Abstract
Introduction
Cardiotocography (CTG) consists of two biophysical signals that are fetal heart rate (FHR) and uterine contraction (UC). In this research area, the computerized systems are usually utilized to provide more objective and repeatable results.
Materials and Methods
Feature selection algorithms are of great importance regarding the computerized systems to not only reduce the dimension of feature set but also to reveal the most relevant features without losing too much information. In this paper, three filters and two wrappers feature selection methods and machine learning models, which are artificial neural network (ANN), k-nearest neighbor (kNN), decision tree (DT), and support vector machine (SVM), are evaluated on a high dimensional feature set obtained from an open-access CTU-UHB intrapartum CTG database. The signals are divided into two classes as normal and hypoxic considering umbilical artery pH value (pH < 7.20) measured after delivery. A comprehensive diagnostic feature set forming the features obtained from morphological, linear, nonlinear, time–frequency and image-based time–frequency domains is generated first. Then, combinations of the feature selection algorithms and machine learning models are evaluated to achieve the most effective features as well as high classification performance.
Results
The experimental results show that it is possible to achieve better classification performance using lower dimensional feature set that comprises of more related features, instead of the high-dimensional feature set. The most informative feature subset was generated by considering the frequency of selection of the features by feature selection algorithms. As a result, the most efficient results were produced by selected only 12 relevant features instead of a full feature set consisting of 30 diagnostic indices and SVM model. Sensitivity and specificity were achieved as 77.40% and 93.86%, respectively.
Conclusion
Consequently, the evaluation of multiple feature selection algorithms resulted in achieving the best results.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Delivery is a critical event having various risky conditions for the fetus and takes a short time when compared to pregnancy period [1]. The undesired and stressful events for the fetus such as hypoxia and asphyxia frequently occur during the labor due to generally the lack of oxygen [2]. Fetus equipped with defense mechanisms struggles against these developments throughout the pregnancy and more importantly throughout labor [3]. At this point, cardiotocography (CTG) consisting of fetal heart rate (FHR) and uterine contraction (UC) signals recorded as simultaneously is the most prevalent used diagnostic technique to enable both determining distress level and continuous monitoring of the fetus [4]. This surveillance technique has been commonly adopted because of the sense of security it provides to observers [5]. Although the usage of CTG is great common, a gold standard has not been still accepted to evaluate the CTG traces. CTG has led to several debates about the increased rate of cesarean sections as well as high inter- and even intra-observer variability [6, 7].
Several parameters of FHR known as morphological features are the reliable and prominent indices to ascertain whether the fetal condition is well-being. More specifically, the baseline level of FHR, its variability both in short and long terms and its temporal transients considering as accelerations and decelerations are the primary indicators regarding the clinical assessment [8]. The various guidelines, such as the International Federation of Gynecology and Obstetrics (FIGO) [6] have been published by different health organizations to provide identifying the morphological features and a consistent CTG interpretation. In fact, the main aim of the guidelines is to decrease the variability among observers while preventing unnecessary interventions as possible. Despite the existing guidelines, unfortunately, the disagreement level among clinicians has remained stable [10]. The possible approaches to tackle these drawbacks were discussed in [11]. As a result, the usage of computerized CTG systems by supporting the decision process of observers has been pointed the most promising solution.
Automated CTG analysis requires several basic steps which are achieving the database, preprocessing, feature extraction, feature selection, and classification [12]. The morphological features mentioned above are extended with diagnostic features that are obtained from linear and nonlinear [13], time–frequency [14,15,16], and recently image-based time–frequency (IBTF) [12, 17,18,19] domains in computerized CTG analysis [20, 21]. Furthermore, FHR signals are classified using numerous machine learning techniques such as artificial neural networks (ANNs) [22, 23], extreme learning machine (ELM) [17, 24], and support vector machine (SVM) [25,26,27]. Feature extraction algorithms are utilized to improve the performance of classifiers and to propose clinically applicable models. For these particular purposes, genetic algorithms (GAs) [28], principal component analysis (PCA) [29], information gain (InfoGain), group of adaptive models evolution (GAME) neural network [30], correlation-based (CFS), Relief, Mutual Information (MI) [31] feature selection methods have been employed.
In this study, the combinations of five feature selection algorithms and machine learning algorithms, which are artificial neural network (ANN), k-nearest neighbor (kNN), decision tree (DT), and support vector machine (SVM), are evaluated on CTG data. To this end, weighted by support vector machine (WSVM), information gain ratio (IGR), relief, backward elimination (BE), and recursive feature elimination (RFE) algorithms are examined. Lastly, the commonly selected features by the related algorithms are used to generate the most relevant feature subset.
Materials and methods
Data description
An open-access intrapartum CTG database was introduced in 2012 [32] and it can be downloaded from Physionet. The database consists of 552 recordings, and these recordings are a subset of 9164 intrapartum CTG recordings that were acquired by the means of STAN S21/S31 and Avalon FM40/FM50 electronic fetal monitoring (EFM) devices. All signals were selected carefully considering the several technical and clinical criteria. Furthermore, the signals were stored in electronic form using OBTraceVue® system.
We use umbilical artery pH value obtained after delivery to separate the signals as normal and hypoxic. It is observed that different values of pH have been used as a borderline for separating FHR signals [30]. 177 recording with umbilical artery pH < 7.20 were considered hypoxic. The rest of the signals have umbilical artery pH ≥ 7.20 and thus were considered as normal.
Signal preprocessing
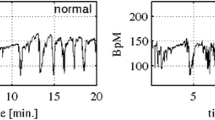
FHR signals can be acquired using either Doppler ultrasound or scalp electrode. In both cases, the signals are contaminated by several factors such as mother and fetal movements, displacement of the transducer and network interference as well. Segment selection, outlier detection, and interpolation are the basic procedures in preprocessing. The experimental study is performed on the signals which last 15 min (3600 sample points due to the 4 Hz sampling frequency). Extreme values (≥ 200 bpm and ≤ 50 bpm) are interpolated, and the long gaps (> 15 s) are not included in the subsequent feature extraction process. FHR signals are detrended using second-order polynomial in the last step of preprocessing since nonlinear signal processing techniques are utilized. After the preprocessing, we achieve 15 min duration segments of the signals that are as close as possible to the labor. Figure 1 demonstrates the state of sample recording before and after the preprocessing. Figure 1 comprises of small squares and large rectangles. Each small square corresponds to 30 s on the horizontal axis and 10 bpm on the vertical axis whereas each large rectangle takes up 3 min on the horizontal axis and 30 bpm on the vertical axis.
Feature transform (feature extraction and selection)
The features used in this study to identify FHR recordings are obtained from an open-access software that is used to analyze CTG recordings called CTG-OAS [29].
The morphological features describing the shape and changes of FHR signals are extracted firstly in accordance with FIGO guidelines. The baseline and the numbers of transient changes called accelerations (ACC) and decelerations (DCC) are taken into consideration [9].
Then, the morphological features are supported using several linear features such as mean (µ), standard deviation (σ), long-term irregularity (LTI), delta, short-term variability (STV), and interval index (II) [22, 33].
The third category of the features is the nonlinear domain. Approximation Entropy (ApEn), Sample Entropy (SampEn) and Lempel–Ziv Complexity (LZC) are the most commonly used features from this domain [34]. Two parameters pairs (embedding dimension, m = 2 with tolerance r = {0.15; 0.20}) are utilized individually in the experiment for ApEn and SampEn.
In the last category of the features is IBTF features involving contrast, correlation, energy, and homogeneity. IBTF features are obtained using a combination of Short-Time Fourier Transform (STFT) and Gray Level Co-occurrence Matrix (GLCM) [19]. GLCM is a directional pattern counter, and IBTF features are extracted according to angle and distance parameters. Distance (δ) and angle (θ) parameters are set to 1 and 90°, respectively. The spectrograms of very low frequency (VLF, 0–0.03 Hz), low frequency (LF, 0.03–0.15 Hz), middle frequency (MF, 0.15–0.50 Hz) and high frequency (HF, 0.50–1 Hz) are used to achieve IBTF features.
At the end of feature extraction stage, a total of 30 features coming from 3 morphological, 6 linear, 5 nonlinear and 16 IBTF (4 features for each specified frequency bandwidths) domains are extracted considering their origins.
In order to determine the most relevant features and to generate an effective subset, we utilize three filters and two wrappers methods. The commonly selected features by the algorithms are added to the final most relevant feature subset, and thus the effective subset is generated.
Artificial neural network (ANN)
ANN is a computational model inspired by the human brain and nervous system [35]. In the ANN architecture, an input layer, one more hidden layer(s) and an output layer are used [25]. Each node in the layers has a connection with the nodes in the subsequent layer, and this connection is represented with the weights [36]. An output of a layer for ANN is represented as follows:
where σ is the activation function, N is the number of input neurons. ωij and b represent the weights and bias value. Levenberg–Marquardt backpropagation algorithm and only one hidden layer with 30 nodes were used in the configuration of ANN. The other parameters were used with their default values.
k-Nearest neighbor (kNN)
kNN is a non-parametric classification method [37]. It is carried out a classification task using a distance metric such as Euclidean as described in Eq. (2). It needs a training set to determine the distribution of the samples. Then the test data is classified using a majority vote of k-nearest neighbors in the training set [38].
We preferred the Euclidean distance metric for kNN and k was searched in the range of 1 and 10. The most efficient results were obtained when k was set to 3.
Decision tree (DT)
DT is a useful machine learning method to generate regression or classification models based on the tree structure. A DT consists of a root node, branch nodes, and leaf nodes [39]. These nodes correspond to an algorithm that is used to control conditional statements. It means that the way from root to a leaf corresponds to a set of classification rules. The root is determined using information gain theory, and growing a DT continues until the leaf nodes are obtained [40]. To achieve an efficient DT model, some hyperparameters such as the depth of the tree, merging criteria of the leaf, the size of parents, and splitting predictor should be chosen properly. In the experiment, we employed hyperparameter optimization based on the Bayesian optimization to optimize all eligible parameters. Gini’s diversity index (GDI) was used as a split criterion.
Support vector machine (SVM)
SVM is an important machine learning concept which can be used for either supervised classification and regression applications or for unsupervised data clustering [30]. SVM aims to find an optimal separating hyperplane between positive and negative samples, where the margin around the hyperplane is maximized. Let’s assume a set of training samples \(\left( {\varvec{x}_{1} ,\varvec{y}_{1} } \right), \ldots ,\left( {\varvec{x}_{\varvec{N}} ,\varvec{y}_{\varvec{N}} } \right)\) are given, where xi shows sample feature vector and yi is the class label. Class labels are either positive or negative. As it was mentioned earlier, the SVM approach runs an optimization algorithm to find an optimum class separation hyperplane, which has the maximal margin. To do so; the following equations are considered;
where αi is the weight vector that is accompanied with xi and C is called as the regulation parameter. K shows the kernel function that is used to calculate the similarity between xi and xj. Gaussian radial function, linear function, and polynomial function can be used for kernel function.
In the experiment, RBF kernel was used and sigma was assessed in the range of 1 and 10. As a result, the most efficient results were yielded when sigma was set to 2. Also, the regulation parameter was evaluated in the range of 1 and 100. It was adjusted to 10.
Results
A total of 30 features were obtained by means of CTG-OAS. The features and marginal histograms are illustrated using the first two principal components in Fig. 2. As shown in Fig. 2, separating the recordings as normal and hypoxic and finding a borderline for this purpose is a quite challenging task. For this reason, we utilized several machine learning models such as ANN, kNN, DT and SVM.
In order to measure the performance of the feature selection algorithms, 10-fold cross-validation (CV) method was used. The several performance metrics, which are accuracy (Acc), sensitivity (Se), Specificity (Sp), quality index (QI) and F-measure, derived from confusion matrix were also considered. Confusion matrix consists of four prognostic indices which are True Positive (TP), True Negative (TN), False Positive (FP) and False Negative (FN). TP and TN represent the number of hypoxic and normal fetuses identified correctly whereas FP and FN represent the number of hypoxic and normal fetuses identified incorrectly, respectively. The aforementioned performances metrics are calculated as follow:
Acc gives the overall efficiency of the model. Se and Sp explain the efficiency of the model on positively and negatively labeled data, respectively. QI is the geometric mean of Se and Sp and it is a very useful metric when the distribution of data is imbalanced among the classes. F-measure expresses the harmonic mean between precision and recall. Furthermore, we used receiver operator (ROC) curve which defines the relationship between Se and Sp. Also, the area under this curve (AUC) was calculated to determine the performance of the classifier.
In the experimental study, feature ranking techniques were examined first. Weighted by SVM, IGR and Relief methods were employed individually with 10-fold cross-validation procedure and machine learning models for this particular purpose. The selected features and classification performances were reported in Tables 1 and 2, respectively. As shown in Table 2, the most efficient results were obtained using a combination of Weighted by SVM and SVM classifier. Then, two wrappers methods, BE and RFE were utilized. Sp values were superior to Se values because of imbalanced data distribution. The features selected by at least 3 of 5 feature selection algorithms were used to generate the most relevant final feature subset. A total of 12 features was determined as the most relevant, and these features and their relationship with each other are illustrated in Fig. 3.
It is observed that there is a high correlation among IBTF features, especially in the features belonging to the VLF band. Figure 3 is examined, a similar relationship can also be seen between Baseline, Mean, and LTI features. It should be noted that the values of IBTF features are normalized in the range of 0 and 1.
In the last step of the experiment, the most relevant features were applied as an input to machine learning models. The aggregate confusion matrices and performance metrics are given in Tables 3 and 4, respectively. Table 4 is compared with Table 2, it can be seen clearly that the best results were obtained using the most informative feature subset. As a result, Se of 77.40% and Sp of 93.86% were achieved. Also, the values of QI (85.23%) and F-measure (81.30%) metrics were quite satisfactory. As mentioned above, another significant tool regarding the model evaluation with two classes is the ROC curve and AUC. The highest AUC (close to 1) shows the highest certainty of the fetal hypoxia detection according to the analyzed feature set. ROC curve of the models with most relevant features are illustrated in Fig. 4, and AUCs were achieved as 0.7890, 0.6777, 0.8591, and 0.8874 for ANN, kNN, DT and SVM, respectively.
Discussion
As underlined in the introduction, CTG has a high disagreement level among observers because of visual inspection and suffers from lacking practicable standards in daily clinical practice [7]. For this reason, automated CTG analysis is admitted as the most promising way to tackle these disadvantages. Features selection algorithms are of great importance in terms of automated CTG analysis. In this paper, we evaluate a total of five feature selection algorithms consisting of three filters and two wrappers methods on CTG data for the fetal hypoxia detection task.
Identification of FHR signals by diagnostic indices obtained from different fields such as morphological, linear, nonlinear, and IBTF enhances the possibility of recognizing fetal hypoxia. A crucial factor is connected with the selection of the most relevant features which are applied as the input to classifiers. The use of multiple feature selection algorithms can produce better results as in our experiment since the most relevant features are determined according to their selection frequency by the feature selection algorithms. Consequently, the most informative feature set which is a subset of the full feature set consisting of 30 features has only 12 diagnostics indices. Moreover, this subset provided the best results.
According to the results of each method used in the experiments, Se values were higher than Sp values due to the imbalanced data distribution. Using either oversampling or downsampling technique to balance data distribution could lead to better results [30]. A further step for improving the classification performance will be using different machine learning techniques. Furthermore, the spectrogram images may be an enormous information source for detection of fetal hypoxia considering deep learning algorithms such as convolution neural network [12, 41]. We believe that we can obtain more successful results by IBTF analysis projection.
In this section, we also present a comparison of the related works considering several parameters such as methods, datasets, the number of features for describing the CTG signals, and performance metrics. However, it is important to be aware that making a one-to-one comparison among the related works is not suitable due to the different parameters as mentioned above. The comparison results are given in Table 5. Subha et al. [42] and Velappan et al. [43] used a public dataset called UCI CTG. This dataset generated using SisPorto software [44] and come up with 21 diagnostic features extracted automatically by the software. On other words, no need to use advanced signal processing techniques on raw CTG signals thanks to the SisPorto software for this dataset. As a result of this situation, high-performance results were achieved. Genetic algorithm, filters, and wrappers methods have been examined on CTU-UHB intrapartum CTG database to reach more consistent diagnosis models. However, because of the different division criteria, and the complex structure of the intrapartum recordings, this area has remained a challenging work. To overcome this issue, we generated a more relevant feature set based on the five feature selection algorithm covering filters and wrappers methods. Each feature in this subset was selected by at least three feature selection algorithms. As a result, we achieved 88.58% classification accuracy.
Conclusion
CTG is one of the fetal surveillance technique used routinely in obstetric clinics to monitor fetal well-being. Basically, it suffers from visual examination, and for this reason, the computerized systems are in demand. In this study, we carried out advanced signal processing techniques to achieve reliable segments and to extract features for describing the signals. Then, the combinations of four machine learning algorithms and five feature selection algorithms were examined. As a result, 12 features were determined as the most relevant from the full feature set consisting of 30 diagnostic features. As a result, we achieved Acc of 88.58%, Se of 77.40% and Sp of 93.86%. This work points out that determining the optimal feature set ensures more consistent and effective diagnosis models.
References
Grivell RM, Alfirevic Z, Gyte GM, Devane D. Antenatal cardiotocography for fetal assessment. Cochrane Database Syst Rev. 2010;9:1–48. https://doi.org/10.1002/14651858.CD007863.pub2.
Strachan BK, Sahota DS, Van Wijngaarden WJ, James DK, Chang AMZ. Computerised analysis of the fetal heart rate and relation to acidaemia at delivery. Br J Obstet Gynaecol. 2001;108:848–52. https://doi.org/10.1016/S0306-5456(00)00195-9.
Pinas A, Chandraharan E. Continuous cardiotocography during labour: analysis, classification and management. Best Pract. Res. Clin. Obstet. Gynaecol. 2016;30:33–47. https://doi.org/10.1016/j.bpobgyn.2015.03.022.
Norén H, Amer-Wåhlin I, Hagberg H, Herbst A, Kjellmer I, Marşál K, Olofsson P, Rosén KG. Fetal electrocardiography in labor and neonatal outcome: data from the Swedish randomized controlled trial on intrapartum fetal monitoring. Am J Obstet Gynecol. 2017;188:183–92. https://doi.org/10.1067/mob.2003.109.
Georgoulas G, Karvelis P, Spilka J, Chudáček V, Stylios CD, Lhotská L. Investigating pH based evaluation of fetal heart rate (FHR) recordings. Health Technol. (Berl). 2017;7:241–54. https://doi.org/10.1007/s12553-017-0201-7.
Bernardes J, Costa-Pereira A, Ayres-De-Campos D, Van Geijn HP, Pereira-Leite L. Evaluation of interobserver agreement of cardiotocograms. Int. J. Gynecol. Obstet. 1997;57:33–7. https://doi.org/10.1016/S0020-7292(97)02846-4.
Donker DK, van Geijn HP, Hasman A. Interobserver variation in the assessment of fetal heart rate recordings. Eur J Obstet Gynecol Reprod Biol. 1993;52:21–8. https://doi.org/10.1016/0028-2243(93)90220-7.
Czabanski R, Jezewski J, Matonia A, Jezewski M. Computerized analysis of fetal heart rate signals as the predictor of neonatal acidemia. Expert Syst Appl. 2012;39:11846–60. https://doi.org/10.1016/j.eswa.2012.01.196.
Ayres-de-Campos D, Spong CY, Chandraharan E. FIGO consensus guidelines on intrapartum fetal monitoring: cardiotocography. Int. J. Gynecol. Obstet. 2015;131:13–24. https://doi.org/10.1016/j.ijgo.2015.06.020.
Rhose S, Heinis AMF, Vandenbussche F, van Drongelen J, van Dillen J. Inter- and intra-observer agreement of non-reassuring cardiotocography analysis and subsequent clinical management. Acta Obstet Gynecol Scand. 2014;93:596–602. https://doi.org/10.1111/aogs.12371.
Ayres-de-Campos D, Ugwumadu A, Banfield P, Lynch P, Amin P, Horwell D, Costa A, Santos C, Bernardes J, Rosen K. A randomised clinical trial of intrapartum fetal monitoring with computer analysis and alerts versus previously available monitoring. BMC Pregnancy Childbirth. 2010;10:71. https://doi.org/10.1186/1471-2393-10-71.
Zhao Z, Zhang Y, Comert Z, Deng Y. Computer-aided diagnosis system of fetal hypoxia incorporating recurrence plot with convolutional neural network. Front. Physiol. 2019;10:255. https://doi.org/10.3389/fphys.2019.00255.
Signorini MG, Magenes G, Cerutti S, Arduini D. Linear and nonlinear parameters for the analysis of fetal heart rate signal from cardiotocographic recordings. IEEE Trans Biomed Eng. 2003;50:365–74. https://doi.org/10.1109/TBME.2003.808824.
Karin J, Hirsch M, Sagiv C, Akselrod S. Fetal autonomic nervous system activity monitoring by spectral analysis of heart rate variations. In: Proceedings of computers in cardiology; 1992. p. 479–482. https://doi.org/10.1109/cic.1992.269517.
Zarmehri MN, Castro L, Santos J, Bernardes J, Costa A, Santos CC. On the prediction of foetal acidaemia: a spectral analysis-based approach. Comput Biol Med. 2019;109:235–41. https://doi.org/10.1016/j.compbiomed.2019.04.041
Romagnoli S, Sbrollini A, Burattini L, Marcantoni I, Morettini M, Burattini L. Digital cardiotocography: what is the optimal sampling frequency? Biomed Signal Process Control. 2019;51:210–5. https://doi.org/10.1016/j.bspc.2019.02.016.
Cömert Z, Kocamaz AF. Cardiotocography analysis based on segmentation-based fractal texture decomposition and extreme learning machine. In: 25th Signal processing and communications applications conference; 2017. p. 1–4. https://doi.org/10.1109/siu.2017.7960397.
Cömert Z, Kocamaz AF, Subha V. Prognostic model based on image-based time–frequency features and genetic algorithm for fetal hypoxia assessment. Comput Biol Med. 2018. https://doi.org/10.1016/j.compbiomed.2018.06.003.
Cömert Z, Kocamaz AF. A study based on gray level co-occurrence matrix and neural network community for determination of hypoxic fetuses. In: International artificial intelligence and data processing symposium, TR, 2016. p. 569–573. https://doi.org/10.13140/rg.2.2.23901.00489.
Usha Sri A, Malini M, Chandana G. Feature extraction of cardiotocography signal. In: Satapathy SC, Raju KS, Shyamala K, Krishna DR, Favorskaya MN, editors. Advances in decision sciences, image processing, security and computer vision. Cham: Springer; 2020. p. 74–81.
Alkhasawneh MS. Hybrid cascade forward neural network with elman neural network for disease prediction. Arab J Sci. Eng. 2019. https://doi.org/10.1007/s13369-019-03829-3.
Cömert Z, Kocamaz AF. Evaluation of fetal distress diagnosis during delivery stages based on linear and nonlinear features of fetal heart rate for neural network community. Int J Comput Appl. 2016;156:26–31. https://doi.org/10.5120/ijca2016912417.
Alsayyari A. Fetal cardiotocography monitoring using Legendre neural networks. Biomed. Eng. Tech. 2019;100000:10000. https://doi.org/10.1515/bmt-2018-0074.
Uzun A, Kızıltas CE, Yılmaz E. Cardiotocography data set classification with extreme learning machine. In: International conference on advanced technologies, computer engineering and science; 2018. p. 224–230.
Cömert Z, Kocamaz AF. Comparison of machine learning techniques for fetal heart rate classification. Acta Phys Pol, A. 2017;132:451–4. https://doi.org/10.12693/aphyspola.131.451.
Stylios CD, Georgoulas G, Karvelis P, Spilka J, Chudáček V, Lhotska L. Least squares support vector machines for FHR classification and assessing the pH based categorization. In: Kyriacou E, Christofides S, Pattichis CS, editors. XIV Mediterranean conference on medical and biological engineering and computing, 2016 (MEDICON 2016), 31 March–2 April 2016, Paphos, Cyprus. Cham: Springer; 2016. p. 1211–1215. https://doi.org/10.1007/978-3-319-32703-7_234.
Spilka J, Frecon J, Leonarduzzi R, Pustelnik N, Abry P, Doret M. Sparse support vector machine for intrapartum fetal heart rate classification. IEEE J. Biomed. Heal. Inform. 2016;10000:10000. https://doi.org/10.1109/JBHI.2016.2546312.
Xu L, Georgieva A, Redman CWG, Payne SJ. Feature selection for computerized fetal heart rate analysis using genetic algorithms. In: 2013 35th Annual international conference of the IEEE Engineering in Medicine and Biology Society; 2013. pp. 445–448. https://doi.org/10.1109/embc.2013.6609532.
Cömert Z, Kocamaz AF. Novel software for comprehensive analysis of cardiotocography signals CTG-OAS. In: Karci A, editor. International conference on artificial intelligence and data processing. Malatya: IEEE; 2017. p. 1–6. https://doi.org/10.1109/idap.2017.8090210.
Chudacek V, Spilka J, Rubackova B, Koucky M, Georgoulas G, Lhotska L, Stylios C. Evaluation of feature subsets for classification of cardiotocographic recordings. In: 35th Annual conference on computers in cardiology 2008. Piscataway, NJ: IEEE; 2008. p. 845–848. https://doi.org/10.1109/cic.2008.4749174.
Menai MEB, Mohder FJ. Al-mutairi, Influence of feature selection on naïve Bayes classifier for recognizing patterns in cardiotocograms. J. Med. Bioeng. 2013;2:66–70. https://doi.org/10.12720/jomb.2.1.66-70.
Chudáček V, Spilka J, Burša M, Janků P, Hruban L, Huptych M, Lhotská L. Open access intrapartum CTG database. BMC Pregnancy Childbirth. 2014;14:16. https://doi.org/10.1186/1471-2393-14-16.
Gonçalves H, Bernardes J, Paula Rocha A, Ayres-de-Campos D. Linear and nonlinear analysis of heart rate patterns associated with fetal behavioral states in the antepartum period. Early Hum Dev. 2007;83:585–91. https://doi.org/10.1016/j.earlhumdev.2006.12.006.
Spilka J, Chudáček V, Koucký M, Lhotská L, Huptych M, Janků P, Georgoulas G, Stylios C. Using nonlinear features for fetal heart rate classification. Biomed Signal Process Control. 2012;7:350–7. https://doi.org/10.1016/j.bspc.2011.06.008.
Huang M-L, Yung-Yan H. Fetal distress prediction using discriminant analysis, decision tree, and artificial neural network. J. Biomed. Sci. Eng. 2012;05:526–33. https://doi.org/10.4236/jbise.2012.59065.
Sahin H, Subasi A. Classification of the cardiotocogram data for anticipation of fetal risks using machine learning techniques. Appl. Soft Comput. 2015;33:231–8. https://doi.org/10.1016/j.asoc.2015.04.038.
Aha DW, Kibler D, Albert MK. Instance-based learning algorithms. Mach. Learn. 1991;6:37–66. https://doi.org/10.1007/BF00153759.
Akbulut Y, Sengur A, Guo Y, Smarandache F. NS-k-NN: neutrosophic set-based k-nearest neighbors classifier. Symmetry (Basel). 2017;9:179.
Safavian SR, Landgrebe D. A survey of decision tree classifier methodology. IEEE Trans. Syst. Man. Cybern. 1991;21:660–74.
Altuntaş Y, Kocamaz AF, Cömert Z, Cengiz R, Esmeray M. Identification of haploid maize seeds using gray level co-occurrence matrix and machine learning techniques. In: 2018 International conference on artificial intelligence and data Processing, Malatya, Turkey; 2018. p. 1–5. https://doi.org/10.1109/idap.2018.8620740
Cömert Z, Kocamaz AF. Fetal hypoxia detection based on deep convolutional neural network with transfer learning approach. In: Silhavy R, editor. Software engineering and algorithms in intelligent systems. Cham: Springer; 2019. p. 239–248. https://doi.org/10.1007/978-3-319-91186-1_25.
Subha V, Murugan D, Boopathi AM, Velappan S, Murugan D, Boopathi M. A hybrid filter-wrapper attribute reduction approach for fetal risk anticipation. Asian J. Res. Soc. Sci. Humanit. 2017;7:1094–106.
Velappan S, Murugan D, Prabha S, Boopathi M. Genetic Algorithm based feature subset selection for fetal state classification. J. Commun. Technol. Electron. Comput. Sci. 2015;2:13–7.
Ayres-de-campos D, Bernardes J, Garrido A, Marques-de-sá J, Pereira-leite L. SisPorto 2.0: a program for automated analysis of cardiotocograms. J. Matern. Fetal. Med. 2000;9:311–8. https://doi.org/10.3109/14767050009053454.
Spilka J, Georgoulas G, Karvelis P, Oikonomou VP, Chudáček V, Stylios C, Lhotská L, Jankru P. Automatic evaluation of FHR recordings from CTU-UHB CTG database. In: Bursa M, Khuri S, Renda ME, editors. Proceedings of 4th international conference on information technology in bio- and medical informatics (ITBAM 2013), Prague, Czech Republic, 28 August 2013. Berlin: Springer, 2013, pp. 47–61. https://doi.org/10.1007/978-3-642-40093-3_4.
Acknowledgements
We would like to thanks to Dr. Sami Güngör due to its comments on the medical background.
Funding
There is no funding source for this article.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that there is no conflict to interest related to this paper.
Ethical approval
This article does not contain any data, or other information from studies or experimentation, with the involvement of human or animal subjects.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cömert, Z., Şengür, A., Budak, Ü. et al. Prediction of intrapartum fetal hypoxia considering feature selection algorithms and machine learning models. Health Inf Sci Syst 7, 17 (2019). https://doi.org/10.1007/s13755-019-0079-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13755-019-0079-z