Abstract
Introduction
Type 2 diabetes mellitus (T2DM) is one of the leading preventable chronic diseases with high rates of morbidity and mortality. It is multi-factorial, polygenic, and characterized by chronic hyperglycemia. Transcription factor 7-like 2 (TCF7L2), a key component of the canonical Wnt signaling pathway, is one of the most susceptible genes with the largest effect on T2DM. Pakistan is populated with the rising trend of T2DM in the adult population, hence to advocate the previous though scarce findings, this case–control study was designed to see T2DM prevalence and its association with TCF7L2 intronic variant, rs12255372.
Methodology
DNA extraction and genotyping through the tetra primer amplification refractory mutation system (T-ARMS-PCR) were done on a total of 162 collected samples of the Pakistani cohort. Data was statistically analyzed by SPSS IBM v20 and the Hardy–Weinberg equilibrium test was applied to see the stratification.
Results
The variant genotyped was in Hardy–Weinberg equilibrium (p > 0.05). The SNP rs12255372 showed an association with T2DM and remained significant after multiple adjustments for age and body mass index. The T allele showed a positive association in study subjects.
Conclusion
The study concluded, TCF7L2 SNP, rs12255372 are associated with T2DM. The risk allele T is more susceptible to T2DM in the later age group (40–70 years) of females and the early age group (20–40 years) of males. This gender effect is a unique finding, and further, more replicated studies need to be carried out to determine the effects of risk factors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diabetes is placed as the 7th disease that affects people and causes death among various ethnicities, as per World Health Organization (WHO). By the end of 2030, this level will raise up to 522 million affecters [1]. Type 2 diabetes mellitus (T2DM), formerly considered as adult-onset, is one of the leading, however, preventable chronic disease with high rates of morbidity and mortality. In the adult population, the estimate of diabetes prevalence is to be 8.5 percent globally and out of which approximately, 90% patients are scourged with T2DM [2]. T2DM is characterized by chronic hyperglycemia due to the mutual interplay of environmental and genetic factors [3, 4]. There are some additional T2DM correlated risk factors including unhealthy eating, lack of exercise, overweight, and family history [5, 6].
Some candidate genes confirmed as T2DM associated genes by genome wide association studies (GWAS) in several studies. However, there are many genes that along with other diseases are also related with T2DM [7]. Planning and construction of genetic studies by using different approaches have progressed over the last decade for the inquisition and identification of complex human disease risk factors [8]. The recent data of polygenic T2DM show that more than four hundred genetic risk variants at two hundred and forty T2DM loci have been identified mainly through GWAS [9].
Transcription factor 7-like 2 (TCF7L2) is one of the most T2DM susceptible genes. It is located on the long arm of chromosome 10 (10q25.3) and started from 114,710,009 bp to 114,927,437 bp on the forward strand [10]. First, Grant et al. in 2006 identified TCF7L2 and its consequence on T2DM was replicated effectively in several populations [11, 12]. It was previously called as T-cell factor 4 (TCF4), not to be confused with transcription factor 4, and its variants are effectively involved in developing T2DM [13]. It is a key component of the canonical Wnt signaling pathway, involved in the development of a wide variety of cell lineages and organs. This pathway is exposed to beta cell dysfunction in T2DM [14].
The total population of Pakistan consists of 207.7 million and comprised of an área, 796095 km2 [15]. The recent findings in Pakistan reported a rise in T2DM prevalence among adults [16]. To interrogate about the causal variant and the risk allele of intronic SNPs within a gene is an integral proxy of genomic diversity. In several replicated studies, the different associations between T2DM and TCF7L2 gene variants were reported; therefore, due to the scarcity of data in South Asian countries and to compare the findings with the previous evidences, this case–control study was conducted. The specific objectives were to study the T2DM prevalence and its association with the TCF7L2 variant. Moreover, to estimate variables (age, BMI, gender) individually and as clustered effects towards T2DM in the case–control subjects.
Material and method
Study subjects and anthropometric measurements
A total of 162 subjects were recruited from the Pakistani Sindhi cohort based in Karachi according to inclusion and exclusion criteria. The exclusion criteria included the subjects with an age group equal or less than 18, controls with a history of any chronic disease or ailment, and cases with a history of any disease other than T2DM. All the healthy controls were selected on the basis of age and gender matching, without any history of medication, diabetes, or any other disease symptoms. Anthropometric measurements including weight in kilograms and height in square meters were noted to calculate BMI [17].
Sample collection, biochemical tests, and DNA extraction
A 10 ml blood sample from each subject was collected from the antecubital vein in the sitting position. Samples were grouped and a total of 84 of 162 samples were diabetic patients (DP), and 78 of 162 samples were diabetic controls (DC). Biochemical fasting blood sugar and lipid tests were done through commercially available kits (Abcam, USA). Purified genomic DNA from the blood of each participant was extracted through the available genomic isolation kit protocol (Promega, USA). Extracted DNA purity check was done through quantification by using a Nanodrop-ND1000 (Thermo Fisher Scientific, Waltham, MA).
Genotyping for TCF7L2 rs12255372 variant
The intronic SNP rs12255372 was chosen based on literature. The genotyping was carried out by tetra-primer amplification, refractory mutation system polymerase chain reaction. (T-ARMS-PCR) in the presence of two pairs of specifically designed primers (Table 1). Primers were verified by the online bioinformatics primer design tool Primer3web, https://primer3plus.com/primer3web/primer3web_input.htm. The SNP possesses G, a reference allele, and T, a T2DM risk allele. This SNP was visualized on a 1% agarose gel and analyzed through a gel documentation system (Bio-rad).
Statistical analyses
Hardy–Weinberg equilibrium (HWE) was applied to find the stratification of study participants. All continuous and binary variables were calculated as mean ± standard deviation (SD) and numbers with percentages, respectively. The case–control groups were compared by an independent sample t-test. The relationship of genotypes with cases and controls was recognized by the chi-square test and obtained an odds ratio by logistic regression for further authentication and to find the association of variables with T2DM. In addition, a hierarchical linear regression was employed by making different models with risk factors. A cut-off p-value ≤ 0.05 was considered statistically significant. All the calculations were carried out using SPSS version 20.
Results
Gender distribution
A total of 41 of 84 DP were males, and the remaining 43 of 84 DP were females; however, 49 of 78 were DC males, and the remaining 29 were DC females (Fig. 1). The results showed a high prevalence of T2DM in female adults which is supporting a previous study. Tang et al. in 2019 reported that T2DM prevalence in females occurs in low as well as in high gross domestic per capita areas where they are at multiple high risks of malnutrition, lack of physical activity, or obesity [18].
Body mass index (BMI)
The BMI of subjects was recorded and analyzed (Table 2). All pooled samples were listed as per WHO BMI Asian criteria as normal (19–24.9), overweight (≥ 25), and obese (> 30), though the later two are abnormal categories. Out of the total subjects, 31 (19.13%) had a normal BMI, 69 (41.97%) were overweight whereas 59 (36.4%) were obese.
Basic demographic characteristics and biochemical tests
The data was analyzed and the results concluded that the T2DM case group had high dyslipidemia, fasting blood sugar, BMI, triglycerides (TG), and low levels of high-density lipoprotein cholesterol (HDL-C) (p < 0.05) (Table 2). Results showed no difference in age and males among both case–control groups, because the controls were individually matched on age and gender. These controls were not on medication and with no history of hypertension. Their total cholesterol (TC) and low-density lipoprotein cholesterol (LDL-C) levels are high as compared to diabetic cases while their BMI is less, HDL-C level is high, and TG is low than the diabetic subjects.
A subset of individuals despite having a high BMI or high-fat deposition have been reported to have metabolically healthy as characterized by high insulin sensitivity, no sign of hypertension, low C-reactive protein, high HDL-C, and low TG [19]. Although insulin sensitivity and C-reactive protein are not tested in our study, in the light of the above reference, our study indicated the same trend. The high level of elevated BMI with a high level of TG and LDL-C and low levels of HDL-C have reported to be associated with cardiovascular diseases, diabetes, or metabolic syndrome; this situation is found in most of our case samples.
Association of subjects, gender, BMI with genotypes
By applying the chi-square test, the association of genotypes (GG, GT, and TT) with case–control subjects, male–female gender, and normal-abnormal BMI of the pooled samples was observed. Results showed a significant association of male–female gender with genotypes (χ2 = 7.806 and p-value = 0.02) (Table 3) in the selected cohort.
Genetic analysis by HWE
The genotype and allele frequencies for the SNP in healthy and control individuals were tested by HWE through the Pearson χ2 test and are detailed in Table 4. The frequency of genotypes in the DC group was 21.88% for GG, 60.3% for GT, and 17.9% for TT; however, the frequency of reference allele (G) was 51.9% and rare allele (T) was 48.1%. Results revealed observed and expected values (χ2 = 3.34 and p-value = 0.06) which are non-significant and indicated that the population of controls is falling in concordance with HWE. The frequency of genotypes in the DP group was 13.1% for GG, 71.4% for GT and 15.5% for TT; however, the frequency of reference allele (G) was 48.8% and of rare allele (T) was 51.1%. Results by HWE indicated observed and expected values providing χ2 = 15.48 and p-value = 8.308E-05, which is significant. This indicates that the population of cases is not falling in concordance with HWE and is being stratified. This stratification may be due to the presence of a lethal allele as our samples were of homogenous ethnicity. The occurrence of the GT genotype was found higher in both groups; however, the TT genotype was higher in DP and the GG genotype was higher in controls. Exclusively controls fall in concordance with the HWE. Deviations of HWE in cases were expected due to the marked association of genotypes with clinical phenotype certainly in homogenous populations.
The Pakistani population is a broadly diverse and genetic admixture of European and Central or West Asian migrants with the gene pools of indigenous South Asian. Pakistanis are organized in different ethnicities grounded on linguistic, cultural, and geographical origin [20]. Though, the genetic data to differentiate various Pakistan ethnicities is scanty, a few studies are done so far based on the microsatellites/short tandem repeats and variants [21, 22]. This study population genetically as well as geographically differs from the previous study conducted by Hameed et al. in the Pashtun linguistic group of Pakistan. The differences in the genotype and allele percentages for the TCF7L2 SNP rs12255372 among Sindhi and Pakhtun linguistic groups reveal no genetic resemblance with the previous study of the Northern cohort [23].
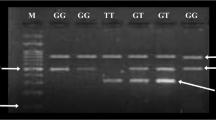
The T-ARMS PCR results were visualized after horizontal agarose gel electrophoresis of DC and DP samples. The Fig. 2 is showing amplicons of various sizes with respect to alleles.
TCF7L2 rs12255372 SNP genotyping of type 2 diabetes mellitus cases and controls by tetra-primer amplification refractory mutation system polymerase chain reaction (T-ARMS PCR). Left: lane 1 contains a 100 bp ladder; lane 2 contains the negative control; lane 3 through 10 wells contain amplicons of diabetic control (DC) samples. Right: lane 1 contains the 100 bp ladder; lane 2 through 10 wells contain amplicons of diabetic patient (DP) samples. The gel was made up of 1.5% agarose assembled on horizontal gel electrophoresis (Sigma) with a comb of 10 wells. Later, bands for nonspecific (760 bp), G-allele specific (494 bp), and T-allele (310bp) specific were visualized under UV light via the Bio-Rad gel documentation system.
Logistic regression analysis for association of risk factors with diabetics and non-diabetics
A crude and adjusted odds ratio with a 95% confidence limit was calculated in cases and controls by using binary logistic regression analysis for estimation of relative risk association (Table 5).
Genotype
In cases, 2.057 increased relative odds of genotype GT found a strong association with diabetes and a less strong association in controls by increased odds of 0.486 with a significant p-value (p < 0.05) showing that the reported effect is solely due to chance, whereas in controls, although there is a 1.77 increase of positive outcome of the genotype TT, this increase is not statistically significant.
Gender
Similarly, the female gender is strongly associated with DP with odds of 1.77 although it is statistically slightly insignificant (p = 0.074). The male gender showed a strong association with DC. The age group from 20 to < 40 years showed a strong association with DP (odds ratio value 1.5); however, the age group from 40 to 70 years showed an association with cases through the same 1.5-fold increased odds while this increase in both of these age groups proved statistically insignificant (p = 0.216). These results suggested that DC and DP subjects are associated with risk allele genotype GT with p value 0.044 (OR = 0.486) and 0.044(OR = 2.057), respectively.
BMI
South Asians are at increased risk of morbidity at the lower values of BMI in comparison to WHO defined cut off values. Obesity and T2D are multifactorial diseases, and a combination of genetic and environmental risk factors is believed to lead these complex diseases [24]. BMI, including normal, overweight, and obese categories, possessed a statistically insignificant association with 95% CI in cases and controls (Table 5).
Comparison between effects of genetic variant and clustering of risk factors on cases and controls
We employed hierarchical linear regression to study the association of the TCF7L2 SNPs with T2DM under three genetic models showing weak and protective effects due to reference of T2DM-associated TCF7L2 SNP [25]. In primary concern of this study, linear regression analysis was conducted to estimate the pooled effects of various variables and to see the significant increase risk of T2DM among recruited cases and controls. This statistical test was analyzed by making models unadjusted for cases and controls and analyzed after adjustment with risk factors including genotype, age, gender, and BMI (Tables 6 and 7). These models provided us the effect size of genetic factors alone and the effect size with clustering of the covariates risk factors.
Gender
The results showed that in controls of both age groups (20 to 40 years; 40 to 70 years), the male showed a statistically significant weak and moderate effect in association with risk allele genotypes, GT and TT. However, females are also significantly associated with both risk allele genotypes but showed a protective effect (Table 6). In cases, males and females of the age group (21 to < 40 years) conclude a significant protective and lethal effect on T2DM, respectively. On the contrary, in cases of an age group > 40 years, males showed a significant weak lethal effect with the GT genotype and a protective effect with the TT genotype, whereas females showed a significantly largest lethal effect size with the GT genotype and a moderate lethal effect with the TT genotype (Table 7).
BMI
BMI of the obese category was not associated with cases and controls by separate binary logistic regression analysis; however, the significant association was found only when it was evaluated together with other covariates. The significant moderate effect size of BMI (> 25 kg/m2) with genotype TT in the age group of > 40 years (p = 0.045) had been observed in female T2DM cases (Table 7).
Genotype
In both cases and controls, genotypes GT and TT showed a lethal effect in males and females of both age groups from 20 to 70 years, (Tables 6 and 7) whereas in cases, the TT genotype showed a lethal effect with BMI in females only of age groups 40 to 70 years (Table 7). Our results are represented in concordance to Cohen’s rule that shows the clustering effect is increased from weak to moderate and strong with the addition of risk factors. In controls and cases with the age group of 40 to 70 years, the only significant and problematic factor left was raised BMI. It was found that the effect size of genetic factors produced less elevation in T2DM cases, but it was significantly more due to other factors particularly by gender and BMI. The significant effect size for risk allele genotype in both case and control subjects came from gender only.
Discussion
Our result provides chief confirmation that variants of the TCF7L2 gene are strongly associated with type 2 diabetes in a selected ethnicity. We found that the minor allele-T of TCF7L2 rs12255372 significantly increases T2D risk. Our data support the multiplicative model of inheritance for the TCF7L2 SNPs proposed by Grant et al., (2006). Results of T-ARMS PCR genotyping found that rare allele-T was higher among cases than controls suggested another evidence of TCF7L2 susceptibility. The association of females and males found significantly associated with genotypes, and the rare allele-T was higher in females (91.7%) as compared to males (75.6%) with odds of 1.77 which are in contrary to the results confirmed in US population [26]. The previous study showed a high percentage of rare allele-T in men as compared to women. However, another study suggested a high mortality rate of females due to uncertain diabetes as compared to males [27].
This chromosome locus has a significance to be considered as an inheritable risk factor for diabetes. Logistic regression analysis of SNP rs12255372 showed odds of 2.057 (95% CI) for GT genotypes with significant p-value and TT genotype showed odds of 0.56 which is statistically insignificant. This outcome is unlikely to the study previously reported by Bodhini et al., 2007 [28]. It is due to the potential limitation of a relatively small number of cases and control subjects. This limitation may resolve by further increasing the study power with the addition of study samples. The present study also highlights the importance of clustering of risk factors with genotypes. Results found that TCF7L2 variant rs12255372 in the later age group of T2DM cases has a more moderate effect to its predisposition which is in contrary to the previous study, which reported that TCF7L2 variants are associated with an early age onset of T2DM [29].
TCF7L2 was associated with T2DM and decreased BMI [30], but some studies showed that it is associated with increased BMI. There was a need to further investigate its relevance in Pakistani cohorts. Hence, our results provide that in addition to later age, higher BMI along with risk allele-T of rs12255372 variant has a more pronounced effect on the T2DM. These results are in concordance with the study of Cauchi et al., 2008 who suggested that TCF7L2 SNP is not a risk factor for obesity in European populations, but its effect on T2DM risk is modulated by obesity [31]. The present study also determined a novel effect of gender on T2DM in association with the risk allele. The study concluded that along with other risk factors, females with risk allele genotype (GT or TT) are more susceptible to T2DM in the later age of 40 to 70 years, whereas males are more susceptible to T2DM in the early age from 20 to 40 years in association with risk allele genotypes. This is a unique finding and further more studies are required to support this outcome.
Conclusion
The study concluded, TCF7L2 variant rs12255372 has associated with T2DM and when combined with various phenotypic risk factors, it enhances the predisposition of individuals to develop T2DM. Moreover, the study also concluded a novel effect of gender that females with risk allele genotypes are more susceptible to T2DM in the later age of 40 to 70 years, whereas males are more susceptible to T2DM in the early age from 20 to 40 years in association with risk allele genotypes. This is a unique finding and more studies are required to determine the environmental risk factors effect in relation to genes implicated in the predisposition of T2DM. Apart from TCF7L2 variants, additional genetic studies can explore T2DM risk for other genes interplaying a role in the pathogenesis of T2DM in different Pakistani cohorts.
References
Mambiya M, Shang M, Wang Y, Li Q, Liu S, Yang L, Zhang Q, Zhang K, Liu M, Nie F, Zeng F, Liu W. The play of genes and non-genetic factors on type 2 diabetes. Front Public Health. 2019. https://doi.org/10.3389/fpubh.2019.00349.
World Health Organization. Classification of diabetes mellitus. 2019. https://www.who.int/publications/i/item/classification-of-diabetes-mellitus. Accessed 21 Apr 2019.
Franks PW. The complex interplay of genetic and lifestyle risk factors in type 2 diabetes: an overview. Scientifica (Cairo). 2012. https://doi.org/10.6064/2012/482186.
Kolb H, Martin S. Environmental/lifestyle factors in the pathogenesis and prevention of type 2 diabetes. BMC Med. 2017. https://doi.org/10.1186/s12916-017-0901-x.
International Diabetes Federation. IDF Diabetes Atlas, 9th edn. Brussels, Belgium: International Diabetes Federation. 2019. http://www.diabetesatlas.org.
Cho NH, Shaw JE, Karuranga S, Huang Y, da Rocha Fernandes JD, Ohlrogge AW, Malanda B. IDF Diabetes Atlas: global estimates of diabetes prevalence for 2017 and projections for 2045. Diabetes Res Clin Pract. 2018;138:271–81.
Lyssenko V, Groop L. Genome-wide association study for type 2 diabetes: clinical applications. Curr Opin Lipidol. 2009;20(2):87–91.
Waqar A, Adnan R, Bushra C, Ikram-ul H. Alterations, complexity and efficient strategies suitable for genetic studies. Biologia (Lahore, Pakistan). 2019;65(II):1–7.
Dziewulska A, Dobosz AM, Dobrzyn A. High-throughput approaches onto uncover (Epi) genomic architecture of type 2 diabetes. Genes. 2018. https://doi.org/10.3390/genes9080374.
Howe KL, Achuthan P, Allen J, Allen J, Alvarez-Jarreta J, Amode MR, Armean IM, Azov AG, Bennett R, Bhai J, Billis K, Boddu S, Charkhchi M, Cummins C, Da Rin Fioretto L, Davidson C, Dodiya K, El Houdaigui B, Fatima R, Gall A, Garcia Giron C, Grego T, Guijarro-Clarke C, Haggerty L, Hemrom A, Hourlier T, Izuogu OG, Juettemann T, Kaikala V, Kay M, Lavidas I, Le T, Lemos D, Gonzalez Martinez J, Marugán JC, Maurel T, McMahon AC, Mohanan S, Moore B, Muffato M, Oheh DN, Paraschas D, Parker A, Parton A, Prosovetskaia I, Sakthivel MP, Salam AIA, Schmitt BM, Schuilenburg H, Sheppard D, Steed E, Szpak M, Szuba M, Taylor K, Thormann A, Threadgold G, Walts B, Winterbottom A, Chakiachvili M, Chaubal A, De Silva N, Flint B, Frankish A, Hunt SE, IIsley GR, Langridge N, Loveland JE, Martin FJ, Mudge JM, Morales J, Perry E, Ruffier M, Tate J, Thybert D, Trevanion SJ, Cunningham F, Yates AD, Zerbino DR, Flicek P. Ensembl. Nucleic Acids Res. 2021;49(1):884–91.
Grant SF, Thorleifsson G, Reynisdottir I, Benediktsson R, Manolescu A, Sainz J, Helgason A, Stefansson H, Emilsson V, Helgadottir A, Styrkarsdottir U, Magnusson KP, Walters GB, Palsdottir E, Jonsdottir T, Gudmundsdottir T, Gylfason A, Saemundsdottir J, Wilensky RL, Reilly MP, Rader DJ, Bagger Y, Christiansen C, Gudnason V, Sigurdsson G, Thorsteinsdottir U, Gulcher JR, Kong A, Stefansson K. Variant of transcription factor 7-like 2 (TCF7L2) gene confers risk of type 2 diabetes. Nat Genet. 2006;38(3):320–3.
Gloyn AL, Braun M, Rorsman P. Type 2 diabetes susceptibility gene TCF7L2 and its role in beta-cell function. Diabetes. 2009;58(4):800–2.
Del Bosque-Plata L, Hernández-Cortés EP, Gragnoli C. The broad pathogenetic role of TCF7L2 in human diseases beyond type 2 diabetes. J Cell Physiol. 2022;237(1):301–12.
Abou Azar F, Lim GE. Metabolic contributions of Wnt signaling: more than controlling flight. Front Cell Dev Biol. 2021. https://doi.org/10.3389/fcell.2021.709823.
Aamir AH, Ul-Haq Z, Mahar SA, Qureshi FM, Ahmad I, Jawa A, Sheikh A, Raza A, Fazid S, Jadoon Z, Ishtiaq O, Safdar N, Afridi H, Heald AH. Diabetes Prevalence Survey of Pakistan (DPS-PAK): prevalence of type 2 diabetes mellitus and prediabetes using HbA1c: a population-based survey from Pakistan. BMJ Open. 2019. https://doi.org/10.1136/bmjopen-2018-025300.
Adnan M, Aasim M. Prevalence of type 2 diabetes mellitus in adult population of Pakistan: a meta-analysis of prospective cross-sectional surveys. Ann Glob Health. 2020. https://doi.org/10.5334/aogh.2679.
Domaradzki J, Rokita A, Koźlenia D, Popowczak M. Optimal values of body composition for the lowest risk of failure in Tabata training’s effects in adolescents: a pilot study. Biomed Res Int. 2021. https://doi.org/10.1155/2021/6675416.
Tang K, Wang H, Liu Y, Tan SH. Interplay of regional economic development, income, gender and type 2 diabetes: evidence from half a million Chinese. J Epidemiol Community Health. 2019;73(9):867–73.
Smith GI, Mittendorfer B, Klein S. Metabolically healthy obesity: facts and fantasies. J Clin Invest. 2019;129(10):3978–89. https://doi.org/10.1172/JCI129186.
Pigeyre M, Saqlain M, Turcotte M, Raja GK, Meyre D. Obesity genetics: insights from the Pakistani population. Obes Rev. 2018;19(3):364–80. https://doi.org/10.1111/obr.12644.
Mehdi SQ, Qamar R, Ayub Q, Khaliq S, Mansoor A, Ismail M, Hammer M, Underhill PA, Cavalli-Sforza LL. The origins of Pakistani populations, evidence from Y hromosome markers. In: Papiha SS, Deka R, Chakraborty R, editors. Genomic diversity: Applications in Human Population Genetics. New York: Kluwer Academic/Plenum Publishers; 1999. p. 83–90.
Anwar I, Hussain S, Rehman AU, Hussain M. Genetic variation among the major Pakistani populations based on 15 autosomal STR markers. Int J Legal Med. 2019;133(4):1037–8. https://doi.org/10.1007/s00414-018-1951-0.
Hameed T, Khan Z, Imran M, Ali S, Albegali AA, Ullah MI, Ejaz H. Associations of transcription factor 7-Like 2 (TCF7L2) gene polymorphism in patients of type 2 diabetes mellitus from Khyber Pakhtunkhwa population of Pakistan. Afr Health Sci. 2021;21(1):15–22. https://doi.org/10.4314/ahs.v21i1.4.
O’Rahilly S, Farooqi IS. Genetics of obesity. Phil Trans Royal Soc Lond Ser B Biol Sci. 2006;361(1471):1095–105.
Cohen J. A power primer. Psychol Bull. 1992;112(1):155–9.
Zhang C, Qi L, Hunter DJ, Meigs JB, Manson JE, Damvan RM, Hu FB. Variant of transcription factor 7-like 2 (TCF7L2) gene and the risk of type 2 diabetes in large cohorts of U.S. women and men. Diabetes. 2006;55(9):2645–8.
Zhu M, Li J, Li Z, Luo W, Dai D, Weaver SR, Stauber C, Luo R, Fu H. Mortality rates and the causes of death related to diabetes mellitus in Shanghai Songjiang District: an 11-year retrospective analysis of death certificates. BMC Endocr Disord. 2015. https://doi.org/10.1186/s12902-015-0042-1.
Bodhini D, Radha V, Dhar M, Narayani N, Mohan V. The rs12255372(G/T) and rs7903146(C/T) polymorphisms of the TCF7L2 gene are associated with type 2 diabetes mellitus in Asian Indians. Metab Clin Exp. 2007;56(9):1174–8.
Silbernagel G, Renner W, Grammer TB, Hügl SR, Bertram J, Kleber ME, Hoffmann MM, Winkelmann BR, März W, Boehm BO. Association of TCF7L2 SNPs with age at onset of type 2 diabetes and proinsulin/insulin ratio but not with glucagon-like peptide 1. Diabetes Metab Res Rev. 2011;27(5):499–505.
Lukacs K, Hosszufalusi N, Dinya E, Bakacs M, Madacsy L, Panczel P. The type 2 diabetes-associated variant in TCF7L2 is associated with latent autoimmune diabetes in adult Europeans and the gene effect is modified by obesity: a meta-analysis and an individual study. Diabetologia. 2012;3:689–93.
Cauchi S, Nead KT, Choquet H, Horber F, Potoczna N, Balkau B, Marre M, Charpentier G, Froguel P, Meyre D. The genetic susceptibility to type 2 diabetes may be modulated by obesity status: implications for association studies. BMC Med Genet. 2008. https://doi.org/10.1186/1471-2350-9-45.
Funding
Institute of Industrial Biotechnology, GCU Lahore and Department of Biological and Biomedical Sciences, AKU Karachi research module fund.
Author information
Authors and Affiliations
Contributions
AW carried out the literature search, data collection, experimental analysis, interpretation, and manuscript writing. BC and IH were involved in finalizing the manuscript and revising it critically for important intellectual content. KS was involved in experimental analysis, and AN was involved in proofreading and submission. The final manuscript was approved by all authors for publication.
Corresponding author
Ethics declarations
Ethics and consent
This study was approved by the institutional bioethics committee in the Government College University, Lahore (GCUL). The study was conducted in accordance with the ethical standards as laid down in the 1964 Declaration of Helsinki and its later amendments or comparable ethical standards. The participants were included in the study after signing a consent proforma.
Ethics approval
An ERC approval (GCU-IIB-153) was obtained, and every participant signed an informed written consent.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Waqar, A., Chaudhry, B., Haq, Iu. et al. Association between type 2 diabetes mellitus and TCF7L2 gene variant in the Pakistani cohort. Int J Diabetes Dev Ctries 43, 807–815 (2023). https://doi.org/10.1007/s13410-022-01138-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13410-022-01138-4