Abstract
Pakistan is located at an important cross-road of human history and has been a passageway for many invaders and dynasties in the past. The historic human migrations across this country have resulted in a blend of ancient civilizations, which are still reflected in the current socio-cultural fabrication of this population. This makes Pakistan an ideal country to study the genetic differentiation and various other genomic aspects of a human population.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
This study encompass a total of 1020 unrelated healthy subjects belonging to four major ethnic groups of Pakistan (Punjabi, 600; Saraiki, 150; Pakhtuns, 140; and Sindhi, 130). After approval of the written informed consent, blood or buccal swab samples were collected from each subject residing in the respective area of their ethnicity. In order to avoid sampling biases and have a reasonable representation of each population, participants were recruited from both genders and all age groups. The selected ethnic groups in this study represent a reasonable proportion of the total Pakistani population. For example, by comparing the two maps shown in Figs. 1 and 2 (Supplementary Files), it is plausible to say that the current sample in our study is a true representation of the Pakistani population in whole.
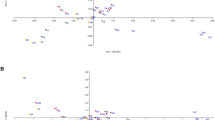
Genomic DNA was extracted from blood samples/buccal swab by organic method. All 15 loci along with amelogenin were co-amplified using the AmpFℓSTR Identifiler® kit (Applied Biosystems). Allele frequencies at each locus were calculated using “Hierfstat package” [1] of R computing language. Observed heterozygosity (Ho) and expected heterozygosity (He) were calculated using the Genepop software version 4 [2]. Parameters of forensic interest were calculated by using the PowerStats software v1.2 [3]. Matching probability (MP), power of discrimination (PD), polymorphic information content (PIC), power of exclusion (PE), and typical paternity index (TPI) were calculated to investigate the admissibility of studied marker set for Pakistani populations. Genetic distances (FST) were calculated by introducing 5000 bootstrap values. These genetic distances were calculated by using the Poptree2 software [4] and phylogenetic tree (neighbor joining method) showing the closest and farthest genetic neighbors of the studied populations was created. Finally, our results were compared with other neighboring populations like Afghanistan [5], Iran [6], China [7], Nepal [8], Bhutan [9], United Arab Emirates [10], and India [11,12,13,14].
Distribution of allele frequencies for the four studied populations (Punjabi, Saraiki, Pakhtun, and Sindhi) is presented in Table S1 to S4 of the supplementary material provided in the online version of this article. A total of 187 different alleles were observed with a range of frequency from 0.001 (D21S11, CSF1PO, D3S1358, TH01, D13S317, D2S1338, D19S433, vWA, TPOX, D18S51, D5S818, FGA) in Punjabi population to 0.471 (TPOX) in Pakhtun population. Observed heterozygosity (Ho) was observed to be the lowest with a value of 0.5615 at locus (CSF1PO) in Sindhi population and was highest with a value of 0.9133 at locus FGA in Saraiki population. Matching probability (MP) calculations also showed a variety from the lowest being at 0.028 at locus D2S1338 in Punjabi population to the highest value at 0.174 at locus TPOX in Pakhtun population, while the combined matching probability was 1.321 × 10−18 for Punjabi population, 2.348 × 10−18 for Saraiki population, 6.327 × 10−18 for Pakhtun population, and 6.715 × 10−18 for Sindhi population. Power of discrimination (PD) ranged from 0.826 (TPOX) in Pakhtun population to 0.972 (D2S1338) in Punjabi population, and the combined power of discrimination for the Punjabi population was observed to be 0.3188, for Saraiki 0.3122, for Pakhtun 0.2822, and for the Sindhi population it was 0.2815. Power of exclusion (PE) spanned from 0.247 at locus CSF1PO in Sindhi population to the highest value of 0.823 at locus FGA in Saraiki population. All four major Pakistani populations were found to be in close proximity when the genetic distances were calculated among them and with the neighboring populations.
Notably, all the studied major Pakistani populations were observed to show close genetic affinity. However, the Punjabi population showed more genetic resemblance to the neighboring Balmiki population of India which lives across the border of the divided Punjab at the time of partition of the British India. Moreover, the previously studied Hazara population of Pakistan [15] showed their genetic neighborhood with the Ouzbek population of Afghanistan and to the Uyghur population of China which borders the northern parts of Pakistan. Balochi population showed their genetic resemblance with the Irani counterpart and also with the Arabs living in United Arab Emirates, while Afridi Pathan population of India showed the greatest genetic distance with the Pakistani populations. Nepali and Bhutani populations were also present at a notable distance. All the calculated genetic distances (FST) can be seen in Table S5 of the Supplementary data.
References
Goudet J (2005) Hierfstat a package for r to compute and test hierarchical F-statistics. Mol Ecol Notes 5(1p):184–186. https://doi.org/10.1111/j.1471-8286.2004.00828.x
Rousset F (2008) Genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Mol Ecol Resour 8(1p):103–106. https://doi.org/10.1111/j.1471-8286.2007.01931.x
Tereba A (1999) Tools for analysis of population statistics, profiles in DNA. Promega Corporation 2:14–16
Takezaki N, Nei M, Tamura K (2009) POPTREE2: software for constructing population trees from allele frequency data and computing other population statistics with Windows Interface. Molecular Biology Evolution 27(4p):747–752. https://doi.org/10.1093/molbev/msp312
Di Cristofaro J, Buhler S, Temori SA, Chiaroni J (2012) Genetic data of 15 STR loci in five populations from Afghanistan. Forensic Sci Int: Genetics 6(1p):e44–e45. https://doi.org/10.1016/j.fsigen.2011.03.004
Shepard EM, Herrera RJ (2006) Iranian STR variation at the fringes of biogeographical demarcation. Forensic Sci Int 158:140–148. https://doi.org/10.1016/j.forsciint.2005.05.012
Dong L, Liu XX, Zhang HH, Bo R, Wei G, Zhang LS (2010) Genetic data for 15 STR loci in Uygur ethnic group of Northwest China. Romanian J Legal Med 18(1). https://doi.org/10.4323/rjlm.2010.59
KraaijenbrinK T, VAN Driem GL, Opgenort JRML, Tuladhar NM, DE Knijff P (2007) Allele frequency distribution for 21 autosomal STR loci in Nepal. Forensic Sci Int 168, no. 2-3p:227–231. https://doi.org/10.1016/j.forsciint.2006.02.014
Kraaijenbrink T, Driem V, George L, Gaselô OF, Tshering K, De Knijff P (2007) Allele frequency distribution for 21 autosomal STR loci in Bhutan. Forensic Sci Int 170(1p):68–72. https://doi.org/10.1016/j.forsciint.2006.04.006
Garcia-Bertrand R, Simms TM, Cadenas AM, Herrera RJ (2014) United Arab Emirates: phylogenetic relationships and ancestral populations. Gene 533(1p):411–419. https://doi.org/10.1016/j.gene.2013.09.092
Ghosh T, Kalpana D, Mukerjee S, Mukherjee M, Sharma AK, Nath S, Rathod VR, Thakar MK, Jha GN (2011) Genetic diversity of autosomal STRs in eleven populations of India. Forensic Sc Int Genetics 5(3p):259–261. https://doi.org/10.1016/j.fsigen.2010.01.005
Chaudhari RR, Dahiya MS (2014) Genetic diversity of 15 autosomal short tandem repeats loci using the AmpFLSTR Identifiler kit in a Bhil Tribe population from Gujarat state India. Indian Human Genetics 20:148–152. https://doi.org/10.4103/0971-6866.142879
Mohapatra, B. K., Kamal Chauhan, U. S., Thakur, Bhuvnesh Yadav and Anupuma Raina. Genetic analysis and evolutionary relationship of Jammu and Kashmir Muslim population with short tandem repeat loci. Int J Current Res 2016. Vol. 8, p. 36398–36401
NOOR S, ALI S, EAASWARKHANTH M, HAQUE I (2009) Allele frequency distribution for 15 autosomal STR loci in Afridi Pathan population of Uttar Pradesh, India. Legal Med 11(6p):308–311. https://doi.org/10.1016/j.legalmed.2009.08.002
CHISHTI HM, MANICA A, ANSAR M, ERIKSSON A, AJMAL M, HAMEED A (2016) Inability of the most commonly used forensic genetic markers to distinguish between samples belonging to different ethnicities of Pakistan with diverse genetic background. Forensic Sci Int: Genetics 22:e7–e8. https://doi.org/10.1016/j.fsigen.2016.01.006
Funding
The authors are thankful for the financial support provided by the Swiss Government Excellence Scholarship for PhD studies of foreign students in Switzerland and by the Centre for Applied Molecular Biology, University of the Punjab, Lahore Pakistan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Anwar, I., Hussain, S., Rehman, A.U. et al. Genetic variation among the major Pakistani populations based on 15 autosomal STR markers. Int J Legal Med 133, 1037–1038 (2019). https://doi.org/10.1007/s00414-018-1951-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-018-1951-0