Abstract
Purpose
Cancer cells with stem cell-like properties may contribute to cancer development and therapy resistance. The advancement of multi-omics technology has sparked interest in exploring cancer stemness from a multi-omics perspective. However, there is a limited number of studies that have attempted to subtype cancer by combining different types of stem cell signatures.
Methods
In this study, 10,323 cancer specimens from 33 TCGA cancer types were clustered based on the enrichment scores of six stemness gene sets, representing two types of stem cell backgrounds: embryonic stem cells (ESCs) and hematopoietic stem cells (HSCs).
Results
We identified four subtypes of pan-cancer, termed StC1, StC2, StC3 and StC4, which displayed distinct molecular and clinical features, including stemness, genome integrity, intratumor heterogeneity, methylation levels, tumor microenvironment, tumor progression, responses to chemotherapy and immunotherapy, and survival prognosis. Importantly, this subtyping method for pan-cancer is reproducible at the protein level.
Conclusion
Our findings indicate that the ESC signature is an adverse prognostic factor in cancer, while the HSC signature and ratio of HSC/ESC signatures are positive prognostic factors. The subtyping of cancer based on ESC and HSC signatures may provide insights into cancer biology and clinical implications of cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Stem cells, characterized by two critical abilities: self-renewal and differentiation, are the foundation for various organs and tissues [1]. There are different types of stem cells, including embryonic stem cells (ESCs), adult stem cells (ASCs), and induced pluripotent stem cells (iPSCs). ESCs originate from early embryos known as blastocysts and exhibit pluripotency. In contrast, ASCs, also known as tissue-specific stem cells or somatic stem cells, generate specific cell types for specific tissues or organs and are not pluripotent [1]. ASCs encompass various subtypes, such as hematopoietic stem cells (HSCs), mesenchymal stem cells (MSCs), neural stem cells (NSCs), epithelial stem cells, and skin stem cells. Among these, HSCs are the most representative ASCs and serve as a standard model for studying tissue-specific stem cells [1]. HSCs play a crucial role in generating all types of blood cells, including red blood cells, white blood cells, and platelets. White blood cells, also known as leukocytes, constitute a vital part of the body’s immune system, including lymphocytes, monocytes, neutrophils, eosinophils, basophils, and macrophages.
Cancer stem cells (CSCs) refer to cancer cells with stem cells-like characteristics. CSCs were initially discovered in hematopoietic malignancies [2] and later were found in various solid tumors, such as pancreatic cancer [3], melanoma [4], breast cancer [5], and head and neck cancer [6]. These CSCs typically constitute a small fraction of the total cancer cell population and are characterized by their ability to continuously self-renew, proliferate, undergo dedifferentiation, promote cancer progression, and confer resistance to therapy [7]. CSCs are often endowed with dysregulated proliferative pathways or oncogenic mutations to maintain tumor growth [8]. Furthermore, different levels of stemness of cancer cells contribute to the intratumor heterogeneity (ITH) [9]. This heterogeneity allows drugs to target a portion but not all of the cancer cells, which is a key factor contributing to therapy resistance and cancer relapse. CSCs express many genes in common with early ESCs, such as OCT4, NANOG, and SOX2 [10]. Many of these genes are transcriptional regulators of ESCs and may synergistically regulate tumor cell self-renewal and proliferation [11]. Furthermore, since HSCs are the origin of all lymphocytes and myeloid cells [12], they play important roles in the immune regulation of various diseases, including cancer [13]. A previous study has suggested that both hematopoietic malignancies and solid tumors may utilize the hematopoietic stem cell niche of the bone marrow to promote tumor growth and suppress antitumor immunity [14]. Another study has shown that hematopoietic stem and progenitor cells (HSPCs) are involved in tumor progression [15]. Considering that uncontrolled cell proliferation and immunosuppression are hallmarks of cancer [16], both ESCs and HSCs-like signatures are significant in cancer biology. However, the majority of previous investigations primarily focused on ESCs, with only limited investigations into cancer stemness through simultaneous analysis of ESC and HSC signatures. To fill the knowledge gap, this study explored both ESCs and HSCs-like signatures in pan-cancer.
With the recent advancement of multi-omics technology, abundant cancer-associated genomics, epigenomics, transcriptomics, and proteomics data have emerged, such as The Cancer Genome Atlas (TCGA, https://cancergenome.nih.gov/) and The International Cancer Genome Consortium (ICGC, https://dcc.icgc.org/). These data have been widely utilized to explore various aspects of tumor characteristics across diverse cancer types, such as tumor immunity [17], tumor stemness [18], tumor metabolism [19], and ITH [20]. For example, Thorsson et al. identified six immune-specific subtypes of the pan-cancer of 33 TCGA cancer types based on a transcriptomic analysis [21]. Another pan-cancer analysis of transcriptomes in these 33 cancer types unveiled three metabolic expression subtypes with distinct clinical outcomes [19]. Additionally, Hoadley et al. performed clustering of around 10,000 cancer samples from the same 33 TCGA cancer types based on data of aneuploidy, DNA hypermethylation, mRNA, miRNA, and protein expression levels. Their work identified 28 clusters significantly related to histology, tissue type, or anatomic origin [22]. The multi-omics-based exploration of tumor stemness has attracted certain interests. For example, multi-omics analyses have shown that tumor stemness is associated with dedifferentiated oncogenic phenotype, immunosuppression, ITH, metastasis, and drug resistance [7, 18]. Transcriptomes-based stemness scores have been utilized to classify cancers, including bladder urothelial carcinoma (BLCA) [23], lung adenocarcinoma (LUAD) [9], and glioblastoma multiforme (GBM) [24]. Despite these previous studies, the omics-based investigation of different stem cell backgrounds in pan-cancer or individual cancer types remains lacking. Furthermore, the omics-based exploration of tumor stemness at the single-cell level remains inadequate, although there is already an abundance of single-cell omics data available for various cancers [25,26,27,28,29].
In this study, we conducted a clustering analysis of 10,323 specimens representing the 33 TCGA cancer types based on transcriptomes-based scores of six stemness gene sets, which represent two types of stem cell backgrounds, namely ESCs and HSCs. This analysis identified four distinct stemness subtypes of pan-cancer. We further comprehensively compared the molecular and clinical features among these subtypes, as well as their associations with the response to immunotherapy and chemotherapy.
2 Materials and methods
2.1 Data acquisition
We downloaded data of gene expression profiles (RSEM normalized and batch effects adjusted), somatic mutation profiles (level 3) and clinical features for the TCGA Pan-Cancer (PANCAN) cohort consisting of 33 TCGA cancer types from UCSC Xena (https://xenabrowser.net/datapages/). The 33 cancer types included adrenocortical carcinoma (ACC), BLCA, breast invasive carcinoma (BRCA), cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC), cholangiocarcinoma (CHOL), colon adenocarcinoma (COAD), lymphoid neoplasm diffuse large B-cell lymphoma (DLBC), esophageal carcinoma (ESCA), GBM, head and neck squamous cell carcinoma (HNSC), kidney chromophobe (KICH), kidney renal clear cell carcinoma (KIRC), kidney renal papillary cell carcinoma (KIRP), acute myeloid leukemia (LAML), brain lower grade glioma (LGG), liver hepatocellular carcinoma (LIHC), LUAD, lung squamous cell carcinoma (LUSC), mesothelioma (MESO), ovarian serous cystadenocarcinoma (OV), pancreatic adenocarcinoma (PAAD), pheochromocytoma and paraganglioma (PCPG), prostate adenocarcinoma (PRAD), rectum adenocarcinoma (READ), sarcoma (SARC), skin cutaneous melanoma (SKCM), stomach adenocarcinoma (STAD), testicular germ cell tumors (TGCT), thyroid carcinoma (THCA), thymoma (THYM), uterine corpus endometrial carcinoma (UCEC), uterine carcinosarcoma (UCS), and uveal melanoma (UVM). In addition, we downloaded 15 normalized protein expression profiles in 12 cancer types (TCGA-BRCA, TCGA-COAD, TCGA-OV, BRCA, COAD, OV, GBM, HNSC, LUAD, LUSC, PAAD, KIRC, UCEC, STAD, LIHC) from the Clinical Proteomic Tumor Analysis Consortium (CPTAC, https://proteomics.cancer.gov/programs/cptac/) and the International Cancer Proteogenome Consortium (ICPC, https://icpc.cancer.gov/portal/). We also downloaded three single-cell RNA sequencing (scRNA-seq) datasets, namely GSE75688 for BRCA [29], GSE131309 for SARC [30], and GSE89567 for GBM [25], from the NCBI gene expression omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo/). Besides, we downloaded a scRNA-seq dataset for renal cell carcinoma (RCC) from a recent publication [31]. We obtained eight immunotherapy-related datasets for eight cancer types (READ, non-small cell lung carcinoma (NSCLC), LIHC, ESCA, urothelial cancer (UC), triple-negative breast cancer (TNBC), STAD, SKCM) from GEO and recent publications [32,33,34,35,36,37,38,39]. The immunotherapy-related datasets included gene expression profiles and clinical information pertaining to immune checkpoint blockade (ICB) treatment in cancer patients. A summary of these datasets is shown in Supplementary Table S1.
2.2 Data processing and quality control
The gene expression values were preprocessed and normalized in the single cell transcriptomes, except the BRCA scRNA-seq dataset, in which they were raw transcripts per million (TPM). For the BRCA scRNA-seq dataset, we performed data preprocessing and quality control prior to subsequent analyses. First, we replaced the TPM values less than one with zero. Second, all TPM values (x) were transformed by log2(x + 1). Third, we removed the genes with expression values of zero across all single cancer cells. For the other three scRNA-seq datasets, we also preprocessed them following the third step.
Proteomics data were processed by the publication [40] at the gene level rather than at the protein isoform level. The names of rows in the protein expression matrix refer to the protein-coding genes. For the pan-caner analysis, we merged the data from 15 protein expression profiles into an expression matrix with the function “merge ()” in the R package “base.” Subsequently, we adjusted for batch effects and normalized combined data using the ‘‘normalizeBetweenArrays” function in the R package “limma”.
2.3 Collection of stemness signatures
We collected six stemness signatures (or gene sets) from the StemChecker webserver (http://stemchecker.sysbiolab.eu). Among these stemness signatures, three are markers of ESCs, including Hs_ESC_Assou, Hs_ESC_Bhattacharya, and Hs_ESC_Wong; the other three are markers of HSCs, including Hs_HSC_Huang, Hs_HSC_Novershtern, and Hs_HSC_Toren. These stemness gene sets are presented in Supplementary Table S2.
2.4 Single-sample gene-set enrichment analysis
To determine the enrichment score of a given gene set, which represents a stemness signature, biological process, pathway, or phenotypic feature, within a tumor bulk or single cancer cell, we employed the single-sample gene-set enrichment analysis (ssGSEA) method [41]. The ssGSEA algorithm calculates a gene set’s enrichment score in a sample based on its expression profile. We implemented this analysis with the “GSVA” R package. The gene sets analyzed are presented in Supplementary Table S2.
2.5 Clustering analysis
We used hierarchical clustering to identify stemness subtypes of cancers based on their enrichment scores of the six stemness signatures. This analysis was conducted using the R package “hclust” with the parameters: method = “ward.D2” and members = NULL. Prior to clustering, the data of ssGSEA scores were transformed by z-score and translated into distance matrices by the “dist()” function with the parameter: method = “Euclidean”.
2.6 Survival analysis
We utilized the Kaplan-Meier (K-M) model to compare overall survival (OS), disease-specific survival (DSS), disease-free interval (DFI), and progression-free interval (PFI) among different subgroups of cancer patients. Log-rank tests were employed to assess the significance of their differences. These analyses were implemented with the function “survfit()” in the R package “survival.” Furthermore, we performed multivariate survival analysis by the Cox proportional hazards model to explore the correlations of the signatures of ESCs and HSCs, and the ratio of HSCs/ESCs with survival prognosis in pan-cancer after correcting for confounding variables, including age, sex, and tumor stage. The “age”, “ESCs”, “HSCs”, and “ratio of HSCs/ESCs” were continuous variables, and the “sex” (male versus female) and “tumor stage” (early (stage I-П) versus late (stage III-IV)) were binary variables. The multivariate survival analysis was conducted using the function “coxph()” in the R package “survival”.
2.7 Evaluation of genomic instability, ITH, and immune scores
Somatic mutations and copy number alterations (CNAs) reflect genomic instability. The tumor mutation burden (TMB) was defined as the total number of somatic mutations in the tumor. We obtained CNA scores in TCGA pan-cancer from the publication by Knijnenburg et al. [42]. We used GISTIC2 [43] to calculate G-scores in tumors with the input of “SNP6” files. The G-score reflects the amplitude of the somatic CNA and the frequency of its occurrence across a group of samples [43]. Besides, we evaluated ITH in tumors utilizing the DEPTH algorithm [20], which measures ITH at mRNA level based on the heterogeneity of gene expression perturbations. In addition, we employed the ABSOLUTE algorithm [44] to evaluate tumor aneuploidy, namely ploidy scores, with the input of “SNP6” files. To gauge immune infiltration levels within tumors, we calculated immune scores using the ESTIMATE algorithm [45] with the input of gene expression matrix.
2.8 Identification of marker genes for subtypes
To identify marker genes for a stemness subtype, we first identified the upregulated genes in the subtype versus each of the other subtypes using the threshold of Student’s t test adjusted P value < 0.01 and mean expression fold change > 2. The marker genes for the stemness subtype were the common genes among the sets of upregulated genes.
2.9 Identification of upregulated proteins for subtypes
To identify upregulated proteins in a stemness subtype, we compared the protein expression profiling in a stemness subtype with that in each of the other subtypes. The upregulated proteins for the stemness subtype were the common proteins identified as upregulated in the stemness subtype compared to each of the other subtypes, with the threshold of Student’s t test adjusted P value < 0.05 and mean expression fold change > 1.
2.10 Logistic regression analysis
We employed logistic regression models to predict tumors with higher immune scores (> median) versus those with lower immune scores (< median) with four predictors (ESC score, HSC score, CNA score, and TMB). In logistic regression analyses, all predictors’ values were normalized by z-score; the R function “glm” was utilized to fit the binary model with the parameter “family” as “binomial” and other parameters as their default values; the standardized regression coefficients (β values) were obtained with the function “lm.beta” in the R package “QuantPsyc”.
2.11 Statistical analysis
In class comparisons, for non-normally distributed data, we utilized Mann–Whitney U tests (for two classes) or Kruskal–Wallis (K–W) tests (for more than two classes); for normally distributed data, we used Student’s t tests (for two classes) or ANOVA tests (for more than two classes). In the analysis of contingency tables, we utilized Fisher’s exact tests or Chi-square tests. For evaluating the correlations between the enrichment of molecular features and stemness scores, we employed the Spearman method. To adjust P values in multiple tests, we utilized the Benjamini-Hochberg method [46] to calculate the false discovery rate (FDR). All of these statistical analyses were conducted within the R programming environment (version 4.2.2).
3 Results
3.1 Stemness signatures-based clustering analysis identifies four subtypes of pan-cancer
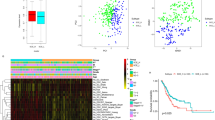
Based on the enrichment scores (ssGSEA scores) of the six stemness signatures, we identified four subtypes of pan-cancer by hierarchical clustering (Fig. 1A). We termed the four subtypes StC1, StC2, StC3, and StC4, respectively. StC1 displayed high enrichment of both ESC and HSC signatures. StC2 showed high enrichment of ESC but low enrichment of HSC signatures. In contrast to StC2, StC3 had low enrichment of ESC but high enrichment of HSC signatures. StC4 displayed the lowest enrichment of both ESC and HSC signatures. We defined the enrichment score of the ESC signature in a tumor as the average enrichment scores of the three ESC signatures in the tumor, denoted as the ESC score. Similarly, the enrichment score of the HSC signature was calculated as the average enrichment scores of the three HSC signatures, termed the HSC score. As expected, ESC and HSC scores were significantly different among the four subtypes: StC1 > StC2 > StC3 > StC4 for ESC and StC3 > StC1 > StC2 > StC4 for HSC (K–W test, P = 0) (Fig. 1B). Additionally, the ratios of HSC to ESC signatures, defined as HSC score divided by ESC score, were also significantly different among the four subtypes: StC3 > StC1 > StC2 > StC4 (P = 0) (Fig. 1B).
Identifying subtypes of pan-cancer based on stemness signatures. A Hierarchical clustering identifies four stemness subtypes of TCGA pan-cancer: StC1, StC2, StC3 and StC4 based on the enrichment scores of six stemness gene sets, representing two types of stem cell backgrounds: embryonic stem cells (ESCs) and hematopoietic stem cells (HSCs). HPV: human papillomavirus, HBV: hepatitis B virus, EBV: Epstein–Barr virus, Ad: adenocarcinomas, GI: gastrointestinal, GYN: gynecological, NA: not available. B Comparisons of HSC scores, ESC scores and ratios of HSC/ESC signatures among the four stemness subtypes of TCGA pan-cancer. The HSC score in a tumor is the average enrichment scores of the three HSC signatures. The ESC score in a tumor is the average enrichment scores of the three ESC signatures. The ratio of HSC/ESC signatures in a tumor is the ratio of HSC score over ESC score. C Kaplan–Meier curves show that StC3 and StC2 likely have the best and worst 10-year OS, and that StC4 and StC2 likely have the best and worst 10-year DSS, DFI and PFI in pan-cancer. The log-rank test P values are shown. OS: overall survival, DSS: disease-specific survival, PFI: progression-free interval, DFI: disease-free interval. D Kaplan–Meier curves show that the tumors with high ESC scores (> median) have significantly worse prognosis than the tumors with low ESC scores (< median) in all four 10-year endpoints (OS, DSS, DFI and PFI) in pan-cancer; In contrast, the tumors with high HSC scores or ratios of HSC/ESC signatures (> median) have significantly better prognosis than the tumors with low HSC scores or ratios of HSC/ESC signatures (< median) in all four 10-year endpoints in pan-cancer. The log-rank test P values are shown. E Multivariate Cox proportional hazards regression analysis show that ESC scores have a significant inverse correlation with OS, and that HSC scores and the ratios of HSC/ESC signatures have a significant positive correlation with OS in pan-cancer. The “age”, “ESC score”, “HSC score”, and “ratio of HSC/ESC” are continuous variables, and the “sex” (male versus female) and “tumor stage” (early (stage I-П) versus late (stage III-IV)) are binary variables
We compared four 10-year endpoints (OS, DSS, DFI, and PFI) among the four subtypes, revealing a nearly consistent pattern: StC3 > StC4 > StC1 > StC2 for OS, and StC4 > StC3 > StC1 > StC2 for the other three endpoints (Fig. 1C). This pattern underscores significant prognostic disparities among these stemness subtypes. Specifically, StC3 exhibited the most favorable OS; StC4 displayed the best PFI outcome. In contrast, StC2 displayed the poorest OS, DSS, DFI, and PFI, with StC1 likely following as the second poorest in these endpoints. Intriguingly, StC2 and StC3 exhibited opposite patterns in the expression of both ESC and HSC signatures, yet StC2 had the worst prognosis compared to StC3, which had the best OS prognosis. These findings indicate that the expression of ESC signatures may serve as an adverse prognostic factor, whereas the expression of HSC signatures appears to be a positive prognostic indicator. Indeed, the tumors with high ESC scores (> median) displayed significantly worse outcomes than those with low ESC scores (< median) across all four endpoints (P < 0.001) (Fig. 1D). Conversely, tumors with high expression of HSC signatures demonstrated significantly improved outcomes in all four endpoints compared to those with low HSC signature expression (P < 0.001) (Fig. 1D). Similarly, tumors with high ratios of HSC/ESC signatures exhibited markedly better prognosis than those with low ratios across all four endpoints (P = 0) (Fig. 1D).
To explore whether the significant correlation between the enrichment of ESC and HSC signatures, or the ratio of HSC/ESC signatures, and prognosis was confounded by other variables, such as age, sex, and tumor stage, we conducted multivariate survival analyses using the multivariate Cox proportional hazards model. These analyses confirmed that the ESC signature remained a significant risk factor, while the HSC signature and ratio of HSC/ESC signatures served as protective factors in pan-cancer (Fig. 1E and Supplementary Fig. S1).
Furthermore, we compared the enrichment levels of ESC and HSC signatures between tumor and normal samples. This analysis revealed that tumor samples exhibited higher enrichment levels of ESC signatures than normal samples, while the enrichment levels of HSC signatures in tumor samples were lower than those in normal samples (Supplementary Fig. S2).
3.2 Distribution of the stemness subtypes across individual cancer types
For individual cancer types, there were the highest proportions of BLCA, BRCA, CESC, COAD, HNSC, LUAD, LUSC, READ, TGCT, THYM, and UCS in StC1, the highest proportions of DLBC, ESCA, LIHC, OV, SKCM, STAD, and UVM in StC2, the highest proportions of ACC, CHOL, GBM, KICH, KIRC, KIRP, LGG, MESO, LUAD, PAAD, PCPG, PRAD, SARC, and THCA in StC3, and the highest proportions of LAML and UCEC in StC4 (Fig. 2A). Notably, most CESC (84.7%), LUSC (86.7%), TGCT (93.0%) and UCS (91.2%) tumors belonged to StC1, indicating the high enrichment of both ESC and HSC signatures in these cancer types. StC2 harbored all DLBC and most of ESCA (85.4%), LIHC (81.2%), OV (94.5%), and UVM (97.5%) tumors, suggesting the high enrichment of ESC but low enrichment of HSC signatures displayed in these cancer types. In contrast, StC3 was dominated by most of KICH (97.0%), KIRC (97.0%), KIRP (94.5%), LGG (95.1%), PAAD (82.7%), PCPG (98.9%), PRAD (93.2%), and THCA (98.8%) tumors, suggesting the low enrichment of ESC but high enrichment of HSC signatures shown in these cancer types. Finally, StC4 contained all LAML cases, indicating the low enrichment of both ESC and HSC signatures in this cancer type. Interestingly, approximately 70% of squamous cell carcinomas, such as CESC, HNSC, and LUSC, were classified into StC1 (Fig. 2B), highlighting their high stemness. Moreover, a majority (96.2%) of kidney cancers, including KIRC, KICH, and KIRP, were grouped into StC3 (Fig. 2B). It suggests that kidney cancers have high enrichment of HSC signatures but low enrichment of ESC signatures.
Distribution of the stemness subtypes across individual cancer types. A Proportions of tumor samples belonging to each stemness subtype in each of the 33 TCGA cancer types. Only the proportions over 10% are shown in the bars. B The Sankey diagram shows the stemness-subtype composition of pan-kidney and pan-squamous cell cancers. C Comparisons of the enrichment scores of ESC and HSC signatures and their ratios across the 33 TCGA cancer types, ordered by the median value
We further compared the enrichment scores of ESC and HSC signatures and their ratios across the 33 TCGA cancer types (Fig. 2C). The analysis demonstrated that LAML, UCEC, PCPG, KICH, LGG, KIRP, and KIRC had the lowest median ESC scores, as compared to TGCT, UCS, CESC, LUSC, COAD, HNSC, and BLCA having the highest median ESC scores. Meanwhile, LAML, UCEC, DLBC, ESCA, STAD, and OV had the lowest median HSC scores, while UCS, KIRC, KICH, PRAD, PAAD, KIRP, and THCA had the highest median HSC scores. The ratios of HSC/ESC signatures exhibited similar trend to HSC scores. This analysis indicated that the cancer types with more favorable prognoses like kidney cancer, prostate cancer, thyroid cancer, and low-grade gliomas, showed a high enrichment of the HSC signature as well as a high ratio of HSC/ESC signatures. Contrastively, upper gastrointestinal cancer (such as esophageal, gastric, and liver cancer) and several gynecologic cancer (such as endometrial and ovarian cancer) displayed low enrichment of the HSC signature and a low ratio of HSC/ESC signatures.
3.3 Correlates of the stemness subtypes with clinicopathologic features in pan-cancer
Among the four stemness subtypes, the proportion of early-stage (stage I-II) tumors followed the pattern: StC3 (74.0%) > StC4 (73.0%) > StC1 (52.4%) > StC2 (29.2%) (Chi-square test, P = 3.0 × 10−59) (Fig. 3A). Additionally, a similar trend was observed in the proportion of low-grade tumors (G1-2) (Fig. 3A). These results conform to the prognostic difference among these subtypes. We further explored the association of the stemness subtypes with therapeutic responses in pan-cancer. Notably, StC4 exhibited the highest response rate (94.4%) to chemotherapy (P = 3.0 × 10−16) (Fig. 3A). It is justified since StC4 has the lowest stemness signatures to facilitate chemotherapy response [47].
Comparisons of clinicopathologic features among the stemness subtypes of pan-cancer. A The proportions of early-stage (stage I-II) tumors and low-grade (G1-2) tumors, and the response (complete or partial response) rates to chemotherapy among the four stemness subtypes in TCGA pan-cancer. Chi-square test, P < 0.001. B Comparisons of the ratios of HSC/ESC signatures between early-stage and late-stage cancer cells in BRCA, low-grade and high-grade cancer cells in GBM, and primary and metastatic cancer cells in SARC. The one-tailed Mann–Whitney U test P values are shown
We further explored the association between stemness and clinicopathologic features in several cancer single-cell datasets. We observed that the ratios of HSC/ESC signatures were elevated in cancer cells of early-stage tumors compared to those in late-stage tumors in BRCA (P = 0.05) (Fig. 3B). In another dataset for GBM, the ratios of HSC/ESC signatures were markedly higher in cancer cells of low-grade tumors than in those of high-grade tumors (P = 3.6 × 10−5) (Fig. 3B). Furthermore, in SARC, the ratios of HSC/ESC signatures were significantly higher in cancer cells of primary tumors compared to those of metastatic tumors (P = 1.8 × 10−4) (Fig. 3B). These findings substantiate the notion of the ratio of HSC/ESC signatures being a positive prognostic factor in tumors.
3.4 Correlates of the stemness subtypes with molecular features in pan-cancer
Both increased TMB and CNAs reflect genomic instability [48]. We found TMB of the four stemness subtypes following the pattern: StC2 > StC1 > StC3 > StC4 (K–W test, P = 2.0 × 10−279) (Fig. 4A), and CNA scores following the pattern: StC2 > StC1 > StC4 > StC3 (K–W test, P = 6.3 × 10−215) (Fig. 4B). Additionally, G-scores of copy number amplifications and deletions followed the same pattern as CNA scores (Fig. 4C). Furthermore, ESC scores showed significant positive correlations with both TMB and CNA scores in pan-cancer, whereas HSC scores demonstrated significant negative correlations with both TMB and CNA scores (Fig. 4D). Moreover, ESC and HSC scores were observed to have significant positive and negative correlations, respectively, with microsatellite instability (MSI) scores evaluated by MSIsensor [49] in pan-cancer(P < 0.001) (Fig. 4D). Collectively, these results indicate that the enrichment of ESC and HSC signatures positively and negatively correlates with genomic instability, respectively. Notably, The ITH scores by DEPTH [20] were the highest in StC2 and the lowest in StC3, suggesting that the enrichment of ESC and HSC signatures positively and negatively correlates with ITH, respectively (P = 0) (Fig. 4E). Indeed, ESC scores displayed a positive correlation with ITH scores in pan-cancer (Spearman correlation, ρ = 0.13; P = 3.8 × 10−33), while HSC scores showed a negative correlation with ITH scores (ρ = -0.44; P = 0) (Fig. 4F).
Comparisons of genome instability and intratumor heterogeneity among the stemness subtypes of pan-cancer. Comparisons of TMB (A), CNA scores (B), G-scores (C), and ITH scores (E) among the four stemness subtypes. The G-scores were calculated by GISTIC2 [43]. Heatmap (D) and scatter plots (F) show that ESC scores have significant positive correlations with MSI (D), TMB (D), CNA (D), and ITH scores (F), while HSC scores have significant negative correlations with MSI, TMB, CNA, and ITH scores in pan-cancer. TMB: tumor mutation burden, CNA: copy number alterations, ITH: intratumor heterogeneity, MSI: microsatellite instability. The Kruskal–Wallis test P values (A, B, E) and the Spearman correlation coefficients (ρ) and P values (D, F) are shown
Furthermore, we compared the somatic mutation profiles among the four stemness subtypes. Figure 5 A displayed top ten genes with the highest mutation rates in each subtype. TP53 showed the highest mutation rate (52%) in StC1 and the lowest mutation rate (20%) in StC3, despite being the most frequently mutated gene in StC3. PTEN, PIK3CA, ARID1A, APC, KRAS, and FAT4 exhibited the highest mutation rates in StC4 among the four subtypes.
Comparisons of molecular features among the stemness subtypes of pan-cancer. A Comparisons of somatic mutation profiles among the four stemness subtypes in pan-cancer. The oncoplot displays top ten genes having the highest mutation rates in each subtype. B Comparisons of the enrichment scores of mismatch repair, homologous recombination, cell cycle, epithelial-mesenchymal transition (EMT) and angiogenesis signatures among the four stemness subtypes. The Kruskal–Wallis test P values are shown. C Heatmap shows the expression levels of marker genes for each of the four stemness subtypes in pan-cancer (two-tailed Student’s t test, FDR < 0.01). The gene ontology (GO) biological process (BP) enriched in the stemness subtypes are shown on the right (FDR < 0.05). D Left: comparisons of global methylation levels among the four stemness subtypes in pan-cancer. The mean global methylation level and error bar of each subtype are shown. Right: the scatter plots show that HSC scores have significant positive correlations with global methylation levels, while ESC scores have significant negative correlations with global methylation levels in pan-cancer. The one-tailed Mann–Whitney U test P values and the Spearman correlation coefficients (ρ) and P values are shown. * P < 0.05, *** P < 0.001, ns not significant
Transcriptome analysis demonstrated that the enrichment scores of the mismatch repair, homologous recombination, and cell cycle signatures followed the pattern: StC1 > StC2 > StC3 > StC4 (P = 0) (Fig. 5B), aligning with the pattern of the enrichment scores of the ESC signature. It is rationalized as increased tumor stemness is linked to heightened genomic instability and cell proliferation potential in cancer [7]. In contrast, the enrichment scores of the epithelial-mesenchymal transition (EMT) and angiogenesis signatures followed the pattern: StC3 > StC1 > StC2 > StC4 (P = 0) (Fig. 5B), consistent with the enrichment pattern of the HSC signature. The positive association between the EMT and angiogenesis signatures and the HSC signature is imaginable since the HSC signature tends to be enriched in the tumor stromal microenvironment. Additionally, transcriptome analysis identified marker genes for each stemness subtype (Fig. 5C and Supplementary Table S3). Based on these markers, we determined biological processes enriched in the subtypes using the GO database by g: Profiler [50]. StC1 showed enrichment in developmental process, embryonic morphogenesis, embryo development, and tissue homeostasis, in line with the high enrichment of both ESC and HSC signatures in this subtype. StC2 was enriched for metabolic process, inflammatory response, regulation of blood coagulation, and humoral immune response. StC3 displayed enrichment in nervous system development, blood circulation, blood vessel development, angiogenesis, regulation of transporter activity, and extracellular matrix organization. StC4 was enriched for immune system process, and embryonic skeletal system development (Fig. 5C).
DNA methylation analysis demonstrated that global methylation levels [51] were the highest in StC3 and the lowest in StC2 (P < 0.001) (Fig. 5D). It suggests that the HSC and ESC signatures correlate positively and negatively with global methylation levels, respectively. We further confirmed this conclusion by Spearman rank correlation tests (Fig. 5D).
3.5 Correlates of the ESC and HSC signatures with antitumor immune responses and immunotherapy responses
We found that HSC scores and ratios of HSC/ESC signatures correlated negatively with immune scores in pan-cancer (Spearman correlation test, P = 9.2 × 10−124 and 1.3 × 10−119, respectively) (Fig. 6A). Furthermore, the ratios of immunostimulatory to immunosuppressive signatures (pro-inflammatory/anti-inflammatory cytokines and M1/M2 macrophages) showed negative correlations with HSC scores and ratios of HSC/ESC signatures, while they displayed positive correlations with ESC scores (Fig. 6A). These results suggest that heightened enrichment of HSC and ESC signatures is associated with decreased and increased antitumor immune responses, respectively. Previous results have demonstrated that the enrichment of HSC and ESC signatures has significant correlations with aneuploidy and TMB, which have been revealed to correlate significantly with antitumor immune responses [17]. Hence, the significant correlations between HSC and ESC signatures and antitumor immune responses could be associated their associations with aneuploidy and TMB. To explore this conjecture, we respectively predicted immune scores, ratios of pro-inflammatory/anti-inflammatory cytokines and ratios of M1/M2 macrophages (high (> median) versus low (< median)) using four variables: ESC score, HSC score, CNA score, and TMB, by logistic regression analyses. These analyses showed that ESC score and HSC score were significant positive and negative predictors of the two ratios of immunostimulatory to immunosuppressive signatures, respectively. HSC score was a significant negative predictor of immune scores (β = -1.05; P = 1.4 × 10−46), while ESC score showed a positive trend in predicting immune scores, albeit not significant (β = 0.13; P = 0.06) (Fig. 6B). These findings imply that the correlations of HSC and ESC signatures with antitumor immune responses are likely independent of their associations with aneuploidy and TMB.
Correlations of ESC and HSC signatures with antitumor immune responses and immunotherapy responses. A The bar charts show the correlations of ESC scores, HSC scores and ratios of HSC/ESC signatures with immune scores, M1/M2 macrophages and pro-inflammatory/anti-inflammatory cytokines in pan-cancer. The Spearman correlation P values are shown. B Logistic regression analysis show that ESC score and HSC score are a positive and negative predictor of ratios of pro-inflammatory/anti-inflammatory cytokines, ratios of M1/M2 macrophages, and immune scores, respectively, after correcting for TMB and CNA. The standardized regression coefficients (β) and P values are shown. C The correlations of HSC and ESC signatures and their ratios with anti-PD-1/PD-L1 immunotherapy responses in several cancer cohorts. NSCLC: non-small cell lung carcinoma, UC: urothelial cancer, TNBC: triple-negative breast cancer. The one-tailed Mann–Whitney U test P values are shown. D The correlations of HSC and ESC signatures and their ratios with ICB responses in a scRNA-seq dataset for RCC. ICB: immune checkpoint blockade. The one-tailed Mann–Whitney U test P values are shown
We further analyzed correlations of HSC and ESC signatures with anti-PD-1/PD-L1 immunotherapy responses in several cancer cohorts. In six cancer cohorts, HSC scores appeared higher in non-responsive than in responsive cancers (P < 0.1) (Fig. 6C). Likewise, ratios of HSC/ESC signatures showed the same trend in five cancer cohorts (P < 0.1) (Fig. 6C). However, ESC scores were likely higher in responsive than in non-responsive cancers in four cancer cohorts (P < 0.1) (Fig. 6C). In a single-cell transcriptome dataset for RCC [31], we also observed higher ESC scores in cancer cells from ICB-responsive tumors than in those from ICB-non-responsive tumors (P < 0.001) (Fig. 6D). Conversely, ratios of HSC/ESC signatures were markedly lower in cancer cells from ICB-non-responsive tumors than in those from ICB-responsive tumors (P < 0.001). However, there was no significant difference of HSC scores between these two groups (P = 0.29) (Fig. 6D). In summary, these results suggest that the enrichment of HSC and ESC signatures is negatively and positively associated with immunotherapy responses, respectively.
3.6 Correlates of the ESC and HSC signatures with clinicopathologic and etiological factors in pan-cancer
We found ESC scores to be higher in late-stage than in early-stage cancers (P = 0.02). In contrast, HSC scores and ratios of HSC/ESC signatures were significantly lower in late-stage than in early-stage cancers (P = 2.8 × 10−10 and 4.0 × 10−9, respectively) (Fig. 7A). In addition, ESC scores were notably higher in high-grade compared to low-grade cancers (P = 2.9 × 10−13), while the ratios of HSC/ESC signatures were significantly lower in high-grade than in low-grade cancers (P = 2.9 × 10−4) (Fig. 7B). These results further support that the ESC signature, HSC signature, and ratio of HSC/ESC signatures is a negative, positive, and positive prognostic factor in cancer, respectively.
Correlations of ESC and HSC signatures with clinicopathologic and etiological factors in pan-cancer. Comparisons of HSC scores, ESC scores and ratios of HSC/ESC signatures between early-stage and late-stage (A), low-grade and high-grade (B), smoker and non-smoker (D), HPV/HBV/EBV-positive and -negative (E), and female and male (F) cancers, respectively. Correlations of HSC and ESC signatures and their ratios with age (C), hypoxia scores (G), TERT mRNA abundance (H), and the expression levels of ESR1, AR and their ratios (I). ESR1: estrogen receptor 1, AR: androgen receptor. The one-tailed Mann–Whitney U test P values are shown in (A, B, D, E, F), and the Spearman correlation coefficients (ρ) and P values are shown in (C, G, H). *** P < 0.001, ns not significant
Aging, tobacco, and viral infection [52,53,54] are common cancer risk factors. Interestingly, ESC and HSC scores exhibited significant, albeit opposing, correlations with ages in pan-cancer (Spearman correlation test, P < 0.001) (Fig. 7C). The ratios of HSC/ESC signatures showed a marked negative correlation with ages in pan-cancer (P = 6.8 × 10−58). Smokers displayed notably higher ESC scores than non-smokers in pan-cancer (P = 4.5 × 10−24), while the ratios of HSC/ESC signatures were significantly higher in non-smoker than in smoker cancers (P = 3.0 × 10−5) (Fig. 7D). Viral infections are important etiological factors for several cancers, such as human papillomavirus (HPV) for HNSC and CESC, hepatitis B virus (HBV) for LIHC, and Epstein–Barr virus (EBV) for lymphoma and nasopharyngeal cancers. Remarkably, HPV/HBV/EBV-positive cancers had higher ESC scores than negative cancers in pan-cancer, while the former group showed lower HSC scores and ratios of HSC/ESC signatures than the latter group (P < 0.1) (Fig. 7E). In addition, females demonstrated significantly higher ESC scores but lower HSC scores and ratios of HSC/ESC signatures than males in pan-cancer (P = 0.01, 1.3 × 10−6 and 1.6 × 10−14, respectively) (Fig. 7F).
Hypoxia, a fundamental trait of cancer, significantly influences cancer development [55]. Of note, hypoxia scores showed a significant positive, negative, and negative correlation with ESC scores, HSC scores, and ratios of HSC/ESC signatures, respectively (Fig. 7G). The expression of TERT (telomerase reverse transcriptase), vital for telomere structure and length maintenance [56], displayed a significant positive, negative, and negative correlation with ESC scores, HSC scores, and ratios of HSC/ESC signatures, respectively (P < 0.001; ρ = 0.35, -0.32 and − 0.50, respectively) (Fig. 7H). It indicates that the ESC-like signatures are associated with telomere lengthening in cancer cells, while the HSC-like signatures are related to telomere shortening. Additionally, the expression of ESR1 (estrogen receptor 1) and AR (androgen receptor) were positively correlated with HSC scores and ratios of HSC/ESC signatures, while AR expression levels were negatively correlated with ESC scores (Fig. 7I). Furthermore, the ratios of ESR1/AR displayed a notable positive correlation with ESC scores and negative correlations with HSC scores and ratios of HSC/ESC signatures. These findings align with the previous observation that females tend to have higher ESC scores and lower HSC scores and ratios of HSC/ESC signatures than males in pan-cancer.
3.7 Validation of the stemness signatures-based subtyping method in proteomics data
Furthermore, we performed a hierarchical clustering analysis of pan-cancer based on the enrichment scores of the six stemness signatures in proteomics data [40]. Consistent with the result from transcriptomics data, we identified four subtypes of pan-cancer, also termed StC1, StC2, StC3, and StC4, respectively (Fig. 8A). Likewise, we identified the marker proteins for each stemness subtypes (Supplementary Table S4). We also identified biological processes enriched in the subtypes based on the marker proteins using the R package “clusterProfiler” [57] (Fig. 8B). Specifically, StC1 was enriched for the biological processes representing active molecular activities and DNA repair, such as DNA replication, regulation of DNA repair, RNA splicing, RNA catabolic process, regulation of translation, and protein folding. It aligns with StC1 having high enrichment of both ESC and HSC signatures. StC2 showed enrichment in immune activities, including positive regulation of innate immune response, lymphocyte proliferation, antigen processing and presentation. It is consistent with the transcriptomics analysis indicating an enrichment of immune signatures in StC2. StC3 was enriched for processes involving regulation of angiogenesis, extracellular matrix organization, and regulation of wound healing, in accordance with the transcriptomics analysis result. StC4 exhibited enrichment in regulation of autophagy, and endosomal transport, similar to the finding from transcriptomics data.
Validation of the stemness signatures-based subtyping method in proteomics data. A Hierarchical clustering identifies four stemness subtypes (StC1, StC2, StC3 and StC4) based on protein expression profiles in pan-cancer. B Enrichment plots show the biological processes enriched in the four subtypes using GO database by the R package “clusterProfiler.” Each protein-coding gene is shown as gray pie. The biological processes were identified using a threshold of FDR < 0.05
Taken together, these analyses suggest that the stemness signatures-based subtyping method for pan-cancer is reproducible at the protein level.
4 Discussion
This study has identified four distinct subtypes of TCGA pan-cancer based on the enrichment of six stemness signatures associated with ESCs and HSCs. Each subtype displays unique molecular and clinical features. The subtype StC3, characterized by low enrichment of ESC but high enrichment of HSC signatures, has the best OS, the lowest aneuploidy level, ITH and TP53 mutation rate, the highest enrichment of the EMT and angiogenesis signatures, and the highest global methylation levels. This subtype contains a majority of kidney, pancreatic, prostate, thyroid, and lower grade glioma tumors. In contrast, the subtype StC2, characterized by high enrichment of ESC but low enrichment of HSC signatures, has the worst prognosis, the highest aneuploidy level, tumor mutation loads and ITH, and the lowest global methylation levels. This subtype involves all DLBC and most of esophageal, liver, ovarian, and uveal melanoma tumors. StC1, characterized by high enrichment of both ESCs and HSCs signatures, has the second worst prognosis, the highest TP53 mutation rate (52%), the highest enrichment of DNA mismatch repair, homologous recombination, and cell cycle signatures, and the second highest levels of genomic instability. StC1 harbors most squamous cell cancers, testicular germ cell tumors, and uterine carcinosarcomas. StC4 is characterized by the lowest enrichment of both ESC and HSC signatures; this subtype has the best progression-free survival, the highest response rate to chemotherapy, the lowest tumor mutation loads, the most frequent mutations in PTEN, PIK3CA, ARID1A, APC, KRAS, and FAT4, and the lowest enrichment of DNA mismatch repair, homologous recombination, cell cycle, EMT, and angiogenesis signatures. StC4 contains all acute myeloid leukemia cases.
This analysis further demonstrated that the ESC signature is a risk factor, while the HSC signature and ratio of HSC/ESC signatures are protective factors in cancer. Factors such as aging, smoking and viral infections appear to be associated with an increased enrichment of ESC signatures but a decreased enrichment of HSC signatures and the ratio of HSC/ESC signatures in cancer. A significant distinction between ESCs and HSCs is their different properties of cell divisions. ESCs expand themselves by symmetrical cell divisions [58], while HSCs maintain a balance between self-renewal and differentiation by asymmetric cell divisions [59]. Disruption of asymmetric cell divisions of stem cells may facilitate neoplastic transformation of stem cells [60]. Hence, the differences in prognostic associations between the ESC and HSC signatures could be attributed to their distinct properties of cell divisions. It has been reported that several proteins play key roles in maintaining asymmetric cell divisions, including Par3, Par6, aPKC, NUMA, LGN, Gαi, and Prospero [61]. Notably, we found that the expression levels of the genes encoding these proteins likely had negative correlations with ESC scores but significant positive correlations with HSC scores and ratios of HSC/ESC signatures in TCGA pan-cancer. In addition, p53 dysfunction can promote a shift from asymmetric to symmetric cell divisions [62]. We observed that TP53-mutated tumors had significantly higher ESC scores but significantly lower HSC scores and ratios of HSC/ESC signatures compared to TP53-wildtype tumors in TCGA pan-cancer. These data support that the ESC and HSC signatures are associated with increased and reduced symmetric cell divisions in cancer. Indeed, MKI67, a marker for proliferation, showed a significant positive expression correlation with ESC scores and negative expression correlations with HSC scores and ratios of HSC/ESC signatures in TCGA pan-cancer (ρ = 0.50, -0.39, and − 0.66, respectively).
Indeed, in addition to HSCs, MSCs and NSCs are tissue-specific stem cell types. MSCs have been reported to induce EMT in cancer cells [63,64,65]. We found that the enrichment levels of MSC signatures were positively correlated with the degree of stemness in the four subtypes, indicating that higher stemness might promote EMT in cancer. NSCs are primarily associated with brain tumors, and our findings showed that the enrichment levels of NSC signatures were the highest in StC3, aligning with the distribution of LGG and GBM primarily in StC3. Furthermore, we incorporated MSC and NSC signatures into clustering analysis, but could not identify stemness-related subtypes clearly. We further explored the impact of MSC and NSC signatures on cancer patients’ prognosis. It was observed that the patients with high MSC scores exhibited poorer survival outcomes compared to those with low MSC scores, while the opposite result was shown for NSC signatures. Nevertheless, the association between MSC or NSC signatures and survival was not as significant as those between HSC and ESC signatures and survival in pan-cancer (Supplementary Fig. S3).
An interesting finding is the negative correlation between HSC scores and tumor immune infiltration levels, while ESC scores exhibit a positive correlation with them. This finding appears inconsistent with reports from previous studies [7, 18, 24]. However, a previous report [14] has demonstrated that solid tumors accumulate immunosuppressive hematopoietic lineages within the tumor microenvironment. Another study has suggested that activation of HSC signatures can promote immunosuppression within the pre-metastatic niche of tumors [66]. Additionally, we utilized the CIBERSORT algorithm [67] to calculate the proportions of tumor immune cell infiltrations. We found that HSC scores and the ratios of HSC/ESC signatures were negatively correlated with the proportions of CD8 T cells, activated memory CD4 T cells, T follicular helper cells, M0 macrophages, M1 macrophages, and activated dendritic cells. However, ESC scores were positively correlated with the proportions of these immune cells. These data to some extent support our finding that high HSC scores or ratios of HSC/ESC signatures are associated with enhanced resistance to immunotherapy in tumors.
To conclude, this study performed a novel classification of pan-cancer in the context of HSC and ESC signatures and identified four stemness subtypes. Besides stemness features, these subtypes display different molecular and clinical characteristics, including genome integrity, ITH, methylation levels, tumor microenvironment, tumor progression phenotypes, chemotherapy and immunotherapy responses, and survival prognosis. The ESC signature is an adverse prognostic factor in cancer, while the HSC signature and ratio of HSC/ESC signatures are favorable prognostic factors. The HSC and ESC signatures-based subtyping of cancer provides new insights into tumor biology and has potential clinical implications for diagnosis, prognosis, and treatment of cancers.
Data availability
The authors declare that all data supporting the findings of this study are available within the paper and its supplementary information files.
References
W. Zakrzewski et al., Stem cells: past, present, and future. Stem. Cell. Res. Ther. 10(1), 68 (2019)
D. Bonnet, J.E. Dick, Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat. Med. 3(7), 730–737 (1997)
C. Li et al., Identification of pancreatic cancer stem cells. Cancer Res. 67(3), 1030–1037 (2007)
N.Y. Frank et al., ABCB5-mediated doxorubicin transport and chemoresistance in human malignant melanoma. Cancer Res. 65(10), 4320–4333 (2005)
M. Al-Hajj et al., Prospective identification of tumorigenic breast cancer cells. Proc. Natl. Acad. Sci .U. S. A. 100(7), 3983–3988 (2003)
M.E. Prince et al., Identification of a subpopulation of cells with cancer stem cell properties in head and neck squamous cell carcinoma. Proc. Natl. Acad. Sci. U. S. A. 104(3), 973–978 (2007)
A. Miranda et al., Cancer stemness, intratumoral heterogeneity, and immune response across cancers. Proc. Natl. Acad. Sci. U. S. A. 116(18), 9020–9029 (2019)
M. Al-Hajj et al., Therapeutic implications of cancer stem cells. Curr. Opin. Genet. Dev. 14(1), 43–47 (2004)
Q. Liu et al., Classification of lung adenocarcinoma based on stemness scores in bulk and single cell transcriptomes. Comput. Struct. Biotechnol. J. 20, 1691–1701 (2022)
B. van Schaijik et al., Subcellular localisation of the stem cell markers OCT4, SOX2, NANOG, KLF4 and c-MYC in cancer: a review. J. Clin. Pathol. 71(1), 88–91 (2018)
I. Ben-Porath et al., An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat. Genet. 40(5), 499–507 (2008)
S. Haas, A. Trumpp, M.D. Milsom, Causes and consequences of hematopoietic stem cell heterogeneity. Cell. Stem. Cell. 22(5), 627–638 (2018)
D. Carroll, D.K. St Clair, Hematopoietic stem cells: normal Versus Malignant. Antioxid. Redox. Signal 29(16), 1612–1632 (2018)
A.J. Giles, C.C. Reid, The functional interplay between systemic cancer and the hematopoietic stem cell niche. Pharmacol. Ther. 168, 53–60 (2016)
W.C. Wu et al., Circulating hematopoietic stem and progenitor cells are myeloid-biased in cancer patients. Proc. Natl. Acad. Sci. U. S. A. 111(11), 4221–4226 (2014)
D. Hanahan, R.A. Weinberg, Hallmarks of cancer: the next generation. Cell 144(5), 646–674 (2011)
T. Davoli et al., Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science. 355(6322), eaaf8399 (2017)
T.M. Malta, S.A. Gentles, Machine learning identifies stemness features Associated with Oncogenic Dedifferentiation. Cell 173(2), 338-354e15 (2018)
X. Peng et al., Molecular characterization and clinical relevance of metabolic expression subtypes in human cancers. Cell Rep 23(1), 255-269e4 (2018)
M. Li, Z.Z. Li, L. Wang, An algorithm to quantify intratumor heterogeneity based on alterations of gene expression profiles. Commun. Biol. 3(1), 505 (2020)
V. Thorsson, G.D. Brown, The Immune Landscape of Cancer. Immunity 48(4), 812-830e14 (2018)
K.A. Hoadley et al., Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of Cancer. Cell 173(2), 291-304e6 (2018)
C. Tang, M.J. Liu, X. Liu, Development and validation of a novel stem cell subtype for bladder cancer based on stem genomic profiling. Stem. Cell Res. Ther. 11(1), 457 (2020)
Z. Wang, W.Y. Yang, Machine learning revealed stemness features and a novel stemness-based classification with appealing implications in discriminating the prognosis, immunotherapy and temozolomide responses of 906 glioblastoma patients. Brief Bioinform. 22(5), bbab032 (2021)
A.S. Venteicher et al., Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq. Science 355(6332), eaai8478 (2017)
S. Chen, Z.G. Yang, Single-cell analysis reveals transcriptomic remodellings in distinct cell types that contribute to human prostate cancer progression. Nat. Cell. Biol. 23(1), 87–98 (2021)
S.V. Puram, T.I. Parikh, Single-cell transcriptomic analysis of primary and metastatic Tumor Ecosystems in Head and Neck Cancer. Cell 171(7), 1611-1624e24 (2017)
I. Tirosh, I.B. Prakadan, Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352(6282), 189–196 (2016)
W. Chung, E.H. Lee, Single-cell RNA-seq enables comprehensive tumour and immune cell profiling in primary breast cancer. Nat. Commun. 8, 15081 (2017)
L. Jerby-Arnon, C. Neftel, M.E. Shore et al., Opposing immune and genetic mechanisms shape oncogenic programs in synovial sarcoma. Nat. Med. 27, 289–300 (2021)
K. Bi, M. He, Z. Bakouny et al., Tumor and immune reprogramming during immunotherapy in advanced renal cell carcinoma. Cancer Cell. 39(5), 649-661e5 (2021)
M. Verstraete, D.A. Dekervel, Combining bevacizumab and chemoradiation in rectal cancer. Translational results of the AXEBeam trial. Br. J. Cancer 112(8), 1314–1325 (2015)
J.Y. Kim, C.J. Jung, Genome-wide methylation patterns predict clinical benefit of immunotherapy in lung cancer. Clin. Epigenetics 12(1), 119 (2020)
C.L. Hsu, O.D. Bai, Exploring markers of exhausted CD8 T cells to predict response to Immune checkpoint inhibitor therapy for Hepatocellular Carcinoma. Liver Cancer 10(4), 346–359 (2021)
van den T. Ende, d., C.N. van Berge Henegouwen et al., MI,., Neoadjuvant Chemoradiotherapy Combined with Atezolizumab for Resectable Esophageal Adenocarcinoma: A Single-arm Phase II Feasibility Trial (PERFECT) Clin Cancer Res. 27(12):3351–3359 (2021)
T.L. Rose, W.W. Mayhew, Fibroblast growth factor receptor 3 alterations and response to immune checkpoint inhibition in metastatic urothelial cancer: a real world experience. Br. J. Cancer 125(9), 1251–1260 (2021)
N.J. Birkbak, L.Y. Pathania, Overexpression of BLM promotes DNA damage and increased sensitivity to platinum salts in triple-negative breast and serous ovarian cancers. Ann. Oncol. 29(4), 903–909 (2018)
S.T. Kim, C.R. Bass, Comprehensive molecular characterization of clinical responses to PD-1 inhibition in metastatic gastric cancer. Nat. Med. 24(9), 1449–1458 (2018)
T.N. Gide, Q.C. Menzies, Distinct Immune cell populations define response to Anti-PD-1 monotherapy and Anti-PD-1/Anti-CTLA-4 combined Therapy. Cancer Cell. 35(2), 238-255e6 (2019)
Y. Zhang, F.C. Darshan, S. Chandrashekar, S. Varambally, C.J. Creighton, Proteogenomic characterization of 2002 human cancers reveals pan-cancer molecular subtypes and associated pathways. Nat. Commun. 13(1), 2669 (2022)
S. Hanzelmann, R. Castelo, J. Guinney, GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform 14, 7 (2013)
R.A. Burrell et al., The causes and consequences of genetic heterogeneity in cancer evolution. Nature 501(7467), 338–345 (2013)
C.H. Mermel et al., GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12(4), R41 (2011)
S.L. Carter et al., Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30(5), 413–421 (2012)
K. Yoshihara et al., Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013)
Y. Benjamini, Y. Hochberg, Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Royal Stat. Soc. B 57, 289–300 (1995)
L.T.H. Phi et al., Cancer Stem Cells (CSCs) in drug resistance and their therapeutic implications in cancer treatment. Stem. Cells. Int. 2018, 5416923 (2018)
S. Negrini, V.G. Gorgoulis, T.D. Halazonetis, Genomic instability–an evolving hallmark of cancer. Nat. Rev. Mol. Cell. Biol. 11(3), 220–228 (2010)
B. Niu et al., MSIsensor: microsatellite instability detection using paired tumor-normal sequence data. Bioinformatics 30(7), 1015–1016 (2014)
U. Raudvere et al., G:profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47(W1), W191–W198 (2019)
H. Jung et al., DNA methylation loss promotes immune evasion of tumours with high mutation and copy number load. Nat. Commun. 10(1), 4278 (2019)
L. Zhang et al., Analysis of clinical features and outcome of 356 triple-negative breast Cancer patients in China. Breast Care (Basel) 7(1), 13–17 (2012)
A.K. Witkiewicz et al., Using the reverse Warburg effect to identify high-risk breast cancer patients: stromal MCT4 predicts poor clinical outcome in triple-negative breast cancers. Cell Cycle. 11(6), 1108–1117 (2012)
D. Tang et al., The expression and clinical significance of the androgen receptor and E-cadherin in triple-negative breast cancer. Med. Oncol. 29(2), 526–533 (2012)
K. Ruan, G. Song, G. Ouyang, Role of hypoxia in the hallmarks of human cancer. J. Cell. Biochem. 107(6), 1053–1062 (2009)
X. Yuan, L.C. Xu, Mechanisms underlying the activation of TERT transcription and telomerase activity in human cancer: old actors and new players. Oncogene 38(34), 6172–6183 (2019)
T. Wu, H.E. Xu, clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innov. (Camb) 2(3), 100141 (2021)
T. Burdon, A. Smith, P. Savatier, Signalling, cell cycle and pluripotency in embryonic stem cells. Trends Cell Biol. 12(9), 432–438 (2002)
B. Giebel, I. Bruns, Self-renewal versus differentiation in hematopoietic stem and progenitor cells: a focus on asymmetric cell divisions. Curr. Stem. Cell. Res. Ther. 3(1), 9–16 (2008)
S. Gomez-Lopez, R.G. Lerner, C. Petritsch, Asymmetric cell division of stem and progenitor cells during homeostasis and cancer. Cell. Mol. Life Sci. 71(4), 575–597 (2014)
J. Zhu et al., LGN/mInsc and LGN/NuMA complex structures suggest distinct functions in asymmetric cell division for the Par3/mInsc/LGN and Galphai/LGN/NuMA pathways. Mol. Cell. 43(3), 418–431 (2011)
J. Bajaj, B. Zimdahl, T. Reya, Fearful symmetry: subversion of asymmetric division in cancer development and progression. Cancer Res. 75(5), 792–797 (2015)
R. Hass, J. von der Ohe, H. Ungefroren, Potential role of MSC/Cancer Cell Fusion and EMT for breast Cancer stem cell formation. Cancers (Basel). 11(10) (2019)
W.H. Lin et al., STAT3 phosphorylation at Ser727 and Tyr705 differentially regulates the EMT-MET switch and cancer metastasis. Oncogene 40(4), 791–805 (2021)
X. Zhang et al., Human colorectal cancer-derived mesenchymal stem cells promote colorectal cancer progression through IL-6/JAK2/STAT3 signaling. Cell Death Dis. 9(2), 25 (2018)
A.J. Giles et al., Activation of hematopoietic Stem/Progenitor cells promotes Immunosuppression within the pre-metastatic niche. Cancer Res. 76(6), 1335–1347 (2016)
A.M. Newman et al., Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12(5), 453–457 (2015)
Funding
This work was supported by the China Pharmaceutical University (grant numbers 3150120001 to XW).
Author information
Authors and Affiliations
Contributions
Jiali Lei: Methodology, Software, Validation, Formal analysis, Investigation, Data curation, Visualization, Writing - original draft, Writing - review & editing. Jiangti Luo: Software, Formal analysis, Data curation, Visualization. Qian Liu: Software, Data curation. Xiaosheng Wang: Conceptualization, Methodology, Resources, Investigation, Writing - original draft, Writing - review & editing, Supervision, Project administration, Funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1.18 MB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lei, J., Luo, J., Liu, Q. et al. Identifying cancer subtypes based on embryonic and hematopoietic stem cell signatures in pan-cancer. Cell Oncol. 47, 587–605 (2024). https://doi.org/10.1007/s13402-023-00886-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-023-00886-7