Abstract
Purpose of Review
This review provides an overview of the role of dysbiosis (imbalanced gut microbiota) in the maintenance of host homeostasis and immune function and summarizes recent evidence connecting gut microbiota dysbiosis to the development of autoimmune diseases (ADs) (such as rheumatoid arthritis, type 1 diabetes, systemic lupus erythematosus, multiple sclerosis, spondyloarthritis, and irritable bowel syndrome). The potential mechanisms that underlie the host-microbiota interaction are also discussed to evaluate the manipulation of the gut microbiota as a potential therapeutic approach to managing ADs. Additionally, this review addresses current challenges in gut microbiota-host research and provides future recommendations.
Recent Findings
Recent findings suggested that the pathogenesis of ADs appears to be multifaceted involving both genetic and environmental factors. Dysbiosis or imbalanced gut microbiota has been increasingly identified as one of the main environmental factors that can modulate immune responses and contribute to the development of ADs.
Summary
New research has highlighted the significance of gut microbial dysbiosis in the etiology of numerous diseases. Understanding the relationship between the gut microbiota and the host, however, goes beyond taxonomic concerns, demanding multidisciplinary efforts to design new therapeutic approaches that take individual variances into account.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Autoimmune diseases are a group of disorders in which the immune system targets and attacks healthy tissues and cells. These diseases show a wide range of clinical manifestations, impacting various organs in the body [1]. Prominent examples of ADs include inflammatory autoimmune diseases like rheumatoid arthritis (RA) and spondyloarthritis (SpA), primarily affecting the musculoskeletal system, leading to joint inflammation [2]. Additionally, systemic lupus erythematosus (SLE) is known for its systemic nature, affecting multiple organs, including the skin, joints, kidneys, and nervous system [3]. Furthermore, type 1 diabetes (T1D) which targets the insulin-producing beta cells in the pancreas and results in insulin deficiency and associated consequences is another example of ADs [4].
Neuroinflammatory autoimmune diseases such as multiple sclerosis (MS) represent another class of ADs that affect the central nervous system, leading to demyelination, neural damage, and a range of neurological symptoms [5]. Moreover, gastrointestinal autoimmune disorders like irritable bowel syndrome (IBS) which affects 15% of the global population can significantly impact the functionality of the gastrointestinal tract and the overall quality of life of individuals by causing abdominal pain, bloating, diarrhea, and constipation among others [6].
Recent studies have highlighted that certain microbial taxon and their metabolites are linked to the development of ADs [7]. This comprehensive review explores the intricate relationship between gut microbiota and ADs, shedding light on the pathways through which the immune system and gut microbiota dysbiosis are associated. This review also investigates the therapeutic methods that have shown promise in mitigating the impact of ADs by modulating the gut microbiota composition. Additionally, the manuscript discusses the existing challenges faced in microbiota studies and presents strategies to address these challenges.
Gut Microbiota
The human gut harbors a complex community of microorganisms known as the gut microbiota, which plays a significant role in maintaining the host’s physiology [8]. The study of the gut microbiota has gained considerable attention in recent years, particularly with the development of new terms such as the gut-brain, gut-skin, gut-mouth, gut-immunity axes, and other yet-to-be-discovered. The gut microbiota contributes to the development of the immune system through different mechanisms, including the maintenance of the intestinal barrier and the maturation and regulation of immune cells through the production of short-chain fatty acids (SCFAs). SCFA-producing bacteria have the ability to regulate immune cell differentiation and the development of regulatory T cells (Tregs), which are critical for maintaining immune homeostasis and controlling immune responses [9].
The initial evidence of the role of gut microbiota dysbiosis in the pathogenesis of ADs developed from germ-free models that lacked gut microbe composition and did not develop ADs [10]. Follow-up studies on transferring fecal microbiota from AD individuals to healthy mice, which triggered the development of autoimmune responses, and antibiotic treatments, which prevented the growth of beneficial bacteria, underlined the significance of gut microbiota in the overall health [11, 12].
Dysbiosis can lead to the loss of immune tolerance, over-activation of T cells, and the production of several pro-inflammatory cytokines. These can activate autoimmune responses and contribute to the development of various diseases, including ADs [13]. The prevalence of ADs has been increasing worldwide in recent decades, with over 80 ADs currently recognized, including RA, T1D, SLE, MS, and SpA [14].
Recent findings have shown an association between specific microbial taxa and their metabolites with the development of ADs. For instance, in patients with RA, an overgrowth of Prevotella spp. (such as P. copri) and a reduction in the abundance of Bacteroides, Bifidobacterium, and butyrate-producing bacteria was associated with the production of pro-inflammatory molecules and activation of autoreactive immune cells [7]. Also, high abundance of Ruminococcus gnavus, a microorganism that degrades mucin, has been detected in stool of individuals affected by RA, SpA, and SLE [15, 16]. Moreover, its presence in the ileum has been linked to RA susceptibility associated with specific HLA-DRB1 alleles [17].
Similarly, patients with inflammatory bowel disease (IBD) exhibit a reduced abundance of anti-inflammatory bacteria, such as Faecalibacterium prausnitzii, and an overgrowth of pro-inflammatory bacteria, such as Escherichia coli [18]. Decreased abundance of Lachnospiraceae and Faecalibacterium (involved in the production of anti-inflammatory molecules) and increased abundance of pro-inflammatory bacteria such as Akkermansia spp. have also been implicated in the pathogenesis of multiple sclerosis (MS) [19]. Induction of pro-inflammatory responses by Akkermansia muciniphila and Acinetobacter calcoaceticus isolated from MS patients in monocolonized mice and stimulation of anti-inflammatory IL-10-expressing human CD4+CD25+ T cells and IL-10+FoxP3+ Tregs by Parabacteroides distasonis have also shown the important role of gut microbiota in modulating immune responses [20].
The abundance of specific microbial taxa may also reflect disease severity and could potentially be used as biomarkers to assess the progression and activity of these ADs. For instance, an increased abundance of Lactobacillus salivarius in RA or SLE has been closely associated with higher clinical disease activity scores [21, 22].
Gut microbiota can also regulate other organs remotely by its signals and metabolites. An example of this is the remote control of gut microbial metabolites on the permeability of blood–brain barrier and the development of neuroinflammation in patients with MS [23]. However, relying only on the taxonomy of bacterial communities is insufficient to understand the complex role of gut microbiota dysbiosis in ADs. Recent findings have highlighted the importance of studying microbial metabolites and their interactions with humans using multi-omics methods. Multi-omics approaches offer detailed insight into the gut microbiota-host crosstalk and its impact on ADs (Table 2). For instance, metabolomic profiling has revealed distinct microbial patterns in RA and MS compared to healthy controls [24]. Combining multi-omics data can also pave the way for designing targeted therapeutic interventions aimed at modulating the gut microbiota and improving the health of patients with ADs.
The Complex Interplay
Our current understanding of gut microbiota-associated diseases indicates a complex network that is not fully understood. While evidence has suggested a correlation between dysbiosis and the development of ADs, it is essential to note that this association does not necessarily establish a “cause and effect” relationship, particularly as many of these studies have been conducted on animal models, and the applicability of these models to humans remains unclear [14].
A summary of the most common pathways through which gut microbiota can contribute to the development of ADs is discussed as follows:
Activation of Immune Responses
The dysregulation of immune pathways is the primary mechanism through which gut microbiota dysbiosis can lead to the development and progression of ADs [25]. The gut-associated lymphoid tissue (GALT), comprising different immune cells such as dendritic cells (DCs), macrophages, and innate lymphocytes, serves as the first line of defense. Dysbiosis can trigger abnormal activation of different immune pathways, resulting in the upregulation of pro-inflammatory cytokines (IL-1β, IL-6, TNF-α, IL-17, IL-12, IFN-γ, etc.) and the reduction of anti-inflammatory cytokines (IL-10, IL-4, IL-13, TGF-β, IL-1ra, etc.) [26].
One common pathway in the development of some ADs is the dysregulated function of Th17 cells and the production of IL-17. This pathway contributes to joint inflammation in RA, glandular inflammation and autoantibody production in Sjögren’s syndrome, and neuroinflammation in MS [27].
In T1D, activation of TNF-α, IL-1β, and IL-6 cytokines can lead to beta-cell destruction and disease progression. Furthermore, autoreactive T cells can target beta-cells in T1D and damage the intestinal epithelial cells in celiac disease [28].
Activation of toll-like receptors (TLRs) on DCs and macrophages, leading to the release of IFN-α, IFN-β, IL-6, and IL-23 cytokines, have also a significant role in the development and progression of SLE [29, 30]. In vitro and ex vivo findings have shown that dysbiosis in SLE patients can promote lymphocyte activation and Th17 differentiation, while Bifidobacterium bifidum supplementation can balance the Treg/Th17/Th1 ratio and prevent over-activation of CD4+ lymphocyte [31]. Also, decreased bacterial diversity with a fivefold greater representation of R. gnavus and serum antibodies against Ruminococcus antigens in SLE patients suggest the contribution of gut microbiota dysbiosis in the pathogenesis of lupus nephritis [32]. Similarly, decreased gut microbial diversity and a high abundance of Collinsella (correlated with pro-inflammatory cytokine IL-17A) were associated with RA duration and autoantibody levels, which suggests the potential application of these variations for predicting RA disease status [33]. Also, in RA patients with positive tests for anti-citrullinated protein antibody (ACPA), a decreased microbial diversity and enrichment of Blautia, Akkermansia, and Clostridiales were noted when compared to ACPA-negative individuals [34].
The potential role of gut microbiota on the immune system can also be seen in healthy animal models that received fecal microbiota transplantation (FMT) from lupus-prone mice which led to abnormal activation of plasmacytoid DCs and to the production of pro-inflammatory cytokines [35].
Gut Barrier Function and Permeability
Dysbiosis can also increase gut permeability by affecting the protein complex in tight junctions (occludins and claudins) between the epithelial cells in the gut [36]. To assess the gut permeability, lactulose/mannitol or lactulose/rhamnose tests are mostly used, which measure the urinary excretion of these molecules. A low lactulose/mannitol or lactulose/rhamnose ratio in healthy individuals shows a well-functioning gut barrier with minimal lactulose passage. While an increased ratio observed in patients with MS, RA, T1D, or celiac disease suggests increased gut permeability and impaired intestinal barrier function [36,37,38]. Increased intestinal permeability, known as leaky gut syndrome, allows the entry of microbial products into the bloodstream, disturbs immune homeostasis, and triggers systemic inflammation [39]. For instance, bacterial lipopolysaccharides (LPS) and flagellin can activate toll-like receptors (TLRs), inducing a pro-inflammatory environment characterized by the production of interleukins (IL-1β, IL-6), tumor necrosis factor-alpha (TNF-α), and an imbalanced Th17/Treg ratio [40,41,42]. This pro-inflammatory environment in the gut can lead to development of different ADs.
It is worth noting that some gut microbial communities involved in ADs may have originated from the oral cavity. For instance, Porphyromonas gingivalis, a significant periodontal pathogen, is known to express peptidylarginine deiminase and to produce citrullinated epitopes, which are recognized by ACPA (anti-citrullinated protein antibodies) in patients with RA. However, the exact mechanism is still unidentified, and further research in this area is being conducted [43].
Molecular Mimicry
Sharing similar structural components between gut microbiota and self-antigens, known as molecular mimicry, can lead to the activation of unnecessary immune responses and cross-reactivity. For instance, some peptides originated from bacterial communities such as Bacteroides fragilis, P. copri, Candida albicans, and Streptococcus sanguis can mimic collagen and synovial/ribosomal peptides, and induce cross-reactive immune responses and lead to the development of ADs [44].
A high degree of resemblance between citrullinated fibrinogen peptides commonly found in synovial tissue and bacterial antigens in RA patients, and demyelination (loss of myelin sheath-protecting nerves) caused by molecular mimicry between myelin and certain microbial communities, have also suggested a significant role of autoimmune responses triggered by microbial communities in the pathogenesis of ADs [18, 45].
Recent studies have shown a growing collection of microbial peptides that exhibit molecular mimicry with host self-antigens, leading to autoimmune responses. For instance, specific peptides originated from gut microbial species such as A. muciniphila and R. gnavus have been found to mimic pancreatic beta-cell antigens and link these bacteria to the development of T1D [46]. Similarly, certain strains of Klebsiella pneumoniae can mimic HLA-B27, having shared structural features with these strains, which can lead to autoimmunity, inflammation, and tissue damage in ankylosing spondylitis [47].
Although dysbiosis can influence host immune responses, host susceptibility determined by HLA-DR genotypes (human leukocyte antigen-DR) seems to be one of the primary factors in the AD development. For instance, the association of HLA-DR2 and HLA-DR3 with SLE [48, 49], HLA-DRB1*04 with RA [50, 51], and HLA-DR3 and HLA-DR4 genotypes with T1D [52] show the significant role of genetic-gut microbiota interaction in AD development.
Epitope Spreading
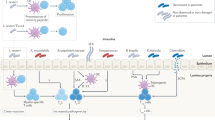
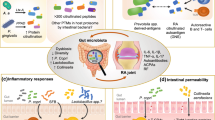
The impact of gut microbiota on host immunity extends beyond molecular mimicry. Specific microbial taxa can convert host self-proteins using their enzymes and generate neoepitopes that are modified version of host antigens. These neoepitopes can be recognized by the immune system and activate autoimmune responses (epitope spreading) [51, 53]. For instance, some bacterial enzymes can convert arginine residues to citrulline and produce citrullinated self-antigens which are associated with RA [54]. A high abundance of P. copri have been associated with increased protein citrullination, induction of pro-inflammatory cytokines, and led to tissue damage and the release of self-antigens. This dysregulation may potentially activate epitope spreading and contribute to the expansion of autoimmune responses in ADs [55] (Fig. 1 and Table 1).
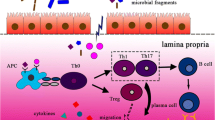
Potential mechanisms by which disturbed gut microbiota contributes to development of ADs. In patients with ADs, a leaky gut barrier can lead to the translocation of microbes and microbial products from the gut lumen into gut tissues and even into the circulation. The imbalanced gut microbiota can promote the activation of both innate and adaptive immunity, and the activation of pro-inflammatory cytokines, resulting in systemic immune dysregulation. Innate immune cells, like plasmacytoid DCs, become activated and secrete inflammatory cytokines, including type I interferons (IFNs). Moreover, microbial antigens can be presented to CD4+ T cells by DCs and macrophages, leading to the differentiation of inflammatory T cell subtypes such as T helper (Th)1, Th17, and T follicular helper (Tfh) cells. B cells can also be activated directly by microbial antigens or with Tfh cells, differentiating into plasma cells that produce protective secretory IgA (sIgA) and pathogenic autoantibodies. Microbial antigens can also trigger autoimmunity by mimicking self-antigens and lead to the development of ADs
Gut Microbiota-Targeted Therapies
The importance of the gut microbiota’s function in host physiology and its role in regulating immune responses have created new opportunities for adapted and targeted therapeutic approaches. Several strategies have been proposed for the treatment of ADs by modulating the composition of the gut microbiota. These include probiotics, prebiotics, postbiotics, diet adjustments, fecal microbiota transplantation, and engineered bacteria, which are discussed as follows:
Probiotics
As per the definition provided by the Food and Agriculture Organization and the World Health Organization (FAO/WHO), probiotics are live microorganisms that, when administered in sufficient quantities, confer a health benefit to the host [110]. These beneficial probiotic strains can play a vital role in restructuring the gut microbiota composition, regulating the immune system, and improving disease outcomes. Several studies have shown promising results of using probiotics in the management of ADs.
According to a randomized double-blind clinical trial, Lactobacillus casei 01 supplementation in women reduced inflammation (decreased IL-10, IL-12, TNF-α, and hs-CRP levels), improved disease activity (tender and swollen joint counts), and a positive treatment effect based on EULAR criteria (European League Against Rheumatism criteria) [111]. Furthermore, patients with RA demonstrated notable improvements after 8 weeks of probiotic supplementation (L. casei, Acidophilus, and Bifidobacterium), with reduced Disease Activity Score, enhanced B cell function, and decreased high-sensitivity C-reactive protein concentrations, suggesting potential benefits of probiotics in managing clinical and metabolic status of RA [112]. However, a meta-analysis of nine studies with 361 patients indicated that although probiotics significantly lowered the pro-inflammatory cytokine IL-6 compared to the placebo group in RA patients, there was no significant difference in disease activity score [113].
Moreover, evidence from animal studies demonstrated that oral administration of L. casei to Lewis rats suppressed RA progression by reducing pro-inflammatory molecules and promoting immunoregulatory IL-10 levels in CD4+ T cells [31]. L. casei intervention in adjuvant-induced arthritis (AIA) rats also inhibited joint swelling, reduced arthritis scores, and prevented bone destruction, suggesting that probiotics like L. casei could be a promising strategy for treating RA, especially in the early stages of the disease [114].
In the context of SLE, an in vitro culture study on microbiota isolated from SLE patients’ stool demonstrated that SLE-associated microbiota promoted lymphocyte activation and Th17 differentiation from naive CD4+ lymphocytes, while probiotics with Treg-inducer strains showed potential in rebalancing Treg/Th17/Th1 ratio [115]. Additionally, in murine experimental autoimmune encephalomyelitis (EAE), a model for MS, probiotic Lactobacillus reuteri DSM 17938 reduced TH1/TH17 cells and other associated cytokines, restored gut microbiota diversity, and altered the abundance of EAE-associated bacterial taxa [116]. Furthermore, the gut commensal bacterium Prevotella histicola suppressed EAE in mice by modulating systemic immune responses, including reducing pro-inflammatory Th1 and Th17 cells, while increasing CD4+FoxP3+ Treg cells, tolerogenic DCs, and suppressive macrophages [117].
Probiotics have also shown promising results in the management of T1D. Early probiotic administration in children at high genetic risk of T1D, particularly in those with the DR3/4 genotype, suggested a beneficial impact in reducing the risk of islet autoimmunity [118].
Despite all the beneficial effects of probiotic administration, it is important to note that the strain-specific effects of probiotics require further research to evaluate the most beneficial and effective strains for each autoimmune condition. Additionally, more research is needed to assess the long-term effects and therapeutic effectiveness of probiotics in human trials.
Prebiotics
Non-digestible dietary fibers that selectively stimulate the growth and activity of beneficial microbes in the gut are known as prebiotics. Prebiotics provides a suitable growth environment for microbial communities which positively influence the gut microbiota structure [119]. Recent studies have shown the potential of prebiotics in modulating immune responses and reducing disease severity.
Studies focusing on RA have shown that prebiotics promoted the growth of beneficial bacteria and reduced the abundance of pro-inflammatory bacteria and disease severity. For instance, the administration of Bacillus coagulans and prebiotic inulin (either alone or in combination) significantly reduced pro-inflammatory cytokines, serum amyloid A (SAA), and fibronectin (Fn); inhibited RA progression; and improved its clinical parameters [120]. Furthermore, the combination of prebiotics, a specific diet (known as synbiotics), and beneficial probiotic strains enhance positive effects through synergistic interactions. For instance, L. casei 01 combined with prebiotic oligofructose-enriched inulin promoted anti-inflammatory responses, reduced colonic damage, increased lactobacilli counts in feces, and improved myeloperoxidase activity in rat colitis models [121]. By providing a favorable environment for probiotic activity, synbiotic can optimize the symbiotic relationship between probiotic strains and the gut microbiota, which ultimately leads to improved health outcomes [122].
Postbiotics
Postbiotics are bioactive compounds produced by the metabolic activity of probiotic strains. These include bacterial lysates and enzymes, cell wall fragments, SCFAs, anti-microbial peptides, exopolysaccharides, and cell-free supernatants [123]. Postbiotics have emerged as potential alternatives to live microorganisms, as they offer a safer and more stable option for therapeutic use [124]. Studies have shown the immunomodulatory effects of postbiotics in several autoimmune diseases. For instance, Propionibacterium freudenreichii MJ2, a bacterium with postbiotic and probiotic properties, could inhibit osteoclast differentiation and improve RA in collagen-induced arthritis mice. It also reduced bone erosion, joint damage, and inflammation, offering potential therapeutic benefits for RA [125]. Similarly, L. casei DG (LC-DG) and its postbiotic (PB) could reduce the inflammatory mucosal response in ex-vivo cultures of mucosa from postinfectious irritable bowel syndrome (PI-IBS) patients. The results showed that LC-DG and PB reduced pro-inflammatory cytokines and TLR-4 expression, while increasing IL-10 levels after stimulation. This indicates the protective role of LC-DG and its PB in regulating the inflammatory response in PI-IBS [126]. Emerging research on postbiotics presents a promising therapeutic strategy for managing ADs. Their stable and well-defined properties, along with their immunomodulatory and anti-inflammatory effects, make them suitable candidates for clinical applications.
Fecal Microbiota Transplantation
Fecal microbiota transplantation (FMT) involves transferring healthy fecal material to a recipient’s gastrointestinal tract to restore a balanced gut microbiota composition. Although FMT has gained significant attention for its high efficiency in the treatment of Clostridium difficile infections [127], the impact of FMT on the management of ADs still is under investigation.
In a recent study, it was found that the FMT from SLE mice could trigger autoimmune responses in germ-free C57BL/6J mice. This was demonstrated by inflammatory responses, production of anti-dsDNA antibodies, and an increased susceptibility to the effects of genes associated with SLE [128]. In a similar study, FMT from SLE patients to germ-free mice (GF C57/B6J) resulted in the development of lupus-like features and pro-inflammatory responses, characterized by elevated levels of SLE-related autoantibodies, serum cytokines (IL-6, IL-8, TNF-α, and INF-γ), increased B-lymphocyte subsets in the intestinal lamina propria, and expanded peripheral Th-17 and CD4+ CXCR3+ cells with reduced immunomodulatory Treg cells compared to mice receiving healthy controls’ gut microbiota [129].
Also, RA fecal samples introduced into arthritis-prone SKG mice increased intestinal Th17 cells and caused severe arthritis when treated with zymosan. Lymphocytes in the colon and regional lymph nodes, but not the spleen, displayed enhanced IL-17 responses to RPL23A. Additionally, naive SKG mouse T cells co-cultured with P. copri-stimulated DCs induced IL-17 production and rapid arthritis development [58].
Mice colonized with IBD donor-derived microbiota also exhibited an abundance of mucosal Th17 cells, a deficit in tolerogenic RORγt+ Treg cells and increased susceptibility to colitis, while transplanting healthy donor-derived microbiota induced RORγt+ Treg cells and improved disease outcomes [130].
Another study on the role of isolated bacteria from individuals with MS in influencing human T cells and exacerbating MS symptoms in mouse models showed that MS fecal bacteria can induce pro-inflammatory responses and T cell differentiation leading to an exacerbation of MS-like symptoms in mice [81].
In the context of metabolic disorders, FMT in recently diagnosed T1D patients could prevent the decline in endogenous insulin production over 12 months. Patients who received autologous FMT (from their fecal samples) had significantly preserved stimulated C peptide levels compared to those who received allogenic FMT (from a healthy donor) with detected associations between specific bacteria taxa, plasma metabolites, and the preservation of residual beta cell function [131].
Alternative methods, such as oral capsules and sterile fecal filtrate transfer, have also shown success in managing microbiota-associated diseases [131, 132]. However, careful screening of these intervention methods targeting gut microbiota composition is crucial to prevent any potential adverse effects in recipients. Also, there are several challenges in terms of donor selection, standardization of FMT procedures, and potential long-term effects that need to be addressed in preclinical and clinical trials [133].
Beside recently introduced gut-microbiota associated therapies, synthetic or genetically modified bacteria, known as engineered bacteria, have also introduced a new therapeutic approach. Modification of microbial genetic properties enables specialized abilities beyond the natural features of microorganisms, which have been utilized across various fields, from agriculture to healthcare and industrial applications and holds great promise for future improvements in biotechnology and medicine [134].
Discussion
Gut microbiota dysbiosis has been linked to the pathophysiology and development of ADs [135]. In this review, the complex interactions between the gut microbiota and the host immune system in several ADs have been discussed.
Dysbiosis can result in an unbalanced immune response that is characterized by increased pro-inflammatory activities and impaired immunological tolerance [44]. Gut microbiota not only affects the immune response but also produces metabolites, such as SCFAs, that can regulate various organs and functions in the body [136]. SCFAs (acetate, propionate, and butyrate), in particular, have immunomodulatory properties which regulate the balance between pro-inflammatory and anti-inflammatory responses. Numerous studies have shown the association between altered gut microbiota composition and lower microbial diversity with autoimmune diseases. These variations have been linked to abnormal microbial translocation, increased intestinal permeability, cross-reactivity of microbial components with autoantigens, inflammatory responses, and dysregulated immune cell activation.
In recent years, studies examining the relationship between gut microbiota and different human disorders have attracted significant attention. However, despite the growing body of research in this area, the field is still in its early stages and have several gaps and challenges which are summarized as follows:
Existing Challenges and Strategies in Microbiota-Associated Studies
Definition of a Healthy Microbiota
Traditionally, researchers have defined a “healthy microbiota” based on the presence or absence of certain microbial taxa or species [137]. However, the gut microbiota structure may vary between individuals without necessarily indicating poor health. Therefore, a healthy gut microbiota should be defined based on the microbial functions and its interaction with the host, rather than merely relying on microbial taxonomical information. Also, the gut microbiota composition is influenced by several confounding factors, such as genetics and environmental factors over time. Therefore, it is difficult to define a “one-size-fits-all” solution for a healthy gut microbiota [138].
Study Design
Inappropriate study design and small sample size, which are seen in many microbiome studies, can significantly influence the conclusions of microbiota findings. The design of study depends on the study question and hypotheses of the research. For instance, cross-sectional studies are useful for identifying associations between microbial communities and clinical outcomes, while case–control studies are suitable for identification of potential biomarkers. Longitudinal studies can provide insight into the changes of the gut microbiota over time and the influence of confounding factors on gut microbiota composition and overall health. Moreover, randomized controlled trials can found causality between the microbiota findings and disease outcomes [139]. Strengths and limitations of each design should also be considered when interpreting the findings.
Methodology
The lack of standardized guidelines regarding sample collection, storage conditions, DNA extraction, sequencing platforms, and data analysis pipelines has made it challenging to draw clear conclusions and identify consistent patterns of gut microbiota involved in human diseases. A standardized approach enables comparability between studies and increases the reliability and reproducibility of results in the microbiota field [140, 141].
Model System
Although mouse models are commonly used in microbiota research, they are a few issues that need to be taken into account. These include the absence of standardized experimental protocols, the genetic similarity of laboratory mice, differences between mouse and human microbiota, and coprophagia habit which can significantly influence the gut microbiota composition in mouse models [142]. To overcome these challenges, it is necessary to develop mouse models that more accurately reflect the human gut microbiota and to adhere to standardized protocols when working with these animal models.
From Laboratory to Human Studies
Since most of our knowledge is limited to preclinical studies on animal models, long-term clinical studies are required to assess the safety, effectiveness, and long-term effects of in vivo and in vitro findings.
Ethics
Manipulation of the gut microbiota using different methods of interventions such as FMT requires considering different ethical considerations, such as informed consent, selection of appropriate donors, and safety of protocols, which might be very challenging in microbiota studies [143]. Transparency and open communication with participants must be prioritized in order to address these challenges, and strict guidelines for donor selection and risk assessments must also be implemented in order to guarantee the health and safety of study participants (Fig. 2).
Conclusion
Dysbiosis has been associated with the development of several gut-microbiota-associated diseases in humans. It has been shown that the abundance of several beneficial microbial communities decreases, while pathogenic and opportunistic pathogens increase. Therefore, targeting the gut microbiota composition through various human interventions with the aim of balancing the ratio of different microbial communities and their influence on host physiology seems to be a promising approach. However, it is important to note that the host-gut microbiota interaction is a complex network, and dysbiosis is not the only contributing factor in the development of several diseases. Recent advancements in omics approaches have shed light on the complex interactions between the host-microbiota interaction. Meta-transcriptomics, metagenomics, proteomics, metabolomics, transcriptomics, glycomics, lipidomics, and epigenomics have revealed that microbial metabolites, genes, and transcripts may provide more comprehensive insights than microbial taxa alone. These advancements have enabled researchers to go beyond the identification of microbial taxonomy and explore microbiota functions and their interactions with the host, which are essential in designing new therapeutical approaches. Also, it is worth pointing out that the non-bacterial communities of the gut, including viruses (virome), fungi (mycobiome), archaea, and protozoa, also play a significant role in shaping gut microbiota composition and influencing host physiology [144]. Although there is little information now available on these topics, gaining an understanding of the intricate relationships that exist between these bacteria and their hosts may help develop novel therapeutic approaches that focus on the entire gut ecology and pave the way for more personalized therapy (Table 2).
Data Availability
All data is available in the manuscript.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Davidson A, Diamond B. Autoimmune diseases. N Engl J Med. 2001;345(5):340–50.
Cici D, Corrado A, Rotondo C, Cantatore FP. Wnt signaling and biological therapy in rheumatoid arthritis and spondyloarthritis. Int J Mol Sci. 2019;20(22):5552.
Basta F, Fasola F, Triantafyllias K, Schwarting A. Systemic lupus erythematosus (SLE) therapy: the old and the new. Rheumatol Ther. 2020;7(3):433–46.
Burn P. Type 1 diabetes. Nat Rev Drug Discov. 2010;9(3):187.
Dobson R, Giovannoni G. Multiple sclerosis—a review. Eur J Neurol. 2019;26(1):27–40.
Ng QX, Soh AY, Loke W, Lim DY, Yeo WS. The role of inflammation in irritable bowel syndrome (IBS). J Inflamm Res. 2018;21:345–9.
Scher JU, Sczesnak A, Longman RS, Segata N, Ubeda C, Bielski C, et al. Expansion of intestinal Prevotellacopri correlates with enhanced susceptibility to arthritis. elife. 2013;2:e01202.
Sadeghpour Heravi F, Hu H. Bifidobacterium: host–microbiome interaction and mechanism of action in preventing common gut-microbiota-associated complications in preterm infants: a narrative review. Nutrients. 2023. https://doi.org/10.3390/nu15030709.
Berer K, Mues M, Koutrolos M, Rasbi ZA, Boziki M, Johner C, et al. Commensal microbiota and myelin autoantigen cooperate to trigger autoimmune demyelination. Nature. 2011;479(7374):538–41.
Wu H-J, Ivanov II, Darce J, Hattori K, Shima T, Umesaki Y, et al. Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity. 2010;32(6):815–27.
Armstrong D, Dregan A, Ashworth M, White P, McGee C, de Lusignan S. Influence of prior antibiotic use on risk of rheumatoid arthritis: case control study in general practice. Rheumatology. 2020;59(6):1281–7.
Singh JA, Furst DE, Bharat A, Curtis JR, Kavanaugh AF, Kremer JM, et al. 2012 update of the 2008 American College of Rheumatology recommendations for the use of disease-modifying antirheumatic drugs and biologic agents in the treatment of rheumatoid arthritis. Arthritis Care Res. 2012;64(5):625–39.
Law SC, Street S, Yu C-HA, Capini C, Ramnoruth S, Nel HJ, et al. T-cell autoreactivity to citrullinated autoantigenic peptides in rheumatoid arthritis patients carrying HLA-DRB1 shared epitope alleles. Arthritis Res Ther. 2012;14(3):1–12.
Cho I, Blaser MJ. The human microbiome: at the interface of health and disease. Nat Rev Genet. 2012;13(4):260–70.
Koh JH, Lee EH, Cha KH, Pan C-H, Kim D, Kim W-U. Factors associated with the composition of the gut microbiome in patients with established rheumatoid arthritis and its value for predicting treatment responses. Arthritis Res Ther. 2023;25(1):32.
Azzouz DF, Chen Z, Izmirly PM, Chen LA, Li Z, Zhang C, Mieles D, Trujillo K, Heguy A, Pironti A, Putzel GG. Longitudinal gut microbiome analyses and blooms of pathogenic strains during lupus disease flares. Ann Rheum Dis. 2023;82(10):1315–27.
Viatte S, Rigby WFC. HLA and other susceptibility genes in rheumatoid arthritis. 2019;1–4.
Berer K, Krishnamoorthy G. Microbial view of central nervous system autoimmunity. FEBS Lett. 2014;588(22):4207–13.
Jangi S, Gandhi R, Cox LM, Li N, Von Glehn F, Yan R, et al. Alterations of the human gut microbiome in multiple sclerosis. Nat Commun. 2016;7(1):12015.
Cekanaviciute E, Yoo BB, Runia TF, Debelius JW, Singh S, Nelson CA, et al. Gut bacteria from multiple sclerosis patients modulate human T cells and exacerbate symptoms in mouse models. Proc Natl Acad Sci. 2017;114(40):10713–8.
Liu X, Zou Q, Zeng B, Fang Y, Wei H. Analysis of fecal Lactobacillus community structure in patients with early rheumatoid arthritis. Curr Microbiol. 2013;67:170–6.
Chen BD, Jia XM, Xu JY, Zhao LD, Ji JY, Wu BX, Fei YY, Yang HX, Chen H, Zuo XX, Li H. Proinflammatory and autoimmunogenic gut microbiome in systemic lupus erythematosus. Biorxiv. 2019;621995.
Braniste V, Al-Asmakh M, Kowal C, Anuar F, Abbaspour A, Tóth M, et al. The gut microbiota influences blood-brain barrier permeability in mice. Sci Transl Med. 2014;6(263):263ra158-263ra158.
Golpour F, Abbasi-Alaei M, Babaei F, Mirzababaei M, Parvardeh S, Mohammadi G, et al. Short chain fatty acids, a possible treatment option for autoimmune diseases. Biomed Pharmacother. 2023;163:114763.
Bellocchi C, Fernández-Ochoa Á, Montanelli G, Vigone B, Santaniello A, Quirantes-Piné R, et al. Identification of a shared microbiomic and metabolomic profile in systemic autoimmune diseases. J Clin Med. 2019;8(9):1291.
Zhang H, Liao X, Sparks JB, Luo XM. Dynamics of gut microbiota in autoimmune lupus. Appl Environ Microbiol. 2014;80(24):7551–60.
Yasuda K, Takeuchi Y, Hirota K. The pathogenicity of Th17 cells in autoimmune diseases. In Seminars in immunopathology. Springer Berlin Heidelberg; 2019; 41, pp. 283–97.
De Groot P, Nikolic T, Pellegrini S, Sordi V, Imangaliyev S, Rampanelli E, et al. Faecal microbiota transplantation halts progression of human new-onset type 1 diabetes in a randomised controlled trial. Gut. 2021;70(1):92–105.
Belkaid Y, Harrison OJ. Homeostatic immunity and the microbiota. Immunity. 2017;46(4):562–76.
Jiao Y, Wu L, Huntington ND, Zhang X. Crosstalk between gut microbiota and innate immunity and its implication in autoimmune diseases. Front Immunol. 2020;11:282.
López P, de Paz B, Rodríguez-Carrio J, Hevia A, Sánchez B, Margolles A, et al. Th17 responses and natural IgM antibodies are related to gut microbiota composition in systemic lupus erythematosus patients. Sci Rep. 2016;6(1):24072.
Azzouz D, Omarbekova A, Heguy A, Schwudke D, Gisch N, Rovin BH, et al. Lupus nephritis is linked to disease-activity associated expansions and immunity to a gut commensal. Ann Rheum Dis. 2019;78(7):947–56.
Chen J, Wright K, Davis JM, Jeraldo P, Marietta EV, Murray J, et al. An expansion of rare lineage intestinal microbes characterizes rheumatoid arthritis. Genome Med. 2016;8(1):1–14.
Chiang H-I, Li J-R, Liu C-C, Liu P-Y, Chen H-H, Chen Y-M, et al. An association of gut microbiota with different phenotypes in Chinese patients with rheumatoid arthritis. J Clin Med. 2019;8(11):1770.
Shin C, Kim Y-K. Autoimmunity in microbiome-mediated diseases and novel therapeutic approaches. Curr Opin Pharmacol. 2019;49:34–42.
Furuse M, Hirase T, Itoh M, Nagafuchi A, Yonemura S, Tsukita S, et al. Occludin: a novel integral membrane protein localizing at tight junctions. J Cell Biol. 1993;123(6):1777–88.
Buscarinu M, Romano S, Mechelli R, Pizzolato Umeton R, Ferraldeschi M, Fornasiero A, et al. Intestinal permeability in relapsing-remitting multiple sclerosis. Neurotherapeutics. 2018;15:68–74.
Harbison JE, Roth-Schulze AJ, Giles LC, Tran CD, Ngui KM, Penno MA, et al. Gut microbiome dysbiosis and increased intestinal permeability in children with islet autoimmunity and type 1 diabetes: a prospective cohort study. Pediatr Diabetes. 2019;20(5):574–83.
Schmidt TS, Hayward MR, Coelho LP, Li SS, Costea PI, Voigt AY, et al. Extensive transmission of microbes along the gastrointestinal tract. Elife. 2019;8:e42693.
Belkaid Y, Hand TW. Role of the microbiota in immunity and inflammation. Cell. 2014;157(1):121–41.
Kim CH. Immune regulation by microbiome metabolites. Immunology. 2018;154(2):220–9.
Boulangé CL, Neves AL, Chilloux J, Nicholson JK, Dumas M-E. Impact of the gut microbiota on inflammation, obesity, and metabolic disease. Genome Med. 2016;8(1):42. https://doi.org/10.1186/s13073-016-0303-2.
Kriebel K, Hieke C, Müller-Hilke B, Nakata M, Kreikemeyer B. Oral biofilms from symbiotic to pathogenic interactions and associated disease—connection of periodontitis and rheumatic arthritis by peptidylarginine deiminase. Front Microbiol. 2018;9:53. https://doi.org/10.3389/fmicb.2018.00053.
Zhang X, Chen BD, Zhao LD, Li H. The gut microbiota: emerging evidence in autoimmune diseases. Trends Mol Med. 2020;26(9):862–73. https://doi.org/10.1016/j.molmed.2020.04.001.
Law SC, Street S, Yu CH, Capini C, Ramnoruth S, Nel HJ, et al. T-cell autoreactivity to citrullinated autoantigenic peptides in rheumatoid arthritis patients carrying HLA-DRB1 shared epitope alleles. Arthritis Res Ther. 2012;14(3):R118. https://doi.org/10.1186/ar3848.
Hansen CH, Krych L, Nielsen DS, Vogensen FK, Hansen LH, Sørensen SJ, et al. Early life treatment with vancomycin propagates Akkermansia muciniphila and reduces diabetes incidence in the NOD mouse. Diabetologia. 2012;55(8):2285–94. https://doi.org/10.1007/s00125-012-2564-7.
Zhang L, Zhang YJ, Chen J, Huang XL, Fang GS, Yang LJ, et al. The association of HLA-B27 and Klebsiella pneumoniae in ankylosing spondylitis: a systematic review. Microb Pathog. 2018;117:49–54. https://doi.org/10.1016/j.micpath.2018.02.020.
Hevia A, Milani C, López P, Cuervo A, Arboleya S, Duranti S, et al. Intestinal dysbiosis associated with systemic lupus erythematosus. mBio. 2014;5(5):e01548-14. https://doi.org/10.1128/mBio.01548-14.
Webber D, Cao J, Dominguez D, Gladman DD, Levy DM, Ng L, et al. Association of systemic lupus erythematosus (SLE) genetic susceptibility loci with lupus nephritis in childhood-onset and adult-onset SLE. Rheumatology (Oxford). 2020;59(1):90–8. https://doi.org/10.1093/rheumatology/kez220.
Asquith M, Sternes PR, Costello ME, Karstens L, Diamond S, Martin TM, et al. HLA alleles associated with risk of ankylosing spondylitis and rheumatoid arthritis influence the gut microbiome. Arthritis Rheumatol. 2019;71(10):1642–50. https://doi.org/10.1002/art.40917.
Vaahtovuo J, Munukka E, Korkeamäki M, Luukkainen R, Toivanen P. Fecal microbiota in early rheumatoid arthritis. J Rheumatol. 2008;35(8):1500–5.
Mejía-León ME, Petrosino JF, Ajami NJ, Domínguez-Bello MG, de la Barca AMC. Fecal microbiota imbalance in Mexican children with type 1 diabetes. Sci Rep. 2014;4(1):3814. https://doi.org/10.1038/srep03814.
Zhang X, Zhang D, Jia H, Feng Q, Wang D, Liang D, et al. The oral and gut microbiomes are perturbed in rheumatoid arthritis and partly normalized after treatment. Nat Med. 2015;21(8):895–905. https://doi.org/10.1038/nm.3914.
Darrah E, Andrade F. Rheumatoid arthritis and citrullination. Curr Opin Rheumatol. 2018;30(1):72–8. https://doi.org/10.1097/bor.0000000000000452.
Scher JU, Sczesnak A, Longman RS, Segata N, Ubeda C, Bielski C, et al. Expansion of intestinal Prevotellacopri correlates with enhanced susceptibility to arthritis. eLife. 2013;2:e01202. https://doi.org/10.7554/eLife.01202.
•• Mei L, Yang Z, Zhang X, Liu Z, Wang M, Wu X, et al. Sustained drug treatment alters the gut microbiota in rheumatoid arthritis. Front Immunol. 2021;12:704089. https://doi.org/10.3389/fimmu.2021.704089. (Studied gut microbiome alterations in the of RA patients (treated with HQT or LEF) and controls.)
• Kitamura K, Shionoya H, Suzuki S, Fukai R, Uda S, Abe C, et al. Oral and intestinal bacterial substances associated with disease activities in patients with rheumatoid arthritis: a cross-sectional clinical study. J Immunol Res. 2022;2022:6839356. https://doi.org/10.1155/2022/6839356. (Exploring multifactorial correlations in autoimmune diseases with demographic considerations.)
Maeda Y, Kurakawa T, Umemoto E, Motooka D, Ito Y, Gotoh K, et al. Dysbiosis contributes to arthritis development via activation of autoreactive T cells in the intestine. Arthritis Rheumatol. 2016;68(11):2646–61. https://doi.org/10.1002/art.39783.
Scher JU, Ubeda C, Equinda M, Khanin R, Buischi Y, Viale A, et al. Periodontal disease and the oral microbiota in new-onset rheumatoid arthritis. Arthritis Rheum. 2012;64(10):3083–94. https://doi.org/10.1002/art.34539.
Chen J, Wright K, Davis JM, Jeraldo P, Marietta EV, Murray J, et al. An expansion of rare lineage intestinal microbes characterizes rheumatoid arthritis. Genome Med. 2016;8(1):43. https://doi.org/10.1186/s13073-016-0299-7.
Krasselt M, Baerwald C. Sex, symptom severity, and quality of life in rheumatology. Clin Rev Allergy Immunol. 2019;56(3):346–61. https://doi.org/10.1007/s12016-017-8631-6.
Mu Q, Tavella VJ, Kirby JL, Cecere TE, Chung M, Lee J, et al. Antibiotics ameliorate lupus-like symptoms in mice. Sci Rep. 2017;7(1):13675. https://doi.org/10.1038/s41598-017-14223-0.
Mikuls TR, Payne JB, Yu F, Thiele GM, Reynolds RJ, Cannon GW, et al. Periodontitis and Porphyromonas gingivalis in patients with rheumatoid arthritis. Arthritis Rheumatol. 2014;66(5):1090–100. https://doi.org/10.1002/art.38348.
Teng F, Klinger CN, Felix KM, Bradley CP, Wu E, Tran NL, et al. Gut microbiota drive autoimmune arthritis by promoting differentiation and migration of Peyer’s patch T follicular helper cells. Immunity. 2016;44(4):875–88. https://doi.org/10.1016/j.immuni.2016.03.013.
•• Alpizar-Rodriguez D, Lesker TR, Gronow A, Gilbert B, Raemy E, Lamacchia C, et al. Prevotella copri in individuals at risk for rheumatoid arthritis. Ann Rheum Dis. 2019;78(5):590–3. https://doi.org/10.1136/annrheumdis-2018-214514. (Determining the microbiome composition and prevalence of Prevotella prior to the development of RA.)
Zegarra-Ruiz DF, El Beidaq A, Iñiguez AJ, Lubrano Di Ricco M, Manfredo Vieira S, Ruff WE, et al. A diet-sensitive commensal Lactobacillus strain mediates TLR7-dependent systemic autoimmunity. Cell Host Microbe. 2019;25(1):113-27.e6. https://doi.org/10.1016/j.chom.2018.11.009.
• Choi SC, Brown J, Gong M, Ge Y, Zadeh M, Li W, et al. Gut microbiota dysbiosis and altered tryptophan catabolism contribute to autoimmunity in lupus-susceptible mice. Sci Transl Med. 2020;12(551). https://doi.org/10.1126/scitranslmed.aax2220. (Investigation the role of gut microbiota, tryptophan metabolism, and host genetic susceptibility in systemic lupus erythematosus pathogenesis through mouse model studies.)
Azzouz D, Omarbekova A, Heguy A, Schwudke D, Gisch N, Rovin BH, Caricchio R, Buyon JP, Alekseyenko AV, Silverman GJ. Lupus nephritis is linked to disease-activity associated expansions and immunity to a gut commensal. Ann Rheum Dis. 2019;78(7):947–56.
van der Meulen TA, Harmsen HJM, Vila AV, Kurilshikov A, Liefers SC, Zhernakova A, et al. Shared gut, but distinct oral microbiota composition in primary Sjögren’s syndrome and systemic lupus erythematosus. J Autoimmun. 2019;97:77–87. https://doi.org/10.1016/j.jaut.2018.10.009.
•• Li Y, Wang HF, Li X, Li HX, Zhang Q, Zhou HW, et al. Disordered intestinal microbes are associated with the activity of systemic lupus erythematosus. Clin Sci (Lond). 2019;133(7):821–38. https://doi.org/10.1042/cs20180841. (Identifying specific microbial associations and providing diagnostic and predictive models for disease activity.)
• Manfredo Vieira S, Hiltensperger M, Kumar V, Zegarra-Ruiz D, Dehner C, Khan N, et al. Translocation of a gut pathobiont drives autoimmunity in mice and humans. Science. 2018;359(6380):1156–61. https://doi.org/10.1126/science.aar7201. (Gut pathobiont Enterococcus gallinarum triggers autoimmunity in genetically predisposed hosts, translocating to systemic tissues; antibiotic treatment and vaccination show preventive potential.)
Greiling TM, Dehner C, Chen X, Hughes K, Iñiguez AJ, Boccitto M, et al. Commensal orthologs of the human autoantigen Ro60 as triggers of autoimmunity in lupus. Sci Transl Med. 2018;10(434). https://doi.org/10.1126/scitranslmed.aan2306
de Goffau MC, Luopajärvi K, Knip M, Ilonen J, Ruohtula T, Härkönen T, et al. Fecal microbiota composition differs between children with β-cell autoimmunity and those without. Diabetes. 2013;62(4):1238–44. https://doi.org/10.2337/db12-0526.
Endesfelder D, zu Castell W, Ardissone A, Davis-Richardson AG, Achenbach P, Hagen M, Pflueger M, Gano KA, Fagen JR, Drew JC, Brown CT. Compromised gut microbiota networks in children with anti-islet cell autoimmunity. Diabetes. 2014;63(6):2006–14.
Vatanen T, Franzosa E, Schwager R, Tripathi S, Arthur T, Vehik K, et al. The human gut microbiome in early-onset type 1 diabetes from the TEDDY study. Nature. 2018;562. https://doi.org/10.1038/s41586-018-0620-2.
de Goffau MC, Fuentes S, van den Bogert B, Honkanen H, de Vos WM, Welling GW, et al. Aberrant gut microbiota composition at the onset of type 1 diabetes in young children. Diabetologia. 2014;57(8):1569–77. https://doi.org/10.1007/s00125-014-3274-0.
Brown CT, Davis-Richardson AG, Giongo A, Gano KA, Crabb DB, Mukherjee N, Casella G, Drew JC, Ilonen J, Knip M, Hyöty H. Gut microbiome metagenomics analysis suggests a functional model for the development of autoimmunity for type 1 diabetes. PloS one. 2011;6(10):e25792. https://doi.org/10.1371/journal.pone.0025792.
Giongo A, Gano KA, Crabb DB, Mukherjee N, Novelo LL, Casella G, et al. Toward defining the autoimmune microbiome for type 1 diabetes. Isme J. 2011;5(1):82–91. https://doi.org/10.1038/ismej.2010.92.
•• Kostic AD, Gevers D, Siljander H, Vatanen T, Hyötyläinen T, Hämäläinen AM, et al. The dynamics of the human infant gut microbiome in development and in progression toward type 1 diabetes. Cell Host Microbe. 2015;17(2):260–73. https://doi.org/10.1016/j.chom.2015.01.001. (Examining infants predisposed to type 1 diabetes, this study identifies stable individualized gut microbiome trajectories during infancy.)
Jangi S, Gandhi R, Cox LM, Li N, von Glehn F, Yan R, et al. Alterations of the human gut microbiome in multiple sclerosis. Nat Commun. 2016;7(1):12015. https://doi.org/10.1038/ncomms12015.
Cekanaviciute E, Yoo BB, Runia TF, Debelius JW, Singh S, Nelson CA, et al. Gut bacteria from multiple sclerosis patients modulate human T cells and exacerbate symptoms in mouse models. Proc Natl Acad Sci U S A. 2017;114(40):10713–8. https://doi.org/10.1073/pnas.1711235114.
• Boziki MK, Kesidou E, Theotokis P, Mentis AA, Karafoulidou E, Melnikov M, et al. Microbiome in multiple sclerosis; where are we, what we know and do not know. Brain Sci. 2020;10(4). https://doi.org/10.3390/brainsci10040234. (Examines the increasing incidence of MS and its potential link to environmental factors.)
Miyake S, Kim S, Suda W, Oshima K, Nakamura M, Matsuoka T, et al. Dysbiosis in the gut microbiota of patients with multiple sclerosis, with a striking depletion of species belonging to Clostridia XIVa and IV clusters. PLOS ONE. 2015;10(9):e0137429. https://doi.org/10.1371/journal.pone.0137429.
Cantarel BL, Waubant E, Chehoud C, Kuczynski J, DeSantis TZ, Warrington J, Venkatesan A, Fraser CM, Mowry EM. Gut microbiota in multiple sclerosis: possible influence of immunomodulators. J Investig Med. 2015;63(5):729–34. https://doi.org/10.1097/jim.0000000000000192.
•• Ventura RE, Iizumi T, Battaglia T, Liu M, Perez-Perez GI, Herbert J, et al. Gut microbiome of treatment-naïve MS patients of different ethnicities early in disease course. Sci Rep. 2019;9(1):16396. https://doi.org/10.1038/s41598-019-52894-z. (This study compares the microbiome of MS subjects in early disease stages across different ethnic groups indicating an increased abundance of Clostridium (potential biomarkers).)
• Saresella M, Marventano I, Barone M, La Rosa F, Piancone F, Mendozzi L, et al. Alterations in circulating fatty acid are associated with gut microbiota dysbiosis and inflammation in multiple sclerosis. Front Immunol. 2020;11:1390. https://doi.org/10.3389/fimmu.2020.01390. (Evaluating serum concentrations of butyric acid, caproic acid (CA) in MS patients reveals a dysregulated SCFA/MCFA ratio, and dysbiosis suggesting a potential biomarker for disease activity and treatment efficacy.)
Tremlett H, Fadrosh DW, Faruqi AA, Zhu F, Hart J, Roalstad S, et al. Gut microbiota in early pediatric multiple sclerosis: a case-control study. Eur J Neurol. 2016;23(8):1308–21. https://doi.org/10.1111/ene.13026.
Miyake S, Kim S, Suda W, Oshima K, Nakamura M, Matsuoka T, et al. Dysbiosis in the gut microbiota of patients with multiple sclerosis, with a striking depletion of species belonging to Clostridia XIVa and IV clusters. PLoS One. 2015;10(9):e0137429. https://doi.org/10.1371/journal.pone.0137429.
Chen J, Chia N, Kalari KR, Yao JZ, Novotna M, Paz Soldan MM, et al. Multiple sclerosis patients have a distinct gut microbiota compared to healthy controls. Sci Rep. 2016;6:28484. https://doi.org/10.1038/srep28484.
Cosorich I, Dalla-Costa G, Sorini C, Ferrarese R, Messina MJ, Dolpady J, et al. High frequency of intestinal T(H)17 cells correlates with microbiota alterations and disease activity in multiple sclerosis. Sci Adv. 2017;3(7):e1700492. https://doi.org/10.1126/sciadv.1700492.
•• Marie V, Béatrice S, Elise L, Vincent M, Stephanie F, Christophe C, et al. Characterisation of gut microbiota composition in patients with axial spondyloarthritis and its modulation by TNF inhibitor treatment. RMD Open. 2023;9(1):e002794. https://doi.org/10.1136/rmdopen-2022-002794. (Evaluating the association between gut microbiota and axial spondyloarthritis, specific bacterial operational taxonomic units at baseline could serve as a novel predictor of TNF inhibitor response.)
•• Wang L, Wang Y, Zhang P, Song C, Pan F, Li G, et al. Gut microbiota changes in patients with spondyloarthritis: a systematic review. Semin Arthritis Rheum. 2022;52:151925. https://doi.org/10.1016/j.semarthrit.2021.11.002. (Consolidate evidence on gut microbiota characteristics in spondyloarthritis (SpA) compared to controls. Significant differences in microbial composition was detected in SpA patients.)
• Gill T, Stauffer P, Asquith M, Laderas T, Martin TM, Davin S, et al. Axial spondyloarthritis patients have altered mucosal IgA response to oral and fecal microbiota. Front Immunol. 2022;13. https://doi.org/10.3389/fimmu.2022.965634. (Exploring altered immunoglobulin A (IgA) responses in the gut and oral microbiota of axial spondyloarthritis (axSpA) patients, revealing distinct populations of immunoreactive microbes.)
Li M, Dai B, Tang Y, Lei L, Li N, Liu C, et al. Altered bacterial-fungal interkingdom networks in the guts of ankylosing spondylitis patients. mSystems. 2019;4(2). https://doi.org/10.1128/mSystems.00176-18.
Breban M, Tap J, Leboime A, Said-Nahal R, Langella P, Chiocchia G, Furet JP, Sokol H. Faecal microbiota study reveals specific dysbiosis in spondyloarthritis. Ann Rheum Dis. 2017;76(9):1614–22. https://doi.org/10.1136/annrheumdis-2016-211064.
Wu GL, Lu HF, Chen YL, Wang Q, Cao H, Li TY. Changes of intestinal microecology in patients with primary Sjogren’s syndrome after therapy of Yangyin Yiqi Huoxue Recipe (养阴益气活血方). Chin J Integr Med. 2019;25:654–62. https://doi.org/10.1007/s11655-019-2939-4.
•• Wang X, Pang K, Wang J, Zhang B, Liu Z, Lu S, et al. Microbiota dysbiosis in primary Sjögren’s syndrome and the ameliorative effect of hydroxychloroquine. Cell Rep. 2022;40(11):111352. https://doi.org/10.1016/j.celrep.2022.111352. (This study reveals dysbiosis in the gut, oral, and vaginal microbiome of individuals with pSS, with the oral microbiome exhibiting the most significant variation.)
•• Mendez R, Watane A, Farhangi M, Cavuoto KM, Leith T, Budree S, et al. Gut microbial dysbiosis in individuals with Sjögren’s syndrome. Microb Cell Fact. 2020;19(1):90. https://doi.org/10.1186/s12934-020-01348-7. (Exploring the gut microbiome in individuals with dry eye (DE) indicating microbial composition alterations, with depletion of Firmicutes and expansion of Proteobacteria, Actinobacteria, and Bacteroidetes.)
• Li Y, Li Z, Sun W, Wang M, Li M. Characteristics of gut microbiota in patients with primary Sjögren’s syndrome in Northern China. PLOS ONE. 2022;17(11):e0277270. https://doi.org/10.1371/journal.pone.0277270. (Investigation of gut microbiota in patients with pSS compared to controls showed significant differences in alpha- and beta-diversity, with increased abundance of opportunistic pathogens.)
• AlmståhI A, Wikström M, Stenberg I, Jakobsson A, Fagerberg-Mohlin B. Oral microbiota associated with hyposalivation of different origins. Oral Microbiol Immunol. 2003;18(1):1–8. https://doi.org/10.1034/j.1399-302X.2003.180101.x. (Distinct oral microbial patterns associated with hyposalivation was detected. The RT group exhibited a marked increase in Lactobacillus and Candida albicans. In the pSS group mutans streptococci were elevated.)
Zhou S, Cai Y, Wang M, Yang WD, Duan N. Oral microbial flora of patients with Sicca syndrome. Mol Med Rep. 2018;18(6):4895–903. https://doi.org/10.3892/mmr.2018.9520.
Mandl T, Marsal J, Olsson P, Ohlsson B, Andréasson K. Severe intestinal dysbiosis is prevalent in primary Sjögren’s syndrome and is associated with systemic disease activity. Arthritis Res Ther. 2017;19(1):237. https://doi.org/10.1186/s13075-017-1446-2.
de Paiva CS, Jones DB, Stern ME, Bian F, Moore QL, Corbiere S, et al. Altered mucosal microbiome diversity and disease severity in Sjögren syndrome. Sci Rep. 2016;6:23561. https://doi.org/10.1038/srep23561.
Machiels K, Joossens M, Sabino J, De Preter V, Arijs I, Eeckhaut V, et al. A decrease of the butyrate-producing species Roseburia hominis and Faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut. 2014;63(8):1275–83. https://doi.org/10.1136/gutjnl-2013-304833.
Sokol H, Pigneur B, Watterlot L, Lakhdari O, Bermúdez-Humarán LG, Gratadoux JJ, et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc Natl Acad Sci U S A. 2008;105(43):16731–6. https://doi.org/10.1073/pnas.0804812105.
•• Knoll RL, Forslund K, Kultima JR, Meyer CU, Kullmer U, Sunagawa S, et al. Gut microbiota differs between children with inflammatory bowel disease and healthy siblings in taxonomic and functional composition: a metagenomic analysis. Am J Physiol Gastrointest Liver Physiol. 2017;312(4):G327–39. https://doi.org/10.1152/ajpgi.00293.2016. (Observed dysbiosis, which distinguishes patients from siblings, highlights such siblings as potential donors for microbiotal remodeling therapy in IBD.)
•• Zhou Y, Xu ZZ, He Y, Yang Y, Liu L, Lin Q, et al. Gut microbiota offers universal biomarkers across ethnicity in inflammatory bowel disease diagnosis and infliximab response prediction. mSystems. 2018;3(1). https://doi.org/10.1128/mSystems.00188-17. (Reporting that human fecal microbiota contains promising and universal biomarkers for the noninvasive evaluation of inflammatory bowel disease severity.)
Frank DN, St Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci U S A. 2007;104(34):13780–5. https://doi.org/10.1073/pnas.0706625104.
Kerckhoffs AP, Samsom M, van der Rest ME, de Vogel J, Knol J, Ben-Amor K, Akkermans LM. Lower Bifidobacteria counts in both duodenal mucosa-associated and fecal microbiota in irritable bowel syndrome patients. World J Gastroenterol. 2009;15(23):2887–92. https://doi.org/10.3748/wjg.15.2887.
Morelli L, Capurso L. FAO/WHO guidelines on probiotics: 10 years later. J Clin Gastroenterol. 2012;46(Suppl):S1-2. https://doi.org/10.1097/MCG.0b013e318269fdd5.
Alipour B, Homayouni-Rad A, Vaghef-Mehrabany E, Sharif SK, Vaghef-Mehrabany L, Asghari-Jafarabadi M, et al. Effects of Lactobacillus casei supplementation on disease activity and inflammatory cytokines in rheumatoid arthritis patients: a randomized double-blind clinical trial. Int J Rheum Dis. 2014;17(5):519–27. https://doi.org/10.1111/1756-185x.12333.
Zamani B, Golkar HR, Farshbaf S, Emadi-Baygi M, Tajabadi-Ebrahimi M, Jafari P, et al. Clinical and metabolic response to probiotic supplementation in patients with rheumatoid arthritis: a randomized, double-blind, placebo-controlled trial. Int J Rheum Dis. 2016;19(9):869–79. https://doi.org/10.1111/1756-185x.12888.
Mohammed AT, Khattab M, Ahmed AM, Turk T, Sakr N, MK A, et al. The therapeutic effect of probiotics on rheumatoid arthritis: a systematic review and meta-analysis of randomized control trials. Clin Rheumatol. 2017;36(12):2697–707. https://doi.org/10.1007/s10067-017-3814-3.
Pan H, Guo R, Ju Y, Wang Q, Zhu J, Xie Y, et al. A single bacterium restores the microbiome dysbiosis to protect bones from destruction in a rat model of rheumatoid arthritis. Microbiome. 2019;7(1):107. https://doi.org/10.1186/s40168-019-0719-1.
López P, de Paz B, Rodríguez-Carrio J, Hevia A, Sánchez B, Margolles A, et al. Th17 responses and natural IgM antibodies are related to gut microbiota composition in systemic lupus erythematosus patients. Sci Rep. 2016;6:24072. https://doi.org/10.1038/srep24072.
He B, Hoang TK, Tian X, Taylor CM, Blanchard E, Luo M, et al. Lactobacillus reuteri reduces the severity of experimental autoimmune encephalomyelitis in mice by modulating gut microbiota. Front Immunol. 2019;10. https://doi.org/10.3389/fimmu.2019.00385
Mangalam A, Shahi SK, Luckey D, Karau M, Marietta E, Luo N, et al. Human gut-derived commensal bacteria suppress CNS inflammatory and demyelinating disease. Cell Rep. 2017;20(6):1269–77. https://doi.org/10.1016/j.celrep.2017.07.031.
Uusitalo U, Liu X, Yang J, Aronsson CA, Hummel S, Butterworth M, Lernmark Å, Rewers M, Hagopian W, She JX, Simell O. Association of early exposure of probiotics and islet autoimmunity in the TEDDY study. JAMA Pediatr. 2016;170(1):20–8. https://doi.org/10.1001/jamapediatrics.2015.2757.
Yadav MK, Kumari I, Singh B, Sharma KK, Tiwari SK. Probiotics, prebiotics and synbiotics: safe options for next-generation therapeutics. Appl Microbiol Biotechnol. 2022;106(2):505–21. https://doi.org/10.1007/s00253-021-11646-8.
Abhari K, Shekarforoush SS, Hosseinzadeh S, Nazifi S, Sajedianfard J, Eskandari MH. The effects of orally administered Bacillus coagulans and inulin on prevention and progression of rheumatoid arthritis in rats. Food Nutr Res. 2016;60:30876. https://doi.org/10.3402/fnr.v60.30876.
Ivanovska TP, Mladenovska K, Zhivikj Z, Pavlova MJ, Gjurovski I, Ristoski T, et al. Synbiotic loaded chitosan-Ca-alginate microparticles reduces inflammation in the TNBS model of rat colitis. Int J Pharm. 2017;527(1–2):126–34. https://doi.org/10.1016/j.ijpharm.2017.05.049.
Kearney SM, Gibbons SM, Erdman SE, Alm EJ. Orthogonal dietary niche enables reversible engraftment of a gut bacterial commensal. Cell Rep. 2018;24(7):1842–51. https://doi.org/10.1016/j.celrep.2018.07.032.
Żółkiewicz J, Marzec A, Ruszczyński M, Feleszko W. Postbiotics—a step beyond pre- and probiotics. Nutrients. 2020;12(8). https://doi.org/10.3390/nu12082189
Wegh CAM, Geerlings SY, Knol J, Roeselers G, Belzer C. Postbiotics and their potential applications in early life nutrition and beyond. Int J Mol Sci. 2019;20(19). https://doi.org/10.3390/ijms20194673
Yeom J, Yim DJ, Ma S, Lim YH. Propionibacterium freudenreichii inhibits rankl-induced osteoclast differentiation and ameliorates rheumatoid arthritis in collagen-induced arthritis mice. Microorganisms. 2021;10(1):48. https://doi.org/10.3390/microorganisms10010048
Compare D, Rocco A, Coccoli P, Angrisani D, Sgamato C, Iovine B, Salvatore U, Nardone G. Lactobacillus casei DG and its postbiotic reduce the inflammatory mucosal response: an ex-vivo organ culture model of post-infectious irritable bowel syndrome. BMC Gastroenterol. 2017;17(1):1–8. https://doi.org/10.1186/s12876-017-0605-x.
Rohlke F, Stollman N. Fecal microbiota transplantation in relapsing Clostridium difficile infection. Therap Adv Gastroenterol. 2012;5(6):403–20. https://doi.org/10.1177/1756283x12453637.
Ma Y, Xu X, Li M, Cai J, Wei Q, Niu H. Gut microbiota promote the inflammatory response in the pathogenesis of systemic lupus erythematosus. Mol Med. 2019;25(1):35. https://doi.org/10.1186/s10020-019-0102-5.
Ma Y, Guo R, Sun Y, Li X, He L, Li Z, et al. Lupus gut microbiota transplants cause autoimmunity and inflammation. Clin Immunol. 2021;233:108892. https://doi.org/10.1016/j.clim.2021.108892.
Britton GJ, Contijoch EJ, Spindler MP, Aggarwala V, Dogan B, Bongers G, et al. Defined microbiota transplant restores Th17/RORγt(+) regulatory T cell balance in mice colonized with inflammatory bowel disease microbiotas. Proc Natl Acad Sci U S A. 2020;117(35):21536–45. https://doi.org/10.1073/pnas.1922189117.
de Groot P, Nikolic T, Pellegrini S, Sordi V, Imangaliyev S, Rampanelli E, et al. Faecal microbiota transplantation halts progression of human new-onset type 1 diabetes in a randomised controlled trial. Gut. 2021;70(1):92–105. https://doi.org/10.1136/gutjnl-2020-322630.
Kao D, Roach B, Silva M, Beck P, Rioux K, Kaplan GG, et al. Effect of oral capsule- vs colonoscopy-delivered fecal microbiota transplantation on recurrent Clostridium difficile infection: a randomized clinical trial. JAMA. 2017;318(20):1985–93. https://doi.org/10.1001/jama.2017.17077.
Borody TJ, Khoruts A. Fecal microbiota transplantation and emerging applications. Nat Rev Gastroenterol Hepatol. 2011;9(2):88–96. https://doi.org/10.1038/nrgastro.2011.244.
Landry BP, Tabor JJ. Engineering diagnostic and therapeutic gut bacteria. Microbiol Spectr. 2017;5(5). https://doi.org/10.1128/microbiolspec.BAD-0020-2017.
Ruff WE, Greiling TM, Kriegel MA. Host–microbiota interactions in immune-mediated diseases. Nat Rev Microbiol. 2020;18(9):521–38. https://doi.org/10.1038/s41579-020-0367-2.
Mizuno M, Noto D, Kaga N, Chiba A, Miyake S. The dual role of short fatty acid chains in the pathogenesis of autoimmune disease models. PLoS One. 2017;12(2):e0173032. https://doi.org/10.1371/journal.pone.0173032.
Berg G, Rybakova D, Fischer D, Cernava T, Vergès M-CC, Charles T, et al. Microbiome definition re-visited: old concepts and new challenges. Microbiome. 2020;8(1):103. https://doi.org/10.1186/s40168-020-00875-0.
Hasan N, Yang H. Factors affecting the composition of the gut microbiota, and its modulation. PeerJ. 2019;7:e7502. https://doi.org/10.7717/peerj.7502.
Qian XB, Chen T, Xu YP, Chen L, Sun FX, Lu MP, et al. A guide to human microbiome research: study design, sample collection, and bioinformatics analysis. Chin Med J (Engl). 2020;133(15):1844–55. https://doi.org/10.1097/cm9.0000000000000871.
Mirzayi C, Renson A, Furlanello C, Sansone S-A, Zohra F, Elsafoury S, et al. Reporting guidelines for human microbiome research: the STORMS checklist. Nat Med. 2021;27(11):1885–92. https://doi.org/10.1038/s41591-021-01552-x.
Heravi FS, Zakrzewski M, Vickery K, Hu H. Host DNA depletion efficiency of microbiome DNA enrichment methods in infected tissue samples. J Microbiol Methods. 2020;170:105856. https://doi.org/10.1016/j.mimet.2020.105856.
Hugenholtz F, de Vos WM. Mouse models for human intestinal microbiota research: a critical evaluation. Cell Mol Life Sci. 2018;75(1):149–60. https://doi.org/10.1007/s00018-017-2693-8.
Rhodes R. Ethical issues in microbiome research and medicine. BMC Med. 2016;14(1):156. https://doi.org/10.1186/s12916-016-0702-7.
Vemuri R, Shankar EM, Chieppa M, Eri R, Kavanagh K. Beyond just bacteria: functional biomes in the gut ecosystem including virome, mycobiome, archaeome and helminths. Microorganisms. 2020;8(4). https://doi.org/10.3390/microorganisms8040483.
Vilanova C, Porcar M. Are multi-omics enough? Nat Microbiol. 2016;1(8):16101. https://doi.org/10.1038/nmicrobiol.2016.101.
Thomas T, Gilbert J, Meyer F. Metagenomics — a guide from sampling to data analysis. Microb Inform Exp 2. 2012;2(1):1–12. https://doi.org/10.1186/2042-5783-2-3.
Aguiar-Pulido V, Huang W, Suarez-Ulloa V, Cickovski T, Mathee K, Narasimhan G. Metagenomics, metatranscriptomics, and metabolomics approaches for microbiome analysis. Evol Bioinform Online. 2016;12(Suppl 1):5–16. https://doi.org/10.4137/ebo.S36436.
Wang Z, Gerstein M, Snyder M. RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet. 2009;10(1):57–63. https://doi.org/10.1038/nrg2484.
Salvato F, Hettich RL, Kleiner M. Five key aspects of metaproteomics as a tool to understand functional interactions in host-associated microbiomes. PLoS Pathog. 2021;17(2):e1009245. https://doi.org/10.1371/journal.ppat.1009245.
Clish CB. Metabolomics: an emerging but powerful tool for precision medicine. Cold Spring Harb Mol Case Stud. 2015;1(1):a000588. https://doi.org/10.1101/mcs.a000588.
Al-Amrani S, Al-Jabri Z, Al-Zaabi A, Alshekaili J, Al-Khabori M. Proteomics: concepts and applications in human medicine. World J Biol Chem. 2021;12(5):57–69. https://doi.org/10.4331/wjbc.v12.i5.57.
Thaysen-Andersen M, Kolarich D, Packer NH. Glycomics & glycoproteomics: from analytics to function. Mol Omics. 2021;17(1):8–10. https://doi.org/10.1039/d0mo90019b.
Yang K, Han X. Lipidomics: techniques, applications, and outcomes related to biomedical sciences. Trends Biochem Sci. 2016;41(11):954–69. https://doi.org/10.1016/j.tibs.2016.08.010.
Callinan PA, Feinberg AP. The emerging science of epigenomics. Hum Mol Genet. 2006;15(Spec No 1):R95-101. https://doi.org/10.1093/hmg/ddl095.
Acknowledgements
I would like to express my gratitude to Professor Camille Locht for critically reviewing and providing valuable advice and comments on this manuscript (figures are original and created using Biorender.com).
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions
Author information
Authors and Affiliations
Contributions
FSH: Conception and design of the study, literature review and data collection, data interpretation, writing and drafting of the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The author declares no competing interests.
Human and Animal Rights and Informed Consent
This article contains no studies with human or animal subjects performed by the author.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sadeghpour Heravi, F. Gut Microbiota and Autoimmune Diseases: Mechanisms, Treatment, Challenges, and Future Recommendations. Curr Clin Micro Rpt 11, 18–33 (2024). https://doi.org/10.1007/s40588-023-00213-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40588-023-00213-6