Abstract
In the era of personalized medicine and targeted therapies for the management of patients with cancer, ultrasensitive detection methods for tumor genotyping, such as next-generation sequencing or droplet digital polymerase chain reaction (ddPCR), play a significant role. In the search for less invasive strategies for diagnosis, prognosis and disease monitoring, the number of publications regarding liquid biopsy approaches using ddPCR has increased substantially in recent years. There is a long list of malignancies in which ddPCR provides a reliable and accurate tool for detection of nucleic acid-based markers derived from cell-free DNA, cell-free RNA, circulating tumor cells, extracellular vesicles or exosomes when isolated from whole blood, plasma and serum, helping to anticipate tumor relapse or unveil intratumor heterogeneity and clonal evolution in response to treatment. This updated review describes recent developments in ddPCR platforms and provides a general overview about the major applications of liquid biopsy in blood, including its utility for molecular response and minimal residual disease monitoring in hematological malignancies or the therapeutic management of patients with colorectal or lung cancer, particularly for the selection and monitoring of treatment with tyrosine kinase inhibitors. Although plasma is the main source of genetic material for tumor genomic profiling, liquid biopsy by ddPCR is being investigated in a wide variety of biologic fluids, such as cerebrospinal fluid, urine, stool, ocular fluids, sputum, saliva, bronchoalveolar lavage, pleural effusion, mucin, peritoneal fluid, fine needle aspirate, bile or pancreatic juice. The present review focuses on these “alternative” sources of genetic material and their analysis by ddPCR in different kinds of cancers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Liquid biopsy in blood is the major application of droplet digital polymerase chain reaction (ddPCR) technology in a wide variety of cancers for diagnostic, predictive, prognostic and monitoring purposes. |
The use of ddPCR for liquid biopsy has increased in recent years in other biologic fluids, including cerebrospinal fluid, urine, stool, ocular fluids, sputum, saliva, bronchoalveolar lavage, pleural effusion, mucin, peritoneal fluid, fine needle aspirate, bile or pancreatic juice. |
By far the most utilized ddPCR platform to date is the Bio-Rad QX100/200 system. However, new ddPCR platforms have been recently developed. |
1 Introduction
Droplet digital polymerase chain reaction (ddPCR) is a molecular biology technique based on sample partitioning into thousands of nanoliter-sized droplets where individual PCR reactions take place, allowing for the detection of very low abundance molecular targets with extremely high sensitivity [1]. Although this technology is nearly 10 years old [1, 2], its use has increased substantially in recent years, particularly in the field of precision oncology, with hundreds of publications demonstrating its clinical utility in many different kinds of malignancies. Liquid biopsy, defined as the analysis of molecular biomarkers in a wide variety of body fluids with diagnostic, predictive, prognostic or monitoring purposes, represents a noninvasive (or minimally invasive) approach with significant relevance in the management of patients with cancer that requires the implementation of extraordinarily accurate detection methods [3]. In this clinical scenario, ddPCR has gained much attention as a powerful tool for the detection of genetic alterations, including single nucleotide variants, copy number variations, genomic rearrangements and methylation biomarkers, mainly in blood (particularly in plasma and serum) but also in many other biologic fluids, such as cerebrospinal fluid, urine or stool, among others [4]. In this updated review, we describe some recent developments in ddPCR platforms and how they are being applied in oncology. Then, we focus our attention on the major applications of ddPCR in liquid biopsy, particularly in other “alternative” biofluids that are still less frequently used than blood but are gaining increasing interest in different types of cancers.
A literature research for this review was performed in PubMed using the following search strategy: ("Polymerase Chain Reaction"[Mesh:NoExp] OR "Multiplex Polymerase Chain Reaction"[Mesh] OR "PCR"[tw] OR "Polymerase Chain Reaction"[tw]) AND (("droplet based" AND "digital") OR "droplet based digital" OR "droplet digital" OR "bio-rad"[tw] OR "biorad"[tw] OR "raindance"[tw] OR "stilla"[tw] OR "digital droplet") AND (cancer[sb])AND "2016/12/01"[Date - Publication] : "3000"[Date - Publication].
2 Recent Developments in Droplet Digital Polymerase Chain Reaction Platforms
By far the most widely used ddPCR platform in the literature is the Bio-Rad platform (Bio-Rad; Hercules, CA, USA) (Fig. 1). This water-in-oil emulsion system for droplet generation has evolved from a manual workflow—where the user had to pipette the PCR reaction mix and the oil into the cartridges—to a more automated system, with the so-called AutoDG droplet generator, along with a change from the QX100 to the QX200 system. Other new ddPCR platforms have been developed in the last few years, such as the Naica Crystal ddPCR (Stilla Technologies; Villejuif, France) or the SAGAsafe® technology (formerly known as IBSAFE®; SAGA Diagnostics, Lund, Sweden). The main advantage of these newly developed systems is the increased multiplexing capabilities and improved sensitivity, with a lower limit of detection (LoD), respectively.
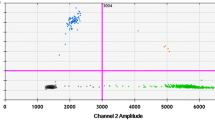
BioRad droplet digital polymerase chain reaction (ddPCR) platform. (1) Preparation of ddPCR reaction mix containing sample, probes and ddPCR master mix; (2) generation of droplets in the droplet generator by water-in-oil emulsion using vendor-specific oil; (3) droplets containing sample and ddPCR reaction mix; (4) transfer of droplets to a 96-well PCR plate; (5) the plate is run with PCR protocol in a ddPCR thermocycler; (6) droplet fluorescence is checked into the droplet reader; (7) analysis of results using QuantaSoft. Positive and negative droplets are plotted in a two-dimensional graph, setting thresholds for discrimination. Created with BioRender.com
The Naica System is based on a principle initially published in 2013 by Dangla et al. [5]. This digital PCR platform relies on a hybrid approach (named crystal digital PCR [cdPCR]) that combines a two-dimensional array of microchambers for partitioning and the use of crystal droplets that are thermocycled and transferred to a fluorescence microscope to detect amplification. All these steps take place in a specifically designed microfluidic chip (the Sapphire chip, containing the preloaded emulsion oil, which represents an advantage because it simplifies the process and prevents contamination [6]), and two different instruments are involved: the Naica Geode for sample partitioning and thermalcycling and a three-color detection system consisting of an automated fluorescence microscope, the Naica Prism3. Results are analyzed using the Crystal Miner software (Stilla Technologies). Madic et al. [7] reported a detailed and comprehensive description of the system and its workflow. This paper also discussed issues related to data analysis and the application of this system to the detection of L858R, L861Q and T790M epidermal growth factor receptor (EGFR) mutations. Thus, the capability of three-color multiplexing of this platform was tested, showing that it did not result in a loss of sensitivity. When compared with massive parallel sequencing (MPS), the cdPCR showed better performance for the detection of known mutations in the plasma of patients with metastatic non-small-cell lung cancer (NSCLC) [8]. In this comparison, the authors found 11 positive plasma samples with cdPCR that could not be detected with MPS, with mutant allele fractions between 0.09 and 7.9%. In longitudinal plasma samples collected for monitoring EGFR along the disease trajectory, MPS also reported six negative samples that digital PCR found to be positive.
In 2018, a customized six-color assay was developed for detecting and quantifying 19 prevalent EGFR sensitizing and resistance mutations using the Naica cdPCR and an inverted Nikon eclipse TI microscope (Nikon Instruments Europe, France) with an appropriate selection of filter sets for reading fluorescence in six different detection channels [9]. The LoD varied, depending on the mutation assayed, between 0.125% (p.C797S c.2389 T > A and p.C797S c.2390 G > C) and 0.0975% (exon 19 ins/del), with 0.25% for EGFR resistance mutation p.T790M and a range of 0.125–0.25% for activating mutations in the Cy5 detection channel. Sensitivity was also tested in tumor and plasma samples from 82 patients, comparing results with those from next-generation sequencing (NGS) and the three-color system. These comparisons showed good correlation, especially for the concentration of mutant copies per microliter and mutant allele frequency (MAF) in plasma samples measured by six- and three-color digital PCR. Longitudinal samples from four patients were also analyzed to monitor the course of disease, and fluctuations in mutant DNA levels were consistent with clinical evolution. This six-color assay for EGFR mutation detection was subsequently optimized on a prototype of a six-color reader instrument and a prototype of six-dimensional Crystal Miner software that are integrated in the new six-color cdPCR prototype platform, with the commercial version due for launch in late 2021 [10].
Song et al. [11] also recently developed an integrated digital PCR assay using the three-color version of the Stilla platform. The assay is called dEGFR39 and is the first that allows the screening and monitoring in plasma of all the EGFR mutations known to be clinically relevant in NSCLC for treatment guidance and prognosis. It simultaneously detects 39 mutations of exons 18–21 of this gene, including not only the most frequently identified variants such as L858R, 19Del and T790M but also other less common mutations including L861Q, S768I, G719X, C797S and 20 insertions [11]. This study analyzed the formalin-fixed paraffin-embedded (FFPE) tumor tissue and plasma of patients with NSCLC (N = 30 and N = 33, respectively) and demonstrated that dEGFR39 could detect EGFR mutations with a sensitivity of 0.308 copies/μL and an accuracy of 88.87% (for dEGFR39 in plasma and amplification-refractory mutation system [ARMS] in FFPE), showing a direct association between mutational load and response to treatment. It also anticipated disease progression by detecting T790M mutations earlier than other methods such as SuperARMS PCR and computed tomography (CT) imaging.
It is evident that the main application of cdPCR to date has been detection of EGFR alterations in NSCLC, but research is also ongoing for detection of PIK3CA mutations in plasma in patients with advanced breast cancer, aimed at the selection of alpelisib treatment. As presented in the European Society for Medical Oncology Breast Cancer virtual meeting in May 2020, the three-color detection system of Stilla was used with a multiplex assay for detection of 26 mutations located in exons 4, 7, 9 and 20 of the PIK3CA gene [12]. Quantification of BRAF V600E has also been performed using a novel DNA reference material that is intended to mimic circulating tumor DNA (ctDNA). Low levels of BRAF V600E ctDNA reference material were tested in eight different laboratories, seven of them with the QX200 ddPCR platform from Bio-Rad and one with the Stilla Naica Crystal ddPCR system [13]. Results from the interlaboratory study showed a significant difference in mutant and wild-type copy number concentration between the only laboratory using the Naica platform and the other seven laboratories. This inconsistency between the two platforms was improved considerably by correcting the droplet volume (the droplet volume measured by the authors of this study was 0.476 ± 0.008 nL vs. the 0.44 nL estimated by the manufacturer; this difference of 8.3% was suspected to be the reason for the overestimation observed in both mutant and wild-type copies).
Finally, the use of this platform has also been reported for chimerism monitoring of post-allogeneic hematopoietic stem cell transplantation for the treatment of hematological malignancies, including acute myeloid leukemia (AML) and acute lymphoblastic leukemia [14]. In this study, cdPCR was compared with NGS, with both methods reaching a sensitivity of 0.1%. The results in terms of percentage of chimerism in cdPCR and NGS showed a high concordance with those obtained by the reference techniques (ddPCR, short tandem repeat and quantitative PCR [qPCR]).
The SAGAsafe® platform is a droplet-based proprietary methodology with an improved LoD of ~ 0.001% MAF. It consists of a two-step process that takes place sequentially within the droplets: linear amplification or copying of the target sequence followed by a limited exponential signal generation. This technology minimizes polymerase base-incorporation errors, enhancing true-positive signals while simultaneously reducing the number of false positives, thereby achieving greater sensitivity and specificity [15, 16].
It has already been used to detect TP53 mutations in liquid-based Pap samples from patients with ovarian cancer [17]. In this study, IBSAFE detected TP53 mutations using a custom-designed assay with very high sensitivity (MAF of 0.0068%) in samples with low DNA input (as little as 0.17 ng). The in-sample LoD was reported to be 1 in 50,000. Bio-Rad ddPCR was used as a control, but the small number of samples tested did not allow a direct comparison between the platforms.
Another study reported that IBSAFE detected somatic mutations in plasma ctDNA from patients with breast cancer, showing a small average increase in ctDNA levels in both peripheral and central blood following mammographic breast compression, with no apparent clinical relevance [18]. Additionally, IBSAFE technology has been applied to the detection of EGFR, KRAS and BRAF mutations in the preoperative plasma of patients with lung adenocarcinoma [15]. Finally, the most recent work using this platform demonstrated its applicability for the detection of minimal residual disease in AML [16]. Between five and nine mutation assays were developed for each patient using this technology. The method was more sensitive in identifying residual disease than multicolor flow cytometry, detecting the targeted mutations in all relapsing patients and allowing the tracking of early recurrence in leukemic subclones, revealing the existence of three different mutational patterns of relapse.
SAGA Diagnostics launched its first in vitro diagnostic (IVD) European Conformity (CE)-marked SAGAsafe® kit for EGFR T790M testing in 2020. The company also presented a combined strategy of NGS of tissue to detect chromosomal rearrangements and a digital PCR fingerprint in plasma, called SAGAsign® (formerly known as KROMA) [19]. Two recent references in the literature to another platform should also be mentioned: the MicroDrop-100 ddPCR system (Forevergen, China) is based on water‐emulsion droplet technology and has been used to detect BRAF V600E mutations in thyroid nodules, with better performance than ARMS-PCR [20], and also to measure the effect of CSNK2A3 expression on hepatitis B virus infection in hepatocarcinoma cells in vitro [21].
Studies reporting clinical applications of these emerging ddPCR platforms in cancer are summarized in Table 1.
3 Applications of ddPCR in Liquid Biopsy
3.1 Liquid Biopsy in Blood
Hundreds of papers published in the last 4 years have supported the clinical utility of analyzing genetic biomarkers using ddPCR in blood in the field of oncology. It is being tested for application in the monitoring of molecular response and minimal residual disease in hematologic malignancies [22,23,24,25,26,27,28]. In many cases, both blood and bone marrow aspirates are used for liquid biopsy, and BCR-ABL is frequently the biomarker of choice [26, 28,29,30,31,32,33,34], and this has also been suggested as a useful tool to predict and assess the outcomes after discontinuation of treatment with tyrosine kinase inhibitors (TKIs) [32, 35, 36]. The QXDx BCR-ABL %IS (Bio-Rad) ddPCR assay is the first commercially available ddPCR-based IVD product with US FDA clearance and the CE mark. This assay can detect the e13a2 and e14a2 fusion transcripts (but not e1a2, e19a2 or other rare transcripts) and has an acceptable analytical performance, with results comparable to those of the CE-IVD-marked ipsogen BCR-ABL1 Mbcr IS-MMR (Qiagen, Hilden, Germany) real-time qPCR (RT-qPCR) assay [37]. The main advantages of ddPCR versus the gold standard RT-qPCR method include a superior sensitivity and accuracy for ddPCR (with a LoD of one copy of transcript), as well as the ability to perform an absolute quantification without standard curves. Disadvantages include the lack of standardized methods and its limited availability in laboratories [38]. Longer turnaround times (due to droplet generation and reading) and the possibility of false positives have also been suggested as potential limitations of this technique [37]. We found no consensus in the literature regarding cost and throughput concerns. Alu methylation status has also been quantified by ddPCR in bone marrow samples from patients with chronic lymphocytic leukemia, myelodysplastic syndromes and myelomonocytic leukemia before and after treatment with 5-azacytidine. Decreased levels of this epigenetic marker have been observed after hypomethylating therapy, suggesting a potential utility of this approach for molecular monitoring of response [39]. Other biomarkers include NPM1 [22] and IDH2 [40] mutations, WT1 levels [25] and immunoglobulin heavy chain (IGH) gene [41] or immunoglobulin kappa-deleting-element (IGK‐Kde) [42] rearrangements. Table 2 summarizes some data on these studies.
As previously mentioned, this technology has also been utilized to quantify engraftment after hematopoietic stem cell transplantation [14, 43,44,45,46], to monitor the expansion and persistence of chimeric antigen receptor (CAR)-T cells in vivo, reflecting response rates and side effects [47,48,49,50,51,52,53], and to detect vector copy number in clinical CAR/T-cell receptor (TCR)-T-cell products [54, 55].
Genetic biomarkers have been analyzed from different sources, including circulating tumor cells (CTCs) [56,57,58], cell-free DNA (cfDNA) [59,60,61] and cell-free RNA or extracellular RNA [62], nucleic acids derived from exosomes [63,64,65,66,67,68] and extracellular vesicles (EVs) [69,70,71], including long noncoding RNAs [72,73,74,75], microRNAs (miRNA) [76,77,78,79,80] and messenger RNA (mRNA) [57, 81,82,83], isolated from whole blood, plasma or serum (Fig. 2). Many studies have analyzed a combination of several of these genetic materials [84,85,86].
Different sources of genetic biomarkers isolated from whole blood, plasma or serum that can be analyzed by droplet digital polymerase chain reaction (ddPCR): circulating tumor cells (CTCs), extracellular RNA (exRNA), cell-free DNA (cfDNA), cell-free RNA (cfRNA), long non-coding RNA (lncRNA) and messenger RNA (mRNA). Created with BioRender.com
ddPCR has been applied in a long list of cancer types, headed by colorectal [60, 87,88,89,90,91,92,93,94,95,96,97,98] and lung cancer, particularly NSCLC, with EGFR and KRAS mutations being the most analyzed markers because of their relevance for therapeutic management of patients, particularly for the selection and monitoring of treatment with TKIs [61, 99,100,101,102,103,104,105,106,107,108,109,110] (Table 3). These are without a doubt the most widespread clinical applications of ddPCR in the field of oncology. Pancreatic cancer [56, 59, 65, 76, 78, 111,112,113,114,115,116,117], breast cancer [82, 118,119,120,121,122,123,124], melanoma [58, 125,126,127,128,129,130,131,132,133,134], prostate cancer [57, 81, 135,136,137,138,139,140,141,142,143] and ovarian cancer [144,145,146,147,148] are also among the most studied neoplasms using ddPCR for clinical purposes. Other less frequent malignancies such as gastrointestinal stromal tumors [149, 150] or peritoneal metastasis from colorectal origin [151, 152] have also benefited from ctDNA profiling in plasma by ddPCR. It is not the aim of this review to cover all the different applications of ddPCR in every cancer type. Notably, in most of these studies, fluctuations in ctDNA levels detected by ddPCR mirrored disease response and predicted recurrence before clinical evidence. Circulating tumor markers detected by ddPCR have also shown a clinical value for early detection of disease and had prognostic implications in many of these studies.
In many studies, ddPCR was applied in combination with NGS assays, which provide a broader perspective of the tumor mutational landscape, whereas ddPCR focuses on single molecular targets that allow confirmation of the presence of these variants and detection of changes in ctDNA levels over the disease course to track tumor burden [23, 106, 151, 153,154,155,156,157]. This strategy gives the clinician the opportunity to monitor the response to treatment and guide therapeutic decisions, anticipating relapse even months ahead of the emergence of clinical symptoms or evidence in imaging techniques. A remarkable number of studies have demonstrated the utility of ddPCR to unveil intratumoral heterogeneity [158] and clonal evolution in response to treatment [61, 150]. Of note, ddPCR has also proven useful for the detection of nonmalignant mutations present in hematopoietic cells (a phenomenon called clonal hematopoiesis) that can give rise to a confounding false-positive plasma result when non-hematopoietic cancers, such as NSCLC, are monitored using liquid biopsy [159]. In recent years, a trend towards multianalyte or multifactorial models has increased, where several biologic markers are simultaneously analyzed, with ddPCR playing a significant role as an accurate and reliable tool for quantitative nucleic acid-based biomarkers [160,161,162].
ddPCR is usually employed as a validation technique when alternative or newly developed methodologies are tested or to solve discordant cases [163–167].
It should also be noted that there are still some challenges and/or limitations to a more widespread use of ddPCR in routine clinical practice. The number of genetic variants that can be analyzed is limited by the amount of sample available. Multiplexing strategies have been developed to overcome this problem, including additional detection channels in the newer ddPCR platforms, as mentioned earlier. Besides, more than one target can also be detected in a single fluorescent channel by varying the concentration of different probes labeled with the same fluorophore or using amplicons of different sizes marked with DNA binding dyes [168].
Another relevant hurdle is the occurrence of false-positive signals, mainly caused by PCR errors in cfDNA samples, where a rare variant is intended to be detected in a high background of a wild-type allele, limiting the fractional abundance that can be detected [168].
First, standardized protocols for sample collection, storage, processing, nucleic acid extraction and modification are lacking. A range of tubes are used for blood collection, along with a range of anticoagulants and/or conservative compounds (EDTA, Streck, CellSave). The time from collection to processing and the temperature and storage conditions also deserve consideration. A huge diversity of protocols is found in the literature, from one-step to two-step centrifugation, with variable speeds, times and temperatures. The starting volume of plasma or serum for extraction varied from < 1 to ≥ 5 mL. Commercial kits specifically developed for circulating nucleic acid isolation are the most frequent choice, with protocols described by the manufacturers, but elution volume or the method of nucleic acid quantification, for example, usually differ between studies. It is not the aim of this review to delve into isolation methods, but heterogeneity is high and, remarkably, many of these preanalytical steps are of the utmost importance and can lead to measurement errors. ddPCR results are severely affected by factors such as DNA purity or concentration, hence all these procedures still require further optimization.
Other associated challenges refer to data analysis, mainly thresholding setting, and particularly when “rain” occurs (the presence of partitions located between positive and negative populations). Several software tools (both platform specific and independent) have been developed in recent years in response to this problem. The Minimum Information for Publication of Digital PCR Experiments (dMIQE) guidelines were published in 2013 [169] and an update presented in 2020 (dMIQE2020) [168]. These guidelines highlight the key experimental information that should be provided by researchers and helps in understanding every step of the experimental process, from assay design to validation and performance, why they require standardization, and how this could be achieved.
3.2 Other Biological Fluids
The vast majority of studies using ddPCR involve the detection of genetic alterations in plasma. However, as previously reviewed [4], other body fluids can also be used for noninvasive or less invasive molecular examination, including cerebrospinal fluid (CSF), urine or stool. Recent studies have investigated the use of additional body fluids for diagnostic purposes, including saliva and ocular fluids, such as vitreous fluid (VF) and aqueous humor (AH). Of note, the use of these biofluids is not yet widespread in the clinic, and many of these approaches are still under development in research studies. Also, the different targets analyzed in these alternative sources of nucleic acids requires the adjustment of isolation procedures. Limiting factors include the amount of sample collected (e.g., saliva or sputum) or the concentration and/or purity obtained (e.g., in stool).
3.2.1 Cerebrospinal Fluid
IDH1 mutations were among the first genetic alterations detected in CSF with ddPCR, and further research has been undertaken, particularly in lower grade gliomas [170]. However, MYD88 L265P (a myeloid differentiation primary response gene 88 single-base substitution at c.794T > C resulting in a leucine to proline amino acid change) is another hotspot mutation that has recently gained much more attention, with a number of publications accumulating evidence about its clinical value. The major application of MYD88 L265P detection by ddPCR in CSF is for minimally invasive confirmation of a diagnosis of primary central nervous system (CNS) lymphoma (PCNSL) [171,172,173] (a case of secondary CNS lymphoma has also been reported [174]) and of Bing–Neel syndrome [171, 175]. Interestingly, in some cases, this mutation has been detected in cfDNA from CSF at a higher frequency than in cellular DNA [172], and this might provide more sensitivity than cytomorphology and NGS in samples with low cellularity and very low DNA content [171]. In a very recent study, CSF testing by ddPCR for MYD88 L265P proved useful for the detection of early relapse in a patient with Bing–Neel syndrome treated with ibrutinib, showing an increase in mutation levels 2 weeks before the appearance of clinical symptoms and without evidence of recurrence on magnetic resonance imaging (MRI), CSF cytology, flow cytometry analysis and immunofixation electrophoresis [175]. In line with these observations, Bobillo et al. [176] recently combined NGS and ddPCR (covering many tumor mutations apart from the aforementioned MYD88 L265P) and reported better detection of CNS lesions by measuring ctDNA in CSF than flow cytometry, cytology and plasma measurements. This work showed the great potential of CSF-based liquid biopsy using ddPCR to monitor CNS involvement of B-cell lymphomas, predicting CNS relapse and detecting residual disease [176].
However, a previous study including three patients with spinal ependymoma suggested that anatomic sequestration or a low grade of intramedullary spinal cord tumors might hamper the utility of CSF-based liquid biopsy in these malignancies [177].
In recent years, liquid biopsy by ddPCR has gained increasing interest as a powerful tool to detect genetic biomarkers in pediatric brain tumors, particularly in pediatric diffuse midline glioma (DMG), as previously reviewed by Lu et al. [178] and Azad et al. [179]. A recurrent actionable mutation in histone 3, affecting either the H3.1 or the H3.3 protein at lysine position 27 (HIST1H3B K27M and H3F3A K27M, respectively, also known as H3K27M mutation), has been detected using ddPCR assays in the CSF of patients with diffuse intrinsic pontine glioma (DIPG) or high-grade glioma [180–183]. In a study by Panditharatna et al. [181], H3K27M and other obligate partner mutations in ACVR1, PIK3R1 and BRAF genes were detected by ddPCR in the CSF of patients with DMG. Their results also suggested that fluctuations in ctDNA levels in serial plasma samples might have clinical utility to monitor treatment response in patients with DIPG, comparable with MRI evaluation [181]. A recent study also showed the potential of ddPCR determinations in CSF for disease monitoring in pediatric patients with medulloblastoma. Again, a combined strategy of NGS and ddPCR allowed the detection of a wide variety of tumor mutations in CSF, highlighting that it both represents a better source of ctDNA than plasma and has superior sensitivity compared with cytology [184]. Thus, mutations identified by NGS and validated by ddPCR efficiently detected minimal residual disease and shed light on genomic tumor evolution, revealing intratumor and interlesion heterogeneity since this approach was able to unveil the existence of two completely different tumors at diagnosis and relapse.
Apart from tumors directly affecting the CNS as a primary target, the analysis of genetic alterations in CSF has also been applied for detection of metastatic disease in the CNS from other tumor origins, such as leptomeningeal or brain metastasis derived from lung adenocarcinoma [185,186,187], breast cancer [188] or melanoma [189].
3.2.2 Urine
Several publications have reported the application of ddPCR to the assessment of molecular biomarkers in urine for prostate and bladder cancer. The detection of methylation biomarkers (GSTP1, APC, RASSF1A, PITX2 and C1orf114) in the cell fraction isolated from urine using a filtration device [190] or the development of a two-gene panel (PCA3, PCGEM1) in urinary exosomal mRNA [191] are some of the strategies that have been explored to improve the noninvasive identification of high-grade prostate cancer. The urinary transcriptome was proposed as a valuable source of biomarkers and later validated by ddPCR in a recent study showing that five protein-coding genes (FTH1, BRPF1, OSBP, PHC3 and UACA) distinguished patients with prostate cancer from cases of benign hyperplasia and healthy subjects, both in the centrifuged and non-centrifuged fraction of small-volume urine samples (1 mL) [192]. Similarly, another novel urinary mRNA signature (including three upregulated genes [PDLIM5, GDF-15 and THBS4] and three downregulated genes [UPK1A, SSTR3 and NPFFR2]) was developed using ddPCR to discriminate prostate cancer from benign prostatic hyperplasia within the “prostate-specific antigen [PSA] gray zone” (3–10 ng/mL total PSA) [193]. The expression of the androgen-receptor splice variant 7 (AR-V7) has also been reliably quantified in urine-derived EVs, with higher levels in patients with castration-resistant prostate cancer than in those with hormone-sensitive tumors [194].
On the other hand, the analysis of hotspot mutations in TERT promoter and FGFR3 by ddPCR in urine has been proven useful for the early detection of urothelial cancer, including upper tract urothelial cell carcinoma and bladder cancer [195,196,197,198,199,200]. Tumor-specific mutations in FGFR3 and PIK3CA hotspot mutations, among many others, have been measured in the plasma and urine of patients with bladder cancer for disease and treatment monitoring, showing a remarkable potential to detect early signs of metastasis [201, 202]. The combination of ddPCR and urine cytology yields a higher sensitivity than cytology alone (UroVysion) for detection and prognosis in urothelial bladder cancer [198]. Specific ddPCR assays for TERT promoter mutations have shown results comparable to those with the UroMuTERT NGS-based assay for detection of MAFs > 2%, both in the urine supernatant cfDNA and the urine pellet cellular DNA, although some discrepancies have been found below this allelic fraction [200]. Previous studies suggested that ddPCR may have a limited sensitivity to detect low-grade tumors harboring very low MAFs [195].
PIK3CA p.H1047R mutation has been detected by ddPCR in the urine of a patient with CLOVES (congenital lipomatous overgrowth with vascular epidermal and skeletal anomalies) syndrome, a subgroup of the PIK3CA-related overgrowth spectrum (PROS), who had a Wilms tumor. The mutation was present not only in the affected tissue but also in urine and in the Wilms tumor, which had been resected upon diagnosis 26 months prior to the urine sample collection. These results suggest that urine may be useful for noninvasive mutation screening by ddPCR in patients with CLOVES syndrome, some of whom are candidates to develop Wilms tumor [203]. This suggestion was supported by another study involving patients with PROS that analyzed several kinds of biologic specimens, including plasma, whole blood, saliva, buccal swabs and urine (at its cellular and cfDNA fraction) [204]. Three different PIK3CA variants [c.3140A > G; p.(His1047Arg), c.3140A > T; p.(His1047Leu) or c.1624G > A; p.(Glu542Lys)] were detected in this wide variety of tissues, with the exception of leukocytes (only one case was detected in saliva and the corresponding buccal swab from a clinically affected cheek). Interestingly, patients who had a positive variant of PIK3CA in DNA extracted from the cellular fraction of urine also presented this variant in urine cfDNA, and these levels were much higher in patients with a history of nephroblastomatosis or Wilms tumor than in individuals without known renal involvement. Thus, urine testing by ddPCR could provide information about the renal involvement in PROS, and multiple tissue analysis may help identify patients at high risk for Wilms tumor.
The usefulness of urine as a suitable source of genetic material for molecular analysis in cancers not related to the genitourinary tract was also further explored in pancreatic ductal adenocarcinoma (PDAC) [205], metastatic colorectal cancer (CRC) [206] and lung cancer, particularly NSCLC [207]. KRAS mutations have been examined in the plasma and urine of patients with PDAC, showing a similar detection rate and sensitivity in both fluids, although they are influenced by renal function degeneration [205]. ddPCR has also found KRAS and BRAF mutations in matched plasma and urine samples from patients with metastatic CRC [206]. EGFR mutations have been measured in the urine of patients with NSCLC by ddPCR at different time points after curative intent surgery for disease monitoring, with the aim of detecting relapse and minimal residual disease [207]. Thus, in this study, the presence of detectable mutant DNA in the urine samples post-treatment was associated with disease recurrence, whereas patients with undetectable levels had better disease-free survival. In another study, matched plasma and urine samples from patients with NSCLC were collected and analyzed for EGFR mutation detection by ddPCR after TKI therapy [208], demonstrating that both body fluids may provide complementary information. Baseline plasma values showed better positive predictive value, whereas urine samples seemed to be more useful for serial monitoring since changes in secondary EGFR T790M mutation levels were detectable earlier. It was concluded from both types of samples that patients with higher post-treatment values than at baseline had poorer outcomes (the majority were T790M positive), but urinary cfDNA performed better at identifying patients with potentially worse outcomes.
3.2.3 Stool
In 2017, Herring et al. [209] published the detection of ITGA6 and ITGA6A transcripts (integrin α6 subunit and its α6A variant) by ddPCR in stool samples obtained from patients with CRC. Patients with colorectal lesions showed statistically significantly elevated levels of ITGA6 transcript in stools with respect to the non-pathological controls (approximately eight times higher for adenomas and 6–11 times higher for CRC, being particularly higher in more advanced stages). Meanwhile, a greater than 40-fold elevation of ITGA6A was found in stools of patients with stage II and III CRC with respect to controls. This study directly compared ddPCR and conventional qPCR, with results being similar in terms of sensitivity and specificity.
Stool-derived DNA has also been analyzed by ddPCR for the presence of KRAS G12D mutation in patients with CRC who presented this hotspot mutation in their tumor tissues, being detectable in 80% of stool samples [210]. More recently, hypermethylation in GRIA4- and VIPR2-associated CpG islands was detected by ddPCR in stool samples from patients with CRC, highlighting their potential as early noninvasive biomarkers for diagnosis of this neoplasia [211]. In this study, ddPCR was compared with Methylight qPCR using the same primers and probes in both assays for detection of the two stool-based methylation biomarkers, demonstrating that the sensitivity of ddPCR was superior.
Apart from alterations in cancer-related genes, another possibility explored using ddPCR in stool samples was the detection of DNA from different strains of bacteria that have been associated with malignancy, such as Fusobacterium nucleatum in CRC [212] and Helicobacter pylori in gastric cancer [213, 214]. In a study performed in a Japanese population, F. nucleatum was significantly elevated in the non-advanced adenoma group, the advanced adenoma/carcinoma in situ group and the CRC group compared with the control group of healthy subjects, suggesting that this ddPCR-based assay could be useful for detecting individuals with CRC [212]. Similarly, H. pylori DNA was detected by ddPCR in patients with gastric cancer in a Chinese population by measuring the copy number of the H. pylori 16S ribosomal RNA (rRNA) gene. These authors found levels six times higher in stool from patients with gastric cancer than in those from healthy controls, in contrast to gastric loads, which were comparable between both groups. Additional cagA detection and cagA EPIYA typing ddPCR assays developed by the same research group [213] were also tested in the stool samples. In this population with a high prevalence of the cagA virulence gene, the East Asian allele was suggested as a risk marker for gastric cancer [214]. Interestingly, stool-based detection of H. pylori clarithromycin resistance-associated genotypes through an assay targeting 23S rRNA mutant alleles (A2143G, A2142G and A2142C) also proved feasible in patients with and without gastric cancer, particularly in cases of heteroresistance, where it seemed to be more sensitive than commonly used methods for testing in routine clinical practice [215].
Along this line, altered microbiota was detected by 16S rRNA analysis using ddPCR in postoperative fecal samples from patients subjected to pancreaticoduodenectomy (with a cancer diagnosis confirmed by surgical pathology in 45 of 50 cases) [216]. A depletion of strict anaerobes and an expansion of some Proteobacteria, with an enrichment in Bacteroides and Klebsiella, was observed in fecal, pancreatic fluid, bile and jejunal samples, deviating from the microbial patterns considered normal in healthy individuals. These results suggest that postoperative fecal microbiota may have a potential predictive value to identify patients at high risk for pancreatic cancer, but this possibility needs to be further explored.
3.2.4 Ocular Fluids
Hiemcke-Jiwa et al. [217] proved that detection of MYD88 L265P mutation in these fluids by ddPCR is feasible and represents a reliable tool for the diagnosis of vitreoretinal lymphoma (VRL) and for treatment monitoring. This hotspot mutation was present in 74% of patients in this study and is a distinguishing mark of VRL. Patients with uveitis were included as a negative control group. The analysis of paired samples from patients with VRL revealed that sensitivity was 75% in VF versus 67% in AH, with positive predictive values and specificities of 100% in both cases. Indeed, ddPCR allowed the detection of MYD88 L265P on cfDNA even in > 100-fold diluted VF samples. Interestingly, the mutation became undetectable in any ocular fluid after intravitreal and systemic treatment [217]. In another recent study, MYD88 L265P mutation (which was present in 75% of patients with VRL, an incidence similar to that in the aforementioned work by Hiemcke-Jiwa et al. [217]) was detected by ddPCR in the VF of patients with diffuse large B-cell lymphomas and in one patient with lymphoplasmacytoid lymphoma [218].
AH can be obtained by paracentesis and is considered a safer and less invasive method of gathering DNA from tumor origin than collecting VF specimens. In particular, taking retinal biopsies by fine needle aspiration (FNA) incurs a high risk of complications, including infection, hemorrhage or retinal detachment [217]. However, the volumes of AH obtained are small and have low DNA content. Thus, a very sensitive technique for analysis is required. VRL diagnosis is extremely complicated and requires the combination of several laboratory tests, including flow cytometry (for detection of clonal B-cell populations), cytomorphology, immunohistochemistry and molecular analysis (determination of cytokine levels, immunoglobulin gene rearrangements and mutational analysis), because no single diagnostic test has sufficient sensitivity and specificity of detection. The use of ddPCR alone is not enough for an accurate diagnosis, but it provides an additional tool that could be integrated into the clinical routine. The analysis of MYD88 L265P mutation in both VF and AH in combination could also contribute to a better diagnosis [173, 217, 218]. Double-side vitrectomy and multisite sampling including CSF have also been demonstrated to improve ddPCR detection efficiency in PCNSL and other primary extranodal lymphomas [173].
3.2.5 Saliva
Recent studies have explored the utility of saliva as a source of DNA for study in different types of cancer. The levels of human papillomavirus (HPV) DNA were previously studied in the plasma of patients with advanced HPV-associated oropharyngeal cancer (OPC) using a ddPCR multiplex assay to detect the most common high-risk HPV subtypes: 16, 18, 31, 33 and 45 [219]. The same authors later hypothesized that viral DNA could also be shed by tumor cells in the oropharynx into the saliva, paving the way for the use of salivary secretions for diagnostic, prognostic and predictive purposes in disease monitoring [220]. To test their hypothesis, they designed an observational study to analyze paired plasma–saliva samples from patients with HPV-OPC. This study confirmed that HPV DNA is detectable in the saliva and correlates with tumor burden and local disease subsite. Interestingly, salivary HPV DNA viral load showed a strong correlation with tumor burden in patients with locoregional disease but not in those with distant disease only, in contrast to plasma, the levels of which were associated with tumor burden among the whole cohort, irrespective of disease site. Furthermore, HPV DNA baseline levels in saliva were almost 20 times higher in patients with clinical and imaging evidence of locoregional disease than in those with distant disease outside the head and neck only, showing a clinically valuable predictive potential. These levels were particularly elevated in those with base-of-tongue tumors compared with tonsil cases. Salivary HPV DNA levels fluctuated in close relationship with disease progression and response, and changes were observed prior to clinical detection in most cases. High HPV DNA levels in plasma, in turn, were associated with worse outcomes, indicating that saliva and plasma provide different and complementary information. The use of both bodily fluids simultaneously in a combined strategy increased the sensitivity of detection up to 100% in this study.
Salivary exosomes have been suggested as an alternative and enriched source of DNA and RNA in patients with HPV-OPC. An acoustofluidic biocompatible platform previously developed for plasma samples [221] was later optimized to isolate salivary exosomes at a high yield, irrespective of the variable viscosity and collection method of saliva samples [222]. This platform consists of a fusion of acoustics and microfluidics that uses standing surface acoustic waves, and ddPCR was used to evaluate the exosome yield obtained by this platform compared with other isolation methods. Thus, the concentration of the two small RNAs (miR-148-a and piR014923) measured by ddPCR was 15 times higher in the exosome fraction isolated by the optimized platform than in that isolated with the differential ultracentrifugation method. These authors also designed a ddPCR assay that could detect HPV16 DNA in 80% of patients with HPV-OPC in a small cohort. Interestingly, the concentration of the target DNA was 12 times higher in the exosome fraction than in microvesicles.
Saliva has also been interrogated for the presence of EGFR mutations in patients with NSCLC using ddPCR technology. Paired plasma and saliva samples showed a concordance of 83.78% and was correlated with clinical response. However, saliva cfDNA concentrations could not distinguish patients with NSCLC from controls (including healthy subjects and patients with pulmonary benign disease). This suggests it is a qualitative rather than a quantitative indicator and thus could not be applied for diagnostic purposes in this malignancy, but it could complement plasma and tissue biopsies [223]. A recent study also compared ddPCR and a novel technology called electric field-induced release and measurement (EFIRM) for the detection of EGFR mutations in paired plasma and saliva from patients with NSCLC. The sensitivity of EFIRM was 100% in both types of samples, whereas ddPCR showed sensitivities of 84.6% in plasma and 15.4% in saliva. This work by Li et al. [224] revealed that ctDNA was more fragmented in saliva than in plasma, and EGFR L858R was present mainly as ultrashort DNA fragments between 40 and 60 bp in size (known as ultrashort ctDNA) that the ddPCR assay was mostly unable to amplify. Of note, the EFIRM test results indicated that the concentration of mutant ctDNA in saliva was higher than in plasma, in stark contrast with the results of the previous study by Ding et al. [223, 224]. Interestingly, the majority of these EGFR mutant sequences were encapsulated within exosomes [224].
Finally, as previously mentioned, a PIK3CA variant was also detected in the saliva of a patients with PROS [204].
3.2.6 Sputum
In 2018, Su et al. [225] applied droplet digital methylation-specific PCR (ddMSP) to the detection of epigenetic biomarkers in sputum to develop a classifier for the early detection of lung cancer. A comparison of ddMSP and the conventional quantitative MSP (qMSP) showed that the former was more sensitive and provided more precise and reproducible results for methylation quantification. The authors built an epigenetic classifier using ddMSP that included four sputum methylation biomarkers (HOXA9, RASSF1A, SOX17 and TAC1) and demonstrated a sensitivity of 86.6% and a specificity of 90.6% for the detection of lung cancer. This method was superior to the clinical gold standard—sputum cytology—in accuracy and sensitivity, although both had a similar specificity. Additionally, the authors intentionally added inhibitors (sodium dodecyl sulfate and heparin) to the PCR reactions and observed that ddMSP better tolerated the presence of these inhibitory substances than did qMSP [225].
More recently, EGFR mutations were detected in the sputum of patients with NSCLC using ddPCR [226, 227]. Isaka et al. [226] reported that the detectability of EGFR mutations in sputum samples by ddPCR was highly sensitive in cases with positive sputum cytology and very low (3.1%) in patients with negative cytology. Based on these and previous observations, these authors suggested that sputum samples should be collected for EGFR mutation analysis in cases where CT tumor size is ≥ 29 mm, which is considered a potential predictive factor for positive sputum cytology [226].
On the other hand, Hackner et al. [227] analyzed paired plasma and sputum samples for the screening of activating and resistance EGFR mutations in patients with NSCLC, showing that the combination of both fluids for detection of T790M mutation increased diagnostic efficiency in patients with progressive disease compared with the single analysis of plasma. Again, conventional cytology was unable to detect tumor cells in any of the sputum specimens, although ctDNA was detectable [227].
3.2.7 Bronchoalveolar Lavage
Another body fluid that has been examined using ddPCR in the oncologic field is bronchoalveolar lavage (BAL) and, more precisely, the cell pellets obtained from this liquid biopsy. In 2017, ddPCR was used to validate an miRNA-based prediction model (including two miRNAs: miR-205-5p and 944) to distinguish squamous cell carcinoma from adenocarcinoma in BAL samples from patients with NSCLC, showing higher diagnostic accuracy than cytology [228]. Similarly, the determination of BRAF V600E mutation in BAL samples by ddPCR was proposed as a complementary tool for the diagnosis of pulmonary Langerhans cell histiocytosis [229].
More recently, a ddPCR method was developed to quantify the CpG methylation levels of TMPRSS4 and SHOX2 promoters in plasma and BAL samples from patients with NSCLC, showing that TMPRSS4 methylation status allows distinction between patients with early-stage NSCLC and healthy individuals. TMPRSS4 was hypomethylated in BAL and plasma of patients with early-stage disease compared with controls, and an inverse correlation was observed between TMPRSS4 and SHOX2 in patients with early-stage NSCLC versus healthy controls in both body fluids, although this correlation was only observed in BAL in all stages. These results support the potential of TMPRSS4 as noninvasive epigenetic biomarker as an indicator of malignancy in early-stage NSCLC [230]. In line with this, Roncarati et al. [231] recently described a four-gene methylation panel (RASSF1A, CDH1, DLC1 and PRPH) in bronchial washings using ddMSP that showed a remarkable diagnostic value in lung cancer, with a sensitivity of 97% and a specificity of 74%. ddPCR has also been used for the detection of EGFR-TKI-sensitizing mutations in bronchial washing fluid and plasma of patients with NSCLC, showing that the former has greater diagnostic efficacy than the latter [232].
3.2.8 Pleural Effusions/Pleural Fluid
Several publications have reported the detection of EGFR mutations in pleural effusions or pleural fluid samples of patients with NSCLC [233–236]. These mutations were analyzed in the cfDNA obtained from the supernatant of pleural fluid [233], in the cell pellet [236] or both [234, 235]. Hummelink et al. [235] demonstrated that KRAS and EGFR mutations were detectable in paired supernatant and cell pellet samples, from not only NSCLC but also colon carcinoma, appendiceal carcinoma and adenocarcinoma of unknown primary. This study suggested that the cell-free fraction of pleural effusions is an excellent source of genetic material for detection of both driver and resistance mutations and that the combined analysis of both fractions reached optimal sensitivity [235].
3.2.9 Mucin
ddPCR was recently applied to the detection of KRAS mutations in cfDNA from mucin obtained from patients with pseudomyxoma peritonei, a rare malignant disorder characterized by the accumulation of huge amounts of this viscous fluid in the abdominal cavity [237]. To date, the gold standard for routine diagnosis is the screening of mucin in search of tumor cells. However, this pilot study proved that acellular mucin contained cfDNA from tumor origin despite the absence of detectable tumor cells. Paired plasma samples from the same patients were also analyzed but were negative in all cases. These results are concordant with the localized nature of this malignancy and pave the way for the use of ddPCR in further studies aimed at elucidating the complex molecular mechanisms leading to recurrence in this acellular neoplasm [237].
3.2.10 Peritoneal Fluid/Ascites
As mentioned, in many cases, ddPCR represents a valuable tool for validation of NGS results from different kinds of biologic samples. The cellular fraction of second-look peritoneal washings from patients with high-grade serous ovarian, fallopian tube, or primary peritoneal cancer was analyzed using ddPCR to validate the results previously obtained by NGS [238]. Tumor-specific mutations (including the most frequent mutated gene TP53 but also PTEN and HNF1A, among others) were detectable in second-look washings, for both primary and recurrent tumors, providing valuable information about tumor heterogeneity and residual disease [238]. In line with this, a recent study employed ddPCR as a validation technique for quantification of miR-593-3p as a prognostic biomarker in pancreatic cancer undergoing staging laparoscopy [239]. Its expression was upregulated in the supernatant of peritoneal lavage fluids with positive cytology and correlated with worse overall survival and disease-free survival, even in cases with localized disease and negative cytology. These results suggest that elevated levels of miR-593-3p could be an indicator of the presence of subclinical intra-abdominal micrometastasis [239]. In agreement with these observations, another recent publication also reported that high levels of peritoneal lavage tumor DNA were associated with poorer outcomes in terms of disease-free survival and overall survival [240]. In this work by Suenaga et al. [240], ddPCR was employed to analyze peritoneal lavage from patients with PDAC in search of KRAS mutations. Interestingly, tumor-derived DNA was detectable not only in patients with positive cytology but also in 40% of the patients with a negative cytology result, showing a remarkably superior sensitivity for prediction of peritoneal recurrence than cytology, despite having lower specificity [240].
The analysis of KRAS mutations in peritoneal cfDNA by ddPCR was very recently proposed as a biomarker for peritoneal surface malignancies [241]. MAF was found to correlate with the surgical peritoneal carcinomatosis index, showing that MAF < 1% were associated with complete cytoreduction in contrast to MAF > 1%. These results suggest that peritoneal ctDNA testing would be useful as a surrogate for disease burden and as an indicator of resectability in peritoneal carcinomatosis. However, in this study, MAF could not be related to survival. Interestingly, mutant KRAS ctDNA was detected in peritoneal fluid in 20% of patients with KRAS wild-type tumors (as determined by tissue analysis using Sanger sequencing and NGS). Some of these patients had received anti-EGFR therapy prior to sample collection. Thus, this discordance may be related to intratumoral heterogeneity and/or clonal selection in response to treatment [241].
Another recent study evaluated the use of plasma and peritoneal fluid in a combined liquid biopsy approach as a prognostic factor in patients with advanced colorectal and appendicular tumors undergoing complete cytoreductive surgery and hyperthermic intraperitoneal chemotherapy (CC-HIPEC) [152]. In this study, KRAS mutations were analyzed in plasma and peritoneal fluid by ddPCR before and after CC-HIPEC. Patients with detectable ctDNA in plasma or peritoneal fluid after treatment had shorter disease-free and overall survival (including the 3-year survival rate). Patients with positive ctDNA in post-treatment plasma experienced systemic relapses, whereas its presence in post-HIPEC peritoneal fluid was associated with peritoneal recurrence and/or systemic relapses. In turn, all patients with negative liquid biopsy after treatment remained disease free. Interestingly, post-HIPEC cytology was negative in all cases, but ctDNA was not neutralized in peritoneal fluid by this treatment, suggesting that its persistence may predict a worse outcome [152].
3.2.11 Fine Needle Aspirate
ddPCR for detection of RAS and BRAF mutation has been proposed as a complementary tool in association with cytology to increase the diagnostic sensitivity and specificity of FNA biopsy of thyroid nodules [242]. In another study, these mutations were detected by NGS and confirmed by ddPCR [243]. In line with this, BRAF V600E mutation was also detected in FNA fluid from patients with papillary thyroid carcinoma [244]. In fact, the performance of ddPCR was better than that of ARMS-PCR when detecting this mutation from thyroid nodule FNA samples [20].
Two different studies have highlighted the utility of analyzing the supernatant cfDNA from FNA, a fluid that is usually discarded after centrifugation to separate the cell pellet for cytospin or FFPE cell block preparations. Guibert et al. [245] detected EGFR, BRAF and KRAS mutations with ddPCR in supernatant cfDNA from patients with suspected lung cancer and adenocarcinomas with acquired EGFR resistance. Similarly, another study confirmed EGFR, KRAS, BRAF, PIK3CA and NRAS mutations by ddPCR in post-centrifuged supernatant from malignant and benign FNA needle rinses, including cases of melanoma and pancreatic, lung, colorectal, breast, urothelial and hepatocellular carcinoma [246]. The same authors later observed a 100% concordance between NGS and ddPCR results in a small subset of FNA samples from patients with NSCLC [247]. More recently, ddPCR has been used to confirm EGFR mutations in samples from patients with NSCLC when inconsistencies were found between ARMS and SuperARMS-PCR [236].
The analysis of KRAS mutations using washes from endoscopic ultrasound-guided FNA also proved useful for the detection of local recurrence in pancreatic cancer [248].
3.2.12 Pancreatic Juice/Cyst Fluid/Bile
Suenaga et al. [249] determined that KRAS codon 12 and GNAS codon 201 mutations are detectable by ddPCR in pancreatic juice, finding higher concentrations when directly collected from the ampulla using an endoscopic distal cap. In addition, they observed that the optimal time for collection of pancreatic juice to increase the likelihood of detecting KRAS mutations in this biofluid was 10 minutes after secretin infusion [250].
KRAS mutations have also been examined with ddPCR in cfDNA from bile samples and plasma of patients with PDAC and cholangiocarcinoma, with results comparable to those with NGS. These results suggest that bile-based liquid biopsy by ddPCR might be a reliable tool for diagnosis of pancreatobiliary cancers [251].
In a different line of research, ddPCR was used to detect clinically relevant bacteria in pancreatic duct fluid and in bile and jejunal contents from patients undergoing pancreaticoduodenectomy [216].
Studies describing the use of ddPCR in biologic fluids other than blood are summarized in Table 4.
3.3 Concluding Remarks
In this updated review, we emphasize that the number of publications evaluating ddPCR has increased considerably in recent years, particularly in the field of liquid biopsy. The clinical utility of this methodology has been reinforced not only by its combination with other techniques, mainly NGS, but also by the integration of different biomarkers in multianalyte or multiparametric approaches. ddPCR is also usually employed as a validation, control or reference system to confirm the results obtained by other technologies, which highlights its reliability for detection of a wide variety of genetic alterations.
Blood is the main source of nucleic acids for noninvasive biomarker analysis by ddPCR, but the range of biologic fluids exploited for this purpose has extensively broadened in the last 4 years, providing a complementary tool for diagnosis and surveillance in many cancer types. ctDNA is the more widespread nucleic acid-based biomarker studied, but there is an increasing trend towards the investigation of RNA signatures, including long non-coding RNAs, miRNAs or mRNAs, frequently derived from exosomes or EVs, alone or in combination with other markers, such as CTCs.
Despite the evident prevalence of the Bio-Rad ddPCR platform in the literature, the development of new ddPCR platforms with additional detection channels paves the way for improved multiplexing strategies. Even so, limitations regarding issues such as scalability and sensitivity in samples with low DNA content and low-DNA-shedding tumors at early stages or anatomically sequestered locations, and the lack of standardized protocols still need to be overcome. Further research on the application of ddPCR in oncology is warranted.
References
Hindson BJ, Ness KD, Masquelier DA, Belgrader P, Heredia NJ, Makarewicz AJ, et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem. 2011;83:8604–10.
Hindson CM, Chevillet JR, Briggs HA, Gallichotte EN, Ruf IK, Hindson BJ, et al. Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat Methods. 2013;10:1003–5.
Siravegna G, Marsoni S, Siena S, Bardelli A. Integrating liquid biopsies into the management of cancer. Nat Rev Clin Oncol. 2017;14:531–48.
Olmedillas-López S, García-Arranz M, García-Olmo D. Current and emerging applications of droplet digital PCR in oncology. Mol Diagn Ther. 2017;21:493–510.
Dangla R, Kayi SC, Baroud CN. Droplet microfluidics driven by gradients of confinement. Proc Natl Acad Sci USA. 2013;110:853–8.
Basu AS. Digital assays part I: partitioning statistics and digital PCR. SLAS Technol. 2017;22:369–86.
Madic J, Zocevic A, Senlis V, Fradet E, Andre B, Muller S, et al. Three-color crystal digital PCR. Biomol Detect Quantif. 2016;10:34–46.
Jovelet C, Madic J, Remon J, Honoré A, Girard R, Rouleau E, et al. Crystal digital droplet PCR for detection and quantification of circulating EGFR sensitizing and resistance mutations in advanced non-small cell lung cancer. PLoS ONE. 2017;12: e0183319.
Madic J, Jovelet C, Lopez J, André B, Fatien J, Miran I, et al. EGFR C797S, EGFR T790M and EGFR sensitizing mutations in non-small cell lung cancer revealed by six-color crystal digital PCR. Oncotarget. 2018;9:37393–406.
Madic J, Jovelet C, Dehri I, Mallory AC. 6-Color crystal digital PCRTM for the high-plex detection of EGFR mutations in non-small cell lung cancer. Methods Mol Biol. 2021;2279:127–44.
Song X, Gong J, Zhang X, Feng X, Huang H, Gao M, et al. Plasma-based early screening and monitoring of EGFR mutations in NSCLC patients by a 3-color digital PCR assay. Br J Cancer. 2020;123:1437–44.
de Rouge TLM, Du FL, Corne J, Godey F, Bourien H, Brunot A, et al. 58P Detection of PIK3CA mutations in plasma ctDNA by crystal digital PCR for the selection of alpelisib treatment in routine clinical practice in advanced breast cancer patients. Ann Oncol. 2020;31:S35.
Dong L, Wang X, Wang S, Du M, Niu C, Yang J, et al. Interlaboratory assessment of droplet digital PCR for quantification of BRAF V600E mutation using a novel DNA reference material. Talanta. 2020;207: 120293.
Pedini P, Cherouat N, Basire A, Simon S, Budon L, Pourtein M, et al. Evaluation of next-generation sequencing and crystal digital PCR for Chimerism monitoring of post-allogeneic hematopoietic stem cell transplantation. Transplant Cell Ther. 2021;27:89.e1-89.e10.
Isaksson S, George AM, Jönsson M, Cirenajwis H, Jönsson P, Bendahl P-O, et al. Pre-operative plasma cell-free circulating tumor DNA and serum protein tumor markers as predictors of lung adenocarcinoma recurrence. Acta Oncol. 2019;58:1079–86.
Pettersson L, Chen Y, George AM, Rigo R, Lazarevic V, Juliusson G, et al. Subclonal patterns in follow-up of acute myeloid leukemia combining whole exome sequencing and ultrasensitive IBSAFE digital droplet analysis. Leuk Lymphoma. 2020;61:2168–79.
Arildsen NS, de la Fuente LM, Måsbäck A, Malander S, Forslund O, Kannisto P, et al. Detecting TP53 mutations in diagnostic and archival liquid-based Pap samples from ovarian cancer patients using an ultra-sensitive ddPCR method. Sci Rep. 2019;9:15506.
Förnvik D, Aaltonen KE, Chen Y, George AM, Brueffer C, Rigo R, et al. Detection of circulating tumor cells and circulating tumor DNA before and after mammographic breast compression in a cohort of breast cancer patients scheduled for neoadjuvant treatment. Breast Cancer Res Treat. 2019;177:447–55.
Bratulic S, Gatto F, Nielsen J. The translational status of cancer liquid biopsies. Regen Eng Transl Med. 2021;7:312–52.
Li X, Du H, Luo J, Ding W, Lai B, He J, et al. Comparison of the clinical validity of droplet digital PCR to ARMS-PCR for BRAF v600e mutation detection in thyroid nodules. J Clin Lab Anal. 2020;34: e23458.
Zhou H, Jia X, Hu K, Mo Z, Xu W, Peng L, et al. TMEM2 binds to CSNK2A3 to inhibit HBV infection via activation of the JAK/STAT pathway. Exp Cell Res. 2021;400:112517.
Bill M, Grimm J, Jentzsch M, Kloss L, Goldmann K, Schulz J, et al. Digital droplet PCR-based absolute quantification of pre-transplant NPM1 mutation burden predicts relapse in acute myeloid leukemia patients. Ann Hematol. 2018;97:1757–65.
Nakamura S, Yokoyama K, Yusa N, Ogawa M, Takei T, Kobayashi A, et al. Circulating tumor DNA dynamically predicts response and/or relapse in patients with hematological malignancies. Int J Hematol. 2018;108:402–10.
Della Starza I, Cavalli M, De Novi LA, Genuardi E, Mantoan B, Drandi D, et al. Minimal residual disease (MRD) in non-Hodgkin lymphomas: interlaboratory reproducibility on marrow samples with very low levels of disease within the FIL (Fondazione Italiana Linfomi) MRD Network. Hematol Oncol. 2019;37:368–74.
Bussaglia E, Pratcorona M, Carricondo M, Sansegundo L, Rubio MA, Monter A, et al. Application of a digital PCR method for WT1 to myeloid neoplasms in CR and deep ELN WT1 molecular response (< 10 copies). Ann Hematol. 2020;99:765–72.
Cortés AA, Olmedillas S, Serrano-López J, Lainez-González D, Castaño T, Iñiguez R, et al. Comparison of droplet digital PCR versus qPCR measurements on the international scale for the molecular monitoring of chronic myeloid leukemia patients. Mol Diagn Ther. 2020;24:593–600.
Drandi D, Alcantara M, Benmaad I, Söhlbrandt A, Lhermitte L, Zaccaria G, et al. Droplet digital PCR quantification of mantle cell lymphoma follow-up samples from four prospective trials of the European MCL network. Hemasphere. 2020;4: e347.
Petiti J, Lo Iacono M, Dragani M, Pironi L, Fantino C, Rapanotti MC, et al. Novel multiplex droplet digital PCR assays to monitor minimal residual disease in chronic myeloid leukemia patients showing atypical BCR-ABL1 transcripts. J Clin Med. 2020;9:1457.
Coccaro N, Anelli L, Zagaria A, Casieri P, Tota G, Orsini P, et al. Droplet digital PCR is a robust tool for monitoring minimal residual disease in adult philadelphia-positive acute lymphoblastic leukemia. J Mol Diagn. 2018;20:474–82.
Wang W-J, Zheng C-F, Liu Z, Tan Y-H, Chen X-H, Zhao B-L, et al. Droplet digital PCR for BCR/ABL(P210) detection of chronic myeloid leukemia: a high sensitive method of the minimal residual disease and disease progression. Eur J Haematol. 2018;101:291–6.
Krumbholz M, Goerlitz K, Albert C, Lawlor J, Suttorp M, Metzler M. Large amplicon droplet digital PCR for DNA-based monitoring of pediatric chronic myeloid leukaemia. J Cell Mol Med. 2019;23:4955–61.
Nicolini FE, Dulucq S, Boureau L, Cony-Makhoul P, Charbonnier A, Escoffre-Barbe M, et al. Evaluation of residual disease and TKI duration are critical predictive factors for molecular recurrence after stopping imatinib first-line in chronic phase CML patients. Clin Cancer Res. 2019;25:6606–13.
Park H, Shin D-Y, Kim I, Sohn S-K, Koh Y, Lee J-H, et al. Use of droplet digital polymerase chain reaction for detecting minimal residual disease: a prospective multi-institutional study. Vivo. 2019;33:2273–80.
Franke G-N, Maier J, Wildenberger K, Cross M, Giles FJ, Müller MC, et al. Comparison of real-time quantitative PCR and digital droplet PCR for BCR-ABL1 monitoring in patients with chronic myeloid leukemia. J Mol Diagn. 2020;22:81–9.
Colafigli G, Scalzulli E, Porrazzo M, Diverio D, Loglisci MG, Latagliata R, et al. Digital droplet PCR at the time of TKI discontinuation in chronic-phase chronic myeloid leukemia patients is predictive of treatment-free remission outcome. Hematol Oncol. 2019;37:652–4.
Atallah E, Schiffer CA, Radich JP, Weinfurt KP, Zhang M-J, Pinilla-Ibarz J, et al. Assessment of outcomes after stopping tyrosine kinase inhibitors among patients with chronic myeloid leukemia: a nonrandomized clinical trial. JAMA Oncol. 2021;7:42–50.
Chung HJ, Hur M, Yoon S, Hwang K, Lim HS, Kim H, et al. Performance evaluation of the QXDx BCR-ABL %IS droplet digital PCR assay. Ann Lab Med. 2020;40:72–5.
Soverini S, Bernardi S, Galimberti S. Molecular testing in CML between old and new methods: are we at a turning point? J Clin Med. 2020;9:E3865.
Orsini P, Impera L, Parciante E, Cumbo C, Minervini CF, Minervini A, et al. Droplet digital PCR for the quantification of Alu methylation status in hematological malignancies. Diagn Pathol. 2018;13:98.
Petiti J, Rosso V, Croce E, Franceschi V, Andreani G, Dragani M, et al. Highly sensitive detection of IDH2 mutations in acute myeloid leukemia. J Clin Med. 2020;9:271.
Takamatsu H, Wee RK, Zaimoku Y, Murata R, Zheng J, Moorhead M, et al. A comparison of minimal residual disease detection in autografts among ASO-qPCR, droplet digital PCR, and next-generation sequencing in patients with multiple myeloma who underwent autologous stem cell transplantation. Br J Haematol. 2018;183:664–8.
Della Starza I, De Novi LA, Cavalli M, Novelli N, Soscia R, Genuardi E, et al. Immunoglobulin kappa deleting element rearrangements are candidate targets for minimal residual disease evaluation in mantle cell lymphoma. Hematol Oncol. 2020;38:698–704.
Waterhouse M, Pfeifer D, Follo M, Duyster J, Schäfer H, Bertz H, et al. Early mixed hematopoietic chimerism detection by digital droplet PCR in patients undergoing gender-mismatched hematopoietic stem cell transplantation. Clin Chem Lab Med. 2017;55:1115–21.
Kliman D, Castellano-Gonzalez G, Withers B, Street J, Tegg E, Mirochnik O, et al. Ultra-sensitive droplet digital PCR for the assessment of microchimerism in cellular therapies. Biol Blood Marrow Transplant. 2018;24:1069–78.
Mika T, Baraniskin A, Ladigan S, Wulf G, Dierks S, Haase D, et al. Digital droplet PCR-based chimerism analysis for monitoring of hematopoietic engraftment after allogeneic stem cell transplantation. Int J Lab Hematol. 2019;41:615–21.
Waterhouse M, Pfeifer D, Duque-Afonso J, Follo M, Duyster J, Depner M, et al. Droplet digital PCR for the simultaneous analysis of minimal residual disease and hematopoietic chimerism after allogeneic cell transplantation. Clin Chem Lab Med. 2019;57:641–7.
Davis L, Riccitelli N, Valencia N, Ch’en IL, Tangri S, Brogdon JL, et al. Monitoring of tisagenlecleucel transgene DNA using a quantitative polymerase chain reaction assay. Mol Ther Methods Clin Dev. 2021;20:535–41.
Fehse B, Badbaran A, Berger C, Sonntag T, Riecken K, Geffken M, et al. Digital PCR assays for precise quantification of CD19-CAR-T cells after treatment with Axicabtagene Ciloleucel. Mol Ther Methods Clin Dev. 2020;16:172–8.
Badbaran A, Berger C, Riecken K, Kruchen A, Geffken M, Müller I, et al. Accurate in-vivo quantification of CD19 CAR-T cells after treatment with Axicabtagene Ciloleucel (Axi-Cel) and Tisagenlecleucel (Tisa-Cel) using digital PCR. Cancers (Basel). 2020;12:1970.
Mika T, Maghnouj A, Klein-Scory S, Ladigan-Badura S, Baraniskin A, Thomson J, et al. Digital-droplet PCR for quantification of CD19-directed CAR T-Cells. Front Mol Biosci. 2020;7:84.
Pabst T, Joncourt R, Shumilov E, Heini A, Wiedemann G, Legros M, et al. Analysis of IL-6 serum levels and CAR T cell-specific digital PCR in the context of cytokine release syndrome. Exp Hematol. 2020;88:7-14.e3.
Wang D, Zeng C, Xu B, Xu J-H, Wang J, Jiang L-J, et al. Anti-CD30 chimeric antigen receptor T cell therapy for relapsed/refractory CD30+ lymphoma patients. Blood Cancer J. 2020;10:8.
Tambaro FP, Singh H, Jones E, Rytting M, Mahadeo KM, Thompson P, et al. Autologous CD33-CAR-T cells for treatment of relapsed/refractory acute myelogenous leukemia. Leukemia. 2021;35:3282–6.
Vedvyas Y, McCloskey JE, Yang Y, Min IM, Fahey TJ, Zarnegar R, et al. Manufacturing and preclinical validation of CAR T cells targeting ICAM-1 for advanced thyroid cancer therapy. Sci Rep. 2019;9:10634.
Lu A, Liu H, Shi R, Cai Y, Ma J, Shao L, et al. Application of droplet digital PCR for the detection of vector copy number in clinical CAR/TCR T cell products. J Transl Med. 2020;18:191.
Amantini C, Morelli MB, Nabissi M, Piva F, Marinelli O, Maggi F, et al. Expression profiling of circulating tumor cells in pancreatic ductal adenocarcinoma patients: biomarkers predicting overall survival. Front Oncol. 2019;9:874.
Tagawa ST, Antonarakis ES, Gjyrezi A, Galletti G, Kim S, Worroll D, et al. Expression of AR-V7 and ARv(567es) in circulating tumor cells correlates with outcomes to taxane therapy in men with metastatic prostate cancer treated in TAXYNERGY. Clin Cancer Res. 2019;25:1880–8.
Aya-Bonilla CA, Morici M, Hong X, McEvoy AC, Sullivan RJ, Freeman J, et al. Detection and prognostic role of heterogeneous populations of melanoma circulating tumour cells. Br J Cancer. 2020;122:1059–67.
Cheng H, Liu C, Jiang J, Luo G, Lu Y, Jin K, et al. Analysis of ctDNA to predict prognosis and monitor treatment responses in metastatic pancreatic cancer patients. Int J Cancer. 2017;140:2344–50.
Lyskjær I, Kronborg CS, Rasmussen MH, Sørensen BS, Demuth C, Rosenkilde M, et al. Correlation between early dynamics in circulating tumour DNA and outcome from FOLFIRI treatment in metastatic colorectal cancer. Sci Rep. 2019;9:11542.
Iwama E, Sakai K, Hidaka N, Inoue K, Fujii A, Nakagaki N, et al. Longitudinal monitoring of somatic genetic alterations in circulating cell-free DNA during treatment with epidermal growth factor receptor-tyrosine kinase inhibitors. Cancer. 2020;126:219–27.
Chen M, Mithraprabhu S, Ramachandran M, Choi K, Khong T, Spencer A. Utility of circulating cell-free RNA analysis for the characterization of global transcriptome profiles of multiple myeloma patients. Cancers (Basel). 2019;11:887.
Helwa I, Cai J, Drewry MD, Zimmerman A, Dinkins MB, Khaled ML, et al. A comparative study of serum exosome isolation using differential ultracentrifugation and three commercial reagents. PLoS ONE. 2017;12: e0170628.
Castillo J, Bernard V, San Lucas FA, Allenson K, Capello M, Kim DU, et al. Surfaceome profiling enables isolation of cancer-specific exosomal cargo in liquid biopsies from pancreatic cancer patients. Ann Oncol. 2018;29:223–9.
Bernard V, Kim DU, San Lucas FA, Castillo J, Allenson K, Mulu FC, et al. Circulating nucleic acids are associated with outcomes of patients with pancreatic cancer. Gastroenterology. 2019;156:108-118.e4.
Gasparello J, Papi C, Allegretti M, Giordani E, Carboni F, Zazza S, et al. A distinctive microRNA (miRNA) signature in the blood of colorectal cancer (CRC) patients at surgery. Cancers (Basel). 2020;12:2410.
Puigdelloses M, González-Huárriz M, García-Moure M, Martínez-Vélez N, Esparragosa Vázquez I, Bruna J, et al. RNU6-1 in circulating exosomes differentiates GBM from non-neoplastic brain lesions and PCNSL but not from brain metastases. Neurooncol Adv. 2020;2:vdaa010.
Wang Z-Y, Wang R-X, Ding X-Q, Zhang X, Pan X-R, Tong J-H. A protocol for cancer-related mutation detection on exosomal DNA in clinical application. Front Oncol. 2020;10: 558106.
Joncas F-H, Lucien F, Rouleau M, Morin F, Leong HS, Pouliot F, et al. Plasma extracellular vesicles as phenotypic biomarkers in prostate cancer patients. Prostate. 2019;79:1767–76.
Yap SA, Münster-Wandowski A, Nonnenmacher A, Keilholz U, Liebs S. Analysis of cancer-related mutations in extracellular vesicles RNA by Droplet DigitalTM PCR. Biotechniques. 2020;69:99–107.
Sun N, Lee Y-T, Zhang RY, Kao R, Teng P-C, Yang Y, et al. Purification of HCC-specific extracellular vesicles on nanosubstrates for early HCC detection by digital scoring. Nat Commun. 2020;11:4489.
Lin Y, Leng Q, Zhan M, Jiang F. A plasma long noncoding RNA signature for early detection of lung cancer. Transl Oncol. 2018;11:1225–31.
Li Y, Li G, Chen X, Huang H, Liao L, Yuan T, et al. A novel lncRNA NONHSAT053785 acts as an independent risk factor for intrahepatic metastasis of hepatocellular carcinoma. Onco Targets Ther. 2020;13:5455–66.
Possieri C, Locantore P, Salis C, Bacci L, Aiello A, Fadda G, et al. Combined molecular and mathematical analysis of long noncoding RNAs expression in fine needle aspiration biopsies as novel tool for early diagnosis of thyroid cancer. Endocrine. 2021;72:711–20.
Manoochehri M, Jones M, Tomczyk K, Fletcher O, Schoemaker MJ, Swerdlow AJ, et al. DNA methylation of the long intergenic noncoding RNA 299 gene in triple-negative breast cancer: results from a prospective study. Sci Rep. 2020;10:11762.
Tavano F, Gioffreda D, Valvano MR, Palmieri O, Tardio M, Latiano TP, et al. Droplet digital PCR quantification of miR-1290 as a circulating biomarker for pancreatic cancer. Sci Rep. 2018;8:16389.
Beheshti A, Stevenson K, Vanderburg C, Ravi D, McDonald JT, Christie AL, et al. Identification of circulating serum multi-MicroRNA signatures in human DLBCL models. Sci Rep. 2019;9:17161.
Mazza T, Gioffreda D, Fontana A, Biagini T, Carella M, Palumbo O, et al. Clinical significance of circulating miR-1273g-3p and miR-122-5p in pancreatic cancer. Front Oncol. 2020;10:44.
Tombolan L, Millino C, Pacchioni B, Cattelan M, Zin A, Bonvini P, et al. Circulating miR-26a as potential prognostic biomarkers in pediatric rhabdomyosarcoma. Front Genet. 2020;11: 606274.
Perdas E, Stawski R, Kaczka K, Zubrzycka M. Analysis of Let-7 family miRNA in plasma as potential predictive biomarkers of diagnosis for papillary thyroid cancer. Diagnostics (Basel). 2020;10:130.
Seitz AK, Thoene S, Bietenbeck A, Nawroth R, Tauber R, Thalgott M, et al. AR-V7 in peripheral whole blood of patients with castration-resistant prostate cancer: association with treatment-specific outcome under abiraterone and enzalutamide. Eur Urol. 2017;72:828–34.
Del Re M, Bertolini I, Crucitta S, Fontanelli L, Rofi E, De Angelis C, et al. Overexpression of TK1 and CDK9 in plasma-derived exosomes is associated with clinical resistance to CDK4/6 inhibitors in metastatic breast cancer patients. Breast Cancer Res Treat. 2019;178:57–62.
Thwin KKM, Ishida T, Uemura S, Yamamoto N, Lin KS, Tamura A, et al. Level of seven neuroblastoma-associated mRNAs detected by droplet digital PCR is associated with tumor relapse/regrowth of high-risk neuroblastoma patients. J Mol Diagn. 2020;22:236–46.
Möhrmann L, Huang HJ, Hong DS, Tsimberidou AM, Fu S, Piha-Paul SA, et al. Liquid biopsies using plasma exosomal nucleic acids and plasma cell-free DNA compared with clinical outcomes of patients with advanced cancers. Clin Cancer Res. 2018;24:181–8.
Buscail E, Alix-Panabières C, Quincy P, Cauvin T, Chauvet A, Degrandi O, et al. High clinical value of liquid biopsy to detect circulating tumor cells and tumor exosomes in pancreatic ductal adenocarcinoma patients eligible for up-front surgery. Cancers (Basel). 2019;11:1656.
Nimir M, Ma Y, Jeffreys SA, Opperman T, Young F, Khan T, et al. Detection of AR-V7 in liquid biopsies of castrate resistant prostate cancer patients: a comparison of AR-v7 analysis in circulating tumor cells, circulating tumor RNA and exosomes. Cells. 2019;8:688.
Gai W, Ji L, Lam WKJ, Sun K, Jiang P, Chan AWH, et al. Liver- and colon-specific dna methylation markers in plasma for investigation of colorectal cancers with or without liver metastases. Clin Chem. 2018;64:1239–49.
Khan K, Rata M, Cunningham D, Koh D-M, Tunariu N, Hahne JC, et al. Functional imaging and circulating biomarkers of response to regorafenib in treatment-refractory metastatic colorectal cancer patients in a prospective phase II study. Gut. 2018;67:1484–92.
Spindler K-LG, Demuth C, Sorensen BS, Johansen JS, Nielsen D, Pallisgaard N, et al. Total cell-free DNA, carcinoembryonic antigen, and C-reactive protein for assessment of prognosis in patients with metastatic colorectal cancer. Tumour Biol. 2018;40:1010428318811207.
Thomsen CB, Hansen TF, Andersen RF, Lindebjerg J, Jensen LH, Jakobsen A. Monitoring the effect of first line treatment in RAS/RAF mutated metastatic colorectal cancer by serial analysis of tumor specific DNA in plasma. J Exp Clin Cancer Res. 2018;37:55.
Bidard F-C, Kiavue N, Ychou M, Cabel L, Stern M-H, Madic J, et al. Circulating tumor cells and circulating tumor DNA detection in potentially resectable metastatic colorectal cancer: a prospective ancillary study to the unicancer prodige-14 trial. Cells. 2019;8:516.
Hamfjord J, Guren TK, Dajani O, Johansen JS, Glimelius B, Sorbye H, et al. Total circulating cell-free DNA as a prognostic biomarker in metastatic colorectal cancer before first-line oxaliplatin-based chemotherapy. Ann Oncol. 2019;30:1088–95.
Lueong SS, Herbst A, Liffers S-T, Bielefeld N, Horn PA, Tannapfel A, et al. Serial circulating tumor DNA mutational status in patients with KRAS-mutant metastatic colorectal cancer from the phase 3 AIO KRK0207 trial. Clin Chem. 2020.
Polivka J, Windrichova J, Pesta M, Houfkova K, Rezackova H, Macanova T, et al. The level of preoperative plasma KRAS mutations and CEA predict survival of patients undergoing surgery for colorectal cancer liver metastases. Cancers (Basel). 2020;12:2434.
Silveira AB, Bidard F-C, Kasperek A, Melaabi S, Tanguy M-L, Rodrigues M, et al. High-accuracy determination of microsatellite instability compatible with liquid biopsies. Clin Chem. 2020;66:606–13.
Thomsen CB, Hansen TF, Andersen RF, Lindebjerg J, Jensen LH, Jakobsen A. Early identification of treatment benefit by methylated circulating tumor DNA in metastatic colorectal cancer. Ther Adv Med Oncol. 2020;12:1758835920918472.
van’t Erve I, Greuter MJE, Bolhuis K, Vessies DCL, Leal A, Vink GR, et al. Diagnostic strategies toward clinical implementation of liquid biopsy RAS/BRAF circulating tumor DNA analyses in patients with metastatic colorectal cancer. J Mol Diagn. 2020;22:1430–7.
Kim JS, Bae GE, Kim S-H, Choi MK, Yeo M-K. Serum-based KRAS(G12/G13) mutation detection using droplet digital PCR: clinical implications and limitations in colorectal adenocarcinoma with tumor heterogeneity. Front Oncol. 2020;10: 604772.
Knebel FH, Bettoni F, Shimada AK, Cruz M, Alessi JV, Negrão MV, et al. Sequential liquid biopsies reveal dynamic alterations of EGFR driver mutations and indicate EGFR amplification as a new mechanism of resistance to osimertinib in NSCLC. Lung Cancer. 2017;108:238–41.
Ni J, Weng L, Liu Y, Sun Z, Bai C, Wang Y. Dynamic monitoring of EGFR mutations in circulating cell-free DNA for EGFR-mutant metastatic patients with lung cancer: early detection of drug resistance and prognostic significance. Oncol Lett. 2017;13:4549–57.
Zhu Y-J, Zhang H-B, Liu Y-H, Zhu Y-Z, Chen J, Li Y, et al. Association of mutant EGFR L858R and exon 19 concentration in circulating cell-free DNA using droplet digital PCR with response to EGFR-TKIs in NSCLC. Oncol Lett. 2017;14:2573–9.
Zhu Y-J, Zhang H-B, Liu Y-H, Zhang F-L, Zhu Y-Z, Li Y, et al. Quantitative cell-free circulating EGFR mutation concentration is correlated with tumor burden in advanced NSCLC patients. Lung Cancer. 2017;109:124–7.
Buder A, Hochmair MJ, Schwab S, Bundalo T, Schenk P, Errhalt P, et al. Cell-free plasma DNA-guided treatment with osimertinib in patients with advanced EGFR-mutated NSCLC. J Thorac Oncol. 2018;13:821–30.
Del Re M, Bordi P, Rofi E, Restante G, Valleggi S, Minari R, et al. The amount of activating EGFR mutations in circulating cell-free DNA is a marker to monitor osimertinib response. Br J Cancer. 2018;119:1252–8.
Ding PN, Becker TM, Bray VJ, Chua W, Ma YF, Lynch D, et al. The predictive and prognostic significance of liquid biopsy in advanced epidermal growth factor receptor-mutated non-small cell lung cancer: a prospective study. Lung Cancer. 2019;134:187–93.
Beagan JJ, Bach S, van Boerdonk RA, van Dijk E, Thunnissen E, van den Broek D, et al. Circulating tumor DNA analysis of EGFR-mutant non-small cell lung cancer patients receiving osimertinib following previous tyrosine kinase inhibitor treatment. Lung Cancer. 2020;145:173–80.
Chan DLH, Toh GLX, Goh LL. Clinical implementation of plasma EGFR T790M testing using droplet digital PCR in TKI-resistant NSCLC patients. Exp Mol Pathol. 2020;116: 104515.
Wehrle J, Philipp U, Jolic M, Follo M, Hussung S, Waldeck S, et al. Personalized treatment selection and disease monitoring using circulating tumor DNA profiling in real-world cancer patient management. Diagnostics (Basel). 2020;10:550.
Nardo G, Carlet J, Marra L, Bonanno L, Boscolo A, Dal Maso A, et al. Detection of low-frequency KRAS mutations in cfDNA from EGFR-mutated NSCLC patients after first-line EGFR tyrosine kinase inhibitors. Front Oncol. 2020;10: 607840.
Spence T, Perera S, Weiss J, Grenier S, Ranich L, Shepherd F, et al. Clinical implementation of circulating tumour DNA testing for EGFR T790M for detection of treatment resistance in non-small cell lung cancer. J Clin Pathol. 2021;74:91–7.
Del Re M, Vivaldi C, Rofi E, Vasile E, Miccoli M, Caparello C, et al. Early changes in plasma DNA levels of mutant KRAS as a sensitive marker of response to chemotherapy in pancreatic cancer. Sci Rep. 2017;7:7931.
Kim MK, Woo SM, Park B, Yoon K-A, Kim Y-H, Joo J, et al. Prognostic implications of multiplex detection of KRAS mutations in cell-free DNA from patients with pancreatic ductal adenocarcinoma. Clin Chem. 2018;64:726–34.
Groot VP, Mosier S, Javed AA, Teinor JA, Gemenetzis G, Ding D, et al. Circulating tumor DNA as a clinical test in resected pancreatic cancer. Clin Cancer Res. 2019;25:4973–84.
Wang Z-Y, Ding X-Q, Zhu H, Wang R-X, Pan X-R, Tong J-H. KRAS mutant allele fraction in circulating cell-free DNA correlates with clinical stage in pancreatic cancer patients. Front Oncol. 2019;9:1295.
Watanabe F, Suzuki K, Tamaki S, Abe I, Endo Y, Takayama Y, et al. Longitudinal monitoring of KRAS-mutated circulating tumor DNA enables the prediction of prognosis and therapeutic responses in patients with pancreatic cancer. PLoS ONE. 2019;14: e0227366.
Hussung S, Akhoundova D, Hipp J, Follo M, Klar RFU, Philipp U, et al. Longitudinal analysis of cell-free mutated KRAS and CA 19–9 predicts survival following curative resection of pancreatic cancer. BMC Cancer. 2021;21:49.
Yamaguchi T, Uemura K, Murakami Y, Kondo N, Nakagawa N, Okada K, et al. Clinical implications of pre- and postoperative circulating tumor DNA in patients with resected pancreatic ductal adenocarcinoma. Ann Surg Oncol. 2021;28:3135–44.
Moynahan ME, Chen D, He W, Sung P, Samoila A, You D, et al. Correlation between PIK3CA mutations in cell-free DNA and everolimus efficacy in HR(+), HER2(−) advanced breast cancer: results from BOLERO-2. Br J Cancer. 2017;116:726–30.
Kodahl AR, Ehmsen S, Pallisgaard N, Jylling AMB, Jensen JD, Laenkholm A-V, et al. Correlation between circulating cell-free PIK3CA tumor DNA levels and treatment response in patients with PIK3CA-mutated metastatic breast cancer. Mol Oncol. 2018;12:925–35.
Hrebien S, Citi V, Garcia-Murillas I, Cutts R, Fenwick K, Kozarewa I, et al. Early ctDNA dynamics as a surrogate for progression-free survival in advanced breast cancer in the BEECH trial. Ann Oncol. 2019;30:945–52.
Jacot W, Dalenc F, Lopez-Crapez E, Chaltiel L, Durigova A, Gros N, et al. PIK3CA mutations early persistence in cell-free tumor DNA as a negative prognostic factor in metastatic breast cancer patients treated with hormonal therapy. Breast Cancer Res Treat. 2019;177:659–67.
Gerratana L, Basile D, Franzoni A, Allegri L, Viotto D, Corvaja C, et al. Plasma-based longitudinal evaluation of ESR1 epigenetic status in hormone receptor-positive HER2-negative metastatic breast cancer. Front Oncol. 2020;10: 550185.
Liu Z-B, Ezzedine NE, Eterovic AK, Ensor JE, Huang HJ, Albanell J, et al. Detection of breast cancer stem cell gene mutations in circulating free DNA during the evolution of metastases. Breast Cancer Res Treat. 2019;178:251–61.
Jeannot E, Darrigues L, Michel M, Stern M-H, Pierga J-Y, Rampanou A, et al. A single droplet digital PCR for ESR1 activating mutations detection in plasma. Oncogene. 2020;39:2987–95.
Garlan F, Blanchet B, Kramkimel N, Puszkiel A, Golmard J-L, Noe G, et al. Circulating tumor DNA measurement by picoliter droplet-based digital PCR and vemurafenib plasma concentrations in patients with advanced BRAF-mutated melanoma. Target Oncol. 2017;12:365–71.
Keller L, Guibert N, Casanova A, Brayer S, Farella M, Delaunay M, et al. Early circulating tumour DNA variations predict tumour response in melanoma patients treated with immunotherapy. Acta Derm Venereol. 2019;99:206–10.
Lee RJ, Gremel G, Marshall A, Myers KA, Fisher N, Dunn JA, et al. Circulating tumor DNA predicts survival in patients with resected high-risk stage II/III melanoma. Ann Oncol. 2018;29:490–6.
Tan L, Sandhu S, Lee RJ, Li J, Callahan J, Ftouni S, et al. Prediction and monitoring of relapse in stage III melanoma using circulating tumor DNA. Ann Oncol. 2019;30:804–14.
Calbet-Llopart N, Potrony M, Tell-Martí G, Carrera C, Barreiro A, Aguilera P, et al. Detection of cell-free circulating BRAF(V) (600E) by droplet digital polymerase chain reaction in patients with and without melanoma under dermatological surveillance. Br J Dermatol. 2020;182:382–9.
Kozak K, Kowalik A, Gos A, Wasag B, Lugowska I, Jurkowska M, et al. Cell-free DNA BRAF V600E measurements during BRAF inhibitor therapy of metastatic melanoma: long-term analysis. Tumori. 2020;106:241–8.
Marsavela G, Johansson PA, Pereira MR, McEvoy AC, Reid AL, Robinson C, et al. The prognostic impact of circulating tumour DNA in melanoma patients treated with systemic therapies-beyond BRAF mutant detection. Cancers (Basel). 2020;12:3793.
Warburton L, Meniawy TM, Calapre L, Pereira M, McEvoy A, Ziman M, et al. Stopping targeted therapy for complete responders in advanced BRAF mutant melanoma. Sci Rep. 2020;10:18878.
Emelyanova MA, Telysheva EN, Orlova KV, Ryabaya OO, Snigiryova GP, Abramov IS, et al. Microarray-based analysis of the BRAF V600 mutations in circulating tumor DNA in melanoma patients. Cancer Genet. 2021;250–251:25–35.
Syeda MM, Wiggins JM, Corless BC, Long GV, Flaherty KT, Schadendorf D, et al. Circulating tumour DNA in patients with advanced melanoma treated with dabrafenib or dabrafenib plus trametinib: a clinical validation study. Lancet Oncol. 2021;22:370–80.
Conteduca V, Wetterskog D, Sharabiani MTA, Grande E, Fernandez-Perez MP, Jayaram A, et al. Androgen receptor gene status in plasma DNA associates with worse outcome on enzalutamide or abiraterone for castration-resistant prostate cancer: a multi-institution correlative biomarker study. Ann Oncol. 2017;28:1508–16.
Buelens S, Claeys T, Dhondt B, Poelaert F, Vynck M, Yigit N, et al. Prognostic and therapeutic implications of circulating androgen receptor gene copy number in prostate cancer patients using droplet digital polymerase chain reaction. Clin Genitourin Cancer. 2018;16:197-205.e5.
Haldrup C, Pedersen AL, Øgaard N, Strand SH, Høyer S, Borre M, et al. Biomarker potential of ST6GALNAC3 and ZNF660 promoter hypermethylation in prostate cancer tissue and liquid biopsies. Mol Oncol. 2018;12:545–60.
Conteduca V, Jayaram A, Romero-Laorden N, Wetterskog D, Salvi S, Gurioli G, et al. Plasma androgen receptor and docetaxel for metastatic castration-resistant prostate cancer. Eur Urol. 2019;75:368–73.
Conteduca V, Castro E, Wetterskog D, Scarpi E, Jayaram A, Romero-Laorden N, et al. Plasma AR status and cabazitaxel in heavily treated metastatic castration-resistant prostate cancer. Eur J Cancer. 2019;116:158–68.
Qu LG, Wardan H, Davis ID, Iddawela M, Sluka P, Pezaro CJ. Circulating oestrogen receptor mutations and splice variants in advanced prostate cancer. BJU Int. 2019;124(Suppl 1):50–6.
Bjerre MT, Nørgaard M, Larsen OH, Jensen SØ, Strand SH, Østergren P, et al. Epigenetic analysis of circulating tumor DNA in localized and metastatic prostate cancer: evaluation of clinical biomarker potential. Cells. 2020;9:1362.
Gurioli G, Conteduca V, Lolli C, Schepisi G, Gargiulo S, Altavilla A, et al. Plasma AR copy number changes and outcome to abiraterone and enzalutamide. Front Oncol. 2020;10: 567809.
Del Re M, Conteduca V, Crucitta S, Gurioli G, Casadei C, Restante G, et al. Androgen receptor gain in circulating free DNA and splicing variant 7 in exosomes predict clinical outcome in CRPC patients treated with abiraterone and enzalutamide. Prostate Cancer Prostatic Dis. 2021;24:524–31.
Färkkilä A, McConechy MK, Yang W, Talhouk A, Ng Y, Lum A, et al. FOXL2 402C > G mutation can be identified in the circulating tumor DNA of patients with adult-type granulosa cell tumor. J Mol Diagn. 2017;19:126–36.
Link T, Kuhlmann JD, Kobelt D, Herrmann P, Vassileva YD, Kramer M, et al. Clinical relevance of circulating MACC1 and S100A4 transcripts for ovarian cancer. Mol Oncol. 2019;13:1268–79.
Cirillo PDR, Margiotti K, Mesoraca A, Giorlandino C. Quantification of circulating microRNAs by droplet digital PCR for cancer detection. BMC Res Notes. 2020;13:351.
Ogasawara A, Hihara T, Shintani D, Yabuno A, Ikeda Y, Tai K, et al. Evaluation of circulating tumor DNA in patients with ovarian cancer harboring somatic PIK3CA or KRAS mutations. Cancer Res Treat. 2020;52:1219–28.
Rusan M, Andersen RF, Jakobsen A, Steffensen KD. Circulating HOXA9-methylated tumour DNA: a novel biomarker of response to poly (ADP-ribose) polymerase inhibition in BRCA-mutated epithelial ovarian cancer. Eur J Cancer. 2020;125:121–9.
Serrano C, Leal A, Kuang Y, Morgan JA, Barysauskas CM, Phallen J, et al. Phase I study of rapid alternation of sunitinib and regorafenib for the treatment of tyrosine kinase inhibitor refractory gastrointestinal stromal tumors. Clin Cancer Res. 2019;25:7287–93.
Serrano C, Vivancos A, López-Pousa A, Matito J, Mancuso FM, Valverde C, et al. Clinical value of next generation sequencing of plasma cell-free DNA in gastrointestinal stromal tumors. BMC Cancer. 2020;20:99.
Beagan JJ, Sluiter NR, Bach S, Eijk PP, Vlek SL, Heideman DAM, et al. Circulating tumor DNA as a preoperative marker of recurrence in patients with peritoneal metastases of colorectal cancer: a clinical feasibility study. J Clin Med. 2020;9:1738.
López-Rojo I, Olmedillas-López S, Villarejo Campos P, Domínguez Prieto V, Barambio Buendía J, Cortés Guiral D, et al. Liquid biopsy in peritoneal fluid and plasma as a prognostic factor in advanced colorectal and appendiceal tumors after complete cytoreduction and hyperthermic intraperitoneal chemotherapy. Ther Adv Med Oncol. 2020;12:1758835920981351.
Boeckx N, de Beeck KO, Beyens M, Deschoolmeester V, Hermans C, De Clercq P, et al. Mutation and methylation analysis of circulating tumor DNA Can Be used for follow-up of metastatic colorectal cancer patients. Clin Colorectal Cancer. 2018;17:e369–79.
Demuth C, Winther-Larsen A, Madsen AT, Meldgaard P, Sorensen BS. A method for treatment monitoring using circulating tumour DNA in cancer patients without targetable mutations. Oncotarget. 2018;9:31066–76.
Moati E, Blons H, Taly V, Garlan F, Wang-Renault S-F, Pietrasz D, et al. Plasma clearance of RAS mutation under therapeutic pressure is a rare event in metastatic colorectal cancer. Int J Cancer. 2020;147:1185–9.
Cavallone L, Aguilar-Mahecha A, Lafleur J, Brousse S, Aldamry M, Roseshter T, et al. Prognostic and predictive value of circulating tumor DNA during neoadjuvant chemotherapy for triple negative breast cancer. Sci Rep. 2020;10:14704.
Kang J-K, Heo S, Kim H-P, Song S-H, Yun H, Han S-W, et al. Liquid biopsy-based tumor profiling for metastatic colorectal cancer patients with ultra-deep targeted sequencing. PLoS ONE. 2020;15: e0232754.
Treger TD, Chagtai T, Butcher R, Cresswell GD, Al-Saadi R, Brok J, et al. Somatic TP53 mutations are detectable in circulating tumor DNA from children with anaplastic wilms tumors. Transl Oncol. 2018;11:1301–6.
Hu Y, Ulrich BC, Supplee J, Kuang Y, Lizotte PH, Feeney NB, et al. False-positive plasma genotyping due to clonal hematopoiesis. Clin Cancer Res. 2018;24:4437–43.
Leng Q, Lin Y, Zhan M, Jiang F. An integromic signature for lung cancer early detection. Oncotarget. 2018;9:24684–92.
Yang Z, LaRiviere MJ, Ko J, Till JE, Christensen T, Yee SS, et al. A Multianalyte panel consisting of extracellular vesicle miRNAs and mRNAs, cfDNA, and CA19-9 shows utility for diagnosis and staging of pancreatic ductal adenocarcinoma. Clin Cancer Res. 2020;26:3248–58.
Del Re M, Cucchiara F, Rofi E, Fontanelli L, Petrini I, Gri N, et al. A multiparametric approach to improve the prediction of response to immunotherapy in patients with metastatic NSCLC. Cancer Immunol Immunother. 2020;70:1667–78.
Colling R, Wang LM, Soilleux E. Validating a fully automated real-time PCR-based system for use in the molecular diagnostic analysis of colorectal carcinoma: a comparison with NGS and IHC. J Clin Pathol. 2017;70:610–4.
Normanno N, Esposito Abate R, Lambiase M, Forgione L, Cardone C, Iannaccone A, et al. RAS testing of liquid biopsy correlates with the outcome of metastatic colorectal cancer patients treated with first-line FOLFIRI plus cetuximab in the CAPRI-GOIM trial. Ann Oncol. 2018;29:112–8.
Prieto-Potin I, Montagut C, Bellosillo B, Evans M, Smith M, Melchior L, et al. Multicenter evaluation of the idylla NRAS-BRAF mutation test in metastatic colorectal cancer. J Mol Diagn. 2018;20:664–76.
Jensen SØ, Øgaard N, Ørntoft M-BW, Rasmussen MH, Bramsen JB, Kristensen H, et al. Novel DNA methylation biomarkers show high sensitivity and specificity for blood-based detection of colorectal cancer-a clinical biomarker discovery and validation study. Clin Epigenetics. 2019;11:158.
Pasquale R, Forgione L, Roma C, Fenizia F, Bergantino F, Rachiglio AM, et al. Targeted sequencing analysis of cell-free DNA from metastatic non-small-cell lung cancer patients: clinical and biological implications. Transl Lung Cancer Res. 2020;9:61–70.
dMIQE Group, Huggett JF. The digital MIQE guidelines update: minimum information for publication of quantitative digital PCR experiments for 2020. Clin Chem. 2020;66:1012–29.
Huggett JF, Foy CA, Benes V, Emslie K, Garson JA, Haynes R, et al. The digital MIQE guidelines: minimum information for publication of quantitative digital PCR experiments. Clin Chem. 2013;59:892–902.
Hirano M, Ohka F, Maeda S, Chalise L, Yamamichi A, Aoki K, et al. A novel high-sensitivity assay to detect a small fraction of mutant IDH1 using droplet digital PCR. Brain Tumor Pathol. 2018;35:97–105.
Hiemcke-Jiwa LS, Minnema MC, Radersma-van Loon JH, Jiwa NM, de Boer M, Leguit RJ, et al. The use of droplet digital PCR in liquid biopsies: a highly sensitive technique for MYD88 p.(L265P) detection in cerebrospinal fluid. Hematol Oncol. 2018;36:429–35.
Hiemcke-Jiwa LS, Leguit RJ, Snijders TJ, Bromberg JEC, Nierkens S, Jiwa NM, et al. MYD88 p.(L265P) detection on cell-free DNA in liquid biopsies of patients with primary central nervous system lymphoma. Br J Haematol. 2019;185:974–7.
Chen K, Ma Y, Ding T, Zhang X, Chen B, Guan M. Effectiveness of digital PCR for MYD88L265P detection in vitreous fluid for primary central nervous system lymphoma diagnosis. Exp Ther Med. 2020;20:301–8.
Zorofchian S, Lu G, Zhu J-J, Duose DY, Windham J, Esquenazi Y, et al. Detection of the MYD88 p.L265P mutation in the CSF of a patient with secondary central nervous system lymphoma. Front Oncol. 2018;8:382.
Shikata H, Kihara H, Kaneko M, Matsukage S, Hattori K. Monitoring of MYD88 L265P mutation by droplet digital polymerase chain reaction for prediction of early relapse in a patient with Bing-Neel syndrome. Int J Hematol. 2021;113:586–91.
Bobillo S, Crespo M, Escudero L, Mayor R, Raheja P, Carpio C, et al. Cell free circulating tumor DNA in cerebrospinal fluid detects and monitors central nervous system involvement of B-cell lymphomas. Haematologica. 2020;106:513–21.
Connolly ID, Li Y, Pan W, Johnson E, You L, Vogel H, et al. A pilot study on the use of cerebrospinal fluid cell-free DNA in intramedullary spinal ependymoma. J Neurooncol. 2017;135:29–36.
Lu VM, Power EA, Zhang L, Daniels DJ. Liquid biopsy for diffuse intrinsic pontine glioma: an update. J Neurosurg Pediatr. 2019;24:593–600.
Azad TD, Jin MC, Bernhardt LJ, Bettegowda C. Liquid biopsy for pediatric diffuse midline glioma: a review of circulating tumor DNA and cerebrospinal fluid tumor DNA. Neurosurg Focus. 2020;48:E9.
Martínez-Ricarte F, Mayor R, Martínez-Sáez E, Rubio-Pérez C, Pineda E, Cordero E, et al. Molecular diagnosis of diffuse gliomas through sequencing of cell-free circulating tumor DNA from cerebrospinal fluid. Clin Cancer Res. 2018;24:2812–9.
Panditharatna E, Kilburn LB, Aboian MS, Kambhampati M, Gordish-Dressman H, Magge SN, et al. Clinically relevant and minimally invasive tumor surveillance of pediatric diffuse midline gliomas using patient-derived liquid biopsy. Clin Cancer Res. 2018;24:5850–9.
Stallard S, Savelieff MG, Wierzbicki K, Mullan B, Miklja Z, Bruzek A, et al. CSF H3F3A K27M circulating tumor DNA copy number quantifies tumor growth and in vitro treatment response. Acta Neuropathol Commun. 2018;6:80.
Bruzek AK, Ravi K, Muruganand A, Wadden J, Babila CM, Cantor E, et al. Electronic DNA analysis of CSF cell-free tumor DNA to quantify multi-gene molecular response in pediatric high-grade glioma. Clin Cancer Res. 2020;26:6266–76.
Escudero L, Llort A, Arias A, Diaz-Navarro A, Martínez-Ricarte F, Rubio-Perez C, et al. Circulating tumour DNA from the cerebrospinal fluid allows the characterisation and monitoring of medulloblastoma. Nat Commun. 2020;11:5376.
Arulananda S, Do H, Rivalland G, Loh Z, Musafer A, Lau E, et al. Standard dose osimertinib for erlotinib refractory T790M-negative EGFR-mutant non-small cell lung cancer with leptomeningeal disease. J Thorac Dis. 2019;11:1756–64.
Huang R, Xu X, Li D, Chen K, Zhan Q, Ge M, et al. Digital PCR-based detection of EGFR mutations in paired plasma and CSF samples of lung adenocarcinoma patients with central nervous system metastases. Target Oncol. 2019;14:343–50.
van Bussel MTJ, Pluim D, Milojkovic Kerklaan B, Bol M, Sikorska K, Linders DTC, et al. Circulating epithelial tumor cell analysis in CSF in patients with leptomeningeal metastases. Neurology. 2020;94:e521–8.
Siravegna G, Geuna E, Mussolin B, Crisafulli G, Bartolini A, Galizia D, et al. Genotyping tumour DNA in cerebrospinal fluid and plasma of a HER2-positive breast cancer patient with brain metastases. ESMO Open. 2017;2: e000253.
Ballester LY, Glitza Oliva IC, Douse DY, Chen MM, Lan C, Haydu LE, et al. Evaluating circulating tumor DNA from the cerebrospinal fluid of patients with melanoma and leptomeningeal disease. J Neuropathol Exp Neurol. 2018;77:628–35.
Larsen LK, Jakobsen JS, Abdul-Al A, Guldberg P. Noninvasive detection of high grade prostate cancer by dna methylation analysis of urine cells captured by microfiltration. J Urol. 2018;200:749–57.
Kohaar I, Chen Y, Banerjee S, Borbiev T, Kuo H-C, Ali A, et al. A urine exosome gene expression panel distinguishes between indolent and aggressive prostate cancers at biopsy. J Urol. 2021;205:420–5.
Solé C, Goicoechea I, Goñi A, Schramm M, Armesto M, Arestin M, et al. The urinary transcriptome as a source of biomarkers for prostate cancer. Cancers (Basel). 2020;12:513.
Kang HW, Lee HY, Byun YJ, Jeong P, Yoon JS, Kim DH, et al. A novel urinary mRNA signature using the droplet digital polymerase chain reaction platform improves discrimination between prostate cancer and benign prostatic hyperplasia within the prostate-specific antigen gray zone. Investig Clin Urol. 2020;61:411–8.
Woo H-K, Park J, Ku JY, Lee CH, Sunkara V, Ha HK, et al. Urine-based liquid biopsy: non-invasive and sensitive AR-V7 detection in urinary EVs from patients with prostate cancer. Lab Chip. 2018;19:87–97.
Russo IJ, Ju Y, Gordon NS, Zeegers MP, Cheng KK, James ND, et al. Toward personalised liquid biopsies for urothelial carcinoma: characterisation of ddPCR and urinary cfDNA for the detection of the TERT 228 G > A/T mutation. Bladder Cancer. 2018;4:41–8.
Borkowska EM, Traczyk-Borszyńska M, Kutwin P, Pietrusiński M, Jabłonowski Z, Borowiec M. Usefulness of droplet digital PCR and Sanger sequencing for detection of FGFR3 mutation in bladder cancer. Urol Oncol. 2019;37:907–15.
Hayashi Y, Fujita K, Matsuzaki K, Matsushita M, Kawamura N, Koh Y, et al. Diagnostic potential of TERT promoter and FGFR3 mutations in urinary cell-free DNA in upper tract urothelial carcinoma. Cancer Sci. 2019;110:1771–9.
Hayashi Y, Fujita K, Matsuzaki K, Eich M-L, Tomiyama E, Matsushita M, et al. Clinical significance of hotspot mutation analysis of urinary cell-free DNA in urothelial bladder cancer. Front Oncol. 2020;10:755.
Hosen MI, Sheikh M, Zvereva M, Scelo G, Forey N, Durand G, et al. Urinary TERT promoter mutations are detectable up to 10 years prior to clinical diagnosis of bladder cancer: evidence from the Golestan Cohort Study. EBioMedicine. 2020;53: 102643.
Hosen MI, Forey N, Durand G, Voegele C, Bilici S, Avogbe PH, et al. Development of sensitive droplet digital PCR assays for detecting urinary TERT promoter mutations as non-invasive biomarkers for detection of urothelial cancer. Cancers (Basel). 2020;12:3541.
Christensen E, Birkenkamp-Demtröder K, Nordentoft I, Høyer S, van der Keur K, van Kessel K, et al. Liquid biopsy analysis of FGFR3 and PIK3CA hotspot mutations for disease surveillance in bladder cancer. Eur Urol. 2017;71:961–9.
Birkenkamp-Demtröder K, Christensen E, Nordentoft I, Knudsen M, Taber A, Høyer S, et al. Monitoring treatment response and metastatic relapse in advanced bladder cancer by liquid biopsy analysis. Eur Urol. 2018;73:535–40.
Michel ME, Konczyk DJ, Yeung KS, Murillo R, Vivero MP, Hall AM, et al. Causal somatic mutations in urine DNA from persons with the CLOVES subgroup of the PIK3CA-related overgrowth spectrum. Clin Genet. 2018;93:1075–80.
Biderman Waberski M, Lindhurst M, Keppler-Noreuil KM, Sapp JC, Baker L, Gripp KW, et al. Urine cell-free DNA is a biomarker for nephroblastomatosis or Wilms tumor in PIK3CA-related overgrowth spectrum (PROS). Genet Med. 2018;20:1077–81.
Terasawa H, Kinugasa H, Ako S, Hirai M, Matsushita H, Uchida D, et al. Utility of liquid biopsy using urine in patients with pancreatic ductal adenocarcinoma. Cancer Biol Ther. 2019;20:1348–53.
Crisafulli G, Mussolin B, Cassingena A, Montone M, Bartolini A, Barault L, et al. Whole exome sequencing analysis of urine trans-renal tumour DNA in metastatic colorectal cancer patients. ESMO Open. 2019;4: e000572.
Hu T, Shen H, Huang H, Song M, Yang Z, Zhou Y, et al. Urinary circulating DNA profiling in non-small cell lung cancer patients following treatment shows prognostic potential. J Thorac Dis. 2018;10:4137–46.
Yu H, Liu M, Qiu H, Yang K. Urinary and plasma cell-free DNA comparison for lung cancer patients treated with epidermal growth factor receptor-thyroxine kinase inhibitors. Am J Med Sci. 2019;357:29–36.
Herring E, Kanaoka S, Tremblay É, Beaulieu J-F. Droplet digital PCR for quantification of ITGA6 in a stool mRNA assay for the detection of colorectal cancers. World J Gastroenterol. 2017;23:2891–8.
Olmedillas-López S, Lévano-Linares DC, Alexandre CLA, Vega-Clemente L, Sánchez EL, Villagrasa A, et al. Detection of KRAS G12D in colorectal cancer stool by droplet digital PCR. World J Gastroenterol. 2017;23:7087–97.
Vega-Benedetti AF, Loi E, Moi L, Orrù S, Ziranu P, Pretta A, et al. Colorectal cancer early detection in stool samples tracing CpG islands methylation alterations affecting gene expression. Int J Mol Sci. 2020;21:4494.
Suehiro Y, Sakai K, Nishioka M, Hashimoto S, Takami T, Higaki S, et al. Highly sensitive stool DNA testing of Fusobacterium nucleatum as a marker for detection of colorectal tumours in a Japanese population. Ann Clin Biochem. 2017;54:86–91.
Talarico S, Safaeian M, Gonzalez P, Hildesheim A, Herrero R, Porras C, et al. Quantitative detection and genotyping of helicobacter pylori from stool using droplet digital PCR reveals variation in bacterial loads that correlates with cagA virulence gene carriage. Helicobacter. 2016;21:325–33.
Talarico S, Leverich CK, Wei B, Ma J, Cao X, Guo Y, et al. Increased H. pylori stool shedding and EPIYA-D cagA alleles are associated with gastric cancer in an East Asian hospital. PLoS ONE. 2018;13: e0202925.
Sun L, Talarico S, Yao L, He L, Self S, You Y, et al. Droplet digital PCR-based detection of clarithromycin resistance in Helicobacter pylori isolates reveals frequent heteroresistance. J Clin Microbiol. 2018;56:e00019–18.
Rogers MB, Aveson V, Firek B, Yeh A, Brooks B, Brower-Sinning R, et al. Disturbances of the perioperative microbiome across multiple body sites in patients undergoing pancreaticoduodenectomy. Pancreas. 2017;46:260–7.
Hiemcke-Jiwa LS, Ten Dam-van Loon NH, Leguit RJ, Nierkens S, Norel JO-v, de Boer JH, et al. Potential diagnosis of vitreoretinal lymphoma by detection of MYD88 mutation in aqueous humor with ultrasensitive droplet digital polymerase chain reaction. JAMA Ophthalmol. 2018;136:1098–104.
Shi H, Zhou X, Chen B, Xiao J, Li Y, Zhou X, et al. Clinical relevance of the high prevalence of MYD88 L265P mutated vitreoretinal lymphoma identified by droplet digital polymerase chain reaction. Ocul Immunol Inflamm. 2021;29:448–55.
Hanna GJ, Supplee JG, Kuang Y, Mahmood U, Lau CJ, Haddad RI, et al. Plasma HPV cell-free DNA monitoring in advanced HPV-associated oropharyngeal cancer. Ann Oncol. 2018;29:1980–6.
Hanna GJ, Lau CJ, Mahmood U, Supplee JG, Mogili AR, Haddad RI, et al. Salivary HPV DNA informs locoregional disease status in advanced HPV-associated oropharyngeal cancer. Oral Oncol England. 2019;95:120–6.
Wu M, Ouyang Y, Wang Z, Zhang R, Huang P-H, Chen C, et al. Isolation of exosomes from whole blood by integrating acoustics and microfluidics. Proc Natl Acad Sci USA. 2017;114:10584–9.
Wang Z, Li F, Rufo J, Chen C, Yang S, Li L, et al. Acoustofluidic salivary exosome isolation: a liquid biopsy compatible approach for human papillomavirus-associated oropharyngeal cancer detection. J Mol Diagn. 2020;22:50–9.
Ding S, Song X, Geng X, Liu L, Ma H, Wang X, et al. Saliva-derived cfDNA is applicable for EGFR mutation detection but not for quantitation analysis in non-small cell lung cancer. Thorac Cancer. 2019;10:1973–83.
Li F, Wei F, Huang W-L, Lin C-C, Li L, Shen MM, et al. Ultra-short circulating tumor DNA (usctDNA) in plasma and saliva of non-small cell lung cancer (NSCLC) patients. Cancers (Basel). 2020;12:2041.
Su Y, Fang HB, Jiang F. An epigenetic classifier for early stage lung cancer. Clin Epigenetics. 2018;10:68.
Isaka T, Yokose T, Ito H, Nakayama H, Miyagi Y, Saito H, et al. Detection of EGFR mutation of pulmonary adenocarcinoma in sputum using droplet digital PCR. BMC Pulm Med. 2021;21:100.
Hackner K, Buder A, Hochmair MJ, Strieder M, Grech C, Fabikan H, et al. Detection of EGFR activating and resistance mutations by droplet digital PCR in sputum of EGFR-mutated NSCLC patients. Clin Med Insights Oncol. 2021;15:1179554921993072.
Li H, Jiang Z, Leng Q, Bai F, Wang J, Ding X, et al. A prediction model for distinguishing lung squamous cell carcinoma from adenocarcinoma. Oncotarget. 2017;8:50704–14.
Pierry C, Caumont C, Blanchard E, Brochet C, Dournes G, Gros A, et al. Assessment of BRAF(V600E) mutation in pulmonary Langerhans cell histiocytosis in tissue biopsies and bronchoalveolar lavages by droplet digital polymerase chain reaction. Virchows Arch. 2018;472:247–58.
Villalba M, Exposito F, Pajares MJ, Sainz C, Redrado M, Remirez A, et al. TMPRSS4: a novel tumor prognostic indicator for the stratification of stage IA tumors and a liquid biopsy biomarker for NSCLC patients. J Clin Med. 2019;8:2134.
Roncarati R, Lupini L, Miotto E, Saccenti E, Mascetti S, Morandi L, et al. Molecular testing on bronchial washings for the diagnosis and predictive assessment of lung cancer. Mol Oncol. 2020;14:2163–75.
Lee SH, Kim EY, Kim T, Chang YS. Compared to plasma, bronchial washing fluid shows higher diagnostic yields for detecting EGFR-TKI sensitizing mutations by ddPCR in lung cancer. Respir Res. 2020;21:142.
Li X, Liu Y, Shi W, Xu H, Hu H, Dong Z, et al. Droplet digital PCR improved the EGFR mutation diagnosis with pleural fluid samples in non-small-cell lung cancer patients. Clin Chim Acta. 2017;471:177–84.
Li Y, Lv J, Wan S, Xin J, Xie T, Li T, et al. High sensitive and non-invasive ctDNAs sequencing facilitate clinical diagnosis and clinical guidance of non-small cell lung cancer patient: a time course study. Front Oncol. 2018;8:491.
Hummelink K, Muller M, Linders TC, van der Noort V, Nederlof PM, Baas P, et al. Cell-free DNA in the supernatant of pleural effusion can be used to detect driver and resistance mutations, and can guide tyrosine kinase inhibitor treatment decisions. ERJ Open Res. 2019;5:00016–2019.
Wu W, Cao Z, Zhang W, Zhang L, Hou L, Wu C. Comparison of the SuperARMS and ARMS for detecting EGFR mutations in liquid-based cytology specimens from NSCLC patients. Diagn Pathol. 2020;15:9.
García-Olmo D, Olmedillas-López S, Cortés-Guiral D, Villarejo P, López Rojo I, Guadalajara H, García Gómez-Heras S, García-Arranz M. The role of mucin cell-free DNA detection as a new marker for the study of acellular pseudomyxoma peritonei of appendicular origin by liquid biopsy. Ther Adv Med Oncol. 2020;12:1–12.
Schwartz M, Camacho-Vanegas O, Wood AM, Dashkoff M, Whitelock C, Harkins TT, et al. Applying precision medicine to ovarian cancer: proof-of-principle for a “Molecular Second Look.” Int J Gynecol Cancer. 2018;28:479–85.
Hata T, Mizuma M, Masuda K, Chiba K, Ishida M, Ohtsuka H, et al. MicroRNA-593-3p expression in peritoneal lavage fluid as a prognostic marker for pancreatic cancer patients undergoing staging laparoscopy. Ann Surg Oncol. 2021;28:2235–45.
Suenaga M, Fujii T, Yamada S, Hayashi M, Shinjo K, Takami H, et al. Peritoneal lavage tumor DNA as a novel biomarker for predicting peritoneal recurrence in pancreatic ductal adenocarcinoma. Ann Surg Oncol. 2021;28:2277–86.
Leick KM, Kazarian AG, Rajput M, Tomanek-Chalkley A, Miller A, Shrader HR, et al. Peritoneal cell-free tumor DNA as biomarker for peritoneal surface malignancies. Ann Surg Oncol. 2020;27:5065–71.
Biron VL, Matkin A, Kostiuk M, Williams J, Cote DW, Harris J, et al. Analytic and clinical validity of thyroid nodule mutational profiling using droplet digital polymerase chain reaction. J Otolaryngol Head Neck Surg. 2018;47:60.
Ye W, Hannigan B, Zalles S, Mehrotra M, Barkoh BA, Williams MD, et al. Centrifuged supernatants from FNA provide a liquid biopsy option for clinical next-generation sequencing of thyroid nodules. Cancer Cytopathol. 2019;127:146–60.
Xu X, Ma X, Zhang X, Cao G, Tang Y, Deng X, et al. Detection of BRAF V600E mutation in fine-needle aspiration fluid of papillary thyroid carcinoma by droplet digital PCR. Clin Chim Acta. 2019;491:91–6.
Guibert N, Tsukada H, Hwang DH, Chambers E, Cibas ES, Bale T, et al. Liquid biopsy of fine-needle aspiration supernatant for lung cancer genotyping. Lung Cancer. 2018;122:72–5.
Roy-Chowdhuri S, Mehrotra M, Bolivar AM, Kanagal-Shamanna R, Barkoh BA, Hannigan B, et al. Salvaging the supernatant: next generation cytopathology for solid tumor mutation profiling. Mod Pathol. 2018;31:1036–45.
Hannigan B, Ye W, Mehrotra M, Lam V, Bolivar A, Zalles S, et al. Liquid biopsy assay for lung carcinoma using centrifuged supernatants from fine-needle aspiration specimens. Ann Oncol. 2019;30:963–9.
Matsumoto K, Kato H, Nouso K, Ako S, Kinugasa H, Horiguchi S, et al. Evaluation of local recurrence of pancreatic cancer by kras mutation analysis using washes from endoscopic ultrasound-guided fine-needle aspiration. Dig Dis Sci. 2020;65:2907–13.
Suenaga M, Sadakari Y, Almario JA, Borges M, Lennon A-M, Shin E-J, et al. Using an endoscopic distal cap to collect pancreatic fluid from the ampulla (with video). Gastrointest Endosc. 2017;86:1152–56.e2.
Suenaga M, Dudley B, Karloski E, Borges M, Irene Canto M, Brand RE, et al. The effect of pancreatic juice collection time on the detection of KRAS mutations. Pancreas. 2018;47:35–9.
Driescher C, Fuchs K, Haeberle L, Goering W, Frohn L, Opitz FV, et al. Bile-based cell-free DNA analysis is a reliable diagnostic tool in pancreatobiliary cancer. Cancers (Basel). 2020;13:39.
Acknowledgments
The authors would like to thank Dr. María García-Puente for her collaboration and advice to build the literature search strategy. The authors gratefully acknowledge the editors and reviewers for their contribution to improve the quality of this article.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work is part of a grant from “FIS-ISCIII-FEDER” (Fondo de Investigaciones Sanitarias-Instituto de Salud Carlos III-Fondo Europeo de Desarrollo Regional [European Regional Development Fund]), Ministry of Health, Spain (grant number PI20/01052).
Conflicts of Interest
Susana Olmedillas-López, Rocío Olivera-Salazar, Mariano García-Arranz, and Damián García-Olmo have no conflicts of interest that are directly relevant to the content of this article.
Author Contributions
Susana Olmedillas-López and Rocío Olivera-Salazar performed the literature search. Susana Olmedillas-López drafted the manuscript. Rocío Olivera-Salazar prepared the figures. Rocío Olivera-Salazar, Mariano García-Arranz, and Damián García-Olmo critically revised the work. All authors agreed on the final version.
Availability of data and material
Not applicable.
Code availability
Not applicable.
Ethics approval
Not applicable.
Consent
Not applicable.
Rights and permissions
About this article
Cite this article
Olmedillas-López, S., Olivera-Salazar, R., García-Arranz, M. et al. Current and Emerging Applications of Droplet Digital PCR in Oncology: An Updated Review. Mol Diagn Ther 26, 61–87 (2022). https://doi.org/10.1007/s40291-021-00562-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-021-00562-2