Abstract
We report on the results of particle-based, coarse-grained molecular dynamics simulations of amphiphilic lipid molecules in aqueous environment where the membrane structures at equilibrium are subsequently exposed to strong shock waves, and their damage is analyzed. The lipid molecules self-assemble from unbiased random initial configurations to form stable bilayer membranes, including closed vesicles. During self-assembly of lipid molecules, we observe several stages of clustering, starting with many small clusters of lipids, gradually merging together to finally form one single bilayer membrane. We find that the clustering of lipids sensitively depends on the hydrophobic interaction \(h_\mathrm{c}\) of the lipid tails in our model and on temperature T of the system. The self-assembled bilayer membranes are quantitatively analyzed at equilibrium with respect to their degree of order and their local structure. We also show that—by analyzing the membrane fluctuations and using a linearized theory— we obtain area compression moduli \(K_\mathrm{A}\) and bending stiffnesses \(\kappa _\mathrm{B}\) for our bilayer membranes which are within the experimental range of in vivo and in vitro measurements of biological membranes. We also discuss the density profile and the pair correlation function of our model membranes at equilibrium which has not been done in previous studies of particle-based membrane models. Furthermore, we present a detailed phase diagram of our lipid model that exhibits a sol–gel transition between quasi-solid and fluid domains, and domains where no self-assembly of lipids occurs. In addition, we present in the phase diagram the conditions for temperature T and hydrophobicity \(h_\mathrm{c}\) of the lipid tails of our model to form closed vesicles. The stable bilayer membranes obtained at equilibrium are then subjected to strong shock waves in a shock tube setup, and we investigate the damage in the membranes due to their interaction with shock waves. Here, we find a transition from self-repairing membranes (reducing their damage after impact) and permanent (irreversible) damage, depending on the shock front speed. The here presented idea of using coarse-grained (CG) particle models for soft matter systems in combination with the investigation of shock-wave effects in these systems is a quite new approach.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Our coarse-grained (CG) particle model of lipid molecules. From left to right: The most common structure of lipid molecules found in biomembranes of eukaryotic cells, Dipalmitoylphosphatidylcholine (DPPC), \(\hbox {C}_{40}\hbox {H}_{80}\hbox {NO}_8\hbox {P}\), as a formula and a space filling model which in essence is built from 5 parts: The hydrophilic head, choline, is linked via a phosphate to glycerol, which in turn is linked to two hydrocarbon chains, forming the hydrophobic tail. In our particle-based, coarse-graining approach, the hydrophobic and hydrophilic parts of the lipid are replaced with a particle-based polymer model, where we can use any number of particles for the head part and also for the hydrocarbon tail. For our investigations in this study, we use a three-particle model that captures the aspect ratio R of real lipids, which can be estimated as \(R = (0.5\cdot \text {bilayer thickness})/ \sqrt{(\text {area per molecule})}\), and which is (for many cases) \(R\approx 2.5 \mathrm{nm} /\sqrt{(0.7\,\mathrm{nm}^2)}\approx 3\). Hence, we use two particles for the tails, indicated in light-colored spheres and one for the head, indicated in dark color. The particles of the lipid model are connected using anisotropic springs according to Eq. (3). The standard excluded volume interaction according to Eq. (1) between all beads is indicated in dark color (blue), and the angle \(\theta \) for the bending potential of Eq. (2) is also drawn. Finally, the attractive potential of Eq. (4) between tail particles of different chains, which mimics the hydrophobic interaction of the lipid tails with the aqueous environment is indicated with light-colored arrows (light-green). (Color figure online)
Every cell on Earth uses a plasma membrane to separate and protect its chemical components from the outside environment. The plasma membrane acts as a barrier to prevent the contents of the cell from escaping and mixing with the surrounding medium. It allows nutrients to pass inward, across the membrane through channels, and waste products to pass out. When a cell grows or changes shape, so does its plasma membrane; it can deform without tearing. If the membrane is pierced, it neither collapses like a ballon, nor remains torn; instead, it quickly reseals [1]. This transient permeability of a cell’s plasma membrane or even complete destruction can also be achieved by exposing cells to shock waves, as we have demonstrated in a recent publication where we introduced a new experimental setup for exposing U87 glioblastoma tumor cells to laser-induced shock waves [2]. Shock-wave treatment of cells might open up new interesting biological and medical applications, e.g., in gene therapy, where the permeabilization of the membrane aims at delivering therapeutic molecules, genes, or other genetic material into the cells’ cytoplasm [3–13].
Eukaryotic cells also contain an abundance of internal membranes that enclose intracellular compartments to form the various organelles of cells. These internal membranes are constructed on the same principles as the plasma membrane, and they, too, serve as highly selective barriers between spaces containing distinct collections of molecules. Subtle differences in the composition of these membranes, especially in their resident proteins, give each organelle its distinctive character [14].
Regardless of their locations, all cell membranes are composed of lipids and proteins and share a common general structure: They are arranged in two closely opposed sheets, forming a lipid bilayer: the lipid bilayer has been firmly established as the universal basis of membrane structure, and its properties are responsible for the general properties of all cell membranes [15]. The lipids in membranes combine two very different properties in a single molecule: each lipid has a hydrophilic head and one or two hydrophobic hydrocarbon tails. The most abundant lipids in eukaryotic cell membranes are the phospholipids which are molecules built around glycerol, a small molecule with three carbons and three associated hydroxyl groups [16]. In order to create a lipid, the cell attaches two long hydrophobic tails to two of these three hydroxyl groups forming a chemical linkage called an ester to which the hydrophilic head (most frequently a sugar or an amino acid) is linked through a phosphate group, see Fig. 1. For our purposes, we use a simple representation of the atomic structure of the lipids where many atoms are lumped into CG soft particles which are connected by entropic springs. This type of generic bead-spring model where several atoms (i.e., atomic degrees of freedom) are mapped onto a specific particle in order to gain computational speed-up is very common in computational polymer physics [17–19].
As reviewed by de Weer, [20], the paradigm of the fluid-mosaic model of membranes by Singer and Nicolson [21] was of great heuristic value, and contributed to our general thinking about membrane organization by providing insight into the assembly of lipids into membranes resulting from diffusion processes. However, subsequent experimental evidence indicated that the lateral motion of membrane components was not free after all, but constraint by various molecular mechanisms, such as direct or indirect interactions with cytoskeleton elements. Many relevant processes in cells that rely on membrane structures, e.g., the docking of proteins or other nanoparticles to the membrane, the exchange of particles through ion-channels, the formation of vesicles, or the self-assembly of a membrane from lipid molecules, are still not fully understood today because they occur on sub-microscopic length scales [16].
Dynamic processes on such a small scale cannot be observed in-vivo in experiments [14, 22, 23]. That is why for the experimental study of membrane properties one actually studies the physical chemistry of idealized in-vitro model-membranes with few, synthetic lipid molecules [6, 20]. One always has to accept here a compromise between the possible spatial resolution of the observation and the temporal resolution of dynamic processes in the membrane structures [24].
The mathematical treatment of the statistical behavior of membranes and their changes in shape is based on their description as two-dimensional surfaces and the introduction of thermodynamic potentials taking into account their geometry and curvature [25, 26]. Such models however, do not reproduce at all the actual microstructure of biological membranes and their composition as double lipid layer, consisting of amphiphilic lipids. Hence, mathematical models are normally restricted to length scales much larger than the actual lateral extension of membranes.
The standard numerical technique for the simulation of phospholipid bilayers is molecular dynamics (MD) [18, 27–30] and most of the existing body of MD simulation studies of biomembrane properties has almost exclusively been performed at or very near at equilibrium [31–37]. All-atom MD simulations of lipid bilayers which resolve the dynamics of individual atoms—treated with classical force fields—are generally limited to very small membrane samples (tens of nanometers in extension), even on the largest computer systems [38, 39]. This is also the case for novel compute unified device architecture (CUDA)-based simulations on NVIDIA graphics processing units (GPU)s. Here, the largest reported simulations using very specific force fields common in biophysics (AMBER and CHARMM) were able to generate trajectories up to the microsecond range, albeit only using 54.000 atoms, corresponding to a 6GB GPU card [40, 41].
In recent years, there has been an increased interest in studying the interaction of biological structures such as biomembranes and whole cells with shock waves [2, 42, 43]. Apart from the interesting aspects of shock-wave physics itself, these investigations are triggered mostly by the potential of new shock-wave-based applications in gene therapy and transfection [5, 8, 11, 44] and the simple practical need for an enhanced fundamental understanding of the physical processes occurring in medical cancer treatments based on shock-wave physics, e.g., High-intensity-focused ultrasound (HIFU). HIFU is a minimal invasive method that has been established for many years, but which is still poorly understood physically and theoretically [10, 45]. The experimental investigation of the physics of cells [46, 47] and in particular the theoretical-numerical modeling of the physical mechanisms of the interaction of cells with shock waves, are still representing emerging new fields [43]. This is why there have only been a handful of MD simulation studies dealing with shock wave interactions of cells, all of which are limited to very small all-atom simulations with explicitly modeled solvent, not taking into account the particular thermodynamic transient conditions of shock waves [48–50].
In fact, the largest atomistic MD simulations of shock-wave interactions with cell membranes until today were done by Koshiama et al. [49] and used system sizes of 128 phospholipids, with simulation box lengths parallel, and perpendicular to the bilayer plane of \(L_{\parallel }\approx 6.5 \, {\mathrm {nm}}\) and \(L_{\bot }\approx 16.0 \, {\mathrm {nm}}\), i.e., comparable to the bilayer thickness \(d_\mathrm{b}\), which is roughly \(d_\mathrm{b}\approx 5\,{{\mathrm {nm}}}\).
However, it seems highly questionable whether such extremely small systems can quantitatively reproduce real damage processes in cell membranes because, first of all, the employed periodic boundary conditions in Ref. [49] impose an artificial stabilization of the membrane patch due to correlation effects. Secondly, membrane rupture will likely originate from a defect, i.e., a deviation from the ideal flat surface. However, undulations of the membrane are strongly suppressed due to the small simulation box. Last but not least, the time evolution of the shocked membrane needs to be studied for as long as possible, requiring a large box length \(L_{\bot }\) along the direction of the shock impulse in order to allow the shock-wave front to travel for a long period of time after contact with the membrane. This last point is important because either a slow, diffusive disintegration of the membrane, or structural recovery from the induced damage can be observed as we will show in this paper.
All in all, atomistic simulations are too limited in their length and time scales to monitor structural instabilities of membranes or their transient permeability due to their interaction with shock waves. In particular, the former phenomenon has been reported experimentally [1, 4] but not yet been understood in detail, despite its importance for potential new applications, e.g., for shock-wave-based transdermal drug delivery or transfection of genes in eukaryotic cells [11, 51–53]. Thus, many relevant dynamic phenomena remain far outside the range of both, experimental observation and all-atom simulations [39].
This situation has led to the increasing use of CG models of biomembranes during the last ten years [17, 30, 54–60]. CG models constitute a class of mesoscale models, in which many atoms are treated by grouping them together into new particles which act as individual interaction sites connected by entropic springs [19, 43, 55, 57, 61, 62]. CG models were originally introduced in macromolecular modeling approaches for globular proteins in 1976 in a pioneering paper by Warshel and Levitt [63]. Since then, CG models have quickly found their way into polymer physics as so-called bead-spring models, taking advantage of universal scaling laws of long polymer chains due to their fractal nature, [64–66] as well as into engineering, biophysics, geophysics and other areas of computational research [19, 67]. As reviewed by Saunders and Voth, [68], CG models are nowadays routinely used in polymer physics and biophysics and provide a description of reduced complexity with respect to the molecular degrees of freedom [69–71].
Sometimes, also the solvent degrees of freedom are entirely removed in a CG description. In such implicit models, the solvent contributions to the interactions are treated through effective interaction terms such as stochastic forces, within an effective potential for the free energy [72–74]. The hydrophobic effects of the lipid tails due to the presence of solvent particles can be taken into account by the use of an attractive interaction potential between the lipids. This simplification leads to a computational speed-up, rendering simulations of large-scale membrane structures and their interaction with shock waves possible.
In this paper, we present an implicit-solvent, coarse-grained MD simulation study of the self-assembly of lipid molecules, which form stable bilayers and which are subsequently exposed to shock waves. In particular, in contrast to previous studies that included aspects of the dynamics of membrane assembly [29, 33, 41, 62, 75], we investigate the successive steps in membrane formation during self-assembly with respect to energy minimization and the degree of order in the systems.
The organization of the paper is as follows: In Sect. 2, details of the simulations performed are given. In Sect. 3, we discuss in detail the self-organization of lipid molecules starting from an unbiased initial gas state. The molecules eventually form stable bilayer membranes or closed vesicles. We proceed to analyze the structures of the bilayers at equilibrium by means of a multitude of physical parameters, such as an order parameter, the density profile, the pair correlation function, a phase diagram of our membrane model, the compression modulus \(K_\mathrm{C}\) and the bending stiffness \(\kappa _\mathrm{B}\). We believe that by providing an in-depth analysis of structural properties, we not only go beyond previous simulation studies, but we provide several new interesting aspects in the discussion of equilibrium membrane properties. Finally, as a novel approach, combining our CG model with shock-wave simulations, we expose the bilayer membranes to shock waves and analyze their damage quantitatively. Section 4 summarizes the main conclusions of this particle-based simulation study.
2 Coarse-grained particle model and computational details
We perform constant-temperature MD simulations using the MD simulation software package MD-CUBE which was developed by Steinhauser [65] for CG macromolecular simulations of single polymers and melts of almost arbitrarily branched monomer connectivity. For studying lipid bilayer membranes, a large variety of CG implicit-solvent models based on either MD [31, 59, 61, 74, 76–78] or Monte Carlo (MC) procedures [38, 72, 73, 79–83] have been developed, each one capturing molecular details at different levels of resolution.
Our approach to coarse-graining lipids is based on a standard CG polymer model, Eqs.(1)–(3), that has successfully been used in several studies of polymers [17–19]. In order to model the specific hydrophobic effect of lipids, we use an additional potential in Eq. (4), akin to one, that was introduced originally by Steinhauser in computational polymer physics to account for the effects of different solvent qualities in polymer solutions [65]. Our CG model is illustrated schematically in Fig. 1.
The complex molecular structure with its hydrophilic head group and hydrophobic tail group is simplified by replacing the lipid structure with spherical soft particles connected by elastic springs. We note that we can use any number of particles in our model for the hydrophilic head and also for the hydrophobic tail. For the principal analysis of our model presented here, we use a particularly simple version of it, where we use only three particles per lipid: The first particle which is color-coded in dark color (red) in Fig. 1, represents the hydrophilic head and the other two particles shown in light color (yellow) are the hydrophobic tails. The structural and amphiphilic properties of all lipids are modeled by the sum of the interaction potentials of Eqs. (1)–(4).
The finite size of a monomer of the lipid molecule is taken into account by the truncated Lennard-Jones (LJ) potential
where \(\sigma \) and \(\varepsilon \) determine the length and energy scale, respectively; r denotes the distance between two (nonbonded) mass points, and the cut-off radius of this short-ranged potential is chosen as \(r_{\text {c}}=\root 6 \of {2}\,\sigma \).
To straighten the lipids, a bending potential depending on the bond angle \(\theta \) (see Fig. 1) is used in the form
where \(\lambda \) determines the stiffness of the lipid molecule.
The mass points of a lipid chain are connected by anharmonic bonds with the potential
where \(\alpha \) is the force constant and the maximum bond length is chosen as \(r_{\infty }=1.5\sigma \)
The typical structures of self-assembling lipids arise from the hydrophobic and hydrophilic interactions of the lipid tails and heads, respectively, with the surrounding water molecules. When using a solvent-free model, the hydrophobic interactions of lipid tails with water can be modeled using an attractive potential between all tail particles of different lipid chains as indicated in Fig. 1 by the light-colored arrows:
where \(h_\mathrm{c}\) determines the range of the attractive potential, i.e., the larger \(h_\mathrm{c}\) the greater the hydrophobicity of the lipid tails, \(\xi \) is the force constant, and \(r_{\text {c}}\) is the same cutoff as in Eq. (1). A tunable potential of this type (involving a cosine function) was introduced by Steinhauser [65] for MD simulation of polymer-solvent interactions with varying solvent qualities. A similar form of the potential of Ref. [65] which we use in Eq. (4) was later used in implicit solvent simulations of lipids (i.e., very short polymer chains) by Cooke et al. [84].
The solvent–particle interaction is taken into account by a stochastic force \(\mathbf {\zeta }_i\) and a friction force, with a damping constant \(\gamma \), acting on each mass point m. The equations of motion of the i-th particle of the system are then given by the Langevin equation:
The force \(\mathbf {F}_{i}\) acting on particle i is obtained by
where \(\varPhi _\mathrm{total}\) is the sum of the potentials in Eqs. (1)–(4) and \(\mathbf {\nabla }_{\mathbf {r}_{i}}\) is a symbolic notation to indicate the partial derivatives with respect to the components of the position vector \(\mathbf {r}\) of the i-th particle, i.e., \(\mathbf {\nabla }_{\mathbf {r}_{i}}=\left( \frac{\partial }{\partial x_{i}}, \frac{\partial }{\partial y_{i}}, \frac{\partial }{\partial z_{i}}\right) \). The stochastic force \(\mathbf {\zeta }_{i}\) is assumed to be stationary, random, and Gaussian (white noise). This ensures proper equilibration of the dilute initial system [18]. The parameters for our model lipid systems are chosen so as to guarantee stable integration conditions: \(\lambda = 10 \varepsilon /\sigma , \,\,\alpha = 30\varepsilon /\sigma , \,\,\xi = \varepsilon , \,\,\gamma = 1.2\).
2.1 Shock-wave simulations
To study the degree of damage in our equilibrated bilayer membranes caused by the mechanical effects of shock waves, we use a recently introduced multiscale coupling method that bridges the gap between the atomistic and the continuum domain by introducing a heat exchange across the continuum-atomistic interface which is based on the dissipative particle dynamics at constant energy (DPDE) thermostat [85]. This scale bridging procedure couples the MD method (using our coarse-grained model of lipid molecules) with a continuum method, SPH (Smooth Particle Hydrodynamics), see Fig. 2. For the shock-wave simulations, we use the SPH method to model the surrounding aqueous environment of the membrane. The shock-wave front propagates in the fluid medium and exchanges energy with the membrane.
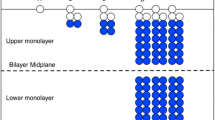
Snapshot of the initial equilibrated state of our solvated CG membrane model used for shock-wave simulations. We use coarse-grained MD particles for the lipid molecules (red hydrophilic head particle, green hydrophobic tail particles) and water as the surrounding SPH continuum. The SPH particles shown are not particles in the classical Newtonian sense, but locations where the hydrodynamic conservation equations are integrated. The thermodynamic coupling of the CG domain and the continuum domain using a DPDE thermostat is indicated with arrows. For details of the MD-SPH coupling algorithm, which we use for shock-wave simulations of membranes, we refer the readers to Ref. [85]. (Color figure online)
Many published MD simulation studies of shock waves propagating in a fluid medium use a simple, microcanonical-based approach. This is suitable for isothermal processes; however, the transient, discontinuous, thermodynamic nature of a shock wave, with a sudden change of thermodynamic variables, is usually not taken into account with this computational approach. Because there are high gradients of temperature and density involved at the shock-wave front, these effects need to be described accurately by a scale-bridging simulation method. The advantage of using SPH for modeling the surrounding fluid domain of the membrane over using ordinary MD particles is a tremendous speed-up of the simulation. With SPH we obtain the correct hydrodynamic behavior of the fluid with a comparably small number of integration points (SPH particles), where the hydrodynamic equations are solved. For details of our employed coupling method, we refer the readers to Ref. [85].
2.2 Initial setup of coarse-grained lipid molecules
At the beginning of the simulation, the lipid chains are generated within a cubic simulation box as follows: the center position of the first particle i of a chain is chosen randomly inside the simulation box. The position of the second particle is chosen in a random direction of the first one, at distance \(r_{i,i+1}=0.97\,\sigma \). The center position of the third particle of a lipid chain is chosen in the same way as the second but with the constraint that the distance to the first particle is \(r_{i,i+2}=1.02\,\sigma \). This is done to prevent the third particle from getting too close to the first one, which would immediately lead to strong repulsive LJ forces inside the chain and subsequent instability of the integration algorithm.
However, this procedure does not prevent particles of different lipid chains from initially overlapping each other. Therefore, before starting the actual simulation run with the full repulsive part of the potential of Eq. (1), we carry out a warm-up procedure, which integrates the system by gradually increasing the repulsive effect of the LJ potential as follows: First, the minimum distance \(r_{\text {min}}\) between all particles is calculated. Then, instead of calculating the LJ interactions between particles of different chains with the actual distance r in step n , a shifted distance of \(r^{\prime }=r+\Delta r - n/n_{\text {warmup}}\times \Delta r\) with \(\Delta r = r_\mathrm{c} -r_{\text {min}}\) and \(n\in \{1,\ldots ,n_{\text {warmup}}\}\) is used. Thus, the full LJ potential is effective only after \(n= n_{\text {warmup}}\) steps, which then allows for a fully stable integration of the equations of motion in Eq. (5). A more detailed description of our warm-up procedure can be found in Ref. [86].
3 Results and discussion
In this section, we discuss the dynamics of the formation and equilibration of bilayer membrane structures based on energy minimization and an order parameter \(S_{2}(\tau )\), derived from the second-order Legendre polynomial. Such a detailed analysis has not been provided in any previous MD simulation study that included aspects of molecular self-assembly [29, 31, 33, 41, 62, 75, 84]. Hence, we believe that we provide here new aspects in the discussion of bilayer formation using CG models, and in the quantitative analysis of structural properties of bilayer membranes at equilibrium.
In our structural analysis, we show that the obtained self-stabilizing equilibrium structures of bilayer membranes exhibit a bending stiffness \(\kappa _\mathrm{B}\) and compression modulus \(K_\mathrm{A}\) which are within the range of experimental measurements under physiological conditions. We further find that the gradient of the order parameter \(S_{2}\) can be used as a measure for major reorientations of the lipids during the process of self-assembly. We also discuss the density profile and the pair correlation function of our model membranes at equilibrium, and we present a calculated phase diagram of our lipid model which includes a sol–gel transition, quasi-liquid and quasi-crystalline phases behavior. In addition, we show in the phase diagram, the precise conditions for temperature T and hydrophobicity \(h_\mathrm{c}\) for stable bilayer and closed vesicle formation. Finally, as a new approach, we expose our membranes to shock waves in a shock-tube setup and analyze quantitatively the resulting damage based on an orientation dependent correlation function \(\varPsi \) which serves as damage parameter.
Simulation snapshots of the self-assembly process of lipid molecules to form bilayer membranes at temperature \(T = 1.0\,\varepsilon /k_\mathrm{B}\). The initial state at simulation time a \(\tau = 0\) is a random gas state. The other snapshots are taken at times b \(\tau = 100\), c \(\tau = 1000\), d \(\tau = 4000\), e \(\tau = 10200\), f \(\tau = 18000\), g \(\tau = 20200\) and h \(\tau = 22400\). As Electronic Supplementary Material (ESM), we provide Movie_S1, exhibiting the self-assembly of the lipid molecules as displayed here, finally aggregating to a single bilayer membrane. We also provide ESM Movie_S2, that shows the dynamics of the formation of closed vesicles with an appropriate choice of temperature T and hydrophobicity parameter \(h_\mathrm{c}\) of the CG lipid tails. The exact values are: \(T = 1.5, \,\,h_\mathrm{c} = 1.6\) and \(N = 2000\) lipids. We note here, that our simulation box is always large enough for the whole membrane to fit in several times in each direction such that vesicle formation can occur
3.1 Self-assembly of lipid molecules
Since biological cells are surrounded by—and filled with—solutions of molecules in water, the structure of cell membranes is mostly a consequence of the interactions of the membrane lipids with the aqueous environment. In Fig. 3, we show a typical example for the self-assembly process from random gas states into bilayer membranes for a system with \(N=3000\) particles (1000 lipid molecules), density \(\rho =0.05/\sigma ^3\), which corresponds to an edge ledge of \(L\approx 39.15\sigma \) of our cubic simulation domain, temperature \(T=1.0\,\varepsilon /k_\mathrm{B}\), timestep \(\Delta t= 0.01\tau \), where \(\tau \) is the time unit of the simulation, \(\alpha =30\varepsilon /\sigma ^2, \,\,r_{\infty }=1.5\sigma , \,\,\lambda =10\varepsilon /\sigma ^2, \,\,\xi = 1\varepsilon \) and \(h_\mathrm{c}=1.5\sigma \). The aggregation dynamics strongly depends on temperature T, and much lesser on the value of \(h_\mathrm{c}\) (the range of the attractive tail potential). The size of the system determines whether or not bilayers or vesicles are formed.
The start time \(\tau = 0\) denotes the beginning of the actual integration after the warm-up procedure described in Sect. 2.2. As can be seen in Fig. 3, starting from an unbiased random distribution of lipid molecules, very quickly the lipids assemble to small clusters with ellipsoidal or spherical shape. The smaller clusters gradually aggregate into bigger clusters, while minimizing the number of hydrophobic tail particles on the surface of these clusters. Finally, due to our particular choice of parameters, and because we run the systems longer than in previous coarse-grained MD studies [62, 75, 76, 82], in the simulations presented in Fig. 3, one single bilayer membrane in fluid phase is formed. The snapshots presented in Fig. 3b and particularly in (e) show one characteristic feature during lipid assembly, namely the occurrence of “lipid bridges“ during merging of different membranes, which looks as though one membrane is “swallowed“ by the other. This is consistent with one of the intermediate structures reported in recent all-atom MC simulations [41].
The time scale for occurrence of different stages in the self-assembly processes presented in Fig. 3 (very rapid initial clustering to form several smaller membranes, then gradual fusing) is consistent with what was reported in a relatively recent CG MD simulation study conducted by Noguchi [62] (Fig. 3 therein), but in contrast to that study which ends with 4 disjunct clusters, we present the dynamics until only one single equilibrated membrane remains. We note here, that the potential used in the study of Ref. [62] (a particle-based CG model of lipids) included orientation and angle-dependent functions of much more complexity and with more free fit-parameters compared to our CG model.
Depending on the particular choice of parameters in the simulations we can also achieve the formation of closed vesicles. For further details of lipid bilayer and vesicle formation dynamics, respectively, see the video sequences in the Electronic Supplementary Material (ESM): Movie_S1 and Movie_S2. The membrane fusion that can be observed in these movies, lies at the heart of important biological processes and of fundamental questions relating to cell evolution and the biological function of membrane curvature [87, 88]. In vivo, membrane fusion is tightly regulated by proteins [89]. The basic mechanism however, is determined by the physics of lipid–lipid interactions.
The progress in the self-assembly of lipid molecules can be tracked by monitoring the total potential energy of the system. When small lipid clusters merge into bigger clusters, the total surface area is decreased, thus reducing the free energy of the clustered system. While the system is approaching equilibrium, one can distinguish different decreasing energy levels in a plot of potential energy vs. time as displayed in Fig 4. The letters (a)–(h) in Fig. 4 indicate the different snapshots in the time interval \(\tau \in [0,\,4.0\times 10^{4}]\) and correspond to the snapshots presented in Fig. 3. After the system has attained a stationary equilibrium state for \(\tau \ge \tau _{eq} \approx 22400\) (snapshot (h) in Fig. 3), the total potential energy eventually oscillates about a constant value. We have checked this behavior in long simulation runs until \(\tau = 5\times 10^5\).
3.2 Order parameter
To characterize the degree of structural order in the system of lipid molecules during self-assembly, we introduce an order parameter \(O_{2}(\tau )\) based on the second-order Legendre polynomial \(P_{2}(\cos \vartheta )\):
Order parameter \(O_2(\tau )\) versus simulation time \(\tau \) during equilibrium phase for the same system that is discussed in Figs. 3 and 4. The indicated letters c–h in this figure correspond to the same points in time as indicated in Figs. 3 and 4. For better presentation of data points, a and b are not labeled
Here, \(\vartheta (\tau )\) denotes the momentary angle between the lipid direction vector and the average normal vector \(\mathbf {n}(\tau )\) of the membrane plane, called director, at time \(\tau \). The lipid direction vector is defined as the vector connecting the center of mass of a head particle with the center of mass of the second tail particle. The brackets \(\langle ...\rangle \) indicate averaging over all lipids. \(O_2(\tau )\) can be averaged over many time steps to obtain one single average value, but, as we show in Fig. 5, it is more interesting to look at the information provided by plotting the development of \(O_2(\tau )\) with simulation time.
\(O_2(\tau )\) provides information about the anisotropy of a system, i.e., the directional orientation of our self-assembled membranes. The values of \(O_2(\tau )\) are in the range of \(S_{2}(\tau )\in [-0.5, \,1.0]\). For \(O_2(\tau )=-0.5\), the molecules exhibit an alignment similar to a bottle brush, and hence, the average normal vector equals 0. \(O_2(\tau )=0.0\) occurs, when the lipids are isotropically oriented at random, like in a nondirectional fluid phase. Finally, \(O_2(\tau )=1.0\) corresponds to a perfect parallel alignment of the lipids. Typical liquid crystals exhibit values between \(O_2(\tau )=0.3\) and \(O_2(\tau )=0.8\).
In Fig. 5, we present a plot of \(O_2(\tau )\) of the system discussed in Figs. 3 and 4. Starting from a completely disordered, random initial state with \(O_2(0)=0.0\), the system slowly develops into an equilibrium state with values of \(O_2(\tau )\) saturating just below \(S_{2}\approx 0.6\) which is within the typical range of liquid crystals for a fluid phase. Thus, our model membranes also exhibit fluid-like behavior. The undulations of the values of \(O_2(\tau )\) in Fig.5 are due to this fluidity property of the simulated membranes.
After simulation time \(\tau = \tau _{eq} \approx 22400\) (snapshot (h) in Figs. 3–5), the system obviously has reached its equilibrium state, as indicated by the fluctuations of the order parameter about a mean value, just like the energy in Fig. 4. There are still strong fluctuations of \(O_2(\tau )\) for \(\tau > \tau _{eq}\), indicating that \(O_2(\tau )\) is a very sensitive quantity for local rearrangements of molecules, even in a stationary state. At snapshot (g), Fig. 3 shows that there are two bilayer structures which are in the process of fusing together. The energy of the overall system in Fig. 4 has decreased at this point compared to the previous snapshot (f), indicating the path of the system towards an equilibrium (minimum energy) state. The order parameter at snapshot (g) is rising from \(O_2\approx 0.1\) at (g) to \(O_2 \approx 0.55\) at (h). Thus, the re-organization of membranes by fusion is reflected by a steep gradient of \(O_2(\tau )\). When the two bilayer membranes are still separated from each other in snapshot (f), the order in the system is much larger (\(O_2 \approx 0.4\)) and then—when the fusion process sets in—\(O_2(\tau )\) decreases, indicating a less ordered state of the system during the re-organization of lipid molecules. This pattern for the values of \(O_2(\tau )\) obviously repeats itself several times during the whole process of self-assembly of lipid molecules: Whenever a rearrangement of lipids due to membrane fusion occurs, \(O_2(\tau )\) drops while the lipids rearrange, and then rises again, indicating increased order in the newly formed bilayer. Hence, the fluctuations of \(O_2(\tau )\) can be used as an indicator for major molecular rearrangements during the self-aggregation of lipids into bilayer membranes.
Order parameter \(O_2(\tau )\) according to Eq. (7) for a system at equilibrium with 1000 lipids. a A system at temperature \(T=0.5\varepsilon /k_\mathrm{B}\) with different ranges of hydrophobic tail interactions, indicated by parameter \(h_\mathrm{c}\). b A system with \(h_\mathrm{c}\,=\,1.3\sigma \) for a range of different temperatures T
Pair correlation function g(r) according to Eq. 8 for a system of 1000 lipids at equilibrium. The analysis is done for the same systems discussed in Fig. 6. a A system with very low temperature \(T=0.5\varepsilon /k_\mathrm{B}\) for different values of hydrophobicity \(h_\mathrm{c}\). b A system with \(h_\mathrm{c}=1.3\) for different temperatures T in units of \(\varepsilon /k_{B}\)
In Fig. 6, we present two striking examples of systems with 1000 lipids for which we performed a detailed analysis of the order parameter \(O_2(\tau )\). In Fig. 6a we show \(O_2(\tau )\) for a system at very low temperature \(T=0.5\varepsilon /k_\mathrm{B}\) for a variety of ranges of the attractive potential \(h_\mathrm{c}\) of the lipid tails which indicates the degree of hydrophobicity. The aggregated systems at such a low temperature are in a crystal-like solid phase, with a very particular arrangement of molecules. Interestingly, the fluctuations of \(O_2(\tau )\) are the largest for a small range of hydrophobic interactions, tailing off for larger \(h_\mathrm{c}\)-values. \(O_2(\tau )\) is generally limited to a rather narrow range of values within the interval [0.5, 0.7] and some fluctuations going down to \(O_2(\tau )=0.4\). Hence, we see that reducing the temperature to a rather low value has a dramatic effect on the overall structures of the aggregates that are formed. However, we also realize that tuning the interaction range \(h_\mathrm{c}\) in our model has an effect on the resulting bilayer structures that is much less pronounced than the effect of varying temperature T and keeping \(h_\mathrm{c}\) at a constant value, which is shown in Fig. 6b.
Phase diagram of our membrane model with respect to temperature T and interaction parameter \(h_\mathrm{c}\) with several typical bilayer membrane structures. Four different phases can be distinguished: I. A gas, where no clustering occurs (red crosses), II. A fluid phase where stable bilayer membranes are formed (blue squares). III. A quasi-crystalline phase where stable bilayers are formed with fluctuations in shape and membrane height (pink upper triangles). IV. A fluid phase where closed membrane vesicles are formed (filled black squares). Typical representative structures of lipid molecules in the four different phases are displayed as well. (Color figure online)
Here, we show the results for \(O_2(\tau )\) for a system with \(h_\mathrm{c} = 1.3\), varying the temperature between \(T=0.5\) and \(T=1.3\). Hence, the smallest value of T in Fig. 6b represents the same conditions as in Fig. 6a for the largest \(h_\mathrm{c}\)-value. We see, that for these two cases, \(O_2(\tau )=0.7\) consistently in (a) and (b) with very small fluctuations. It should be noted, that in (b) the range of \(O_2(\tau )\) is much larger than in (a), ranging from random configurations with \(O_2(\tau )\approx 0\) for the largest temperatures, where no clustering of lipids occurs, up to values pertaining to strongly aggregated bilayer membrane systems, which is confirmed by visual inspection of the self-assembled structures. The intermediate range of temperatures in Fig. 6b pertain to stable bilayer membrane structures in a fluid phase which lead to distinct fluctuations of the order parameter \(O_2(\tau )\). Hence, from Fig. 6 we can induce that the degree of order obtained in the self-assembled lipid aggregates for our model is much more sensitive to the variation of temperature rather than the variation of interaction range \(h_\mathrm{c}\).
3.3 Pair correlation function
Another quantity providing information about the local molecular structure of the aggregated lipids is the pair correlation function
In Fig. 7, we analyze g(r) for the same systems discussed in Fig. 6, where, in the case of (a), the temperature is set to a very low value \(T=0.5\varepsilon /k_\mathrm{B}\), and in the case of (b), \(h_\mathrm{c}\) is set to a constant value of \(h_\mathrm{c}=1.3\sigma \). In Fig. 7a, g(r) exhibits pronounced peaks for all values of the hydrophobicity parameter which is varied in the range \(h_\mathrm{c}\in [0.7,1.3]\). These peaks indicate a very strong overall aggregation of the lipids and are all the more pronounced with increasing scope of the hydrophobic interaction. The peaks are clearly visible at least until a distance of \(r=6\sigma \). Thus, the structure of these systems is that of a crystal-like solid with a pronounced long-range order in accordance with our conclusions drawn for the order parameter \(O_2(\tau )\) in Fig. 6a. In Fig. 7b where the temperature is varied, we can see that for g(r) the resulting aggregated membrane structures at equilibrium are more sensitive to a change in temperature T than to a change of the hydrophobic interaction range. For the lowest temperatures, the system is practically frozen into a quasi-crystalline state, exhibiting long-range order even at distances beyond \(r=6\sigma \). With increasing temperature the system attains a different structure with only local order up to distances of roughly \(r=3\sigma \). Hence, in this range of temperature, the structural features of the aggregated lipids are similar to fluid behavior with fluctuations in the bilayer membrane height and shape. Finally, for very large temperatures, only the first- and the second-nearest neighbour peaks remain which reflects in essence the local structure of single lipid molecules. This means that no aggregation of lipids occurs for this range of temperatures and the given value of \(h_\mathrm{c}\), and no bilayer membranes are formed. All of these results are consistent with our discussion of the order parameter \(O_2(\tau )\) in Fig. 6b and also with the phase diagram of our model which is presented in the next section.
3.4 Phase diagram of our CG lipid model
In this section, we present a phase diagram for our lipid model introduced in Sect. 2 for a large range of temperature T and interaction parameter \(h_\mathrm{c}\), in which we can distinguish at least four different phases of system behavior:
-
I:
In a gas phase, denoted with crosses in Fig. 8 no self-assembly of lipids occurs. Here, for a given attractive interaction range \(h_\mathrm{c}\) of the lipid tails, the temperature and consequently, the thermal motion of the lipids are too large to allow for membrane formation.
-
II:
In a fluid phase, we observe formation of stable membranes, denoted with blue squares in Fig. 8. The self-assembled membranes of this domain are the ones for which we proceed to show in Sect. 3.5 that here we obtain values for the bending stiffness \(\kappa _\mathrm{B}\) and the area compression modulus \(K_\mathrm{A}\) which are within the range of typical experimental values for membranes in vivo.
-
III:
In a third quasi-solid phase, clustering of lipids leads to stable bilayer membranes with a solid-like molecular structure with long-range order as can be seen in the distinctive peaks of g(r) in Fig. 7. This area in the phase diagram is denoted with upper triangles in Fig. 8. The bilayer structures in this region tend to be rather rigid and exhibit much less fluctuations in shape and height.
-
IV:
In a fourth vesicle phase, for larger values of both, \(h_\mathrm{c}\) and T, we observe the formation of closed membrane vesicles within the fluid domain of the phase diagram, indicated in Fig. 8 with filled black squares.
Each data point in Fig. 8 is obtained by analyzing at least five different realizations of fully equilibrated systems with the respective parameter pair \((h_\mathrm{c},T)\) and simulation times of at least \(\tau =10^{5}\) after equilibration. The solid-fluid transition, as well as the transition between fluid phase and gas phase, where no aggregation of lipids occurs, can be distinguished rather unambiguously based on using a combination of evaluation methods: Visual inspection of the simulated systems, the pair correlation function g(r), and the order parameter \(O_{2}(\tau )\). Whether or not closed membrane vesicles in the lipid domain of the phase diagram are formed, additionally depends on the size of the simulated system, i.e., on the number of lipid molecules and on the size of the simulation box, which both have to be large enough to allow for vesicle formation. With our model, we observe vesicle formation in the corresponding parameter domain, as soon as the number of lipids exceeds \(\sim 900\).
3.5 Elastic modulus and density profile of lipid membranes
In this section, we show that our lipid model leads to the aggregation of double lipid membranes which exhibit properties of typical biological membranes under physiological conditions. We show this by calculating the area compression modulus \(K_\mathrm{A}\) and the bending elasticity modulus \(\kappa _\mathrm{B}\) of the membranes obtained in our computer simulations.
The bending elasticity modulus \(\kappa _\mathrm{B}\) determines by how much an applied external force deforms a lipid bilayer from its perfectly flat energetic ground state. It can be obtained from linear response theory by analyzing the bilayer height fluctuation spectrum. From Helfrich’s linearized continuum theory [90] one can derive the following relation for a two-dimensional elastic sheet: [91, 92].
where
is a Fourier component of the deviation of the local bilayer height \(z_{j}\) with respect to its average height \(z_0\). The sum in Eq. (10) runs over all bilayer particles with coordinates \(x_j, \,\,y_j\), and \(z_j\), where \(\mathbf {n}\) is a two-dimensional vector with components \(n_x\) and \(n_y\) in reciprocal space, which relates to a wave-vector \(\mathbf {q}\) through
The quantity t in Eq. (9) is the surface tension
which is related to the diagonal components of the pressure tensor \(p_{\alpha \beta }\). In our MD simulations, the pressure is isotropic with \(p_{zz} = p_{xx} = p_{yy}\), i.e., the surface tension vanishes. Expressing Eq. (9) in terms of wave-vectors, we finally obtain for the regime of small wave vectors
As shown in Fig. 9, the asymptotic \(q^{-4}\) scaling regime is recovered at our system size, which implies that the characteristic lipid bilayer bending modes can be accommodated with this size of the simulation box. We obtained the bending rigidities by fitting Eq. (9) to the simulation data. Continuum theory [91] relates \(\kappa _\mathrm{B}\) to the area compression modulus via
with \(d_\mathrm{b}\) being the bilayer thickness, which we assume to be 5 nm. Results of simulations for these quantities are provided in Table 1. The values obtained from our simulations are well within the typical experimental range of these observables, which is \(K_\mathrm{A} \simeq 234 \, \mathrm {mN/m}\), and \(\kappa _\mathrm{B} = 10 - 40 \, k_BT\), depending on the technique used, such as micropipet aspiration or various forms of optical measurements [93, 94].
3.6 Shock-wave simulations of CG bilayer membranes
In this section, we present shock-wave simulations of our CG model of lipids, introduced in Sect. 2. Our shock-wave simulations are based on a multiscale coupling scheme presented in detail in Ref. [85]. For presenting the properties of our model when interacting with shock waves, we map our reduced model units onto physical units. The units are connected by the timescale \(\tau \), the mass m of the particles, the length scale \(\sigma \) and the energy scale \(\varepsilon \). The connection between reduced and absolute units has been described in standard literature [27, 28, 30] and for further details we refer to these references. In order to achieve a mapping between units, it is easiest to make a decision about the size of our model particles and the membranes they are supposed to represent, respectively. We assume a bilayer thickness of \(5\,\)nm. We have a three-particle CG model for a lipid that corresponds to a DPPC molecule, the mass of which is known due to its chemical formula provided in the caption of Fig. 1. With this, we can obtain a timestep t in absolute units according to \(t = \sqrt{m\sigma ^2/\varepsilon }\cdot \tau \). As a result, time and length are mapped to absolute units.
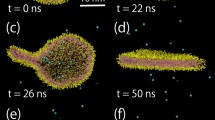
Snapshots of a large scale shock-wave simulation with a lipid bilayer in a shock tube at two different times: a, b correspond to a simulation time of \(0.72\,ps\) and c, d correspond to a simulation time of \(8.61\,ps\). The simulation box measures \(\left( 62.90\times 15.01\times 15.01 \right) \, {\mathrm {nm}}^3\), comprising 1,117,556 particles, 90,000 of which are MD particles, i.e., 30,000 lipid molecules, and the other ones are SPH particles, i.e integration points for the hydrodynamic equations. A piston, coming from the left and moving with a velocity of \(4730 \, \mathrm {m}\mathrm {s}^{-1}\), compresses the material ahead of it and induces a shock wave. In a) and d), the particles are colored according to type (blue water, red lipid head, green lipid tail). In b, c, the pressure distribution is shown, with blue and red signifying low and high pressures, respectively. Peak pressures in b are 38.4 GPa and 1.9 GPa in c, respectively. The lipid bilayer has a momentum to the right during the simulation, as the compressing piston introduces a finite linear momentum in shock compression direction. (Color figure online)
Three-dimensional perspective view of the large-scale simulation snapshots presented in Fig. 10. This is a simulation with \(1.1\times 10^6\) particles. In a, the system is displayed just after the shock wave has been initiated. In b, the system is shown when the shock wave has passed the membrane. One can see a rather complete destruction of the membrane. To get a better viewpoint, the first two layers of fluid particles surrounding the membrane are displayed in blue. The hydrophobic tail particles are colored in red and the hydrophilic lipid heads are colored in green. In b, one can clearly recognize that fluid particles are penetrating the membrane, that is, its initial structure is completely destroyed. This process can be seen in the provided Electronic Supplementary Material (ESM) movie, Movie_S3, from which the two snapshots presented in this figure have been taken. (Color figure online)
As the lipid bilayer is not a crystalline solid, the shock-wave does not experience a major change in impedance as it traverses the interface between water and bilayer, and it is a priori unclear by which mechanisms damage is caused, and how it depends on physical parameters such as shock-wave velocity, shock pulse duration, or shock pulse shape.
An interesting principal objection against the necessity of using CG models vs. atomistic models for shock-wave research might be the following argument: The largest atomistic simulation of phospholipid molecules organized in a bilayer membrane to this day was done by Koshiyama [49] as discussed in our introduction. So, if we assume for the moment that Moore’s law will still be valid in the future (in fact, it has ceased to be valid already today), we could in principle wait until the computers are powerful enough to simulate shock-wave damage in a membrane which has the size of a typical eukaryotic cell, i.e., \(\approx 30-50\mu \)m and then simply upscale the atomistic model. However, a best case estimation of the time needed leaves us with a time span of at least 40 years. Thus, mere brute force computational speed is not a useful option. This is why we have to come up with more clever ways to bridge the length scales from the nano to the micro regime and beyond. Thus, in the following, we use the MD-SPH coupling scheme of Ref. [85] and our CG model of lipids in the MD regime to investigate the damage of shock waves in membranes, positioned in a shock tube geometry. The shock tube setup has been used successfully in experimental studies of shock-wave effects on biological cells [2, 5, 43, 95, 96].
We perform shock-wave simulations on square lipid bilayer patches that were aggregated in the fluid phase for values of the lipid interaction parameter in the interval \(h_\mathrm{c} \in \left[ 0.8, 1.4\right] \) and temperature \(T\in \left[ 0.5,1.0\right] \). For \(h_\mathrm{c} < 0.7\), the bilayer dissolves, while it is crystalline for \(h_\mathrm{c} >1.5\). Thus, the investigated range is representative of a wide spectrum of lipid bilayers with different mechanical stabilities. Initial configurations are taken from equilibrium runs as described previously. The driving speed of the piston that generates the shock wave is varied in the range \(v_p \in \left[ 1892, 5676\right] \mathrm {ms}^{-1}\) in order to produce shock waves with different supersonic velocities. Snapshots of an exemplary large-scale shock-wave simulation are shown in Fig. 10. Here, the shock wave is initiated by a piston with velocity \(v_p=4730 \, \mathrm {m}\mathrm {s}^{-1}\). After 200 time steps of \(\delta t=2.87 \, \mathrm {fs}\), the piston is stopped, while the initiated shock wave continues to travel further along \(L_{\bot }\). The lipid bilayer is placed \(9.435 \, {\mathrm {nm}}\) in front of the initial piston position, far enough away not to be hit by the piston.
In Fig. 11, we show an enlarged three-dimensional section of the two simulation snapshots of Fig. 10a and c, displaying the membrane CG particles and the SPH water particles. In Fig. 11a the shock-wave front has just started to propagate through the shock tube. In b), the shock front has already passed the bilayer membrane causing damage. As one can see, the lipid bilayer membrane is surrounded by SPH particles which are modeled using an equation of state for \(\hbox {H}_{2}\hbox {O}\). Membrane molecules are color-coded in green (hydrophillic head part), red (hydrophobic tail) and one layer of water molecules is displayed in blue. Note that after the shock wave has passed the membrane, water molecules penetrate into the lipid bilayer, the structure of which is destroyed. The whole process of shock-wave propagation displayed in the two snapshots of Fig. 11 can be watched in the movie Movie_S3 which is provided as Electronic Supplementary Material (ESM).
For a thorough analysis of shock-wave experiments, it is necessary to follow the shock front velocity and peak pressure, as it travels through the simulation box and dissipates. Consequently, we discretize the simulation volume in thin slabs of width \(0.5 \, {\mathrm {nm}}\) which are oriented parallel to the xy-plane, while the z-direction is the direction of shock-wave propagation. By computing the average velocity and pressure in each slab for each simulation timestep individually, the evolution of the shock-wave front can be studied in detail.
The pressure profile corresponding to the system displayed in Figs. 10 and 11 is presented in Fig. 12. At \(t=0 \, \mathrm {ps}\), one can identify the location of the lipid bilayer by the interfacial pressure variations near \(z \simeq 9.5 \, {\mathrm {nm}}\). The profiles at \(t=1.00 \, \mathrm {ps}\) and \(t=12.50\, \mathrm {ps}\) clearly show the shock-wave front with an associated pressure peak. The fast dissipation of the shock wave is demonstrated by the quickly diminishing peak pressure heights. The minor pressure peak near \(z \simeq 20 \, {\mathrm {nm}}\) in the \(t=12.5 \mathrm {ps}\) profile is due to the bilayer membrane, which has moved to the right during the simulation, as the compressing piston introduces a finite linear momentum. Interestingly, the bilayer retains a relatively high pressure after the shock-wave front has passed over it. This can be explained by noting that, in contrast to the fluid particles, the CG lipid molecules contain anharmonic springs according to Eq. (3), which can store energy in oscillatory motions.
3.6.1 Damage analysis
There are a number of different phenomena associated with lipid bilayer damage:
-
The bilayer can tear along well-localized paths leaving the majority of its area intact.
-
Single lipids can move out of their equilibrium positions and orientations via diffusive mechanisms.
-
The two bilayer leaflets can interpenetrate upon compression.
We adopt a pragmatic approach to combine all of these effects into a single-scalar order parameter, \(\varPsi \), which is based on the projection of the particle pair distribution function on rotational invariants [97]. To do so, we define the orientation dependent correlation function
where the sum runs over all pairs of lipids i and \(j, \,\,r_{ij}\) is the pair distance as measured between the central beads, \(\delta \) is Dirac‘s delta distribution, and \(\mathbf {e}_i\) and \(\mathbf {e}_j\) are orientation vectors (the normalized distance vectors between the first and the third bead of each lipid). This function was introduced by one of the authors (M. O. Steinhauser) in Ref. [98]. C(r) is similar to the familiar radial distribution function, cf. Eq. (8), except that it is additionally weighted by relative orientations. To convert this distance dependent correlation function into a single scalar, we integrate C(r) up to a distance \(r_c=1.0 \, {\mathrm {nm}}\), which is chosen such as to include the first coordination shell, i.e., the first peak in C(r), in the undamaged bilayer structure:
Disruption of the equilibrium bilayer configuration affects this order parameter in two different ways: If the mutual orientation of neighboring lipids deviates from parallel alignment, or if lipids are separated from each other beyond their equilibrium distance \(r_c\), the value of \(\varPsi \) is reduced, thus providing a quantitative means to characterize lipid bilayer damage.
Analysis of membrane damage due to shockwaves. a Plot of membrane damage \(\varPsi \) induced by shock waves of different incident velocities \(v_\mathrm{s}\). The data shown are for our CG lipid model with interaction parameter \(h_\mathrm{c}\,=\,1.4\) and two different system sizes: 1000 lipids (filled circles) and 10000 lipids (open squares), respectively. b Effect of lipid attraction parameter \(h_\mathrm{c}\) on bilayer damage induced by shock waves of different incident velocities \(v_\mathrm{s}\). Filled circles exhibit results for the CG lipid model with interaction parameter \(h_\mathrm{c}\,=\,1.4\), being representative of a very stable lipid bilayer. Open squares show the induced damage for lipids with \(h_\mathrm{c}\,=\,0.8\), which is close to the lower end of the fluid bilayer regime in the phase diagram in Fig. 8
In Fig. 13, we present the results of our bilayer damage analysis. Figure 13a displays an investigation of finite size effects in two systems containing 1000 and 10000 lipids, respectively, and for a whole range of different shock front velocities \(v_{s}\). The lipid bilayer was chosen to be of medium stability, with an interaction parameter \(h_\mathrm{c}=1\). A clear trend emerges from the result, with higher impact velocities causing a stronger decrease of \(\varPsi \). The initial steep decrease in \(\varPsi \) is almost instantaneous for all shock-wave speeds considered, however, the following behavior is dependent on \(v_s\): for \(v_s \le 3000\, \, \mathrm {m}\mathrm {s}^{-1}, \,\,\varPsi \) increases within a few ps, indicating reversible damage. For \(v_s \ge 3920 \, \mathrm {m}\mathrm {s}^{-1}\), no such recovery can be observed. We note that the two system sizes considered here show no significant difference in the resulting variation of the damage parameter \(\varPsi \). Hence, we conclude that these system sizes are already large enough to study simple aggregate order parameters which are computed by averaging over one configuration. We also note that the data points for faster \(v_s\) in Fig. 13 extend to shorter times than those for slower \(v_s\), as the simulation box size is fixed and a faster shock-wave takes less time to traverse the box.
Our CG model introduced in Sect. 2 is capable of simulating lipid bilayers of different stability, ranging from unstable bilayers over a transition state to almost solid bilayers, close to the crystallization transition to closed vesicles. We exploit this feature in order to study the relation of bilayer stability and shock-wave-induced damage. Thus, in Figure 13 b), we compare the variation of \(\varPsi \) for two bilayers with \(h_\mathrm{c}=0.8\) and \(h_\mathrm{c}=1.4\), which are representative of the extreme ends of the stable bilayer range, cf. Fig. 8. Here, we use systems with 1000 lipids. We justify our small choice of system size—1000 lipids—by noting that it is large enough to capture the characteristic bilayer undulations, which are driven by thermal fluctuations. This has been verified before in Fig. 9, where we have shown that the Fourier transform of the lipid bilayer height fluctuations reaches the asymptotic scaling regime predicted by continuum theory. We therefore assume that such a lipid bilayer patch is a representative element of a real membrane and that the periodic boundary conditions which we impose do not introduce large artificial errors.
The graph shows a clear difference in the resulting damage of these two systems, however, it is surprising that bilayer stability leads to a quantitatively small effect, especially at long times after shock-wave impact. A possible explanation for this result is that the difference in cohesive energy density between these two systems is smaller by orders of magnitude than the energy densities introduced temporarily by the shock wave. In this analysis, we see again that for for \(v_s \le 3000 \, \mathrm {m}\mathrm {s}^{-1}, \,\,\varPsi \) increases within a few ps, indicating reversible damage which is not the case for shock velocities above \(v_s \ge 3920 \, \mathrm {m}\mathrm {s}^{-1}\).
4 Summary and concluding remarks
We have presented in this paper MD simulations of a CG solvent-free model that is derived from a well-tested polymer model. We analyze the dynamics of the lipid molecules when self-organizing into numerous, small clusters of lipid molecules, driven by the particular interaction potentials of our CG model that mimics the amphiphilic properties of phospholipid molecules. We observe several stages of clustering while the system approaches equilibrium and minimizes its total energy, until finally all individual clusters merge into a single stable structure. The most common stable structure produced with our model is a fluid bilayer membrane, either as flat membrane or as closed vesicle, depending on the specific choice of the simulation parameters temperature T and lipid interaction potential \(h_\mathrm{c}\). In the present study, we vary the parameter range of our lipid interaction potential such that mechanically barely stable bilayers as well as very stable bilayers can be simulated. For stable bilayer membranes at equilibrium, we are able to show that with a proper choice of our model parameters we obtain values for the compression modulus \(K_C\) and the bending stiffness \(\kappa _\mathrm{B}\) within the range of typical experimental values. Therefore, our numerical experiments are representative of a wide range of different real phospholipid bilayers.
We investigate the local structure of the bilayer membranes at equilibrium by means of the pair correlation function which exhibits a sensitive dependence on the temperature with a given lipid tail interaction range \(h_\mathrm{c}\), and we introduce an order parameter \(O_{2}\) based on the Legendre polynomials to further elucidate quantitatively the transition from a random initial gas of unconnected lipid chains to highly ordered fluid or quasi-solid phases. For \(O_{2}\), we also find a much greater sensitivity to a variation of temperature than to a variation of interaction range \(h_\mathrm{c}\). Furthermore, we determine the parameter combinations for temperature T and lipid tail interaction \(h_\mathrm{c}\) for the different phases and for the aggregation of lipids in closed membrane vesicles. With the detailed analyses of the membranes properties presented in this simulation study, we have added some interesting aspects to the discussion of MD-based CG membrane models that were not included in similar previous studies [31, 59, 61, 74, 76–78].
The results of our multiscale shock-wave simulations which bridge the gap of a detailed description of the lipids using discrete MD particles and the continuum SPH particles, indicate the existence of a threshold shock front velocity \(v_s\approx 3000 \, \mathrm {ms}^{-1}\), below which the bilayer recovers from shock-wave-induced damage on the timescale of at least several tens of ps. Above this velocity, no such recovery can be observed. Most of the damage is caused instantaneously while the shock-wave front traverses the lipid bilayer, with only little more damage occurring afterwards. A strong correlation between \(v_s\) and the induced damage is observed, however, the dependence of the induced damage on the lipid attraction parameter \(h_\mathrm{c}\) is found to be weak.
We suggest that new shock-wave experiments with standard cell lines should be performed that investigate the damage in the cells to see whether the existence of a shock front velocity threshold, separating reversible and irreversible (permanent) membrane damage can be verified experimentally. Such experiments would require high-speed camera equipment and the possibility of performing single cell shock-wave experiments, e.g., by including an AFM (atomic force microscope) with the capability to select and measure the properties of single cells before and after shock-wave treatment.
In future applications of our model, we plan to investigate the effect of the interaction of shock waves with closed vesicles on a length scale comparable to that of biological cells (tens of micrometers), which might be used as simple mechanical models of the plasma membrane of eukaryotic cells. The larger context of this research program is the attempt to enhance our fundamental understanding of shock-wave interaction with soft matter systems, in particular with cellular systems and biomolecules.
To the best of our knowledge, the here presented idea of using coarse-grained particle models for soft matter systems in combination with the investigation of shock-wave effects in these systems is a fairly new approach. Revealing the mechanisms by which shock-wave effects lead to mechanical damage in soft biological matter could further enhance our understanding of tumor therapies that are based on shock-wave interactions with neoplasia.
References
McNeil PL, Terasaki M (2001) Coping with the inevitable: how cells repair a torn surface membrane. Nat Cell Biol 3(5):E124–E129
Schmidt M, Kahlert U, Wessolleck J, Maciaczyk D, Merkt B, Maciaczyk J, Osterholz J, Nikkhah G, Steinhauser MO (2014) Characterization of s setup to test the impact of high-amplitude pressure waves on living cells. Nat Sci Rep 4:3849-1–3849-9
Gambihler S, Delius M, Ellwart JW (1992) Transient increase in membrane permeability of L1210 cells upon exposure to lithotripter shock waves in vitro. Die Naturwissenschaften 79:328–329
Gambihler S, Delius M, Ellwart JW (1994) Permeabilization of the plasma membrane of L1210 mouse leukemia cells using lithotripter shock waves. J Membr Biol 141:267–275
Kodama T, Doukas AG, Hamblin MR (2002) Shock wave-mediated molecular delivery into cells. Biochim Biophys Acta 1542:186–194
Bao G, Suresh S (2003) Cell and molecular mechanics of biological materials. Nat Mater 2:715–725
Tieleman DP, Leontiadou H, Mark AE, Marrink S-J (2003) Simulation of pore formation in lipid bilayers by mechanical stress and electric fields. J Am Chem Soc 125:6382–6383
Sundaram J, Mellein BR, Mitragotri S (2003) An experimental and theoretical analysis of ultrasound-induced permeabilization of cell membranes. Biophys J 84:3087–3101
Doukas AG, Kollias N (2004) Transdermal drug delivery with a pressure wave. Adv Drug Deliv Rev 56:559–579
Coussios C-C, Roy RA (2008) Applications of acoustics and cavitation to noninvasive therapy and drug delivery. Ann Rev Fluid Mech 40:395–420
Prausnitz MR, Langer R (2008) Transdermal drug delivery. Nat Biotechnol 26:1261–1268
Ashley CE, Carnes EC, Phillips GK, Padilla D, Durfee PN, Brown PA, Hanna TN, Liu J, Phillips B, Carter MB, Carroll NJ, Jiang X, Dunphy DR, Willman CL, Petsev DN, Evans DG, Parikh AN, Chackerian B, Wharton W, Peabody DS, Brinker CJ (2011) The targeted delivery of multicomponent cargos to cancer cells by nanoporous particle-supported lipid bilayers. Nat Mater 10:389–397
Koshiyama K, Wada S (2011) Molecular dynamics simulations of pore formation dynamics during the rupture process of a phospholipid bilayer caused by high-speed equibiaxial stretching. J Biomech 44:2053–2058
Phillips R, Kondev J, Theriot J (2009) Physical biology of the cell. Garland Science. Taylor and Francis Group, New York
Alberts B, Bray D, Hopkin K, Jonson A, Lewis J, Raff M, Roberts K, Walter P (2010) Essential cell biology. Garland Science. Taylor and Francis Group, New York
Alberts B, Bray D, Johnson A, Lewis J, Morgan D, Raff M, Roberts K, Walter P (2000) Molecular biology of the cell. Garland Science. Taylor and Francis Group, New York
Steinhauser MO (2006) Computational methods in polymer physics. Recent Res Dev Phys 7:59–97
Steinhauser MO (2008) Computational multiscale modeling of fluids and solids—theory and applications. Springer, Berlin
Steinhauser MO, Schneider J, Blumen A (2009) Simulating dynamic crossover behavior of semiflexible linear polymers in solution and in the melt. J Chem Phys 130:164902-1–164902-8
De Weer P (2000) A century of thinking about cell membranes. Annu Rev Physiol 62:919–926
Singer SJ, Nicolson GL (1972) The fluid mosaic model of the structure of cell membranes. Science 175:720–731
Yang K, Ma YQ (2010) Computer simulation of the translocation of nanoparticles with different shapes across a lipid bilayer. Nat Nanotechnol 5:579–583
Das C, Sheikh KH, Olmsted PD, Connell SD (2010) Nanoscale mechanical probing of supported lipid bilayers with atomic force microscopy. Phys Rev E 82:041920-1–041920-6
Edidin M (2003) Lipids on the frontier: a century of cell-membrane bilayers. Nat Rev Mol Cell Biol 4:414–418
Bowick MJ, Travesset A (2001) The statistical mechanics of membranes. Phys Rep 344:255–308
Nelson D, Piran T, Weinberg S (2004) Statistical mechanics of membranes and surfaces. World Scientific Publishing, Singapore
Allen MP, Tildesley DJ (1987) Computer simulation of liquids. Clarendon Press, Oxford
Rapaport DC (2004) The art of molecular dynamics simulation. Cambridge University Press, Cambridge
Marrink SJ, Risselada HJ, Yefimov S, Tieleman DP, de Vries AH (2007) The MARTINI force field: coarse grained model for biomolecular simulations. J Phys Chem B 111:7812–7824
Steinhauser MO (2013) Computer simulation in physics and engineering. Walter de Gruyter, Berlin
Drouffe JM, Maggs AC, Leibler S (1998) Computer simulations of self-assembled membranes. Science 254:1353–1356
Goetz R, Lipowsky R (1998) Computer simulations of bilayer membranes: self-assembly and interfacial tension. J Chem Phys 108:7397–7409
Noguchi H, Takasu M (2001) Self-assembly of amphiphiles into vesicles: a Brownian dynamics simulation. Phys Rev E 64:041913-1–041913-7
Bourov GK, Bhattacharya A (2005) Brownian dynamics simulation study of self-assembly of amphiphiles with large hydrophilic heads. J Chem Phys 122:44702-1–44702-6
Noguchi H, Gompper G (2006) Dynamics of vesicle self-assembly and dissolution. J Chem Phys 125:164908-1–164908-13
Tribet C, Vial F (2008) Flexible macromolecules attached to lipid bilayers: impact on fluidity, curvature, permeability and stability of the membranes. Soft Matter 4:68–81
Yang S, Qu J (2014) Coarse-grained molecular dynamics simulations of the tensile behavior of a thermosetting polymer. Phys Rev E 90:012601-1–012601-8
Brannigan G, Lin LCL, Brown FLH (2006) Implicit solvent simulation models for biomembranes. Eur Biophys J 35:104–124
Dror RO, Jensen MO, Borhani DW, Shaw DE (2010) Perspectives on: molecular dynamics and computational methods: exploring atomic resolution physiology on a femtosecond to millisecond timescale using molecular dynamics simulations. J Gen Physiol 135:555–562
Götz AW, Williamson MJ, Xu D, Poole D, Le Grand S, Walker RC (2012) Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. Generalized born. J Chem Theory Comput 8:1542–1555
Skjevik AA, Madej BD, Dickson CJ, Teigen K, Walker RC, Gould IR (2015) All-atom lipid bilayer self-assembly with the AMBER and CHARMM lipid force fields. Chem Commun 51:4402–4405
Steinhauser MO (2012) Modeling dynamic failure behavior in granular and biological materials: emerging new applications. IJATEMA 1:15–29
Steinhauser MO, Schmidt M (2014) Destruction of cancer cells by laser-induced shock waves: recent developments in experimental treatments and multiscale computer simulations. Soft Matter 10:4778–4788
Huber PE, Jenne J, Debus J, Wannenmacher MF, Pfisterer P (1999) A comparison of shock wave and sinusoidal-focused ultrasound-induced localized transfection of HeLa cells. Ultrasound Med Biol 25:1451–1457
Dubinsky TJ, Cuevas C, Dighe MK, Kolokythas O, Hwang JH (2008) High-intensity focused ultrasound: current potential and oncologic applications. Am J Roentgenol 190:191–199
Gevaux D (2010) Physics and the cell. Nat Phys 6:725
Fritsch A, Höckel M, Kiessling T, Nnetu KD, Wetzel F, Zink M, Käs JA (2010) Are biomechanical changes necessary for tumour progression? Nat Phys 6:730–732
Koshiyama K, Kodama T, Yano T, Fujikawa SS (2006) Structural change in lipid bilayers and water penetration induced by shock waves: molecular dynamics simulations. Biophys J 91:2198–2205
Koshiyama K, Kodama T, Yano T, Fujikawa SS (2008) Molecular dynamics simulation of structural changes of lipid bilayers induced by shock waves: effects of incident angles. Biochim Biophys Acta 1778:1423–1428
Lechuga J, Drikakis D, Pal S (2008) Molecular dynamics study of the interaction of a shock wave with a biological membrane. Int J Numer Mech Fluids 57:677–692
Sridhar A, Srikanth B, Kumar A, Dasmahapatra AK (2015) Coarse-grain molecular dynamics study of fullerene transport across a cell membrane. J Chem Phys 143:024907-1–024907-9
Hofmann A, Ritz U, Rompe JD, Tresch A, Rommens PM (2015) The effect of shock wave therapy on gene expression in human osteoblasts isolated from hypertrophic fracture non-unions. Shock Waves 25:91–102
Hoon Ha C, Cheol Lee S, Kim S, Chung J, Bae H, Kwon K (2015) Novel mechanism of gene transfection by low-energy shock wave. Nat Sci Rep 5:12843-1–12843-13
Zhang Z, Lu L, Noid WG, Krishna V, Pfaendtner J, Voth GA (2008) A systematic methodology for defining coarse-grained sites in large biomolecules. Biophys J 95:5073–5083
Noguchi H (2009) Membrane simulation models from nanometer to micrometer scale. J Phys Soc Jpn 78:041007
Murtola T, Karttunen M, Vattulainen I (2009) Systematic coarse graining from structure using internal states: application to phospholipid/cholesterol bilayer. J Chem Phys 131:055101-1–055101-15
Müller M (2011) Studying amphiphilic self-assembly with soft coarse-grained models. J Stat Phys 145:967–1016
Lyubartsev AP, Rabinovich AL (2010) Recent development in computer simulations of lipid bilayers. Soft Matter 7:25–39
Huang MJ, Kapral R, Mikhailov AS, Chen HY (2012) Coarse-grain model for lipid bilayer self-assembly and dynamics: multiparticle collision description of the solvent. J Chem Phys 137:055101-1–055101-10
May A, Pool R, van Dijk E, Bijlard J, Abeln S, Heringa J, Feenstra KA (2014) Coarse-grained versus atomistic simulations: realistic interaction free energies for real proteins. Bioinformatics 30:326–334
Yuan H, Huang C, Li J, Lykotrafitis G, Zhang S (2010) One-particle-thick, solvent-free, coarse-grained model for biological and biomimetic fluid membranes. Phys Rev E 82:011905-1–011905-8
Noguchi H (2011) Solvent-free coarse-grained lipid model for large-scale simulations. J Chem Phys 134:055101-1–055101-12
Warshel A, Levitt M (1976) Theoretical studies of enzymic reactions: dielectric, electrostatic and steric stabilization of the carbonium ion in the reaction of lysozyme. J Mol Biol 103:227–249
Dünweg B, Reith D, Steinhauser MO, Kremer K (2002) Corrections to scaling in the hydrodynamic properties of dilute polymer solutions. J Chem Phys 117:914–924
Steinhauser MO (2005) A molecular dynamics study on universal properties of polymer chains in different solvent qualities. Part I. A review of linear chain properties. J Chem Phys 122:094901-1–094901-13
Stevens MJ (2004) Coarse-grained simulations of lipid bilayers. J Chem Phys 2004(121):11942–11948
Steinhauser MO, Grass K, Strassburger E, Blumen A (2009) Impact failure of granular materials—non-equilibrium multiscale simulations and high-speed experiments. Int J Plast 25:161–182
Saunders MG, Voth GA (2012) Coarse-graining methods for computational biology. Annu Rev Biophys 42:73–93
Nielsen SO, Lopez CF, Srinivas G, Klein ML (2004) Topical review: coarse grain models and the computer simulation of soft materials. J Phys-Condens Matter 16:481–R512
Pogodin S, Baulin VA (2010) Coarse-grained models of phospholipid membranes within the single chain mean field theory. Soft Matter 6:2216–2226
Wang Y, Sigurdsson JK, Brandt E, Atzberger PJ (2013) Dynamic implicit-solvent coarse-grained models of lipid bilayer membranes: fluctuating hydrodynamics thermostat. Phys Rev E 88:023301-1–023301-5
Farago O (2003) ”Water-free” computer model for fluid bilayer membranes. J Chem Phys 119:596–605
Brannigan G, Philips PF, Brown FLH (2005) Flexible lipid bilayers in implicit solvent. Phys Rev E 72:011915-1–011915-4
Sodt AJ, Head-Gordon T (2010) An implicit solvent coarse-grained lipid model with correct stress profile. J Chem Phys 132:205103-1–205103-8
Shih AY, Arkhipov A, Freddolino PL, Schulten K (2006) Coarse grained protein-lipid model with application to lipoprotein particles. J Phys Chem B 110:3674–3684
Noguchi H, Takasu M (2002) Adhesion of nanoparticles to vesicles: a brownian dynamics simulation. Biophys J 83:299–308
Cooke IR, Deserno M (2005) Solvent-free model for self-assembling fluid bilayer membranes: stabilization of the fluid phase based on broad attractive tail potentials. J Chem Phys 123:224710-1–224710-13
Izvekov S, Voth GA (2009) Solvent-free lipid bilayer model using multiscale coarse-graining. J Phys Chem B 113:4443–4455
Wang ZJ, Frenkel D (2005) Modeling flexible amphiphilic bilayers: a solvent-free off-lattice Monte Carlo study. J Chem Phys 122:234711-1–234711-8
Shelley JC, Shelley MY, Reeder RC, Bandyopadhyay S, Klein ML (2001) A coarse grain model for phospholipid simulations. J Phys Chem B 105:4464–4470
Shelley JC, Shelley MY, Reeder RC, Bandyopadhyay S, Moore PB, Klein ML (2001) Simulations of phospholipids using a coarse grain model. J Phys Chem B 105:9785–9792
Marrink SJ, de Vries AH, Mark AE (2003) Coarse grained model for semiquantitative lipid simulations. J Phys Chem B 108:750–760
Brannigan G, Brown FLH (2004) Solvent-free simulations of fluid membrane bilayers. J Chem Phys 120:1059–1071
Cooke IR, Kremer K, Deserno M (2005) Tunable generic model for fluid bilayer membranes. Phys Rev E 72:011506-1–011506-4
Ganzenmüller GC, Hiermaier S, Steinhauser MO (2012) Energy-based coupling of smooth particle hydrodynamics and molecular dynamics with thermal fluctuations. Eur Phys J Spec Top 206:51–60
Steinhauser MO (2008) Static and dynamic scaling of semiflexible polymer chains—a molecular dynamics simulation study of single chains and melts. Mech Time Depend Mater 12:291–312
Griffiths G (2007) Cell evolution and the problem of membrane topology. Nat Rev Mol Cell Biol 8:1018–1024
Yang K, Ma YQ (2012) Computer simulations of fusion, fission and shape deformation in lipid membranes. Soft Matter 8:606–618
Chernomordirk L, Kozlov MM, Zimmerberg J (1995) Lipids in biological membrane-fusion. J Membr Biol 146:1–14
Helfrich W (1973) Elastic properties of lipid bilayers: theory and possible experiments. Z Naturforsch C 28:693–703
Seifert U (1997) Configurations of fluid membranes and vesicles. Adv Phys 46:13–137
Goetz R, Gompper G, Lipowsky R (1999) Mobility and elasticity of self-assembled membranes. Phys Rev Lett 82:221–224
Olbrich K, Rawicz W, Needham D, Evans E (2000) Water permeability and mechanical strength of polyunsaturated lipid bilayers. Biophys 79:321–327
Lee CH, Lin WC, Wang J (2001) All-optical measurements of the bending rigidity of lipid-vesicle membranes across structural phase transitions. Phys Rev E 64:020901-1–02091-4
Doukas AG, McAuliffe DJ, Lee S, Venugopalan V, Flotte TJ (1995) Physical factors involved in stress-wave-induced cell injury: the effect of stress gradient. Ultrasound Med Biol 21:961–967
Kodama T, Hamblin MR, Doukas AG (2000) Cytoplasmic molecular delivery with shock waves: importance of impulse. Biophys J 79:1821–1832
Hansen JP, McDonald IR (2006) Theory of simple liquids. Academic Press, Oxford
Ganzenmüller GC, Hiermaier S, Steinhauser MO (2011) Shock-wave induced damage in lipid bilayers: a dissipative particle dynamics simulation study. 7:4307–4317
Funding
This study was funded by the Fraunhofer-Gesellschaft, e.V., Germany, under grant no. 400017 “Extracorporeal, Focused Ultrasound Therapy: Effectiveness, Simulation, and Planning of New Therapies” and grant no. 600016, Vintage Class Program: “Shock Wave Induced Destruction of Tumor Cells”.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 2 (mp4 22464 KB)
Rights and permissions
About this article
Cite this article
Steinhauser, M.O., Schindler, T. Particle-based simulations of bilayer membranes: self-assembly, structural analysis, and shock-wave damage. Comp. Part. Mech. 4, 69–86 (2017). https://doi.org/10.1007/s40571-016-0126-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40571-016-0126-3