthermore, mutations do not happen linearly; many mutations frequently occur simultaneously [193]. It will present a more significant challenge to the vaccine at this time or will evade the vaccine’s immunological protection. The Alpha, Delta and newly discovered Omicron viruses are among the most well known [194]. Although vaccine research and development has always been a priority for various countries, SARS-CoV-2 is easy to transcribe during replication. It frequently results in some surface proteins or vaccines targeting regions that seem changeable, and they cannot keep up with the virus mutation rate. Therefore, SARS-CoV-2 cannot be regulated effectively or consistently.
Unfortunately, few anti-virus vaccinations or medicines have been licensed to treat illnesses caused by SARS-CoV-2. Clinical management includes infection control, prevention and supportive care, including mechanical ventilation and supplemental oxygen when required. Although many nations are working on a SARS-CoV-2 vaccine, it is almost certain that none of the vaccines will be prepared by the end of the year. Currently, four types of vaccines are undergoing clinical trials: protein subunit, whole virus, nucleic acid (RNA and DNA) and viral vector [195]. Each of these vaccinations protects humans by inducing immunity [29].
Furthermore, certain vaccinations are dangerous and cannot be used. Thus, more dangers during testing make vaccine development a complicated path. Vaccine research is essential from this point of view, but finding one or two relatively stable targets to produce a drug that can work for a long time without fear of SARS-CoV-2 mutation is even more critical in the long run [196, 197].
As a result, there has been increased demand to develop another medicine that can effectively combat the infection. The focus of this effort has primarily been on repurposing existing medications. According to WHO officials, many drugs have successfully treated COVID-19.
8 Discussion
Initially established as a respiratory tract pathogen, subsequent studies have demonstrated that this viral infection can impact other organs and systems. The RdRp protein, crucial for RNA viruses, has been identified as a promising target for creating antiviral treatments. Repurposing existing RdRp inhibitors for SARS-CoV-2 is still a viable option to explore the correlation between the drug-binding sites of SARS-CoV, SARS-CoV-2 and MERS-CoV RdRps. Remdesivir, with its complex structure that inhibits RdRp, has provided insight into the mechanisms of viral RNA replication and a rationale for designing drugs to combat novel viral infections. It was discovered that remdesivir is covalently inserted into the primer strand, leading to chain elongation termination. The current list of potential medications includes EIDD-2801, sofosbuvir, galidesivir, remdesivir, AT-527, IDX-184, favipiravir and ribavirin, all of which have demonstrated anti-SARS-CoV-2 activity in cellular studies or clinical trials. These nucleoside analogues, such as remdesivir, can concentrate within the inner cavity of viral RdRps and block it via RNA chain termination. This approach involves converting the parent compound to the triphosphate-activated state. EIDD-2801, galidesivir, remdesivir, favipiravir and ribavirin maintain the entire ribose group and build a strong hydrogen-bond network resembling natural substrates. We assessed the pros and cons of nucleoside inhibitors (NIs) to evaluate potential medications better. The biochemical properties of NIs target the active site of viral RdRps. Most NI antiviral activity is broad-spectrum. Several effective anti-HCV NIs suggest that the active binding sites of these inhibitors are relatively conserved [198, 199]. The challenge in developing nucleoside inhibitors is the high cellular natural NTP concentration level. Triphosphorylated NIs require higher doses to achieve their antiviral effect due to competition with high levels of cellular NTP, increasing the risk of drug toxicity.
Cryo-electron microscopy frameworks of RdRp have elucidated the inhibitory mechanism of remdesivir. RdRp inhibitors are typically modified nucleosides with structurally modified nucleobase and ribose groups, which can be recognized by RdRp and function as their substrates after undergoing tri-phosphorylation by kinases. Subsequently, they can be integrated into a growing viral RNA strand. Hence, the chemical modifications in the structure of nucleoside inhibitors of RdRp impede the incorporation of a new nucleotide in the ever-increasing chain of viral RNA, efficiently causing RNA chain termination [200] (Fig. 25).
After entering the body, remdesivir undergoes metabolism, hydrolysis,and phosphorylation to form its active form, remdesivir monophosphate (RDV-MP) [201, 202].
RDV-MP then boosts the active triphosphate metabolite and interacts with the basic amino acid residues LYS545 and ARG555 of RdRp. Magnesium ions and pyrophosphate create two ionic bond contacts close to RDV-MP, which are absent in all other configurations of the SARS-CoV-2 RdRp complex with RNA. The triphosphate occupying the nucleotide input site may prevent nucleotide triphosphates (NTP) from entering the active site, providing a possible mechanism for remdesivir’s inhibitory effect [203, 204]. Other nucleotides, including favipiravir, ribavirin, IDX-184, galidesivir, EIDD-2801 and molnupiravir, effectively hinder SARS-CoV-2 replication by targeting RdRp [205] (Fig. 26).
Moreover, structural modifications enabling covalent integration of phosphate at the growing terminus are more favourable, and the development of derivatives of pyrophosphate bonded with additional functional groups, such as fluorine, can enhance their hydrogen bond acceptance and improve interactions with RdRp [206, 207]. Virtual screening analysis has demonstrated that newly designed medicines containing phenyl pyrazole or tetrazole functionalities increase the likelihood of inhibiting RdRp [208]. Aryl di-keto acids have been proven to be highly effective and temporary inhibitors of HCV’s RdRp. They act as product-like mimics, binding the two Mg2+ ions (divalent cations) at the active site of HCV, similar to pyrophosphate mimetic inhibitors. The investigation showed that bioisosteres tetrazoles or triazoles, such as ribavirin, could replace the free carboxylic acid moiety of aryl di-keto acids since the free acidic functional group causes biological and chemical instability and low membrane permeability of drug molecules [209, 210] (Fig. 27).
The discussion in this review indicates that targeting RdRp could be the most effective therapeutic strategy for combating COVID-19 infection since it is the primary receptor for viral replication. Small-molecule inhibitors are considered the best option during development due to their research progress, safety and efficacy, compared with alternative treatments like oligonucleotide, monoclonal antibody-based, peptide and plasma therapies. However, there is a need to discover target-specific lead compounds, and expert investigation on SARS-CoV-2 infections is ongoing. To develop novel medications, it is crucial to understand the pharmacophore characteristics that are essential for all substances.
9 Conclusion and Future Aspects
SARS-CoV-2 has rapidly spread to nearly 199 countries worldwide, with no specific treatment available to combat this wild-type virus. Developing drugs for COVID-19 is a complex yet necessary process due to the virus’s genetic alterations and medication resistance, which pose challenges in treatment strategies. One of the major problems in drug design is to enhance the drug’s potency against genetic variants, for which adding a suitable pharmacophore to a newly designed molecule is preferred. Although pharmacophore modelling and docking studies have successfully recognized remdesivir as an effective inhibitor of the RdRp enzyme, clinical trials are necessary to confirm these findings. Apart from remdesivir, several nucleotides like ribavirin, EIDD-2801, favipiravir and galidesivir have shown efficacy in blocking SARS-CoV-2 replication. Additionally, some FDA-approved small molecules for clinical use have demonstrated effectiveness against SARS-CoV-2. This review summarizes existing antiviral drugs and their structure–activity relationship, and comparing or incorporating their characteristics is crucial to optimize or to design more effective drugs in the future.
Data availability statement
The data supporting this work is available in this paper, and all other details are present in the supporting information. Furthermore, the data supporting this study are available from the corresponding author upon reasonable request.
Abbreviations
- EUA:
-
Emergency Use Authorization
- FDA:
-
Food and Drug Administration
- ACE2:
-
Angiotensin-converting enzyme 2
- NHC:
-
N4-hydroxycytidine
- RSV:
-
Respiratory syncytial virus
- TNF-α:
-
Tumour necrosis factor α
- ExoN:
-
Exoribonuclease
- CTP:
-
Cytidine triphosphate
- HBD:
-
Hydrogen-bond donors
- HBA:
-
Hydrogen-bond acceptors
- NiRAN:
-
Nidovirus RdRp-associated nucleotidyl transferase
References
Park SE (2020) Epidemiology, virology, and clinical features of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2; coronavirus disease-19). Pediatr Infect Vaccine 27(1):1–10
Ilyas M, Muhammad S, Iqbal J, Amin S, Al-Sehemi AG, Algarni H, Alarfaji SS, Alshahrani MY, Ayub K (2022) Insighting isatin derivatives as potential antiviral agents against NSP3 of COVID-19. Chem Pap 76(10):6271–6285
Wu W, Wang A, Liu M (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395(10223):497–506
Dyall J, Gross R, Kindrachuk J, Johnson RF, Olinger GG Jr, Hensley LE, Frieman MB, Jahrling PB (2017) Middle East respiratory syndrome and severe acute respiratory syndrome: current therapeutic options and potential targets for novel therapies. Drugs 77(18):1935–1966
Deng S-Q, Peng H-J (2020) Characteristics of and public health responses to the coronavirus disease 2019 outbreak in China. J Clin Med 9(2):575
Allan M, Lièvre M, Laurenson-Schaefer H, de Barros S, Jinnai Y, Andrews S, Stricker T, Formigo JP, Schultz C, Perrocheau A (2022) The World Health Organization COVID-19 surveillance database. Int J Equity Health 21(Suppl 3):167
Agostini ML, Andres EL, Sims AC, Graham RL, Sheahan TP, Lu X, Smith EC, Case JB, Feng JY, Jordan R (2018) Coronavirus susceptibility to the antiviral remdesivir (GS-5734) is mediated by the viral polymerase and the proofreading exoribonuclease. MBio 9(2):e00221-18
Harrison AG, Lin T, Wang P (2020) Mechanisms of SARS-CoV-2 transmission and pathogenesis. Trends Immunol 41(12):1100–1115
Gordon CJ, Tchesnokov EP, Feng JY, Porter DP, Götte M (2020) The antiviral compound remdesivir potently inhibits RNA-dependent RNA polymerase from Middle East respiratory syndrome coronavirus. J Biol Chem 295(15):4773–4779
Yin W, Mao C, Luan X, Shen D-D, Shen Q, Su H, Wang X, Zhou F, Zhao W, Gao M (2020) Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir. Science 368(6498):1499–1504
Parvez MSA, Karim MA, Hasan M, Jaman J, Karim Z, Tahsin T, Hasan MN, Hosen MJ (2020) Prediction of potential inhibitors for RNA-dependent RNA polymerase of SARS-CoV-2 using comprehensive drug repurposing and molecular docking approach. Int J Biol Macromol 163:1787–1797
Jordheim LP, Durantel D, Zoulim F, Dumontet C (2013) Advances in the development of nucleoside and nucleotide analogues for cancer and viral diseases. Nat Rev Drug Discov 12(6):447–464
Schuler J, Falls Z, Mangione W, Hudson ML, Bruggemann L, Samudrala R (2022) Evaluating the performance of drug-repurposing technologies. Drug Discov Today 27(1):49–64
Zhang J, Xiao T, Cai Y, Chen B (2021) Structure of SARS-CoV-2 spike protein. Curr Opin Virol 50:173–182
Li G, De Clercq E (2020) Therapeutic options for the 2019 novel coronavirus (2019-nCoV). Nat Rev Drug Discov 19(3):149–150
Aronskyy I, Masoudi-Sobhanzadeh Y, Cappuccio A, Zaslavsky E (2021) Advances in the computational landscape for repurposed drugs against COVID-19. Drug Discov Today 26(12):2800–2815
Muhammad S, Amin S, Iqbal J, Al-Sehemi AG, Alarfaji SS, Ilyas M, Atif M, Ullah S (2022) Insighting the therapeutic potential of fifty (50) shogaol derivatives against mpro of SARS-CoV-2. J Comput Biophys Chem 21(05):555–568
McBride R, Van Zyl M, Fielding BC (2014) The coronavirus nucleocapsid is a multifunctional protein. Viruses 6(8):2991–3018
Hilgenfeld R (2014) From SARS to MERS: crystallographic studies on coronaviral proteases enable antiviral drug design. FEBS J 281(18):4085–4096
Li F (2016) Structure, function, and evolution of coronavirus spike proteins. Annu Rev Virol 3:237–261
Morse JS, Lalonde T, Xu S, Liu WR (2020) Learning from the past: possible urgent prevention and treatment options for severe acute respiratory infections caused by 2019-nCoV. Chembiochem 21(5):730–738
Hillen HS (2021) Structure and function of SARS-CoV-2 polymerase. Curr Opin Virol 48:82–90
Te Velthuis AJ, Van Den Worm SH, Snijder EJ (2012) The SARS-coronavirus nsp7+ nsp8 complex is a unique multimeric RNA polymerase capable of both de novo initiation and primer extension. Nucleic Acids Res 40(4):1737–1747
Muhammad S, Qaisar M, Iqbal J, Khera RA, Al-Sehemi AG, Alarfaji SS, Adnan M (2022) Exploring the inhibitory potential of novel bioactive compounds from mangrove actinomycetes against nsp10 the major activator of SARS-CoV-2 replication. Chem Pap 76(5):3051–3064
Kaushik D, Bhandari R, Kuhad A (2021) TLR4 as a therapeutic target for respiratory and neurological complications of SARS-CoV-2. Expert Opin Ther Targets 25(6):491–508
Li X, Yang Y, Liu L, Yang X, Zhao X, Li Y, Ge Y, Shi Y, Lv P, Zhang J (2020) Effect of combination antiviral therapy on hematological profiles in 151 adults hospitalized with severe coronavirus disease 2019. Pharmacol Res 160:105036
Tompa DR, Immanuel A, Srikanth S, Kadhirvel S (2021) Trends and strategies to combat viral infections: a review on FDA approved antiviral drugs. Int J Biol Macromol 172:524–541
Singhal T (2020) A review of coronavirus disease-2019 (COVID-19). Indian J Pediatr 87(4):281–286
Bobrowski T, Melo-Filho CC, Korn D, Alves VM, Popov KI, Auerbach S, Schmitt C, Moorman NJ, Muratov EN, Tropsha A (2020) Learning from history: do not flatten the curve of antiviral research! Drug Discov Today 25(9):1604–1613
Bank PD (2021) RCSB protein data bank: integrated searching and efficient access to macromolecular structure data from the PDB archive. Found Crystallogr 77:a253
Kouranov A, Xie L, de la Cruz J, Chen L, Westbrook J, Bourne PE, Berman HM (2006) The RCSB PDB information portal for structural genomics. Nucleic Acids Res 34(Suppl 1):D302–D305
Tian W, Chen C, Liang J (2018) CASTp 30: computed atlas of surface topography of proteins and beyond. Biophys J 114(3):50a
Chan JF-W, Kok K-H, Zhu Z, Chu H, To KK-W, Yuan S, Yuen K-Y (2020) Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg Microbes Infect 9(1):221–236
Grellet E, Goulet A, Imbert I (2022) Replication of the coronavirus genome: a paradox among positive-strand RNA viruses. J Biol Chem 298:101923
Gao Y, Yan L, Huang Y, Liu F, Zhao Y, Cao L, Wang T, Sun Q, Ming Z, Zhang L (2020) Structure of the RNA-dependent RNA polymerase from COVID-19 virus. Science 368(6492):779–782
Shu B, Gong P (2016) Structural basis of viral RNA-dependent RNA polymerase catalysis and translocation. Proc Natl Acad Sci 113(28):E4005–E4014
Kirchdoerfer RN, Ward AB (2019) Structure of the SARS-CoV nsp12 polymerase bound to nsp7 and nsp8 co-factors. Nat Commun 10(1):2342
Furuta Y, Takahashi K, Kuno-Maekawa M, Sangawa H, Uehara S, Kozaki K, Nomura N, Egawa H, Shiraki K (2005) Mechanism of action of T-705 against influenza virus. Antimicrob Agents Chemother 49(3):981–986
Furuta Y, Komeno T, Nakamura T (2017) Favipiravir (T-705), a broad spectrum inhibitor of viral RNA polymerase. Proc Jpn Acad Ser B 93(7):449–463
Furuta Y, Gowen BB, Takahashi K, Shiraki K, Smee DF, Barnard DL (2013) Favipiravir (T-705), a novel viral RNA polymerase inhibitor. Antivir Res 100(2):446–454
Abdelnabi R, de Morais ATS, Leyssen P, Imbert I, Beaucourt S, Blanc H, Froeyen M, Vignuzzi M, Canard B, Neyts J (2017) Understanding the mechanism of the broad-spectrum antiviral activity of favipiravir (T-705): key role of the F1 motif of the viral polymerase. J Virol 91(12):e00487-17
Graci JD, Cameron CE (2006) Mechanisms of action of ribavirin against distinct viruses. Rev Med Virol 16(1):37–48
Deval J, Fung A, Stevens SK, Jordan PC, Gromova T, Taylor JS, Hong J, Meng J, Wang G, Dyatkina N (2016) Biochemical effect of resistance mutations against synergistic inhibitors of RSV RNA polymerase. PLoS One 11(5):e0154097
Deval J, Hong J, Wang G, Taylor J, Smith LK, Fung A, Stevens SK, Liu H, Jin Z, Dyatkina N (2015) Molecular basis for the selective inhibition of respiratory syncytial virus RNA polymerase by 2′-fluoro-4′-chloromethyl-cytidine triphosphate. PLoS Pathog 11(6):e1004995
Wang G, Deval J, Hong J, Dyatkina N, Prhavc M, Taylor J, Fung A, Jin Z, Stevens SK, Serebryany V (2015) Discovery of 4′-chloromethyl-2′-deoxy-3′, 5′-di-O-isobutyryl-2′-fluorocytidine (ALS-8176), a first-in-class RSV polymerase inhibitor for treatment of human respiratory syncytial virus infection. J Med Chem 58(4):1862–1878
Nilsson M, Kalayanov G, Winqvist A, Pinho P, Sund C, Zhou X-X, Wähling H, Belfrage A-K, Pelcman M, Agback T (2012) Discovery of 4′-azido-2′-deoxy-2′-C-methyl cytidine and prodrugs thereof: a potent inhibitor of hepatitis C virus replication. Bioorg Med Chem Lett 22(9):3265–3268
Rondla R, Coats SJ, McBrayer TR, Grier J, Johns M, Tharnish PM, Whitaker T, Zhou L, Schinazi RF (2009) Anti-hepatitis C virus activity of novel β-d-2′-C-methyl-4′-azido pyrimidine nucleoside phosphoramidate prodrugs. Antivir Chem Chemother 20(2):99–106
Deutsch M, Hadziyannis S (2008) Old and emerging therapies in chronic hepatitis C: an update. J Viral Hepat 15(1):2–11
Gerber L, Welzel TM, Zeuzem S (2013) New therapeutic strategies in HCV: polymerase inhibitors. Liver Int 33:85–92
Andreou A, Trantza S, Filippou D, Sipsas N, Tsiodras S (2020) COVID-19: the potential role of copper and N-acetylcysteine (NAC) in a combination of candidate antiviral treatments against SARS-CoV-2. In Vivo 34(3 suppl):1567–1588
Painter WP, Holman W, Bush JA, Almazedi F, Malik H, Eraut NC, Morin MJ, Szewczyk LJ, Painter GR (2021) Human safety, tolerability, and pharmacokinetics of molnupiravir, a novel broad-spectrum oral antiviral agent with activity against SARS-CoV-2. Antimicrob Agents Chemother 65(5):e02428-20
Cox RM, Wolf JD, Plemper RK (2021) Therapeutically administered ribonucleoside analogue MK-4482/EIDD-2801 blocks SARS-CoV-2 transmission in ferrets. Nat Microbiol 6(1):11–18
Celik I, Tallei TE (2022) A computational comparative analysis of the binding mechanism of molnupiravir’s active metabolite to RNA-dependent RNA polymerase of wild-type and Delta subvariant AY 4 of SARS-CoV-2. J Cell Biochem 123(4):807–818
Painter GR, Bowen RA, Bluemling GR, DeBergh J, Edpuganti V, Gruddanti PR, Guthrie DB, Hager M, Kuiper DL, Lockwood MA (2019) The prophylactic and therapeutic activity of a broadly active ribonucleoside analog in a murine model of intranasal Venezuelan equine encephalitis virus infection. Antivir Res 171:104597
Agostini ML, Pruijssers AJ, Chappell JD, Gribble J, Lu X, Andres EL, Bluemling GR, Lockwood MA, Sheahan TP, Sims AC (2019) Small-molecule antiviral β-d-N 4-hydroxycytidine inhibits a proofreading-intact coronavirus with a high genetic barrier to resistance. J Virol 93(24):e01348-19
Julander JG, Demarest JF, Taylor R, Gowen BB, Walling DM, Mathis A, Babu Y (2021) An update on the progress of galidesivir (BCX4430), a broad-spectrum antiviral. Antivir Res 195:105180
Aschenbrenner DS (2021) Remdesivir approved to treat COVID-19 amid controversy. Am J Nurs 121(1):22–24
Young B, Tan TT, Leo YS (2021) The place for remdesivir in COVID-19 treatment. Lancet Infect Dis 21(1):20–21
Hendaus MA (2021) Remdesivir in the treatment of coronavirus disease 2019 (COVID-19): a simplified summary. J Biomol Struct Dyn 39(10):3787–3792
Shannon A, Fattorini V, Sama B, Selisko B, Feracci M, Falcou C, Gauffre P, El Kazzi P, Delpal A, Decroly E (2022) A dual mechanism of action of AT-527 against SARS-CoV-2 polymerase. Nat Commun 13(1):621
Good SS, Westover J, Jung KH, La Colla P, Collu G, Moussa A, Canard B, Sommadossi J-P (2020) AT-527 is a potent in vitro replication inhibitor of SARS-CoV-2, the virus responsible for the COVID-19 pandemic. Biorxiv 2020:242834
Elfiky AA, Elshemey WM, Gawad WA (2015) 2′-Methylguanosine prodrug (IDX-184), phosphoramidate prodrug (sofosbuvir), diisobutyryl prodrug (R7128) are better than their parent nucleotides and ribavirin in hepatitis C virus inhibition: a molecular modeling study. J Comput Theor Nanosci 12(3):376–386
Elfiky AA (2016) Zika viral polymerase inhibition using anti-HCV drugs both in market and under clinical trials. J Med Virol 88(12):2044–2051
Sadeghi A, Ali Asgari A, Norouzi A, Kheiri Z, Anushirvani A, Montazeri M, Hosamirudsai H, Afhami S, Akbarpour E, Aliannejad R (2020) Sofosbuvir and daclatasvir compared with standard of care in the treatment of patients admitted to hospital with moderate or severe coronavirus infection (COVID-19): a randomized controlled trial. J Antimicrob Chemother 75(11):3379–3385
Vicenti I, Zazzi M, Saladini F (2021) SARS-CoV-2 RNA-dependent RNA polymerase as a therapeutic target for COVID-19. Expert Opin Ther Patents 31(4):325–337
Shannon A, Canard B (2023) Kill or corrupt: mechanisms of action and drug-resistance of nucleotide analogues against SARS-CoV-2. Antivir Res 210:105501
Moeller NH, Shi K, Demir Ö, Belica C, Banerjee S, Yin L, Durfee C, Amaro RE, Aihara H (2022) Structure and dynamics of SARS-CoV-2 proofreading exoribonuclease ExoN. Proc Natl Acad Sci 119(9):e2106379119
Jia X, Schols D, Meier C (2020) Lipophilic triphosphate prodrugs of various nucleoside analogues. J Med Chem 63(13):6991–7007
Mackman RL (2022) Phosphoramidate prodrugs continue to deliver, the journey of remdesivir (GS-5734) from RSV to SARS-CoV-2. ACS Med Chem Lett 13(3):338–347
Elfiky AA (2020) Anti-HCV, nucleotide inhibitors, repurposing against COVID-19. Life Sci 248:117477
Wang Z, Yang L, Zhao X-E (2021) Co-crystallization and structure determination: an effective direction for anti-SARS-CoV-2 drug discovery. Comput Struct Biotechnol J 19:4684–4701
Zhang W-F, Stephen P, Theriault J-F, Wang R, Lin S-X (2020) Novel coronavirus polymerase and nucleotidyl-transferase structures: potential to target new outbreaks. J Phys Chem Lett 11(11):4430–4435
Celik I, Erol M, Duzgun Z (2021) In silico evaluation of potential inhibitory activity of remdesivir, favipiravir, ribavirin and galidesivir active forms on SARS-CoV-2 RNA polymerase. Mol Divers 26:279–292
Du YX, Chen XP (2020) Favipiravir: pharmacokinetics and concerns about clinical trials for 2019-nCoV infection. Clin Pharmacol Ther 108(2):242–247
Naydenova K, Muir KW, Wu L-F, Zhang Z, Coscia F, Peet MJ, Castro-Hartmann P, Qian P, Sader K, Dent K (2021) Structure of the SARS-CoV-2 RNA-dependent RNA polymerase in the presence of favipiravir-RTP. Proc Natl Acad Sci 118(7):e2021946118
Pilkington V, Pepperrell T, Hill A (2020) A review of the safety of favipiravir – a potential treatment in the COVID-19 pandemic? J Virus Erad 6(2):45–51
Chen C, Huang J, Yin P, Zhang Y, Cheng Z, Wu J, Chen S, Zhang Y, Chen B, Lu M (2020) Favipiravir versus arbidol for COVID-19: a randomized clinical trial. MedRxiv 2020:20037432
Blattman N (2015) Management of hepatitis C in patients with HIV care for patients with HIV-HCV co-infection is evolving as new medications are introduced that will provide simpler, more accessible treatment regimens. Fed Pract 32(Suppl 2):15S
Arabi YM, Shalhoub S, Mandourah Y, Al-Hameed F, Al-Omari A, Al Qasim E, Jose J, Alraddadi B, Almotairi A, Al Khatib K (2020) Ribavirin and interferon therapy for critically ill patients with Middle East respiratory syndrome: a multicenter observational study. Clin Infect Dis 70(9):1837–1844
Parker WB (2005) Metabolism and antiviral activity of ribavirin. Virus Res 107(2):165–171
Unal MA, Bitirim CV, Summak GY, Bereketoglu S, CevherZeytin I, Besbinar O, Gurcan C, Aydos D, Goksoy E, Kocakaya E (2021) Ribavirin shows antiviral activity against SARS-CoV-2 and downregulates the activity of TMPRSS2 and the expression of ACE2 in vitro. Can J Physiol Pharmacol 99(5):449–460
Bylehn F, Menendez CA, Perez-Lemus GR, Alvarado W, De Pablo JJ (2021) Modeling the binding mechanism of remdesivir, favilavir, and ribavirin to SARS-CoV-2 RNA-dependent RNA polymerase. ACS Cent Sci 7(1):164–174
Uddin R, Jalal K, Khan K (2022) Re-purposing of hepatitis C virus FDA approved direct acting antivirals as potential SARS-CoV-2 protease inhibitors. J Mol Struct 1250:131920
Elfiky AA (2020) Ribavirin, remdesivir, sofosbuvir, galidesivir, and tenofovir against SARS-CoV-2 RNA dependent RNA polymerase (RdRp): a molecular docking study. Life Sci 253:117592
Crotty S, Andino R (2002) Implications of high RNA virus mutation rates: lethal mutagenesis and the antiviral drug ribavirin. Microbes Infect 4(13):1301–1307
Leong HN, Ang B, Earnest A, Teoh C, Xu W, Leo YS (2004) Investigational use of ribavirin in the treatment of severe acute respiratory syndrome, Singapore, 2003. Trop Med Int Health 9(8):923–927
Booth CM, Matukas LM, Tomlinson GA, Rachlis AR, Rose DB, Dwosh HA, Walmsley SL, Mazzulli T, Avendano M, Derkach P (2003) Clinical features and short-term outcomes of 144 patients with SARS in the greater Toronto area. JAMA 289(21):2801–2809
Lee N, Hui D, Wu A, Chan P, Cameron P, Joynt GM, Ahuja A, Yung MY, Leung C, To K (2003) A major outbreak of severe acute respiratory syndrome in Hong Kong. N Engl J Med 348(20):1986–1994
Tan EL, Ooi EE, Lin C-Y, Tan HC, Ling AE, Lim B, Stanton LW (2004) Inhibition of SARS coronavirus infection in vitro with clinically approved antiviral drugs. Emerg Infect Dis 10(4):581
Muller MP, Dresser L, Raboud J, McGeer A, Rea E, Richardson SE, Mazzulli T, Loeb M, Louie M (2007) Adverse events associated with high-dose ribavirin: evidence from the Toronto outbreak of severe acute respiratory syndrome. Pharmacotherapy 27(4):494–503
Barlow A, Landolf KM, Barlow B, Yeung SYA, Heavner JJ, Claassen CW, Heavner MS (2020) Review of emerging pharmacotherapy for the treatment of coronavirus disease 2019. Pharmacotherapy 40(5):416–437
Chang C-H, Chen K, Lai M-Y, Chan K (2002) Meta-analysis: ribavirin-induced haemolytic anaemia in patients with chronic hepatitis C. Aliment Pharmacol Ther 16(9):1623–1632
Knowles SR, Phillips EJ, Dresser L, Matukas L (2003) Common adverse events associated with the use of ribavirin for severe acute respiratory syndrome in Canada. Clin Infect Dis 37(8):1139–1142
Kaur K, Gandhi MA, Slish J (2015) Drug-drug interactions among hepatitis C virus (HCV) and human immunodeficiency virus (HIV) medications. Infect Dis Ther 4:159–172
Zandi K, Amblard F, Musall K, Downs-Bowen J, Kleinbard R, Oo A, Cao D, Liang B, Russell OO, McBrayer T (2020) Repurposing nucleoside analogs for human coronaviruses. Antimicrob Agents Chemother 65(1):e01652-20
Goswami D (2021) Comparative assessment of RNA-dependent RNA polymerase (RdRp) inhibitors under clinical trials to control SARS-CoV2 using rigorous computational workflow. RSC Adv 11(46):29015–29028
Mandal M, Chowdhury SK, Khan AA, Baildya N, Dutta T, Misra D, Ghosh NN (2021) Inhibitory efficacy of RNA virus drugs against SARS-CoV-2 proteins: an extensive study. J Mol Struct 1234:130152
Chang J (2022) 4′-Modified nucleosides for antiviral drug discovery: achievements and perspectives. Acc Chem Res 55(4):565–578
Clark JL, Hollecker L, Mason JC, Stuyver LJ, Tharnish PM, Lostia S, McBrayer TR, Schinazi RF, Watanabe KA, Otto MJ (2005) Design, synthesis, and antiviral activity of 2′-deoxy-2′-fluoro-2′-C-methylcytidine, a potent inhibitor of hepatitis C virus replication. J Med Chem 48(17):5504–5508
Fung A, Jin Z, Dyatkina N, Wang G, Beigelman L, Deval J (2014) Efficiency of incorporation and chain termination determines the inhibition potency of 2′-modified nucleotide analogs against hepatitis C virus polymerase. Antimicrob Agents Chemother 58(7):3636–3645
Deore R, Chern JW (2010) NS5B RNA dependent RNA polymerase inhibitors: the promising approach to treat hepatitis C virus infections. Curr Med Chem 17(32):3806–3826
Nishiyama T, Kobayashi T, Jirintai S, Nagashima S, Primadharsini PP, Nishizawa T, Okamoto H (2019) Antiviral candidates against the hepatitis E virus (HEV) and their combinations inhibit HEV growth in in vitro. Antivir Res 170:104570
Qu C, Xu L, Yin Y, Peppelenbosch MP, Pan Q, Wang W (2017) Nucleoside analogue 2′-C-methylcytidine inhibits hepatitis E virus replication but antagonizes ribavirin. Adv Virol 162:2989–2996
Lee J-C, Tseng C-K, Wu Y-H, Kaushik-Basu N, Lin C-K, Chen W-C, Wu H-N (2015) Characterization of the activity of 2′-C-methylcytidine against dengue virus replication. Antivir Res 116:1–9
Rocha-Pereira J, Jochmans D, Dallmeier K, Leyssen P, Cunha R, Costa I, Nascimento M, Neyts J (2012) Inhibition of norovirus replication by the nucleoside analogue 2′-C-methylcytidine. Biochem Biophys Res Commun 427(4):796–800
Elfiky AA (2021) SARS-CoV-2 RNA dependent RNA polymerase (RdRp) targeting: an in silico perspective. J Biomol Struct Dyn 39(9):3204–3212
Jena N (2020) Identification of potent drugs and antiviral agents for the treatment of the SARS-CoV-2 infection. J Biol Med Chem
Pierra C, Amador A, Benzaria S, Cretton-Scott E, d’Amours M, Mao J, Mathieu S, Moussa A, Bridges EG, Standring DN (2006) Synthesis and pharmacokinetics of valopicitabine (NM283), an efficient prodrug of the potent anti-HCV agent 2′-C-methylcytidine. J Med Chem 49(22):6614–6620
Kuntzen T, Timm J, Berical A, Lennon N, Berlin AM, Young SK, Lee B, Heckerman D, Carlson J, Reyor LL (2008) Naturally occurring dominant resistance mutations to hepatitis C virus protease and polymerase inhibitors in treatment-naive patients. Hepatology 48(6):1769–1778
Kumar R, Mishra S, Maurya SK (2021) Recent advances in the discovery of potent RNA-dependent RNA-polymerase (RdRp) inhibitors targeting viruses. RSC Med Chem 12(3):306–320
Tian L, Qiang T, Liang C, Ren X, Jia M, Zhang J, Li J, Wan M, YuWen X, Li H (2021) RNA-dependent RNA polymerase (RdRp) inhibitors: the current landscape and repurposing for the COVID-19 pandemic. Eur J Med Chem 213:113201
Gordon CJ, Tchesnokov EP, Schinazi RF, Götte M (2021) Molnupiravir promotes SARS-CoV-2 mutagenesis via the RNA template. J Biol Chem 297(1):100770
Wang M, Wu C, Liu N, Zhang F, Dong H, Wang S, Chen M, Jiang X, Gu L (2021) SARS-CoV-2 RdRp is a versatile enzyme with proofreading activity and ability to incorporate NHC into RNA by using diphosphate form molnupiravir as a substrate. BioRxiv 2021:468737
Urakova N, Kuznetsova V, Crossman DK, Sokratian A, Guthrie DB, Kolykhalov AA, Lockwood MA, Natchus MG, Crowley MR, Painter GR (2018) β-D-N 4-hydroxycytidine is a potent anti-alphavirus compound that induces a high level of mutations in the viral genome. J Virol 92(3):e01965-17
Hashemian SMR, Pourhanifeh MH, Hamblin MR, Shahrzad MK, Mirzaei H (2022) RdRp inhibitors and COVID-19: is molnupiravir a good option? Biomed Pharmacother 146:112517
Sheahan TP, Sims AC, Zhou S, Graham RL, Pruijssers AJ, Agostini ML, Leist SR, Schäfer A, Dinnon KH III, Stevens LJ (2020) An orally bioavailable broad-spectrum antiviral inhibits SARS-CoV-2 in human airway epithelial cell cultures and multiple coronaviruses in mice. Sci Transl Med 12(541):eabb5883
Mestres J (2020) The target landscape of N4-hydroxycytidine based on its chemical neighborhood. BioRxiv 2020:016485
Imran M, Kumar Arora M, Asdaq SMB, Khan SA, Alaqel SI, Alshammari MK, Alshehri MM, Alshrari AS, Mateq Ali A, Al-Shammeri AM (2021) Discovery, development, and patent trends on molnupiravir: a prospective oral treatment for COVID-19. Molecules 26(19):5795
Stevaert A, Groaz E, Naesens L (2022) Nucleoside analogs for management of respiratory virus infections: mechanism of action and clinical efficacy. Curr Opin Virol 57:101279
Zarenezhad E, Marzi M (2022) Review on molnupiravir as a promising oral drug for the treatment of COVID-19. Med Chem Res 31:232–243
Gangadharan S, Ambrose JM, Rajajagadeesan A, Kullappan M, Patil S, Gandhamaneni SH, Veeraraghavan VP, Nakkella AK, Agarwal A, Jayaraman S (2022) Repurposing of potential antiviral drugs against RNA-dependent RNA polymerase of SARS-CoV-2 by computational approach. J Infect Public Health 15(11):1180–1191
Kabinger F, Stiller C, Schmitzová J, Dienemann C, Kokic G, Hillen HS, Höbartner C, Cramer P (2021) Mechanism of molnupiravir-induced SARS-CoV-2 mutagenesis. Nat Struct Mol Biol 28(9):740–746
Fischer WA, Eron JJ Jr, Holman W, Cohen MS, Fang L, Szewczyk LJ, Sheahan TP, Baric R, Mollan KR, Wolfe CR (2021) A phase 2a clinical trial of molnupiravir in patients with COVID-19 shows accelerated SARS-CoV-2 RNA clearance and elimination of infectious virus. Sci Transl Med 14(628):eabl7430
Ju J, Kumar S, Li X, Jockusch S, Russo JJ (2020) Nucleotide analogues as inhibitors of viral polymerases. BioRxiv 2020:927574
Abuo-Rahma GE-DA, Mohamed MF, Ibrahim TS, Shoman ME, Samir E, Abd El-Baky RM (2020) Potential repurposed SARS-CoV-2 (COVID-19) infection drugs. RSC Adv 10(45):26895–26916
Jena N (2020) Role of different tautomers in the base-pairing abilities of some of the vital antiviral drugs used against COVID-19. Phys Chem Chem Phys 22(48):28115–28122
Taylor R, Bowen R, Demarest JF, DeSpirito M, Hartwig A, Bielefeldt-Ohmann H, Walling DM, Mathis A, Babu YS (2021) Activity of galidesivir in a hamster model of SARS-CoV-2. Viruses 14(1):8
Gandeepan P, Ackermann L (2018) Transient directing groups for transformative C–H activation by synergistic metal catalysis. Chem 4(2):199–222
Romano M, Ruggiero A, Squeglia F, Maga G, Berisio R (2020) A structural view of SARS-CoV-2 RNA replication machinery: RNA synthesis, proofreading and final capping. Cells 9(5):1267
Hasan MK, Kamruzzaman M, Manjur OHB, Mahmud A, Hussain N, Mondal MSA, Hosen MI, Bello M, Rahman A (2021) Structural analogues of existing anti-viral drugs inhibit SARS-CoV-2 RNA dependent RNA polymerase: a computational hierarchical investigation. Heliyon 7(3):e06435
Ataei M, Hosseinjani H (2020) Molecular mechanisms of galidesivir as a potential antiviral treatment for COVID-19. J Pharm Care 2020:150–151
Uzunova K, Filipova E, Pavlova V, Vekov T (2020) Insights into antiviral mechanisms of remdesivir, lopinavir/ritonavir and chloroquine/hydroxychloroquine affecting the new SARS-CoV-2. Biomed Pharmacother 131:110668
Wang M, Cao R, Zhang L, Yang X, Liu J, Xu M, Shi Z, Hu Z, Zhong W, Xiao G (2020) Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res 30(3):269–271
Pizzorno A, Padey B, Julien T, Trouillet-Assant S, Traversier A, Errazuriz-Cerda E, Fouret J, Dubois J, Gaymard A, Lescure F-X (2020) Characterization and treatment of SARS-CoV-2 in nasal and bronchial human airway epithelia. Cell Rep Med 1(4):100059
Sun G-Q, Wang S-F, Li M-T, Li L, Zhang J, Zhang W, Jin Z, Feng G-L (2020) Transmission dynamics of COVID-19 in Wuhan, China: effects of lockdown and medical resources. Nonlinear Dyn 101:1981–1993
Singh AK, Singh A, Singh R, Misra A (2020) Remdesivir in COVID-19: a critical review of pharmacology, pre-clinical and clinical studies. Diabetes Metab Syndr 14(4):641–648
Amirian ES, Levy JK (2020) Current knowledge about the antivirals remdesivir (GS-5734) and GS-441524 as therapeutic options for coronaviruses. One Health 9:100128
Li Y, Zhang D, Gao X, Wang X, Zhang L (2022) 2′-and 3′-ribose modifications of nucleotide analogues establish the structural basis to inhibit the viral replication of SARS-CoV-2. J Phys Chem Lett 13(18):4111–4118
Wakchaure PD, Ghosh S, Ganguly B (2020) Revealing the inhibition mechanism of RNA-dependent RNA polymerase (RdRp) of SARS-CoV-2 by remdesivir and nucleotide analogues: a molecular dynamics simulation study. J Phys Chem B 124(47):10641–10652
Ionescu MI (2020) An overview of the crystallized structures of the SARS-CoV-2. Protein J 39(6):600–618
Padhi AK, Shukla R, Saudagar P, Tripathi T (2021) High-throughput rational design of the remdesivir binding site in the RdRp of SARS-CoV-2: implications for potential resistance. Iscience 24(1):101992
Moirangthem DS, Surbala L (2021) Remdesivir (GS-5734) in COVID-19 therapy: the fourth chance. Curr Drug Targets 22(12):1346–1356
Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, Spitters C, Ericson K, Wilkerson S, Tural A (2020) First case of 2019 novel coronavirus in the United States. N Engl J Med 382:929–936
Sanchez-Codez MI, Rodriguez-Gonzalez M, Gutierrez-Rosa I (2021) Severe sinus bradycardia associated with remdesivir in a child with severe SARS-CoV-2 infection. Eur J Pediatr 180(5):1627–1627
Saqrane S, El Mhammedi M, Lahrich S, Laghrib F, El Bouabi Y, Farahi A, Bakasse M (2021) Recent knowledge in favor of remdesivir (GS-5734) as a therapeutic option for the COVID-19 infections. J Infect Public Health 14(5):655–660
Ko W-C, Rolain J-M, Lee N-Y, Chen P-L, Huang C-T, Lee P-I, Hsueh P-R (2020) Arguments in favour of remdesivir for treating SARS-CoV-2 infections. Int J Antimicrob Agents 55:105933
Belhadi D, Peiffer-Smadja N, Lescure F-X, Yazdanpanah Y, Mentré F, Laouénan C (2020) A brief review of antiviral drugs evaluated in registered clinical trials for COVID-19. MedRxiv 2020:20038190
Warren TK, Jordan R, Lo MK, Ray AS, Mackman RL, Soloveva V, Siegel D, Perron M, Bannister R, Hui HC (2016) Therapeutic efficacy of the small molecule GS-5734 against Ebola virus in rhesus monkeys. Nature 531(7594):381–385
Anastasiou IA, Eleftheriadou I, Tentolouris A, Tsilingiris D, Tentolouris N (2020) In vitro data of current therapies for SARS-CoV-2. Curr Med Chem 27(27):4542–4548
Qiao Y, Wotring JW, Zhang CJ, Jiang X, Xiao L, Watt A, Gattis D, Scandalis E, Freier S, Zheng Y (2023) Antisense oligonucleotides to therapeutically target SARS-CoV-2 infection. PLoS One 18(2):e0281281
Su X, Ma W, Feng D, Cheng B, Wang Q, Guo Z, Zhou D, Tang X (2021) Efficient inhibition of SARS-CoV-2 using chimeric antisense oligonucleotides through RNase L activation. Angew Chem 133(40):21830–21835
Sahu B, Behera SK, Das R, Dalvi T, Chowdhury A, Dewangan B, Kalia K, Shard A (2022) Design and in-silico screening of peptide nucleic acid (PNA) inspired novel pronucleotide scaffolds targeting COVID-19. Curr Comput Aided Drug Des 18(1):26–40
Good SS, Westover J, Jung KH, Zhou X-J, Moussa A, La Colla P, Collu G, Canard B, Sommadossi J-P (2021) AT-527, a double prodrug of a guanosine nucleotide analog, is a potent inhibitor of SARS-CoV-2 in vitro and a promising oral antiviral for treatment of COVID-19. Antimicrob Agents Chemother 65(4):e02479-20
Basu D, Chavda VP, Mehta AA (2022) Therapeutics for COVID-19 and post COVID-19 complications: an update. Curr Res Pharmacol Drug Discov 2022:100086
Mungur O, Berliba E, Bourgeois S, Cardona M, Jucov A, Good S, Moussa A, Pietropaolo K, Zhou X-J, Brown N (2020) A combination of AT-527, a pan-genotypic guanosine nucleotide prodrug, and daclatasvir was well-tolerated and effective in HCV-infected subjects. J Hepatol 73:S357
Berliba E, Bogus M, Vanhoutte F, Berghmans P-J, Good SS, Moussa A, Pietropaolo K, Murphy RL, Zhou X-J, Sommadossi J-P (2019) Safety, pharmacokinetics, and antiviral activity of AT-527, a novel purine nucleotide prodrug, in hepatitis C virus-infected subjects with or without cirrhosis. Antimicrob Agents Chemother 63(12):e01201-19
Good SS, Moussa A, Zhou X-J, Pietropaolo K, Sommadossi J-P (2020) Preclinical evaluation of AT-527, a novel guanosine nucleotide prodrug with potent, pan-genotypic activity against hepatitis C virus. PLoS One 15(1):e0227104
Wang X, Sacramento CQ, Jockusch S, Chaves OA, Tao C, Fintelman-Rodrigues N, Chien M, Temerozo JR, Li X, Kumar S (2021) Combination of antiviral drugs to inhibit SARS-CoV-2 polymerase and exonuclease as potential COVID-19 therapeutics. bioRxiv
Eltayb WA, Abdalla M, Rabie AM (2023) Novel investigational anti-SARS-CoV-2 agent Ensitrelvir “S-217622”: a very promising potential universal broad-spectrum antiviral at the therapeutic frontline of coronavirus species. ACS Omega 8(6):5234–5246
Canard B, Shannon A, Fattorini V, Sama B, Selisko B, Feracci M, Falcou C, Gauffre P, El-Kazzi P, Decroly E (2021) A dual mechanism of action of AT-527 against SARS-CoV-2 polymerase
Shannon A, Fattorini V, Sama B, Selisko B, Feracci M, Falcou C, Gauffre P, Kazzi PE, Decroly E, Rabah N (2021) Protein-primed RNA synthesis in SARS-CoVs and structural basis for inhibition by AT-527. bioRxiv 2021:436564
Mayhoub AS (2012) Hepatitis C RNA-dependent RNA polymerase inhibitors: a review of structure–activity and resistance relationships; different scaffolds and mutations. Bioorg Med Chem 20(10):3150–3161
Elfiky AA, Elshemey WM (2016) IDX-184 is a superior HCV direct-acting antiviral drug: a QSAR study. Med Chem Res 25:1005–1008
Elfiky AA (2017) Zika virus: novel guanosine derivatives revealed strong binding and possible inhibition of the polymerase. Future Virol 12(12):721–728
Elfiky AA, Ismail AM (2018) Molecular docking revealed the binding of nucleotide/side inhibitors to Zika viral polymerase solved structures. SAR QSAR Environ Res 29(5):409–418
Noor R (2021) Antiviral drugs against severe acute respiratory syndrome coronavirus 2 infection triggering the coronavirus disease-19 pandemic. Tzu-Chi Med J 33(1):7
Elfiky AA, Ismail AM (2017) Molecular modeling and docking revealed superiority of IDX-184 as HCV polymerase inhibitor. Future Virol 12(7):339–347
Zhou X-J, Pietropaolo K, Chen J, Khan S, Sullivan-Bólyai J, Mayers D (2011) Safety and pharmacokinetics of IDX184, a liver-targeted nucleotide polymerase inhibitor of hepatitis C virus, in healthy subjects. Antimicrob Agents Chemother 55(1):76–81
Elfiky AA (2022) Dual targeting of RdRps of SARS-CoV-2 and the mucormycosis-causing fungus: an in silico perspective. Future Microbiol 17(10):755–762
Murakami E, Niu C, Bao H, MicolochickSteuer HM, Whitaker T, Nachman T, Sofia MA, Wang P, Otto MJ, Furman PA (2008) The mechanism of action of β-d-2′-deoxy-2′-fluoro-2′-C-methylcytidine involves a second metabolic pathway leading to β-d-2′-deoxy-2′-fluoro-2′-C-methyluridine 5′-triphosphate, a potent inhibitor of the hepatitis C virus RNA-dependent RNA polymerase. Antimicrob Agents Chemother 52(2):458–464
Zhang C (2022) Fluorine in medicinal chemistry: in perspective to COVID-19. ACS Omega 7(22):18206–18212
Ma H, Jiang W-R, Robledo N, Leveque V, Ali S, Lara-Jaime T, Masjedizadeh M, Smith DB, Cammack N, Klumpp K (2007) Characterization of the metabolic activation of hepatitis C virus nucleoside inhibitor β-d-2′-deoxy-2′-fluoro-2′-C-methylcytidine (PSI-6130) and identification of a novel active 5′-triphosphate species. J Biol Chem 282(41):29812–29820
Sofia MJ, Furman PA (2019) The discovery of sofosbuvir: a liver-targeted nucleotide prodrug for the treatment and cure of HCV. HCV I:141–169
Sayad B, Sobhani M, Khodarahmi R (2020) Sofosbuvir as repurposed antiviral drug against COVID-19: why were we convinced to evaluate the drug in a registered/approved clinical trial? Arch Med Res 51(6):577–581
Murakami E, Tolstykh T, Bao H, Niu C, Steuer HMM, Bao D, Chang W, Espiritu C, Bansal S, Lam AM (2010) Mechanism of activation of PSI-7851 and its diastereoisomer PSI-7977. J Biol Chem 285(45):34337–34347
Kirby BJ, Symonds WT, Kearney BP, Mathias AA (2015) Pharmacokinetic, pharmacodynamic, and drug-interaction profile of the hepatitis C virus NS5B polymerase inhibitor sofosbuvir. Clin Pharmacokinet 54:677–690
Gentile I, Maraolo AE, Buonomo AR, Zappulo E, Borgia G (2015) The discovery of sofosbuvir: a revolution for therapy of chronic hepatitis C. Expert Opin Drug Discov 10(12):1363–1377
Bhatia R, Narang RK, Rawal RK (2020) Repurposing of RdRp inhibitors against SARS-CoV-2 through molecular docking tools. Coronaviruses 1(1):108–116
Wang Y, Anirudhan V, Du R, Cui Q, Rong L (2021) RNA-dependent RNA polymerase of SARS-CoV-2 as a therapeutic target. J Med Virol 93(1):300–310
Chien M, Anderson TK, Jockusch S, Tao C, Li X, Kumar S, Russo JJ, Kirchdoerfer RN, Ju J (2020) Nucleotide analogues as inhibitors of SARS-CoV-2 polymerase, a key drug target for COVID-19. J Proteome Res 19(11):4690–4697
Nourian A, Khalili H, Ahmadinejad Z, Kouchak HE, Jafari S, Manshadi SAD, Rasolinejad M, Kebriaeezadeh A (2020) Efficacy and safety of sofosbuvir/ledipasvir in treatment of patients with COVID-19; a randomized clinical trial. Acta Bio Medica Atenei Parmensis 91(4)
Jockusch S, Tao C, Li X, Chien M, Kumar S, Morozova I, Kalachikov S, Russo JJ, Ju J (2020) Sofosbuvir terminated RNA is more resistant to SARS-CoV-2 proofreader than RNA terminated by remdesivir. Sci Rep 10(1):16577
Choudhury C, Narahari Sastry G (2019) Pharmacophore modelling and screening: concepts, recent developments and applications in rational drug design. In Structural bioinformatics: applications in preclinical drug discovery process, pp 25–53
Mosayebnia M, HajiaghaBozorgi A, Rezaeianpour M, Kobarfard F (2022) In silico prediction of SARS-CoV-2 main protease and polymerase inhibitors: 3D-pharmacophore modelling. J Biomol Struct Dyn 40(14):6569–6586
Giordano D, Biancaniello C, Argenio MA, Facchiano A (2022) Drug design by pharmacophore and virtual screening approach. Pharmaceuticals 15(5):646
Schaller D, Šribar D, Noonan T, Deng L, Nguyen TN, Pach S, Machalz D, Bermudez M, Wolber G (2020) Next generation 3D pharmacophore modeling. Wiley Interdiscip Rev Comput Mol Sci 10(4):e1468
Aouidate A, Ghaleb A, Chtita S, Aarjane M, Ousaa A, Maghat H, Sbai A, Choukrad MB, Bouachrine M, Lakhlifi T (2021) Identification of a novel dual-target scaffold for 3CLpro and RdRp proteins of SARS-CoV-2 using 3D-similarity search, molecular docking, molecular dynamics and ADMET evaluation. J Biomol Struct Dyn 39(12):4522–4535
Singh S, Banavath HN, Godara P, Naik B, Srivastava V, Prusty D (2022) Identification of antiviral peptide inhibitors for receptor binding domain of SARS-CoV-2 omicron and its sub-variants: an in-silico approach. 3 Biotech 12(9):198
Pundir H, Joshi T, Pant M, Bhat S, Pandey J, Chandra S, Tamta S (2022) Identification of SARS-CoV-2 RNA dependent RNA polymerase inhibitors using pharmacophore modelling, molecular docking and molecular dynamics simulation approaches. J Biomol Struct Dyn 40(24):13366–13377
Aziz S, Waqas M, Mohanta TK, Halim SA, Iqbal A, Ali A, Khalid A, Abdalla AN, Khan A, Al-Harrasi A (2023) Identifying non-nucleoside inhibitors of RNA-dependent RNA-polymerase of SARS-CoV-2 through per-residue energy decomposition-based pharmacophore modeling, molecular docking, and molecular dynamics simulation. J Infect Public Health 16(4):501–519
Brunt D, Lakernick PM, Wu C (2022) Discovering new potential inhibitors to SARS-CoV-2 RNA dependent RNA polymerase (RdRp) using high throughput virtual screening and molecular dynamics simulations. Sci Rep 12(1):19986
Qaisar M, Muhammad S, Iqbal J, Khera RA, Al-Sehemi AG, Alarfaji SS, Khalid M, Hussain F (2022) Identification of marine fungi-based antiviral agents as potential inhibitors of SARS-CoV-2 by molecular docking, ADMET and molecular dynamic study. J Comput Biophys Chem 21(02):139–153
Velavan TP, Meyer CG (2020) The COVID-19 epidemic. Trop Med Int Health 25(3):278
Lauring AS, Tenforde MW, Chappell JD, Gaglani M, Ginde AA, McNeal T, Ghamande S, Douin DJ, Talbot HK, Casey JD (2022) Clinical severity of, and effectiveness of mRNA vaccines against, COVID-19 from Omicron, Delta, and Alpha SARS-CoV-2 variants in the United States: prospective observational study. BMJ 376
Ndwandwe D, Wiysonge CS (2021) COVID-19 vaccines. Curr Opin Immunol 71:111–116
Ng WH, Liu X, Mahalingam S (2020) Development of vaccines for SARS-CoV-2. F1000Research 9
Kremer EJ (2020) Pros and cons of adenovirus-based SARS-CoV-2 vaccines. Elsevier, Amsterdam, pp 2303–2304
Wang G, Dyatkina N, Prhavc M, Williams C, Serebryany V, Hu Y, Huang Y, Wan J, Wu X, Deval J (2019) Synthesis and anti-HCV activities of 4′-fluoro-2′-substituted uridine triphosphates and nucleotide prodrugs: discovery of 4′-fluoro-2′-C-methyluridine 5′-phosphoramidate prodrug (AL-335) for the treatment of hepatitis C infection. J Med Chem 62(9):4555–4570
Wang G, Dyatkina N, Prhavc M, Williams C, Serebryany V, Hu Y, Huang Y, Wu X, Chen T, Huang W (2020) Synthesis and anti-HCV activity of sugar-modified guanosine analogues: discovery of AL-611 as an HCV NS5B polymerase inhibitor for the treatment of chronic hepatitis C. J Med Chem 63(18):10380–10395
Tchesnokov EP, Feng JY, Porter DP, Götte M (2019) Mechanism of inhibition of Ebola virus RNA-dependent RNA polymerase by remdesivir. Viruses 11(4):326
Zhang L, Zhou R (2020) Structural basis of the potential binding mechanism of remdesivir to SARS-CoV-2 RNA-dependent RNA polymerase. J Phys Chem B 124(32):6955–6962
Qudsiani KS, Rahmasari R (2021) Polyamidoamine-remdesivir conjugate: physical stability and cellular uptake enhancement. Biomed Pharmacol J 14(4):2073–2084
Ning S, Yu B, Wang Y, Wang F (2021) SARS-CoV-2: origin, evolution, and targeting inhibition. Front Cell Infect Microbiol 11:676451
Serpi M, Pertusati F (2021) An overview of ProTide technology and its implications to drug discovery. Expert Opin Drug Discov 16(10):1149–1161
Menéndez JC (2022) Approaches to the potential therapy of COVID-19: a general overview from the medicinal chemistry perspective. Molecules 27(3):658
Hassanipour S, Arab-Zozani M, Amani B, Heidarzad F, Fathalipour M, Martinez-de-Hoyo R (2021) The efficacy and safety of favipiravir in treatment of COVID-19: a systematic review and meta-analysis of clinical trials. Sci Rep 11(1):11022
Malone B, Campbell EA (2021) Molnupiravir: coding for catastrophe. Nat Struct Mol Biol 28(9):706–708
Mishra A, Rathore AS (2022) Pharmacophore screening to identify natural origin compounds to target RNA-dependent RNA polymerase (RdRp) of SARS-CoV2. Mol Divers 2022:1–17
Gao C, Chang L, Xu Z, Yan X-F, Ding C, Zhao F, Wu X, Feng L-S (2019) Recent advances of tetrazole derivatives as potential anti-tubercular and anti-malarial agents. Eur J Med Chem 163:404–412
Pokhodylo N, Finiuk N, Klyuchivska O, Stoika R, Matiychuk V, Obushak M (2023) Bioisosteric replacement of 1H–1, 2, 3-triazole with 1H-tetrazole ring enhances anti-leukemic activity of (5-benzylthiazol-2-yl) benzamides. Eur J Med Chem 250:115126
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (32201053) and the Beijing Institute of Technology Research Fund Program for Young Scholars (3100012222222). The image of the graphical abstract was created with tools obtained from BioRender.com.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest, financial or otherwise.
Consent for Publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shehzadi, K., Saba, A., Yu, M. et al. Structure-Based Drug Design of RdRp Inhibitors against SARS-CoV-2. Top Curr Chem (Z) 381, 22 (2023). https://doi.org/10.1007/s41061-023-00432-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s41061-023-00432-x