Abstract
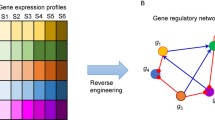
Transcriptional regulatory networks specify the regulatory proteins of target genes that control the context-specific expression levels of genes. With our ability to profile the different types of molecular components of cells under different conditions, we are now uniquely positioned to infer regulatory networks in diverse biological contexts such as different cell types, tissues, and time points. In this chapter, we cover two main classes of computational methods to integrate different types of information to infer genome-scale transcriptional regulatory networks. The first class of methods focuses on integrative methods for specifically inferring connections between transcription factors and target genes by combining gene expression data with regulatory edge-specific knowledge. The second class of methods integrates upstream signaling networks with transcriptional regulatory networks by combining gene expression data with protein–protein interaction networks and proteomic datasets. We conclude with a section on practical applications of a network inference algorithm to infer a genome-scale regulatory network.
The authors Alireza Fotuhi Siahpirani and Deborah Chasman contributed equally.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Markowetz F, Spang R (2007) Inferring cellular networks–a review. BMC Bioinf 8(Suppl 6):S5
Kim HD, Shay T, O’Shea EK, Regev A (2009) Transcriptional regulatory circuits: predicting numbers from alphabets. Science 325(5939):429–432
Thompson D, Regev A, Roy S (2015) Comparative analysis of gene regulatory networks: from network reconstruction to evolution. Annu Rev Cell Dev Biol 31:399–428
Gasch AP, Spellman PT, Kao CM, Carmel-Harel O, Eisen MB, Storz G, Botstein D, Brown PO (2000) Genomic expression programs in the response of yeast cells to environmental changes. Mol Biol Cell 11(12):4241–4257
Ideker T, Krogan NJ (2012) Differential network biology. Mol Syst Biol 8:565
Lee TI, Young RA (2013) Transcriptional regulation and its misregulation in disease. Cell 152(6):1237–1251
Voss TC, Hager GL (2014) Dynamic regulation of transcriptional states by chromatin and transcription factors. Nat Rev Genet 15(2):69–81
de Jong H (2002) Modeling and simulation of genetic regulatory systems: a literature review. J Comput Biol J Comput Mol Cell Biol 9:67–103
Huang S, Kauffman SA (2009) Complex gene regulatory networks – from structure to biological observables: cell fate determination. In: Encyclopedia of complexity and systems science. Springer New York, pp 1180–1213
Carpenter AE, Sabatini DM (2004) Systematic genome-wide screens of gene function. Nat Rev Genet 5(1):11–22
Giaever G, Nislow C (2014) The yeast deletion collection: a decade of functional genomics. Genetics 197(2):451–465
Ren B, Robert F, Wyrick J, Aparicio O, Jennings E, Simon I, Zeitlinger J, Schreiber J, Hannett N, Kanin E, Volkert T, Wilson C, Bell S, Young R (2000) Genome-wide location and function of DNA binding proteins. Science 290(5500):2306–2309
Furey TS (2012) ChIP-seq and beyond: new and improved methodologies to detect and characterize protein-DNA interactions. Nat Rev Genet 13(12):840–852
Song L, Crawford GE (2010) DNase-seq: a high-resolution technique for mapping activegene regulatory elements across the genome from mammalian cells. Cold Spring Harb Protoc 2010(2):pdb.prot5384–pdb.prot5384
Buenrostro JD, Giresi PG, Zaba LC, Chang HY, Greenleaf WJ (2013) Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat Methods 10:1213–1218
MacGilvray ME, Shishkova E, Chasman D, Place M, Gitter A, Coon JJ, Gasch AP (2018) Network inference reveals novel connections in pathways regulating growth and defense in the yeast salt response. PLoS Comput Biol 13(5):1–28
Figeys D (2008) Mapping the human protein interactome. Cell Res 18:716–724
Braun P (2012) Interactome mapping for analysis of complex phenotypes: insights from benchmarking binary interaction assays. Proteomics 12:1499–1518
Friedman N, Nachman I, Peér D (1999) Learning bayesian network structure from massive datasets: The “sparse candidate” algorithm. In: Proceedings of the fifteenth conference on uncertainty in artificial intelligence, UAI’99. Morgan Kaufmann Publishers Inc., San Francisco, CA, pp 206–215
Segal E, Shapira M, Regev A, Pe’er D, Botstein D, Koller D, Friedman N (2003) Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data. Nat Genet 34(2):166–176
Margolin AA, Nemenman I, Basso K, Wiggins C, Stolovitzky G, Favera RD, Califano A (2006) ARACNE: an algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. BMC Bioinf 7(Suppl 1):S7+
Faith JJ, Hayete B, Thaden JT, Mogno I, Wierzbowski J, Cottarel G, Kasif S, Collins JJ, Gardner TS (2007) Large-scale mapping and validation of Escherichia coli transcriptional regulation from a compendium of expression profiles. PLoS Biol 5(1):e8+
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinf 9:559
Joshi A, De Smet R, Marchal K, Van de Peer Y, Michoel T (2009) Module networks revisited: computational assessment and prioritization of model predictions. Bioinformatics 25(4):490–496
Huynh-Thu VA, Irrthum A, Wehenkel L, Geurts P (2010) Inferring regulatory networks from expression data using Tree-Based methods. PLoS One 5(9):e12776+
Haury ACC, Mordelet F, Vera-Licona P, Vert JPP (2012) TIGRESS: trustful inference of gene REgulation using stability selection. BMC Syst Biol 6(1):145+
Tibshirani R (1996) Regression shrinkage and selection via the lasso. J R Stat Soc Ser B (Methodol) 58(1):267–288
Meinshausen N, Bühlmann P (2010) Stability selection. J R Stat Soc Ser B (Stat Methodol) 72(4):417–473
Roy S, Lagree S, Hou Z, Thomson JA, Stewart R, Gasch AP (2013) Integrated module and Gene-Specific regulatory inference implicates upstream signaling networks. PLoS Comput Biol 9(10):e1003252+
Marbach D, Costello JC, Küffner R, Vega NM, Prill RJ, Camacho DM, Allison KR, Aderhold A, Allison KR, Bonneau R, et al (2012) Wisdom of crowds for robust gene network inference. Nat Methods 9(8):796–804
Friedman N (2004) Inferring cellular networks using probabilistic graphical models. Science 303(5659):799–805
De Smet R, Marchal K (2010) Advantages and limitations of current network inference methods. Nat Rev Microbiol 8(10):717–729
Friedman N, Linial M, Nachman I, Pe’er D (2000) Using Bayesian networks to analyze expression data. J Comput Biol 7(3–4):601–620
Pe’er D, Regev A, Tanay A (2002) Minreg: inferring an active regulator set. Bioinformatics (Oxford, England) 18(Suppl 1):S258–S267
Heckerman D, Chickering DM, Meek C, Rounthwaite R, Kadie C (2001) Dependency networks for inference, collaborative filtering, and data visualization. J Mach Learn Res 1:49–75
Werhli AV, Husmeier D (2007) Reconstructing gene regulatory networks with Bayesian networks by combining expression data with multiple sources of prior knowledge. Stat Appl Genet Mol Biol 6(1): Article15
Hill SM, Lu Y, Molina J, Heiser LM, Spellman PT, Speed TP, Gray JW, Mills GB, Mukherjee S (2012) Bayesian inference of signaling network topology in a cancer cell line. Bioinformatics 28(21):2804–2810
Bonneau R, Reiss D, Shannon P, Facciotti M, Hood L, Baliga N, Thorsson V (2006) The inferelator: an algorithm for learning parsimonious regulatory networks from systems-biology data sets de novo. Genome Biol 7(5):R36+
Greenfield A, Hafemeister C, Bonneau R (2013) Robust data-driven incorporation of prior knowledge into the inference of dynamic regulatory networks. Bioinformatics 29(8):1060–1067
Koller D, Friedman N (2009) Probabilistic graphical models: principles and techniques. MIT Press, Cambridge, MA
Grzegorczyk M, Husmeier D, Werhli AV (2008) Reverse engineering gene regulatory networks with various machine learning methods. Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, pp 101–142
Lee SI, Dudley AM, Drubin D, Silver PA, Krogan NJ, Pe’er D, Koller D (2009) Learning a prior on regulatory potential from eQTL data. PLoS Genet 5(1):e1000358
Siahpirani AF, Roy S (2017) A prior-based integrative framework for functional transcriptional regulatory network inference. Nucleic Acids Res 45:e21
Zou H, Hastie T (2005) Regularization and variable selection via the elastic net. J R Stat Soc Ser B Stat Methodol 67(2):301–320
Novershtern N, Regev A, Friedman N (2011) Physical module networks: an integrative approach for reconstructing transcription regulation. Bioinformatics 27(13):i177–i185
Gitter A, Carmi M, Barkai N, Bar-Joseph Z (2013) Linking the signaling cascades and dynamic regulatory networks controlling stress responses. Genome Res 23(2):365–376
Gitter A, Bar-Joseph Z (2013) Identifying proteins controlling key disease signaling pathways. Bioinformatics 29(13):i227–i236
Schulz MH, Devanny WE, Gitter A, Zhong S, Ernst J, Bar-Joseph Z (2012) Drem 2.0: improved reconstruction of dynamic regulatory networks from time-series expression data. BMC Syst Biol 6:104
Chasman D, Walters KB, Lopes TJS, Eisfeld AJ, Kawaoka Y, Roy S (2016) Integrating transcriptomic and proteomic data using predictive regulatory network models of host response to pathogens. PLoS Comput Biol 12:e1005013
Ernst J, Vainas O, Harbison CT, Simon I, Bar-Joseph Z (2007) Reconstructing dynamic regulatory maps. Mol Syst Biol 3:74
Gitter A, Klein-Seetharaman J, Gupta A, Bar-Joseph Z (2011) Discovering pathways by orienting edges in protein interaction networks. Nucleic acids Res 39:e22
Bengio Y, Frasconi P (1996) Input-output HMMs for sequence processing. IEEE Trans Neural Netw 7:1231–1249
Yuan M, Lin Y (2006) Model selection and estimation in regression with grouped variables. J R Stat Soc Ser B (Stat Methodol) 68(1):49–67
Obozinski G, Taskar B, Jordan M (2006) Multi-task feature selection, Technical report 2. Statistics Department, UC Berkeley
Ourfali O, Shlomi T, Ideker T, Ruppin E, Sharan R (2007) SPINE: a framework for signaling-regulatory pathway inference from cause-effect experiments. Bioinformatics 23(13):i359–i366
Silverbush D, Elberfeld M, Sharan R (2011) Optimally orienting physical networks. J Comput Biol J Comput Mol Cell Biol 18:1437–1448
Chasman D, Gancarz B, Hao L, Ferris M, Ahlquist P, Craven M (2014a) Inferring host gene subnetworks involved in viral replication. PLoS Comput Biol 10(5):e1003626
Chasman D, Ho Y, Berry DB, Nemec CM, MacGilvray ME, Hose J, Merrill AE, Lee MV, Will JL, Coon JJ, Ansari AZ, Craven M, Gasch AP (2014b) Pathway connectivity and signaling coordination in the yeast stress-activated signaling network. Mol Syst Biol 10(11):759+
Danna E, Fenelon M, Gu Z, Wunderling R (2007) Generating multiple solutions for mixed integer programming problems. In: Integer programming and combinatorial optimization. Springer, Berlin/Heidelberg, pp 280–294
Stark C, Breitkreutz BJ, Reguly T, Boucher L, Breitkreutz A, Tyers M (2006) BioGRID: a general repository for interaction datasets. Nucleic Acids Res 34(Suppl 1):D535–D539
Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering C, Jensen LJ (2013) String v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res 41(Database issue):D808–D815
Edgar R, Domrachev M, Lash AE (2002) Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30(1):207–210
Leinonen R, Sugawara H, Shumway M, Collaboration INSD (2010) The sequence read archive. Nucleic acids Res 39(Suppl 1):D19–D21
Cahan P, Li H, Morris SA, Lummertz da Rocha E, Daley GQ, Collins JJ (2014) Cellnet: network biology applied to stem cell engineering. Cell 158(4):903–915
Collado-Torres L, Nellore A, Kammers K, Ellis SE, Taub MA, Hansen KD, Jaffe AE, Langmead B, Leek JT (2017) Reproducible RNA-seq analysis using recount2. Nat Biotechnol 35:319–321
Lachmann A, Torre D, Keenan AB, Jagodnik KM, Lee HJ, Silverstein MC, Wang L, Ma’ayan A (2017) Massive mining of publicly available RNA-seq data from human and mouse. bioRXiv preprint
Bernstein MN, Doan A, Dewey CN (2017) MetaSRA: normalized human sample-specific metadata for the sequence read archive. Bioinformatics (Oxford, England) 33:2914–2923
Leek JT, Scharpf RB, Bravo HC, Simcha D, Langmead B, Johnson WE, Geman D, Baggerly K, Irizarry RA (2010) Tackling the widespread and critical impact of batch effects in high-throughput data. Nat Rev Genet 11:733–739
Goh WWB, Wang W, Wong L (2017) Why batch effects matter in omics data, and how to avoid them. Trends Biotechnol 35:498–507
Leek JT, Johnson WE, Parker HS, Fertig EJ, Jaffe AE, Storey JD (2015) SVA: Surrogate Variable Analysis. R package version 3.18.0
Johnson WE, Li C, Rabinovic A (2007) Adjusting batch effects in microarray expression data using empirical bayes methods. Biostatistics (Oxford, England) 8:118–127
Leek JT, Storey JD (2007) Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genet 3:1724–1735
Apweiler R, Bairoch A, Wu CH, Barker WC, Boeckmann B, Ferro S, Gasteiger E, Huang H, Lopez R, Magrane M, et al (2004) Uniprot: the universal protein knowledgebase. Nucleic acids Res 32(Suppl 1):D115–D119
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al (2000) Gene ontology: tool for the unification of biology. Nat Genet 25(1):25–29
Ravasi T, Suzuki H, Cannistraci CV, Katayama S, Bajic VB, Tan K, Akalin A, Schmeier S, Kanamori-Katayama M, Bertin N, et al (2010) An atlas of combinatorial transcriptional regulation in mouse and man. Cell 140(5):744–752
Jin J, Tian F, Yang DC, Meng YQ, Kong L, Luo J, Gao G (2017) Planttfdb 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic acids Res 45(D1):D1040–D1045
Mathelier A, Fornes O, Arenillas DJ, Chen CY, Denay G, Lee J, Shi W, Shyr C, Tan G, Worsley-Hunt R, Zhang AW, Parcy F, Lenhard B, Sandelin A, Wasserman WW (2016) Jaspar 2016: a major expansion and update of the open-access database of transcription factor binding profiles. Nucleic Acids Res 44:D110–D115
Weirauch MT, Yang A, Albu M, Cote AG, Montenegro-Montero A, Drewe P, Najafabadi HS, Lambert SA, Mann I, Cook K, Zheng H, Goity A, van Bakel H, Lozano JC, Galli M, Lewsey MG, Huang E, Mukherjee T, Chen X, Reece-Hoyes JS, Govindarajan S, Shaulsky G, Walhout AJM, Bouget FY, Ratsch G, Larrondo LF, Ecker JR, Hughes TR (2014) Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158(6):1431–1443
Grant CE, Bailey TL, Noble WS (2011) Fimo: scanning for occurrences of a given motif. Bioinformatics 27(7):1017–1018
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, Nusbaum C, Myers RM, Brown M, Li W, Liu XS (2008) Model-based analysis of ChiP-Seq (MACS). Genome Biol 9:R137
Gusmao EG, Allhoff M, Zenke M, Costa IG (2016) Analysis of computational footprinting methods for DNase sequencing experiments. Nat Methods 13(4):303–309
Ritz A, Poirel CL, Tegge AN, Sharp N, Simmons K, Powell A, Kale SD, Murali TM (2016) Pathways on demand: automated reconstruction of human signaling networks. npj Syst Biol Appl 2:16002+
Tuncbag N, Gosline SJC, Kedaigle A, Soltis AR, Gitter A, Fraenkel E (2016) Network-based interpretation of diverse high-throughput datasets through the omics integrator software package. PLOS Comput Biol 12(4):e1004879+
Almozlino Y, Atias N, Silverbush D, Sharan R (2017) Anat 2.0: reconstructing functional protein subnetworks. BMC Bioinf 18:495
Gurobi Optimization, Inc (2016) Gurobi optimizer reference manual
Davis J, Goadrich M (2006) The relationship between Precision-Recall and ROC curves. In: Proceedings of the 23rd international conference on machine learning (ICML 2006), ICML ’06. ACM, New York, NY, pp 233–240
Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K (2017) KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45:D353–D361
Liberzon A, Subramanian A, Pinchback R, Thorvaldsdóttir H, Tamayo P, Mesirov JP (2011) Molecular signatures database (MSigDB) 3.0. Bioinformatics (Oxford, England) 27:1739–1740
Noble WS (2009) How does multiple testing correction work? Nat Biotechnol 27:1135–1137
Marbach D, Roy S, Ay F, Meyer PE, Candeias R, Kahveci T, Bristow CA, Kellis M (2012) Predictive regulatory models in drosophila melanogaster by integrative inference of transcriptional networks. Genome Res 22(7):1334–1349
Bonnet E, Calzone L, Michoel T (2015) Integrative multi-omics module network inference with Lemon-Tree. PLoS Comput Biol 11:e1003983
Chen S, Witten DM, Shojaie A (2014) Selection and estimation for mixed graphical models. Biometrika https://doi.org/10.1093/biomet/asu051
Žitnik M, Zupan B (2015) Gene network inference by fusing data from diverse distributions. Bioinformatics (Oxford, England) 31:i230–i239
Acknowledgements
This work is supported in part by US Environmental Protection Agency grant 83573701, NIH NIGMS grant 1R01GM117339, and NSF CAREER award to Sushmita Roy.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Siahpirani, A.F., Chasman, D., Roy, S. (2019). Integrative Approaches for Inference of Genome-Scale Gene Regulatory Networks. In: Sanguinetti, G., Huynh-Thu, V. (eds) Gene Regulatory Networks. Methods in Molecular Biology, vol 1883. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8882-2_7
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8882-2_7
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8881-5
Online ISBN: 978-1-4939-8882-2
eBook Packages: Springer Protocols