Abstract
The sucrose non-fermenting 1-related protein kinases (SnRKs) mediate plant signalling pathways in response to different cues by phosphorylating target proteins. While full complement of these genes has been reported in several families, such information remains obscure in Solanaceae to date. Through a genome-wide identification approach, we report 40 SnRK members in the tomato genome. Multiple alignments and phylogenetic analysis grouped the identified genes in SnRK1, SnRK2, and SnRK3 subfamilies. SnRK1 subfamily comprises only two members, whereas SnRK3 being the largest category, is constituted by 30 members in tomato. Ten members of the SnRK3 subfamily are present in four tandemly duplicated chromosomal blocks. Transcript profiling of these genes during tomato development identified SlSnRK2.9 and SlSnRK3.3 with root-specific and SlSnRK2.7b with ripening-preferential mRNA abundance. The mRNA profiling of these genes by RNA-sequencing and qPCR revealed that SlSnRK3.10a, SlSnRK3.15a, and SlSnRK3.26 are activated at varying levels under phosphate (Pi) starvation. In-silico analysis anticipated a common set of proteins interacts with the selected phosphorus starvation inducible (PSI) SlSnRK genes. Overall, the present study identifies a set of PSI SnRKs in tomato and provides a comprehensive framework for their functional characterization in Solanaceae in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The sucrose non-fermenting 1 (SNF1)-related protein kinases (SnRKs) belong to Ser/Thr protein kinase group of enzymes that regulate different stress responses and physiological processes by regulating downstream gene expression via activating different protein phosphorylation pathways (Hrabak et al. 2003). These proteins are highly conserved and mainly involved in abiotic and biotic stress responses in eukaryotes. Based on their gene structure and sequence similarity, these genes are classified into three subfamilies (SnRK1, SnRK2, and SnRK3) in plants (Halford and Hardie 1998; Hrabak et al. 2003). SnRK1, homologous to yeast and mammals, constitutes the smallest group among them and is universally present in eukaryotes. SnRK1 subfamily members are characterized by the presence of N-terminal protein kinase (Pkinase) domain, ubiquitin-associated (UBA) domain, and kinase-associated 1 (KA1) domain (Celenza and Carlson 1986). In yeast, SnRK1 plays a vital role in regulating carbon metabolism and energy status (Celenza and Carlson 1986). In contrast, both SnRK2 and SnRK3 subfamilies are unique to plants. SnRK2 members harbour one N-terminal Pkinase domain and second a C-terminal regulatory domain. This domain is further comprised of two subdomains, i.e., domain I and II. These domains are known to mediate protein–protein interaction (PPI) with type 2C protein phosphatases (PP2Cs) (Kulik et al. 2011). SnRK3 genes are the most diverse and constitute the largest group among the three subfamilies. Generally, SnRK3 subfamily members, which are also designated as CIPKs (CBL-interacting protein kinases), harbour three domains, including the N-terminal Pkinase domain, a NAF domain, and a PPI domain in the C-terminal regulatory region. Members of this subfamily are known to interact with calcium sensor calcineurin B-like proteins (CBLs) to mediate the calcium signalling pathway (Wang et al. 2015).
Besides their much explored regulatory roles in sugar starvation responses, SnRK1 members also control the developmental plasticity and resilience under diverse environmental conditions in plants (Jamsheer et al. 2021). On the other hand, SnRK2s and SnRK3 members primarily participate in abiotic stress responses, including drought, salt, and osmotic stress, and also regulate the abscisic acid (ABA) signal transduction pathway in plants (Kobayashi et al. 2004; Boudsocq et al. 2007; Fujita et al. 2009; Jia et al. 2015; Cheng et al. 2017). Manipulation of expression of SnRK2 members such as AtSnRK2.8 or OsSAPK4 using overexpression approach conferred enhanced tolerance to drought and salt, respectively, in transgenic plants (Umezawa et al. 2004; Diédhiou et al. 2008). SnRK3 kinases are also known to participate in the regulation of Na+, K+, and NO3− transportation. The famous salt overly sensitive (SOS) genes, involved in the maintenance of ionic homeostasis in plants though its Na+/H+ antiporter activity, belong to SnRK3 subfamily in plants (Qiu et al. 2002).
Despite their known roles in regulating the plant responses to several abiotic stresses, SnRK members have not been well-studied for their roles in phosphorus starvation response (PSR). Plants employ a wide range of adaptive responses, including physiological, morphological, biochemical, and molecular levels, to mitigate Pi deficiency. One of such mechanisms involves PHOSPHATE RESPONSE1 (PHR1) transcription-mediated transcriptional activation of phosphorus starvation inducible (PSI) genes (Rubio et al. 2001; Puga et al. 2017). Such response leads to activation of PSI genes such as high-affinity phosphate transporters, ribonucleases, acid phosphatases, and those involved in glycerolipid metabolism to salvage Pi from its reserves, including organic P. Sugar is a critical metabolite that regulates the extent of PSR. It is required for robust transcriptional activation of several PSI genes in plants (Hammond and White 2008; Akash et al. 2021). Recently, the AtSnRK1 gene has been reported to interact with two Pi transporters (Pht1;4 and PHO1) in Arabidopsis (Carianopol et al. 2020) and provided crucial evidence on the involvement of these genes in Pi starvation response in plants. However, the entire regulatory spectrum of SnRK proteins under Pi starvation remains to be elucidated in plants.
Based on the already published evidence, the SnRK family consists of 39 members in Arabidopsis (Hrabak et al. 2003; Kolukisaoglu et al. 2004), 48 in rice (Kobayashi et al. 2004; Kolukisaoglu et al. 2004), 44 in Brachypodium (Wang et al. 2015), 34 in Eucalyptus (Wang et al. 2019), and 114 in Brassica napus (Zhu et al. 2020). However, such information on SnRK members in tomato, a model plant for studying fruit development and ripening in Solanaceae, is missing. Further, knowledge of PSI SnRK genes in plants remains unclear. Such understanding is paramount, considering the critical role of sugars in determining the scale of transcriptional regulation of PSI genes in plants (Karthikeyan et al. 2007; Hammond and White 2008; Akash et al. 2021). Through a genome-wide search, we identified 40 SnRK members in the tomato genome. Phylogeny of these proteins grouped them in three subfamilies, SnRK3 being the largest group among them. Transcript profiling during plant growth and development and under Pi starvation revealed the identity of tissue/organ/developmental stage-specific and PSI SnRKs in tomato. We also observed transcriptional activation of PSI SnRKs upon withdrawal of sucrose from the growth medium. Overall, the present study provides a valuable resource for functional elucidation of SnRKs under nutrient stress in tomato and other Solanaceae crops in the future.
Materials and methods
Identification, structure, and phylogenetic analyses of tomato SnRKs
The non-redundant amino acid sequences of the 39 Arabidopsis SnRKs were retrieved from UniprotKB (https://www.uniprot.org/). These sequences were used as queries for BLASTP against the tomato genome proteins database (ITAG release 4.0) with the e-value threshold (≤ 1e-40, score > 150), yielding 88 putative SnRK members. All the potential tomato SnRK (SlSnRK) proteins were validated for the presence of conserved domains such as STKc_SnRK, CIPK_C, AMPKA_C, UBA, STKc_AMPK_alpha, and PKc_like superfamily with NCBI Conserved Domains Database (CDD) database (http://www.ncbi.nlm.nih.gov/cdd/). Based on the presence of the characteristic domains, the overall analysis confirmed the presence of at least 40 SnRK genes in tomato. Further motif analysis in these predicted proteins were done using Multiple Expectation Maximization for Motif Elicitation (MEME Suite 5.3.3; https://meme-suite.org/meme/tools/meme) tool. For motif analysis, parameters such as classic discovery mode, with site distribution zero or one occurrence (of a contributing motif site) per sequence and motif count for 10 motifs with a width between 6 and 50 wide (inclusive) were chosen. The motifs were used as a query sequence in NCBI-CDD tool to check their molecular functions. Protein features of the identified members were analysed using Expasy Protparam (https://www.expasy.org/). Information on the intron–exon architecture was obtained from the tomato genome database (https://solgenomics.net/). For phylogenetic analysis, complete amino acid sequences of 39 AtSnRKs and 40 SlSnRKs (identified in this study) were used for multiple alignments using ClustalW in MEGA7 (Tamura et al. 2013). The evolutionary history was inferred by using the Maximum Likelihood method and Poisson correction. A phylogenetic tree was constructed with bootstrap values for 1000 replicates. The cis-acting regulatory element was identified in the promoter of the selected genes by retrieving a 2 kb upstream sequence SGN tomato genome database. PlantPAN3.0 (http://plantpan.itps.ncku.edu.tw/index.html) was used to predict the phosphate starvation response 1-binding sequence (P1BS; GnATATnC) and W-box elements in the selected promoters.
Data mining for transcript profiling of SlSnRKs
To investigate the mRNA abundance of the identified SlSnRK genes during tomato development, we mined the RNA-sequencing data, published by (Sato et al. 2012), as described previously (Kumar et al. 2015, 2016). Log10 values of the retrieved average RPKM values were used to create a heat map using MORPHEUS (https://software.broadinstitute.org/morpheus/). For the identification of PSI SlSnRKs, the data published by (Pfaff et al. 2020) and in-house unpublished RNA-seq data were analysed, as described previously (Akash et al. 2021). The log10 values of the fold change of transcripts per million (low Pi versus high Pi) for each transcript were used to develop a heat map as described earlier.
Quantitative real-time PCR (qPCR) of selected SlSnRKs under nutrient deficiency
To validate the activation of selected SlSnRKs under Pi deficiency, we used the cDNA samples for + P + S (a half-strength MS media with 2% sucrose), −P + S (a half-strength MS media without KH2PO4 but with 2% sucrose), and + P − S (a half-strength MS media without 2% sucrose) prepared previously (Akash et al. 2021). Gene-specific primers were designed using Primer3plus (https://primer3plus.com/). Primers used in the present study are listed in Supplementary Table S1. The qPCR reactions were carried out using KAPA SYBR FAST qPCR Master Mix (2X) (Roche, Basel, Switzerland) in the StepOnePlus™ Real-Time PCR System (Applied Biosystems, USA). The ΔΔCt method was selected to calculate SnRKs genes' relative expression level (Livak and Schmittgen 2001). For normalization of qPCR data, the Glyceraldehyde-3-phosphate dehydrogenase (Solyc05g014470; GAPDH) gene was used. Error bars in the qPCR data show the mean ± standard deviation of two biological and three technical replicates.
In-silico protein–protein interaction
The functional protein interaction potential of the selected PSI SlSnRKs was investigated using STRING (https://string-db.org/). The amino acid sequence of the candidate SlSnRK was used as a query sequence. The output file was saved as 'svg' and manually modified for the names of the interacting proteins. The list of all interactors with the predicted score is provided in Supplementary Table S2.
Results and discussion
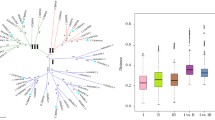
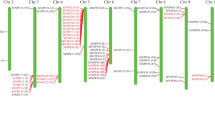
The gene search identified 40 putative SnRK members in the tomato genome (Fig. 1; Table 1). The number of the identified members in tomato are close to the already reported SnRKs homologs in Arabidopsis (Hrabak et al. 2003; Kolukisaoglu et al. 2004), rice (Kobayashi et al. 2004; Kolukisaoglu et al. 2004), Eucalyptus grandis (Wang et al. 2019) and Brachipodium distachyon (Wang et al. 2015), suggesting conservation in the number of these genes in many monocot and dicot species. These genes were found to be non-uniformly distributed on all 12 chromosomes. Chromosome 6 harboured a maximum of eight SlSnRKs (Table 1). Tandem duplication events seem to have moderately contributed to the expansion of this family as four such blocks of duplicated genes, distributed at chromosome 2 and 9 (one block each) and 6 (two blocks), were observed (Table 1). Most of these proteins were predicted to be localized to the cytoplasm, followed by chloroplast and nucleus. In line with previous studies, the size of full SlSnRK proteins ranged from 303 (SlSnRK3.15b) to 542 (SlSnRK1.1), with molecular weight ranging between 40 and 60 kDa (Hrabak et al. 2003; Wang et al. 2019). Analysis of their genomic loci structure revealed members both with and without introns. In total, 18 members, constituting 45% of the identified SnRKs, lacked any intron in their genomic sequence, whereas the genomic loci of SnRK1.1 harboured a maximum of 14 introns (Table 1). Phylogenetic analysis was carried out to understand the relatedness of tomato SnRKs with their Arabidopsis homologs. This scrutiny subgrouped these proteins into three subfamilies. SnRK1 subfamily (group I) comprises two members, whereas eight members belong to the SlSnRK2 subfamily (Fig. 1). In common with earlier published results, the SnRK3 subfamily remained the largest group among the three subfamilies and comprised 30 members in tomato (Hrabak et al. 2003; Kolukisaoglu et al. 2004). MEME motif analysis clearly revealed the absence of motifs 6, 9 and 10 from SnRK1 and SnRK2 members (Fig. 2). While motifs 6 and 9 correspond to AMPKA_C_alpha subunits with kinase function, no function for the novel motif 10 could be established. Motifs 1–5, 7 and 8 were predicted to have PKC protein kinase activity.
Phylogenetic analysis of Arabidopsis and tomato SnRK proteins. The multiple alignment of 39 AtSnRKs and 40 SlSnRKs were done using ClustalW. A Maximum Likelihood tree was created using MEGA7 with bootstrap values for 1000 iterations. Phylogeny grouped these genes in three clades, including group I (SnRK1), II (SnRK2) and III (SnRK3). Purple, green and red colours represent group I, II and III, respectively
Analysis of conserved motifs in SnRK proteins. The motifs (1–10) were identified using MEME search tool. The length and order of the boxes represents the actual length and position of the conserved domain in the protein sequence. Symbol and consensus sequence of each motif is presented at the bottom of the figure
RNA-sequencing is a powerful approach to study gene expression in tomato (Kumar and Khurana 2014). Next, we studied the mRNA abundance of the identified SlSnRKs during the vegetative and reproductive stages of tomato development using RNA-sequencing data available online. While most of the genes belonging to all subfamilies remained ubiquitously expressed, transcripts of SlSnRK2.9 and SlSnRK3.3 were found to be root-specific. Similarly, SlSnRK3.6b remained floral organ-specific, and SlSnRK2.7b showed ripening-preferential transcript accumulation (Fig. 3). Transcripts of three tandemly duplicated genes, including SlSnRK3.1c, SlSnRK3.1d, and SlSnRK3.1e on chromosome 6 remained undetected, indicating that these genes might be still undergoing selection. Most of the tandemly duplicated genes in a block exhibited different expression profiles from each other, indicating that the duplicated gene might have evolved for a new function. After obtaining proof of their expression, we examined transcript accumulation of these genes in tomato root, shoot, and whole seedlings under Pi deficiency, as described previously (Akash et al. 2021). This investigation identified seven genes, including SlSnRK2.7a, SlSnRK3.1a, SlSnRK3.10b, SlSnRK3.15a, SlSnRK3.24, SlSnRK3.25, and SlSnRK3.26, with moderate to strongly activated transcripts at multiple time points in the least two of the three tissues (Fig. 4a). Activation of a few genes such as SlSnRK2.8a, SlSnRK3.14a, SlSnRK3.22, and SlSnRK3.23, only in one of the three tissues, was also observed. In contrast, SlSnRK3.3, SlSnRK3.8, SlSnRK3.12, and SlSnRK3.19 were strongly down-regulated in at least two of the three tissues under Pi starvation. The SnRK1 subfamily members remained mainly unaffected, whereas SnRK3 members were the most affected genes concerning their altered transcriptional regulation under Pi starvation (Fig. 4a). Next, we selected five genes (SlSnRK2.7a, SlSnRK3.1a, SlSnRK3.10b, SlSnRK3.15a, and SlSnRK3.26) for validation of their PSI nature, as observed in RNA-seq data, by qPCR. Among the selected five genes, qPCR confirmed the activation of only three genes, including SlSnRK3.10b, SlSnRK3.15a, and SlSnRK3.26 under Pi starvation (Fig. 4b). One reason for the failure of activation of SlSnRK2.7a and SlSnRK3.1a in the qPCR analysis could be attributed to their spatiotemporally restricted transcriptional regulation under Pi deficiency. For example, SlSnRK2.7a exhibited the highest induction after 1-day of Pi starvation treatment in roots in the RNA-seq data. However, we validated their PSI nature in whole seedlings at 7-day after Pi deficiency, which might have contributed to the observed variation between the RNA-seq and qPCR data. Overall, confirmation of the PSI nature of the SlSnRK2.7a and SlSnRK3.1a warrants further investigations, especially in more tissues and at additional time points of the Pi starvation. Upon further analysis of their promoters for the presence of atypical P1BS elements, we noticed that SlSnRK2.7a, SlSnRK3.10b, and SlSnRK3.26 harboured one to three elements while SlSnRK3.1a and SlSnRK3.15a lacked any of it (Fig. 4c). Overall, SlSnRK3.10b and SlSnRK3.26 showed a positive correlation between the P1BS element and activation of these genes under Pi starvation. Concerning SlSnRK3.15a, the PSI nature could be attributed to other additional cis-acting regulatory element such as W-box (TTGACY), as suggested previously (Srivastava et al. 2020). The putative promoters of SlSnRK3.3, SlSnRK3.10a, SlSnRK3.15a, SlSnRK3.24, and SlSnRK3.25 PSI genes, lacking a P1BS element, were predicted to have one to six W-box elements (Fig. 4c; Supplementary figure Sf1). Interestingly, besides their PSI nature, transcript levels of the selected five genes were found to be upregulated in the absence of exogenous sucrose supply (Fig. 4b). This observation is common with their earlier known roles and places these members in sugar starvation in tomato (Jamsheer et al. 2021 and references therein).
Transcript profiling of SlSnRKs during tomato development. In this analysis, average RPKM values of the identified SlSnRK genes were retrieved from Sato et al. (2012). Log10 of the RPKM values for each stage/tissue/organ was used to generate a heatmap using MORPHEUS (https://software.broadinstitute.org/morpheus/). Hierarchical clustering with Euclidean distance method was performed to group genes with similar profiles together. R, root; L, leaf; Fl, flower, Ft-1, -2, and -3 represent young fruits of 1 cm, 2 cm, or 3 cm diameter, respectively; MG, mature green fruit; B, breaker stage fruit; RR, red-ripe stage fruit. Gene names are given on the right side of the heatmap and colour bar at the bottom represents the range of mRNA abundance
Transcript accumulation of SlSnRK genes under Pi starvation. a In-house normalized and the retrieved RNA-seq data, published recently by Pfaff et al. (2020), were mined to investigate mRNA levels of SlSnRKs under Pi deficiency. Heatmap was generated by taking the log10 fold change of relative transcripts per million (TPM) values (treatment vs. control) for each gene using MORPHEUS. Number 1d, 3d, 7d and 15d represents the respective days of Pi starvation. L, leaf; R, root; WS, whole seedling. The colour bar at the bottom represents the minimum and maximum values of gene expression. b Validation of PSI nature of the selected SlSnRK genes using qPCR. The mRNA levels of the selected PSI SlSnRKs were in + P + S, − P + S or + P − S grown seedlings, as described earlier (Akash et al. 2021). The relative expression of each gene in − P + S or + P − S was calculated against their expression in + P + S grown seedlings. (C) Analysis of P1BS and W-box cis-regulatory elements in the 2-kb upstream promoter region. PlantPAN3.0 (http://plantpan.itps.ncku.edu.tw/index.html) was used to predict these elements in the selected promoters
In Arabidopsis, SnRK1 members have been recently reported to interact with Phosphate transporter 1;4 (PHT1; 4) and PHO1 homolog 7 (PHO1; H7) (Carianopol et al., 2020). Although moderate but consistent downregulation of two SlSnRK1 members, SlSnRK1.2 and SlSnRK3.18 in Pi-starved tissues, is interesting and demands further investigation into their plausible regulatory roles in tomato. Due to kinase activity, these proteins are likely to phosphorylate other proteins to regulate stress response in plants. Recently, using the yeast-2-hybrid approach, several ABA-responsive interactors of SnRKs members have been identified in Arabidopsis (Cheng et al., 2017; Carianopol et al., 2020). Therefore, we predicted interacting partners of the selected SlSnRK members (SlSnRK1.2; SlSnRK2.7a, SlSnRK3.1a, SlSnRK3.3, SlSnRK3.15a, and SlSnRK3.26) from each class. Interestingly, all members included in this analysis were anticipated to interact with Solyc01g099280 (SNF4-homolog), Solyc05g018300 (a protein serine/threonine phosphatases), Solyc03g118070 (an unannotated protein), and Solyc11g007830 (a protein tyrosine phosphatase) (Fig. 5). SlSnRK1.2 and SlSnRK2.7a may interact with additional protein phosphatase 2C (PP2C) as well. In common with earlier studies, only SnRK3 subfamily members showed their interaction potential with calmodulin (CAM) proteins (Wang et al. 2015). SnRK3.1a interacted with CAM such as Solyc03g098050 and Solyc10g077010. Similarly, SnRK3.15a interacted with Solyc06g051970 (Calcium-binding EF-hand family protein). Cheng et al. (2017) have recently demonstrated the involvement of an SCF-complex, which regulates ABA signalling in Arabidopsis by interacting with SnRK2.6. During ABA signalling, a pyrabactin resistance (PYR)/PYR1-like (PYL)/regulatory components of ABA receptor (RCAR) proteins interact with co-receptor protein phosphatase type 2Cs for the activation of SnRK2 kinases. Upon activation, SnRK2 members switch on ABA signalling by phosphorylating various target proteins (Jia et al. 2015; Cheng et al. 2017). The authors demonstrated that the protein turnover of SnRK2 member, SnRK2.3, is regulated by its interaction with a phloem protein2-B (PP2-B) protein PP2-B11 in Arabidopsis. Therefore, interactions between protein phosphatase 2C, F-box protein, and SnRK2 members control ABA signalling in Arabidopsis. The predicted protein interaction data for SlSnRKs is interesting, especially after the recent identification of PP2C interacting PSI F-box genes (Akash et al. 2021), and demands further investigations to validate these anticipated interactions and confirm if a similar SCF-SnRKs-PP2C regulatory module exists to control PSR in tomato. Given the importance of sucrose in determining the amplitude of transcriptional activation of many PSI genes (Hammond and White 2008), we speculate the existence of a similar regulatory module controlling PSRs in plants.
Identification of putative interactors of the selected SlSnRK members. In-silico protein- protein interaction was studied using STRING (https://string-db.org/). This analysis identified several putative interactors for each protein. Only SnRK3 members were anticipated to interact with calmodulin (CAM) proteins. The colours of the lines represent the different modes by which the interaction was predicted. Light green, text mining; red, gene fusions; purple, experimentally determined; cyan, from curated databases
In conclusion, we, for the first time, report the identification of 40 SnRK members in the tomato genome. Gene structure analysis grouped the identified members in three subfamilies. Transcript profiling revealed the ubiquitous or stage/tissue/organ-specific nature of SnRKs. qPCR analysis confirmed SlSnRK3.10b, SlSnRK3.15a, and SlSnRK3.26 as the PSI SnRKs. The unavailability of exogenous sucrose supply also activated the transcription of SlSnRK2.7a, SlSnRK3.1a, SlSnRK3.10b, SlSnRK3.15a, and SlSnRK3.26. The in-silico protein-interaction analysis anticipated a common set of proteins, including SNF4-homolog, PP2C protein, and a protein tyrosine phosphatase, interacting with all the selected tomato SnRK members. Altogether, the data presented here identify PSI SlSnRK genes and provide a scaffold for their functional elucidation in tomato in the future.
Abbreviations
- ABA:
-
Abscisic acid
- B:
-
Breaker
- CBL:
-
Calcineurin B-like protein
- CAM:
-
Calmodulin
- CIPK:
-
CBL-interacting protein kinase
- RCAR:
-
Components of ABA receptor
- CDD:
-
Conserved Domains Database
- Ct:
-
Threshold cycle
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- K:
-
Potassium
- kDa:
-
Kilo-Dalton
- KA1:
-
Kinase-associate 1
- MG:
-
Mature green
- MEGA:
-
Molecular Evolutionary Genetics Analysis
- Na:
-
Sodium
- NO3 − :
-
Nitrate
- qPCR:
-
Quantitative real-time PCR
- P:
-
Phosphorus
- Pi:
-
Phosphate
- PHR1:
-
PHOSPHATE RESPONSE 1
- PSI:
-
Phosphorus starvation inducible
- PPI:
-
Protein–protein interaction
- PSR:
-
Phosphorus starvation response
- Pht1:
-
Phosphate transporter 1
- PHO1:
-
PHOSPAHTE1
- P1BS:
-
Phosphate starvation response 1-binding sequence
- PP2-B:
-
Phloem protein 2-B
- PP2C:
-
Protein phosphatase 2C
- PYL:
-
Pyrabactin resistance, PYR; PYR1-like
- RR:
-
Red-ripe
- RNA-seq:
-
RNA sequencing
- RPKM:
-
Reads per kilo base per million mapped reads
- SOS:
-
Salt overly sensitive
- S:
-
Sucrose
- Ser/Thr:
-
Serine/Threonine
- SNF1:
-
Sucrose non-fermenting 1
- SnRKs:
-
Sucrose non-fermenting 1-related protein kinases
- TPM:
-
Transcripts per million
- UBA:
-
Ubiquitin-associated
References
Akash, Parida AP et al (2021) Identification, evolutionary profiling, and expression analysis of F-box superfamily genes under phosphate deficiency in tomato. Plant Physiol Biochem 162:349–362
Boudsocq M, Droillard MJ, Barbier-Brygoo H, Laurière C (2007) Different phosphorylation mechanisms are involved in the activation of sucrose non-fermenting 1 related protein kinases 2 by osmotic stresses and abscisic acid. Plant Mol Biol 63:491–503
Carianopol CS, Chan AL, Dong S et al (2020) An abscisic acid-responsive protein interaction network for sucrose non-fermenting related kinase1 in abiotic stress response. Commun Biol 3:1–5
Cheng C, Wang Z, Ren Z et al (2017) SCFAtPP2-B11modulates ABA signaling by facilitating SnRK2.3 degradation in Arabidopsis thaliana. PLoS Genet 13:e1006947
Diédhiou CJ, Popova OV, Dietz KJ, Golldack D (2008) The SNF1-type serine-threonine protein kinase SAPK4 regulates stress-responsive gene expression in rice. BMC Plant Biol 8:49
Fujita Y, Nakashima K, Yoshida T et al (2009) Three SnRK2 protein kinases are the main positive regulators of abscisic acid signaling in response to water stress in arabidopsis. Plant Cell 50:2123–2132
Halford NG, Hardie DG (1998) SNF1-related protein kinases: Global regulators of carbon metabolism in plants? . Plant Mol Biol 37:735–748
Hammond JP, White PJ (2008) Sucrose transport in the phloem: Integrating root responses to phosphorus starvation. J Exp Bot 59:93–109
Hrabak EM, Chan CWM, Gribskov M et al (2003) The Arabidopsis CDPK-SnRK superfamily of protein kinases. Plant Physiol 132:666–680
Jamsheer K M, Kumar M, Srivastava V (2021) SNF1-related protein kinase 1: the many-faced signaling hub regulating developmental plasticity in plants. J Exp Bot erab079
Jia F, Wang C, Huang J et al (2015) SCF E3 ligase PP2-B11 plays a positive role in response to salt stress in Arabidopsis. J Exp Bot 66:4683–4697
Karthikeyan AS, Varadarajan DK, Jain A et al (2007) Phosphate starvation responses are mediated by sugar signaling in Arabidopsis. Planta 225:907–918
Kobayashi Y, Yamamoto S, Minami H et al (2004) Differential activation of the rice sucrose nonfermenting1-related protein kinase2 family by hyperosmotic stress and abscisic acid. Plant Cell 16:1163–1177
Kolukisaoglu Ü, Weinl S, Blazevic D et al (2004) Calcium sensors and their interacting protein kinases: genomics of the Arabidopsis and rice CBL-CIPK signaling networks. Plant Physiol 134:43–58
Kulik A, Wawer I, Krzywińska E et al (2011) SnRK2 protein kinases - key regulators of plant response to abiotic stresses. Omi A J Integr Biol 15:859–872
Kumar R, Khurana A (2014) Functional genomics of tomato: opportunities and challenges in post-genome NGS era. J Biosci 39:917–929
Kumar R, Agarwal P, Pareek A et al (2015) Genomic survey, gene expression, and interaction analysis suggest diverse roles of ARF and Aux/IAA proteins in solanaceae. Plant Mol Biol Rep 33:1552–1572
Kumar R, Chauhan PK, Khurana A (2016) Identification and expression profiling of DNA methyltransferases during development and stress conditions in solanaceae. Funct Integr Genom 16:513–528
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408
Pfaff J, Denton AK, Usadel B, Pfaff C (2020) Phosphate starvation causes different stress responses in the lipid metabolism of tomato leaves and roots. Biochim Biophys Acta Mol Cell Biol Lipids 1865:158763
Puga MI, Rojas-Triana M, de Lorenzo L et al (2017) Novel signals in the regulation of Pi starvation responses in plants: facts and promises. Curr Opin Plant Biol 39:40–49
Qiu QS, Guo Y, Dietrich MA et al (2002) Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc Natl Acad Sci U S A 99:8436–8441
Rubio V, Linhares F, Solano R et al (2001) A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev 15:2122–2133
Sato S, Tabata S, Hirakawa H et al (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485:635–641
Srivastava R, Akash PAP et al (2020) Identification, structure analysis, and transcript profiling of purple acid phosphatases under Pi deficiency in tomato (Solanum lycopersicum L.) and its wild relatives. Int J Biol Macromol 165:2253–2266
Tamura K, Stecher G, Peterson D et al (2013) MEGA6: molecular evolutionary genetics analysis version 60. Mol Biol Evol 30:2725–2729
Umezawa T, Yoshida R, Maruyama K et al (2004) SRK2C, a SNF1-related protein kinase 2, improves drought tolerance by controlling stress-responsive gene expression in Arabidopsis thaliana. Proc Natl Acad Sci U S A 101:17306–17311
Wang L, Hu W, Sun J et al (2015) Genome-wide analysis of SnRK gene family in Brachypodium distachyon and functional characterization of BdSnRK2.9. Plant Sci 237:33–45
Wang Y, Yan H, Qiu Z et al (2019) Comprehensive analysis of SNRK gene family and their responses to salt stress in Eucalyptus grandis. Int J Mol Sci 20:2786
Zhu W, Wu D, Jiang L, Ye L (2020) Genome-wide identification and characterization of SnRK family genes in Brassica napus. BMC Plant Biol 20:287
Acknowledgements
AK acknowledges Dr. Rahul Kumar, Department of Plant Sciences, University of Hyderabad, for the critical reading of the manuscript. AK also thank him for sharing his lab resources and recognize his funding institutions, the Science and Engineering Research Board (SERB), Government of India (CRG/2018/001033), and IoE, MHRD (RC1-20-018), under which the resources used in the present study were generated. AK thanks Dr. Masroor Ahmad Beg, the Principal, Zakir Husain Delhi College, University of Delhi for his support. Akash thanks the Council of Scientific and Industrial Research (CSIR), Government of India, for JRF and SRF fellowships. AR thanks the Science and Engineering Research Board (SERB), Government of India (CRG/2018/001033), for his project assistant position and fellowship.
Author information
Authors and Affiliations
Contributions
AK conceived the idea and wrote the original draft. Akash performed transcript profiling experiments. AR performed the promoter architecture and motif analyses. AK, Akash and AR together carried out all bioinformatics analyses. Both authors read and approve the final draft.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Khurana, A., Akash & Roychowdhury, A. Identification of phosphorus starvation inducible SnRK genes in tomato (Solanum lycopersicum L.). J. Plant Biochem. Biotechnol. 30, 987–998 (2021). https://doi.org/10.1007/s13562-021-00701-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-021-00701-0