Abstract
In the era of genomic-assisted breeding for crop improvement, developing new molecular markers and validating them for use in breeding programs are the prelude. Dolichos bean is one of the most important vegetable legume crops owing to its nutrient-rich green pods used as vegetables. Limitations in genomic resources, including molecular markers, restrict the accelerated improvement of the crop. In the present investigation, a set of 430 new simple sequence repeat markers was developed from sequence information of a reference variety. These markers included di- and tri-nucleotide repeats. The markers were assayed on an association panel, which was evaluated for green pod yield over 5 years. A multi-locus model, FarmCPU, was used to assess the marker-trait association analysis. A total of 106 marker-trait associations were identified using an efficient mixed-model approach. Tri-nucleotide repeats were more informative and predominantly associated with trait. Among these markers, 17 were associated with a high level of significance. Markers LP-D-68 and LP-D-14 were identified with a high level of significance in 5-year pooled data and explained 12.70% and 12% of the phenotypic variance, respectively. These markers associated with a high level of confidence have significant scope for use in marker-assisted selection programmes. Other associated markers may be utilized for improving parents through marker-assisted recurrent selection or genomic selection programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Field bean, or dolichos bean (Lablab purpureus L. Sweet), is an ancient legume vegetable crop grown under rainfed conditions in semi-arid regions of India and Africa. It serves as a major source of protein for people in India and developing countries in Africa (Ramesh and Byregowda 2016). The dolichos bean extracts are proven to inhibit viral infections caused by influenza and the SARS-CoV-2 virus (Liu et al. 2020). The fresh green pods are the harvestable and consumable economic product of the dolichos bean. Fresh green pods emit a fragrance that is preferred by consumers (Udaykumar et al. 2016), and fresh seeds are known to have 28% protein, 2% fat, 8.5% fiber, and 60% carbohydrates. Moreover, it is a rich source of micronutrients like calcium, phosphorous, and iron (Kay 1975). Despite its multi-purpose utility, the dolichos bean is considered an underutilized orphan legume crop owing to its low yielding abilities. The development of high yielding cultivars with consumer-favored traits and wide adaptability to different growing climates is expected to enhance production. Breeding efforts are being made to increase the productivity of green pod yield with higher adaptability of the crop both in India and Africa (Ramesh and Byregowda 2016).
Dolichos bean is a self-pollinated crop. Thus, natural variation is being exploited for varietal development programs. The variation in green pod yield caused by genetic (Ajaykumar et al. 2021; Spoorthi et al. 2022) and non-genetic factors was extensively investigated and reported as a quantitative characteristic. Considering predominant self-pollination mechanisms (Kukade and Tidke 2014), pure-line varieties are the only potential cultivar option in the crop. With the improvements in breeding methodologies, it is important to use new molecular markers to increase precision. The development of new genomic resources, including genome-wide markers, is important to adapt new breeder approaches like genomic selection (Anilkumar et al. 2022) and accelerate the breeding process for rapidly achieving genetic gains in orphan crops. However, the physical map information and chromosome-level genome sequence information of the dolichos bean were not available until recently. Njaci et al. (2023) reported a high-quality genome assembly highlighting the genome organization of anti-nutritional factors like trypsin inhibitor. The genome sequence also helped in delineating two events of dolichos domestication in Africa. The new molecular markers help to unearth the crop domestication events and exploit them in genetic improvement programs. Exploring genomic regions associated with economic traits and deploying them in marker-assisted breeding programs enhance the efficiency of dolichos breeding. There is only limited literature available on the identification of genetic determinants for harvestable economic products, such as green pod yield. However, generating knowledge on quantitative trait loci (QTL) for green pod yield will serve as a prelude to the strategic planning of breeding programs (Vaijayanthi et al. 2019).
Among the QTL mapping approaches in plants, bi-parental populations are frequently used to detect and quantify the effect of QTL. However, the variation explained by the QTL discovered in bi-parental populations is limited to variation that existed between two parents used for population development. On the other hand, the genome-wide association study (GWAS) approach utilizes genetic variation from diverse/natural populations to detect all possible genetic components responsible for the expression of a quantitative trait (Korte and Farlow 2013). Diverse populations are excellent sources for QTL mapping and allele mining (Qiu et al. 2013). GWAS is found to be promising for identifying causative alleles for quantitative traits by exploiting ancestral linkage disequilibria between genome-wide marker alleles and genes responsible for the trait. Natural dissimilarities that exist between diverse dolichos bean populations help exploit historical recombination frequencies and increase the resolution of QTL mapping (Rafalski 2010). Furthermore, multi-season precision phenotype information and evenly distributed genome-wide marker information increase the precision of the identification of QTL for complex traits. Hence, the GWAS approach can be considered highly efficient and effective for the identification of all possible causative alleles for green pod yield in dolichos bean.
In the present investigation, a statistically strong association panel evaluated over five years and assayed with newly developed simple sequence repeat (SSR) markers was used to identify marker-trait associations for green pod yield in dolichos bean with a multi-locus model. The results of this study will have significant scope for adapting marker-assisted breeding strategies for improving dolichos varieties with high green pod yield.
Materials and methods
Genetic material and experiment location
The association panel was constituted of 96 diverse genotypes, which were collected from major dolichos growing regions in India and some accessions in Kenya. The experimental material covers most of the genetic diversity available in India. The information on geographic origin of these genotypes is presented in Fig. S1 and Table S1. The panel included two advanced breeding lines and one released variety from the University of Agricultural Sciences, Bangalore. The seed material of these genotypes was maintained at the Department of Genetics and Plant Breeding, University of Agricultural Sciences, Bangalore. Experimental genotypes were evaluated for green pod yield during the rainy seasons of 2012, 2014, 2015, 2018, and 2019 at the experimental plots of the University of Agricultural Sciences, Bangalore. The experimental location represents ideal climatic conditions for dolichos cultivation.
Evaluation for green pod yield
The experimental genotypes were planted in a simple lattice design with two replications in all the years. Ten square lattice design was used to evaluate 96 experimental genotypes and two dummy genotypes in two sets to complete the design. Each genotype was planted in 4 mt rows with 20-cm spacing between plants and 45 cm between rows. Recommended agronomic and protection practices were followed to raise a healthy crop. Briefly, the crop was grown in wet seasons under rainfed conditions, and supplementary irrigation was provided to save the crop during dry spells. The experiment was conducted in fertile soils, and recommended dose of fertilizers (20 N:60 P:60 K) were applied every year while sowing. The weeding was done manually, and chemical pest control measures were taken to protect the crop till the harvesting. The green pod yield was recorded from ten plants in each row, leaving border plants. The green pod yield was recorded at multiple pickings at the mature green-filled pod stage. Cumulative yield over multiple pickings was considered as green pod yields per plant.
Marker development and marker assay
A set of 413 new simple sequence repeat markers was developed from sequence information generated by double digest restriction-site associated DNA (ddRAD-seq) sequencing of the popular variety HA-4. The ddRAD approach was chosen considering the non-availability of reference genome in the crop. The young leaves were supplied to Eurofins Scientific® for extraction of pure-quality DNA and use it for sequencing. The SSR markers were developed from the sequence information using SSR identification tool (Temnykh et al. 2001) for finding potential repeat sequences and Primer 3 software to select potential SSRs (Rozen and Skaletsky 1999). The markers included tri-nucleotide and di-nucleotide repeats. The genome size of dolichos is 367 mb with 2n = 22 chromosomes (Iwata et al. 2013). Young leaves of 35-day-old seedlings were collected, and genomic DNA was extracted following the cetyl-tri-methyl ammonium bromide (CTAB) method (Doyle and Doyle 1990). The quality of DNA was examined using gel electrophoresis and quantified using NanoDrop (Thermo Fisher Scientific). The genomic DNA was diluted to 20 ng/µL before use in amplification. A polymerase reaction mixture was prepared with 1 µL of 20 ng/µL of genomic DNA, forward and reverse primers each with 1 µL of 10 pmol/µL, standard PCR master mix (Takara®) of 4 µL and a volume of 10 µL by adding 3 µL of nuclease-free water. The PCR program was adjusted at 95 °C for 5 min for the initial hot start, followed by 40 cycles of denaturation at 95 °C for 30 s, primer annealing for 1 min with the respective annealing temperature of the marker, and primer extension for 1 min at 72 °C, followed by a final extension for 8 min at 72 °C. The amplified fragments were visualized on a gel electrophoresis unit with 3.4% agarose gel (combination of metaphor agarose and low EEO normal agarose in 1:1 ratio was used for high resolution) containing 0.05 µL/ml ethidium bromide immersed in a 0.5 × Tris borate EDTA (TBE) solution at a constant voltage of 85 V for 4 h. Visualized amplicons were scored in a binary format and used as genotype information for further analysis.
Statistical analysis
Variation in green pod yield
The green pod yield per plant was subjected to preliminary analysis in all years. The summary statistics, including distribution and range, skewness, and kurtosis parameters, were estimated considering mean values for green pod yield over replications. An analysis of variance using a lattice design was performed for all the years separately to verify the significant variation between genotypes in the study panel. All these preliminary analyses were completed using RStudio version 1.4.17. The normal distribution of green pod yield in the population in all the years was tested by estimating Shipir-Wilk’s p. The measures of descriptive statistics and distribution pattern of the panel with respect to green pod yield were estimated to ensure the suitability of the population for genome-wide association analysis.
Population structure and allele diversity analysis
Information generated using 413 new markers was used to estimate allelic diversity and population structure. Allelic information on the population was estimated using PowerMarker V3.25 (Liu and Muse 2005). The population structure of the panel was described using three different approaches: (a) principal component analysis (PCA) to differentiate genotypes into possible subgroups based on marker information. The PCA was performed using “factoextra” package in RStudio (Kassambara and Mundt 2017). (b) Phylogenetic distances between genotypes in the panel were delineated using the neighbor-joining approach in DARwin software, which was further represented as an unrooted tree with clusters using the iTOL online tool (Letunic and Bork 2021). (c) The familial relationships between genotypes in the association population were estimated using the VanRaden kinship algorithm (VanRaden 2008) and presented in the form of a heatmap. A familial relationship index between + 1 and − 1 was considered an indication of a good population fit for genome-wide association analysis.
Genome-wide association analysis
Genome-wide association analysis was performed to associate the marker alleles with green pod yield variation in the population. Marker alleles of new markers developed were assayed in the population and used to identify markers associated with green pod yield in all 5 years. Analysis was performed using the GAPIT package in RStudio (Lipka et al. 2012). While analyzing genome-wide association, a multi-locus model, fixed and random model circulating probability unification (FarmCPU), was used (Liu et al. 2016). The multi-locus model was preferred over the single-locus model considering their efficiency in detecting marginal QTL effects for quantitative traits in diverse backgrounds (Wang et al. 2016; Cui et al. 2018). The GAPIT package is in-built with an efficient mixed model approach (EMMA) for simultaneous correction of population structure while detecting marker trait associations (Yu et al. 2006). The threshold used to decide the significance of marker-trait associations was false discovery rate (FDR) corrected p < 0.05 values. The marker-trait associations with p < 0.05 were considered significant associations.
Results
Variation in green pod yield
The green pod yield per plant significantly varied in all the experimental years (Table S2). The variation explained by genotypes was significant in all the years, as indicated by the significance in the analysis of variance. Yield of green pods per plant was ranged between 39.64 g (2012) and 332.76 g (2018) when considering all the years. The mean performance varied between 105.20 g (2012) and 164.87 g (2018). Considering the non-significance of the homogeneity test, the data over different years was pooled. Within the pooled data, the green pod yield ranged from 94.76 to 179.93 g, with an average of 129.33 g (Table 1). The green pod yield was distributed normally in all the years with negligible skewness and kutrosis less than three, indicating a platykurtic distribution of the data in all the years (Fig. 1). Skewness and kurtosis support the quantitative inheritance of green pod yield in dolichos bean, with several minor genes controlling the trait with additive effects. The broad-sense heritability coefficient estimated and averaged over all the years of experimentation was significantly high (0.65). The non-significant estimates for Shipiro-Wilk’s p values indicate the normal distribution of green pod yield in the population in all the years and in the pooled data.
Allele diversity and population structure
Among the 413 new markers, some markers failed to amplify in all the genotypes, and some were monomorphic among the genotypes of the association panel. Out of 413 markers, 370 were amplified and exhibited polymorphism among the individuals on the association panel (Table S3 and Fig S2). Most of the markers were bi-allelic in the population, and the maximum number of alleles per individual in the population was three, totaling 783 alleles amplified in the population. Among them, 562 alleles were identified as effective alleles that are significantly contributing to the diversity of the population. The allele diversity index in the population ranged between 0.021 and 0.65, with the Shannon information index ranging from 0.06 to 1.06. The polymorphic information content (PIC) of the new markers used in the study ranged from 0.02 to 0.38, with an average of 0.24 (Table S4). Di- and tri-nucleotide repeat markers together accounted for about 98.11% of the polymorphism in the population. The allelic patterns of di- and tri-nucleotide repeats for estimated population parameters are presented in Table 2.
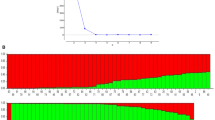
To address the population structure and identify subpopulations within the association population, different approaches were followed. Principal component analysis classified the main population into three subpopulations (Fig. 2a). The existence of these subpopulations was further envisaged by a phylogenetic tree constructed based on genetic distances between genotypes (Fig. 2b). Based on familial relationships between individuals in the association panel, the VanRaden kinship algorithm also classified the main population into three subpopulations (Fig. 2c). The structure of the population was accounted for by considering three principal components while analyzing for marker-trait associations, and an efficient mixed model approach was used to correct the population structure.
Population structure analysis: a 3D representation of principle component analysis showing three subpopulations, b phylogenetic tree classifying the genotypes into three groups based on genetic distances, and c heat map of kinship matrix; the heat map shows the level of relatedness among the population. The darker areas show the level of relatedness between varieties, and dendrogram drawn at the top represents the phylogenetic grouping/clustering of genotypes in the association panel
Marker-trait associations for green pod yield
The FarmCPU model identified a total of 106 markers associated with green pod yield across different years-wise data and pooled data used for association analysis at the threshold probability of 0.05 (p < 0.05) (Fig. 3; Table S5). Among the years, 18 markers were found to be significantly associated with the trait in 2012, 16 markers in 2014, 16 markers in 2015, 24 markers in 2018, and 11 markers in 2019. There were 21 markers associated with the trait in pooled data over the years. Among the 21 markers identified as significant in the pooled data, two markers, LP-D-68 and LP-D-14, were associated with a high level of significance and a high level of explained variance. Tri-nucleotide repeats were more prevalent among the associated markers, with a wide range of phenotypic variation explained. The phenotypic variation explained by the associated markers ranged between 5.10% and 12.70%. Some associations were significant at a probability < 0.01 with high explained train phenotypic variance (Table 3). A total of 17 markers over years and pooled data were associated with green pod yield at high level of significance and explained phenotypic variation. Most of the associated markers were consistent at least over 2 or 3 years, and many of them were significant in pooled analysis (Table S6).
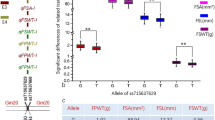
The allelic effects of associated markers ranged from significant negative effects to significant positive effects. Two major associations identified in pooled analysis with high phenotypic variance explained showed highly positive allelic effects. Marker LP-D-14 was significantly positively associated with green pod yield (12% PVE), where a 260-bp allele was associated with high green pod yield and a 280-bp allele was associated with lower green pod yields (Fig. 4). Similarly, marker LP-D-68 was significantly associated with 12.70% phenotype variance explained, and allele 250 bp was associated with high green pod yields, whereas allele 200 bp was associated with lower green pod yields. The markers associated with higher levels of significance may be used in marker-assisted improvement of green pod yield in dolichos bean.
Discussion
Improving the green pod yields of field beans or dolichos beans enhances the farmers’ income for those who grow them for vegetable purposes. The dolichos bean has its demand in the southern states of India and most parts of Africa, owing to its fragrant green pods used as vegetables. The pods are proven to be highly nutritional vegetables rich in micronutrients (Udaykumar et al. 2016). Conventional breeding strategies are being used to develop improved varieties. However, the genetic gain achieved is slow, and the time requirement for development is relatively high (Byregowda et al. 2015). The genetic improvement of dolichos bean for enhanced yields is limited considering the lack of molecular markers. Developing new genomic resources, including molecular markers and their association with economic traits, has significant importance in accelerating breeding programs. In this context, a set of new marker resources was developed and associated those markers with genomic regions for economic traits like green pod yield.
Proper phenotyping and understanding the genetic diversity that exists in the population are prerequisites in any genetic study, including strategizing an appropriate selection program. More specifically, association mapping demands highly precise phenotype information to tag molecular markers to the respective genomic regions (Alqudah et al. 2020). Phenotype information from the association panel over years/environments is more important for precise results (Campbell et al. 2003). In the present investigation, the association panel genotypes were evaluated over 5 years for their green pod yields. The analysis of variance year-wise and pooled data over the years indicated significant variation in the genotypes’ performance for green pod yield. The significance of Shipiro-Wilk’s test in all the experimental years indicated continued variation for green pod yields following normal distribution, which is a characteristic of typically quantitative traits. Substantial variability in green pod yield in dolichos bean over a multi-year evaluation was documented earlier by Vaijayanthi et al. (2015) and Spoorthi et al. (2022). Further, the negligible skewness and playkurtic distribution of green pod yields in all the years demonstrate the involvement of several minor genes acting additively in the expression of the trait. Phenotypic diversity and distribution suggest the suitability of a selected population for association mapping to identify the markers associated with QTL for green pod yield in dolichos bean.
The new molecular markers developed and utilized in the present investigation were found superior for delineating population diversity and population structure. Among di- and tri-nucleotide repeats, tri-nucleotide repeats accounted for the maximum diversity in the population due to their high informativeness compared to di-nucleotide repeats. Hence, these markers may help overcome the impediment of low marker availability in dolichos bean breeding (Spoorthi et al. 2024). Similar efforts to assess the genetic diversity using simple sequence repeat markers were made for chickpea (Upadhyaya et al. 2008) and common bean (Blair et al. 2009). Additionally, the new markers explained the familial relationship between genotypes in the association panel. The population was classified into three subpopulations based on genetic distances between genotypes. This suggests an unequal distribution of alleles within the population, and differences arise due to ancestral linkage disequilibrium among genotypes. This variation in allele distribution directly affects the linkage disequilibrium throughout the genome. Hence, three principal components were considered before analyzing marker-trait associations to account for population structure. Similar numbers of subpopulations were detected in dolichos bean core collection population used for association mapping (Vaijayanthi et al. 2019). Addressing the population structure by considering a significant number of principal components and an effective model like the effective mixed model algorithm (EMMA) ensures the precision of marker-trait associations identified (Mather et al. 2004).
The phenotype information of green pod yield in different years and genotype data obtained by assaying 370 new simple sequence repeat markers helped in the identification of 106 significant marker-trait associations. The multi-locus model used in the present investigation identified all those markers associated with traits, even with marginal levels of phenotypic variation. Among them, 17 markers were associated with a high level of significance. The markers associated with high explained phenotypic variance may be directly used in marker-assisted selection programs for improving green pod yield in dolichos bean. Similarly, remaining markers may be utilized in marker-assisted recurrent selection programs for parental population improvement. Furthermore, the markers associated with traits may be best utilized in genomic selection program for improving prediction accuracy (Anilkumar et al. 2023). Hence, all the markers associated with green pod yield may be utilized in genomic selection for green pod yield in dolichos bean. Overall, the results of this study have significant application in the strategic improvement of dolichos bean through marker-assisted selection or genomic selection programs.
Conclusion
Dolichos bean is considered one of the most underutilized and highly nutritious orphan legume vegetables. The crop is popular in southern Indian states and many African countries due to its nutritional quality and pod fragrance. Breeding efforts to improve this orphan vegetable legume are being made continuously. However, the lack of genome-wide molecular markers and molecular tools limits the improvement of this crop. In the present investigation, a set of 413 new molecular markers developed from sequence information was utilized for genetic diversity and marker-trait association analysis. The study identified new molecular markers associated with green pod yields. The findings of this investigation have scope for marker-assisted improvement of dolichos beans for higher green pod yield. Further, the associated markers may be utilized in genomic selection programs for accelerated improvement of dolichos bean green pod yield.
Data availability
The data generated during the experimentation is presented as tables and supplementary tables in the manuscript.
References
Ajaykumar H, Ramesh S, Sunitha NC, Anilkumar C (2021) Assessment of natural DNA methylation variation and its association with economically important traits in dolichos bean (Lablab purpureus L.Var. Lignosus) using AMP-PCR assay. J Appl Genet 62(4):571–83. https://doi.org/10.1007/s13353-021-00648-x
Alqudah AM, Sallam A, Baenziger PS, Börner A (2020) GWAS: fast-forwarding gene identification and characterization in temperate cereals: lessons from barley–a review. J Adv Res 22:119–135. https://doi.org/10.1016/j.jare.2019.10.013
Anilkumar C, Sunitha NC, Harikrishna DNB, Ramesh S (2022) Advances in integrated genomic selection for rapid genetic gain in crop improvement: a review. Planta 256(5):87. https://doi.org/10.1007/s00425-022-03996-y
Anilkumar C, Muhammed Azharudheen TP, Sah RP, Sunitha NC, Devanna BN, Marndi BC, Patra BC (2023) Gene based markers improve precision of genome-wide association studies and accuracy of genomic predictions in rice breeding. Heredity 130(5):335–345. https://doi.org/10.1038/s41437-023-00599-5
Blair MW, Díaz LM, Buendía HF, Duque MC (2009) Genetic diversity, seed size associations and population structure of a core collection of common beans (Phaseolus vulgaris L.). Theor Appl Genet 119:955–972. https://doi.org/10.1007/s00122-009-1064-8
Byregowda M, Girish G, Ramesh S, Mahadevu P, Keerthi CM (2015) Descriptors of Dolichos bean (Lablab purpureus L.). J Food Legumes 28:203–214
Campbell BT, Baenziger PS, Gill KS, Eskridge KM, Budak H, Erayman M, Dweikat I, Yen Y (2003) Identification of QTLs and environmental interactions associated with agronomic traits on chromosome 3A of wheat. Crop Sci 43:1493–1505. https://doi.org/10.2135/cropsci2003.1493
Cui Y, Zhang F, Zhou Y (2018) The application of multi-locus GWAS for the detection of salt-tolerance loci in rice. Front Plant Sci 9:1464. https://doi.org/10.3389/fpls.2018.01464
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Iwata A, Greenland CM, Jackson SA (2013) Cytogenetics of legumes in the Phaseoloid clade. The Plant Genome Plant Genome 6:2013–2103. https://doi.org/10.3835/plantgenome2013.03.0004
Kassambara A, Mundt F (2017) Factoextra: extract and visualize the results of multivariate data analyses. R Package Version 1:337–354
Kay DE (1975) Food legumes. In: TPI crop and product digest No. 3. Lablab purpureus. Hyacinth bean, pp 184–196
Korte A, Farlow A (2013) The advantages and limitations of trait analysis with GWAS: a review. Plant Methods 9:1–9. https://doi.org/10.1186/1746-4811-9-29
Kukade SA, Tidke JA (2014) Reproductive biology of Dolichos lablab L. (Fabaceae). Indian J Plant Sci 3:22–25
Letunic I, Bork P (2021) Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49:W293–W296. https://doi.org/10.1093/nar/gkab301
Lipka AE, Tian F, Wang Q, Peiffer J, Li M, Bradbury PJ, Gore MA, Buckler ES, Zhang Z (2012) GAPIT: genome association and prediction integrated tool. Bioinformatics 28:2397–2399. https://doi.org/10.1093/bioinformatics/bts444
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129. https://doi.org/10.1093/bioinformatics/bti282
Liu X, Huang M, Fan B, Buckler ES, Zhang Z (2016) Iterative usage of fixed and random effect models for powerful and efficient genome-wide association studies. PLoS Genet 12(2):e1005767. https://doi.org/10.1371/journal.pgen.1005767
Liu YM, Shahed-Al-Mahmud M, Chen X, Chen TH, Liao KS, Lo JM, Wu YM, Ho MC, Wu CY, Wong CH, Jan JT (2020) A carbohydrate-binding protein from the edible lablab beans effectively blocks the infections of influenza viruses and SARS-CoV-2. Cell Rep 32:108016. https://doi.org/10.1016/j.celrep.2020.108016
Mather D, Hayes P, Chalmers K, Eglinton J, Matus I, Richardson K, Von Zitzewitz J, Marquez-Cedillo L, Hearnden P, Pal N (2004) Use of SSR marker data to study linkage disequilibrium and population structure in Hordeum vulgare: prospects for association mapping in barley. Czech J Genet Plant Breed 302–307. https://hdl.handle.net/2440/39842. Accessed 20 Sept 2023
Njaci I, Waweru B, Kamal N, Muktar MS, Fisher D, Gundlach H, Muli C, Muthui L, Maranga M, Kiambi D, Maass BL (2023) Chromosome-level genome assembly and population genomic resource to accelerate orphan crop lablab breeding. Nat Commun 14:1915. https://doi.org/10.1038/s41467-023-37489-7
Qiu LJ, Xing LL, Guo Y, Wang J, Jackson SA, Chang RZ (2013) A platform for soybean molecular breeding: the utilization of core collections for food security. Plant Mol Biol 83:41–50. https://doi.org/10.1007/s11103-013-0076-6
Rafalski JA (2010) Association genetics in crop improvement. Curr Opin Plant Biol 13:174–180. https://doi.org/10.1016/j.pbi.2009.12.004
Ramesh S, Byregowda M (2016) Dolichos bean (Lablab purpureus L. Sweet var. Lignosus) genetics and breeding–present status and future prospects. Mysore J Agric Sci 50:481–500
Rozen S, Skaletsky H (1999) Primer3 on the WWW for general users and for biologist programmers. In: Misener S, Krawetz SA (eds) Bioinformatics Methods and Protocols, Methods in Molecular Biology TM. Humana Press: Totowa, NJ, pp 365–386.
Spoorthi V, Ramesh S, Sunitha NC, Vaijayanthi PV (2022) Prediction of genotype performance for untested years based on additive main effects and multiplicative interaction and linear mixed models: an illustration using dolichos bean (Lablab purpureus (L.) Sweet) multiyear data. Ann Appl Biol 180:224–235. https://doi.org/10.1111/aab.12726
Spoorthi V, Ramesh S, Sunitha NC, Anilkumar C, Vedashree MS (2024) Revisiting population genetics with new genomic resources in dolichos bean (Lablab purpureus L. Sweet): an orphan crop. Genet Resour Crop Evol 71(2):761–771. https://doi.org/10.1007/s10722-023-01655-z
Temnykh S, DeClerck G, Lukashova A, Lipovich L, Cartinhour S, McCouch S (2001) Computational and experimental analysis of microsatellites in Rice (Oryza sativa L.): frequency, length variation, transposon associations, and genetic marker potential. Genome Res 11:1441–1452. https://doi.org/10.1101/gr.184001
Udaykumar HR, Byregowda M, Ramesh S, Vasundhara M (2016) An objective method of phenotyping pod fragrance in dolichos bean Lablab purpureus L. Sweet var. lignosus. Int J Agric Sci 8:1867–1869
Upadhyaya HD, Sastry GCLL, DVSSR (2008) Plant genetic resources management: collection, characterization, conservation and utilization. J SAT Agril Res 6:1–15
Vaijayanthi PV, Ramesh S, Gowda MB, Rao AM, Keerthi CM, Reena GA (2015) Genetic variability for morpho-metric traits in Dolichos bean (Lablab purpureus L. Sweet) germplasm. J Food Legumes 28:5–10
Vaijayanthi PV, Ramesh S, Gowda MB, Rao AM, Keerthi CM (2019) Genome-wide marker-trait association analysis in a core set of Dolichos bean germplasm. Plant Genet Resour 17:1–11. https://doi.org/10.1017/S1479262118000163
VanRaden PM (2008) Efficient methods to compute genomic predictions. J Dairy Sci 91:4414–4423. https://doi.org/10.3168/jds.2007-0980
Wang SB, Feng JY, Ren WL, Huang B, Zhou L, Wen YJ, Zhang J, Dunwell JM, Xu S, Zhang YM (2016) Improving power and accuracy of genome-wide association studies via a multi-locus mixed linear model methodology. Sci Rep 6:19444. https://doi.org/10.1038/srep19444
Yu J, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208. https://doi.org/10.1038/ng1702
Funding
The study was financially supported by the Indian Council of Agricultural Research (ICAR)—World Bank funded project to carry out part of the research (ICAR_NAHEP; F. No./NAHEP/CAAST/2018–19/08).
Author information
Authors and Affiliations
Contributions
SR and AC conceptualized the experiment. VS, MSV, and PVV: evaluation of association panel over years. VS and MSV genotyped the panel. SNC and AC: formal analysis and drafting the manuscript. SR edited the final version of the manuscript. All authors read, revised, and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Communicated by: Izabela Pawłowicz
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Spoorthi, V., Ramesh, S., Sunitha, N.C. et al. Genetic dissection of green pod yield in dolichos bean, an orphan vegetable legume, using new molecular markers. J Appl Genetics 65, 429–438 (2024). https://doi.org/10.1007/s13353-024-00865-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-024-00865-0