Abstract
We aimed to explore the cellular action of micro-RNAs that are non-coding-RNAs modulating gene expression, whose expression is dysregulated in myotonic dystrophy (DM1). Basic procedure was to measure the levels of muscle-specific myo-miRNAs (miR-1, miR-133a/b, miR-206) in muscle of 12 DM1 patients. Muscle fiber morphometry and a new grading of histopathological severity score were used to compare specific myo-miRNA level and fiber atrophy. We found that the levels of miR-1 and miR-133a/b were significantly decreased, while miR-206 was significantly increased as compared to controls. The histopathological score did not significantly correlate with the levels of myo-miRNAs, even if the lowest levels of miRNA-1 and miRNA-133a/b, and the highest levels of miRNA-206 were observed in patients with either severe histopathological scores or long disease duration. The histopathological score was inversely correlated with disease duration. Nowadays that DM1 muscle biopsies are scanty, since patients are usually diagnosed by genetic analysis, our study offers a unique opportunity to present miRNA expression profiles in muscle and correlate them to muscle morphology in this rare multisystem disorder. Our molecular and morphologic data suggest a post-transcriptional regulatory action of myo-miRNA in DM1, highlighting their potential role as biomarkers of muscle plasticity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Myotonic dystrophy type 1 (DM1) is a dominant disorder affecting mostly skeletal and cardiac muscle. The most common clinical signs are myotonia, distal muscle weakness and atrophy. DM1 is caused by an expansion of (CTG)n repeats within the 3′ non-coding region of the DMPK gene. The mutant RNA, which contains 50 to several hundreds CTG repeats, is retained in nuclear aggregates that sequester the muscleblind-like-1 RNA-binding protein, which is essential for mRNA splicing, leading to a toxic effect on many functional processes in the nucleus.
Micro-RNAs (miRNAs), which are short non-coding RNA molecules, have been identified in various physiological and pathological conditions, in affected tissues (e.g., skeletal and cardiac muscle) and in body fluids [1]. MiRNAs exert a complex network of negative gene regulation by targeting messenger RNAs for translational repression or cleavage [1, 2], so that upregulation of a specific miRNA leads to a reduced expression of the corresponding protein product.
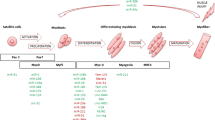
A subset of about 25 skeletal and cardiac muscle-specific miRNAs (myo-miRNAs) regulate muscle function and adaptation during development (proliferation, differentiation, quiescence, regeneration) and disease [3, 4]. Muscle gene expression is regulated by a transcriptional network, involving serum response factor (SRF), myocyte enhancer factor-2 (MEF2), myogenic regulatory factor differentiation-1 (MyoD), Myf5, and myogenin. SRF and MEF2 cooperate with MyoD and myogenin to transcriptionally activate the expression of myo-miRNAs [1, 5].
Upregulation of miRNA-1 and miRNA-206 reduces myoblasts proliferation, increases myogenic differentiation and regeneration of muscle fibers [6–8]. Inhibition of miRNA-1 and miRNA-206 leads to delayed myogenesis. MiRNA-1 is a crucial regulator of gap junction proteins and muscle cells excitability (channels) and its deregulation has been suggested to contribute to the heart conduction defects seen in DM1 patients [9, 10]. MiRNA-133a/b increase myoblasts proliferation, promote cell growth, and decrease differentiation [8, 11].
In DM1, abnormal levels of myo-miRNAs have been found in muscle [5, 12, 13] but their action has not been investigated in relation to muscle pathology.
The aim of this study was to measure the levels of myo-miRNAs in muscle and compare the results with morphometric data of muscle fiber atrophy in biopsies.
Materials and methods
Muscle biopsy
A diagnostic vastus lateralis muscle biopsy was obtained after written consent and agreement from the local ethical committee. Muscle specimens were frozen in isopentane chilled in liquid nitrogen, and then stored at −80 °C until used.
Muscle morphometry and morphology grading
We assigned a histopathological score to each biopsy according to the degree of changes observed by microscopic inspection, as follows (Fig. 1): score 1 = increased fiber size variability, increased internal nuclei; score 2 = as in score 1 and atrophic fibers, splitting of fibers; score 3 = as in score 2 and nuclear clumps, fibrosis, ring fibers, degenerating fibers.
Different extent of muscle histopathological changes observed in DM1 patients. Histopathological score 1 a increased fiber size variability, occasional internal nuclei. Score 2 b atrophic fibers (arrow). Score 3 c interstitial connective tissue increase, nuclear clumps, atrophic fibers. The lower panel shows a correlation between the histopathological severity score and years of disease duration (Rank Spearman test: ρ = 0.68, p < 0.02)
Morphometric analysis was done on haematoxylin–eosin-stained sections, used to digitalize 3–5 non-overlapping random microscope fields. ImageJ software (v.1.34) was used to outline and measure fiber cross-sectional area and diameter of fibers. We also calculated the “fiber atrophy and hypertrophy factors” [14], which express the proportion of abnormally small or large fibers in the section. Briefly, the fibers in normal adult muscle are 40–80 μm diameter in males and 30–70 μm in females. Few fibers in the range of 30–40 μm would have less significance than the same number of fibers in the range of 10–20 μm. This is taken into account by multiplying the number of fibers with a diameter between 30 and 40 μm by 1, the number of fibers with a diameter between 20 and 30 by 2, the number of those from 10 to 20 by 3, and the number in the group <10 μm by 4. These products are then added together and divided by the total number of fibers. The resulting number is then multiplied by 1000 to obtain the “atrophy factor”.
In a subset of six patients in whom acid and alkaline ATP-ase-stained sections were available for the study, we measured also the morphometric parameters in type 1 and type 2 fibers.
miRNAs profiling and validation
miRNAs were purified from muscle using miRNeasy mini kit (Qiagen, Hilden, Germany). Reverse transcription was performed using TaqMan micro-RNA reverse transcription kit (Applied Biosystems, Carlsbad, CA, USA). The resulting cDNA was amplified by real-time PCR using TaqMan micro-RNA assay primers (hsa-miRNA-1, hsa-miRNA-133a, hsa-miRNA-133b, hsa-miRNA-206) with the TaqMan universal mastermix-II and analyzed with an ABI PRISM 7000 Sequence Detector System. Primer sequences are available on request. After median Ct values normalization, relative miRNA expression was calculated with the 2−ΔΔCt method. The relative levels of miRNA expression were normalized to the housekeeping hsa-let-7a. TaqMan-PCR experiments were performed in triplicate. Values in DM1 patients are expressed as fold-change of controls.
Statistical analysis
Values in patients and controls were analyzed using two-tailed Student’s t tests and the Spearman rank correlation test. The level of significance was set at p < 0.05.
Results
Patients
The study population consisted of 12 DM1 patients (11 males, 1 female) in whom the clinical diagnosis of DM1 was confirmed by genetic testing (Table 1). Additional 8 DM1 patients have been investigated for muscle fiber morphometry in both type 1 and type 2 fibers (Supplementary Table 1), but these samples have not been used for miRNA analysis since no enough tissue was available for this investigation.
The age of patients at biopsy ranged from 19 to 52 years (mean 32.2). The age at onset of symptoms ranged from 3 to 48 years, and the disease duration (time elapsed from onset to biopsy) ranged from 1 to 24 years (Table 1). As controls we used the vastus lateralis muscle biopsy obtained from 6 age-matched male subjects (aged 18–38 years, mean 29.3) who showed no signs of neuromuscular disorders and have been considered healthy controls.
Muscle morphometry and histopathology
In controls, the fiber diameter ranged from 48 to 64 μm (mean 55), the fiber cross-sectional area ranged from 2968 to 5118 μm2 (mean 3792), the atrophy factor ranged from 0 to 114 units (mean 43), and the hypertrophy factor ranged from 0 to 61 units (mean 12).
Histopathological analysis of muscle showed a variable degree of pathological features, including increased number of internal nuclei, increased fiber size variability with atrophic fibers and nuclear clumps, and proliferation of connective tissue (fibrosis).
Morphometric analysis showed that five male DM1 patients had a mild reduction of the mean fiber diameter (<50 μm), and increased values of the atrophy factor (260–727 units) (Table 2). All of these five cases were classified as severe or intermediate histopathological scores (score 3 or 2), as the result of chronic muscle changes which are possibly due to long disease duration (>20 years) in two cases (Fig. 1). One male patient had a mean fiber diameter below the normal range (37 μm) and the highest value of atrophy factor (829 units), indicating both a generalized fiber hypotrophy and scattered fiber atrophy, despite a short disease duration and mild histopathological score (score 1). The remaining male patients had normal values of fiber size and atrophy factor.
Increased values of hypertrophy factors were found in three male patients, two of whom had normal atrophy factor and 1 had mildly increased atrophy factor.
In the six cases in whom a morphometric analysis of type 1 and type 2 fibers was done, a severe atrophy of type 1 fibers was found in two cases who also showed a moderate atrophy of type 2 fibers. Hypertrophy factor was never increased in type 1 fibers, while it was increased in type 2 fibers in 2 cases (Table 2). Type 1 fiber atrophy was also found in 6 of 8 additional patients (Supplementary Table 1), confirming this histopathological hallmark of the disease.
Although the histopathological score was not correlated with morphometric parameters of fiber size, it was inversely correlated (ρ = 0.68, p < 0.02) with disease duration (Fig. 1): more severe score (score 3) was observed in four patients who had either a long disease duration (>20 years) and/or a large CTG expansion, indicating that this qualitative score offers further information than the data provided by quantitative parameters of fiber size.
miRNA levels in muscle, comparison with histopathological features
In this study we investigated the expression in DM1 muscle of the four myo-miRNAs which are the most frequently analyzed in human muscle disease: we found a significant decrease of miRNA-1, miRNA-133a, and miRNA-133b, while the levels of miRNA-206 were significantly higher in patients than controls (Fig. 2).
Histograms showing myo-miRNA expression levels in muscle of DM1 patients. The values are expressed as fold-change of control mean (set at 1): as compared to controls, miRNA-1 and miRNA-133a were significantly decreased; miRNA-133b was not significantly reduced and miRNA-206 was significantly elevated. The asterisks show a significant difference (p < 0.05). Individual values in the 12 DM1 patients are shown in the panels on the right, showing individual variability probably related to different age of patients and different muscle adaptation to the disease
Although the lowest levels of miRNA-1 and miRNA-133a and miRNA-133b, and the highest levels of miRNA-206 were observed in patients with either severe histopathological score or long disease duration, the histopathological score did not significantly correlate with the levels of myo-miRNAs (Fig. 2; Table 2).
Discussion
Muscle myo-miRNAs and fiber morphometry
The predominant change in muscle biopsies in the majority of our series of DM1 patients was a mild fiber atrophy involving several scattered fibers.
We observed a reduced expression of miRNA-1, miRNA-133a, miRNA-133b in the muscle of DM1 patients, confirming previous results obtained on muscle tissues of DM1 patients [9, 13, 15]; other studies on DM1 muscles have shown either no changes in miRNA-133a/b [5, 12] or an overexpression of miRNA-1 [5]. This discrepancy with our data may be explained by the different muscles analyzed, the different method for normalization of results, the number of CTG repeats, or the age of patients.
MiRNA-1 is considered to be a member of the group of “degenerative miRNAs”, which may be critical mediators of cell death, contributing to apoptotic/necrotic myofiber loss [13]. The levels of miRNA-1 and miRNA-133 in DM1 muscle should, therefore, be markers of the residual muscle mass [16]; their levels were found to be reduced by 10% in the muscle of healthy subjects after 7 days of bed rest, reflecting the metabolic and molecular response of muscle to lack of exercise [17].
On the other hand, we found an overexpression of miRNA-206 in DM1 muscle, as previously observed in the muscle of both DM1 [12] and DMD patients [18], and in activated satellite cells and proliferating myoblasts. MiRNA-206 belongs to the group of “regenerative miRNAs” [18], and its levels in muscle can be used to establish regeneration potential [19]. MiRNA-206 has also been implicated in promoting regeneration of neuromuscular synapses and preventing muscle atrophy.
An interesting issue arising from our results is the disease specificity of the abnormal myo-miRNA expression found in DM1 patients. A comparison of myo-miRNA plasma levels in DM1 patients and in patients affected with a series of different muscular dystrophies (i.e., DMD, BMD, LGMD, FSHD1) showed that myo-miRNA were significantly increased in progressive muscular dystrophies but not in DM1 [20], and their increase was suggested to be possibly related to an active regeneration process occurring in progressive muscular dystrophies, while it is almost absent in DM1.
Muscle weakness and wasting are the main clinical features of DM1, and although they are progressive over time, the rate of muscle deterioration varies between patients. Disease severity can be assessed by physical examination, but the progression of muscle wasting can only be monitored by muscle imaging (CT, MRI). Myo-miRNA levels appear to be reduced when muscle wasting is stabilized and increased in muscle atrophy [21].
Pathological involvement and dysregulation of myo-miRNA seem to be related to muscle atrophy that could differently affect the slow or fast fiber types. Since it is well known that type 1 fibers are selectively affected in DM1, it is likely that such changes are due not only to activity but also to other cellular processes.
The originality of our study is to investigate these regulatory molecules and compare them with histopathological features. Nowadays, DM1 muscle biopsies are scanty, since most patients are diagnosed by genetic analysis; therefore, our study offers a unique opportunity to present these data.
We demonstrated an altered expression of myo-miRNAs in DM1 patients. These data highlight the value of myo-miRNAs regulation in DM1 and investigate muscle adaptation and pathological features in this multisystem disorder. The original approach used in this study might be prospectively used in a number of different neuromuscular patients to monitor both disease progression and effectiveness of new drugs or novel rehabilitative treatments.
References
Chen JF, Mandel EM, Thomson JM, Wu Q, Callis TE, Hammond SM, Conlon FL, Wang DZ (2006) The role of microRNAs-1 and microRNAs-133 in skeletal muscle proliferation and differentiation. Nat Genet 38:228–233
He L, Hannon GJ (2004) MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5:522–531
Eisenberg I, Alexander MS, Kunkel LM (2009) MiRNAs in normal and diseased skeletal muscle. J Cell Mol Med 13:2–11
Horak M, Novak J, Bienertova-Vasku J (2016) Muscle-specific microRNAs in skeletal muscle development. Dev Biol 410:1–13
Perbellini R, Greco S, Sarra-Ferraris G, Cardani R, Capogrossi MC, Meola G, Martelli F (2011) Dysregulation and cellular mislocalization of specific miRNAs in myotonic dystrophy type 1. Neuromuscul Disord 21:81–88
McCarthy JJ (2008) MicroRNA-206: the skeletal muscle-specific myo-miR. Biochim Biophys Acta 1779:682–691
Kim HK, Lee YS, Sivaprasad U, Malhotra A, Dutta A (2006) Muscle-specific microRNAs miR-206 promotes muscle differentiation. J Cell Biol 174:677–687
Wang XH (2013) MicroRNA in myogenesis and muscle atrophy. Curr Opin Clin Nutr Metab Care 16:258–266
Rau F, Freyermuth F, Fugier C, Villemin JP, Fischer MC, Jost B, Dembele D, Gourdon G, Nicole A, Duboc D, Wahbi K, Day JW, Fujimura H, Takahashi MP, Auboeuf D, Dreumont N, Furling D, Charlet-Berguerand N (2011) Misregulation of miR-1 processing is associated with heart defects in myotonic dystrophy. Nat Struct Mol Biol 18:840–846
Zhao Y, Ransom JF, Li A, Vedantham V, von Drehle M, Muth AN, Tsuchihashi T, McManus MT, Schwartz RJ, Srivastava D (2007) Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell 129:303–317
Mitchelson KR, Qin WY (2015) Roles of the canonical myo-miRs miR-1, -133 and -206 in cell development and disease. World J Biol Chem 6:162–208
Gambardella S, Rinaldi F, Lepore SM, Viola A, Loro E, Angelini C, Vergani L, Novelli G, Botta A (2010) Over-expression of microRNAs-206 in the skeletal muscle from myotonic dystrophy type 1 patients. J Transl Med 8:48
Fernandez-Costa JM, Garcia-Lopez A, Zuñiga S, Fernandez-Pedrosa V, Felipo-Benavent A, Mata M, Jaka O, Hernandez-Torres F, Aguado B, Perez-Alonso M, Vilchez JJ, Lopez de munain A, Artero RD (2013) Expanded CTG repeats trigger miRNA alterations in Drosophila that are conserved in myotonic dystrophy type 1 patients. Hum Mol Genet 22:704–716
Dubowitz V, Sewry C (2007) Normal muscle. In: Dubowitz V, Sewry C (eds) Muscle biopsy. A practical approach, 3rd edn. Saunders, London, pp 41–74
Kalsotra A, Singh RK, Gurha P, Ward AJ, Creighton CJ, Cooper TA (2014) The Mef2 transcription network is disrupted in myotonic dystrophy heart tissue, dramatically altering miRNA and mRNA expression. Cell Rep 6:336–345
Zaharieva IT, Calissano M, Scoto M, Preston M, Cirak S, Feng L, Collins J, Kole R, Guglieri M, Straub V, Bushby K, Ferlini A, Morgan JE, Muntoni F (2013) Dystromirs as serum biomarkers for monitoring the disease severity in Duchenne muscular dystrophy. PLoS One 8:e80263
Ringholm S, Bienso RS, Kiilerich K, Guadalupe-Grau A, Aachmann-Andersen NJ, Saltin B, Plomgaard P, Lundby C, Wojtaszewski JF, Calbet JA, Pilegaard H (2011) Bed rest reduces metabolic protein content and abolishes exercise-induced mRNA responses in human skeletal muscle. Am J Physiol Endocrinol Metab 301:E649–E658
Greco S, De Simone M, Colussi C, Zaccagnini G, Fasanaro P, Pescatori M, Cardani R, Perbellini R, Isaia E, Sale P, Meola G, Capogrossi MC, Gaetano C, Martelli F (2009) Common micro-RNA signature in skeletal muscle damage and regeneration induced by Duchenne muscular dystrophy and acute ischemia. FASEB J 23:3335–3346
Cacchiarelli D, Legnini I, Martone J, Cazzella V, D’Amico A, Bertini E, Bozzoni I (2011) MiRNAs as serum biomarkers for Duchenne muscular dystrophy. EMBO Mol Med 3:258–265
Matsuzaka Y, Kishi S, Aoki Y, Komaki H, Oya Y, Takeda S, Hashido K (2014) Three novel serum biomarkers, miR-1, miR-133a, and miR-206 for limb girdle muscular dystrophy, facioscapulohumeral muscular dystrophy, and Becker muscular dystrophy. Environ Health Prev Med 19:452–458
Koutsoulidou A, Kyriakides TC, Papadimas GK, Christou Y, Kararizou E, Papanicolaou EZ, Phylactou LA (2015) Elevated muscle-specific miRNAs in serum of myotonic dystrophy patients relate to muscle disease progress. PLoS One 10:e0125341
Acknowledgements
We acknowledge the Telethon Genetic Biobank Network and the EuroBioBank Network for providing muscle tissues.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors’ conflict of interest: none declared.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fritegotto, C., Ferrati, C., Pegoraro, V. et al. Micro-RNA expression in muscle and fiber morphometry in myotonic dystrophy type 1. Neurol Sci 38, 619–625 (2017). https://doi.org/10.1007/s10072-017-2811-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10072-017-2811-2