Abstract
Skeletal muscle pathology is thought to have an important role in the onset and/or progression of amyotrophic lateral sclerosis (ALS), which is a neurodegenerative disorder characterized by progressive muscle weakness. Since miRNAs are recognized as important regulatory factors of essential biological processes, we aimed to identify differentially expressed miRNAs in the skeletal muscle of sporadic ALS patients through the combination of molecular-omic technologies and bioinformatic tools. We analyzed the miRnome profiles of skeletal muscle biopsies acquired from ten sALS patients and five controls with Affymetrix GeneChip miRNA 4.0 Array. To find out differentially expressed miRNAs in patients, data were analyzed by The Institute for Genomic Research-Multi Experiment Viewer (MeV) and miRNAs whose expression difference were statistically significant were identified as candidates. The potential target genes of these miRNAs were predicted by miRWalk 2.0 and were functionally enriched by gene ontology (GO) analysis. The expression level of priority candidates was validated by quantitative real-time PCR (qRT-PCR) analysis. We identified ten differentially expressed miRNAs in patients with a fold change threshold ≥ 2.0, FDR = 0. We identified ten differentially expressed miRNAs in patients with a fold change threshold ≥ 2.0, FDR = 0. Nine out of the ten miRNAs were found to be related to top three enriched ALS-related terms. Based on the qRT-PCR validation of candidate miRNAs, patients were separated into two groups: those with upregulated miR-4429 and miR-1825 expression and those with downregulated miR-638 expression. The different muscle-specific miRNA profiles in sALS patients may indicate the involvement of etiologic heterogeneity, which may allow the development of novel therapeutic strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amyotrophic lateral sclerosis (ALS) is a progressive and fatal neuromuscular disorder, particularly affecting especially upper and lower motor neurons. It is the most common motor neuron disease in adults with a worldwide prevalence of 4–6 per 100,000. The majority of cases of ALS are sporadic (90–95%). The remaining cases (5–10%) exhibit a Mendelian pattern of inheritance, usually inherited in an autosomal dominant manner. Both forms are thought to have common pathogenic mechanisms as they share similar clinical and pathological features [1].
The disease causes progressive and severe muscle weakness as muscle function is largely controlled by motor neurons. It is known that skeletal muscle is not only the end-organ that is affected by motor neuron degeneration but also contributes actively to the onset and/or progression of the disease [2]. It has been suggested that skeletal muscle pathology plays a role in the pathogenesis of ALS by activating a retrograde signaling cascade that degrades motor neurons [3]. It is also commonly accepted that neuromuscular junction (NMJ) dismantlement is one of the earliest pathogenic signature occurred before disease symptoms and independently of motor neuron degeneration [4]. Both before the clinical onset and during the disease progression, ineffective cycles of reinnervation and denervation are observed during motor neuron degeneration in the skeletal muscle of ALS patients [2]. Therefore, understanding the molecular and pathological changes in skeletal muscle fibers can lead to the identification of the pathogenic mechanism of the disease and can be exploited to develop new treatment strategies.
MicroRNAs (miRNAs) are short, single-stranded endogenous non-coding RNA molecules that play an essential role in mediating RNA-interference through epigenetic regulation of gene expression [5, 6]. The ability of miRNAs to fine-tune the activity of biological pathways supports their potential as therapeutic targets for the treatment of various diseases [7]. In this study, we presented comprehensive and large-scale data with regard to dysregulated miRNAs in skeletal muscle tissue of sALS patients.
Materials and methods
Patients
All participants provided written informed consent to participate in this study and ethical approval was granted by the Hacettepe University Faculty of Medicine Ethical Review Board (GO 16/552-37). Skeletal muscle biopsies from the left biceps brachii muscle were obtained from all six control subjects (age range 22–54 years, mean age 33.2 years) and ten definite ALS patients (age range 26–59 years, mean age 42.8 years) according to Revised El-Escorial criteria with no family history of ALS (sporadic ALS, sALS). Muscle biopsies with no diagnostic pathology as indicated by normal histomorphology were used as controls. Tissue specimens were rapidly frozen in isopentane cooled in liquid nitrogen and stored at −80 °C until further use. Standard histological and histochemical techniques were applied to all cryostat tissue sections, including haematoxylin and eosin (H&E), modified Gomori trichrome, ATP (pH: 9.4), nicotinamide adenine dinucleotide (NADH), succinate dehydrogenase (SDH), periodic acid–Schiff (PAS), Oil-red-O, and myophosphorylase.
Total RNA extraction from skeletal muscle biopsies of sALS patients and controls
Total RNA was extracted from the skeletal muscle biopsies (~ 30 mg) using miRCURY RNA Isolation Kit—Cell & Plant (Qiagen, Valencia, California, USA), according to the manufacturer’s instructions. Quantity and integrity of total RNA were determined using a NanoDrop ND-1000 spectrophotometer (Thermo Scientific, Waltham, MA, USA) and agarose gel electrophoresis, respectively.
miRNA microarray analysis
Differential analysis of the miRNAs was performed using Affymetrix GeneChip miRNA array technology version 4.0 (Thermo Fisher Scientific, Santa Clara, CA, USA), according to the manufacturer's directions on total RNA extracted from skeletal muscle biopsies of ten sALS patients and six controls. The array contains probe sets for 2578 mature human miRNAs. Total RNA (300 ng) including miRNAs was biotin-labeled using the FlashTagTM Biotin HSR RNA Labeling kit (Affymetrix, Genisphere, Hatfield, PA, USA) and the samples were then hybridized overnight using the GeneChip Hybridization Oven 640 (Affymetrix, Santa Clara, CA, USA) at 48 °C. The arrays were then washed and stained in the GeneChip Fluidics Station 450 (Affymetrix, Santa Clara, CA, USA). The arrays were scanned using a GeneChip Scanner 3000 7G (Affymetrix, Santa Clara, CA, USA) and the signal values were evaluated using the Expression Console Software (EC) v1.2 (Affymetrix by Thermo Fisher Scientific). Intensity values (presence/absence values) and signal histograms of each hybridization were quality checked (Supplementary Fig. 1). To identify the differentially expressed miRNAs between sALS patients and controls, acquired data were analyzed using Multi Experiment Viewer (MeV v4.9.0; The Institute for Genomic Research). miRNAs whose differential expression was statistically significant [with a fold change ≥ 2.0 and false discovery rate (FDR) = 0 for MeV-SAM analysis] were identified as the potential miRNA candidates.
miRNA target prediction and functional annotation/gene enrichment analysis of miRNA targets
The potential target genes of candidate miRNAs were investigated in-silico using the miRWalk 2.0 database, a comprehensive atlas of predicted and validated miRNA-target interactions [8, 9]. Subsequently, using the EnrichR tool, Gene ontology (GO) enrichment analysis was performed to gain insights into the biological functions of all the target genes of each candidate miRNA based on biological processes [10, 11].
Quantitative real-time PCR (qRT-PCR) validation of candidate miRNAs
The expression of differentially expressed miRNA candidates in sALS patients was determined by qRT-PCR. Reverse transcription was carried out using the miRNA-specific stem-loop RT primers (TaqMan™ MicroRNA Assays, Thermo Fisher Scientific, Waltham, MA, USA) and the TaqMan® MicroRNA Reverse Transcription Kit (Thermo Fisher Scientific, Waltham, MA, USA). qRT-PCR was carried out with TaqMan probes (TaqMan® MicroRNA Assays, Thermo Fisher Scientific, Waltham, MA, USA), TaqMan® Universal PCR Master Mix, and no AmpErase® UNG (Thermo Fisher Scientific, Waltham, MA, USA). Amplifications were performed using Bio-Rad IQ5 Real-time PCR Detection Systems (Bio-Rad, Hercules, CA, USA). The 2−ΔΔCt method was then used to calculate the relative expression levels of candidate miRNAs. All reactions were performed in triplicate for each sample, and as with other ALS-related studies [12,13,14] the U6 snRNA was used for normalization of miRNA RT-qPCR data.
Sorting out ALS-related target genes and the evaluation of miRNA-binding sites
ALS-related target genes were sorted out from the target gene lists of three differentially expressed miRNAs using the Venny 2.1.0 tool. Subsequently, validated ALS-related target genes were examined in MirTarBase, an experimentally validated microRNA-target interactions database and binding sites of ALS-related target genes were identified in TargetScan 7.2, a tool for sequence-based miRNA-target prediction [15], which has classified target complementary base pairing with the miRNA seed region as 8mer, 7mer-m8, 7mer-A1, and 6mer according to efficiency and the conservation level among species. For the expression analysis of ALS-related target genes, total RNA was used as a template for cDNA synthesis using iScript™ cDNA Synthesis Kit ((Bio-Rad Lab, Hercules, CA, USA). qRT-PCR was carried out by iTaq Universal SYBR Green Supermix using the iQ5 Real-Time PCR Detection System (Bio-Rad Lab, Hercules, CA, USA). The reactions were run in triplicate. The PCR cycling conditions were as follows: 2 min at 95 °C; followed by 40 cycles of 95 °C for 10 s, and 56 °C for 20 s. Post-amplification melt-curve analysis was run after each reaction. Relative quantities of UBQLN2 and FUS mRNAs were calculated using the 2−ΔΔCt method after normalization to the GAPDH. The experiments were repeated three times.
Statistical analysis
Significance analysis of microarrays (SAM) was applied to identify the differentially expressed miRNAs in the MeV program. For the qRT-PCR experiments, fold changes in the expression level of miRNAs in MD groups were evaluated by Mann–Whitney U test with the p value of < 0.05 being considered significant. Statistical analyses were performed using the GraphPad Prism 5.0 software (GraphPad Software, Inc., San Diego, CA).
Results
Clinical evaluation of patients
Muscle biopsies from ten sALS patients (five females and five males; age range 26–59 years, mean age 42.8 years) and six controls (three females and three males; age range 27–56 years, mean age 44.1 years) were included in the study. The average duration of the disease was 1.3 (0.5–4) years while the average ALS functional rating scale was 38.8 (27–46). The clinical and histopathological findings of sALS patients are listed in Table 1.
Microarray-based miRNA expression profile analysis revealed ten differentially expressed miRNAs in sALS patients
miRNA profiling was performed by miRNA microarray analysis of skeletal muscle biopsies of sALS patients and controls. Since the concentration of RNA extracted from Control 3 was below the threshold (61.8 ng/µl), it was not included in our study. Background correction, data normalization, and the computation of the probe-set level expression were performed using the Expression Console Software v.1.2 (Affymetrix) with Robust Multichip Average (RMA) and Detection Above Background (DABG) algorithms. After comparing all patients with all controls, ten miRNAs were found to be dysregulated in ALS patients (Table 2). Nine of ten miRNAs (miR-933, miR-191-3p, miR-4310, miR-4750-5p, miR-1825, miR-371b-5p, miR-638, miR-940, and miR-572) as well as a small nucleolar RNA, U48, were upregulated in patients. However, miR-4429 was downregulated in ALS patients. Raw data of our microarray experiment can be accessed through ArrayExpress database at EMBL-EBI (www.ebi.ac.uk/arrayexpress) under accession number E-MTAB-9140.
Potential target genes of the ten dysregulated miRNAs were associated with ALS-related terms
We obtained potential target gene lists of the dysregulated miRNAs from miRWalk 2.0. Subsequently, Gene Ontology (GO) enrichment analysis was performed according to the biological process domain on target genes of candidate miRNAs. All dysregulated miRNAs were involved in disease associated pathways; however, only nine of them, excluding miR-572, were associated with the top three enriched ALS-related terms. As shown in Figs. 1 and 2, the top three enriched GO terms were mainly related to the regulation of transcription [positive (GO 0045893) and negative (GO 0045892)], DNA-templated regulation of transcription from RNA polymerase II promoter [positive (GO 0045944) and negative (GO 0000122)], and regulation of gene expression [positive (GO 0010628) and negative (GO 0010629)]. The other ALS-related biological processes are ubiquitin-dependent protein catabolic process (GO 0006511), intrinsic apoptotic signaling pathway (GO 0097193), skeletal muscle tissue development (GO 0007519), dendrite morphogenesis (GO 0048813), regulation of cytoskeleton organization (GO 0051493), and positive regulation of lipid storage (GO 0010884).
Gene ontology (GO) term enrichment analysis of the target genes of differentially expressed six miRNAs (miR-191-3p, miR-371b-5p, miR-572, miR-638, miR-933, and miR-940). Analysis was performed within the ‘Biological Process’ category. Bar graphs are sorted by p value (p < 0.05). The longer bars and lighter colored bars mean that the term/gene-set is more significant
Gene ontology (GO) term enrichment analysis of the target genes of differentially expressed four miRNAs (miR-1825, miR-4310, miR-4429, and miR-4750-5p). Analysis was performed within the ‘Biological Process’ category. Bar graphs are sorted by p value (p < 0.05). The longer bars and lighter colored bars mean that the term/gene-set is more significant
qRT-PCR analysis revealed three candidate miRNAs were differentially expressed in sALS patients
All ten candidate miRNAs were possibly related to ALS-related biological processes. Since miR-572 was not associated with the top three GO enriched processes and a miRNA assay for miR-940 was not available, they were excluded from validation experiments. Validation experiments of the remaining eight miRNAs showed no significant difference between all ten patients and controls. However, three out of the eight miRNAs (miR-1825, miR-4429 and miR-638) were differentially expressed in half of the sALS patients. These results divided patients into two groups. Among miRNAs that were differentially expressed, miR-4429 and miR-1825 were significantly upregulated in five of the sALS patients (ALS1-ALS5), while miR-638 was downregulated in the other half of the sALS patients (ALS6-ALS10) (Fig. 3).
qRT-PCR validation of miR-1825, miR-638, and miR-4429 expression levels. miRNA scores were derived from differential expression analyses comparing all sALS patients to controls (left panels) or half of the patients versus controls (right panels). The relative expression of miRNAs was normalized to U6 snRNA, using the 2−∆∆CT method. Data are expressed as means ± SEM. Statistical significance was determined using the Mann–Whitney U test, *p < 0.05, **p < 0.01
All three dysregulated miRNAs may target ALS-related genes that have at least one miRNA-binding sites
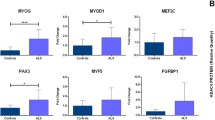
After investigation of potential target genes in miRWalk 2.0 database, we aimed to identify miRNA-binding sites of ALS-related target genes. We checked for the presence of validated/predicted target genes that are associated with ALS using possible target gene lists of candidate miRNAs and found that there were no disease-related target genes validated by strong experimental evidence and all three miRNAs (miR-1825, miR-638, miR-4429) can target predicted ALS-related genes. As seen in Table 3, miRNA-binding site analysis of ALS-related target genes using TargetScan database revealed that all target genes have at least one miRNA-binding site in their 3’UTR. The main results of our study are summarized in the form of a flowchart representation in Fig. 4. After analysing miRNA-binding sites of ALS-related target genes, expression analysis of the most potential target genes, UBQLN2 and FUS, was performed. A statistically significant downregulation of UBQLN2 (p = 0.0212) and FUS (p = 0.0079) was detected in half of the patients (ALS1-ALS5) who showed increased miR-1825 and miR-4429 expression (Fig. 5).
qRT-PCR validation of UBQLN2 and FUS expression levels. The left panel shows differential expression analysis comparing all sALS patients to controls (left panels) or half of the patients versus controls (right panels). The relative expression of mRNAs was normalized to GAPDH, using the 2−∆∆CT method. Data are expressed as mean ± SEM. Statistical significance was determined using the Mann–Whitney U test, *p < 0.05, **p < 0.01
Discussion
In this study we evaluated the differential expression levels of skeletal muscle miRNAs in sALS patients and controls. A growing number of scientific studies have revealed important roles for miRNAs including muscle-specific miRNAs (myomiRs) in skeletal muscle function such as, growth, regeneration, and metabolism [16, 17]. miRNAs are recognized as important regulatory factors in different diseases, and large-scale microarray analysis has provided evidence of the dysregulation of miRNAs in a variety of neuromuscular diseases, such as Duchenne muscular dystrophy (DMD), Facioscapulohumeral muscular dystrophy (FSHD), and Limb-girdle muscular dystrophies (LGMDs) [18]. Also, dysregulated miRNAs in skeletal muscle tissue of sALS patients were investigated in some studies, most of which were focusing only on myomiRs [13, 19, 20]. Two high-throughput studies identified differentially expressed miRNAs in the skeletal muscle of sALS patients. However, one study was performed with a small sample size using an array system containing low miRNA coverage, and the other was completed without any validation experiment and pathway analysis [21, 22]. On the contrary, our study is a more detailed study with a comprehensive miRNA array coverage, sufficient sample size, validation experiments, and pathway analysis.
We identified one downregulated (miR-4429) and 9 upregulated miRNAs (miR-933, miR-191-3p, miR-4310, miR-4750-5p, miR-1825, miR-371b-5p, miR-638, miR-940, miR-572) in skeletal muscle of sALS patients. To understand the putative roles of these miRNAs, we identified potential targets using bioinformatics approaches. According to GO term enrichment analysis, the target genes of these miRNAs, except for miR-572, were mainly enriched in the top three enriched ALS-related terms, such as DNA-templated regulation of transcription, regulation of transcription from RNA polymerase II promoter, and regulation of gene expression. Consistent with our findings, gene expression regulation at the post-transcriptional level has also been reported as an ALS-related term [12, 23]. Terms apart from the most enriched top three pathways (ubiquitin-dependent protein catabolic process, intrinsic apoptotic signaling pathway, skeletal muscle tissue development, and regulation of cytoskeleton organization) have also been associated with ALS in previous studies [24,25,26,27]. These GO terms are apparently not specific to ALS, and can also be related to other neuromuscular diseases. This may promote the opportunity for the identification of novel and common therapeutic options for many different neuromuscular diseases that are associated with these terms.
Candidate miRNAs were selected for further verification using qRT-PCR, and the expression levels of miR-1825, miR-638, and miR-4429 were statistically significant in sALS patients compared to controls. The discrepancy between the miRNA microarray and qRT-PCR results might be due to the difference in data normalization methods. Also, different results can be obtained on different experimental platforms, due to the technological differences [28]. These three differentially expressed miRNAs have the potential to target the 3′UTR of some of the ALS-related genes that play an important role in the ubiquitin–proteasome system (UPS), ribostasis, regulation of cytoskeleton, and vesicular trafficking [29]. Interestingly, miR-1825 and miR-4429 were found to be upregulated in half of our patients (ALS1-ALS5), whereas miR-638 was downregulated in the other half (ALS6-ALS10). However, we could not find difference between the two groups in terms of age, sex, clinical features, such as disease severity or progression, pathological findings. Further, previous studies have shown that both miR-1825 and miR-638 are involved in sALS. Contrary to our findings, the expression level of miR-1825, which causes motor axon defects by targeting tubulin-folding cofactor b (TBCB), was reduced in plasma and post-mortem skeletal muscle tissue of sALS patients [14, 30]. In that study, the mean age of the sALS patients whose post-mortem tissues were used was 63.2 ± 9.5. Consequently, different results in our study could be because miRNA profiles of skeletal muscle or serum may change with age, and distinct expression patterns of miRNAs can be detected in different biological samples [31,32,33]. Moreover, miRNAs can regulate different cellular pathways by targeting different genes. Therefore, we suggest that the upregulation of miR-1825 causes a defect in proteostasis by targeting the Ubiquilin 2 (UBQLN2) gene from three sites in the 3’UTR. In our study, potential target gene expression analysis revealed that there was statistically significant downregulation of UBQLN2 (p = 0.0366) in half of the patients (ALS1-ALS5), who showed increased miR-1825 expression (data not shown). This result will allow us to examine the validity of this target gene of miR-1825.
Interestingly, our study determined that miR-638 was downregulated in skeletal muscle of sALS patients. Similarly, miR-638 was shown to be downregulated in leukocytes of sALS patients in one study [12], and in another study it was significantly decreased in serum of patients with familial ALS and pre-symptomatic ALS mutation carriers [34].
There is no prior study yet on a direct association between sALS and miR-4429. Therefore, our study has revealed miR-4429 as a promising candidate for further exploration. It was previously associated with different medical conditions, such as stroke and biliary atresia [35, 36]. According to the miRNA-binding site analysis performed in the TargetScan database, miR-4429 has the potential to target the fused in sarcoma (FUS) gene, which encodes a DNA/RNA binding protein that plays roles in various cellular processes, such as transcription regulation, RNA splicing, and RNA transport [37,38,39,40]. Therefore, FUS expression was analyzed in our patient group and statistically significant downregulation of FUS gene was observed in half of the sALS patients (ALS1-ALS5) with increased miR-4429 expression. This observation also supports the division of our patients into two groups (as ALS1-ALS5 and ALS6-ALS10) in qRT-PCR experiments.
As mentioned above, variations in qRT-PCR analysis indicated that the skeletal muscle miRNA profile was very heterogeneous among our patients and the results of our study clustered patients into two groups. However, we did not find a significant relationship between miRNA expression profiles and age, sex or clinical/histopathological findings of the patients, as well as array hybridization levels. This could be explained by molecular heterogeneity of sALS. Moreover, undefined mutations in sALS patients may contribute to a heterogeneous miRNA profile. Similar results were also obtained in both miRNA and gene expression profiling studies related to sALS. The miRNA expression profiles are heterogeneous within patients and they are generally divided into two groups [22, 24, 34]. Our patient group seems heterogeneous but compared to other studies reported in the literature the biopsies of our patients were obtained from the same muscle. In addition, when we examined similar studies in the literature, we found that the patient age ranges, ALSFRS-R scores were wider and also disease symptoms and duration were more heterogeneous than our study. They also included both female and male patients [13, 20, 22]. In addition, different miRNAs have been found to be involved in disease pathogenesis in various studies [12, 21, 22, 41]. Although few studies have investigated their functions [14], identification of muscle-specific miRNA profiles associated with sALS will shed light on the etiopathogenesis of the disease.
Apart from differential miRNA expression, microarray and qPCR results showed that RNU48, a small nucleolar RNA, was upregulated in sALS patients. This upregulation was also reported in a small RNA profiling of sALS patients. This upregulation was also reported in a small RNA profiling of patients with sALS [22]. The differential expression of U48, which is used as a normalizer in many studies [42, 43], is also an indicator of perturbed RNA metabolism in ALS pathogenesis.
Despite intensive research efforts, the molecular pathogenic mechanism of ALS and the underlying causes of motor neuron degeneration have not been fully elucidated yet. Various pathogenic mechanisms, such as environmental factors, oxidative stress, impaired axonal transport, aggregation of ubiquitinated proteins, mitochondrial dysfunction, and altered RNA metabolism, have been described as potential contributors to neurodegeneration and ALS progression, but none have proven to be causative [44,45,46,47,48,49]. In this descriptive study, we identified a muscle-specific miRNA profile associated with the etiopathogenesis of sALS. These three miRNAs that were identified as potential candidates might be the most important factors related to skeletal muscle damage observed in the pathogenesis of sALS. Further exploring the effect of these common miRNAs on disease-related pathways will provide an opportunity for the identification of novel therapeutic options. Our ongoing in vitro functional analysis studies are being carried out with these miRNAs to identify the most affected pathways and target genes of these miRNAs.
Availability of data and material
Raw data of our microarray experiment can be accessed through ArrayExpress database at EMBL-EBI (www.ebi.ac.uk/arrayexpress) under accession number E-MTAB-9140.
Code availability
Not applicable.
References
Grad LI, Rouleau GA, Ravits J, Cashman NR (2017) Clinical spectrum of amyotrophic lateral sclerosis (ALS). Cold Spring Harb Perspect Med. https://doi.org/10.1101/cshperspect.a024117
Loeffler JP, Picchiarelli G, Dupuis L, Gonzalez De Aguilar JL (2016) The role of skeletal muscle in amyotrophic lateral sclerosis. Brain Pathol 26(2):227–236
Fitzsimonds RM, Poo MM (1998) Retrograde signaling in the development and modification of synapses. Physiol Rev 78(1):143–170
Moloney EB, de Winter F, Verhaagen J (2014) ALS as a distal axonopathy: molecular mechanisms affecting neuromuscular junction stability in the presymptomatic stages of the disease. Front Neurosci 8:252
He L, Hannon GJ (2004) MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5(7):522–531
Wu L, Fan J, Belasco JG (2006) MicroRNAs direct rapid deadenylation of mRNA. Proc Natl Acad Sci USA 103(11):4034–4039
Jackson A, Linsley PS (2010) The therapeutic potential of microRNA modulation. Discov Med 9(47):311–318
Dweep H, Gretz N (2015) miRWalk2.0: a comprehensive atlas of microRNA-target interactions. Nat Methods 12(8):697
Dweep H, Sticht C, Pandey P, Gretz N (2011) miRWalk–database: prediction of possible miRNA binding sites by “walking” the genes of three genomes. J Biomed Inform 44(5):839–847
Chen EY, Tan CM, Kou Y, Duan Q, Wang Z, Meirelles GV, Clark NR, Ma’ayan A (2013) Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform 14:128
Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, Koplev S, Jenkins SL, Jagodnik KM, Lachmann A et al (2016) Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res 44(W1):W90-97
De Felice B, Guida M, Guida M, Coppola C, De Mieri G, Cotrufo R (2012) A miRNA signature in leukocytes from sporadic amyotrophic lateral sclerosis. Gene 508(1):35–40
Di Pietro L, Baranzini M, Berardinelli MG, Lattanzi W, Monforte M, Tasca G, Conte A, Logroscino G, Michetti F, Ricci E et al (2017) Potential therapeutic targets for ALS: MIR206, MIR208b and MIR499 are modulated during disease progression in the skeletal muscle of patients. Sci Rep 7(1):9538
Helferich AM, Brockmann SJ, Reinders J, Deshpande D, Holzmann K, Brenner D, Andersen PM, Petri S, Thal DR, Michaelis J et al (2018) Dysregulation of a novel miR-1825/TBCB/TUBA4A pathway in sporadic and familial ALS. Cell Mol Life Sci 75(23):4301–4319
Agarwal V, Bell GW, Nam JW, Bartel DP (2015) Predicting effective microRNA target sites in mammalian mRNAs. Elife. https://doi.org/10.7554/eLife.05005
Wang J, Yang LZ, Zhang JS, Gong JX, Wang YH, Zhang CL, Chen H, Fang XT (2018) Effects of microRNAs on skeletal muscle development. Gene 668:107–113
Zhang S, Chen N (2018) Regulatory role of microRNAs in muscle atrophy during exercise intervention. Int J Mol Sci 19(2):405
Alexander MS, Kunkel LM (2015) Skeletal muscle MicroRNAs: their diagnostic and therapeutic potential in human muscle diseases. J Neuromuscul Dis 2(1):1–11
Jensen L, Jorgensen LH, Bech RD, Frandsen U, Schroder HD (2016) Skeletal muscle remodelling as a function of disease progression in amyotrophic lateral sclerosis. Biomed Res Int 2016:5930621
Russell AP, Wada S, Vergani L, Hock MB, Lamon S, Leger B, Ushida T, Cartoni R, Wadley GD, Hespel P et al (2013) Disruption of skeletal muscle mitochondrial network genes and miRNAs in amyotrophic lateral sclerosis. Neurobiol Dis 49:107–117
de Andrade HM, de Albuquerque M, Avansini SH, de Cistiane RS, Dogini DB, Nucci A, Carvalho B, Lopes-Cendes I, França MC (2016) MicroRNAs-424 and 206 are potential prognostic markers in spinal onset amyotrophic lateral sclerosis. J Neurol Sci 368:19–24
Kovanda A, Leonardis L, Zidar J, Koritnik B, Dolenc-Groselj L, Ristic Kovacic S, Curk T, Rogelj B (2018) Differential expression of microRNAs and other small RNAs in muscle tissue of patients with ALS and healthy age-matched controls. Sci Rep 8(1):5609
Raman R, Allen SP, Goodall EF, Kramer S, Ponger LL, Heath PR, Milo M, Hollinger HC, Walsh T, Highley JR et al (2015) Gene expression signatures in motor neurone disease fibroblasts reveal dysregulation of metabolism, hypoxia-response and RNA processing functions. Neuropathol Appl Neurobiol 41(2):201–226
Aronica E, Baas F, Iyer A, ten Asbroek ALMA, Morello G, Cavallaro S (2015) Molecular classification of amyotrophic lateral sclerosis by unsupervised clustering of gene expression in motor cortex. Neurobiol Dis 74:359–376
Baloh RH, Rakowicz W, Gardner R, Pestronk A (2007) Frequent atrophic groups with mixed-type myofibers is distinctive to motor neuron syndromes. Muscle Nerve 36(1):107–110
Brockington A, Ning K, Heath PR, Wood E, Kirby J, Fusi N, Lawrence N, Wharton SB, Ince PG, Shaw PJ (2013) Unravelling the enigma of selective vulnerability in neurodegeneration: motor neurons resistant to degeneration in ALS show distinct gene expression characteristics and decreased susceptibility to excitotoxicity. Acta Neuropathol 125(1):95–109
Guégan C, Vila M, Rosoklija G, Hays AP, Przedborski S (2001) Recruitment of the mitochondrial-dependent apoptotic pathway in amyotrophic lateral sclerosis. J Neurosci 21(17):6569–6576
Morey JS, Ryan JC, Van Dolah FM (2006) Microarray validation: factors influencing correlation between oligonucleotide microarrays and real-time PCR. Biol Proc Online 8:175–193
Chia R, Chiò A, Traynor BJ (2018) Novel genes associated with amyotrophic lateral sclerosis: diagnostic and clinical implications. Lancet Neurol 17(1):94–102
Takahashi I, Hama Y, Matsushima M, Hirotani M, Kano T, Hohzen H, Yabe I, Utsumi J, Sasaki H (2015) Identification of plasma microRNAs as a biomarker of sporadic amyotrophic lateral sclerosis. Mol Brain 8(1):67
Mercken EM, Majounie E, Ding J, Guo R, Kim J, Bernier M, Mattison J, Cookson MR, Gorospe M, de Cabo R et al (2013) Age-associated miRNA alterations in skeletal muscle from rhesus monkeys reversed by caloric restriction. Aging (Albany NY) 5(9):692–703
Noren Hooten N, Fitzpatrick M, Wood WH 3rd, De S, Ejiogu N, Zhang Y, Mattison JA, Becker KG, Zonderman AB, Evans MK (2013) Age-related changes in microRNA levels in serum. Aging (Albany NY) 5(10):725–740
Soriano-Arroquia A, House L, Tregilgas L, Canty-Laird E, Goljanek-Whysall K (2016) The functional consequences of age-related changes in microRNA expression in skeletal muscle. Biogerontology 17(3):641–654
Freischmidt A, Muller K, Zondler L, Weydt P, Mayer B, von Arnim CA, Hubers A, Dorst J, Otto M, Holzmann K et al (2015) Serum microRNAs in sporadic amyotrophic lateral sclerosis. Neurobiol Aging 36(9):2660.e2615–2620
Dong R, Shen Z, Zheng C, Chen G, Zheng S (2016) Serum microRNA microarray analysis identifies miR-4429 and miR-4689 are potential diagnostic biomarkers for biliary atresia. Sci Rep 6(1):21084
Jickling GC, Ander BP, Zhan X, Noblett D, Stamova B, Liu D (2014) microRNA expression in peripheral blood cells following acute ischemic stroke and their predicted gene targets. PLoS ONE 9(6):e99283
Crozat A, Aman P, Mandahl N, Ron D (1993) Fusion of CHOP to a novel RNA-binding protein in human myxoid liposarcoma. Nature 363(6430):640–644
Perrotti D, Bonatti S, Trotta R, Martinez R, Skorski T, Salomoni P, Grassilli E, Lozzo RV, Cooper DR, Calabretta B (1998) TLS/FUS, a pro-oncogene involved in multiple chromosomal translocations, is a novel regulator of BCR/ABL-mediated leukemogenesis. Embo J 17(15):4442–4455
Yang L, Embree LJ, Tsai S, Hickstein DD (1998) Oncoprotein TLS interacts with serine-arginine proteins involved in RNA splicing. J Biol Chem 273(43):27761–27764
Zinszner H, Sok J, Immanuel D, Yin Y, Ron D (1997) TLS (FUS) binds RNA in vivo and engages in nucleo-cytoplasmic shuttling. J Cell Sci 110(Pt 15):1741–1750
Freischmidt A, Muller K, Zondler L, Weydt P, Volk AE, Bozic AL, Walter M, Bonin M, Mayer B, von Arnim CA et al (2014) Serum microRNAs in patients with genetic amyotrophic lateral sclerosis and pre-manifest mutation carriers. Brain 137(Pt 11):2938–2950
Akkaya-Ulum YZ, Balci-Peynircioglu B, Karadag O, Eroglu FK, Kalyoncu U, Kiraz S, Ertenli AI, Özen S, Yilmaz E (2017) Alteration of the microRNA expression profile in familial Mediterranean fever patients. Clin Exp Rheumatol 108(6):90–94
Han IB, Kim M, Lee SH, Kim JK, Kim SH, Chang JH, Teng YD (2016) Down-regulation of MicroRNA-126 in glioblastoma and its correlation with patient prognosis: a pilot study. Anticancer Res 36(12):6691–6697
Barber SC, Shaw PJ (2010) Oxidative stress in ALS: key role in motor neuron injury and therapeutic target. Free Radic Biol Med 48(5):629–641
Blokhuis AM, Groen EJ, Koppers M, van den Berg LH, Pasterkamp RJ (2013) Protein aggregation in amyotrophic lateral sclerosis. Acta Neuropathol 125(6):777–794
Bogaert E, d’Ydewalle C, Van Den Bosch L (2010) Amyotrophic lateral sclerosis and excitotoxicity: from pathological mechanism to therapeutic target. CNS Neurol Disord Drug Targets 9(3):297–304
Cozzolino M, Carri MT (2012) Mitochondrial dysfunction in ALS. Prog Neurobiol 97(2):54–66
Fischer-Hayes LR, Brotherton T, Glass JD (2013) Axonal degeneration in the peripheral nervous system: implications for the pathogenesis of amyotrophic lateral sclerosis. Exp Neurol 246:6–13
Ling SC, Polymenidou M, Cleveland DW (2013) Converging mechanisms in ALS and FTD: disrupted RNA and protein homeostasis. Neuron 79(3):416–438
Acknowledgements
Advanced English Language editing service was supplied by Hacettepe Teknokent Technology Transfer Center (HT-TTM).
Funding
No funding was received for conducting this study.
Author information
Authors and Affiliations
Contributions
EA performed experiments and bioinformatics analysis, analyzed the data, and drafted the manuscript. BB designed and conceptualized the study, analyzed the data; interpreted the results, and drafted the manuscript. CEB performed clinical classification, characterization and follow-up of patients with sALS; interpreted the results. ATA performed qRT-PCR experiments. SE performed clinical classification, characterization and follow-up of patients with sALS; revised the manuscript for intellectual content. ET performed clinical classification, characterization and follow-up of patients with sALS; revised the manuscript for intellectual content.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
This study was performed in line with the principles of the Declaration of Helsinki. Ethical approval was granted by the Hacettepe University Faculty of Medicine Ethical Review Board (GO 16/552-37).
Consent to participate
All participants provided written informed consent to participate in this study.
Consent for publication
Patients signed informed consent regarding publishing their data.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13760_2021_1743_MOESM1_ESM.tif
Supplementary Fig 1. Quality control of miRNA array. (A)Histogram plot of the signal values for the all probe sets. (B) Principal component analysis (PCA) of probe cell intensity for all samples (TIF 2517 KB)
Rights and permissions
About this article
Cite this article
Aksu-Menges, E., Balci-Hayta, B., Bekircan-Kurt, C.E. et al. Two distinct skeletal muscle microRNA signatures revealing the complex mechanism of sporadic ALS. Acta Neurol Belg 122, 1499–1509 (2022). https://doi.org/10.1007/s13760-021-01743-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13760-021-01743-w