Abstract
Transcription factors (TFs) are pivotal players in plant stress signaling and signal transduction pathways. Among the key TFs, NAC, ZF-HD, AP2-EREBP, WRKY and bHLH proteins play crucial roles in the regulation of reprogramming the transcriptome and associated responses in stress. Considering this, genome-wide identification of NAC, ZF-HD, AP2-EREBP, WRKY and bHLH TF families were performed in the C3 model plant, Oryza sativa. The computational study identified 144 NAC, 15 ZF-HD, 164 AP2-EREBP, 103 WRKY, and 135 bHLH proteins and their physicochemical properties and, expression profiling by computational analysis. Genome-wide in silico expression analysis of NAC, ZF-HD, AP2-EREBP, WRKY, bHLH genes showed their differential expression profiling in different tissues. Expression patterns, gene structure, subcellular localization, gene ontology of 17 NAC, 3 ZF-HD, 13 AP2-EREBP, 11 WRKY, 8 bHLH key genes suggested the putative novel variants in stress and signal transduction. These key players are needed to be studied in order to categorize and outline their functional roles in AbS signaling network.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A large number of the world’s population depend on rice for future food requirement. Unlike animals, plants are exposed to diverse environmental stresses. Especially abiotic stresses (AbS’s) are predominant, which negatively affects their growth and development of plants thereby reduction in productivity. Drought, cold, salinity, flood, submergence are the major stressors responsible for substantial yield loss. At the molecular and physiological level, expression and signalling of several genes are affected by these stresses, which known as abiotic stress responsive (AbSR) genes (Hirayama and Shinozaki 2010; Sharoni et al. 2011; Hadiarto and Tran 2011; Lata et al. 2015). Among many AbSR genes, transcription factors (TFs) are the crucial targets to understand the molecular cross-talks of AbS responses and to conducting overexpression studies in plants for AbS tolerance, they can act as key regulators overdriving the expression of several target genes (Nakashima et al. 2009; Urano et al. 2010; Yang et al. 2010; Golldack et al. 2011; Mickelbart et al. 2015). Plant genome contains around 7% TFs (Udvardi et al. 2007), which are categorized into 58 TF families (Jin et al. 2014). Among these, NAC, ZF-HD, AP2-EREBP, WRKY, bHLH are the largest TF families, excluding ZF-HD TFs (http://grassius.org/grasstfdb.php) (Singh et al. 2002; Gong et al. 2004, Xiong et al. 2005).
Various investigation and available literatures reveal the regulatory role of NAC, ZF-HD, AP2-EREBP, WRKY and bHLH TF families in signal transduction and inflection of several physiological and molecular processes including somatic embryogenesis (Toledo-Ortiz et al. 2003; Albrecht et al. 2005; Sonnenfeld et al. 2005; Mittler et al. 2006; Xu et al. 2008), plant development (Shigyo and Ito 2004), pollen development and function (Guan et al. 2014), internode elongation (Hattori et al. 2009), biosynthesis of secondary metabolites (Ma et al. 2009; Suttipanta et al. 2011), seed development (Johnson et al. 2002; Luo et al. 2005), seed dormancy (Rushton et al. 2010; Ding et al. 2014), biomass (Yu et al. 2013), flowering period and plant height (Cai et al. 2014), leaf senescence (Miao et al. 2004; Guo and Gan 2006), anther and ovule development (Zhao et al. 2008) and hormone signaling (Rashotte and Goertzen 2010; Feller et al. 2011; Hu et al. 2013). Significantly NAC, ZF-HD, AP2-EREBP, WRKY, bHLH TF families are activated in response to diverse biotic stress (Shinozaki et al. 2003; Cao et al. 2006; Singh et al. 2010; Xia et al. 2010; Muthamilarasan and Prasad 2013) including fungal invasion (Zheng et al. 2006; Marchive et al. 2007), virus attack (Huh et al. 2012), bacterial infection (Du and Chen 2000; Deslandes et al. 2002; Kim et al. 2008), disease resistance (Gutterson and Reuber 2004; Oh et al. 2005; Nakashima et al. 2007) and as well as AbS (Shinozaki et al. 2003; Cao et al. 2006; Singh et al. 2010; Xia et al. 2010; Tang et al. 2013) like drought and heat (Rizhsky et al. 2002; Kiribuchi et al. 2004, 2005; Sakuma et al. 2006; Wu et al. 2009; Zheng et al. 2009; Qiu and Yu 2009; Ren et al. 2010), salinity (Mukhopadhyay et al. 2004; Gutha and Reddy 2008; Qiu and Yu 2009; Zheng et al. 2009), cold (Pnueli et al. 2002; Wang et al. 2003; Mukhopadhyay et al. 2004; Kume et al. 2005; Qiu and Yu 2009), desiccation, submergence, heavy metals and wounding (Mukhopadhyay et al. 2004; Kiribuchi et al. 2004, 2005), osmotic stress (Gutha and Reddy 2008), low temperatures (Zheng et al. 2009; Sharoni et al. 2011) and phosphate starvation (Yi et al. 2005).
Therefore, the significant role of NAC, ZF-HD, AP2-EREBP, WRKY, bHLH TF families in several biological, molecular and physiological process were studied in assorted crop plants (Ledent and Vervoort 2001; Toledo-Ortiz et al. 2003; Ulker and Somssich 2004; Zheng et al. 2009; Rushton et al. 2010; Sharoni et al. 2011; Figueiredo et al. 2012; Chen et al. 2016). However, only minimum reports are available in the C3 model plant, O. sativa. For that reason, it is important to expedite the functional genomic approaches in the panicoideae family, especially C3 photosynthesis and AbS tolerance.
Based on the above addressed issues, the present study aimed at, in silico approaches to identify the potential AbSR encoding genes from rice NAC, ZF-HD, AP2-EREBP, WRKY, bHLH TF families. This is the first wide range investigation on these TFs O. sativa. The current study provides better insights about the functional aspects of these AbSR TFs, and unravels key genes for further validation toward describing their functional role in AbS dynamism.
Materials and methods
Database search for the identification of transcription factor family in O. sativa (L.)
Rice transcription factors (TF) like NAC, ZF-HD, AP2-EREBP, WRKY, bHLH and their protein sequences retrieved from the GRASSIUS Grass Regulatory Information Server (http://grassius.org/grasstfdb.html; Yilmaz et al. 2009).
Mining and meta-analysis of rice transcription factors
TF family members and their RAP-DB ID/Gene locus ID were collected. It was exposed to Rice Oligonucleotide Array Database (ROAD) meta-analysis search tool (Cao et al. 2012) for analyzing the tissues specific expression profile in different tissues in rice. The AbSR TF genes (Gene ID’s) were used to retrieve the corresponding genomic transcript, coding sequences with their chromosomal localization and protein sequence from RiceSRTFDB (Priya and Jain 2013).
Gene structure prediction
The genomic sequences and coding sequences of potential AbSR TFs like 17 NAC, 3 ZF-HD, 13 AP2-EREBP, 11 WRKY, 8 bHLH proteins were analyzed by GSDS web server v2.0 (Hu et al. 2015) to identify the position of exons and introns.
Physicochemical properties of identified proteins and phylogenetic analysis
The physicochemical properties including amino acid length, molecular weight, isoelectric point (pI), the number of positive/negatively charged residues, the instability index, and aliphatic index were predicted using the protparam tool of ExPASy (http://web.expasy.org/protparam/; Gasteiger et al. 2005). The 17 NAC, 3 ZF-HD, 13 AP2-EREBP, 11 WRKY, 8 bHLH protein sequences were imported into Phylogeny.fr (http://www.phylogeny.fr/; Dereeper et al. 2008) to construct a phylogenetic tree by maximum likelihood method.
Analysis of subcellular localization in TF families
The subcellular localization of TF family proteins of O. sativa was predicted using CELLO2GO (http://cello.life.nctu.edu.tw/cello2go/; Yu et al. 2014), Wolf Psort 2 (http://wolfpsort.org/; Horton et al. 2007), Bacello (http://gpcr.biocomp.unibo.it/bacello/; Pierleoni et al. 2006), ESLPred2 (http://www.imtech.res.in/raghava/eslpred2/; Garg and Raghava 2008), SubLoc (http://www.bioinfo.tsinghua.edu.cn/SubLoc/; Hua and Sun 2001).
Gene ontology analysis
Identified potential TF family members (Table 1) were loaded into CELLO2GO (http://cello.life.nctu.edu.tw/cello2go/; Yu et al. 2014) to find the Gene Ontology (GO) annotation against eukaryote. TF genes were characterized as per biological process, molecular function and cellular components according to CELLO2GO GO annotation.
Results
Identification of potent transcription factors (TF) in rice
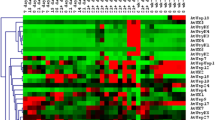
A total of 144 NAC, 15 ZF-HD, 164 AP2-EREBP, 103 WRKY and 135 bHLH TFs of rice with their gene ID’s were analyzed computationally for tissues specific (TS) gene expression studies. Seventeen NAC, 3 ZF-HD, 13 AP2-EREBP, 11 WRKY, 8 bHLH were potential abiotic stress responsive (AbSR) TF genes (Table 1), which have been involved in TS gene expression in all the 22 different tissues (Fig. 1a–e). This higher expression heatmap profiling was also evidenced among 5 different TF families, thereby reasonably delineating their function in tissue-specific manner.
Differential expression patterns of rice TF family genes. Heat map representing expression profiling of a NAC, b ZF-HD, c AP2-EREBP, d WRKY, e bHLH potent AbSR TF genes with respect to specific tissues. Yellow color indicates up regulation; Blue color indicates down regulation; Black color indicates unchanged expression level of AbSR TF genes. The colored scale bar at right side denotes relative expression value, where 5.0 and 13.0 represent down regulation and up regulation respectively (color figure online)
Structure of AbSR TF genes
Positions of exons and introns within the 17 NAC, 3 ZF-HD, 13 AP2-EREBP, 11 WRKY and 8 bHLH genes were predicted. Gene structure determination showed the numbers and arrangements of exons and introns (Fig. 2a–e and Table 1). The majority of AbSR TF genes (11; 21.153%) were found to contain a single intron, while 7 genes (13.46%) have two introns. Six AbSR TF genes (11.538%) have three introns; whereas 8 genes (15.38%) have four introns. Three AbSR TF genes (5.769%) were found to contain six introns; while 4 genes (7.692%) contained five and 8 introns, respectively. One AbSR TF gene (1.92%) has 16 introns. Among these, 12 AbSR TF genes (23.07%) were found intronless (Fig. 2a–e and Table 1).
Gene ontology annotation
Gene characteristic features of TF families were analyzed by protein sequence using CELLO2GO and showed the putative involvement of these proteins in different molecular function, and biological process (Table 2). A large number of proteins were predicted to be involved in stress response, embryo development, anatomical structure development, metabolic and biosynthetic process, and signal transduction (Fig. 3a–e). The molecular functions of these proteins paralleled to transcriptional regulator activity, protein binding and DNA binding activity (Fig. 4a–e).
Protein properties of potentially expressed TFs
All five different TF families and their well-expressed protein properties were analyzed. Smallest and biggest amino acids with respect to molecular weight, pI ranging, stability index, aliphatic index and grand average of hydropathicity (GRAVY) of potential TF genes were determined (Table 3).
Phylogenetic analysis of TFs
To study the phylogenetic organization of the 17 NAC, 3 ZF-HD, 13 AP2-EREBP, 11 WRKY and 8 bHLH family, the imputed protein sequences were used to generate an unrooted phylogenetic tree (Fig. 5). The unrooted tree divided the potentially expressed TF family genes into 5 major groups (groups I–V) based on the conserved NAC, ZF-HD, AP2-EREBP, WRKY, bHLH domains and homology of TF family gene sequences. Twenty proteins belongs to group—I (9 NAC; 1 AP2 –EREBP; 5 WRKY; 5 bHLH), 8 to group—II (2 NAC; 1 WRKY; 2 AP2-EREBP; 3 ZF-HD), 5 to group—III (1 NAC; 4 AP2-EREBP), 4 to group—IV (4 NAC); 15 to group—V (1 NAC; 6 AP2-EREBP; 5 WRKY; 3 bHLH) (Fig. 5).
Subcellular localization of rice TFs
The 4 programes were classed into two based on their resolution as high resolution and low-resolution predictions. The prediction principles and the competences of the four programs described in literature (Horton et al. 2007; Garg and Raghava 2008; Pierleoni et al. 2006; Hua and Sun 2001). The prediction results for the key TF family genes among the five different TF families and summarized in Table 4. It revealed the localization of these key players and their products in nucleus.
Discussion
NAC, ZF-HD, AP2-EREBP, WRKY, and bHLH type of TF’s have been reported to play vital roles in regulating various plant processes and physiological responses like plant development, normal growth, response to environmental stimuli and involved in plant defenses and so on (Ledent and Vervoort 2001; Toledo-Ortiz et al. 2003; Ulker and Somssich 2004; Zheng et al. 2009; Rushton et al. 2010; Sharoni et al. 2011; Figueiredo et al. 2012; Chen et al. 2016). This class of TFs are one of the well-studied proteins whose mode of action, cross regulation in signaling, auto regulation and evolution have been reported (Singh et al. 2002; Shimono et al. 2007; Puranik et al. 2012; Bakshi and Oelmüller 2014; Chen et al. 2016). These TFs play a crucial role in conferring tolerance to various AbS’s that includes cold, salinity, drought (Hu et al. 2006; Nakashima et al. 2007; Hu et al. 2008; Zheng et al. 2009), low temperature (Zheng et al. 2009) in NAC; cold, drought, salinity, metal, submergence (Kreps et al. 2002; Mukhopadhyay et al. 2004; Vij and Tyagi 2006; Kilian et al. 2007; Figueiredo et al. 2012; Giri et al. 2013) in ZF-HD; salinity, drought (Sharoni et al. 2010; Hsieh et al. 2013), low temperature (Sharoni et al. 2010), submergence, flooding (Sharoni et al. 2010), osmotic stress (Hsieh et al. 2013) in AP2—EREBP; salinity, drought, heat (Li et al. 2009, 2011), cold (Zou et al. 2010), H2O2 (Song et al. 2009),ozone oxidative stress, UV radiation (Jiang and Deyholos 2009), sugar starvation (Song et al. 2010), phosphate deprivation (Chen et al. 2009) in WRKY; cold, drought (Shinozaki et al. 2003) in bHLH TF families.
Expression of NAC, ZF-HD, AP2-EREBP, WRKY, bHLH TFs in response to diverse AbS decodes their putative involvement in the regulation of signaling mechanisms associated with transcriptional reprogramming during physiological stress. In silico identification of NAC, ZF-HD, AP2-EREBP, WRKY, bHLH TFs has been analyzed in various crop plants and their expression profiling in response to many AbS’s have been well studied. To the best of our knowledge, so far no such study has been reported in rice which is a panicoid C3 model crop with potential tolerance to diverse AbS.
The current study, 144 NAC, 15 ZF-HD, 164 AP2-EREBP, 103 WRKY, 135 bHLH from O. sativa TF family members were identified (Yilmaz et al. 2009). According to the heat map data, well expressed 17 NAC, 3 ZF-HD, 13 AP2-EREBP, 11 WRKY, 8 bHLH potential abiotic stress responsive (AbSR) TF genes were retrieved and subjected to ROAD. These AbSR TFs genes were chosen for expression profiling under AbS. Publically available microarray hybridization of ROAD expression values showed tissue-specific expression patterns in the 27 tissues of all the identified (17 NAC, 3 ZF-HD, 13 AP2-EREBP, 11 WRKY, 8 bHLH) potential AbSR TF genes. High level expression of these TFs expressed genes from multiple tissues and their expression during the individual AbS’s, delineates their multiple roles in diverse biological process and molecular crosstalks. This data could be further exploited for selecting key genes showing distinct expression pattern for explaining their functional roles. It highlighted the genome-wide analysis of genes expressed in different tissues. We outlined information on co-regulation among TF genes under abiotic stress conditions (Cao et al. 2012; Muthuramalingam et al. 2017). Identified each AbSR TF genes involved in the molecular crosstalks of the plant stress response was predicted. Further, this heatmap data could pave the way for conducting over-expression studies of key genes in different plant tissues in order to develop the AbSR protein content in rice.
Phylogenetic analysis of the potentially expressed TFs genes in rice showed the genes are present in the same subclades/subgroups. In addition, a joined evolutionary tree was constructed from expressed rice 17 NAC, 3 ZF-HD, 13 AP2-EREBP, 11 WRKY, 8 bHLH proteins involved in diverse aspects of plant growth, development and involved in multiple abiotic stress tolerance mechanisms. The potent TF genes with the same functions revealed a tendency to cluster into one subgroup, which provided an vital resource to explore the functions of the TF genes. It implied that these potent TF genes may be involved in the responses to unique and combined AbS, and this hypothesis was also supported using computational tissues specific expression and gene ontology annotation.
These AbSR TFs encoding proteins have huge variations in length of amino acid, isoelectric point, molecular weight, aliphatic index, instability index and GRAVY values of these proteins. Additionally, subcellular localization of these proteins at independent organelles can be attributed to the presence of putative novel variants, which are prerequisite for further validation.
Emerging advancement of ultra-high- throughput omics tools and approaches, including molecular physiology and computational strategies, the pivotal role of NAC, ZF-HD, AP2-EREBP, WRKY, bHLH TFs in gene regulation and signal transduction mechanisms has been studied well in all major crops and tree species. Though, no such reports on these TFs have been conducted in O. sativa, considered as model systems for scrutinizing C3 photosynthesis and AbS tolerance mechanisms. Considering the significance of this crop and NAC, ZF-HD, AP2-EREBP, WRKY, bHLH TFs, the current study used a wide range of computational approaches to categorizing and characterize these TF gene family members. The identified TF genes were used for gene ontology annotation, gene structure prediction, physicochemical properties and subcellular localization prediction analysis. In toting, in silico expression profiling of 17 NAC, 3 ZF-HD, 13 AP2-EREBP, 11 WRKY, 8 bHLH genes were used to understand the different tissues specific differential expression profiling. As a result proving an important indication of their controlling and regulatory functions in AbS conditions.
Abbreviations
- AbS:
-
Abiotic stress
- AbSR:
-
Abiotic stress responsive
- GO:
-
Gene ontology
- TFs:
-
Transcription factors
References
Albrecht C, Russinova E, Hecht V, Baaijens E, de Vries S (2005) The Arabidopsis thaliana somatic embryogenesis receptor like kinases1 and 2 control male sporogenesis. Plant Cell 17:3337–3349
Bakshi M, Oelmüller R (2014) WRKY transcription factors: Jack of many trades in plants. Plant Signal. Behav 9:e27700
CaiY Chen X, Xie K, Xing Q, WuY Li J, Du C, Sun Z, Guo Z (2014) Dlf1, a WRKY transcription factor, is involved in the control of flowering time and plant height in rice. PLoS One 9(7):e102529
Cao Y, Song F, Goodmand RM, Zheng Z (2006) Molecular characterization of four rice genes encoding ethylene-responsive transcriptional factors and their expressions in response to biotic and abiotic stress. J Plant Physiol 163:1167–1178
Cao P, Jung KH, Choi D, Hwang D, Zhu J, Ronald PC (2012) The rice oligonucleotide array database: an atlas of rice gene expression. Rice. 5:17
Chen YF, Li L, Xu Q, Kong YH, Wang H, Wu WH (2009) The WRKY6 transcription factor modulates PHOSPHATE1 expression in response to low Pi stress in Arabidopsis. Plant Cell 21:3554–3566
Chen L, Han J, Deng X, Tan S, Li L, Li L, Zhou J, Peng H, Yang G, He G, Zhang W (2016) Expansion and stress responses of AP2/EREBP superfamily in Brachypodium Distachyon. Scientific reports 6:21623
Dereeper A, Guignon V, Blanc G, Audic S, Buffet S, Chevenet F, Dufayard JF, Guindon S, Lefort V, Lescot M, Claverie JM, Gascuel O (2008) Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res 36:W465–W469
Deslandes L, Olivier J, Theulieres F, Hirsch J, Feng DX, Bittner- Eddy P, Beynon J, Marco Y (2002) Resistance to Ralstonia solanacearum in Arabidopsis thaliana is conferred by the recessive RRS1-R gene, a member of a novel family of resistance genes. Proc Natl Acad Sci USA 99:2404–2409
Ding ZJ, Yan JY, Li GX, Wu ZC, Zhang SQ, Zheng SJ (2014) WRKY41 controls Arabidopsis seed dormancy via direct regulation Of ABI3 transcript levels not downstream of ABA. Plant J 79:810–823
Du L, Chen Z (2000) Identification of genes encoding receptor-like protein kinases as possible targets of pathogen-and salicylic acid- induced WRKY DNA-binding proteins in Arabidopsis. Plant J 24:837–847
Feller A, Machemer K, Braun EL, Grotewold E (2011) Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J 66:94–116
Figueiredo DD, Barros PM, Cordeiro AM, Serra TS, Lourenc T, Chander S, Oliveira MM, Saibo NJ (2012) Seven zinc-finger transcription factors are novel regulators of the stress responsive gene OsDREB1B. J Exp Bot 63(10):3643–3656
Garg A, Raghava GP (2008) ESLpred2: improved method for predicting subcellular localization of eukaryotic proteins. BMC Bioinform 9:503
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: Walker JM (ed) The proteomics protocols handbook. Springer, NewYork, pp 571–607
Giri J, Dansana PK, Kothari KS, Sharma G, Vij S, Tyagi AK (2013) SAPs as novel regulators of abiotic stress response in plants. BioEssays 35:639–648
Golldack D, Lu’king I, Yang O (2011) Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep 30:1383–1391
Gong W, Shen YP, Ma LG, Pan Y, Du YL, Wang DH, Yang JY, Hu LD, Liu XF, Dong CX, Ma L, Chen YH, Yang XY, Gao Y, Zhu D, Tan X, Mu JY, Zhang DB, Liu YL, DineshKumar SP, Li Y, Wang XP, Gu HY, Qu LJ, Bai SN, Lu YT, Li JY, Zhao JD, Zuo J, Huang H, Deng XW, Zhu YX (2004) Genome-wide ORFeome cloning and analysis of Arabidopsis transcription factor genes. Plant Physiol 135:773–782
GuanY Meng X, Khanna R, LaMontagne E, Liu Y, Zhang S (2014) Phosphorylation of a WRKY transcription factor by MAPKs is required for pollen development and function in Arabidopsis. PLoS Genet 10:e1004384
Guo Y, Gan S (2006) AtNAP, a NAC family transcription factor, has an important role in leaf senescence. Plant J 46:601–612
Gutha LR, Reddy AR (2008) Rice DREB1B promoter shows distinct stress-specific responses, and the overexpression of cDNA in tobacco confers improved abiotic and biotic stress tolerance. Plant Mol Biol 68:533–555
Gutterson N, Reuber TL (2004) Regulation of disease resistance pathways by AP2/ERF transcription factors. Curr Opin Plant Biol 7:465–471
Hadiarto T, Tran LS (2011) Progress studies of drought-responsive genes in rice. Plant Cell Rep 30:297–310
Hattori Y, Nagai K, Furukawa S, Song XJ, Kawano R, Sakakibara H, Wu J, Matsumoto T, Yoshimura A, Kitano H (2009) The ethylene response factors SNORKEL1 and SNORKEL2 allow rice to adapt to deep water. Nature 460:1026–1030
Hirayama T, Shinozaki K (2010) Research on plant abiotic stress responses in the post-genome era: past, present and future. Plant J. 61:1041–1052
Horton P, Park KJ, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K (2007) WoLF PSORT: protein localization predictor. Nucl Acids Res 35:W585–W587
Hsieh EJ, Cheng MC, Lin TP (2013) Functional characterization of an abiotic stress-inducible transcription factor AtERF53 in Arabidopsis thaliana. Plant Mol Biol 82:223–237
Hu H, Dai M, Yao J, Xiao B, Xianghua L, Zhang Q, Xiong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci USA 103:12987–12992
Hu H, You J, Fang Y, Zhu X, Qi Z, Xiong L (2008) Characterization of transcription factor gene SNAC2 conferring cold and salt tolerance in rice. Plant Mol Biol 67:169–181
Hu Y, Jiang L, Wang F, Yu D (2013) Jasmonate regulates the inducer of cbf expression-C-repeat binding factor/DRE binding factor1 cascade and freezing tolerance in Arabidopsis. Plant Cell 25:2907–2924
Hu B, Jin J, Guo A-Y, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31(8):1296–1297
Hua S, Sun Z (2001) Support vector machine approach for protein subcellular localization prediction. Bioinformatics 17:721–728
Huh SU, Choi LM, Lee GJ, Kim YJ, Paek KH (2012) Capsicum annuum WRKY transcription factor (CaWRKYd) regulates hypersensitive response and defense response upon Tobacco mosaic virus infection. Plant Sci 197:50–58
Jiang Y, Deyholos MK (2009) Functional characterization of Arabidopsis NaCl-inducible WRKY25 and WRKY33 transcription factors in abiotic stresses. Plant Mol Biol 69:91–105
Jin J, Zhang H, Kong L, Gao G, Luo J (2014) PlantTFDB3.0: a portal for the functional and evolutionary study of plant transcription factors. Nucl Acids Res 42:D1182–D1187
Johnson CS, Kolevski B, Smyth DR (2002) Transparent testa glabra2, a trichome and seed coat development gene of Arabidopsis, encodes a WRKY transcription factor. Plant Cell 14:1359–1375
Kilian J, Whitehead D, Horak J, Wanke D, Weinl S, Batistic O, D’Angelo C, Bornberg-Bauer E, Kudla J, Harter K (2007) The AtGenExpress global stress expression data set: protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J 50:347–363
Kim KC, Lai Z, Fan B, Chen Z (2008) Arabidopsis WRKY38 and WRKY62 transcription factors interact with histone deacetylase 19 in basal defense. Plant Cell 20:2357–2371
Kiribuchi K, Sugimori M, Takeda M, Otani T, Okada K, Onodera H, Ugaki M, Tanaka Y, Tomiyama-Akimoto C, Yamaguchi T, Minami E, Shibuya N, Omori T, Nishiyama M, Nojiri H, Yamane H (2004) RERJ1, a jasmonic acid-responsive gene from rice, encodes a basic helixloop—helix protein. Biochem Biophys Res Commun 325:857–863
Kiribuchi K, Jikumaru Y, Kaku H, Minami E, Hasegawa M, Kodama O, Seto H, Okada K, Nojiri H, Yamane H (2005) Involvement of the basic helix-loop-helix transcription factor RERJ1 in wounding and drought stress responses in rice plants. Biosci Biotechnol Biochem 69:1042–1044
Kreps JA, Wu Y, Chang HS, Zhu T, Wang X, Harper JF (2002) Transcriptome changes for Arabidopsis in response to salt, osmotic, and cold stress. Plant Physiol 130:2129–2141
Kume S, Kobayashi F, Ishibashi M, Ohno R, Nakamura C, Takumi S (2005) Differential and coordinated expression of Cbf and Cor/Lea genes during long-term cold acclimation in two wheat cultivars showing distinct levels of freezing tolerance. Genes Genet Syst 80:185–197
Lata C, Muthamilarasan M, Prasad M (2015) Drought stress responses and signal transduction in plants. In: Pandey GK (ed) Elucidation of abiotic stress signaling in plants. Springer, New York, pp 195–225
Ledent V, Vervoort M (2001) The basic helix-loop-helix protein family: comparative genomics and phylogenetic analysis. Genome Res 11(5):754–770
Li S, Fu Q, Huang W, Yu D (2009) Functional analysis of an Arabidopsis transcription factor WRKY25 in heat stress. Plant Cell Rep 28:683–693
Li S, Fu Q, Chen L, Huang W, Yu D (2011) Arabidopsis thaliana WRKY25, WRKY26, and WRKY33 coordinate induction of plant thermotolerance. Planta 233:1237–1252
Luo M, Dennis ES, Berger F, Peacock WJ, Chaudhury A (2005) Miniseed3 (MINI3), a WRKY family gene, and HAIKU2 (IKU2), a leucine- rich repeat (LRR) Kinase gene, are regulators of seed size in Arabidopsis. Proc Natl Acad Sci USA 102:17531–17536
Ma D, Pu G, Lei C, Ma L, Wang H, GuoY Chen J, Du Z, Wang H, Li G, Ye H, Liu B (2009) Isolation and characterization of AaWRKY1, an Artemisia annua transcription factor that regulates the amorpha- 4,11-diene synthase gene, a key gene of artemisin in biosynthesis. Plant Cell Physiol 50:2146–2161
Marchive C, Mzid R, Deluc L, Barrieu F, Pirrello J, Gauthier A, Corio-Costet MF, Regad F, Cailleteau B, Hamdi S, Lauvergeat V (2007) Isolation and characterization of a Vitis vinifera transcription factor, VvWRKY1, and its effect on responses to fungal pathogens in transgenic tobacco plants. J Exp Bot 58:1999–2010
MiaoY Laun T, Zimmermann P, Zentgraf U (2004) Targets of the WRKY53 transcription factor and its role during leaf senescence in Arabidopsis. Plant Mol Biol 55:853–867
Mickelbart MV, Hasegaw PM, Bailey-Serres J (2015) Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nat Rev Genet 16:237–251
Mittler R, Kim Y, Song LH, Coutu J, Coutu A, Ciftci-Yilmaz S, Lee H, Stevenson B, Zhu JK (2006) Gain- and loss-of-function mutations in Zat10 enhance the tolerance of plants to ablotic stress. FEBS Lett 580:6537–6542
Mukhopadhyay A, Vij S, Tyagi AK (2004) Overexpression of a zinc finger protein gene from rice confers tolerance to cold, dehydration, and salt stress in transgenic tobacco. Proc Natl Acad Sci USA 101:6309–6314
Muthamilarasan M, Prasad M (2013) Plant innate immunity: an updated insight into defense mechanism. J Bio Sci 38:433–449
Muthuramalingam P, Krishnan SR, Pothiraj R, Ramesh M (2017) Global transcriptome analysis of combined abiotic stress signaling genes unravels key players in Oryza sativa L.: an in silico approach. Front. Plant Sci 8:759
Nakashima K, Tran LS, Van Nguyen D, Fujita M, Maruyama K, Todaka D, Ito Y, Hayashi N, Shinozaki K, Yamaguchi-Shinozaki K (2007) Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress responsive gene expression in rice. Plant J 51:617–630
Nakashima K, Ito Y, Yamaguchi-Shinozaki K (2009) Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol 149:88–95
Oh SJ, Song SI, Kim YS, Jang HJ, Kim SY, Kim M, Kim YK, Nahm BH, Kim JK (2005) Arabidopsis CBF3/DREB1A and ABF3 in transgenic rice increased tolerance to abiotic stress without stunting growth. Plant Physiol 138:341–351
Pierleoni A, Martelli PL, Fariselli P, Casadio R (2006) BaCelLo: a balanced subcellular localization predictor. Bioinformatics 22:e408–e416
Pnueli L, Hallak-Her E, Rozenberg M, Cohen M, Goloubinoff P, Kaplan A, Mittler R (2002) Molecular and biochemical mechanisms associated with dormancy and drought tolerance in the desert legume Retama raetam. Plant J. 31:319–330
Priya P, Jain M (2013) RiceSRTFDB: a database of rice transcription factors containing comprehensive expression, cis-regulatory element and mutant information to facilitate gene function analysis. Database bat027
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17:369–381
Qiu Y, Yu D (2009) Over-expression of the stress-induced OsWRKY45 enhances disease resistance and drought tolerance in Arabidopsis. Env. Exp Bot 65:35–47
Rashotte AM, Goertzen LR (2010) The CRF domain defines cytokinin response factor proteins in plants. BMC Plant Biol 10:74
Ren X, Chen Z, Liu Y, Zhang H, Zhang M, Liu Q, Hong X, Zhu JK, Gong Z (2010) ABO3, a WRKY transcription factor, mediates plant responses to abscisic acid and drought tolerance in Arabidopsis. Plant J. 63:417–429
Rizhsky L, Liang H, Mittler R (2002) The combined effect of drought stress and heat shock on gene expression in tobacco. Plant Physiol 130:1143–1151
Rushton PJ, Somssich IE, Ringler P, Shen QJ (2010) WRKY transcription factors. Trends Plant Sci 15:247–258
Sakuma Y, Maruyama K, Qin F, Osakabe Y, Shinozaki K, Yamaguchi-Shinozaki K (2006) Dual function of an Arabidopsis transcription factor DREB2A in water-stress-responsive and heat-stress-responsive gene expression. Proc Natl Acad Sci USA 103:18822–18827
Sharoni AM, Nuruzzaman M, Satoh K, Shimizu T, Kondoh H, Sasaya T, Choi IR, Omura T, Kikuchi S (2010) Gene structures, classification and expression models of the AP2/EREBP transcription factor family in rice. Plant and cell physiol 52:344–360
Sharoni AM, Nuruzzaman M, Satoh K, Shimizu T, Kondoh H, Sasaya T, Choi IR, Omura T, Kikuchi S (2011) Gene structures, classification and expression models of the AP2/EREBP transcription factor family in rice. Plant Cell Physiol 52(2):344–360
Shigyo M, Ito M (2004) Analysis of gymnosperm two-AP2-domain-containing genes. Dev Genes Evol 214:105–114
Shimono M, Sugano S, Nakayama A, Jiang CJ, Ono K, Toki S, Takatsuji H (2007) Rice WRKY45 plays a crucial role in benzothiadiazole inducible BLAST resistance. Plant Cell 19:2064–2076
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6(5):410–417
Singh KB, Foley RC, Oñate-Sánchez L (2002) Transcription factors in plant defense and stress responses. Curr Opin Plant Biol 5(5):430–436
Singh AK, Sopory SK, Wu R, Singla-Pareek SL (2010) Transgenic approaches. In: Pareek A, Sopory SK, Bohnert HJ, Govindjee (eds.) Abiotic stress adaptation in plants: physiological, molecular and genomic foundation, Springer, Berlin, Germany, pp 417–450
Song Y, Jing SJ, Yu DQ (2009) Overexpression of the stress induced OsWRKY08 improves the osmotic stress tolerance in Arabidopsis. Chin Sci Bull 54:4671–4678
Song Y, Chen LG, Zhang LP, Yu DQ (2010) Overexpression of OsWRKY72 gene interferes in the ABA signal and auxin transport pathway of Arabidopsis. J Biosci 35:459–471
Sonnenfeld MJ, Delvecchio C, Sun X (2005) Analysis of the transcriptional activation domain of the Drosophila tango bHLH-PAS transcription factor. Dev Genes Evol 215:221–229
Suttipanta N, Pattanaik S, Kulshrestha M, Patra B, Singh SK, Yuan L (2011) The transcription factor CrWRKY1 positively regulates the terpenoid indole alkaloid biosynthesis in Catharanthus roseus. Plant Physiol 157:2081–2093
Tang J, Wang F, Wang Z, Huang Z, Xiong A, Hou X (2013) Characterization and co-expression analysis of WRKY orthologs involved in responses to multiple abiotic stresses in Pak-choi (Brassicacampestris ssp. chinensis). BMC Plant Biol 13:188
Toledo-Ortiz G, Huq E, Quail PH (2003) The Arabidopsis basic/helixloop- helix transcription factor family. Plant Cell 15:1749–1770
Udvardi MK, Kakar K, Wandrey M, Montanari O, Murray J, Andriankaja A, Zhang JY, Benedito V, Hofer JMI, Chueng F, Town CD (2007) Legume transcription factors: global regulators of plant Development and response to the environment. Plant Physiol 144:538–549
Ulker B, Somssich IE (2004) WRKY transcription factors: from DNA binding towards biological function. Curr Opin Plant Biol 7:491–498
Urano K, Kurihara Y, Seki M, Shinozaki K (2010) ‘Omics’ analyses of regulatory networks in plant abiotic stress responses. Curr Opin Plant Biol 13:1–7
Vij S, Tyagi AK (2006) Genome-wide analysis of the stress associated protein (SAP) gene family containing A20/AN1 zinc-finger(s) in rice and their phylogenetic relationship with Arabidopsis. Mol Genet Genomics 276:565–575
Wang YJ, Zhang ZG, He XJ, Zhou HL, Wen YX, Dai JX, Zhang JS, Chen SY (2003) A rice transcription factor OsbHLH1 is involved in cold stress response. Theor Appl Genet 107:1402–1409
Wu X, Shiroto Y, Kishitani S, Ito Y, Toriyama K (2009) Enhanced heat and drought tolerance in transgenic rice seedlings overexpressing OsWRKY11 under the control of HSP101 promoter. Plant Cell Rep 28:21–30
Xia N, Zhang G, Liu XY, Deng L, Cai GL, Zhang Y, Wang XJ, Zhao J, Huang LL, Kang ZS (2010) Characterization of a novel wheat NAC transcription factor gene involved in defense response against stripe rust pathogen infection and abiotic stresses. Mol Biol Rep 37:3703–3712
Xiong Y, Liu T, Tian C, Sun S, Li J, Chen M (2005) Transcription factors in rice: a genome wide comparative analysis between monocots and eudicots. Plant Mol Biol 59:191–203
Xu DQ, Huang J, Guo SQ, Yang X, Bao YM, Tang HJ, Zhang HS (2008) Overexpression of a TFIIIA-type zinc finger protein gene ZFP252 enhances drought and salt tolerance in rice (Oryza sativa L.). FEBS Lett 582:1037–1043
Yang S, Vanderbeld B, Wan J, Huang Y (2010) Narrowing down the targets: towards successful genetic engineering of drought-tolerant crops. Mol Plant 3:469–490
Yi K, Wu Z, Zhou J, Du L, Guo L, Wu Y, Wu P (2005) OsPTF1, a novel transcription factor involved in tolerance to phosphate starvation in rice. Plant Physiol 138:2087–2096
Yilmaz A, Nishiyama MY, Fuentes BG, Souza GM, Janies D, Gray J, Grotewold E (2009) GRASSIUS: a platform for comparative regulatory genomics across the grasses. Plant Physiol 149:171–180
Yu Y, Hu R, Wang H, Cao Y, He G, Fu C, Zhou G (2013) MlWRKY12, a novel Miscanthus transcription factor, participates in pith secondary cellwall formation and promotes flowering. Plant Sci 212:1–9
Yu CS, Cheng CW, Su WC, Chang KC, Huang SW, Hwang JK, Lu CH (2014) CELLO2GO: a web server for protein subcellular localization prediction with functional gene ontology annotation. PLoS One 9:e99368
Zhao X, de Palma J, Oane R, Gamuyao R, Luo M, Chaudhury A, Hervé P, Xue Q, Bennett J (2008) OsTDL1A binds to the LRR domain of rice receptor kinase MSP1, and is required to limit sporocyte numbers. Plant J 54:375–387
Zheng Z, Qamar SA, Chen Z, Mengiste T (2006) Arabidopsis WRKY33 transcription factor is required for resistance to necrotrophic fungal pathogens. Plant J 48:592–605
Zheng X, Chen B, Lu G, Han B (2009) Overexpression of a NAC transcription factor enhances rice drought and salt tolerance. Biochem Biophys Res Commun 379:985–989
Zou CS, Jiang WB, Yu DQ (2010) Male gametophyte-specific WRKY34 transcription factor negatively mediates cold stress tolerance of mature pollen in Arabidopsis. J Exp Bot 14:3901–3914
Acknowledgements
PM acknowledges Alagappa University Research Fund (Ph.D./1215/AURF Fellowship/2015 dated 25.11.2015), Karaikudi, Tamil Nadu, India for providing Research Fellowship. The authors gratefully acknowledge the use of Bioinformatics Infrastructure Facility, Alagappa University funded by Department of Biotechnology, Ministry of Science and Technology, Government of India Grant (No.BT/BI/25/015/2012).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Muthuramalingam, P., Krishnan, S.R., Saravanan, K. et al. Genome-wide identification of major transcription factor superfamilies in rice identifies key candidates involved in abiotic stress dynamism. J. Plant Biochem. Biotechnol. 27, 300–317 (2018). https://doi.org/10.1007/s13562-018-0440-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-018-0440-3