Abstract
Purpose
Pancreatic ductal adenocarcinoma (PDAC) is a highly aggressive malignancy. Up till now, the patient’s prognosis remains poor which, among others, is due to the paucity of reliable early diagnostic biomarkers. In the past, candidate diagnostic biomarkers and therapeutic targets have been delineated from genes that were found to be differentially expressed in normal versus tumour samples. Recently, new systems biology approaches have been developed to analyse gene expression data, which may yield new biomarkers. As of yet, the weighted gene co-expression network analysis (WGCNA) tool has not been applied to PDAC microarray-based gene expression data.
Methods
PDAC microarray-based gene expression datasets, listed in the Gene Expression Omnibus (GEO) database, were analysed. After pre-processing of the data, we built two final datasets, Normal and PDAC, encompassing 104 and 129 patient samples, respectively. Next, we constructed a weighted gene co-expression network and identified modules of co-expressed genes distinguishing normal from disease conditions. Functional annotations of the genes in these modules were carried out to highlight PDAC-associated molecular pathways and common regulatory mechanisms. Finally, overall survival analyses were carried out to assess the suitability of the genes identified as prognostic biomarkers.
Results
Using WGCNA, we identified several key genes that may play important roles in PDAC. These genes are mainly related to either endoplasmic reticulum, mitochondrion or membrane functions, exhibit transferase or hydrolase activities and are involved in biological processes such as lipid metabolism or transmembrane transport. As a validation of the applied method, we found that some of the identified key genes (CEACAM1, MCU, VDAC1, CYCS, C15ORF52, TMEM51, LARP1 and ERLIN2) have previously been reported by others as potential PDAC biomarkers. Using overall survival analyses, we found that several of the newly identified genes may serve as biomarkers to stratify PDAC patients into low- and high-risk groups.
Conclusions
Using this new systems biology approach, we identified several genes that appear to be critical to PDAC development. As such, they may represent potential diagnostic biomarkers as well as therapeutic targets with clinical utility.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Pancreatic ductal adenocarcinoma (PDAC) is a common and aggressive malignancy, with a 5-year survival rate of 4–6 % and a median survival time of less than 6 months. The poor prognosis and clinical outcome are due to the inability to detect PDAC in an early stage and to the poorly effective therapeutic options currently available. Therefore, a further understanding of the molecular mechanisms underlying PDAC development is necessary to identify new early diagnostic biomarkers and therapeutic targets [1]. In the past, it has e.g. been postulated that epithelial to mesenchymal transition (EMT) processes may yield therapeutic targets for PDAC and other solid tumours [2, 3]. KRAS, CDKN2A, TP53 and SMAD4 are the most frequently mutated genes in PDAC, whereas several other genes are mutated at relatively low frequencies [4]. Also, several expression-related biomarkers at both the RNA and protein level with diagnostic, prognostic and predictive value have been identified [5], including carbohydrate antigen 19–9 (CA 19–9) and carcinoembryonic antigen (CEA, currently named CEACAM5) [5], whereas other biomarkers have been excluded as PDAC prognosticators [6]. In attempts to unravel the molecular mechanisms underlying the pathogenesis of PDAC, microarray-based gene expression profiling studies have in the past been carried out. By doing so, Badea et al. identified 65 over-expressed genes in PDAC tumour epithelia and, among them, KRT7, LAMC2, SFN, PFKP, ANXA2, MAP4K4 and MBOAT2 were found to be inversely related to patient survival [7]. Donahue et al. identified 171 genes by which, on basis of their expression levels, two PDAC prognostic subgroups could be defined. In particular, high levels of PIK3R1 expression were found to be strongly associated with improved survival rates, whereas high levels of SRC expression were found to be associated with poorer survival rates [8]. Zhang et al. identified 2620 differentially expressed genes in PDACs, including 277 genes that were found to be associated with clinical outcome. Among them, DPEP1 reached the strongest association [9].
More recently, systems biology approaches have been applied to microarray-based gene expression datasets, aiming to extract new information besides a simple list of differentially expressed genes. Further in-depth analyses of the data generated by Zhang et al. [9] suggested important roles for the SP1 and TK1 genes in the progression of PDAC [10]. Additional approaches include the analysis of transcriptome maps to reveal genomic regions enriched in over/under-expressed genes [11]. Finally, although widely employed in the field of oncology, the analysis of microarray-based gene expression data by the weighted gene co-expression network analysis (WGCNA) systems biology approach [12] has, to our knowledge, so far not been applied to PDAC-derived data [13–15]. WGCNA allows a global interpretation of gene expression data by constructing gene networks based on similarities in expression profiles among samples. Highly co-expressed genes are connected in the network and, therefore, can be grouped into modules (i.e., highly connected network regions). Since these modules often consist of functionally related genes, different modules are involved in individual functions [16]. Within the modules, WGCNA also allows the identification of the most central and connected genes, that is the so called “hub” genes. When this approach is applied to two groups of samples, for example healthy and diseased samples, it is possible to obtain a network for each group and, therefore, to identify common modules and modules that differ between networks. These latter modules and their key genes may be involved in e.g. pathological processes and, thus, may have important clinical implications as potential diagnostic and prognostic biomarkers or therapeutic targets. As such, they warrant validation. In order to improve our understanding of the biological mechanisms underlying PDAC, we analysed existing PDAC gene expression datasets by applying an advanced network analysis strategy to detect key genes potentially involved in the pathogenesis of PDAC.
2 Materials and methods
2.1 Gene expression data and pre-processing
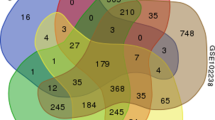
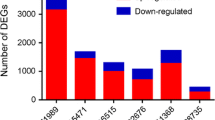
Raw CEL files of five microarray-based gene expression datasets (GSE15471 [7], GSE32676 [8], GSE28735 [9], GSE41368 [17] and GSE71989 (unpublished)) containing expression data from in total 105 normal pancreatic and 129 PDAC tissue samples were downloaded from the NCBI Gene Expression Omnibus (GEO) (Table 1). Data elaboration and figures presenting the results were obtained using the R 3.1.2 statistical environment (www.r-project.org) and Bioconductor (version 2.14) (www.bioconductor.org). Raw data from each microarray dataset were pre-processed identically with the R package affy using the Robust Multichip Average (RMA) function for background correction and normalization with the quantiles method [18]. Previous comparison studies have reported that RMA outperforms other normalization methods for analysis of microarray-based gene expression data [19]. In order to be able to merge the five microarray datasets, which were derived from two different platforms, they were made compatible. To this end, we first mapped the array probes to the respective Entrez Gene ID, a cross-platform common identifier, using the array annotation data hgu133plus2.db or hugene10sttranscriptcluster.db, depending on the platform used. Since the expression of a given gene is usually measured by multiple probes, we next summarized the expression values using the function collapseRows implemented in the R package WGCNA [20]. We selected the parameter “MaxMean” as method for collapsing rows, which chooses the probe with the highest mean value among samples, since this generally produces the most robust results [21]. In order to limit further analyses to genes common to all datasets, we finally created an overlapping gene set by selecting the rows with the Entrez Gene ID present in both platforms using the WGCNA function intersect, resulting in a total of 17,536 common genes. Since in the Affymetrix Human Gene 1.0 ST and Human Genome U133 Plus 2.0 Arrays 19,878 and 19,851 unique Entrez Gene IDs are represented, respectively, the percentages of common genes included in our analyses are considered to be very high (88.2 and 88.3 %). In order to remove cross-platform batch effects, we used the ComBat method [22], which is implemented in the R package sva and carries out another normalization step using an empirical Bayes approach. Among the available methods for batch correction, ComBat reaches the highest precision, accuracy and overall performance [23]. After this, we were able to merge all normalized microarray-based data into two global datasets, i.e., Normal and PDAC, that we used for the subsequent WGCNA analyses. The identification of outlier samples (to be excluded) was performed by hierarchical cluster analysis using the hclust function in WGCNA on each global dataset.
2.2 Dataset comparability analyses
In order to assess the comparability of the Normal and PDAC datasets, necessary for subsequent analyses, we correlated the gene expression levels and the overall connectivity, i.e., the co-expression level correlation, between the datasets. In case the correlations are positive, a higher correlation value indicates a higher comparability between the Normal and PDAC datasets. To this end, we used the function softConnectivity from package WGCNA, with the “randomly selected genes” parameter set at 5000, other parameters set as default, and the power parameter pre-calculated by the pickSoftThreshold function of WGCNA. This function provides the appropriate soft-thresholding power for network construction by calculating the scale-free topology fit index for several powers. If the scale-free topology fit index for the reference dataset reaches values above 0.8 for low powers (<30), as defined in [12], it means that the topology of the network is scale-free and, therefore, that there are no batch-effects.
2.3 Construction of weighted gene co-expression networks and identification of modules
Using standard WGCNA procedures [12], we created two weighted gene co-expression networks based on the Normal and PDAC expression data, respectively. Briefly, in each dataset we first created a matrix of adjacencies using the WGCNA function adjacency, by calculating Pearson correlations between each gene pair to determine concordances of gene expression, after which this matrix was transformed into a Topological Overlap Matrix (TOM) using the function TOMsimilarity. The resulting topological overlap is a biologically meaningful measure of gene similarity based on co-expression relationships between two genes [12]. Each TOM was used as input for hierarchical clustering analysis, which was performed with the function flashClust. Finally, in the resulting dendrograms we identified network modules present in the Normal dataset (used here as reference dataset) with the function cutreeHybrid from the R package dynamicTreeCut, using a relatively large minimum module size (minClusterSize = 30), and a medium sensitivity (deepSplit = 2), with other parameters set as default.
2.4 Module preservation analyses
The preservation levels of Normal network modules in the PDAC network were assessed first by plotting the two networks, thereby imposing modules from the Normal network onto the PDAC network. Secondly, by the function modulePreservation from the WGCNA package, a permutation test was carried out that assesses the preservation of the connectivity and density between each couple of modules, each belonging to the Normal and PDAC networks. This function provides a summary preservation Z-score for each module. The higher the Z-score, the higher the module preservation is, whereas values below 10 indicate a moderate to low preservation. Note that, since the grey module is the module of genes not assigned to any module, and the gold module contains random genes used for statistical aims by the modulePreservation function, they should have low Z-scores [24]. For the modulePreservation function we set some parameters (nPermutations = 30, maxGoldModuleSize = 100, maxModuleSize = 400), whereas others were left as default.
2.5 Detection of hub genes and their functional annotations
Module hub genes, which are highly connected intra-modular genes, have the highest Module Membership (MM) scores to the respective module [25]. The MM of each gene was calculated by the WGCNA function signedKME that correlates the expression profile of a gene with the Module Eigengene (ME) of a module, so it quantifies how close a gene is to a given module. ME is the representation of a module in one synthetic expression profile, obtained by the WGCA function moduleEigengenes. Next, we mapped these hub genes to the associated Gene Ontology (GO) terms and KEGG pathways using the DAVID tool (http://david.abcc.ncifcrf.gov/) [26]. Functional enrichment analysis of the identified hub genes consists of statistically highlighting the most over-represented (enriched) GO terms and KEGG pathways (p < 0.05), in order to facilitate the interpretation of the biological mechanisms related to a given gene list. Next, we used the Enrichr tool (http://amp.pharm.mssm.edu/Enrichr/) [27] to perform enrichment analyses of our hub gene lists. These enrichment analyses were carried out on predicted transcription factor binding sites using the “TRANSFAC_and_JASPAR_PWMs” section, on predicted miRNA binding sites using the “TargetScan_microRNA” section, and on chromosomal regions where these genes are located using the “Chromosome_Location” tool section. Only statistically significant results are reported (p < 0.02).
2.6 Survival analyses
Survival analyses were carried out using the SurvExpress tool [28], through which comparisons and validations of candidate cancer biomarkers using patient survival data present in the microarray datasets are made. The SurvExpress tool divides samples into two groups (high-risk and low-risk) through the median of the prognostic index obtained via a Cox regression model. After this, it generates risk hazard ratios (HR), relative confidence intervals (CI) and p-values. Our survival analyses were performed on an independent GEO dataset (GSE21501), containing gene expression and survival data derived from 132 PDAC patients.
3 Results
3.1 Pre-processing of the Normal and PDAC datasets
In order to increase the sample size and thus enhance the reliability of our analyses, five microarray datasets containing raw gene expression data of both normal and tumour pancreatic samples were pre-processed and merged into two global datasets, i.e., Normal and PDAC (Table 1; see Section 2). This data pre-processing step was needed since weighted gene co-expression network analysis (WGCNA) is sensitive to batch effects, i.e., to systematic and technical differences between different platforms and datasets and to the presence of outlier samples [16]. Besides removal of batch effects, we discarded a clear outlier, the normal sample GSM388111, from the GSE15471 dataset, as indicated by the dendrogram of sample clustering (Supplementary Fig. 1). As a result, the Normal dataset consisted of 104 normal pancreas samples and the PDAC dataset of 129 tumour samples. Subsequently, we assessed whether our datasets were comparable, since highly comparable datasets provide better chances of finding similarities and, therefore, also differences among them during subsequent analyses. In Supplementary Fig. 2 we show that our datasets are indeed comparable, since the correlation of gene expression between datasets was found to be 0.97 (p < 1e-200) and that of gene connectivity was found to be 0.43 (p < 1e-200). The latter parameter reflects the weighted co-expression level correlation, which indicates how strong a gene is connected to all other genes in the network. Next, we verified whether the networks to be constructed had a scale-free topology, as is required for WGCNA. A scale-free topology is a fundamental property of metabolic and signalling networks in which some nodes (here genes) are more connected than others, that is, some nodes are central (hub nodes) and others are peripheral. To this end, we applied the R function pickSoftThreshold and found that the scale-free topology fit index correctly reached values above 0.8 for a low power of 10 in the Normal dataset, here used as a control dataset (Supplementary Fig. 3). This result is also an indirect indication that we have efficiently removed batch-effects. In order to confirm this latter notion, we applied the function pickSoftThreshold to the Normal dataset not corrected for batch-effects, and found that it indeed failed to reach 0.8 at low powers (data not shown). As a further confirmation of efficient removal of batch-effects, the hclust function was applied to the non-corrected Normal and PDAC datasets, yielding several clusters of samples corresponding to the original datasets, whereas the hclust function applied to batch-effect corrected datasets yielded clustering dendrograms in which the samples were correctly sorted in no specific order (Supplementary Fig. 4).
3.2 Identification of gene co-expression networks and modules
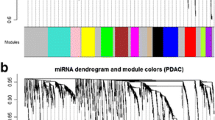
In the weighted gene co-expression network deriving from the Normal dataset, we identified via hierarchical clustering a total of 27 modules of different size in terms of gene number, which were labelled by different colours according to WGCNA package functions. In WGCNA analyses, a module is a group of genes with strongly shared co-expression relationships and, therefore, these genes are more connected than other genes in the network. Next, we set out to evaluate how well the characteristics of the modules that we identified in the reference network (Normal) are reproduced in the test network (PDAC). By doing so, we may be able to identify non-preserved modules, i.e., modules whose network properties are altered between the Normal and PDAC networks. These modules may, in turn, be related to the development of PDAC. To this end, we first plotted the two networks and imposed the modules from the Normal network onto the PDAC network (Supplementary Fig. 5). Since we found that the module-coloured labels still cluster together in the PDAC network, this indicates that the module preservation between the two datasets is good. In order to subsequently quantify the module preservation and to identify lowly preserved modules, we used the modulePreservation function from the WGCNA package, resulting in Z-scores for each module (Supplementary Table 1). Modules with a Z-score higher than 10 are highly preserved, i.e., they have similar network characteristics in the PDAC network, whereas the grey60 (Z-score = 9.6) and lightgreen (Z-score = 8.4) modules are lowly preserved between the datasets, and they may thus distinguish normal from pathological conditions. The Grey and Gold modules are special WGCNA modules that should not be considered here (see Section 2).
3.3 Identification of hub genes and their functional annotations
For each identified module, the hub nodes (that is hub genes) common to both networks were identified (Supplementary Table 2). Next, we focused on the grey60 and lightgreen modules, since these were found to be lowly preserved between the Normal and PDAC networks (see above) and, as such, can potentially distinguish PDAC from normal samples. In order to identify the central (hub) nodes that well represent these modules, we analysed these modules in further detail. In Table 2 we have listed the top 20 hub genes identified in the PDAC network. Since these hub genes were found exclusively in the PDAC network, they may play important roles in the pathogenesis of PDAC and, therefore, warrant further validation. In order to provide an interpretation of the biological mechanisms associated with these hub genes, we used the DAVID tool for a functional enrichment analysis. In Table 3 over-represented (enriched) Gene Ontology (GO) terms with p-values < 0.05 are listed. In the grey60 module, genes related to endoplasmic reticulum were found to be significantly over-represented among the hub genes. In the lightgreen module, several GO terms were found, some of which are related to cellular compartments, such as “cytoplasmic part”, “membrane” and “mitochondrion”, while other terms are related to biological processes such as “lipid metabolic process”, “transferase activity”, “hydrolase activity” and “transmembrane transport”.
In order to identify common elements involved in gene expression regulation, we next performed a gene enrichment analyses using the Enrichr tool. In Table 4 the transcription factors that are most over-represented in the grey60 and lightgreen modules are listed, of which RBPJ and FOXO3A were the most statistically significant ones. Regarding microRNAs, only miR-202, predicted to target the BCL7A and MANEAL genes, was found to be enriched in the grey60 module (p = 0.0112). Finally, we found that the chromosomal regions 7q21 and 3q28 were enriched in genes belonging to the lightgreen module, i.e., PON2 and SLC25A13 (p = 0.0012) and B3GNT5 (p = 0.0115), respectively. In the grey60 module, the chromosomal region 20q13, in which the STAU1 and ZNF334 genes are located, was found to be enriched (p = 0.0064).
3.4 Stratification of PDAC patients into high- and low-risk groups based on novel candidate biomarkers
Finally, we assessed whether the grey60 and lightgreen modules, i.e., the very lowly preserved modules between the Normal and PDAC datasets, are associated with the overall survival (OS) of PDAC patients [14]. For each module, we performed both single-gene and multi-gene (top 20 hub genes) survival analyses using the SurvExpress tool on an independent PDAC dataset (GSE21501). In Supplementary Fig. 6 the Kaplan-Meier survival plots for OS are shown using as input either the top 20 hub genes or each gene individually. By doing so, we found that both the grey60 and the lightgreen top 20 hub gene signatures successfully stratified patients into high- and low-risk groups, with OS times of the high-risk group patients being more than three-fold shorter than those of patients in the low-risk group (HR 3.83 [95 % CI 2.26–6.5] p = 6.474e-07 for the grey60 module and HR 3.41 [95 % CI 1.95–5.85] p = 8.813e-06 for the lightgreen module). In addition, we carried out a single-gene analysis in order to reveal which genes were most significantly associated with OS. We found that an increased expression of the CAMKMT (HR 1.76 [95 % CI 1.07–2.89] p = 0.02552), PON2 (HR 1.97 [95 % CI 1.19–3.27] p = 0.008798) and SLC25A13 (HR 1.65 [95 % CI 1.01–2.7] p = 0.04636) genes was associated with a poor OS. Conversely, increased expression levels of the TBC1D24 (HR 1.8 [95 % CI 1.1–2.95] p = 0.02005) and CASK (HR 1.81 [95 % CI 1.11–2.96] p = 0.01742) genes were found to correlate with a better OS.
4 Discussion
Here we identified, for the first time, candidate genetic biomarkers for pancreatic ductal adenocarcinoma (PDAC) by applying weighted gene co-expression network analysis (WGCNA), a systems biology approach, on expression data derived from five microarray-based datasets of PDAC and normal samples. We found that two modules of co-expressed genes differed significantly between the Normal and PDAC networks, suggesting a role in the pathogenesis of PDAC. Subsequently, we narrowed down the list of genes within these modules by identifying only the hub genes, i.e., the most PDAC-related genes according to WGCNA. Functional enrichment analysis of these genes revealed that they are related to either endoplasmic reticulum (ER), mitochondrion or membrane functions, exhibit transferase or hydrolase activities, and are related to biological processes such as lipid metabolism or transmembrane transport.
In particular, we identified the ERLIN2 gene , coding for an ER lipid raft associated protein, that was previously found to be over-expressed in PanIN-3, a pancreatic intraepithelial neoplasia-derived cell line [29]. Moreover, we identified TXNDC12, a gene that was previously found to inhibit ER stress-induced apoptosis of cancer cells [30]. It is well known that high proliferation rates of cancer cells can lead to ER stress, since these cells require increased protein folding activity in the ER [31]. Under stress conditions, increased levels of unfolded and misfolded proteins can induce an unfolded protein response (UPR) that can either mitigate ER damage or induce apoptosis [31]. However, cancer cells may develop effective escape strategies to overcome and prevent UPR-induced apoptosis [31]. Recent work has shown that these strategies are also used by PDAC cells and has yielded therapeutic molecules that induce apoptosis by acting on the ER stress response system [32, 33].
Regarding mitochondria, we identified the VDAC1 gene coding for a mitochondrial protein that belongs to a tethering protein complex that allows physical contact between the ER and mitochondria, i.e., the mitochondria-associated membrane (MAM) [34]. Many proteins regulating the MAM have been associated with cancer [34], of which the VDAC1 protein has very recently been found to be highly expressed in PDAC samples and has been suggested to serve as a reliable biomarker [35]. In addition, siRNA-mediated silencing of VDAC1 gene expression has been found to inhibit the growth of pancreas cancer-derived cell lines and xenograft models [36], highlighting a pivotal role of VDAC1 in PDAC development. Moreover, it has been reported that during apoptosis VDAC1 allows the release from mitochondria of the apoptogenic factor cytochrome c (CYCS) [34], which we identified as another key factor and which was previously found to be highly expressed in invasive ductal adenocarcinomas [37]. Additionally, since it was recently reported that PDAC cells rely on mitochondrial oxidative phosphorylation for survival [38], CYCS, which is a central element in this pathway, may play a critical role. Whereas VDAC1 is an outer mitochondrial membrane protein, the hub gene MCU that we also identified codes for a calcium uniporter localized at the mitochondrial inner membrane. MCU has been proposed to act as a tumour suppressor gene, along with other calcium-related genes, since it shows loss of heterozygosity (LOH) in PDACs [39]. Besides MCU, our hub gene list includes additional calcium-related genes, such as CASK, KCNMB3, PON2, SLC25A13 and ZDHHC3. It should be noted here that alteration of the calcium pathway plays an important role in the initiation and progression of PDAC via the Ca2+/calmodulin, PI3Kα/Akt and Raf/MEK/ERK pathways [40, 41]. Moreover, it has recently been reported that higher serum calcium levels are associated with a poor prognosis in PDAC patients [42]. Among the calcium-related genes, PON2 seems to be interesting since it was found to be over-expressed in tumour epithelia [7] and since the PON1 gene, which is another member of the paraoxonase gene family, has been found to serve as a candidate biomarker for pancreatic cancer [43]. Also the ZDHHC3 gene warrants further investigation since, like its related hub gene ZDHHC4, it codes for a palmitoyltransferase. Recently, integrin palmitoylation levels have been correlated with breast cancer invasiveness [44].
Besides alterations in the calcium pathway, it is well known that cancer cells can reprogram their metabolism to support rapid proliferation, including a metabolic switch to anaerobic glycolysis (Warburg effect) [45, 46]. Another relevant altered metabolic pathway is lipid synthesis, which strongly increases [47]. It has been found that inhibition of lipogenesis, decrease of the lipoprotein lipase and enhancement the biosynthesis of ceramide, a pro-apoptotic molecule, may inhibit cancer growth [47]. Moreover, it has been found that defects in ceramide metabolism may contribute to tumour cell survival and chemoresistance [48]. Among the key genes that we identified, ELOVL6 codes for a fatty acid elongase and NAPEPLD codes for a lipase, whereas CERS6 and B3GNT5 play a role in the ceramide synthesis pathway. In fact, CERS6 has been found to be strongly involved in several tumour types [49, 50]. Moreover, it was recently found that changes in lipid metabolism play a role especially in pancreatic carcinogenesis, since cholesterol uptake is increased in PDAC tissues and LDL receptor silencing reduces cell proliferation and enhances the chemotherapeutic efficacy in PDAC cells [51].
An additional key gene that should be further investigated is CEACAM1, a member of the carcinoembryonic antigen (CEA) gene family. In fact, it is considered as a PDAC biomarker, since it was found at higher expression levels in PDAC samples compared to noncancerous pancreatic samples [52], to be highly expressed in metastatic PDAC-derived cell lines [53] and to be present at high levels in serum of PDAC patients [54]. Moreover, its related protein CEA (also known as CEACAM5), is currently used as a conventional PDAC biomarker. Finally, some of the hub genes belonging to the grey60 module have been found by others to be highly expressed in PDAC, such as the C15ORF52 [55], TMEM51 [29] and LARP1 [37] genes.
We also sought to identify genes potentially related to PDAC by searching for gene expression regulatory elements that were enriched in our hub gene lists. Through this analysis, several common transcription factors were identified that have previously been related to PDAC development, including the well-studied SP1, HIF1A, FOXO3A and TP63 proteins (Supplementary Table 3). Through our miRNA enrichment analysis we identified miR-202, which has been proposed to serve as a therapeutic target since, when expressed at a low level, it has been found to induce apoptosis of PDAC cells [56]. Finally, the genomic regions 7q21-q22 and 20q13, that we identified through chromosomal enrichment analysis, are known to be frequently altered in PDAC and, thus, may play a causal role in its tumorigenesis [57, 58].
Moreover, patient overall survival analyses showed that the grey60 and lightgreen modules could effectively stratify high- and low-risk PDAC patients in an independent gene expression dataset. In particular, five genes (CAMKMT, CASK, PON2, SLC25A13 and TBC1D24) were found to be the major contributors to these gene signatures and, therefore, they may serve as prognostic biomarkers for PDAC.
Overall, the systems biology approach that we adopted in this study (WGCNA) allowed us to identify two modules of co-expressed genes related to pancreatic cancer, including some genes already known to serve as candidate PDAC biomarkers, which confirms the reliability of our results. Our results do, however, differ from those of three previously reported studies, performed on single PDAC microarray datasets [7–9]. This could be due to differences in input data and the adopted method. In fact our analysis has been performed on 5 PDAC datasets, including the three above mentioned ones, since large sample sizes are required for WGCNA. Moreover, WGCNA seeks hub genes in a co-expression network, in contrast to the cited studies in which primarily differentially expressed genes were identified. The previously reported studies yielded different results even among each other both in terms of differentially expressed genes and biomarker genes related to overall survival. These differences could be due to patient variables such as treatment, stage, gender, demographics, ethnicity, etc. Interestingly, several physiologic, hormonal and genomic differences between males and females may be relevant to gender-specific cancer susceptibility [59] or even to differential sensitivities of cells to anti-neoplastic agents [60]. It has been reported that significant differences in PDAC incidences do exist among different ethnic groups, correlating both with disparities in access to care and with genetic and other unknown risk factors [61, 62]. Unfortunately, we cannot assess these variables since this patient information is lacking in the microarray datasets used in this and in the three other cited studies [7–9]. Therefore, a further analysis and validation of the candidate PDAC biomarkers reported here is necessary, including those that have not yet been associated with PDAC.
References
C. Yu, M. Wang, Z. Li, J. Xiao, F. Peng, X. Guo, Y. Deng, J. Jiang, C. Sun, MicroRNA-138-5p regulates pancreatic cancer cell growth through targeting FOXC1. Cell. Oncol. 38, 173–181 (2015)
M. Beuran, I. Negoi, S. Paun, A.D. Ion, C. Bleotu, R.I. Negoi, S. Hostiuc, The epithelial to mesenchymal transition in pancreatic cancer: a systematic review. Pancreatology 15, 217–225 (2015)
F. Piva, M. Giulietti, M. Santoni, G. Occhipinti, M. Scarpelli, A. Lopez-Beltran, L. Cheng, G. Principato, R. Montironi, Epithelial to mesenchymal transition in renal cell carcinoma: implications for cancer therapy. Mol. Diagn. Ther. 20, 111–117 (2016)
E. Takai, S. Yachida, Genomic alterations in pancreatic cancer and their relevance to therapy. World J. Gastrointest. Oncol. 7, 250–258 (2015)
S. Majumder, S.T. Chari, D.A. Ahlquist, Molecular detection of pancreatic neoplasia: current status and future promise. World J. Gastroenterol. 21, 11387–11395 (2015)
K. Andrikou, M. Santoni, F. Piva, A. Bittoni, A. Lanese, C. Pellei, A. Conti, C. Loretelli, A. Mandolesi, M. Giulietti, M. Scarpelli, G. Principato, M. Falconi, S. Cascinu, Lgr5 expression, cancer stem cells and pancreatic cancer: results from biological and computational analyses. Future Oncol. 11, 1037–1045 (2015)
L. Badea, V. Herlea, S.O. Dima, T. Dumitrascu, I. Popescu, Combined gene expression analysis of whole-tissue and microdissected pancreatic ductal adenocarcinoma identifies genes specifically overexpressed in tumor epithelia. Hepatogastroenterology 55, 2016–2027 (2008)
T.R. Donahue, L.M. Tran, R. Hill, Y. Li, A. Kovochich, J.H. Calvopina, S.G. Patel, N. Wu, A. Hindoyan, J.J. Farrell, X. Li, D.W. Dawson, H. Wu, Integrative survival-based molecular profiling of human pancreatic cancer. Clin. Cancer Res. 18, 1352–1363 (2012)
G. Zhang, A. Schetter, P. He, N. Funamizu, J. Gaedcke, B.M. Ghadimi, T. Ried, R. Hassan, H.G. Yfantis, D.H. Lee, C. Lacy, A. Maitra, N. Hanna, H.R. Alexander, S.P. Hussain, DPEP1 inhibits tumor cell invasiveness, enhances chemosensitivity and predicts clinical outcome in pancreatic ductal adenocarcinoma. PLoS One 7, e31507 (2012)
B.S. Wang, Z. Liu, S.L. Sun, Y. Zhao, Identification of genes and candidate agents associated with pancreatic cancer. Tumour Biol. 35, 81–88 (2014)
L. Lenzi, F. Facchin, F. Piva, M. Giulietti, M.C. Pelleri, F. Frabetti, L. Vitale, R. Casadei, S. Canaider, S. Bortoluzzi, A. Coppe, G.A. Danieli, G. Principato, S. Ferrari, P. Strippoli, TRAM (Transcriptome Mapper): database-driven creation and analysis of transcriptome maps from multiple sources. BMC Genomics 12, 121 (2011)
B. Zhang, S. Horvath, A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol Biol. 4, Article17 (2005)
R. Liu, C.X. Guo, H.H. Zhou, Network-based approach to identify prognostic biomarkers for estrogen receptor-positive breast cancer treatment with tamoxifen. Cancer Biol. Ther. 16, 317–324 (2015)
C. Clarke, S.F. Madden, P. Doolan, S.T. Aherne, H. Joyce, L. O’Driscoll, W.M. Gallagher, B.T. Hennessy, M. Moriarty, J. Crown, S. Kennedy, M. Clynes, Correlating transcriptional networks to breast cancer survival: a large-scale coexpression analysis. Carcinogenesis 34, 2300–2308 (2013)
Y.S. Lee, S.G. Hwang, J.K. Kim, T.H. Park, Y.R. Kim, H.S. Myeong, J.D. Choi, K. Kwon, C.S. Jang, Y.T. Ro, Y.H. Noh, S.Y. Kim, Identification of novel therapeutic target genes in acquired lapatinib-resistant breast cancer by integrative meta-analysis. Tumour Biol. 37, 2285–2297 (2015)
M.C. Oldham, G. Konopka, K. Iwamoto, P. Langfelder, T. Kato, S. Horvath, D.H. Geschwind, Functional organization of the transcriptome in human brain. Nat. Neurosci. 11, 1271–1282 (2008)
A.E. Frampton, L. Castellano, T. Colombo, E. Giovannetti, J. Krell, J. Jacob, L. Pellegrino, L. Roca-Alonso, N. Funel, T.M. Gall, A. De Giorgio, F.G. Pinho, V. Fulci, D.J. Britton, R. Ahmad, N.A. Habib, R.C. Coombes, V. Harding, T. Knosel, J. Stebbing, L.R. Jiao, MicroRNAs cooperatively inhibit a network of tumor suppressor genes to promote pancreatic tumor growth and progression. Gastroenterology 146, 268–277 (2014). e218
R.A. Irizarry, B. Hobbs, F. Collin, Y.D. Beazer-Barclay, K.J. Antonellis, U. Scherf, T.P. Speed, Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003)
B.M. Bolstad, R.A. Irizarry, M. Astrand, T.P. Speed, A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003)
P. Langfelder, S. Horvath, WGCNA: an R package for weighted correlation network analysis. BMC Bioinf. 9, 559 (2008)
J.A. Miller, C. Cai, P. Langfelder, D.H. Geschwind, S.M. Kurian, D.R. Salomon, S. Horvath, Strategies for aggregating gene expression data: the collapseRows R function. BMC Bioinf. 12, 322 (2011)
W.E. Johnson, C. Li, A. Rabinovic, Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007)
C. Chen, K. Grennan, J. Badner, D. Zhang, E. Gershon, L. Jin, C. Liu, Removing batch effects in analysis of expression microarray data: an evaluation of six batch adjustment methods. PLoS One 6, e17238 (2011)
P. Langfelder, R. Luo, M.C. Oldham, S. Horvath, Is my network module preserved and reproducible? PLoS Comput. Biol. 7, e1001057 (2011)
S. Horvath, J. Dong, Geometric interpretation of gene coexpression network analysis. PLoS Comput. Biol. 4, e1000117 (2008)
D.W. Huang, B.T. Sherman, Q. Tan, J. Kir, D. Liu, D. Bryant, Y. Guo, R. Stephens, M.W. Baseler, H.C. Lane, R.A. Lempicki, DAVID bioinformatics resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 35, W169–175 (2007)
E.Y. Chen, C.M. Tan, Y. Kou, Q. Duan, Z. Wang, G.V. Meirelles, N.R. Clark, A. Ma’ayan, Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinf. 14, 128 (2013)
R. Aguirre-Gamboa, H. Gomez-Rueda, E. Martinez-Ledesma, A. Martinez-Torteya, R. Chacolla-Huaringa, A. Rodriguez-Barrientos, J.G. Tamez-Pena, V. Trevino, SurvExpress: an online biomarker validation tool and database for cancer gene expression data using survival analysis. PLoS One 8, e74250 (2013)
M. Buchholz, M. Braun, A. Heidenblut, H.A. Kestler, G. Kloppel, W. Schmiegel, S.A. Hahn, J. Luttges, T.M. Gress, Transcriptome analysis of microdissected pancreatic intraepithelial neoplastic lesions. Oncogene 24, 6626–6636 (2005)
W. Jeong, D.Y. Lee, S. Park, S.G. Rhee, ERp16, an endoplasmic reticulum-resident thiol-disulfide oxidoreductase: biochemical properties and role in apoptosis induced by endoplasmic reticulum stress. J. Biol. Chem. 283, 25557–25566 (2008)
M. Wang, R.J. Kaufman, The impact of the endoplasmic reticulum protein-folding environment on cancer development. Nat. Rev. Cancer 14, 581–597 (2014)
H.W. Chiu, S.W. Lin, L.C. Lin, Y.H. Hsu, Y.F. Lin, S.Y. Ho, Y.H. Wu, Y.J. Wang, Synergistic antitumor effects of radiation and proteasome inhibitor treatment in pancreatic cancer through the induction of autophagy and the downregulation of TRAF6. Cancer Lett. 365, 229–239 (2015)
N. Mujumdar, S. Banerjee, Z. Chen, V. Sangwan, R. Chugh, V. Dudeja, M. Yamamoto, S.M. Vickers, A.K. Saluja, Triptolide activates unfolded protein response leading to chronic ER stress in pancreatic cancer cells. Am. J. Physiol. Gastrointest. Liver Physiol. 306, G1011–1020 (2014)
A. Raturi, T. Simmen, Where the endoplasmic reticulum and the mitochondrion tie the knot: the mitochondria-associated membrane (MAM). Biochim. Biophys. Acta 1833, 213–224 (2013)
K.K. Kuo, C.J. Kuo, C.Y. Chiu, S.S. Liang, C.H. Huang, S.W. Chi, K.B. Tsai, C.Y. Chen, E. Hsi, K.H. Cheng, S.H. Chiou, Quantitative proteomic analysis of differentially expressed protein profiles involved in pancreatic ductal adenocarcinoma. Pancreas 45, 71–83 (2016)
T. Arif, L. Vasilkovsky, Y. Refaely, A. Konson, V. Shoshan-Barmatz, Silencing VDAC1 expression by siRNA inhibits cancer cell proliferation and tumor growth in vivo. Mol. Ther. Nucleic Acids 3, e159 (2014)
T. Nakamura, Y. Furukawa, H. Nakagawa, T. Tsunoda, H. Ohigashi, K. Murata, O. Ishikawa, K. Ohgaki, N. Kashimura, M. Miyamoto, S. Hirano, S. Kondo, H. Katoh, Y. Nakamura, T. Katagiri, Genome-wide cDNA microarray analysis of gene expression profiles in pancreatic cancers using populations of tumor cells and normal ductal epithelial cells selected for purity by laser microdissection. Oncogene 23, 2385–2400 (2004)
A. Viale, P. Pettazzoni, C.A. Lyssiotis, H. Ying, N. Sanchez, M. Marchesini, A. Carugo, T. Green, S. Seth, V. Giuliani, M. Kost-Alimova, F. Muller, S. Colla, L. Nezi, G. Genovese, A.K. Deem, A. Kapoor, W. Yao, E. Brunetto, Y. Kang, M. Yuan, J.M. Asara, Y.A. Wang, T.P. Heffernan, A.C. Kimmelman, H. Wang, J.B. Fleming, L.C. Cantley, R.A. DePinho, G.F. Draetta, Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature 514, 628–632 (2014)
J. Long, Z.B. Zhang, Z. Liu, Y.H. Xu, C.L. Ge, Loss of heterozygosity at the calcium regulation gene locus on chromosome 10q in human pancreatic cancer. Asian Pac. J. Cancer Prev. 16, 2489–2493 (2015)
G. Dawelbait, C. Winter, Y. Zhang, C. Pilarsky, R. Grutzmann, J.C. Heinrich, M. Schroeder, Structural templates predict novel protein interactions and targets from pancreas tumour gene expression data. Bioinformatics 23, i115–124 (2007)
R. Nussinov, S. Muratcioglu, C.J. Tsai, H. Jang, A. Gursoy, O. Keskin, The key role of calmodulin in KRAS-Driven adenocarcinomas. Mol. Cancer Res. 13, 1265–1273 (2015)
Q. Dong, Y. Zhang, X.H. Yang, W. Jing, L.Q. Zheng, Y.P. Liu, X.J. Qu, Z. Li, Serum calcium level used as a prognostic predictor in patients with resectable pancreatic ductal adenocarcinoma. Clin. Res. Hepatol. Gastroenterol. 38, 639–648 (2014)
J. Kodydkova, L. Vavrova, B. Stankova, J. Macasek, T. Krechler, A. Zak, Antioxidant status and oxidative stress markers in pancreatic cancer and chronic pancreatitis. Pancreas 42, 614–621 (2013)
D.T. Coleman, Y.H. Soung, Y.J. Surh, J.A. Cardelli, J. Chung, Curcumin prevents palmitoylation of integrin beta4 in breast cancer cells. PLoS One 10, e0125399 (2015)
H.R. Oh, C.H. An, N.J. Yoo, S.H. Lee, Somatic mutations of amino acid metabolism-related genes in gastric and colorectal cancers and their regional heterogeneity--a short report. Cell. Oncol. 37, 455–461 (2014)
K. Goetze, C.G. Fabian, A. Siebers, L. Binz, D. Faber, S. Indraccolo, G. Nardo, U.G. Sattler, W. Mueller-Klieser, Manipulation of tumor metabolism for therapeutic approaches: ovarian cancer-derived cell lines as a model system. Cell. Oncol. 38, 377–385 (2015)
J. Swierczynski, A. Hebanowska, T. Sledzinski, Role of abnormal lipid metabolism in development, progression, diagnosis and therapy of pancreatic cancer. World J. Gastroenterol. 20, 2279–2303 (2014)
S.A. Morad, M.C. Cabot, Ceramide-orchestrated signalling in cancer cells. Nat. Rev. Cancer 13, 51–65 (2013)
S.A. Jensen, A.E. Calvert, G. Volpert, F.M. Kouri, L.A. Hurley, J.P. Luciano, Y. Wu, A. Chalastanis, A.H. Futerman, A.H. Stegh, Bcl2L13 is a ceramide synthase inhibitor in glioblastoma. Proc. Natl. Acad. Sci. U. S. A. 111, 5682–5687 (2014)
M. Suzuki, K. Cao, S. Kato, Y. Komizu, N. Mizutani, K. Tanaka, C. Arima, M.C. Tai, K. Yanagisawa, N. Togawa, T. Shiraishi, N. Usami, T. Taniguchi, T. Fukui, K. Yokoi, K. Wakahara, Y. Hasegawa, Y. Mizutani, Y. Igarashi, J. Inokuchi, S. Iwaki, S. Fujii, A. Satou, Y. Matsumoto, R. Ueoka, K. Tamiya-Koizumi, T. Murate, M. Nakamura, M. Kyogashima, T. Takahashi, Targeting ceramide synthase 6-dependent metastasis-prone phenotype in lung cancer cells. J. Clin. Invest. 126, 254–265 (2016)
F. Guillaumond, G. Bidaut, M. Ouaissi, S. Servais, V. Gouirand, O. Olivares, S. Lac, L. Borge, J. Roques, O. Gayet, M. Pinault, C. Guimaraes, J. Nigri, C. Loncle, M.N. Lavaut, S. Garcia, A. Tailleux, B. Staels, E. Calvo, R. Tomasini, J.L. Iovanna, S. Vasseur, Cholesterol uptake disruption, in association with chemotherapy, is a promising combined metabolic therapy for pancreatic adenocarcinoma. Proc. Natl. Acad. Sci. U. S. A. 112, 2473–2478 (2015)
D.M. Simeone, B. Ji, M. Banerjee, T. Arumugam, D. Li, M.A. Anderson, A.M. Bamberger, J. Greenson, R.E. Brand, V. Ramachandran, C.D. Logsdon, CEACAM1, a novel serum biomarker for pancreatic cancer. Pancreas 34, 436–443 (2007)
T. Kalinina, C. Gungor, S. Thieltges, M. Moller-Krull, E.M. Penas, D. Wicklein, T. Streichert, U. Schumacher, V. Kalinin, R. Simon, B. Otto, J. Dierlamm, H. Schwarzenbach, K.E. Effenberger, M. Bockhorn, J.R. Izbicki, E.F. Yekebas, Establishment and characterization of a new human pancreatic adenocarcinoma cell line with high metastatic potential to the lung. BMC Cancer 10, 295 (2010)
F. Gebauer, D. Wicklein, J. Horst, P. Sundermann, H. Maar, T. Streichert, M. Tachezy, J.R. Izbicki, M. Bockhorn, U. Schumacher, Carcinoembryonic antigen-related cell adhesion molecules (CEACAM) 1, 5 and 6 as biomarkers in pancreatic cancer. PLoS One 9, e113023 (2014)
D. Cao, S.R. Hustinx, G. Sui, P. Bala, N. Sato, S. Martin, A. Maitra, K.M. Murphy, J.L. Cameron, C.J. Yeo, S.E. Kern, M. Goggins, A. Pandey, R.H. Hruban, Identification of novel highly expressed genes in pancreatic ductal adenocarcinomas through a bioinformatics analysis of expressed sequence tags. Cancer Biol. Ther. 3, 1081–1089 (2004). discussion 1090–1081
L. Farhana, M.I. Dawson, J.A. Fontana, Down regulation of miR-202 modulates Mxd1 and Sin3A repressor complexes to induce apoptosis of pancreatic cancer cells. Cancer Biol. Ther. 16, 115–124 (2015)
J.W. Liang, Z.Z. Shi, T.Y. Shen, X. Che, Z. Wang, S.S. Shi, X. Xu, Y. Cai, P. Zhao, C.F. Wang, Z.X. Zhou, M.R. Wang, Identification of genomic alterations in pancreatic cancer using array-based comparative genomic hybridization. PLoS One 9, e114616 (2014)
Y. Tabach, I. Kogan-Sakin, Y. Buganim, H. Solomon, N. Goldfinger, R. Hovland, X.S. Ke, A.M. Oyan, K.H. Kalland, V. Rotter, E. Domany, Amplification of the 20q chromosomal arm occurs early in tumorigenic transformation and may initiate cancer. PLoS One 6, e14632 (2011)
M.T. Dorak, E. Karpuzoglu, Gender differences in cancer susceptibility: an inadequately addressed issue. Front. Genet. 3, 268 (2012)
L.M. Nunes, E. Robles-Escajeda, Y. Santiago-Vazquez, N.M. Ortega, C. Lema, A. Muro, G. Almodovar, U. Das, S. Das, J.R. Dimmock, R.J. Aguilera, A. Varela-Ramirez, The gender of cell lines matters when screening for novel anti-cancer drugs. AAPS J. 16, 872–874 (2014)
R. Ali, I. Barnes, B.J. Cairns, A.E. Finlayson, N. Bhala, M. Mallath, V. Beral, Incidence of gastrointestinal cancers by ethnic group in England, 2001–2007. Gut 62, 1692–1703 (2013)
V. Singal, A.K. Singal, Y.F. Kuo, Racial disparities in treatment for pancreatic cancer and impact on survival: a population-based analysis. J. Cancer Res. Clin. Oncol. 138, 715–722 (2012)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animal rights and informed consent
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1154 kb)
Rights and permissions
About this article
Cite this article
Giulietti, M., Occhipinti, G., Principato, G. et al. Weighted gene co-expression network analysis reveals key genes involved in pancreatic ductal adenocarcinoma development. Cell Oncol. 39, 379–388 (2016). https://doi.org/10.1007/s13402-016-0283-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-016-0283-7