Abstract
Introduction
Agricultural commodities contaminated by molds and mycotoxins can be considered as public health problems in less developed countries, particularly in Iran. Hence the main purpose of this study was to identify mold fungi and molecular analysis of the most important species of aflatoxin-B1-producing Aspergillus species in some dried nuts and grains in local markets in Tehran.
Materials and methods
Two hundred fifty samples of wheat, rice, corn, pistachios, and peanuts were collected from the five different locations of Tehran between January 2018 and January 2019. The samples were analyzed by using direct seed inoculation method and grain crushing method. Fungal strains were identified as Aspergillus spp. on the basis of morphological characters and further confirmed by using of β-tubulin gene sequencing. To differentiate between aflatoxigenic and non-aflatoxigenic Aspergillus spp., the isolates were screened for the presence of aflatoxigenic genes (nor-1, ver-1, omtA, and aflR).
Results
One-handed forty-eight aflatoxigenic Aspergillus isolates (144 A. flavus and 4 A. parasiticus) were identified and aflR gene was the most frequent gene in these species. Five isolates (4 A. flavus, 1 A. parasiticus) had quadruplet pattern, 64 isolates (63 A. flavus, 1 A. parasiticus) had more than 1 gene and 39 isolates (38 A. flavus,1 A. parasiticus) did not have any genes.
Conclusion
According to the contamination of dried nuts and grains by some aflatoxigenic fungi, an extensive surveillance is necessary to provide a wider view on these products. Moreover, effective and efficient aflatoxin control program requires identifying and managing key elements that are effective in reducing mycotoxin production at farm level or in storage conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In all over the world, especially in Iran, food gradients including wheat, corns, pistachio nuts, and peanuts are considered as the main nutrient sources [1]. These products have the high susceptibility to mycotoxin contamination which is a serious threat for animal and human health [2]. Mycotoxins can be produced by several species of molds, such as Fusarium, Penicillium and Aspergillus species [3]. Improper pre-harvest or post-harvest conditions of agricultural products are the key factors in the contamination with these fungi [4]. The percentage of cereals contaminated by mycotoxins has been estimated approximately 25 % around the world [5]. Aflatoxins are produced widely by Aspergillus flavus and Aspergillus parasiticus belonged to Aspergillus section Flavi [6]. There are reports from different parts of the world confirming that aflatoxicosis, as a food-borne disease, causes poisoning, sickness, and death in humans and animals [7]. Among different forms of aflatoxin (B1, B2, G1 and G2), International Agency for Research on Cancer (IARC) has considered aflatoxin B1 as the first group of human carcinogen [8]. Several structural genes in the aflatoxin gene cluster comprising nor-1, ver-1, and omtA encode enzymes of norsolorinic acid reductase converting norsolorinic acid to averantin, versicolorin A dehydrogenase converting versicolorin A to sterigmatocystin and sterigmatocystin-o methyltransferase converting dimethyl sterigmatocystin to sterigmatocystin, respectively. Furthermore, the aflR gene encodes a regulatory factor activating transcript of some structural genes in aflatoxin gene cluster [9]. Screening and identification of fungal isolates on the basis of macroscopic and microscopic characteristics are difficult, especially in the case of morphologically-similar species [10]. In recent years, although advantage of the phenotypic identification method is undeniable, molecular methods due to their rapidity, sensitivity and specificity are employed for identification of fungi. The present study was conducted to investigate the distribution pattern of aflatoxigenic Aspergillus species and their status regarding the presence of aflatoxigenic genes, in wheat, rice, corn, pistachio, and peanut in local markets of Tehran, Iran.
Material and method
Collection of samples
In this descriptive cross-sectional study, 250 samples comprising of wheat (n = 50), rice (n = 50), corn (n = 50), pistachios (n = 50), and peanuts (n = 50) were collected from the five different locations of Tehran (North, South, East, West, Center) between January 2018 and January 2019.
Cultivation, isolation, and identification
Collected samples were cultured by using two methods namely direct implantation and seed grinding on Sabouraud dextrose agar (SDA) plates. In the direct method, 3–5 out of each of samples were inoculated on the SDA containing antibiotic chloramphenicol. In the method of seed grinding, 1 g of grounded samples of wheat, rice, corn, pistachio, and peanut separately was mixed with 10 mL of sterile distilled water then 100 µl from prepared suspension was transferred to SDA containing chloramphenicol. Culture plates of two methods were incubated at 26 °C for 7 days. During this time, the growth of Aspergillus spp. was controlled daily. All strains were preliminarily identified to the species level based on morphological characteristics; subsequently, their identity was confirmed by molecular approaches.
DNA extraction
The isolation of chromosomal DNA from filamentous fungi was done according to the used technique of Yelton et al. with some modification [11]. The quality and quantity of DNA was analyzed by resolving the samples on gel electrophoresis [12].
Molecular identification
All A. flavus and A. parasiticus isolates were identified by microscopic and macroscopic characteristics; subsequently, their identity was confirmed by DNA sequencing of the partial β-tubulin (BTU) gene, using the primers Bt2a (5'-GGT AAC CAA ATC GGT GCT TTC-3') and Bt2b (5'- ACC CTC AGT GTA GTG ACC CTT GGC-3') [13]. A typical PCR mixture contained: 2 µL DNA template, 5 µL Taq polymerase buffer, 1.25 µL primer (120 pmol mL− 1 each), 25 µL H2O and 0.1 µL Taq polymerase (5 U µL− 1). Briefly, amplification of the BTU gene was performed with 2 min for initial denaturation in the start of one cycle at 94 °C, followed by 35 cycles at 94 °C for 60 s, 54 °C for 60 s, and 68 °C for 60 s, with a final 10-min extension at 68 °C. Sequence data obtained were compared with those deposited in the NBLAST algorithm (http://blast.ncbi.nlm.nih.gov) and the local database at the CBS-KNAW Fungal Biodiversity Centre (Utrecht, The Netherlands).
Multiplex PCR reaction
The sequences of the primers of aflR, nor-1, ver-1, and omtA genes are shown in Table 1 [14]. Each PCR reaction mixture with a total volume of 20 µL was contained 10 ng genomic DNA extracted from fungal isolates, 0.4 µL sense and 0.4 µL antisense primers of aflR, nor-1, ver-1, and omtA genes, 2x (dye mix) ready to use PCR master mix containing Taq DNA polymerase (Merck). The amplification of desired genomic regions was done by thermal cycler (Techne) according to the temperature-time program optimized including 6 min for initial denaturation in the start of one cycle at 95 °C, followed by 6 cycles of 45 s at 95˚C, 45 s at 58.2˚C and 80 s at 72˚C, then 30 cycles of 60 s at 95˚C, 60 s at 56˚C, 60 s at 72˚C. While final extension of 10 min, were given at 72˚C. Amplified DNA fragments were electrophoresed on 1.2 % agarose gel stained with ethidium bromide [15].
Results
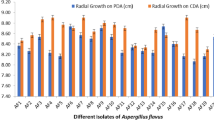
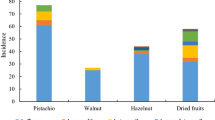
In general, 473 fungal isolates were recovered from the 250 collected samples (Table 2). From this fungi, 285 isolates were identified as Aspergillus spp., 148 of them were confirmed to be aflatoxigenic speices, i.e. A. flavus (144, 50.5 %), and A. parasiticus (4, 1.4 %) by molecular method. These strains isolated from 39 (26.35 %) corn, 36 (24.32 %) rice pieces, 29 (19.60 %) pistachio, 26 (17.57 %) peanut and 18 (12.16 %) wheat samples. Moreover, all A. parasiticus isolates were identified in wheat samples in center of Tehran. The DNA fragments amplified by using multiplex PCR corresponding to aflR, nor-1, ver-1, and omtA genes have given four bands of 400, 537, 797 and 1032 bp on gel electrophoresis (Fig. 1). Regarding the frequency of aflatoxin biosynthesis genes, aflR gene was detected in 86 (58 %) A. flavus and 3 (2 %) A. parasiticus isolates, nor-1 gene was detected in 43 (29 %) A. flavus and 1 (0.7 %) A. parasiticus isolates, ver-1 gene was detected in 32 (22 %) A. flavus and 1 (0.7 %) A. parasiticus isolates and omtA gene was detected in 30 (20 %) A. flavus and 1 (0.7 %) A. parasiticus isolates (Fig. 2). The investigation of gene patterns indicated that the presence of four genes of aflatoxin biosynthetic pathway was observed in five isolates (1 A. parasiticus and 4 A. flavus), 64 isolates (63 A. flavus, 1 A. parasiticus) had more than 1 gene and 39 isolates (38 A. flavus,1 A. parasiticus) did not have these four genes.
Discussion
Aflatoxins, known as secondary metabolites, are produced by some Aspergillus spp. These compounds are the public-health challenge due to their importance in mutagenicity, carcinogenicity on human and animals. The analysis of some agricultural foodstuffs in different production processes has indicated the presence of aflatoxins in these products leading to the consignment rejection of these products via consumers [16, 17]. In this investigation, we demonstrated that distribution pattern of mold fungi associated with them from five different areas of Tehran city. Our findings showed the highest and lowest prevalence was related to corn and wheat samples. By contrast, the report of Lutfullah et al. observation on some cereals and beans of Pakistan indicated that wheat samples had the highest prevalence of contamination. The cause of this difference may be inappropriate conditions of pre-harvest and post-harvest of crops [18]. In studies conducted in Sanandaj and Sari (Iran) on peanuts and pistachio nuts according to morphological profiles indicated that the A. flavus species was more common than other Aspergillus spp., confirming our results [19, 20]. While, in other studies in Isfahan, Rafsanjan and Kerman cites of Iran on pistachio samples, the most predominant Aspergillus spp. isolated were A. niger and A. flavus, respectively [21]. Also in the study of Furlung et al. the genus Aspergillus was not present among the fungi isolated from wheat in Brazil [22].
In order to reduce the risk of aflatoxin consumption, the isolation of agricultural commodities contaminated by aflatoxin- producing fungi is necessary. From this regard, because of remarkable merits of PCR-based methods such as simplicity, accuracy, sensitivity and rapidity, they are useful for the detection of four effective genes in aflatoxin biosynthesis pathway [23]. In the study of Criseo et al. in contrast to our study, the nor-1 gene was the most representative and the aflR gene had the lowest incidence among the four aflatoxin genes tested [24]. Studies conducted to evaluate the prevalence of four aflatoxin genes (aflR, nor-1, ver-1, and omtA) with the use of multiplex PCR method indicated that in non-aflatoxigenic A. flavus strains isolated from food and feed samples in Italy and A. flavus and A. parasiticus strains isolated from maize samples in Kenya, the nor-1 gene was known as the most prevalent gene [25]. Another study showed that omtA gene was the most representative gene in A. flavus and A, parasiticus isolates in Egyption chili [9, 15, 23]. According to previous investigations, although all aflatoxigenic strains had complete pattern of four genes, some strains with compete pattern of four tested genes due to molecular defects at levels of transcription and protein translation were non-aflatoxigenic strains [24]. From this regard, it is unable to take the presence of complete pattern into account a difference between aflatoxigenic and non-aflatoxigenic strains [15, 24].
In conclusion, according to the contamination of dried nuts and grains by some aflatoxigenic fungi, an extensive surveillance is necessary to control of these products. Multiplex PCR method can be used for screening and primary isolation when faced with large numbers of samples.
Change history
19 October 2021
A Correction to this paper has been published: https://doi.org/10.1007/s40201-021-00745-3
References
Gholami-Shabani M, Shams-Ghahfarokhi M, Razzaghi-Abyaneh M. Aflatoxins and aflatoxigenic fungi in Iran: A systematic review of the past, present, and future. Mycol Iran. 2017;4(2):65–84.
Shanakht H, Ali Shahid A, Waseem Ali S. Characterization of fungal microbiota on rice grains from local markets of Lahore. J Hyg Eng Des. 2014;9:35–40.
Gherbawy YA, Elhariry HM, Bahobial AAS. Mycobiota and mycotoxins (aflatoxins and ochratoxin) associated with some Saudi date palm fruits. Foodborne Pathog Dis. 2012;9(6):561–7.
Schmidt-Heydt M, Rüfer CE, Abdel-Hadi A, Magan N, Geisen R. The production of aflatoxin B1 or G1 by Aspergillus parasiticus at various combinations of temperature and water activity is related to the ratio of aflS to aflR expression. Mycotoxin Res. 2010;26(4):241–6.
Iram W, Anjum T, Iqbal M, Ghaffar A, Abbas M. Structural eluci-dation and toxicity assessment of degraded products of aflatoxin B1and B2 by aqueous extracts of Trachyspermum ammi. Front Microbiol. 2016;7:346.
Akinola SA, Ateba CN, Mwanza M. Polyphasic assessment of aflatoxin production potential in selected aspergilli. Toxins (Basel). 2019;11(12):692.
Hamzeh Pour S, Mahmoudi S, Masoumi S, Rezaie S, Barac A, Ranjbaran M, et al. Aflatoxin M1 contamination level in Iranian milk and dairy products: A systematic review and meta-analysis. World Mycotoxin J. 2020;13;67–82. Wageningen Academic Publishers.
Sardiñas N, Vázquez C, Gil-Serna J, González-Jaén MT, Patiño B. Specific detection and quantification of Aspergillus flavus and Aspergillus parasiticus in wheat flour by SYBR® Green quantitative PCR. Int J Food Microbiol. 2011;145(1):121–5.
Chebon S, Wanyoike W, Bii C, Gathumbi J, Ogoyi D. Prevalence of aflatoxin biosynthesis genes according to aflatoxin levels in maize of different varieties in Kenya. Biotechnol J Int. 2017;19(2):1–21.
Hedayati MT, Khodavaisy S, Alialy M, Omran SM, Habibi MR. Invasive aspergillosis in intensive care unit patients in Iran. Acta Medica (Hradec Kralove). 2013;56(2):52-6
Yelton MM, Hamer JE, Timberlake WE. Transformation of Aspergillus nidulans by using a trpC plasmid. Proc Natl Acad Sci U S A. 1984;81:1470–1474.
Hussain A, Afzal A, Irfan M, Malik KA. Molecular detection of aflatoxin producing strains of aspergillus flavus from peanut (Arachis Hypogaea). Turkish J Agric - Food Sci Technol. 2015;3(5):335.
Khodavaisy S, Badali H, Hashemi S.J, et al. In vitro activities of five antifungal agents against 199 clinical and environmental isolates of Aspergillus flavus, an opportunistic fungal pathogen. Journal de Mycologie Médicale. 2016;26:116–21.
Degola F, Berni E, Dall’Asta C, et al. A multiplex RT-PCR approach to detect aflatoxigenic strains of Aspergillus flavus. J Appl Microbiol. 2007;103(2):409–17.
Gherbawy YA, Shebany YM, Hussein MA, Maghraby TA. Molecular detection of mycobiota and aflatoxin contamination of chili. Arch Biol Sci. 2015;67(1):223–34.
Manonmani HK, Anand S, Chandrashekar A, Rati ER. Detection of aflatoxigenic fungi in selected food commodities by PCR. 2005;40:2859–64.
Price MS, Yu J, Nierman WC, Kim HS, Pritchard B, Jacobus CA, et al. The aflatoxin pathway regulator AflR induces gene transcription inside and outside of the aflatoxin biosynthetic cluster. FEMS Microbiol Lett. 2006;255(2):275-9.
Lutfullah G, Hussain A. Studies on contamination level of aflatoxins in some cereals and beans of Pakistan. Food Control [Internet]. 2012;23(1):32–6. Available from: https://doi.org/10.1016/j.foodcont.2011.06.004.
Hedayati MT, Kaboli S, Mayahi S. Mycoflora of pistachio and peanut kernels from sari, iran. Jundishapur J Microbiol. 2010;3:114–20.
Khodavaisy S, Maleki A, Hossainzade B, Rezai S, Ahmadi F. Occurrence of fungal contamination in pistachio and peanut samples from retail shops in Sanandaj province, 2012;6(39):6781–4.
Rahimi P, Sharifnabi B, Bahar M. Detection of aflatoxin in Aspergillus species isolated from pistachio in Iran. J Phytopathol. 2008;156(1):15–20.
Furlung EB, Soares LM, Lasca CC, Kohara EY. Mycotoxins and fungi in wheat harvested during 1990 in test plots in the state of Sao Paulo, Brazil. Mycopathologia. 1995;131(3):185–90.
Criseo G, Bagnara A, Bisignano G. Differentiation of aflatoxin-producing and non-producing strains of Aspergillus flavus group. Lett Appl Microbiol. 2001;33(4):291–5.
Criseo G, Racco C, Romeo O. High genetic variability in non-aflatoxigenic A. flavus strains by using Quadruplex PCR-based assay. Int J Food Microbiol. 2008;125(3):341–3.
Okoth S, De Boevre M, Vidal A, Di Mavungu JD, Landschoot S, Kyallo M, et al. Genetic and toxigenic variability within Aspergillus flavus population isolated from maize in two diverse environments in Kenya. Front Microbiol. 2018;9(JAN):57.
Acknowledgements
This study was part of a M.S. thesis supported by Tehran University of Medical Sciences (grant No. 9411399003). We are grateful to the colleagues of Molecular Biology Laboratory of Medical Mycology, Tehran University of Medical Sciences (Mohammad Reza Safari and Azar Berahmeh) for their assistance in conducting this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest between authors in this study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: "The original article has been updated"
Rights and permissions
About this article
Cite this article
Rahimi, A., Sasani, E., Rezaie, S. et al. Molecular identification of aflatoxigenic Aspergillus species in dried nuts and grains collected from Tehran, Iran. J Environ Health Sci Engineer 19, 1795–1799 (2021). https://doi.org/10.1007/s40201-021-00734-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40201-021-00734-6