Abstract
Chitin is the main component of the cell wall of plant pathogenic fungi. Chitinase 42 (Chit42) from Trichoderma atroviride (PTCC5220) plays a significant role in the biocontrol activity of this fungus against fungal pathogens. This enzyme lacks a chitin binding domain (ChBD) which is involved in its binding to crystalline chitin. In this research, a chimeric chitinase (Chit42+ ChBD) containing a strong chitin binding capacity was constructed by fusing a ChBD from chitinase 18-10 to Chit42 both from isolate PTCC5220. The construct was cloned and overexpressed in Escherichia coli BL21 (DE3). The fusion of ChBD improved the affinity to crystalline and colloidal chitin and also the thermal and chemical stability of the chimeric chitinase, when compared with the native Chit42. In vitro assays indicated that the chimeric chitinase showed higher antifungal activity toward plant pathogenic fungi.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chitin, the second most plentiful polysaccharide in nature, is an insoluble β-1,4-linked polymer of N-acetylglucosamine and the main component of insect exoskeletons, crustacean shells and the cell walls of fungi (Aam et al. 2010). It is degraded by chitinases (Duo-Chuan 2006) that are found in bacteria, fungi, viruses and higher plants (Aronson et al. 2003; Duo-Chuan 2006). Strains from the genus Trichoderma have been described as antagonists able to control a wide range of plant pathogenic fungi such as Botrytis cinerea, Fusarium spp., Pythium spp., Rhizoctonia solani, Sclerotium rolfsii, and Sclerotinia sclerotiorum (Chet et al. 1998; Howell 2003). Trichoderma acts mainly through mycoparasitism (Howell 2003). Mycoparasites produce cell wall hydrolytic enzymes (De la Cruz et al. 1995) amongst which chitinases are considered as one of the most important in the degradation of the fungal cell walls, with a key role in biocontrol (Guthrie et al. 2005; Limón and Codón 2004). Some chitinases contain a chitin-binding domain (ChBD) linked to the catalytic site via a linker region (Limon et al. 2004; Arakane et al. 2003). The ChBD is a tunnel-like structure that simplifies binding to non-colloidal chitin, thus improving the degradation of chitin (Hardt and Laine 2004; Van Aalten et al. 2001). Only a few fungal chitinases have been shown to contain a ChBD (Seidl et al. 2005; Fan et al. 2007). The most important chitinase secreted by Trichoderma sp. is Chit42, which is considered as such due to its role in biocontrol (Hayes et al. 1994). This enzyme lacks a ChBD and has no efficient effect on crystalline chitin. The activity of this enzyme can be improved by addition of a ChBD via protein engineering (Blaak and Schrempf 1995). Previous studies have shown that the fusion of a ChBD to chitinase improves the mycoparasitic activity (Boldo et al. 2009; Fan et al. 2007; Limon et al. 1999).
In this research, we have constructed a chimeric chitinase by adding a ChBD from T. atroviride chitinase 18–10 to the N-terminal end of T. atroviride Chit42, so as to improve its enzyme activity. Native Chit42 and the chimeric chitinase were cloned and overexpressed in Escherichia coli, and the expression of the chimeric chitinase was then optimized by the Taguchi method. In order to study the effect of ChBD on the novel chimeric chitinase, some features such as enzyme activity, pH and optimal temperature, thermal and chemical stability were investigated. The activity of both native and chimeric chitinases was studied in vitro against different plant pathogenic fungi.
Materials and methods
Plasmids, fungal and bacterial isolates
Escherichia coli strain DH5α and cloning vector pJET 1.2 (Novagen) were used for cloning and E. coli BL21 (DE3) strain and expression vector pET26b (+) (Novagen) were used for the prokaryotic expression experiments. The five fungal plant pathogens: R. solani, F. oxysporum, S. sclerotiurum, Alternaria solani, and Verticilium dahliae were provided by H. Afshari-Azad, Iranian Research Institute of Plant Protection, Tehran, Iran. T. atroviride (PTCC5220) from the Persian Type Culture Collection and was identified at NIGEB, Tehran, I.R, as an overproducer of chitinase among 31 different isolates of Trichoderma sp. (Harighi et al. 2006a, b). The fungi were grown on potato dextrose agar (PDA) medium and sub-cultured as needed. All E. coli strains were grown in Luria-Bertani (LB) broth at 37 °C and the media were supplemented with ampicillin and kanamycin (SIGMA, 100 and 50 mg/ml), respectively.
Construction of recombinant plasmid
The Chit42 from T. atroviride (access number DQ022674) fused to ChBD of chitinase 18-10 (AAZ23945.1) was amplified using Pfu polymerase (Fermentas) with specific primers (AchBD42, 5′-GGAAGACAACATGAAATGCGGTCCTCAGGT-3′ and CDP42, 5′-CGCTCGAGGTTGAGACCGCTTCGGA-3′) which contain BpiI and XhoI sites at their 5′ ends respectively. The amplified fragment was purified with the PCR fragment Recovery Kit (iNtRON Biotechnology) and cloned into pET-26b(+) at the NcoI/XhoI restriction sites, in frame with His-tag and under the control of T7 promoter to yield pETAT1 (Fig. 1a). The sequence encoding the chimeric chitinase was confirmed by sequencing.
(a) Schematic representation of the pETSM2 vector (containing the Chit42 gene) and pETAT1 vector (containing the chimeric chitinase). pETAT1 was constructed by amplifying the ChBD, linker region and the mature Chit42. The arrows indicate specific primers (AchBD42 and CDP42) of the chimeric chitinase. (b) SDS-PAGE analysis of chimeric chitinase expression in recombinant E. coli. 1: Chimeric chitinase 2: Protein extraction from E. coli BL21 (DE3) harboring the empty pET-26b(+) as a negative control. (c) Western blot analysis of the chimeric chitinase using anti His-tag, 1: Chimeric chitinase 2: Protein extraction from E. coli BL21 (DE3) harboring the empty pET-26b(+) as a negative control. M: molecular weight marker. The arrow indicates the chimeric chitinase (52 KDa)
Expression of the chimeric chitinase in Escherichia coli
Recombinant plasmid pETAT1 was transformed into E. coli BL21 (DE3) and selected on LB medium containing 50 μg/ml of kanamycin. An overnight pre-culture of a single colony was used to inoculate 100 ml of LB media containing the appropriate antibiotics and grown at 37 °C until an optical density (OD600) of 0.6 was reached. Protein expression was induced by adding 1 mM isopropyl-1-thio-β-D-galactopyranoside (IPTG) to the culture and the cells were shaken at 37 °C for 2, 4, 6, and 16 h. Expression was visualized by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE, 12 %).
Western blotting analysis
For immunodetection of the expressed chimeric chitinase, total protein was extracted from induced E. coli BL21 (DE3). Protein samples were electrophoresed, followed by electrotransfer to a polyvinylidene fluoride (PVDF) membrane. The immunoblots were developed with antibody against His-tag, according to the manufacturer’s instructions (Roche).
Protein expression optimization by Taguchi’s method
For optimization of the chimeric Chit42 expression in the prokaryotic system, the experiments were designed using the statistical Taguchi method. In this method, the effect of three factors including IPTG concentration (mM), incubation temperature (°C), and incubation time (h) were studied at four levels on protein production. According to the considered factors, 16 independent experiments were designed. After these tests were carried out, the total protein was extracted from the bacteria and visualized using SDS-PAGE. The content of the overexpressed chimeric chitinase in each fraction was quantified with a Bio-Rad GS-800 gel densitometer. The density of chimeric protein expression was calculated relative to the reference band (total cell protein of E. coli). After analyzing and quantifying the data, the results were analyzed using the Qualitek-4 software (version 08.10; http://www.nutek-us.com) and the most important effect of different levels of each factor and the optimal expression conditions were then determined. Analysis of variance (ANOVA) was used for statistical analysis.
Enzyme assay
The activity of the chimeric chitinase and Chit42 was assayed with a calorimetric method using colloidal and non-colloidal chitin as substrates. A reaction mixture (total volume of 500 μl) containing chitin as a substrate (3.8 mg), citrate buffer (pH 5), and the appropriate amount of enzymes was incubated at 37 °C for 1 h. Thereafter, the reaction was stopped by immersing the mixture in boiling water for 10 min. The amount of reducing sugar produced was evaluated using N-acetyl-D-glucosamine as a standard. The production of 1 μmol of the product/min was considered as one unit of enzyme activity (Ulhoa and Peberdy 1992).
pH and temperature optima and thermal stability
Optimized pH was determined by incubating the enzymes with colloidal chitin in the buffer at different pHs (3 to 9). Buffers used were sodium citrate (pH 3 to 7), sodium phosphate (pH 8), and sodium carbonate (pH 9). The optimum temperature was determined by carrying out standard assays with colloidal chitin at temperatures ranging from 10 to 90 °C. Thermal stability was also determined by incubating the enzymes for 1 h at temperatures ranging from 10 to 70 °C in sodium acetate buffer (50 mM, pH 5) and then measuring the remaining activity at 37 °C. Each experiment was carried out with three replicates.
Protein denaturation studies
Denaturation curves of the chimeric chitinase and Chit42 (0.2 mg/ml) in the presence of dodecyltrimethylamonnium bromide (DTAB) were performed by measuring the change in optical density at 280 nm. All measurements were carried out after 5 min of DTAB incubation with enzymes in a 1-cm cuvette, thermostable at 20 ± 0.1 °C, when there were no further changes in the optical density. The standard Gibbs free energy of denaturation (∆G°) was used as a criterion of conformational stability (Gheibi et al. 2006).
Antifungal activity
Radial diffusion assays
Fungal growth assays were used to evaluate the antifungal activity of the chimeric chitinase and Chit42. Radial diffusion assays were performed using 100 × 15 mm plates containing 25 ml of PDA. Equal aliquots (90 μg/ml) of chimeric chitinase and Chit42 were added to 5 mm diameter holes punched on the agar surface at 1 cm from the edge of fungal colonies grown for 24 h at 28 °C. The plates were incubated at 28 °C for 24 h until the fungal colonies reached the negative control (10 min boiled enzyme, protein free buffer, and empty pET-26b(+) total protein). The fungal species included S. sclerotiorum, F. oxysporum, R. solani, and V. dahliae.
Spore germination assays
Spore suspensions of A. solani, F. oxysporum, and V. dahliae (2 × 104 cell/ml) in half strength potato dextrose broth (PDB) containing native (70 μg/ml) and chimeric chitinases (70 μg/ml) were incubated at 28 °C with shaking (150 rpm) for 48 h. Crude protein of the empty vector (pET-26b(+) without the chimeric chitinase gene) was used as control. After 48 h of incubation, fungal growth was determined by measuring OD values at 595 nm. The experiments were conducted three times.
Results
Heterologous expression in E. coli
The main goal of this study was to express a new chimeric chitinase containing a ChBD and the native chitinase (Chit42) in E. coli to evaluate the role of ChBD in the antifungal enzyme activity. The ChBD from T. atroviride chitinase 18-10 was fused to Chit42 of T. atroviride. Expressed protein was tagged with 6 × His-tag at the C-terminal end to facilitate detection by Western blotting. The molecular weight of the chimeric chitinase was calculated as a band close to 52 kDa. Analysis of the SDS-PAGE revealed the expected band in the induced conditions (Fig. 1b).
Western blotting
Western blot analysis using anti His-tag confirmed the recombinant protein containing 6 × His with the expected molecular weight for the chimeric chitinase (52 KDa) in its crude form (Fig. 1c). Total cell protein of E. coli BL21 (DE3) harboring empty pET-26b (+) was used as a negative control.
Protein expression optimization
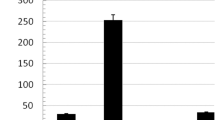
In order to optimize the recombinant protein expression, an LM16 orthogonal experimental design was used to study the effect of incubation temperature, time and IPTG concentration on the amount of protein expression. The 52 kDa protein band was observed by SDS-PAGE and the amount of expressed protein was estimated using densitometry (Fig. 2). The effect of different levels of each factor on protein expression was determined by the Qualitek-4 software. According to ANOVA, incubation temperature was the most effective in protein production and IPTG concentration and incubation time were placed in second and third. Optimal conditions as determined by Qualitek-4 were 4 h of induction at 33 °C by 0.5 mM IPTG.
Enzyme assay
The activities of the chimeric and native Chit42 were compared in the presence of insoluble and colloidal chitin. Chitinase activities for the same concentration of chimeric and Chit42 in the presence of colloidal chitin were 11.7 ± 0.2 and 13.02 ± 0.1 U/ml and in the presence of insoluble chitin were 6.1 ± 0.3 and 12.35 ± 0.2 U/ml, respectively.
pH, temperature profiles, and stability
The highest binding activity of the chimeric chitinase, which is higher than that of Chit42, was detected at 40 °C and pH 4 (Fig. 3a and b). The binding activity of both enzymes significantly decreased with increasing pHs and temperatures above or below 40 oC.
Enzyme kinetics of Chit42 and Chimeric chitinase. (a) Profiles of chitinase activity at different temperatures and (b) different pH values. (c) Enzyme activity after incubation of each enzyme at various temperatures for 1 h. (d) Free energy changes for the chemical denaturation of the chimeric chitinase and Chit42 in the presence of different concentrations of DTAB (mM). Colloidal chitin was used as a substrate in all experiments. Enzyme activities of Chit42 and chimeric chitinase were measured in citrate buffer (0.2 M, pH 5)
Chemical and thermal stability assays demonstrated that the chimeric chitinase had a stronger structure and higher stability (Fig. 3c and d). As a result of these assays, the chimeric chitinase maintained ~25 % of its activity at 70 °C, but Chit42 retained only ~1 %. Also, denaturation studies based on DTAB demonstrated that the chimeric enzyme could tolerate a higher concentration of DTAB when compared with Chit42.
In vitro antifungal activity assays
Total cell proteins were used in the radial diffusion assay. It was revealed that both enzymes showed antifungal activity against S. sclerotiorum, R. solani, F. oxysporum, and V. dahliae (Fig. 4a). The antifungal activity of the chimeric chitinase was shown to be higher than that of native Chit42. Significant differences were observed between the chitinase enzymes and the negative controls [total cell protein from E. coli BL21 (DE3) containing empty pET26b (+)] on spore inhibition. These results showed that spore inhibition in A. solani, F. oxysporum, and V. dahliae using extracted enzymes containing the chimeric chitinase was 58, 37, and 57 % and that of Chit42 was 44, 27, and 38 %, respectively (Fig. 4b).
(a) In vitro antifungal activities of the chimeric chitinase and Chit42 against (a) S. sclerotiorum, (b) F. oxysporum, (c) R. solani, (d) V. dahliae. Approximately 90 μg/ml of Chit42 and chimeric chitinase were applied to the wells numbered 1 and 2, respectively. Inactivated chimeric chitinase, extraction buffer and total cell protein from E. coli BL21 (DE3) harboring the empty pET-26b (+) were tested as controls in the wells numbered 3, 4 and 5, respectively. (b) Growth inhibition (%) of Chit42 and chimeric chitinase on spore germination. Growth of A. solani, F. oxysporum and V. dahliae in crude protein extracts from E. coli BL21 (DE3) harboring chimeric chitinase and Chit42. Absorbance of the reaction mixture (crude protein extract + spore suspension + PDB) after 48 h of incubation was measured at 595 nm. Bars represent means and standard errors and those with different lower case letters are significantly different (p > 0.05) according to the least significant difference (LSD) test. Results represent the average of three experiments
Discussion
In the present work, in order to improve activity of Trichoderma atroviride PTCC5220 Chit42, a ChBD from chitinase 18-10 belonging to the same Trichoderma strain, was fused to the native Chit42. Chitinase 18-10 naturally contains a ChBD at its N-terminal (Seidl et al. 2005). To study the properties of Chit42 and the chimeric chitinase, such as enzyme activity, pH and optimal temperature, thermal and chemical stability, and antifungal activity, both enzymes were successfully cloned and produced in E. coli. This prokaryotic system has been used to produce several recombinant proteins, because of easy handling, inexpensive media, and large-scale production (Makrides 1996). Expression of the chimeric chitinase protein was optimized and improved using the Taguchi method and optimal expression conditions were found to be 4 h, 0.5 mM IPTG, and 33 °C. The expected band (52 KDa) was observed by SDS-PAGE and confirmed by Western blotting.
In this research, the chimeric chitinase showed higher activity than the native enzyme, especially in the presence of crystalline chitin. In a similar study, the effect of a ChBD on chitin binding was also described by Hashimoto et al. (2000). They showed that deletion of the ChBD from chitinase A1 decreased the efficacy of chitin degradation. The effect of ChBD on chitin binding was also reported by Matroodi et al. (2013a, b)) where they showed that fusion of a ChBD from Serratia marcescens to Chit42 greatly increased the activity of the chimeric enzyme on hydrolyzed insoluble chitin. The same finding was also reported by Limon et al. (2001) who demonstrated that the addition of a ChBD from Nicotinia tabacum to Chit42 of T. harzianum improved the activity of the native enzyme.
Earlier reports demonstrated that fungal chitinase activity at an acidic pH is higher than at a basic pH, for example, pH 4 for chitinases of T. harzianum ATCC74058 (Harman et al. 1993), pH 4 to 6 for Gliocladium virens ATCC20906 (Di Pietro et al. 1993), pH 4 for Verticillium lecanii A3 (Fenice et al. 1998), pH 6.2 for Pyrus communis (Sakurada et al. 1996), pH 5.0 for Penicillium oxalicum (Rodriguez et al. 1995) and pH 5.2 for A. carneus (Abdel-Naby et al. 1992), which are similar to the results found in this study. With regard to temperature, similar results have been reported, such as the optimum temperature of 40 °C for chitinases from Bacillus cereus CH2 and Monascus purpureus CCRC31499 (Li et al. 2008; Wang et al. 2002). Similar pH and temperature optima for both Chit42 and chimeric chitinase might be considered as an additional evidence for the hypothesis that the addition of ChBD does not affect the Chit42 active site (Matroodi et al. 2013a, b). Also the reports on thermal and chemical stability of Chit42 containing ChBD indicated certain structural changes. Consequently, its catalytic efficiency, thermal and chemical stability were found to improve (Matroodi et al. 2013a, b).
Chimeric and native Chit42 reported in this study showed antifungal activity against A. solani, F. oxysporum, V. dahliae, S. sclerotirum, and R. solani. However, the chimeric chitinase that is more active towards crystalline chitin displayed higher antifungal activity than Chit42, which seems to result from the subsided structure at the binding cleft of the enzyme (Sasaki et al. 2002). In another research, deletion of ChBD from Streptomyces griseus HUT6037 chitinase C led to moderate reduction in the hydrolyzing activity toward crystalline chitin substrate, however, most of the antifungal activity against Trichoderma reesei was suppressed by this deletion. It appears that the ChBD plays a more important role in antifungal properties than in catalytic activities (Itoh et al. 2002). It was also shown that the antifungal activity of these chitinases was different among phytopathogenic fungi. This variation may be due to the natural variability of chitin in fungal cell walls (Aranaz et al. 2009; Van de Velde and Kiekens 2004).
Our previous work showed that the antifungal activity of a T. harzianum transformed with a Chit42- ChBD against several phytopathogenic fungi was significantly improved relative to the wild type and transformants harboring Chit42 (Kowsari et al. 2014).
The results of the current study showed that the novel chimeric chitinase containing a fungal ChBD is thermochemically stable and active at acidic pH, and thus could be used in industrial processes. This engineered chitinase gene might be able to provide the efficient and cost-effective chitinolytic machinery, which can be expressed in different hosts for field applications and utilized to generate recombinant fungi resistant plants to reduce dependence on chemical fungicides.
References
Aam BB, Heggset EB, Norberg AL, Sørlie M, Vårum KM, Eijsink VG (2010) Production of chitooligosaccharides and their potential applications in medicine. Mar Drugs 8:1482–1517
Abdel-Naby MA, El-Shayeb NM, Sherief A (1992) Purification and some properties of chitinase from Aspergillus carneus. Appl Biochem Biotechnol 37:141–154
Arakane Y, Zhu Q, et al (2003) Properties of catalytic, linker and chitin-binding domains of insect chitinase. Insect Biochem Mol Biol 33(6): 631–648
Aranaz I, Mengíbar M, Harris R, Paños I, Miralles B, Acosta N, Galed G, Heras Á (2009) Functional characterization of chitin and chitosan. Curr Chem Biol 3:203–230
Aronson Jr N, Halloran B, Alexyev M, Amable L, Madura J, Pasupulati L, Worth C, Vanroey P (2003) Family 18 chitinase–oligosaccharide substrate interaction: subsite preference and anomer selectivity of Serratia marcescens chitinase A. Biochem J 376:87–95
Blaak H, Schrempf H (1995) Binding and substrate specificities of a Streptomyces olivaceoviridis chitinase in comparison with its proteolytically processed form. Eur J Biochem 229:132–139
Boldo JT, Junges A, do Amaral KB, Staats CC, Vainstein MH, Schrank A (2009) Endochitinase CHI2 of the biocontrol fungus Metarhizium anisopliae affects its virulence toward the cotton stainer bug Dysdercus peruvianus. Curr Genet 55:551–560
Chet I, Benhamou N, Haran S (1998) Mycoparasitism and lytic enzymes. In: Kubicek Geha CP (ed) Trichoderma and Gliocladium, vol 2. Taylor & Francis, London, pp 153–172
De la Cruz J, Pintor-Toro JA, Benitez T, LLobell A (1995) Purification and characterization of an endo-beta-1, 6-glucanase from Trichoderma harzianum that is related to its mycoparasitism. J Bacteriol 177:1864–1871
Di Pietro A, Lorito M, Hayes C, Broadway R, Harman G (1993) Endochitinase from Guocladium virens: isolation, characterization, and synergistic antifungal activity in combination with cliotoxin. Phytopathology 83:308–313
Duo-Chuan L (2006) Review of fungal chitinases. Mycopathologia 161:345–360
Fan Y, Fang W, Guo S, Pei X, Zhang Y, Xiao Y, Li D, Jin K, Bidochka B, Pei Y (2007) Increased insect virulence in Beauveria bassiana strains overexpressing an engineered chitinase. Appl Environ Microbiol 73:295–302
Fenice M, Selbmann L, Di Giambattista R, Federici F (1998) Chitinolytic activity at low temperature of an Antarctic strain (A3) of Verticillium lecanii. Res Microbiol 149:289–300
Gheibi N, Saboury A, Haghbeen K, Moosavi-Movahedi A (2006) The effect of some osmolytes on the activity and stability of mushroom tyrosinase. J Biosci 31:355–362
Guthrie JL, Khalif S, Castle AJ (2005) An improved method for detection and quantification of chitinase activities. Can J Microbiol 51:491–495
Hardt M, Laine RA (2004) Mutation of active site residues in the chitin-binding domain ChBD from chitinase A1 of Bacillus circulans alters substrate specificity: use of a green fluorescent protein binding assay. Arch Biochem Biophys 426:286–297
Harighi MJ, Motallebi M, Zamani MR (2006a) Antifungal activity of heterologous expressed chitinase 42 (Chit42) from Trichoderma atroviridae PTCC5220. Iran J Biotechnol 4:95–103
Harighi MJ, Motallebi M, Zamani MR (2006b) Purification of chitinase 42 from Trichoderma atroviridae PTCC5220. Iran J Biol 19:203–214
Harman G, Hayes C, Lorito M, Broadway R, Di Pietro A, Peterbauer C, Tronsmo A (1993) Chitinolytic enzymes of Trichoderma harzianum: purification of chitobiosidase and endochitinase. Phytopathology 83:313–318
Hashimoto M, Ikegami T, Seino S, Ohuchi N et al (2000) Expression and characterization of the chitin-binding domain of chitinase A1 from Bacillus circulans WL-12. J Bacteriol 182:3045–3054
Hayes CK, Klemsdal S, et al (1994) Isolation and sequence of an endochitinase-encoding gene from a cDNA library of Trichoderma harzianum." Gene 138(1-2):143–148
Howell C (2003) Mechanisms employed by Trichoderma species in the biological control of plant diseases: the history and evolution of current concepts. Plant Dis 87:4–10
Itoh Y, Kawase T, Nikaidou N, Fukada H, Mitsutomi M, Watanabe T, Itoh Y (2002) Functional analysis of the chitin-binding domain of a family 19 chitinase from Streptomyces griseus HUT6037: substrate-binding affinity and cis-dominant increase of antifungal function. Biosci Biotechnol Biochem 66:1084–1092
Kowsari M, Motallebi M, Zamani M (2014) Protein engineering of Chit42 towards improvement of chitinase and antifungal activities. Curr Microbiol 68:495–502
Li J-G, Jiang Z-Q, Xu L-P, Sun F-F, Guo J-H (2008) Characterization of chitinase secreted by Bacillus cereus strain CH2 and evaluation of its efficacy against Verticillium wilt of eggplant. Biocontrol 53:931–944
Limón MC, Codón AC (2004) Biocontrol mechanisms of Trichoderma strains. Int Microbiol 7:249–260
Limon MC, Pintor-Toro JA, Benítez T (1999) Increased antifungal activity of Trichoderma harzianum transformants that overexpress a 33-kDa chitinase. Phytopathology 89:254–261
Limon MC, Margolles‐Clark E, Benı́tez TA, Penttilä M (2001) Addition of substrate‐binding domains increases substrate‐binding capacity and specific activity of a chitinase from Trichoderma harzianum. FEMS Microbiol Lett 198:57–63
Limon MC, Chacon MR, Mejias R, Delgado-Jarana J, Rincon AM, Codon AC, Benitez T (2004) Increased antifungal and chitinase specific activities of Trichoderma harzianum CECT 2413 by addition of a cellulose binding domain. Appl Microbiol Biotechnol 64:675–685
Makrides SC (1996) Strategies for achieving high-level expression of genes in Escherichia coli. Microbiol Rev 60:512–538
Matroodi S, Motallebi M, Zamani M, Moradyar M (2013a) Designing a new chitinase with more chitin binding and antifungal activity. World J Microbiol Biotechnol 29:1517–1523
Matroodi S, Zamani M, Haghbeen K, Motallebi M, Aminzadeh S (2013b) Physicochemical study of a novel chimeric chitinase with enhanced binding ability. Acta Biochim Biophys Sin 45:845–856
Rodriguez J, Copa‐Patiño J, Pérez‐Leblic M (1995) Purification and properties of a chitinase from Penicillium oxalicum autolysates. Lett Appl Microbiol 20:46–49
Sakurada M, Morgavi DP, Komatani K, Tomita Y, Onodera R (1996) Purification and characteristics of cytosolic chitinase from Piromyces communis OTS1. FEMS Microbiol Lett 137:75–78
Sasaki C, Yokoyama A, Itoh Y, Hashimoto M, Watanabe T, Fukamizo T (2002) Comparative study of the reaction mechanism of family 18 Chitinases from plants and microbes. J Biochem 131:557–564
Seidl V, Huemer B, Seiboth B, Kubicek CP (2005) A complete survey of Trichoderma chitinases reveals three distinct subgroups of family 18 chitinases. FEBS J 272:5923–5939
Ulhoa CJ, Peberdy JF (1992) Purification and some properties of the extracellular chitinase produced by Trichoderma harzianum. Enzym Microb Technol 14:236–240
Van Aalten D, Komander D, Synstad B, Gåseidnes S, Peter M, Eijsink V (2001) Structural insights into the catalytic mechanism of a family 18 exo-chitinase. Proc Natl Acad Sci 98:8979–8984
Van de Velde K, Kiekens P (2004) Structure analysis and degree of substitution of chitin, chitosan and dibutyrylchitin by FT-IR spectroscopy and solid state 13 C NMR. Carbohydr Polym 58:409–416
Wang S-L, Hsiao W-J, Chang W-T (2002) Purification and characterization of an antimicrobial chitinase extracellularly produced by Monascus purpureus CCRC31499 in a shrimp and crab shell powder medium. J Agric Food Chem 50:2249–2255
Acknowledgments
The financial assistance from National Institute of Genetic Engineering and Biotechnology (NIGEB) is acknowledged for the support of this work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Section Editor: Jorge T. de Souza
Rights and permissions
About this article
Cite this article
Ataei, A., Zamani, M., Motallebi, M. et al. Increased antifungal activity of Chit42 from Trichoderma atroviride by addition of a chitin binding domain. Trop. plant pathol. 41, 350–356 (2016). https://doi.org/10.1007/s40858-016-0103-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40858-016-0103-7