Abstract
Background
Human nasopharyngeal carcinoma (NPC) is a malignant type of cancer with an increasing incidence. As yet, however, molecular biomarkers with a strong diagnostic impact and a major therapeutic promise have remained elusive. Here, we identified the esophageal carcinoma related gene 4 (ECRG4) as a novel candidate tumor suppressor gene and a promising therapeutic target for NPC.
Methods
RT-PCR, Western blotting, methylation-specific PCR and bisulfite sequencing were performed to assess the expression and methylation status of the ECRG4 gene in primary NPC samples, NPC-derived cell lines and patient-derived peripheral blood samples. The NPC-derived cell line CNE1 was selected for treatment with a methylation inhibitor to restore ECRG4 expression. In addition, cell proliferation, invasion and colony formation assays were performed to assess the inhibitory effects of exogenous ECRG4 expression in CNE1 cells.
Results
Down-regulated ECRG4 expression was found to occur in 82.5 % (33/40) of the primary NPC biopsies tested. This down-regulation was significantly correlated with its tumor-specific promoter methylation status (72.5 %, 29/40) and was also observed in the matching peripheral blood samples from the NPC patients (57.5 %, 23/40). Pharmacologic demethylation through 5-aza-dC treatment led to gene reactivation in ECRG4 methylated and silenced NPC cell lines. Moreover, exogenous expression of ECRG4 in the CNE1 cell line strongly inhibited its growth and invasive capacities, as well as its enhanced chemosensitivity to cisplatin through autophagy induction.
Conclusion
Our data suggest that methylation-mediated suppression of the ECRG4 gene occurs frequently in NPC and that restoration of its expression may have therapeutic benefits.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Nasopharyngeal carcinoma (NPC) is a common malignancy, in particular in Southeastern China, and undifferentiated carcinoma is the main histological sub-type [1]. Despite improvements in both surgical techniques and radio- and chemotherapy regimens, the prognosis of this malignancy is still poor [2–4]. Increasing evidence has revealed that molecular subtyping of cancers is crucial for a better understanding of their clinical behavior and the selection of suitable targets for therapy [5]. However, the diagnostic and therapeutic implications of various molecular biomarkers that have been identified in other cancers (e.g. those of the breast and esophagus) have remained unclear in NPC [6, 7].
Hence, it is urgent to find reliable molecular biomarkers that are measurable, not only in terms of their altered level in tumors versus normal tissues, but also for their contribution to tumor initiation and progression. These genes and/or their products can be used for diagnosis, outcome prediction and therapy surveillance or, even more importantly, as direct targets for therapy [8]. Due to the multi-step nature of cancer, the better candidates should be central to a variety of oncogenic networks that are activated during tumor development.
The esophageal carcinoma related gene 4 (ECRG4), also called C2ORF40, is located on chromosome 2q12.2 and was originally identified as a putative tumor suppressor gene (TSG) in esophageal carcinoma [9, 10]. ECRG4 acts as an integrator and indispensable modulator of major growth factor signaling pathways, such as the NF-κB signaling pathway, and can mediate downstream signaling events that are involved in cell cycle progression, cell migration and senescence, progenitor cell survival and differentiation and inflammatory events at multiple mechanistic levels [10–14]. Although its exact impact on oncogenesis remains to be elucidated, ECRG4 has been reported to act as a candidate TSG in several cancers [10, 12, 15–19]. Encouraging data show that ECRG4 may not only contribute to prognosis prediction, but may also offer new tailor-made therapeutic options [11, 17–19]. Yet, despite these promises, the role of ECRG4 in NPC has remained largely unknown. Here, we investigated the promoter hypermethylation-mediated inactivation of the ECRG4 gene and assessed its potential for clinical application in NPC. Additionally, the anti-tumor effects and therapeutic implications of ECRG4 expression restoration are addressed.
2 Materials and methods
2.1 Cell lines and tissue specimens
Seven NPC cell lines used in this study, including HNE1, HONE1, CNE1, SUNE1, CNE2, 6-10B and C666-1, were maintained and treated with the demethylating agent 5-aza-2′-deoxycytidine (5-aza-dC) (Sigma, St. Louis, MO), the chemotherapeutic drug cisplatin (Sigma) and the autophagy inhibitor 3-methyladenine (3-MA) (Sigma) as described before [20, 21]. With informed consent, 40 primary tumor biopsies and their matched peripheral blood samples from pathologically verified NPC patients were obtained at the Affiliated Hospital of Bethune Military Medical College. The tumor samples were isolated, immediately frozen in liquid nitrogen and, subsequently, stored at −80 °C until use. Peripheral venous blood samples were collected in EDTA-containing tubes and immediately centrifuged at 2500 × g for 15 min to prepare plasma. Additionally, 10 chronic nasopharyngitis tissues and 10 peripheral blood samples from healthy volunteers were included as controls.
2.2 Western blot and RT-PCR analyses
Cells were harvested after the indicated treatments and analyzed by Western blotting, as described previously [4]. Briefly, proteins (50 μg in RIPA lysis buffer) were separated by SDS-PAGE, transferred to PVDF membranes (Millipore, Billerica, MA) and then subjected to Western blotting using the indicated antibodies, i.e., rabbit polyclonal antibodies directed against ECRG4, cleaved caspase-3 and LC3 (Santa Cruz Biotechnology, Santa Cruz, CA), and a mouse monoclonal antibody directed against PARP (Cell Signaling, Beverly, MA). The blots were re-probed with an anti-β-actin monoclonal antibody (Abcam, Cambridge, MA) to confirm equal loading of the different samples. For the detection of secreted ECRG4 protein, cell culture media were replaced with fresh media without serum. Twenty-four hours later, the culture media were collected and a complete protease inhibitor cocktail (Roche, Mannheim, Germany) was added. Cellular debris was removed by centrifugation at 700 × g and the medium was concentrated 50-fold using Amicon-15 centrifugal filter units (Millipore). Concentrated media were used directly or stored at 4 °C. For semi-quantitative RT-PCR analysis, total RNA was extracted from each sample using a TRIzol reagent (Invitrogen, Carlsbad, CA), and RT-PCR was performed as described before [4]. Gene-specific primers used for the amplification of ECRG4 and β-actin are listed in Table 1. Quantification of intensities was performed using the Bio-Rad Quantity One quantification software package.
2.3 Bisulfite modification and promoter methylation analysis
Genomic DNAs from cell lines, primary tumors and plasma samples were extracted using a TIANamp Extraction Kit (Tiangen, Beijing, China). Bisulfite modification of DNA, methylation-specific PCR (MSP) and bisulfite genome sequencing (BGS) were carried out as described before [4, 20, 21]. The primer sequences used for MSP and BGS are listed in Table 1.
2.4 Vector construction and transfection
An ECRG4 expression vector (pcDNA3.1-ECRG4) was constructed by sub-cloning the full-length wild-type ECRG4 coding sequence into pcDNA3.1(+), and confirmed by sequencing. Cell transfections were conducted using Lipofectamine 2000 reagent (Invitrogen) following the manufacturer’s protocol. Stable ECRG4 transfectants were generated under G418 (Gibco, Paisley, UK) selection as described before [4].
2.5 Colony formation assays
For a colony formation assay using a monolayer culture, 2 × 105 cells per well were seeded in a 12-well plate and transfected with the ECRG4 expression plasmid or the empty control vector. Cells were collected and seeded in 6-well plates 48 h post-transfection, and selected for 2 weeks in G418 (800 mg/l). Surviving colonies (≥ 50 cells/colony) were counted after staining with crystal violet reagent (Sigma). For a colony formation assay using a soft agar culture, stably transfected cells were suspended in RPMI-1640 medium containing 0.35 % agar, 10 % fetal bovine serum (FBS) and 800 mg/l G418 and layered on RPMI-1640 medium containing 0.5 % agar, 10 % FBS and G418 in a 6-well plate. Colonies were counted and photographed. All the experiments were performed in triplicate and repeated three times.
2.6 Cell proliferation and invasion assays
The cell proliferative and invasive capacities were determined using a 3-(4,5-dimethylthiazol-2-yl)-2 5-diphenyl-2H-tetrazolium bromide (MTT) colorimetric assay and a Matrigel invasion chamber assay, respectively, according to standard methods described before [4]. Each experiment was performed three times in replicates of six wells.
2.7 Fluorescence microscopy
Cells mounted on glass slides were permeabilized with PBS containing 0.1 % Triton X-100 and 0.1 M glycine for 15 min, blocked with 10 % BSA in PBS for 30 min at room temperature and then incubated with a rabbit anti-LC3 antibody (Santa Cruz), followed by a FITC-labeled secondary antibody directed against rabbit IgG. Finally, the cells were washed twice with PBS and images were taken under a fluorescence microscope.
2.8 Cell apoptosis analysis
Cells were collected and stained with annexin V/PI using an Annexin V-FITC Apoptosis Detection Kit (Tiangen, Beijing, China), according to the manufacturer’s instructions. Cells were then analyzed by flow cytometry (Beckman Coulter, Cytomics FC 500, CA) and apoptosis profiles were calculated using a ModFit LT software package. Each experiment was repeated three times.
2.9 Statistical analysis
The results are expressed as values of means ± standard deviation (SD). Statistical analysis was carried out with Student’s t-test. P values < 0.05 were considered as statistically significant.
3 Results
3.1 ECRG4 expression down-regulation occurs frequently in NPC
Tumor suppressor genes (TSGs) are often subject to down-regulation during tumorigenesis. Therefore, we first set out to assess ECRG4 gene expression levels in 40 primary NPC biopsies and 10 chronic nasopharyngitis tissues using semi-quantitative RT-PCR. In Fig. 1a representative RT-PCR results are shown. We found that the ECRG4 gene was highly expressed in all 10 chronic nasopharyngitis samples. In contrast, its transcript level was dramatically reduced or totally undetectable in 82.5 % (33/40) of the primary NPC biopsies. Further quantitative analyses showed that the ECRG4 transcript levels in tumor specimens were significantly lower than those in the non-cancerous tissues (p < 0.001). We also observed down-regulated ECRG4 expression in 85.7 % (6/7) of NPC-derived cell lines, i.e., a reduced expression in SUNE1, HNE1, 6-10B and CNE2, and an undetectable expression in CNE1 and HONE1 (Fig. 1b and c). These results clearly indicate that ECRG4 inactivation is a common and tumor-specific event in NPC.
The expression of ECRG4 is downregulated in NPC. Total RNA isolated from primary NPC tissues (a) and NPC cancer cell lines (b) was subjected to RT-PCR using ECRG4 specific primers. Chronic nasopharyngitis (CNPG) tissues were used as positive controls and β-actin was used as an internal loading control. ***P < 0.001. c ECRG4 protein levels determined by Western blot analysis in NPC cell lines
3.2 ECRG4 promoter methylation reversely correlates with its expression
TSGs may be inactivated through epigenetic mechanisms, mainly aberrant methylation of promoter CpG Islands (CGIs) that are generally unmethylated in normal tissues [4]. A typical 528 bp CGI was predicted upstream of ECRG4 exon 1 using a Methprimer CpG Island Prediction tool (http://www.urogene.org/methprimer/), by applying the following criteria: GC content > 50 %, Obs CpG/Exp CpG > 0.60, and length > 200 bp (Fig. 2a), indicating that ECRG4 is most likely vulnerable to methylation-mediated silencing. We next analyzed the methylation status of the ECRG4 gene promoter using MSP and, by doing so, observed this anomaly in 72.5 % (29/40) of the primary NPC biopsies tested. In contrast, virtually no methylation was detected in the chronic nasopharyngitis (CNPG) tissues tested (Fig. 2b). Hypermethylation was only observed in the tumor samples exhibiting down-regulated ECRG4 expression. Interestingly, ECRG4 gene transcription was also found to be down-regulated in four NPC biopsies devoid of methylation, which may be due to other mechanisms of transcriptional repression. Our results also revealed that the NPC-derived cell lines in which ECRG4 expression was reduced or undetectable showed concomitant promoter hypermethylation (Fig. 2c).
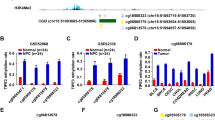
Frequent promoter methylation of ECRG4 in NPC. a Schematic structure of the ECRG4 CGI. The exon 1, MSP and BGS regions are marked. Each short vertical line represents one CpG site. b Representative ECRG4 MSP results in primary NPC tissues. M, methylated; U, unmethylated. c MSP analysis of ECRG4 methylation in NPC cell lines. d High-resolution methylation analysis of the ECRG4 promoter by BGS. A region spanning the core promoter with 23 CpG sites was analyzed. Four or five colonies of cloned BGS-PCR products from each bisulfite-treated DNA sample were sequenced. Each clone is shown as an individual row, representing a single allele of the promoter CpG Islands (CGI) analyzed. One square indicates one CpG site. Dark filled or open squares represent methylated or unmethylated CpG sites, respectively. e Representative MSP result of the ECRG4 gene in peripheral blood samples from NPC patients and normal healthy individuals
We further examined in more detail the ECRG4 gene promoter methylation status through high resolution bisulfite genome sequencing (BGS) analysis of 23 CpG sites within the CGI, including several CpGs analyzed by MSP (Fig. 2a). The presence of cytosines in a CpG context after bisulfite conversion indicates that the genomic DNA of the sample analyzed is methylated. Overall, the BGS results were consistent with our MSP results (Fig. 2d). A heavily methylated CGI was detected in two NPC biopsies (#7 and #17) with undetectable ECRG4 expression levels. In contrast, the chronic nasopharyngitis tissues, exhibiting abundant ECRG4 expression, were almost totally free of methylation. Similarly, we observed a step-wise decrease of methylated CpG distribution in the cell lines in which ECRG4 expression was silenced (CNE1), reduced (HNE1) or unaffected (C666-1). These observations thus suggest a strong correlation between ECRG4 gene promoter hypermethylation and expression silencing.
3.3 ECRG4 promoter methylation in peripheral blood samples
Encouraged by the above data suggesting tumor-specific ECRG4 hypermethylation in primary NPC biopsies, we next explored the feasibility of detecting this anomaly in 40 paired peripheral blood samples from NPC patients and normal control samples from 10 healthy volunteers. Representative MSP results are illustrated in Fig. 2e. We found that 57.5 % (23/40) of the matched peripheral blood samples showed ECRG4 promoter hypermethylation. Of note, this anomaly was only observed in blood samples from patients exhibiting ECRG4 promoter methylation in the primary NPC biopsies. No methylation was detected in any of the normal control samples from healthy individuals.
3.4 Pharmacologic demethylation reactivates ECRG4 expression
To determine whether CGI methylation directly mediates ECRG4 silencing and whether ECRG4 may serve as a molecular target for epigenetic therapy in NPC, we treated NPC cancer cell lines with the DNA methyltransferase inhibitor 5-Aza-dC, and assessed their ECRG4 expression status using RT-PCR and Western blotting. As shown in Fig. 3a and b, exposure to 5-Aza-dC resulted in a marked increase in ECRG4 mRNA and protein expression levels in CNE1 cells, whereas those in HNE1 cells were less affected. C666-1 cells were included as unaffected control (Fig. 3a). Concomitantly, 5-Aza-dC treatment led to relative increases in unmethylated ECRG4 alleles as determined by MSP (Fig. 3a) and BGS (Fig. 3c) analyses. These data confirm that promoter hypermethylation plays an essential role in ECRG4 transcription silencing in NPC cells.
ECRG4 expression restoration through pharmacologic demethylation. a Treatment with 5-aza-dC restored the expression of ECRG4 in CNE1 and HNE1 cells as determined by RT-PCR and MSP analyses, respectively. M, methylated; U, unmethylated. b Western blot analysis confirming the restored ECRG4 expression at the protein level after 5-aza-dC treatment. c High-resolution methylation mapping of CpG sites by BGS confirming the pharmacologic demethylation of ECRG4
3.5 Exogenous ECRG4 expression inhibits tumor cell growth and invasion
The observation that ECRG4 is frequently silenced by promoter methylation in primary NPC tumor samples and NPC-derived cell lines, but not in normal nasopharyngeal epithelial tissues, indicates that ECRG4 most likely acts a tumor suppressor. To further test this hypothesis, we transfected an ECRG4 expression construct into CNE1 cells, in which the ECRG4 gene was found (see above) to be completely silenced by promoter hypermethylation. Subsequently, colony forming efficiencies were evaluated using a monolayer and a soft-agar culture assay, respectively. By doing so, we found that exogenous ECRG4 expression dramatically reduced the colony formation efficiencies of CNE1 cells both in the monolayer (Fig. 4a) and soft-agar (Fig. 4b) assays. These findings were further substantiated using a cell proliferation assay. To this end, two stable ECRG4 transfectants (ECRG4c1 and ECRG4c2) were generated and tested for endogenous expression (Fig. 4c). Of note, we also found that secreted ECRG4 protein was detectable in the cell culture medium (Fig. 4d). We found that CNE1 cells exogenously expressing ECRG4 grew significantly slower than the empty vector-transfected cells or the parental cells (Fig. 4e). Next, we examined the effect of ECRG4 expression on the invasion-suppressing capacity of CNE1 NPC cells using a Matrigel assay. By doing so, a significant decrease in the number of invading cells was observed in the exogenously ECRG4 expressing CNE1 cells (Fig. 4f). Together, these data strongly indicate that ECRG4 does indeed act as a TSG in NPC cells.
Exogenous ECRG4 expression inhibits the growth and invasion of NPC cells. The effect of exogenous ECRG4 expression on CNE1 cell clonogenicity investigated by monolayer colony formation assay (a) and soft agar assay (b), respectively. Quantitative analyses are shown in the right panels deduced from at least three independent experiments. c Exogenous ECRG4 expression in stably transfected CNE1 cells analyzed by Western blotting. β-actin was used as a loading control. ECRG4c1 and ECRG4c2 are two independent clones. d The culture media of transfected (ECRG4c1) and empty vector control cells were collected and concentrated for Western blot analysis using an anti-ECRG4 antibody. e Growth curves of stably ECRG4-transfected CNE1 cells deduced from a MTT assay. The values are shown as means ± SD from three independent experiments. **P < 0.01, compared to the vector control. f Invasive properties of indicated cells analyzed through an invasion assay in a Boyden chamber coated with Matrigel. Quantitative numbers of invaded cells are plotted from three different experiments. ***P < 0.001, as compared to the control group
3.6 ECRG4 enhances chemosensitivity through autophagy induction
Previously, it has been reported that ECRG4 over-expression may enhance chemosensitivity in gastric cancer cells [19]. So far, such an effect has not been tested in NPC. To this end, we first evaluated the viability of stable ECRG4 transfected cells (ECRG4c1) as well as empty vector-transfected cells 48 h after treatment with different concentrations of cisplatin. We found that the ECRG4c1 cells were more sensitive to a range of concentrations (1 to 8 μM) of this drug as compared to the empty vector control (Fig. 5a). Using a relatively low concentration of cisplatin (2 μM), we next determined whether apoptosis might be responsible for the increased cell death observed after exogenous ECRG4 over-expression. This dose was also used as pretreatment condition for all subsequent analyses. Flow cytometry analyses revealed a greater induction of the PI+ cell population in stably transfected CNE1 cells (34.5 % in ECRG4c1 versus 6.7 % in empty vector control) (Fig. 5b). However, exogenous ECRG4 over-expression did not significantly increase the percentage of apoptotic cells as compared to the empty vector control (27.8 % versus 25.5 %). Moreover, this over-expression failed to increase cisplatin-induced poly(ADP-ribose) polymerase (PARP) cleavage and caspase-3 activation (Fig. 5c). Thus, exogenous ECRG4 expression significantly enhanced cisplatin-induced cell death in CNE1 cells, predominantly in a caspase-independent manner.
Enhanced chemosensitivity to cisplatin by ECRG4 overexpression in CNE1 cells is mediated by autophagy induction. a Stably ECRG4-transfected CNE1 cells (ECRG4c1) and empty vector control cells were treated with cispatin (dose range from 1 to 8 μM). Cell viability was assessed using a MTT assay. b Cells were pretreated with cisplatin (2 μM) and subjected to Annexin V/PI staining and analyzed by flow cytometry. c After pretreatment with vehicle control or cisplatin, cell lysates were prepared and subjected to SDS-PAGE, followed by Western blot analysis using anti-PARP and anti-caspase-3 antibodies. d After incubation with cisplatin (2 μM), cells were fixed, stained with DAPI to visualize nuclei (blue) and then immunolabeled using an anti-LC3 antibody to detect autophagic vacuoles (green). A quantitative analysis of the percentage of autophagic cells is shown in the right panel. ***P < 0.001, as compared to the control group. e Western blot analysis of LC3-I/LC3-II levels. Cell lysates were separated by SDS-PAGE and analyzed by Western blotting using anti-LC3 or anti-β-actin antibodies. f Cells were treated with 3-MA for 2 h prior to cisplatin exposure, after which cell viabilities were determined using a MTT assay. **P < 0.01, as compared to the control group
Since we found that caspase-dependent cell death was not a prominent feature of CNE1 cells exogenously expressing ECRG4, we set out to test whether the chemosensitizing effect of ECRG4 might be due to autophagy. As shown in Fig. 5d, stable exogenous over-expression of ECRG4 significantly increased autophagocytic microtubule-associated light chain protein 3 (LC3)-positive puncta in CNE1 cells (ECRG4c1), as compared to the empty vector control. We also examined the cleavage of soluble LC3 (LC3-I) to LC3-II and found that its conversion to LC3-II was markedly induced by cisplatin in ECRG4c1 cells, but not in the control empty vector cells (Fig. 5e), which agrees with the results of the immunostaining (Fig. 5d). To further confirm that autophagy plays a role in chemosensitivity to cisplatin in ECRG4 over-expressing CNE1 cells, we pretreated the cells with the autophagy inhibitor 3-MA [22] and then treated the cells with cisplatin. By doing so, we found that pretreatment with 3-MA significantly prevented cell death in the cisplatin exposed ECRG4c1 cells (Fig. 5f), whereas the survival of empty vector control cells was almost not affected under the same conditions. A similar increase in chemosensitivity concomitant with autophagy was also observed in ECRG4c2 cells (data not shown). Together, these findings support the notion that autophagy induction may serve as at least one of the main mechanisms underlying chemosensitivity enhancement of NPC cells through ECRG4 expression.
4 Discussion
As of yet, there is no accurate clinical screening method for nasopharyngeal carcinoma (NPC). Conventional diagnostic methods, such as radiological imaging and endoscopic biopsy-based histological examination, are characterized by a lack of non-invasiveness and convenience. In addition, they cannot be repeated serially for monitoring residual tumor masses or tumor recurrences after treatment [20]. It is conceivable that the measurement of circulating methylated tumor suppressor gene (TSG) promoter DNA may offer the potential to circumvent these problems and to serve as an epigenetic biomarker for the efficient detection and monitoring of malignancies [21, 23]. As a novel candidate TSG, ECRG4 has frequently been found to be inactivated by promoter hypermethylation in different cancer types, including esophageal cancer, prostate cancer, gastric cancer, colorectal carcinoma and glioma [11, 16, 24, 25]. In the present study, a reduced expression of ECRG4 was noted in primary NPC tumors relative to normal tissues and this reduced expression was found to be associated with its promoter methylation status. We also found that ECRG4 promoter hypermethylation occurred frequently in primary NPC specimens, and that its concomitant detection in peripheral blood samples revealed a high sensitivity and specificity for cancer diagnosis and monitoring. To the best of our knowledge, our study is the first to provide evidence of ECRG4 promoter hypermethylation in peripheral blood samples from NPC patients. Considering the fact that the collection of plasma is easy and non-invasive, the detection of ECRG4 promoter hypermethylation in peripheral blood may be developed into an auxiliary screening tool to detect primary NPC within high-risk individuals, as well as to detect residual disease and recurrence.
Based on the clinical implication that restoring ECRG4 expression through the application of demethylating agents may represent a promising therapeutic approach, in vitro functional studies were performed to uncover the potential role of ECRG4 in NPC development. We found that exogenous ECRG4 expression strongly inhibited NPC cell proliferation, colony formation and invasion. These data suggest that ECRG4 may act as a functional TSG that holds therapeutic promise. ECRG4 is thought to be a 148 amino acid secreted 17 KDa protein that can be processed into 14, 10, 8, 6, 4 and 2 KDa peptides, depending on the cell type in which the gene is expressed [9, 16, 26, 27]. To substantiate our ECRG4 expression data, the ECRG4c1 culture medium was collected and concentrated for Western blot analysis. By doing so, we indeed found secreted ECRG4 protein of approximately 14 kDa and 8 kDa in size. The higher molecular weight (MW) band on the Western blot would fit to the MW of the ECRG4 protein without signal peptide. The lower MW band presumably represents the mature ECRG4 peptide with a processed N-terminus.
Although radiotherapy is the most common approach for NPC therapy, chemotherapeutic drugs are also used to treat advanced carcinoma and to enhance radiosensitivity [28]. The success of chemotherapy depends on the sensitivity of the tumor cells to the anti-cancer agents. Cisplatin, a DNA-damaging agent, is one of the most active drugs for the treatment of NPC patients. Cisplatin-based chemotherapy, however, frequently results in acquired resistance, which has been associated with a failure to induce apoptosis [29–32]. In the past, ECRG4 over-expression was found to enhance the chemosentitivity to 5-fluorouracil in gastric cancer cells, mainly through apoptosis induction [19]. Here, we failed to observe an increase in apoptosis, even after exogenous over-expression of ECRG4 in NPC-derived CNE1 cells with an increased chemosensitivity to cisplatin. This discrepancy may be due to the distinct cellular contexts of the different tumor types or, alternatively, to the involvement of distinct cell death pathway(s).
Although apoptosis is the primary mechanism of chemotherapy-induced cell death, autophagy, which can drive the programmed type II cell death pathway, has emerged as another important mechanism of tumor cell death induction [33–35]. Autophagy entails a recycling system that maintains cellular homeostasis and has been implicated in various diseases, including cancer [35]. Whether autophagy promotes or suppresses neoplasia remains, however, controversial since autophagy can be both protective and detrimental to the viability of cancer cells [33–41]. In the present study, we demonstrated that ECRG4 over-expression can significantly increase tumor cell death in NPC-derived CNE1 cells when exposed to cisplatin. Our data also suggest that ECRG4 can enhance the toxicity of cisplatin by augmentation of autophagy rather than apoptosis. The underlying mechanism may depend on the function of ECRG4 to suppress NF-κB activation, which results in an accumulation of reactive oxygen species (ROS) and a stimulation of autophagy [38]. This suggested mechanism is in contrast to that proposed in other studies in which blocking of autophagy synergizes with chemotherapeutics through enhanced apoptosis [39–41]. Also, the consequences of autophagy induction seem to vary between studies, including ours in which we find enhanced type II cell death, whereas others report a protective effect [33, 42–44].
Taken together, we have employed multiple approaches to obtain insight into the role of ECRG4 in NPC development. We observed frequent methylation-mediated silencing of the ECRG4 gene in NPC tumors. The subsequent detection of ECRG4 gene promoter methylation in peripheral blood samples underscores its potential value as a non-invasive biomarker for NPC diagnosis and recurrence monitoring. Our results also revealed that restoring ECRG4 expression may hold therapeutic promises for NPC. Of note, we found that exogenous ECRG4 expression significantly enhances cisplatin-induced cell death through autophagy induction. We conclude that ECRG4 may serve as a NPC biomarker and therapeutic target.
References
H. Li, Y. You, C. Lin, M. Zheng, C. Hong, J. Chen, D. Li, W.W. Au, Z. Chen, XRCC1 codon 399Gln polymorphism is associated with radiotherapy-induced acute dermatitis and mucositis in nasopharyngeal carcinoma patients. Radiat Oncol. 8, 31 (2013)
G. Sanguineti, F.B. Geara, A.S. Garden, S.L. Tucker, K.K. Ang, W.H. Morrison, L.J. Peters, Carcinoma of the nasopharynx treated by radiotherapy alone: determinants of local and regional control. Int J Radiat Oncol. Biol. Phys. 37, 985–996 (1997)
Y. Ran, S. Wu, Y. You, Demethylation of E-cadherin gene in nasopharyngeal carcinoma could serve as a potential therapeutic strategy. J Biochem. 149, 49–54 (2011)
Y. You, W. Yang, Z. Wang, H. Zhu, H. Li, C. Lin, Y. Ran, Promoter hypermethylation contributes to the frequent suppression of the CDK10 gene in human nasopharyngeal carcinomas. Cell Oncol. 36, 323–331 (2013)
A. Geurts van Kessel. The cancer genome: from structure to function. Cell Oncol. 37, 155–165 (2014)
B. Ramaswamy, S. Majumder, S.K. Roy, H. Ghoshal, J. Kutay, J. Datta, M. Younes, C.L. Shapiro, T. Motiwala, S.T. Jacob, Estrogen-mediated suppression of the gene encoding protein tyrosine phosphatase PTPRO in human breast cancer: mechanism and role in tamoxifen sensitivity. Mol Endocrinol. 23, 176–187 (2009)
T. Nozoe, T. Oyama, M. Takenoyama, T. Hanagiri, K. Sugio, K. Yasumoto, Significance of immunohistochemical expression of estrogen receptors alpha and beta in squamous cell carcinoma of the esophagus. Clin Cancer Res. 13, 4046–4050 (2007)
Z. Li, X. Zou, L. Xie, H. Dong, Y. Chen, Q. Liu, X. Wu, D. Zhou, D. Tan, H. Zhang, Prognostic importance and therapeutic implications of PAK1, a drugable protein kinase, in gastroesophageal junction adenocarcinoma. PLoS One. 8, e80665 (2013)
A. Baird, J. Lee, S. Podvin, A. Kurabi, X. Dang, R. Coimbra, T. Costantini, V. Bansal, B.P. Eliceiri, Esophageal cancer-related gene 4 at the interface of injury, inflammation, infection, and malignancy. Gastrointest Cancer. 2014, 131–142 (2014)
T. Su, H. Liu, S. Lu, Cloning and identification of cDNA fragments related to human esophageal cancer. China J Oncol. 20, 254–257 (1998)
L.W. Li, X.Y. Yu, Y. Yang, C.P. Zhang, L.P. Guo, S.H. Lu, Expression of esophageal cancer related gene 4 (ECRG4), a novel tumor suppressor gene, in esophageal cancer and its inhibitory effect on the tumor growth in vitro and in vivo. Int J Cancer. 125, 1505–1513 (2009)
T. Xu, D. Xiao, X. Zhang, ECRG4 inhibits growth and invasiveness of squamous cell carcinoma of the head and neck in vitro and in vivo. Oncol Lett. 5, 1921–1926 (2013)
A. Kurabi, K. Pak, X. Dang, R. Coimbra, B.P. Eliceiri, A.F. Ryan, A. Baird, Ecrg4 attenuates the inflammatory proliferative response of mucosal epithelial cells to infection. PLoS One. 8, e61394 (2013)
A.M. Gonzalez, S. Podvin, S.Y. Lin, M.C. Miller, H. Botfield, W.E. Leadbeater, A. Roberton, X. Dang, S.E. Knowling, E. Cardenas-Galindo, J.E. Donahue, E.G. Stopa, C.E. Johanson, R. Coimbra, B.P. Eliceiri, A. Baird, Ecrg4 expression and its product augurin in the choroid plexus: impact on fetal brain development, cerebrospinal fluid homeostasis and neuroprogenitor cell response to CNS injury. Fluids Barriers CNS. 8, 6 (2011)
R. Sabatier, P. Finetti, J. Adelaide, A. Guille, J.P. Borg, M. Chaffanet, L. Lane, D. Birnbaum, F. Bertucci, Down-regulation of ECRG4, a candidate tumor suppressor gene, in human breast cancer. PLoS One. 6, e27656 (2011)
S. Götze, V. Feldhaus, T. Traska, M. Wolter, G. Reifenberger, A. Tannapfel, C. Kuhnen, D. Martin, O. Müller, S. Sievers, ECRG4 is a candidate tumor suppressor gene frequently hypermethylated in colorectal carcinoma and glioma. BMC Cancer. 9, 447 (2009)
Y. Mori, H. Ishiguro, Y. Kuwabara, M. Kimura, A. Mitsui, H. Kurehara, R. Mori, K. Tomoda, R. Ogawa, T. Katada, K. Harata, Y. Fujii, Expression of ECRG4 is an independent prognostic factor for poor survival in patients with esophageal squamous cell carcinoma. Oncol Rep. 18, 981–985 (2007)
W. Li, X. Liu, B. Zhang, D. Qi, L. Zhang, Y. Jin, H. Yang, Overexpression of candidate tumor suppressor ECRG4 inhibits glioma proliferation and invasion. J Exp Clin Cancer Res. 29, 89 (2010)
C.P. Jiang, B.H. Wu, B.Q. Wang, M.Y. Fu, M. Yang, Y. Zhou, F. Liu, Overexpression of ECRG4 enhances chemosensitivity to 5-fluorouracil in the human gastric cancer SGC-7901 cell line. Tumour Biol. 34, 2269–2273 (2013)
Y. You, Y. Chen, X. Zheng, J. Meltzer, H. Zhang, Aberrant methylation of the PTPRO gene in peripheral blood as a potential biomarker in esophageal squamous cell carcinoma patients. Cancer Lett. 315, 138–144 (2012)
Y. You, J. Liu, Z. Wang, Y. Zhang, Y. Ran, X. Guo, H. Liu, H. Wang, The enhancement of radio-sensitivity in human esophageal squamous cell carcinoma cells by zoledronic acid and its potential mechanism. Cytotechnology. 66, 17–25 (2014)
A. Coker-Gurkan, E.D. Arisan, P. Obakan, E. Guvenir, N.P. Unsal, Inhibition of autophagy by 3-MA potentiates purvalanol-induced apoptosis in Bax deficient HCT 116 coloncancer cells. Exp Cell Res. 328, 87–98 (2014)
P. Ulivi, R. Silvestrini, Role of quantitative and qualitative characteristics of free circulating DNA in the management of patients with non-small cell lung cancer. Cell Oncol. 36, 439–448 (2013)
Y.B. Wang, C.F. Ba, Promoter methylation of esophageal cancer-related gene 4 in gastric cancer tissue and its clinical significance. Hepatogastroenterology. 59, 1696–1698 (2012)
D.K. Vanaja, M. Ehrich, D. Van den Boom, J.C. Cheville, R.J. Karnes, D.J. Tindall, C.R. Cantor, C.Y. Young, Hypermethylation of genes for diagnosis and risk stratification of prostate cancer. Cancer Invest. 27, 549–560 (2009)
Y. Kujuro, N. Suzuki, T. Kondo, Esophageal cancer-related gene 4 is a secreted inducer of cell senescence expressed by aged CNS precursor cells. Proc Natl Acad Sci U S A. 107, 8259–8264 (2010)
X. Dang, S. Podvin, R. Coimbra, B. Eliceiri, A. Baird, Cell-specific processing and release of the hormone-like precursor and candidate tumor suppressor gene product, Ecrg4. Cell Tissue Res. 348, 505–514 (2012)
S.Y. Low, B.S. Tan, H.L. Choo, K.H. Tiong, A.S. Khoo, C.O. Leong, Suppression of BCL-2 synergizes cisplatin sensitivity in nasopharyngeal carcinoma cells. Cancer Lett. 314(2), 166–175 (2012)
J. Wang, H. Wang, L. Zhao, S. Fan, Z. Yang, F. Gao, L. Chen, G.G. Xiao, J. Molnár, Q. Wang, Down-regulation of P-glycoprotein is associated with resistance to cisplatin and VP-16 in human lung cancer cell lines. Anticancer Res. 30, 3593–3598 (2010)
Z.H. Siddik, Cisplatin: mode of cytotoxic action and molecular basis of resistance. Oncogene. 22, 7265–7279 (2003)
X. Wang, J.R. Masters, Y.C. Wong, A.K. Lo, S.W. Tsao, Mechanism of differential sensitivity to cisplatin in nasopharyngeal carcinoma cells. Anticancer Res. 21, 403–408 (2001)
I.A. Voutsadakis, The chemosensitivity of testicular germ cell tumors. Cell Oncol. 37, 79–94 (2014)
T.R. O'Donovan, G.C. O'Sullivan, S.L. McKenna, Induction of autophagy by drug-resistant esophageal cancer cells promotes their survival and recovery following treatment with chemotherapeutics. Autophagy. 7, 509–524 (2011)
M.J. Nyhan, T.R. O'Donovan, B. Elzinga, L.C. Crowley, G.C. O'Sullivan, S.L. McKenna, The BH3 mimetic HA14-1 enhances 5-fluorouracil-induced autophagy and type II cell death in oesophageal cancer cells. Br J Cancer. 106, 711–718 (2012)
J.M. Yuk, D.M. Shin, K.S. Song, K. Lim, K.H. Kim, S.H. Lee, J.M. Kim, J.S. Lee, T.H. Paik, J.S. Kim, E.K. Jo, Bacillus calmette-guerin cell wall cytoskeleton enhances colon cancer radiosensitivity through autophagy. Autophagy. 6, 46–60 (2010)
Y. Wei, T. Kadia, W. Tong, M. Zhang, Y. Jia, H. Yang, Y. Hu, F.P. Tambaro, J. Viallet, S. O'Brien, G. Garcia-Manero, The combination of a histone deacetylase inhibitor with the Bcl-2 homology domain-3 mimetic GX15-070 has synergistic antileukemia activity by activating both apoptosis and autophagy. Clin. Cancer Res. 16, 3923–3932 (2010)
B. Sirichanchuen, T. Pengsuparp, P. Chanvorachote, Long-term cisplatin exposure impairs autophagy and causes cisplatin resistance in human lung cancer cells. Mol. Cell. Biochem. 364, 11–18 (2012)
W. Hu, S.S. Chen, J.L. Zhang, X.E. Lou, H.J. Zhou, Dihydroartemisinin induces autophagy by suppressing NF-κB activation. Cancer Lett. 343, 239–248 (2014)
J. Pan, C. Cheng, S. Verstovsek, Q. Chen, Y. Jin, Q. Cao, The BH3-mimetic GX15-070 induces autophagy, potentiates the cytotoxicity of carboplatin and 5-fluorouracil in esophageal carcinoma cells. Cancer Lett. 293, 167–174 (2010)
Y. Sun, J.H. Liu, L. Jin, Y.X. Sui, L. Lai, Y. Yang, Inhibition of Beclin 1 expression enhances cisplatin-induced apoptosis through a mitochondrial-dependent pathway in human ovarian cancer SKOV3/DDP cells. Oncol. Res. 21, 261–269 (2014)
X.L. Guo, D. Li, F. Hu, J.R. Song, S.S. Zhang, W.J. Deng, K. Sun, Q.D. Zhao, X.Q. Xie, Y.J. Song, M.C. Wu, L.X. Wei, Targeting autophagy potentiates chemotherapy-induced apoptosis and proliferation inhibition in hepatocarcinoma cells. Cancer Lett. 320, 171–179 (2012)
S. Daido, A. Yamamoto, K. Fujiwara, R. Sawaya, S. Kondo, Y. Kondo, Inhibition of the DNA-dependent protein kinase catalytic subunit radiosensitizes malignant glioma cells by inducing autophagy. Cancer Res. 65, 4368–4375 (2005)
C.E. Zois, M.I. Koukourakis. Radiation-induced autophagy in normal and cancer cells: towards novel cytoprotection and radio-sensitization policies? Autophagy 5, 442–450 (2009)
A. Apel, I. Herr, H. Schwarz, H.P. Rodemann, A. Mayer, Blocked autophagy sensitizes resistant carcinoma cells to radiation therapy. Cancer Res. 68, 1485–1494 (2008)
Acknowledgments
We thank Mr. Zhen Zhang (Department of Biochemistry and Molecular Biology, University of Kansas Medical Center) for his careful reading of this manuscript and kind suggestions. This work was supported by the Science and Technology Planning Project of Henan Province, China (142102310464), the 2015 Annual Natural Science Foundation of Luohe Medical College (Y.-J. You), the Natural Science Foundation of Hubei Province (2014CFC1154), and the Foundation of Medical College of Hubei University of Arts and Science (YXKY 201402).
Conflicts of Interest
The authors have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Yanjie You, Wenjun Yang and Xin Qin have contributed equally to this work.
Rights and permissions
About this article
Cite this article
You, Y., Yang, W., Qin, X. et al. ECRG4 acts as a tumor suppressor and as a determinant of chemotherapy resistance in human nasopharyngeal carcinoma. Cell Oncol. 38, 205–214 (2015). https://doi.org/10.1007/s13402-015-0223-y
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-015-0223-y