Abstract
Potassium ions are widely involved in a series of physiological and biochemical processes of plants, which is of great significance to plant growth. High-affinity K+ (HAK) transporter mainly absorbed and transported potassium in plants, but there are few studies on HAK gene in sugarcane. In this study, the coding region of high-affinity K+ transporter genes were cloned from sugarcane and designated as ScHAK9, ScHAK10, ScHAK11 (GenBank accession number: MG564720, MG564721, MG564722). Phylogenetic analysis results confirmed that ScHAK9 and ScHAK11 had the closest relationship with ZmHAK9 and ZmHAK11(Zea mays), ScHAK10 had the closest relationship with SbHAK10 (Sorghum bicolor). In subcellular localization experiments, the fusion protein of ScHAK9 and ScHAK11 with green fluorescent protein was specifically localized in the cell membrane, but ScHAK10 green fluorescent protein was not detected, it was speculated to be expressed in Golgi apparatus. The gene expression level of ScHAK in different tissues of sugarcane at the growth periods was different, and the gene expression level of ScHAK genes were up-regulated by the low-potassium and salt stress. Through the functional characterization experiments of ScHAK genes in K+ uptake-deficient yeasts, it was founded that ScHAK genes possessed K+ transporter activity. The study indicated that ScHAK genes might mediate K+ absorption through the cell membrane and might be participate in maintaining Na+/K+ homeostasis in sugarcane under the adversity stress, and the development of plant organs is regulated by the potassium ions transport of ScHAK genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Potassium is an essential element in plants and its activity is related to enzyme activity regulation, protein synthesis, osmotic adjustment, and photosynthesis (Ma et al. 2012; Amtmann et al. 2005) as well as stress response of plants (Anschutz et al. 2014). The absorption, transport and distribution of K+ are primarily realized via potassium ion channels and transport carriers, which can be classified as the KUP/HAK/KT family, HKT/TPK family, CHX family, and KEA family (Maser et al. 2001). High-affinity K+ (HAK) transporter gene belongs to KUP/HAK/KT family and is important in the absorption of plant potassium (Gierth et al. 2005). Identification of HAK genes encoding were first identified from barley (Santa-Maria et al. 1997), several HAK genes were cloned from various plants including maize, rice, mesembryanthemum crystallinum and poplar (Zhang et al. 2012; Yang et al. 2009; Su et al. 2002; He et al. 2012). HAK genes were founded to exist widely in plant tissues. NtHAK1 gene was expressed in various organs of tobacco, particularly in roots (Lu and Yang 2011). After the gene was knocked out, root hair elongation was damaged, indicating that the root growth was affected by HAK genes (Zou et al. 2012). Arabidopsis mutants (AtHAK5) germinated slowly and their root growth was limited under the low-potassium stress, indicating that HAK genes were involved in the physiological processes of seed germination and root elongation (Nieves-Cordones et al. 2010).The KUP/HAK/KT family can be divided into three types such as high-affinity, low-affinity and mixed-affinity potassium transporters according to the affinity of potassium absorption (Rubio et al. 2000). High-affinity potassium transporters participate in potassium absorption under low-potassium conditions (Banuelos et al. 2002). Low-affinity potassium transporters are widely distributed in plant tissues and work together with potassium channels to absorb potassium under a high potassium concentration (Song et al. 2013). Mixed-affinity potassium transporters function as K+ and Na+ bidirectional transporters (Maathuis 2006; Takahashi et al. 2007).
In summary, it is reported that many plants contain HAK family genes, the research on the physiological function of HAK family genes is still in the initial stage, the known functional of HAK genes had been identified in a few plants, such as Arabidopsis, rice and maize, but their physiological functions and regulatory mechanisms in sugarcane remain unknown. Sugarcane is allopolyploid, and it is speculated that the HAK family in sugarcane will also be a large family with complex functions. The study on the phylogenetic evolution, structure and function of the HAK genes would lay a foundation for the mechanism of potassium absorption and transport in sugarcane. In this study, three members of sugarcane high-affinity K+ transporter (ScHAK) gene family coding sequence were cloned, and the primary features and structures of ScHAK genes along with its relationship with HAK from other genera were characterized. The sugarcane HAK gene expression characteristics were analyzed in the roots, stems and leaves at the physiological maturity stage, and under low-potassium stress treatment (0.1 mmol/L) and salt stress treatment (1% NaCl) in sugarcane leaves, subcellular localization analyses revealed that ScHAK9 and ScHAK11 protein were localized to the plasma membrane in tobacco epidermal cells. When expressed in K+ uptake-deficient yeast cells, ScHAK genes were clearly improve K+ acquisition of the yeast and tolerance to low K+ environment. The functional and expression analysis of ScHAK genes in sugarcane may provide the way to improve the potassium absorption efficiency in sugarcane.
Materials and methods
Plant material
Sugarcane variety “ROC22” in China was used for gene cloning and gene expression studies. Sugarcane (Saccharum officinarum) was planted with hoagland nutrient solution added in water in the greenhouse (average temperature, 25 °C, relative humidity, 80 ± 5%). One group was set as the control and the other two treatments included low-potassium stress treatment (0.1 mmol/L) and salt stress treatment (1% NaCl). Leaf samples were harvested at different intervals (24 h, 48 h, 72 h and 96 h) at the seedling stage, stored at − 80 °C for RNA isolation. Choose consistent and healthy cane stems for planting in a greenhouse (average temperature, 25 °C, soil water content at 25% ± 5%). The complete plants (roots, stems and leaves) were also collected at the age of seedling stage, elongation stage and maturation stage.
Gene cloning and sequence bioinformatics analysis of ScHAK genes
RNA was extracted from leaves with RNA extraction kit (Omega Bio-Tek, Guangzhou, China). First-strand cDNA template for RT-PCR amplification was obtained by using One-Step gDNA Removal and cDNA Synthesis Super-Mix kit (Transgen, Beijing, China). According to the data of sugarcane cDNA library, primer pair were designed from the conserved sequence region. Then PCR was performed with the template of first-strand cDNA. The used primer pair was designed as:
-
ScHAK9-F: 5'-ATGGATCCCGAGTTCGGCGT-3'.
-
ScHAK9-R: 5'-TCACAGCACGTACACCATGC-3'.
-
ScHAK10-F: 5'-ATGAAGAGCCCCCCTGTCAT-3'.
-
ScHAK10-R: 5'-TCAGATATAGTACATCATGCCG-3'.
-
ScHAK11-F: 5'-ATGGCATCGCTGTCAGAAAGT-3'.
-
ScHAK11-R: 5'-TCAGATGTAGTATATCTGGCCG-3'.
The PCR reaction system contained 12.5 μL of PCR Buffer, 5 μL of dNTP mixture (2 mmol/L), 0.75 μL of upstream primer(10 μM/L), 0.75 μL of downstream primer(10 μM/L), 0.5 μL of KOD FX DNA polymerase, and 3.0 μL of first-strand cDNA template and was diluted to a volume of 25.0 μL with double-distilled water. PCR program was composed of initial denaturation at 94 °C for 2 min, 35 cycles of 98 °C for 10 s, 61 °C for 30 s, and 72 °C for 3 min, and final extension at 72 °C for 7 min. Then, PCR products(2 μL) were electrophoresed on 1.2% agarose gel. Remaining PCR products were purified with the gel extraction kit (Biomed Technology, Beijing, China), then cloned into pEASY-Blunt Simple Cloning vector, transformed into E. coli T1, and sequenced.
The sequence of ScHAK genes were analyzed with Prot Param (https://web.expasy.org/protparam/) and Protter software (http://wlab.ethz.ch/protter/start/) to predict the amino acid sequence, hydropathy, protein structure, molecular weight, and isoelectric point of putative protein. An alignment of ScHAK protein and other plant HAK proteins retrieved from NCBI was carried out in Clustal W. Then with the alignment results, a phylogenetic tree was constructed in MEGA 6.0 according to the neighbor-joining method.
Construction of GFP fusion carrier and transfection
To construct ScHAK-GFP fusion fragment for the expression in tobacco epidermal cells, PCR products were digested with NcoI and SpeI, and then ligated to pCAMBICA1302 vector. The recombinant plasmids (pCAMBICA1302- ScHAK) were confirmed by double digestion with NcoI and SpeI and sequencing and then transferred to TOP10 clone strain.
According to the previous method (Xu et al. 2014), the empty plasmid pCAMBIA1302 was used as the control. A positive monoclonal antibody containing recombinant plasmid pCAMBIA-ScHAK was amplified in YEP liquid culture (containing 50 mg/mL rifampicin and 25 mg/mL Kanamycin) and cultivated to OD600 nm = 1.5–2.0 at 28 °C. After centrifugation, Agrobacterium bacteria were collected. Then, 50 mL of Agrobacterium suspension was added, mixed, injected into tobacco epidermis, and then cultured for 72 h under normal illumination conditions. Tobacco epidermis was observed with a confocal laser scanning microscope (excitation wavelength, 488 nm; emission wavelength, 625–725 nm).
Quantitative real-time PCR analysis
Total RNA from roots, stems and leaves were extracted with Spin Column Plant total RNA Purification Kit (Omega Bio-Tek, Guangzhou, China). Then the first-strand cDNA was synthesized with AMV Reverse Transcriptase First Strand cDNA Synthesis Kit (TransGen Biotech, Beijing, China). Specific primers for qRT-PCR of ScHAK and 25S rRNA (as house-keeping gene) genes are shown in Table 1.
Real-time quantitative PCR was performed on ABI 7500 with the 20-μL reaction system composed of 7.0 μL of 2 × SYBR Green qPCR Master Mix, 0.5 μL of 10 μM upstream primer, 0.5 μL of 10 μM downstream primer, 3 μL of first-strand cDNA template, and 9 μL of double-distilled water. The amplification program was set as follows: initial denaturation at 95 °C for 10 min, 40 cycles of denaturation and annealing extension at 95 °C for 10 s and 60 °C for 34 s, dissolution curve collection at 95 °C for 30 s and 60 °C for 15 s. Triplicate technical replicates were analyzed for each biological replicate.
Complementation test of yeast mutants
Complementation test of yeast mutants was performed according to the method of Li et al. (2014). According to the ORF sequence of ScHAK, primers containing restriction sites were designed (Table 2). After double digestion, ScHAK11 was connected to the expression vector pYES2 with the same digestion site by T4 DNA ligase. The fused plasmid was composed of the GAL1 promoter, the CYC1 terminator and a selective marker URA3. The vector pYES2 was used as a negative control. The transformants were selected on Glc-containing SC-agar plates with 100 mm K+ and zero uracil. AP (arginine and phosphoric acid) plates containing 0.5, 5, 10, 20, and 50 mM K+ were used for subsequent growth assays according to the previous method (Horie et al. 2011). There clones were used in growth tests with solid plates under 28 °C for 48–72 h.
Results
Genetic relationship analysis of ScHAK
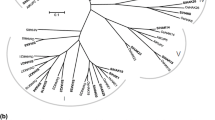
In order to study the evolutionary relationship of ScHAK9, ScHAK10 and ScHAK11 proteins, there related HAK proteins from NCBI protein database were downloader for constructing phylogenetic tree by MEGA 6.0. The phylogenetic relationship suggested that they might have the similar functions.
The phylogenetic results revealed that ScHAK genes can be divided into 3 groups, the sequence of ScHAK9 and ScHAK11 had the highest identity with Zea mays HAK, but the ScHAK10 had the highest identity with Sorghum bicolor HAK genes (Fig. 1). The research on the phylogenetic tree of genes can speculate its functional characteristics.
Protein domain analysis of ScHAK
A coding sequence (CDs) of ScHAK9, ScHAK10 and ScHAK11 were obtained from sugarcane leaves and then submitted in NCBI (GenBank accession number: MG564720, MG564721,MG564722).There amino acid sequences encoded by the cloned ScHAK genes were analyzed by BLASTP of NCBI, and it was found that ScHAK9, ScHAK10 and ScHAK11 sequences had a conserved domain of potassium ion transport subfamily, which belonged to the HAK/ KUP /KT protein family, The basic physicochemical properties and function prediction of the coding proteins were analyzed, and the results were shown in Table 3.
The prediction analysis of the transmembrane structure of ScHAK proteins showed that the transmembrane structure of proteins can be divided into two models, the first model has a long ring structure between the second and third transmembrane regions, and the ScHAK9 gene belongs to this model, the second model has a long ring structure between the first and second transmembrane regions, and the ScHAK10, ScHAK11 genes belongs to this model. The transmembrane times of different proteins were different, but their structures were similar, and no signal peptide structure was found, then indicating that it was a non-secretory protein as a transport vector and could not be transported after it was synthesized in the cytoplasm, but might exist as a transport carrier (Fig. 2).
Subcellular localization analysis of ScHAK
In order to confirm the subcellular localization of ScHAK protein, the vector containing ScHAK-GFP fusion genes were constructed. The fusion protein expression vector pCAMBIA1302 was constructed by fusing the C-terminus of ScHAK protein to the green fluorescence protein (GFP) with the 35S promoter of Cauliflower Mosaic Virus (CaMV). Then, the DNA plasmids of pCAMBIA1302-ScHAK and the GFP control were transform into tobacco epidermal cells by Agrobacterium-mediated transformation. The cellular location of ScHAK protein was confirmed through the fluorescence and confocal analysis of laser microscopy under ultraviolet light. The result showed that the GFP protein was distributed throughout the cell membrane and nucleus with the pCAMBIA1302 vector. However, green fluorescence was specifically detected in the cell membrane transiently transfected with the pCAMBIA-ScHAK9 and pCAMBIA-ScHAK11, and green fluorescence with the pCAMBIA-ScHAK9 was aslo detected in the cell nucleus (Fig. 3b and d), the ScHAK10-GFP fusion protein was not detected which speculated that the protein is expressed in Golgi apparatus (Fig. 3c). These results further proved that ScHAK9 and ScHAK11 were membrane protein.
Expression patterns of ScHAK in sugarcane tissues and abiotic stresses
The tissue-specific expression of genes was related to the specific biological function of genes, so the tissue expression specificity of three genes were analyzed by qRT-PCR. The gene expression profile of ScHAK in sugarcane was confirmed, by further investigation in its expression patterns in leaves, roots and stems at different growth stages (seedling, elongating, and mature stages) by qRT-PCR (Fig. 4a, b and c). In the seedling stage, all ScHAK genes were expressed in leaves, roots and stems, ScHAK9 gene was highly expressed in root at elongation stage, ScHAK10 gene was highly expressed in leaf and stem at maturation stage, ScHAK11 gene was highly expressed in roots and leaves at elongation stage. These results indicated that expression of ScHAK genes were spatiotemporal specific in growth process of sugarcane.
Moreover, the expression profiles of ScHAK were also detected under low-potassium and salt stress treatments. In our observations, the expression of ScHAK genes were induced under multiple stresses (Fig. 4d, e). The expression of ScHAK genes reached the maximum level in 48 h after low potassium treatment, and rapidly induced after 24 h under salt stress and its expression reached the highest level after 96 h. The experimental results suggested that ScHAK genes might play an important role in the regulation under salt and low-potassium stress.
Functional analysis of ScHAK genes in yeast cells
In order to explore the transporter functional of ScHAK protein, the transformants were inoculated on AP medium respectively containing 0.5, 5, 10, 20 and 50 mM KCl. On AP medium containing 0.5 mM KCl, the empty pYES2 vector could not grow, but the strain CY162 transformed with ScHAK genes rescued its growth defect on AP medium containing 5–50 mM KCl. All yeast cells transformed with ScHAK genes grew better than the empty vector-transformed cells, and yeast cells transformed with ScHAK10 grew better than other Transferred ScHAK genes yeast strain (Fig. 5).The studies indicate that ScHAK genes improves the potassium absorption activity of yeasts mutant under low K+ concentrations, it was speculated that ScHAK has the function of transporting K+.
Discussion
As a C4 plant, sugarcane has a long growth cycle and requires a lot of fertilizers, especially potassium (Huang et al. 2013). HAK genes exist in many crops and involved in the absorption and transport of potassium ions in plants. But the HAK genes has not been cloned and characterized in sugarcane. Researches discovered that HAK gene family contains multiple transmembrane regions with a long cytoplasmic ring structure between second and third transmembrane regions (Sato et al. 2014). In our study, ScHAK9, ScHAK10 and ScHAK11 gene was first isolated from sugarcane. By comparing the deduced protein sequence of ScHAK proteins in the Protter software, we determined that ScHAK proteins were highly similar and all contained a long cytoplasmic ring structure, but the transmembrane structure model of proteins were different. Sequence phylogenetic analysis results suggested that ScHAK9 and ScHAK11 was highly homologous to Zea mays potassium transporter, and ScHAK10 was highly homologous to sorghum bicolor HAK10, this result indicates that ScHAK genes belongs to the HAK superfamily, and this result also may explain why ScHAK has similar functions to HAK, which plays an important role in potassium transport.
Previous research found that 25 HAK genes of maize were located in the plasma membrane, and two HAK genes of maize were located in the vesicle membrane and endoplasmic reticulum. In this study, the localization of ScHAK9 and ScHAK11 protein was obeserved in epidermal cells of tobacco. A transient expression of the ScHAK9 and ScHAK11 protein in tobacco was located in the plasma membrane, what is more, the green fluorescence signal of gene sufficiently proofed that the protein of ScHAK9 aslo existed in nucleus, subcellular localization indicated that ScHAK9 and ScHAK11 protein was transmembrane protein and have similar transmembrane transporting (Song et al. 2014). The localization of ScHAK10 protein was not observed in epidermal cells of tobacco, it was speculated that the protein was expressed in Golgi apparatus, and the transcriptional regulation mechanism is different from that of other crop homologous proteins.
In many plants, the expression patterns of HAK genes are obviously different in various tissues. In Arabidopsis, the hight expression level of AtHAK5 gene was detected in roots and its expression level in other tissues was low (Gierth et al. 2005). In soybean, the expression level of GmHAK30 gene in roots was 500 times higher than that in other tissues, but the expression levels of GmHAK10 and GmHAK25 were the highest in pods and seeds (Chao et al. 2017). However, there is no evidence to show the expression pattern of HAK genes in plant growth cycle. In this study, the expression pattern of ScHAK genes in various stages and tissues was analyzed by qRT-PCR, expression patterns of ScHAK genes in sugarcane were different from those of other crop HAK genes, ScHAK genes expression were related to growth period, ScHAK9, ScHAK10 and ScHAK11 gene was expressed throughout the plants, but gene expression levels were different in various growth stages, the relative expression of ScHAK9 in root was higher than that in leaf and stems at the elongating stage, the relative expression of ScHAK10 in leaf and stems were higher than that in root at the maturity stage, and the relative expression of ScHAK11 in leaf and root was higher than that in stems at the elongating and maturity stage, but the gene expression is lower than the other two genes. These results indicated that based on the demand characteristics of sugarcane growth, the members of sugarcane potassium transporter family play different roles, ScHAK genes might be involved in regulatory and developmental processes of K+ absorption and distribution in sugarcane, and the hight expression of genes in different tissues is conducive to promote the development of sugarcane (Feng et al. 2020).
The expression levels of HAK genes were affected by abiotic stresses. AlHAK1 was up-regulated remarkably in the roots of Aeluropus littoralis under the low-potassium stress and the gene expression in leaves lagged behind. It was presumed that the plant injury was reduced by the redistribution of AlHAK1 gene (Liu et al. 2015). Su et al. (2002) reported that the expression level of McHAK1 gene in Mesembryanthemum crystallinum was up-regulated to maintain stable K+/Na+ ratio by enhancing K+ uptake efficiency under the high-salt stress. In Oryza, OsHAK10 was expressed in root and other tissue, but its expression was not induced by low potassium stress and salt stress (Banuelos et al. 2002). In this study, the expression level of ScHAK genes was increased after 24 h under the salt stress and the relative expression of ScHAK genes reached the maximum value after 48 h under the low-potassium conditions. The results indicated that ScHAK9, ScHAK10 and ScHAK11 displayed upregulated expression, suggesting that they may play important roles in maintaining normal growth and mediating potassium acquisition under K+ deficiency. In addition, the expression level of ScHAK genes could improve the potassium uptake efficiency and tolerance of sugarcane under salt stress.
Most of the plant HAK proteins belongs to high-affinity potassium ion transporter which play an important role in potassium acquisition under potassium deficiency conditions (Santa-María et al. 1997; Banuelos et al. 2002; Gierth et al. 2005; Nieves-Cordones et al. 2010; Boscari et al. 2009; Yang et al. 2014). For example, the growth of OsHAK1 invert was detected in the mutant strain of yeasts under different potassium concentration gradients and the functions of three rice potassium transporters in yeasts were identified. (Banuelos et al. 2002) showed that OsHAK1 had the ability to transport K+ in Saccharomyces cerevisiae. After knocking out OsHAK1 in rice, Chen et al. (2015) found that the total K+ uptake was significantly decreased under low-K+ conditions. In this study, we found that the expression level of ScHAK proteins could rescue the growth defect of yeast cells (R5421) in both low-K+ (0.05 and 0.1 mM) media. In the media with high concentration of K+ (20 mM, 50 mM), yeast cells transformed with ScHAK or empty vector yeast strain showed that no obvious growth difference (Fig. 5), and in the medium with low-K+ concentration (0.05 mM), transgenic yeast cells grew better than empty carrier yeasts. The results showed that ScHAK genes could improve the K+ uptake activity of K+-deficient yeast mutant. It is suggested that ScHAK genes might have the similar function in sugarcane.
Conclusions
In the present study, three high-affinity K+ transporter genes cDNA, ScHAK9, ScHAK10 and ScHAK11 from sugarcane have been identified. The transmembrane structure of ScHAK proteins can be divided into two models. The qRT-PCR study showed that ScHAK genes induced by low-potassium and salt stress, and the expression level of genes in sugarcane tissues were different in growth process. Subcellular localizations and K+ uptake-deficient yeast revealed that ScHAK proteins may mediate K+ absorption by the plasma membrane and other organelle, they might be play crucial role in the maintenance of the K+ homeostasis in sugarcane under low-potassium situations.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- HAK :
-
High-affinity K+ transporter genes
- GFP:
-
Green fluorescence protein
- qRT-PCR:
-
Quantitative real-time PCR
- CaMV:
-
Cauliflower mosaic virus
- URA3:
-
Orotidine-5'-phosphate decarboxylase
- GAL1:
-
Recombinant galectin 1
- CYC1:
-
Cytochrome c-1
- AP:
-
Arginine and phosphoric acid
References
Amtmann A, Hammond JP, Armengaud P, White PJ (2005) Nutrient sensing and signalling in plants: potassium and phosphorus. Adv Bot Res 43:209–257. https://doi.org/10.1016/S0065-2296(05)43005-0
Anschutz U, Becker D, Shabala S (2014) Going beyond nutrition: regulation of potassium homoeostasis as a common denominator of plant adaptive responses to environment. J Plant Physiol 171(9):670–687. https://doi.org/10.1016/j.jplph.2014.01.009
Banuelos MA, Garciadeblas B, Cubero B, Navarro AR (2002) Inventory and functional characterization of the HAK potassium transporters of rice. Plant Physiol 130(2):784–795. https://doi.org/10.1104/pp.007781
Boscari A, Clément M, Volkov V, Golldack D, Hybiak J, Miller AJ, Amtmann A, Fricke W (2009) Potassium channels in barley: cloning, functional characterization and expression analyses in relation to leaf growth and development. Plant Cell Environ 32(12):1761–1777. https://doi.org/10.1111/j.1365-3040.2009.02033.x
Chao M, Zhang Z, Song H, Li C, Zhang X, Hu G, Zhang J, Wang Q (2017) Genome-wide identification and expression analysis of Pht1 family genes in cotton (Gossypium hirsutum L.). Cotton Sci 29(1):59–69. https://doi.org/10.11963/issn.1002-7807.201701007
Chen G, Hu Q, Luo LE, Yang T, Zhang S, Hu Y, Yu L, Xu G (2015) Rice potassium transporter OsHAK1 is essential for maintaining potassium-mediated growth and functions in salt tolerance over low and high potassium concentration ranges. Plant Cell Environ 38(12):2747–2765. https://doi.org/10.1111/pce.12585
Feng XM, Wang YJ, Zhang NN, Wu ZL, Zeng QY, Wu JY, Wu XB, Wang L, Zhang JS, Qi YW (2020) Genome-wide systematic characterization of the HAK/KUP/KT gene family and its expression profile during plant growth and in response to low-K+ stress in Saccharum. BMC Plant Biol 20(1):1–17. https://doi.org/10.1186/s12870-019-2227-7
Gierth M, Maser P, Schroeder JI (2005) The potassium transporter AtHAK5 functions in K+ deprivation-induced high-affinity K+ uptake and AKT1 K+ channel contribution to K+ uptake kinetics in Arabidopsis roots. Plant Physiol 137(3):1105–1114. https://doi.org/10.1104/pp.104.057216
He C, Cui K, Duan A, Zeng Y, Zhang J (2012) Genome-wide and molecular evolution analysis of the Poplar KT/HAK/KUP potassium transporter gene family. Ecol & Evol 2(8):1996–2004. https://doi.org/10.1002/ece3.299
Horie T, Sugawara M, Okada T, Taira K, Kaothien-Nakayama P, Katsuhara M, Shinmyo A, Nakayama H (2011) Rice sodium-insensitive potassium transporter, OsHAK5, confers increased salt tolerance in tobacco BY2 cells. J Biosci Bioeng 111(3):346–356. https://doi.org/10.1016/j.jbiosc.2010.10.014
Huang Y, Zeng QY, Ao JH, Chen DW, Zhou WL, Lu YL, Jiang Y, Huang ZR, Li QW (2013) Effects of different potassium levels on nutrient absorption and utilization of N, P K in sugarcane. Guangdong Agric Sci 40(10):50–53. https://doi.org/10.3969/j.issn.1004-874X.2013.10.016
Li J, Long Y, Qi GN, Li J, Xu ZJ, Wu WH, Wang Y (2014) The Os-AKT1channel is critical for K+ uptake in rice roots and is modulated by the rice CBL1-CIPK23 complex. Plant Cell 26(8):3387–3402. https://doi.org/10.1105/tpc.114.123455
Liu JF, Zhang SL, Tang HL, Wu LZ, Dong LJ, Liu LD, Che WL (2015) Overexpression of an Aeluropus littoralis Parl. potassium transporter gene, AlHAK1, in cotton enhances potassium uptake and salt tolerance. Euphytica 203:197–209. https://doi.org/10.1007/s10681-014-1310-2
Lu LM, Yang TZ (2011) Cloning and expression profile analysis of a putative potassium transporter gene NtHAK1 in tobacco. J Nucl Agric Sci 25(03):469–476
Ma TL, Wu WH, Wang Y (2012) Transcriptome analysis of rice root responses to potassium deficiency. BMC Plant Biol 12:161–174. https://doi.org/10.1186/1471-2229-12-161
Maathuis FJM (2006) The role of monovalent cation transporters in plant responses to salinity. J Exp Bot 57(5):1137–1147. https://doi.org/10.1093/jxb/erj001
Maser P, Thomine S, Schroeder JI, Ward JM, Hirschi K, Sze H, Talke IN, Amtmann A, Maathuis FJ, Sanders D, Harper JF, Tchieu J, Gribskov M, Persans MW, Salt DE, Kim SA, Guerinot ML (2001) Phylogenetic relationships within cation transporter families of Arabidopsis. Plant Physiol 126(4):1646–1667. https://doi.org/10.1104/pp.126.4.1646
Nieves-Cordones M, Aleman F, Martinez V, Rubio F (2010) The Arabidopsis thaliana HAK5 K+ transporter is required for plant growth and K+ acquisition from low K+ solutions under saline conditions. Mol Plant 3(2):326–333. https://doi.org/10.1093/mp/ssp102
Rubio F, Santa-Maria GE, Rodriguez-Navarro A (2000) Cloning of Arabidopsis and barley cDNAs encoding HAK potassium transporters in root and shoot cells. Physiol Plantarum 109(1):34–43. https://doi.org/10.1034/j.1399-3054.2000.100106.x
Santa-Maria GE, Rubio F, Dubcovsky J, Rodriguez-Navarro A (1997) The HAK1 gene of barley is a member of a large gene family and encodes a high-affinity potassium transporter. Plant Cell 9(12):2281–2289. https://doi.org/10.2307/3870585
Sato Y, Nanatani K, Hamamoto S, Shimizu M, Takahashi M, Tabuchi-Kobayashi M, Mizutani A, Schroeder JI, Souma S, Uozumi N (2014) Defining membrane spanning domains and crucial membrane-localized acidic amino acid residues for K+ transport of a KUP/HAK/KT-type Escherichia coli potassium transporter. Biochem J 155(5):315–323. https://doi.org/10.1093/jb/mvu007
Song YF, Zhang L, Dong LH, Jin YR, Shi SJ, Bai Y, Liu CK, Feng GL, Feng XG, Wang Q, Liu HB (2013) Research progress on KUP /HAK /KT potassium transporter family in plant. J Agric Sci Technol 15(06):92–98. https://doi.org/10.3969/j.issn.1008⁃0864.2013.0
Song YF, Liu HB, Dong LH, Jin YR, Shi SJ, Zhang L, Liu CK, Feng XG, Hu XM, Wang Q (2014) Subcellular localization and expression analysis of Nicotiana sylvestris KUP/HAK/KT family K+ transporter gene NsHAK11. Sci Agric Sin 47(6):1058–1071. https://doi.org/10.3864/j.issn.0578-1752.2014.06.003
Su H, Golldack D, Zhao C, Bohnert HJ (2002) The expression of HAK-type K+ transporters is regulated in response to salinity stress in common ice plant. Plant Physiol 129(4):1482–1493. https://doi.org/10.1104/pp.001149
Takahashi R, Nishio T, Ichizen N, Takano T (2007) High-affinity K+ transporter PhaHAK5 is expressed only in salt-sensitive reed plants and shows Na+ permeability under NaCl stress. Plant Cell Rep 26(9):1673–1679. https://doi.org/10.1007/s00299-007-0364-1
Yang Z, Gao Q, Sun C, Li W, Gu S, Xu C (2009) Molecular evolution and functional divergence of HAK potassium transporter gene family in rice (Oryza sativa L.). J Genet Genom 36(3):161–172. https://doi.org/10.1016/S1673-8527(08)60103-4
Yang TY, Zhang S, Hu YB, Wu FC, Hu QD, Chen G, Cai J, Wu T, Moran N, Yu L, Xu GH (2014) The role of a potassium transporter OsHAK5 in potassium acquisition and transport from roots to shoots in rice at low potassium supply levels. Plant Physiol 166(2):945–959. https://doi.org/10.1104/pp.114.246520
Zhang Z, Zhang J, Chen Y, Li R, Wang H, Wei J (2012) Genome-wide analysis and identification of HAK potassium transporter gene family in maize (Zea mays. L). Mol Biol Rep 39(8):8465–8473. https://doi.org/10.1007/s11033-012-1700-2
Zou N, Li B, Dong G, Kronzucker HJ, Shi W (2012) Ammonium-induced loss of root gravitropism is related to auxin distribution and TRH1 function, and is uncoupled from the inhibition of root elongation in Arabidopsis. J Exp Bot 63(10):3777–3788. https://doi.org/10.1093/jxb/ers068
Funding
This work was supported by Natural Science Foundation of Guangxi Province (2018GXNSFBA281103, 2020GXNSFAA259061).
Author information
Authors and Affiliations
Contributions
HL, CH, and YW designed the research. HL, HZ, HC, SJ, and LX performed the experiments and data analysis. HL wrote the manuscript. ZD, KW and YW contributed with valuable discussions. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Consent for publication
Not applicable.
Ethics approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Luo, Hb., Huang, Cm., Wei, Yw. et al. Uncovering expression and functional analysis of newly discovered high-affinity K+ transporter family members from sugarcane. J. Plant Biochem. Biotechnol. 31, 826–836 (2022). https://doi.org/10.1007/s13562-021-00726-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-021-00726-5