Abstract
Biliary atresia (BA) is a progressive inflammatory process of the biliary tree resulting in biliary obstruction. No single known genetic or environmental factor has been established to cause BA. Cystic fibrosis (CF) is a rare cause of neonatal cholestasis, and it has never been described in familial BA cases. Here, we investigate two siblings of first-degree consanguineous parents presenting with neonatal BA. Shortly after the Kasai operation, the proband developed severe respiratory symptoms attributable to a missed CF diagnosis. This was discovered after re-investigating the family history, which revealed a first-degree cousin with CF who did not manifest BA. Afterwards, we identified a pathogenic variant (DeltaF508) in CFTR in both BA-affected siblings along with their cousin. This intrigued us to study the molecular etiology behind the familial BA presentations, which exclusively contributed to BA-pathogenesis in BA-CF-affected siblings and not in their CF-only affected cousin. We applied a multistep approach to investigate the variant profile of both siblings’ and their cousin’s exomes. We curated the genes whose variants were shared by the BA-CF siblings but absent or heterozygous in their CF-only-affected cousin. Consequently, we identified three candidate genes (SNAPC4, UCK1, and ZHX2) besides CFTR. We propose that these genes act cumulatively or individually in inducing BA-pathogenesis—either by aggravating the biliary damage in the context of CF or increasing the susceptibility of BA as a separate CF-comorbidity. To our knowledge, this is the first report of DeltaF508 in CFTR with familial neonatal BA cases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biliary atresia (BA) is a necro-inflammatory disease of the biliary tree that progresses to fibro-obliteration and biliary obstruction during the neonatal period (Haber and Russo 2003). BA is the leading cause of pediatric hepatic transplantation worldwide. The worldwide BA incidence ranges between 1 per 10,000 and 1 per 20,000 (Ruuska et al. 2019), while the incidence in the Middle East was not estimated.
The pathogenesis of BA is not well understood. Viral infections, immune dysregulations, and toxic exposures were proposed in inducing BA; nevertheless, inconsistencies in each of those proposed models were recorded (Vij and Rela 2020). A few chromosomal aberrations and other congenital anomalies were linked to inducing BA (M. Sergi 2019).
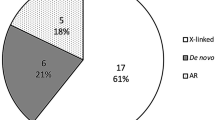
Familial BA recurrences were recorded. Both autosomal recessive (AR) and dominant modes of inheritance were described (Lachaux et al. 1988; Gunasekaran et al. 1992). These findings supported the hypothesis of BA’s genetic propensity. Despite decades of research into the genetic basis of BA, no compelling evidence for a single genetic etiology implicated in BA pathogenesis has been established.
Cystic fibrosis (CF) is an AR multisystemic disorder with an incidence rate of 1 per 2500 in Caucasians and 1 per 2560 in Jordan (Nazer 1992; Kerem 1996). Around 70–90% of CF cases are caused by the DeltaF508 variant in CFTR gene (Ooi and Durie 2016). The main CF manifestations are present in the lungs, pancreas, and gastrointestinal tract. Uncommonly, a few CF cases developed biliary obstructions; however, the majority of them can be either asymptomatic or incidental (Greenholz et al. 1997; Sheth et al. 2003; Pall et al. 2007; Ooi and Durie 2016).
Here, we studied a consanguineous family consisting of two siblings with BA and CF and their cousin, who is only affected by CF without developing BA. We proposed that the BA pathogenesis could have been caused by one of the following hypotheses: (i) The presence of one or more modifier gene(s) increase the risk of developing BA as an atypical manifestation of CF. (ii) BA could be a CF co-comorbidity emerged by bi-allelic causative variants only shared by the CF-BA-affected siblings. Therefore, we explored the genetic background of those familial BA cases by contrasting their whole-exome sequence (WES) data using different tiered approaches. We identified three candidate genes with the potential to play a role in inducing BA pathogenesis.
Materials and methods
Enrollment and clinical evaluation
The Jordan University of Science and Technology (JUST) Research Committee and the Faculty of Medicine’s institutional review board (IRB) approved this study, which is in concordance with the Declaration of Helsinki’s principles (approval code: 27/112/2018; approval date: 31/8/2018). A consanguineous family consisting of three patients with BA and/or CF was recruited to participate in this study (Fig. 1). Prior to the enrolment, the participant (IV-4) and the legal guardian of the siblings (IV-1 and IV-2) signed written informed consents. Unfortunately, the patient (IV-1) died prior to our study at the age of 7 months (Fig. 1). Peripheral blood samples were withdrawn using EDTA tubes from IV-1 and IV-4 at the King Abdullah University Hospital’s pediatric department, Irbid, Jordan.
The pedigree of the participating family. Two siblings (IV-1, and IV-2) of consanguineous parents presented with both cystic fibrosis (CF; filled symbols) and biliary atresia (BA + sign), while their cousin (IV-4) only presented with CF without BA (BA-). Roman numbers, the generation number; + BA, affected by biliary atresia; – BA, unaffected by biliary atresia; filled symbols, affected by cystic fibrosis; white symbols, unaffected by cystic fibrosis; square, male; circle, female
Histopathology evaluation
The glass slides of the liver biopsies from the two patients (IV-1, and IV-2; Fig. 1) were retrieved from the Histopathology Department archives at Jordan University Hospital and reviewed by a gastrointestinal histopathologist. These slides were stained with routine hematoxylin and eosin (H&E) stain as well as a special liver stain panel performed routinely in the Histopathology Department. These include Periodic Acid Schiff (PAS) stain and PAS-diastase to assess glycogen content and alpha-1 antitrypsin globules, iron stain, orcein stain for copper, reticulin for liver architecture, and Masson trichrome for fibrosis.
DNA isolation and WES
gDNA was isolated from IV-1 and IV-4 using QIAprep Spin Mini-prep Kit. For the deceased older sibling (IV-2), the block of liver biopsy was retrieved, and DNA was extracted by QIAamp DNA FFPE Tissue Kit. The DNA extraction protocols were performed according to the manufacturers’ instructions. DNA from the proband (IV-1), her brother (IV-2), and their cousin (IV-4) underwent WES as described by Azab et al. (Azab et al. 2020) with minor differences—the sequence reads were mapped to the NCBI reference sequence (GRCh38) by bwa-0.7.17. Variants’ calling was generated utilizing GATKv4.0.5.1. The following databases were used for variants’ annotation: dbSNP version 151 (https://www.ncbi.nlm.nih.gov/snp/), Clinvar 07/2018 (https://www.ncbi.nlm.nih.gov/clinvar/), 1000Genome Phase3 (https://www.internationalgenome.org/1000-genomes-browsers/index.html), ESP version ESP6500SI_V2 (https://evs.gs.washington.edu/EVS/), and dbNSFP version dbNSFPv3.5c (http://database.liulab.science/dbNSFP). The chromosomal coordinates of the variants have been remapped to GRCh37/hg19 using the Broad Institute LiftOver tool (https://liftover.broadinstitute.org).
Variants’ filtration and prioritization
In order to identify the candidate disease-associated variants, we followed a multistep filtration approach using Variant Annotation and Filter Tool (VarAFT; version 2.17–1) (Desvignes et al. 2018). Rare variants with a minor allele frequency (MAF) of ≤ 1%, as well as variants potentially impacting the protein structure (i.e., missense and loss-of-function variants) in the exonic and the flanking regions, were chosen to be further investigated. The WES analysis was performed utilizing two approaches: focused-gene list filtering and all-genes analysis. Our first-tiered approach was conducted by filtering the variants based on cholestasis/BA-related genes. These genes were compiled from relevant literature and HGMD database (Table S1). In our second-tiered approach, we compared the exomes of the three sequenced individuals (IV-1, IV-2, and IV-4) according to the workflow depicted in Fig. 2.
Results
Overview of clinical history
A 7.5-month-old girl (proband IV-1; Fig. 1) was born to a Syrian consanguineous parent (first-degree cousins) through cesarean section at term, due to breech presentation. At the age of 2 months, she (IV-1) presented with persistent jaundice and a clay-colored stool and was diagnosed with an extrahepatic BA. Consequently, the patient (IV-1) underwent Kasai operation at the age of 2.5 months. Upon her regular follow-ups, she was found jaundiced and failing to thrive. As a result of the suspicion of a failed Kasai procedure, she was referred to the pediatric hepatology clinic for the evaluation of liver transplantation.
The proband (IV-1) had a positive family history of BA as her consanguineous parents gave her two siblings. One of whom (IV-2) had been diagnosed with BA, underwent Kasai operation, and had, unfortunately, died postoperatively secondary to massive bleeding at the age of seven months. The other sibling (IV-3) is in good health.
The mother stated that her child (the proband; IV-1) was feeding well on amino acid formula with no vomiting. She had a frequent, loose, yellow-green, oily, and foul-smelling stool. The mother denied any dark urine. She required frequent hospitalizations over the last few months due to recurrent cough and significant wheezes and was recognized as having reactive airway disease.
At presentation at 7.5 months, her physical examination showed non-dysmorphic, pale, awake, with mild jaundice. She looked malnourished, and her growth parameters were: length 60 cm (< 3rd centile), weight 4.5 kg (< 3rd centile), and head circumference 39 cm (< 3rd centile). She also had mild respiratory distress. Her chest showed expanded costochondral junctions (rosary beads), scattered wheezes, and some coarse crepitations. She had a subcostal incision of previous Kasai, complicated by an incisional hernia. The liver edge was palpable but not the spleen. No significant sacral or lower limb edema was noticed.
Her liver panel showed direct hyperbilirubinemia, transaminitis, and prolonged prothrombin time (Table 1). The patient started vitamin supplementation (A, D, E, and K), and ursodiol at a dose of 20 mg/kg divided into two doses. In addition, MCT oil was added to her feeding routine. After 2 weeks, the patient was re-evaluated and showed minimal weight gain, but her respiratory symptoms worsened. She required multiple visits to the emergency room for nebulizers and was discharged on antibiotics.
Two months later, due to the patient’s respiratory manifestations and her minimal improvement in treatment, the suspicion of having another pathology, namely, CF, was raised. However, her initial workup showed normal sweat chloride levels. Thus, the case was thoroughly re-evaluated, and the family’s medical history was retaken and revealed a first-degree cousin (IV-4; Fig. 1) with CF (sweat chloride test of 128 mmol/L; reference range < 60 mmol/L). On the contrary, the cousin (IV-4) did not develop BA. Moreover, her brother (IV-2), prior to his death, showed high levels of sweat chloride; nevertheless, he did not show any respiratory symptoms.
Therefore, sweat chloride testing was re-ordered for the proband (IV-1) and returned as 113 mmol/L. The patient started on pancreatic replacement therapy (PRT) and respiratory support therapy with chest physiotherapy and home nebulizers. Her weight started to improve. Also, her respiratory symptoms improved significantly. Her liver enzymes stayed elevated, although her bilirubin normalized (Table 1).
Histopathological analysis of the liver
The histopathological slides of the two liver wedge biopsies obtained from the two affected siblings (IV-1, and IV-2) showed essentially similar histopathological features. The sections exhibited preserved liver architecture with normal cell plates composed of one or two cell layers. The proband’s (IV-1) liver histopathology; the portal triads were expanded by a mixed chronic inflammatory infiltrate (Fig. 3A and Fig. 4A) along with bile ductular proliferation, which was more pronounced in the proband’s brother (IV-2) (Fig. 4B) and associated with marked bile plugging (Fig. 4C). This ductular proliferation was less evident in the proband on the H&E section, and as such, a CK7 stain was used to highlight the biliary epithelium (Fig. 3B). Iron stain demonstrated increased iron deposition in macrophages (Fig. 3F and Fig. 3G), a feature seen in cholestasis-associated changes. There were no granulomas and no alpha-1 antitrypsin globules. These features were consistent with an extrahepatic BA associated with fibrosis. As mentioned, these histological features were more pronounced in the proband’s brother (IV-2), in whom the biopsy was taken at 6 months of age, whereas in the proband (IV-1), the biopsy was taken at 2 months of age. The increased severity of the histological findings reflects the fact that the features and clinical symptoms of BA worsen with time.
A sample of liver biopsy for proband IV-1 shows features of extra-hepatic biliary atresia with associated fibrosis. A Expanded portal triads by a mixed chronic inflammatory infiltrate, H&E 200 × . B Bile ductular reaction, CK 7 stain 200 × . C Feathery degeneration of hepatocytes, H&E 200 × . D Giant cells, H&E 400 × . E Periportal bridging fibrosis, Masson Trichrome stain 200 × . F Iron deposition, Pearl’s Iron Stain 200 × . G Spotty necrosis, H&E 400
A sample of liver biopsy for the proband’s older brother (IV-2) shows more extensive features of extra-hepatic biliary atresia with fibrosis. A Portal inflammation, H&E 200 × . B Hepatic ductular proliferation and bile plugs formation, H&E stain 20 × . C Ductular reaction, H&E 400 × . D Feathery degeneration, H&E 200 × . E Giant cells, H&E 400 × . F Extensive Fibrosis, Masson Trichrome 200 × . G Iron deposition, Pearl’s iron stain
Genetic analysis
In pursuit of seeking the cause of the patients’ pathologic picture, WES was performed to investigate possible underlying genetic etiology. In all the CF-affected patients (IV-1, IV-2, and IV-4), we identified a disease-causing variant (DCV) (c.1521_1523delCTT; p.Phe508del) in CFTR (Table 2; Fig. S1). Notably, the variant (p.Phe508del) in CFTR has never been previously discovered in any neonatal BA cases. Also, their CF-only-affected cousin, harboring the same CFTR variant, did not develop BA (Fig. S1B). Therefore, we hypothesized that there might be other genetic variants that might drive the BA pathogenesis in the BA-CF-affected siblings.
To uncover the underlying etiology of BA, our first-tiered approach was to filter the two BA-affected siblings’ exomes (IV-1, and IV-2; Fig. 1) for any candidate disease-causing variants in genes previously reported to be associated with cholestasis and/or BA (Table S1), but no candidate variants were found.
Then, as our second-tiered approach, we attempted to compare the WES data of the three CF-affected patients. This was premised on the possibility that the familial neonatal BA presentation in the two BA-CF-affected siblings would be induced by a unique genetic profile that is lacking in their BA-free, CF-only-affected cousin (Fig. 2). First, we filtered the homozygotes in both BA-CF-affected siblings that are either absent or heterozygous in the BA-free, CF-only-affected cousin (Tables S2-S5). Then, we searched for any candidate genes that have possible combinations of compound heterozygotes that are present in both siblings, but not in their cousin (Tables S6-S8). Thereafter, we compiled a total of 77 genes that fulfill the aforementioned criteria (Table S9). After thoroughly curating the collated sib-shared genes (Fig. 2), we identified three candidate genes that could play a role in the patients’ BA pathogenesis (SNAPC4, UCK1, and ZHX2). Each of these genes harbors a rare candidate homozygous variant that is shared by the BA-CF-affected siblings but absent or heterozygous in their CF- affected cousin, who does not have BA (Table 2; Figs S2-S4).
Discussion
In this study, two siblings of first-degree consanguineous parents presented with neonatal BA. Establishing a BA clinical diagnosis is challenging, owing to several overlapping causes of cholestasis that mimic the BA features (Haber and Russo 2003). Liver biopsy is considered the “gold-standard” for revealing the differential diagnosis (Vij and Rela 2020). Here, liver biopsies were performed for both BA-affected siblings. Their histopathology findings showed ductular proliferation, bile plugs’ formation, and periportal fibrosis. These features are exclusively evident in BA cases (Russo et al. 2011). Nonetheless, the histologic findings were more pronounced in the proband’s older brother (IV-2; 6 M.O) than in the proband herself (IV-1; 2 M.O)—implying that BA presentation can deteriorate over time.
Shortly after Kasai surgery, the proband failed to thrive, which was initially attributed to a failed Kasai procedure. The proband followed aggressive nutritional management, consisting of a high-caloric formula. This treatment is usually used to overcome the excessive caloric needs of malabsorption (Sundaram et al. 2017). Despite slightly gaining weight, the proband experienced unexplained pulmonary symptoms in addition to her persistent failure to thrive. Hence, the suspicion of her suffering from CF was raised, albeit her sweat chloride test was normal. After thoroughly re-taking the family history, her cousin’s (IV-4) CF incident was revealed. Consequently, the proband began a CF treatment regimen, and her respiratory symptoms improved, indicating that she also suffered from CF. The proband’s CF diagnosis was later confirmed with an elevated sweat chloride test. The proband’s proper diagnosis of CF was missed due to an initial false-negative sweat chloride test. In compliance, previous CF cases also had initially normal results (Stewart et al. 1995). Notably, the proband’s brother also showed elevated levels of sweat chloride; however, he did not develop any CF-related manifestations prior to his death.
After utilizing genetic testing to study the three CF-affected relatives (IV-1, IV-2, and IV-4), we identified a shared pathogenic homozygous variant in CFTR (p.Phe508del; formerly known as DeltaF508). While it is the most identified DCV in CF-patients (https://cftr2.org), it has never been reported in any neonatal BA cases. This raised our suspicion of other molecular etiologies driving the BA pathogenesis in two of the three CF-affected individuals.
The determination of the underlying etiology of BA is obscure. In concordance with our study, albeit rare, familial BA occurrences have been previously described, suggesting the presence of a predisposing genetic background (Smith et al. 1991). Recent advances in next-generation sequencing (NGS) have aided in the identification of novel putative genes associated with BA. Nevertheless, due to the paucity of familial cases, and the complexity of the disease, identifying BA-causative genes is yet-to-be-established (Girard and Panasyuk 2019).
Although CF might lead to secondary cholestasis at advanced stages of the disease, CF has rarely been described as causing BA in infancy and has never been reported in BA-familial cases (Vij and Rela 2020). Therefore, the presence of BA can be an atypical presentation of CF. Since the two BA-CF-affected siblings in our study are descending from consanguineous marriage and the absence of BA in their CF-affected cousin (IV-4), the suspicion of other genetic contributing factors (besides the pathogenic variant in CFTR) was raised. This prompted us to study the underlying genetic background of BA. We hypothesized that one of the two following scenarios could have driven the pathogenesis of BA: the presence of CF with one or more mutated modifier gene(s) increased the susceptibility to developing BA as an atypical CF presentation. On the other hand, the BA could have been exerted as an independent CF co-morbidity, triggered by other causative variants in one or more genes.
Consanguinity increases the inheritance of AR disorders (Hamamy 2012; Shawky et al. 2013; Bhinder et al. 2019). Therefore, to explore our hypotheses, we filtered the genes with a possible bi-allelic genotype that are only shared between the BA-CF-affected siblings’ (IV-1, and IV-2) exomes and are absent or heterozygous from their BA-free CF-affected cousin’s (IV-4) exome (Fig. 2). Afterwards, we curated the list of sib-shared genes (Table S9) and identified three candidate genes to be associated with BA (SNAPC4, UCK1, and ZHX2; Figs S1-S4). Nonetheless, this does not rule out the possibility of other heterozygous or intronic modifying variants being involved in driving BA pathogenesis. Further studies are needed to explore the validity of these possibilities.
We identified a novel missense variant (p.Gln217Arg) in SNAPC4 (Table 2; Fig. S2). The product of this gene, SNAPC4, is the largest subunit of at least five snRNA-activating protein complex (SNAPc) subunits, which is essential for activating the transcription of both RNA II and III snRNA genes. SNAPC4 serves as an assembly for the other SNAPc subunits (Ma and Hernandez 2001). The expression of SNAPC4 is ubiquitous; nevertheless, it is moderately expressed in the liver, gallbladder, and gastrointestinal tract (https://www.proteinatlas.org).
In an in vivo zebrafish study by Schaub et al., mutant snapc4 caused intrahepatic bile duct degradation. Initially, the bile ducts underwent complete differentiation. Afterwards, they were subjected to apoptosis and vanished (Schaub et al. 2012). Nevertheless, the gene’s impact on the extrahepatic biliary network remains to be elucidated. Interestingly, biliary cells’ apoptosis has been explored as a disease-causing mechanism inducing BA pathogenesis (Erickson et al. 2008). The clinical implications of SNAPC4 in humans have also been described. For instance, a homozygous missense variant in SNAPC4 has been reported in an adolescent patient with benign recurrent intrahepatic cholestasis (IC) (Munther Odeh et al. 2018). Also, downregulation of SNAPC4 expression was detected in women with IC of pregnancy (Floreani et al. 2013). Taken together, SNAPC4 might have contributed to BA pathogenesis in both BA-siblings. Perhaps by encouraging the intrahepatic cholangiocyte apoptosis and hindering the secretion of bile salts from the hepatocytes.
The second candidate gene is UCK1. We identified a homozygous frameshift variant (p.His103AlafsTer19; Table 2; Fig. S3) in BA-CF-affected siblings and not in their CF-only cousin (IV-4). UCK1 plays a role in the nucleotide metabolism pathway. This gene has two isoforms; one is ubiquitously expressed, while the other is enriched in the kidney, heart, liver, and skeletal muscle (Van Rompay et al. 2001).
Noteworthy, frameshift variants in UCK1 have been discovered in 18 patients with neonatal sclerosing cholangitis (NSC) (Grammatikopoulos 2015). NSC is a serious hepatic disease that prompts inflammation and multi-focal bile duct structuring (Amedee-Manesme et al. 1987; Karlsen et al. 2017). Even though the NSC’s clinical presentation may share some similarities with BA, both can be distinguished by cholangiography and/or liver biopsy (Amedee-Manesme et al. 1987; Lindor et al. 2015). NSC is frequently associated with unique hepatic histopathological features such as “onion skin” fibrosis and periductal fibrosis (Karlsen et al. 2017). Neither of the findings was evident in our BA cases (Fig. 2 and Fig. 3). Instead, UCK1 might have rather induced the development of cholangiopathy in both siblings, leading to their BA presentation.
The third candidate contributing variant in the BA-CF siblings (p.Val388Ile) was in ZHX2 (Table 2; Fig. S4). This gene is a transcription repressor that is known to regulate the expression of several hepatic genes, including alpha-fetoprotein, H19, and glypican 3. ZHX2’s targeted genes are active throughout the perinatal period and become suppressed after delivery. These genes are reactivated during some conditions, such as hepatic development and cancer (Morford et al. 2007).
H19 is involved in the process of bile homeostasis and its expression is upregulated in a variety of liver diseases in both human and mouse models (Li et al. 2018). The elevated H19 expression in liver biopsies retrieved from BA patients was associated with increased BA severity (Xiao et al. 2019). Overexpression of H19 seems to induce essential bile acid receptor signaling, thus leading to bile acid accumulation, cholestatic injury, and cholangiocyte proliferation (Li et al. 2018; Xiao et al. 2019). Moreover, H19 knockout mice had inhibited cholangiocyte proliferation along with reduced liver injury and inflammation (Xiao et al. 2019; Tian et al. 2021). Our BA-CF-patients harbor a variant in ZHX2 that is known to suppress H19 expression, possibly leading to abnormal H19 upregulation that can ultimately induce cholangiocyte proliferation, a hallmark of BA-presentation. The maximum MAF of this variant (p.Val388Ile) in the population database gnomAD is 0.7%, with 5 reported homozygotes. Even though this variant’s frequency is rare in the population, it is deemed too common to be disease-causing in BA. Also, the high number of reported homozygotes reduces the likelihood of this variant being a candidate in BA. However, this variant in ZHX2 might have played a role in increasing the susceptibility to BA development along with the other two variants identified in SNAPC4 and UCK1.
In conclusion, we presented familial BA-CF-cases descending from first-degree consanguineous parents. Despite having a positive CF family history, their BA-free, CF-only affected cousin did not develop BA. This raised the possibility of other shared genetic etiologies between the BA-CF-affected siblings, absent from their BA-free CF-cousin, contributing to the familial BA pathogenesis. We applied multistep filtering approaches to find the genes with variants exclusively found in the BA-CF-affected siblings and not in their CF-only affected cousin. We curated the list of the sib-shared genes and consequently identified three candidate genes (SNAPC4, UCK1, and ZHX2). Based on previous evidence, the described genes can be implicated in inducing BA pathogenesis. We propose that these genes might have cumulatively or individually contributed to accelerating the biliary damage in the setting of CF. Alternatively, these genes might have increased the susceptibility to BA pathogenesis as a separate CF-comorbidity. Our research paves the way for future studies to explore the described genes as candidate molecular etiologies of BA. To the best of our knowledge, this is the first study to report the DeltaF508 variant in CFTR with familial neonatal BA presentation.
Availability of data and materials
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Amedee-Manesme O, Bernard O, Brunelle F et al (1987) Sclerosing cholangitis with neonatal onset. J Pediatr 111:225–229. https://doi.org/10.1016/S0022-3476(87)80072-0
Azab B, Dardas Z, Rabab’h O, et al (2020) Enteric anendocrinosis attributable to a novel Neurogenin-3 variant. Eur J Med Genet 63:103981. https://doi.org/10.1016/j.ejmg.2020.103981
Bhinder MA, Sadia H, Mahmood N et al (2019) Consanguinity: A blessing or menace at population level? Ann Hum Genet 83:214–219. https://doi.org/10.1111/AHG.12308
Desvignes JP, Bartoli M, Delague V et al (2018) VarAFT: A variant annotation and filtration system for human next generation sequencing data. Nucleic Acids Res 46:W545–W553. https://doi.org/10.1093/nar/gky471
Erickson N, Mohanty SK, Shivakumar P et al (2008) Temporal-spatial activation of apoptosis and epithelial injury in murine experimental biliary atresia. Hepatology 47:1567–1577. https://doi.org/10.1002/hep.22229
Floreani A, Caroli D, Lazzari R et al (2013) Intrahepatic cholestasis of pregnancy: New insights into its pathogenesis. J Matern Neonatal Med 26:1410–1415. https://doi.org/10.3109/14767058.2013.783810
Girard M, Panasyuk G (2019) Genetics in biliary atresia. Curr Opin Gastroenterol 35:73–81
Grammatikopoulos T (2015) Genetic investigation of neonatal sclerosing cholangitis. King’s College London
Greenholz SK, Krishnadasan B, Marr C, Cannon R (1997) Biliary obstruction in infants with cystic fibrosis requiring Kasai portoenterostomy. In: Journal of Pediatric Surgery. J Pediatr Surg, pp 175–180
Gunasekaran TS, Hassall EG, Steinbrecher UP, Yong SL (1992) Recurrence of extrahepatic biliary atresia in two half sibs. Am J Med Genet 43:592–594. https://doi.org/10.1002/ajmg.1320430317
Haber BA, Russo P (2003) Biliary atresia. Gastroenterol Clin North Am 32:891–911
Hamamy H (2012) Consanguineous marriages: Preconception consultation in primary health care settings. J Community Genet 3:185. https://doi.org/10.1007/S12687-011-0072-Y
Karlsen TH, Folseraas T, Thorburn D, Vesterhus M (2017) Primary sclerosing cholangitis – a comprehensive review. J Hepatol 67:1298–1323
Kerem B (1996) The molecular basis for disease variability in cystic fibrosis. Eur J Hum Genet 4:65–73
Lachaux A, Descos B, Plauchu H et al (1988) Familial extrahepatic biliary atresia. J Pediatr Gastroenterol Nutr 7:280–283. https://doi.org/10.1097/00005176-198803000-00020
Li X, Liu R, Huang Z et al (2018) Cholangiocyte-derived exosomal long noncoding RNA H19 promotes cholestatic liver injury in mouse and humans. Hepatology 68:599–615. https://doi.org/10.1002/hep.29838
Lindor KD, Kowdley KV, Harrison ME (2015) ACG clinical guideline: Primary sclerosing cholangitis. Am J Gastroenterol 110:646–659. https://doi.org/10.1038/ajg.2015.112
M. Sergi C (2019) Genetics of Biliary Atresia: A Work in Progress for a Disease with an Unavoidable Sequela into Liver Cirrhosis following Failure of Hepatic Portoenterostomy. In: Liver Cirrhosis - Debates and Current Challenges. IntechOpen
Ma B, Hernandez N (2001) A Map of Protein-Protein Contacts within the Small Nuclear RNA-activating Protein Complex SNAPc. J Biol Chem 276:5027–5035. https://doi.org/10.1074/jbc.M009301200
Morford LA, Davis C, Jin L et al (2007) The oncofetal gene Glypican 3 is regulated in the postnatal liver by zinc fingers and homeoboxes 2 and in the regenerating liver by alpha-fetoprotein regulator 2. Hepatology 46:1541–1547. https://doi.org/10.1002/hep.21825
MuntherOdeh A, Safieh YA, Kamal L et al (2018) A Novel Small Nuclear RNA-Activating Protein (SNPC4) Gene Mutation is Associated with Benign Recurrent Intrahepatic Cholestasis (BRIC). EC Gastroenterol Dig Syst 9:741–745
Nazer HM (1992) Early diagnosis of cystic fibrosis in Jordanian children. J Trop Pediatr 38:113–115. https://doi.org/10.1093/tropej/38.3.113
Ooi CY, Durie PR (2016) Cystic fibrosis from the gastroenterologist’s perspective. Nat Rev Gastroenterol Hepatol 13:175–185. https://doi.org/10.1038/nrgastro.2015.226
Pall H, Zielenski J, Jonas MM et al (2007) Primary Sclerosing Cholangitis in Childhood is Associated with Abnormalities in Cystic Fibrosis-Mediated Chloride Channel Function. J Pediatr 151:255–259. https://doi.org/10.1016/j.jpeds.2007.03.062
Russo P, Magee JC, Boitnott J, et al (2011) Design and Validation of the Biliary Atresia Research Consortium Histologic Assessment System for Cholestasis in Infancy. ClinGastroenterol Hepatol 9 https://doi.org/10.1016/j.cgh.2011.01.003
Ruuska SM, Lääperi MT, Hukkinen M et al (2019) Growth of children with biliary atresia living with native livers: impact of corticoid therapy after portoenterostomy. Eur J Pediatr 178:341–349. https://doi.org/10.1007/s00431-018-3302-z
Schaub M, Nussbaum J, Verkade H et al (2012) Mutation of zebrafish Snapc4 is associated with loss of the intrahepatic biliary network. Dev Biol 363:128–137. https://doi.org/10.1016/j.ydbio.2011.12.025
Shawky RM, Elsayed SM, Zaki ME et al (2013) Consanguinity and its relevance to clinical genetics. Egypt J Med Hum Genet 14:157–164. https://doi.org/10.1016/J.EJMHG.2013.01.002
Sheth S, Shea JC, Bishop MD et al (2003) Increased prevalence of CFTR mutations and variants and decreased chloride secretion in primary sclerosing cholangitis. Hum Genet 113:286–292. https://doi.org/10.1007/s00439-003-0963-z
Smith BM, Laberge JM, Schreiber R et al (1991) Familial biliary atresia in three siblings including twins. J Pediatr Surg 26:1331–1333. https://doi.org/10.1016/0022-3468(91)90613-X
Stewart B, Zabner J, Shuber AP et al (1995) Normal Sweat Chloride Values Do Not Exclude the Diagnosis of Cystic Fibrosis. Am J Respir Crit Care Med 151:899–903. https://doi.org/10.1164/ajrccm/151.3_pt_1.899
Sundaram SS, Mack CL, Feldman AG, Sokol RJ (2017) Biliary atresia: Indications and timing of liver transplantation and optimization of pretransplant care. Liver Transplant 23:96–109
Tian X, Wang Y, Lu Y et al (2021) Conditional depletion of macrophages ameliorates cholestatic liver injury and fibrosis via lncRNA-H19. Cell Death Dis 12:1–13. https://doi.org/10.1038/s41419-021-03931-1
Van Rompay AR, Norda A, Lindén K et al (2001) Phosphorylation of uridine and cytidine nucleoside analogs by two human uridine-cytidine kinases. Mol Pharmacol 59:1181–1186. https://doi.org/10.1124/mol.59.5.1181
Vij M, Rela M (2020) Biliary atresia: Pathology, etiology and pathogenesis. Futur. Sci. OA 6
Xiao Y, Liu R, Li X et al (2019) Long Noncoding RNA H19 Contributes to Cholangiocyte Proliferation and Cholestatic Liver Fibrosis in Biliary Atresia. Hepatology 70:1658–1673. https://doi.org/10.1002/hep.30698
Funding
This work was supported by Deanship of Research of Jordan University of Science and Technology, Irbid, Jordan (Grant number 83/2018).
Author information
Authors and Affiliations
Contributions
Conceptualization, B.A, and E.A. Data curation, D.A., Z.D., and L.S. Formal analysis, D.A., O.R., and H.A. Funding acquisition, E.T. Methodology, O.R., D.A., Z.D., and L.S. Project administration, B.A., and E.A. Supervision, B.A., and E.A. Validation, B.A., H.A., and E.A. Visualization, D.A., O.R., Z.D., and L.S. Writing—original draft, B.A., D.A., and E.A. Writing—review and editing, B.A., O.R., L.A., H.A., and E.A. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Ethical approval
The Jordan University of Science and Technology (JUST) Research Committee and the Faculty of Medicine's institutional review board (IRB) approved this study which is in concordance with the Declaration of Helsinki's principles (approval code: 27/112/2018; approval date: 31/8/2018).
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Consent to publish
The authors affirm that human research participants provided informed consent for publication of the images in Figs. 3 and 4.
Competing interests
The authors declare no competing interests.
Additional information
Communicated by Michal Witt.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13353_2022_729_MOESM1_ESM.pdf
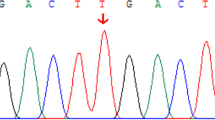
Supplementary file1 Fig. S1. NGS pile-ups of the variant (c.1521_1523delCTT; p.Phe508del) in CFTR. a Homozygous deletion of three nucleotides in CF-BA affected sibling (IV-1). b Homozygous deletion of three nucleotides in CF-only affected cousin (IV-4) (PDF 478 KB)
13353_2022_729_MOESM2_ESM.pdf
Supplementary file2 Fig. S2. NGS pile-ups of the variant (c.650A > G p.Gln217Arg) in SNAPC4. a Homozygous substitution of c.650A to G in CF-BA affected sibling (IV-1). b Heterozygous substitution of c.650A to G in CF-only affected cousin (IV-4). The shown chromosomal coordinates are on hg38 (PDF 334 KB)
13353_2022_729_MOESM3_ESM.pdf
Supplementary file3 Fig. S3. NGS pile-ups of the variant (c.306dupG; p.His103AlafsTer19) in UCK1. a Homozygous duplication of one nucleotide (c.306G) in CF-BA affected sibling (IV-1). b Heterozygous duplication of one nucleotide (c.306G) in CF-only affected cousin (IV-4). The shown chromosomal coordinates are on hg38 (PDF 305 KB)
13353_2022_729_MOESM4_ESM.pdf
Supplementary file4 Fig. S4. NGS pile-ups of the variant (c.1162G > A; p.Val388Ile) in ZHX2. a Homozygous substitution of c.1162G > to A in CF-BA affected sibling (IV-1). b No variant was detected at c.1162G in CF-only affected cousin (IV-4). The shown chromosomal coordinates are on hg38 (PDF 266 KB)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Altamimi, E., Rabab’h, O., Aburizeg, D. et al. Investigating the genetic profile of familial atypical cystic fibrosis patients (DeltaF508-CFTR) with neonatal biliary atresia. J Appl Genetics 64, 71–80 (2023). https://doi.org/10.1007/s13353-022-00729-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-022-00729-5