Abstract
Cotton that serves natural fiber for the textile industry is an important industrial crop. However, abiotic stress imposed a significant negative impact on yield and quality of cotton fiber. Carotenoid cleavage oxygenases (CCOs) that specifically catalyze the cleavage of carotenoid are essential for plant growth and development and abiotic stress response. While information of cotton CCOs and their potential functions in abiotic stress is still far from satisfactory, which imposes restrictions on application in genetic breeding for stress resistance. In this study, 15, 15, and 30 CCOs were identified from Gossypium arboreum, Gossypium raimondii, and Gossypium hirsutum, respectively. Phylogenetic relationship indicated that CCO genes could be classified into two groups (NCEDs and CCDs). Cis-elements prediction showed that there were 18 types of stress-related cis-elements in promoter regions. Analysis with transcriptome data revealed tissue-specific expression pattern of cotton CCOs. qRT-PCR analysis revealed only that GhNCED3a_A/D and GhNCED3c_A/D had strong response to drought, salt, and cold stress, while GhCCD1_A/D and GhCCD4_A showed relatively slight expression changes. Virus-induced gene silencing of GaNCED3a, the ortholog gene of GhNCED3a_A/D, suggested that silenced plants exhibited decreased resistance not only to drought but also to salt, with significantly reduced proline content, and high malondialdehyde content and water loss rate. In addition, stress response genes RD29A, DREB1A, and SOS1 significantly downregulated under drought and salt stress in silenced plants compared to control plants, indicating that GaNCED3a played an important role in drought and salt response. The results provided valuable insights into function analysis of cotton CCOs in abiotic stress response, and suggested potential benefit genes for stress-resistant breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Carotenoids are compounds that typically composed of several isoprene units and had multiple conjugated double bonds. They are ubiquitous in plants, fungi, bacteria, and animals and performed important biological functions. Carotenoid cleavage oxygenase (CCO) are vital for the metabolism of carotenoids, which specifically catalyzes the cleavage of carotenoid to produce various types of cleavage products. The products known as apocarotenoids and their derivatives possess essential biological functions in plant development and physiological response (Akemi et al. 2009; Walter and Strack 2011; Kloer et al. 2005). Based on whether the substrates are forming epoxy structure, CCO can be further classified into nine-cis-epoxycarotenoid dioxygenases (NCEDs) and carotenoid cleavage dioxygenases (CCDs) subfamily (Giuliano et al. 2003).
The vp14 gene isolated from maize was the first identified CCO gene (Schwartz et al. 1997). Subsequently, Tan et al. (2003) identified a total of nine homologs of vp14 in Arabidopsis thaliana. Four of them encoded CCDs (CCD1, CCD4, CCD7, and CCD8), while five encoded NCEDs (NCED2, NCED3, NCED5, NCED6, and NCED9) (Auldridge et al. 2006a). The orthologs of CCO identified in other plants were then named typically according to orthologs in Arabidopsis (Kim et al. 2016; Wang et al. 2013). Recently, CCD-like, a new subgroup of CCDs, was identified in tomato, strawberry, apple, tobacco, and sugarcane (Chen et al. 2018; Su et al. 2021; Wang et al. 2017; Wei et al. 2016; Zhou et al. 2019). To date, CCO gene family had been comprehensively analyzed in several species (Chen et al. 2018; Kim et al. 2016; Su et al. 2021; Tan et al. 2003; Vallabhaneni et al. 2010; Wang et al. 2013; Wei et al. 2016; Zhao et al. 2021; Zhou et al. 2019).
CCDs cleave a variety of trans-carotenoid substrates (Bouvier et al. 2003; Sun et al. 2008). CCD1 and CCD4 mainly contributed to the cleavage of apocarotenoids to generate compounds related to scent of flowers and fruits (Adami et al. 2013; Rubio et al. 2008; Ilg et al. 2009; Simkin et al. 2004; Song et al. 2016). Wang et al. (2013) reported that CCD1 and CCD4 genes in soybean showed strong responses to abiotic stress. CCD7 and CCD8 take part in metabolism of strigolactone which is involved in axillary shoot growth as well as drought and salt response in plants (Alder et al. 2012; Ha et al. 2014; Kim et al. 2016; Kloer and Schulz 2006; Seo and Koshiba 2002; Umehara et al. 2008). So far, little attention had been paid on specific biological functions of CCD-like genes.
NCEDs are key enzymes that catalyze the rate-limiting step in the biosynthesis of abscisic acid (ABA) (Gavassi et al. 2021; Schwartz et al. 2004; Vishwakarma et al. 2017). Wang et al. (2021) revealed that both PpNCED1 and PpNCED5 genes cooperatively regulate ABA biosynthesis in peach fruits. AcNCED1 might serve a major function in the early fruits softening of kiwifruit (Gan et al. 2020). AtNCED3, an ortholog of the best-characterized maize VP14, played a prominent role in drought tolerance (Tan et al. 2003; Iuchi et al. 2000). Overexpression of TaNCED increased drought resistance and delayed seed germination in Arabidopsis (Tong et al. 2017). NCED3 and NCED5 of rice have taken part in abiotic stress response by regulating ABA content (Huang et al. 2018, 2019). AtNCED6 and AtNCED9 were involved in seed development by regulating ABA biosynthesis (Lefebvre et al. 2006). Overall, NCED genes play an essential role in several biological processes in plants, including seed dormancy, plant growth and reproduction development, and abiotic stress response (Chernys and Zeevaart 2000; Leng et al. 2014).
Cotton, as a kind of important industrial crops for many countries around the world, serves natural fiber for the textile industry. Abiotic stress imposes a significant negative impact on yield and quality of cotton fiber. Genetic engineering provided an effective way for cotton genetic improvement on stress resistance. However, information on cotton CCO genes is still unknown and identification of stress responsive CCO genes has received little attention. In this study, characterization of CCO genes from three cotton species was conducted. Phylogenetic relationship, gene structure, putative cis-elements, collinearity relationship, and gene expression patterns under multiple stress conditions were further comprehensively performed. GaNCED3a, the ortholog gene of multiple stress-responsive gene GhNCED3a_A/D, was selected to perform functional analysis under drought and salt stress by VIGS in G. arboreum. Results of GaNCED3a silencing showed evidence that NCED3 in cotton not only played a role in drought stress but also taken part in salt stress. The results shed light on deep understanding of characteristics of cotton CCO genes and taking advantage of them in cotton stress-responsive genetic improvement.
Materials and methods
Identification of cotton CCO genes
The genomic data of the diploid cotton Gossypium raimondii (D5), Gossypium arboreum (A2), and tetraploid cotton Gossypium hirsutum L. acc. TM-1 (AD1) were downloaded from https://www.cottongen.org/. A hidden Markov model (HMM) profile of CCO domain (Pfam accession number: PF03055) was downloaded from Pfam (El-Gebali et al. 2019) and used to scan the protein databases with HMMER v3.0 (http://hmmer.org/) with E value < 0.01. The screened proteins were further confirmed against SMART (Schultz et al. 1998) and InterPro (Mitchell et al. 2019) databases. The identified cotton CCO genes were named according to their homology with Arabidopsis genes. Physiochemical characteristics were identified using ExPASy on http://www.expasy.org/tools/.
Multiple alignment and phylogenetic analysis
Multiple alignment was performed by DNAMAN software. Phylogenetic trees were conducted by MEGA 7.0 using the neighbor-joining (NJ) method. Sequences of CCO proteins of Arabidopsis thaliana, Zea mays, and Solanum lycopersicum were published by Tan et al. (2003), Vallabhaneni et al. (2010), and Wei et al. (2016).
Structure analysis and cis-element prediction
Motif analysis was carried out with MEME Suite (Bailey et al. 2009) and displayed using TBtools software (Chen et al. 2020). Gene structure was retrieved from GFF3 files of the genomes. The sequences 1500 bp upstream of the translation initiation site were extracted, and cis-elements were predicted by PlantCARE on http://bioinformatics.psb.ugent.be/webtools/plantcare/html/. Gene Structure Display Server (GSDS) 2.0 (Hu et al. 2015) was employed to show gene structure and cis-element distribution.
Chromosomal location and collinearity analysis
Chromosomal distribution of cotton CCO genes was mapped by MapInspect software. Gene duplication was identified using the MCScanX program. The nonsynonymous substitution rates (Ka) to synonymous substitutions rates (Ks) (Yadav et al. 2015) were calculated with TBtools software.
Gene expression pattern analysis of cotton CCOs
Data of the accession codes PRJNA248163 of G. hirsutum L. acc. TM-1 were fetched from SRA databases (Zhang et al. 2015). Various tissues (leaf, stem, root, stamen, petal, ovule of − 3, 0, and 3 DPA (days post-anthesis), fiber of 5, 10, 20, and 25 DPA) and abiotic stress (drought, salt, and cold stress for 1, 3, 6 and 12 h) transcriptome datasets were employed. The fragments per kilobase of exon model per million mapped reads (FPKM) were estimated with cufflinks and normalized to evaluate the expression levels. Heat maps were generated using the TBtools software.
Plant materials and treatments
Seedlings of upland cotton cultivar “Jimian 2016” were cultivated in Hogland solution in the artificial incubator under controlled conditions (25℃ for 16-h light/22℃ for 8-h dark). After 3 weeks, seedlings at trefoil stage were subjected to drought (17% PEG6000), salt (200 mM NaCl), and cold (4℃), respectively. Samples were collected at desired time points and stored at − 80℃. Samples from untreated seedlings were selected as control.
Quantitative real-time PCR analysis
Total RNA extraction, cDNA synthesis, and quantitative real-time PCR (qRT-PCR) were performed according to the manufacturer’s instruction of the RNAprep pure plant kit (TIANGEN, China), the PrimeScript™ RT reagent kit (TaKaRa, China), and the TB Green® Premix Ex Taq™II (TaKaRa, China), respectively. qRT-PCR reactions were conducted with three replicates on CFX96 Real-Time PCR System (Bio-Rad, USA). The histone3 (AF024716) was amplified as internal control. Relative expression levels were evaluated using 2−ΔΔCt method (Livak and Schmittgen 2001). Specific primers designed by Primer-BLAST (http://www.ncbi.nlm.nih.gov/tools/primer-blast/) are listed in Table S1.
Virus-induced gene-silencing assay
GaNCED3a, the ortholog gene of stress-related gene GhNCED3a_A/D, was selected for functional analysis under drought and salt stress by VIGS in G. arboreum. A 300-bp PCR product of GaNCED3a was amplified using cDNA from G. arboreum cv. “Shixiya-1” and inserted to the TRV vector. Primer used for pTRV2:GaNCED3a vector conduction is listed in Table S1. Plasmid with three vectors, pTRV2:00, pTRV2:CLA1, and pTRV2:GaNCED3a, were transformed to Agrobacterium tumefaciens strain GV3101. VIGS injection was performed according to Yang et al. (2019). Samples were collected from plants 14 days after inoculation and qRT-PCR analysis was conducted to evaluate the efficiency of VIGS silencing. Plants with expression level of GaNCED3a less than 40% of control were used to exam phenotypic and physiological variations. TRV2:00-tageted plants were taken as control. At 21 days post-inoculation, plants were subjected to drought and salt stress, and leaves were sampled for malondialdehyde (MDA), proline (PRO), and water loss assay according to the instruction of corresponding kit (Suzhou Comin Biotechnology Co., Ltd.). Water loss was evaluated by weighing the leaves per hour and water loss rate represented the leaf water loss weight to the leaf fresh weight. For drought and salt stress treatment, plants were stopped watering and watered with 300-mM NaCl instead, respectively. Plants irrigated with water were taken as mock.
Results
Identification of cotton CCO members
A total of 15, 15, and 30 CCO members were identified from G. arboreum, G. raimondii, and G. hirsutum, respectively. The number of CCO of tetraploid cotton G. hirsutum was twice of diploid cotton G. arboreum and G. raimondii, which was consistent with their corresponding genomes. Length of encoded amino acid residues of cotton CCOs varied from 353 to 647. Molecular weight of cotton CCOs proteins ranged from 39.00 to 72.13 kDa, and isoelectric point (PI) was distributed from 5.39 to 8.94. Information of cotton CCO genes is detailed in Table 1.
Phylogenetic analysis of CCO proteins
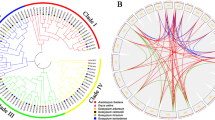
Multiple sequence alignments indicated that CCO proteins in Gossypium showed high sequence identity (Fig. S1). To further analyze the evolutionary relationship of CCO family proteins, a phylogenetic tree was constructed using sequences of CCO from monocot Z. mays, dicot A. thaliana, S. Lycopersicum, and Gossypium (Fig. 1). CCO proteins from six plant species were divided into NCED and CCD groups, and the CCD group was further classified into five subgroups, CCD1, CCD4, CCD-like, CCD7, and CCD8, as previously reported (Chen et al. 2018; Kim et al. 2016).
In NCED group, NCED9 did not exist in cotton and S. lycopersicum. NCED3 in cotton were clustered with AtNCED3 and SlNCED, while NCED5 in cotton were clustered with SlNCED2 but not AtNCED5. Furthermore, NCED6 in cotton were orthologous to AtNCED6 and SlNCED3. Orthologous of NCED5 were not found in maize. Five members of maize NCEDs, ZmNCED1, ZmNCED2, ZmNCED3a, ZmNCED3b, and ZmNCED9, clustered in a single branch, indicating a conservation of NCED sequences in monocot maize. Compared with the fact that other plants contained only one copy of NCED3 and NCED5 (Zhou et al, 2019; Wang et al. 2013; Tan et al. 2003), there are three copies of NCED3 and NCED5 in cotton, NCED3a/3b/3c and NCED5a/5b/5c. This suggested that a duplication event had occurred during the evolution process.
In CCD group, orthologous of CCD-like were found in Gossypium, S. lycopersicum (SlCCD-like), and Z. mays (ZmCCD8b), and were not found in A. thaliana. Obviously, ZmCCD8b were not clustered into CCD8 subgroup but CCD-like subgroup in the present study. There were two CCD4 subgroup members, CCD4a and CCD4b, in Z. mays, S. lycopersicum, and Gossypium. Only one member was found in A. thaliana. These results implied that CCO members in different plant species had great divergency in the evolution process. CCD1, CCD4, and NCEDs had more close relationship in the NJ phylogenetic tree, suggesting their divergence from a common ancestor.
Analysis of gene structure and motif
Exon/intron structures were analyzed to further understand the evolution of CCO gene family in cotton (Fig. 2). NCEDs in cotton shared a simple gene structure. All NCEDs contained only one exon with conserved length except for GrNCED5a and GrNCED5b. While exon number of cotton CCDs varied greatly, with the range of 1 to 15. CCD4 in cotton contained 1–2 exons. CCD 7 and CCD8 contained 6 exons (except for GaCCD8a and GrCCD8a), but with different distribution of exon and intron length. CCD1 contained variable numbers of exons from 12 to 15. CCD-likea and CCD-likeb in cotton shared similar gene structures with different exon numbers and intron lengths. The results revealed that genes grouped in the same clades processed a similar gene structure, indicating a close correlation between the phylogeny and exon/intro structure.
Ten conserved motifs were analyzed by MEME software (Fig. S2, Table S2). The conserved motif in all 60 cotton CCO protein sequence was plotted by TBtools software (Fig. 2). It is clearly that CCOs grouped in the same cluster had similar conserved motifs, which was consistent with the results of the phylogenetic analysis. Cotton NCED3 and NCED5a proteins contained all ten conserved motifs and their relative positions were also conserved. Motif 9 and motif 10 were not found in GhNCED5b_A/D, GhNCED5c_A, and GrNCED5c. Only four motifs were detected in GhNCED5c_D, which might result from the gene fragment loss in the process of evolution or genome assembly errors. Cotton NCED6 protein did not contain motif 7, implying the potential divergence functions of NCEDs in cotton. In cotton CCD subgroup, CCD1, CCD4, and CCD-like had nine conserved motifs except motif 7. Motif 7 were only found in N-terminal of CCD8 members. CCD8 had seven motifs (motif 2–8) and CCD7 had only four motifs (motif 2, motif 5, motif 8, and motif 9). Both CCD7 and CCD8 did not contain motif 1 and motif 10. Motif numbers and distributions of CCD7 and CCD8 were highly different from other CCOs, suggesting that there was a high sequence variation. On the whole, cotton NCEDs contained more conserved motifs than CCDs.
Promoter analysis of cotton CCO genes
According to the potential functions of cotton CCO, cis-elements involved in plant hormone responsiveness and biotic and abiotic stress were analyzed. The distribution of putative cis-elements were drawn by GSDS (Fig. 3, Fig. S3). Eighteen types of cis-elements related to hormone and stress were found in 60 cotton CCO promoter regions. The information is briefly summarized in Table S3. Seven types of stress-related cis-elements including MBS involved in drought stress, LTR, and DRE core that involved in cold response, TC-rich repeats involved in defense and stress response, W box involved in abiotic stress, WUN-motif involved in wound-response, and ARE involved in anaerobic induction were found in the promoters. These results of cis-elements identification implied that cotton CCO genes take a great part in plant multiple stress response. Phytohormones response cis-elements, such as ABA-responsive element (ABRE), ET responsive element (ERE), MeJA-responsive elements (TGACG-motif and CGTCAmotif), SA-responsive element (TCA-element), and gibberellin-responsive element (P-box) were also found abundantly in the promoter sequence. There were 12 out of 15 promoters contained ABA-responsive element (ABRE), indicating cotton CCO’s potential role in ABA-mediated signals. Overall, the diverse cis-regulatory elements in the promoters related to stress and plant hormone may imply their diverse functions in cotton growth.
Chromosomal distribution and synteny analysis
The chromosomal distribution of CCO from three cotton species were determined based on their corresponding genomic information (Fig. 4). The chromosomal distribution of cotton CCOs was uneven. For G. hirsutum, 30 CCOs were distributed on 18 chromosomes and two CCOs (GhNCED5b_A and GhNCED3b_D) were located on two scaffolds. The distribution of CCO on chromosomes of A sub-genome exhibited a good correspondence with those on chromosomes of D sub-genome. There was no CCO gene located on Ga_chr2, Ga_chr4, Ga_chr12, Gr_chr3, Gr_chr5, Gr_chr7, and Gr_chr12. The localization of CCO genes between G. raimondii and G. arboreum did not exhibit a good correspondence, which might be owing to the evolution of different species.
To understand the expansion patterns of cotton CCO genes, gene duplication events were investigated by genome synteny analysis (Fig. 5). Only one pair of tandem duplication cluster (GhCCD-likea_D and GhCCD-likec_D) was identified. One segmental duplication pair was found within the genome of G. raimondii, 14 pairs within G. hirsutum, and 29, 26, and 14 pairs between G. raimondii and G. hirsutum, G. arboreum and G. hirsutum, and G. arboreum and G. raimondii, respectively (Table S4). These results implied polyploidization and segmental duplication were the major processes during gene expansion. The ortholog genes of GaNCED5b and GaCCD-likea were not found in G. hirsutum, while 3 genes in tetraploid cotton (GhNCED5b_A, GhCCD8b_A, and GhCCD-likec_D) had no orthologous gene in the two diploid cottons. These results implied that some CCO genes might undergo gene loss, gain, or rearrangement during the polyploidization process. To explore the selection constraints, the Ka/Ks ratio was calculated (Table S4). Only one pair (GaNCED6 and GhNCED6_A) had a positive pressure (Ka/Ks > 1), while the other pairs experienced a purifying selection process.
Syntenic analysis of CCO genes between G. raimondii, G. arboreum, and G. hirsutum. Green curves denote syntenic regions between G. arboreum and G. raimondii, red curves between G. arboreum and G. hirsutum, blue curves between G. raimondii and G. hirsutum, brown curve within G. raimondii, and orange curve within G. hirsutum
Genomic comparison of different organisms is a way to understand the origins, evolutionary history, and new gene functions (Lyons et al. 2008). Arabidopsis as a model plant had been well characterized, and some CCO genes had been given deep research. To better explore the origin and evolutionary history of CCO genes in G. hirsutum, a syntenic map between G. hirsutum and Arabidopsis was generated (Fig. S4). Sixteen pairs of ortholog between G. hirsutum and Arabidopsis were syntenic genes. Eight pairs of orthologous genes (AtNCED6/GhNCED6_A, AtNCED6/GhNCED6_D, AtNCED5/GhNCED5a_A, AtNCED5/GhNCED5a_D, AtCCD7/GhCCD7_A, AtCCD7/GhCCD7_D, AtCCD4/GhCCD4b_A, AtCCD4/GhCCD4b_D) were single one-to-one correspondence between Arabidopsis and A-subgenome or D-subgenome of G. hirsutum, indicating that they may be derived from a common ancestor. Such results laid foundation on understanding the roles of cotton CCO genes.
Expression profiles of CCO genes in G. hirsutum
Transcription levels of CCO genes in various tissues varied greatly (Fig. 6a). In CCD subgroup, GhCCD1_A/D has shown high expression level in all tested tissues (FPKM > 15). GhCCD4a_A and GhCCD4b_A/D exhibited high expression level in stamen; other CCDs showed high expression in stem. GhNCED3a_A/D exhibited high expression level in root; GhNCED3b_A/D, GhNCED3c_A/D, and GhNCED6_A/D have shown high expression level in leaf, and GhNCED5a_A and GhNCED5c_A/D in petal.
Dynamic changes in expression level of CCO genes in different stages of fiber development were observed (Fig. 6b). GhNCED3c_A/D was high-expressed in fiber initiation stage of 3 DPA ovules and 5 DPA fibers, while GhNCED3b_A/D high-expressed in late stage of fiber development of 25 DPA fibers. The expression profiles of GhNCED3a_A/D were dynamic, which with relatively higher expression in 3 DPA ovules and 25 DPA fibers. Only one CCD member, GhCCD4b_A, had high expression level in 10, 20, and 25 DPA fibers. Other CCD genes showed a sustained low expression level in different fiber developmental stages (FPKM < 1). The specific expression patterns of cotton CCO in fiber implied their potential functions in different fiber developmental stages.
To explore their functions under abiotic stress, a comprehensive analysis of expression profiles under drought, salt, and cold conditions was conducted (Fig. 6c). Notably, GhNCED3a_A/D and GhNCED3c_A/D were upregulated by all imposed stress, and peaked at 12 h after treatments. GhCCD4b_A was induced strongly during the first 6 h and then decreased under drought and salt, while was repressed under cold condition. GhCCD4b_D showed a similar expression pattern with GhCCD4b_A under drought and cold. GhCCD1_A/D were slightly downregulated under all treatments. Almost no expression was detected in the other genes (FPKM < 1).
Seven CCO genes that significantly responded to three abiotic stress were selected for qRT-PCR to validate the differential expression patterns (Fig. 7). The results of qRT-PCR for candidate CCO genes basically coincide with the results from RNA-seq data. Of note, the results indicated the selected NCED genes were upregulated at each time points under various treatments. GhNCED3a_A/D under drought and salt stress were upregulated, with a maximum 11-fold increase. Moreover, GhNCED3c_A/D were strongly induced by all imposed adversity stress and the expression level peaked at 6 h under salt, whereas peaked at 12 h under drought and cold. It is deserved to be mentioned that the expression level of GhNCED3c_D under cold condition at 12 h was up to 389-fold than in control. The high fold-change expression levels indicated their important roles in stress response.
VIGS assay of GaNCED3a in G. arboreum under drought and salt stress
To further identify the function of abiotic stress responsive gene, we selected GaNCED3a, the ortholog gene of GhNCED3a_A/D, for reverse genetics in diploid cotton G. arboreum via VIGS method. About 2 weeks after VIGS injection, plant leaves that inoculated with pTRV2:CLA1 vector (VIGS-CLA1) were observed to turn white (Fig. 8a), which was coincident with previous findings (Li et al. 2018; Yang et al. 2019). Meanwhile, the plant leaves inoculated with pTRV2:00 and pTRV2:GaNCED3a were sampled for qRT-PCR to evaluate the silencing efficiency of GaNCED3a. Figure 8b shows that silencing of GaNCED3a was highly effective. Plants of VIGS-NCED3a-1/2/3/4/5/7/10/11/12 with expression level of GaNCED3a less than 40% of control plants were employed for further study on phenotypic and physiological variations.
Function validation of GaNCED3a by VIGS. a Phenotypes of seedlings after 14d inoculation. b Silencing efficiency of GaNCED3 gene via VIGS. c–d Phenotypes of WT, VIGS-TRV2, and VIGS-GaNCED3a seedlings under drought and salt treatment, respectively. e Water loss rate, MDA, and proline assay of VIGS-TRV2 and VIGS-GaNCED3a seedlings. f Expression analysis of biotic stress responsive genes RD29A, DREB1A, and SOS1 in WT, VIGS-TRV2, and VIGS-GaNCED3a seedlings under drought and salt stress. * indicated significant difference between treatment and its corresponding CK (p < 0.05); different lowercase letter a and b represented significant difference between the expression levels of genes in seedlings under different treatments (p < 0.05)
Then, plants of TRV2:GaNCED3a (VIGS-GaNCED3a), TRV2:00 (VIGS-TRV2), and wild type (WT) were exposed to drought and salt stress. GaNCED3a-silenced plants showed more obvious symptoms of wilting and drooping under both drought and salt stress than control plants (Fig. 8c–d). Water loss rate and MDA content in silenced plants were significantly higher than control plants (P < 0.05), while PRO content in silenced plants was significantly lower than control plants (P < 0.05) (Fig. 8e). The results of VIGS assay of GaNCED3a indicated decreased drought and salt stress tolerance in silenced plants.
To learn how GaNCED3a responded to drought and salt stress in the silenced plants, expression patterns of abiotic stress response genes DREB1A, RD29A, and SOS1 were evaluated. Under drought and salt stress, DREB1A and RD29A exhibited significantly high expression in both silenced and control plants (P < 0.5) (Fig. 8f). However, all three stress response genes showed significantly downregulated expression in VIGS-silenced plants. Silencing GaNCED3a reduced drought and salt tolerance, which may be related to the inhibition of expression activity of abiotic stress response genes DREB1A, RD29A, and SOS1.
Discussion
Characterization of cotton CCO genes
Prior to this study, CCO gene family in rice, soybean, pepper, Brassica, and apple had been systematically identified and named according to Arabidopsis orthologs (Chen et al. 2018; Kim et al. 2016; Tan et al. 2003; Vallabhaneni et al. 2010; Wang et al. 2013). A total of 15, 15, and 30 CCO were identified from G. arboreum, G. raimondii, and G. hirsutum, respectively. Cotton CCO genes could be divided into two groups, NCEDs and CCDs, which was in consistent with other plants (Giuliano et al. 2003). In NCED subgroup, there was one orthologous gene to AtNCED6 in cotton. NCED3 and NCED5 each had three copies in cotton (NCED3a, 3b, 3c, and NCED5a, 5b, 5c), while no orthologs of AtNCED2 and AtNCED9 genes were found in cotton. According to these results, it is speculated that cotton NCED3 and NCED5 had experienced a gene duplication event in the evolution process and their duplication may be the primary factor for the expansion of CCO gene family. Meanwhile, the specific enzymatic role of cotton NCED members remained elusive. Two members of cotton CCD-like genes clustered together with SlCCD-like (Fig. 1), a new subgroup found in some plant species (Chen et al. 2018; Wang et al. 2017; Wei et al. 2016; Zhou et al. 2019), while CCD-like homologs were not found in Arabidopsis. CCD1 and CCD7 had only one member each in cotton, while CCD4 and CCD8 had two members each (CCD4a, 4b, and CCD8a, 8b), which was consistent with previous studies (Vallabhaneni et al. 2010; Wang et al. 2013, 2017).
Numbers of exons and introns of cotton CCO members varied significantly. NCEDs and CCD4 in cotton were found intron deficient (intron ≤ 1), in accordance with other species, such as Rosa damascene, Osmanthus fragrans, and Fragaria vesca (Huang et al. 2009; Wang et al. 2017). This intron-deficient structure used to be considered essential for rapid response to stress through ABA synthesis and ABA-mediated signal transductions in plants (Wang et al. 2017). NCEDs in cotton had only one exon and encoded more conserved motifs than other CCO members, and was similar to the results described in other reports (Zhou et al. 2019).
The number of CCO genes in cotton was more than that in most of other plants, indicating that cotton CCO gene experienced extensive expansion in the evolution process. Polyploidization contributed a great part in duplication, while segment duplication also made valuable contributions to the expansion of gene families (Li et al. 2015; Paterson et al. 2012). Eighty-four pairs of segmental duplicated genes were observed in this study, implying that segmental duplication had profound effects on evolution of CCO gene family in cotton. Sixteen pairs of ortholog between G. hirsutum and Arabidopsis were detected, suggesting that they may derive from a common ancestor and have similar functions (Xu et al. 2012).
Expression patterns of CCO genes
Tissue-specific expression patterns implied a high functional diversification within CCO family. GhCCD1_A/D has shown high expression in all samples (Table S5, Fig. 6), which was in line with the results in other plants (Auldridge et al. 2006b; Chen et al. 2018; Simkin et al. 2004). This result suggests the multiple effects of CCD1 and its orthologs in plant growth and development (Ohmiya et al. 2006; Walter et al. 2007). In contrast with the fact that maize CCD genes preferentially high expressed in leaves (Vallabhaneni et al. 2010), prevalent CCD genes of upland cotton predominantly high-expressed in stem. However, GhCCD4b_A showed highest expression level in petal, stamen, and samples of fiber elongate stage (10 and 20 DPA), suggesting a putative function in flower and fiber development. The predominant expression of CCD4 in floral organs were also found in other plants (Brandi et al. 2011; Adami et al. 2013). The function of GhCCD4a_A, which expressed slightly in stamen, followed by root and stem, and no expression in any other tissues, remained subtle. GhCCD7_A/D barely expressed in petal, stamen, and fibers, but exhibit high expression in stem and leaf, indicating CCD7 genes might mainly take part in vegetative growth instead of reproduction growth in upland cotton, while this was inconsistent with the results in early studies that root and fruit preferentially accumulated CCD7 (Vogel et al. 2010; Wei et al. 2016; Chen et al. 2018). GhNCED3a_A/D has shown high expression level in roots, while GhNCED3b_A/D and GhNCED3c_A/D in leaves, suggesting that NCED3 genes in upland cotton exhibited diverged functions involved in plant growth. The GhNCED5c_A/D had the highest expression in the petal, and followed by the stamen, but barely expressed in ovule and fiber samples. Early studies reported that AtNCED5 induce seed dormancy (Frey et al. 2012), while our results implied specific functions of GhNCED5c_A/D involving in floral organ development.
The expression pattern analysis under abiotic stress provided useful information to gain a deep understanding of the roles of cotton CCO. Based on the results of RNA-seq data analysis and qRT-PCR, NCED3a and NCED3c in cotton were probably responsible for stress response. It is well known that abiotic stress, such as drought, salt, cold, and heat, can induce accumulation of ABA level in plants (Roychoudhury et al. 2013; Zhang et al. 2006) and NCEDs are mainly involved in ABA-mediated stress response and plant growth by regulating ABA synthesis (Huang et al. 2019; Hwang et al. 2018; Pedrosa et al. 2017; Yang et al. 2018). NCED3, which is the key player in ABA biosynthesis pathway, have been widely proved to take a prominent part in abiotic stress response, especially drought stress (Frey et al. 2012; Gavassi et al. 2021; Iuchi et al. 2000; Tan et al. 2003). GhNCED3a_A/D and GhNCED3c_A/D exhibited strong stress responses, indicating their potential roles in stress tolerance. In this study, NCED3 and NCED5 genes in upland cotton exhibited different expression pattens, implying that they had experienced divergency in function. This phenomenon was also found in other plant species (Iuchi et al. 2000; Wang et al. 2013). Frey et al. (2012) demonstrated that there was an interaction between NCED3 and NCED5 in drought stress response. Whether cotton NCED5 interacted together with NCED3 involving in ABA biosynthesis in stress response deserved to be further determined.
Silencing of GaNCED3a by VIGS reduced drought and salt stress resistance
NCED3 was demonstrated to be greatly involved in drought stress response in previous studies (Tan et al. 2003; Fujita et al. 2011; Huang et al. 2018). In this study, results of silencing GaNCED3a in G. arboreum indicated that NCED3 not only are involved in drought stress tolerance but also take an important role in salt stress response.
DREB1 (dehydration-responsive element-binding protein 1) transcription factors are involved in responsiveness to drought, salt, and low temperature stress by regulating a series of downstream genes related to abiotic stress (Gilmour et al. 2000; Yamaguchi-Shinozaki and Shinozaki 1994). RD29A is proved to have a positive effect on enhancing abiotic stress tolerance in plants (Msanne et al. 2011). SOS1, which encodes a plasma membrane Na+/H+ antiporter, is assumed to be involved in salt tolerance improvement (Yuesen et al. 2012; OlÍas et al. 2009). In this study, stress response genes DREB1A, RD29A, and SOS1 were demonstrated to be significantly downregulated in GaNCED3a-silenced cotton plants, which implied that GaNCED3a had a potential function in enhancing stress tolerance. Previous studies showed that some stress-responsive genes were upregulated by ABA under stress condition (Agarwal and Jha 2010; Gavassi et al. 2021; Ingram and Bartels 1996). What is more, mutants of nced3 and nced5 in rice were sensitive to salt and drought stress, and OsNCED3 and OsNCED5 were proved to be responsible for endogenous ABA accumulation (Huang et al. 2018, 2019). Therefore, our findings indicated that inhibited transcription level of GaNCED3 in the silenced plants may lead to decreased ABA biosynthesis, and further caused the inhibition of stress-responsive genes expression, which finally resulted in decreased drought and salt stress tolerance in GaNCED3-silenced plants. It is speculated that GaNCED3, as the orthologous of NCED3 genes in other plants, played an essential role in regulation of ABA content in abiotic stress response. However, detailed molecular mechanism of GaNCED3a in the regulation of multiple abiotic stress still deserved to be evaluated in further study.
Conclusions
We performed a genome-wide characterization of CCO gene family in three cotton species. CCO genes from A. thaliana, Z. mays, and S. lycopersicum and three species of cotton were divided into 6 subgroups. The proteins were found to be conserved. Gene expansion analysis indicated that polyploidization and segmental duplication were the major processes during cotton CCO genes expansion. Expression profile analysis revealed that a certain number of CCO genes, such as GhCCD1_A/D, GhNCED3a_A/D, and GhNCED3c_A/D may be closely involved in stress response. Silencing of GaNCED3a, the ortholog gene of stress-related gene GhNCED3a_A/D, by VIGS reduced drought and salt stress resistance. Our findings paved the way for the researches focused on clarifying the function of cotton CCO genes.
Data availability
All the data generated in the experiments are presented in manuscript and its supplementary files.
References
Adami M, De Franceschi P, Brandi F, Liverani A, Giovannini D, Rosati C, Dondini L, Tartarini S (2013) Identifying a carotenoid cleavage dioxygenase (CCD4) gene controlling yellow/white fruit flesh color of peach. Plant Mol Biol Rep 31:1166–1175. https://doi.org/10.1007/s11105-013-0628-6
Agarwal PK, Jha B (2010) Transcription factors in plants and ABA dependent and independent abiotic stress signalling. Biol Plantarum 54(2):201–202. https://doi.org/10.1007/s10535-010-0038-7
Akemi O, Katsuhiko S, Ryutaro A (2009) “Yellow Jimba”: suppression of carotenoid cleavage dioxygenase (CmCCD4a) expression turns white chrysanthemum petals yellow. J Jpn Soc Hortic Sci 78(4):450–455. https://doi.org/10.2503/jjshs1.78.450
Auldridge ME, McCarty DR, Klee HJ (2006a) Plant carotenoid cleavage oxygenases and their apocarotenoid products. Curr Opin Plant Bio 9(3):315–321. https://doi.org/10.1016/j.pbi.2006.03.005
Auldridge ME, Block A, Vogel JT, Dabney-Smith C, Mila I, Bouzayen M, Magallanes-Lundback M, DellaPenna D, McCarty DR, Klee HJ (2006b) Characterization of three members of the Arabidopsis carotenoid cleavage dioxygenase family demonstrates the divergent roles of this multifunctional enzyme family. Plant J 45:982–993. https://doi.org/10.1111/j.1365-313x.2006.02666.x
Alder A, Jamil M, Marzorati M, Bruno M, Vermathen M, Bigler P, Ghisla S, Bouwmeester H, Beyer P, Al-Babili S (2012) The path from β-carotene to carlactone, a strigolactone-like plant hormone. Science 335(6074):1348–1351. https://doi.org/10.1126/science.1218094
Bailey TL, Bodén M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202-208. https://doi.org/10.1093/nar/gkp335
Bouvier F, Suire C, Mutterer J, Camara B (2003) Oxidative remodeling of chromoplast carotenoids: identification of the carotenoid dioxygenase CsCCD and CsZCD genes involved in Crocus secondary metabolite biogenesis. Plant Cell 15:47–62. https://doi.org/10.1105/tpc.006536.model
Brandi F, Bar E, Mourgues F, Horváth G, Turcsi E, Giuliano G, Liverani A, Tartarini S, Lewinsohn E, Rosati C (2011) Study of ‘Redhaven’ peach and its white-fleshed mutant suggests a key role of CCD4 carotenoid dioxygenase in carotenoid and norisoprenoid volatile metabolism. BMC Plant Biol 11:24. https://doi.org/10.1186/1471-2229-11-24
Chen H, Zuo Z, Shao H, Fan S, Ma J, Zhang D, Zhao C, Yan X, Liu X, Han M (2018) Genome-wide analysis of carotenoid cleavage oxygenase genes and their responses to various phytohormones and abiotic stresses in apple (Malus domestica). Plant Physiol Biochem 123:81–93. https://doi.org/10.1016/j.plaphy.2017.12.001
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Chernys JT, Zeevaart JAD (2000) Characterization of the 9-cis-epoxycarotenoid dioxygenase gene family and the regulation of abscisic acid biosynthesis in Avocado. Plant Physiol 124:343–353. https://doi.org/10.1104/pp.124.1.343
El-Gebali S, Mistry J, Bateman A, Eddy SR, Luciani A, Potter SC, Qureshi M, Richardson LJ, Salazar GA, Smart A, Sonnhammer ELL, Hirsh L, Paladin L, Piovesan D, Tosatto SCE, Finn RD (2019) The Pfam protein families database in 2019. Nucleic Acids Res 47:427–432. https://doi.org/10.1093/nar/gky995
Frey A, Effroy D, Lefebvre V, Seo M, Perreau F, Berger A, Sechet J, To A, North HM, Marion-Poll A (2012) Epoxycarotenoid cleavage by NCED5 fine-tunes ABA accumulation and affects seed dormancy and drought tolerance with other NCED family members. Plant J 70(3):501–512. https://doi.org/10.1111/j.1365-313X.2011.04887.x
Fujita Y, Fujita M, Shinozaki K, Yamaguchi-Shinozaki K (2011) ABA-mediated transcriptional regulation in response to osmotic stress in plants. J Plant Res 124:509–525. https://doi.org/10.1007/s10265-011-0412-3
Gan Z, Shan N, Fei L, Wan C, Chen J (2020) Isolation of the 9-cis-epoxycarotenoid dioxygenase (NCED) gene from kiwifruit and its effects on postharvest softening and ripening. Sci Hortic 261:109020. https://doi.org/10.1016/j.scienta.2019.109020
Gavassi MA, Silva GS, Silva CMS, Thompson AJ, Macleod K, Oliveira PMR, Cavalheiro MF, Domingues DS, Habermann G (2021) NCED expression is related to increased ABA biosynthesis and stomatal closure under aluminum stress. Environ Exp Bot 185:104404. https://doi.org/10.1016/j.envexpbot.2021.104404
Gilmour SJ, Sebolt AM, Salazar MP, Everard JD, Thomashow MF (2000) Overexpression of the Arabidopsis CBF3 transcriptional activator mimics multiple biochemical changes associated with cold acclimation. Plant Physiol 124:1854–1865. https://doi.org/10.1104/pp.124.4.1854
Giuliano G, Al-Babili S, Lintig JV (2003) Carotenoid oxygenases: cleave it or leave it. Trends Plant Sci 8(4):145–149. https://doi.org/10.1016/S1360-1385(03)00053-0
Ha CV, Leyva-González MA, Osakabe Y, Tran UT, Nishiyama R, Watanabe Y, Tanaka M, Seki M, Yamaguchi S, Dong NV, Yamaguchi-Shinozaki K, Shinozaki K, Herrera-Estrella L, Tran LSP (2014) Positive regulatory role of strigolactone in plant responses to drought and salt stress. Proc Nati Acad Sci 111(2):851–856. https://doi.org/10.1073/pnas.1322135111
Hwang SG, Lee CY, Tseng CS (2018) Heterologous expression of rice 9-cis-epoxycarotenoid dioxygenase 4 (OsNCED4) in Arabidopsis confers sugar oversensitivity and drought tolerance. Bot Stud 59(1):2. https://doi.org/10.1186/s40529-018-0219-9
Hu B, Jin J, Guo AY, Zhang H, Lu J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31(8):1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Huang FC, Molnar P, Schwab W (2009) Cloning and functional characterization of carotenoid cleavage dioxygenase 4 genes. J Exp Bot 60:3011–3022. https://doi.org/10.1093/jxb/erp137
Huang Y, Guo Y, Liu Y, Zhang F, Wang Z, Wang H, Wang F, Li D, Mao D, Luan S, Liang M, Chen L (2018) 9-cis-epoxycarotenoid dioxygenase 3 regulates plant growth and enhances multi-abiotic stress tolerance in rice. Front Plant Sci 9:162. https://doi.org/10.3389/fpls.2018.00162
Huang Y, Jiao Y, Xie N, Guo Y, Zhang F, Xiang Z, Wang R, Wang F, Gao Q, Tian L, Li D, Chen L, Liang M (2019) OsNCED5, a 9-cis-epoxycarotenoid dioxygenase gene, regulates salt and water stress tolerance and leaf senescence in rice. Plant Sci 287:110188. https://doi.org/10.1016/j.plantsci.2019.110188
Ilg A, Beyer P, Al-Babili S (2009) Characterization of the rice carotenoid cleavage dioxygenase 1 reveals a novel route for geranial biosynthesis. FEBS J 276:736–747. https://doi.org/10.1111/j.1742-4658.2008.06820.x
Ingram J, Bartels D (1996) The molecular basis of dehydration tolerance in plants. Annu Rev Plant Physiol Plant Mol Biol 47:377–403. https://doi.org/10.1146/annurev.arplant.47.1.377
Iuchi S, Kobayashi M, Yamaguchi-Shinozaki K, Shinozaki K (2000) A stress-inducible gene for 9-cis-epoxycarotenoid dioxygenase involved in abscisic acid biosynthesis under water stress in drought-tolerant cowpea. Plant Physiol 123:553–562. https://doi.org/10.1128/JCM.01852-08
Kim YH, Wang I, Jung HJ, Park JI, Kang JG, Nou IS (2016) Genome-wide classification and abiotic stress-responsive expression profiling of carotenoid oxygenase genes in Brassica rapa and Brassica oleracea. J Plant Growth Regul 35:202–214. https://doi.org/10.1007/s00344-015-9520-y
Kloer DP, Schulz GE (2006) Structural and biological aspects of carotenoid cleavage. Cell Mol Life Sci 63(19–20):2291–2303. https://doi.org/10.1007/s00018-006-6176-6
Kloer DP, Ruch S, Al-Babili S, Beyer P, Schulz GE (2005) The structure of a retinal-forming carotenoid oxygenase. Science 308:267–269. https://doi.org/10.1126/science.1108965
Lefebvre V, North H, Frey A, Sotta B, Seo M, Okamoto M, Nambara E, Marion-Poll A (2006) Functional analysis of Arabidopsis NCED6 and NCED9 genes indicates that ABA synthesized in the endosperm is involved in the induction of seed dormancy. Plant J 45:309–319. https://doi.org/10.1111/j.1365-313X.2005.02622.x
Leng P, Yuan B, Guo Y (2014) The role of abscisic acid in fruit ripening and responses to abiotic stress. J Exp Bot 65:4577–4588. https://doi.org/10.1093/jxb/eru204
Li F, Fan G, Lu C et al (2015) Genome sequence of cultivated upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat Biotechnol 33(5):524–530. https://doi.org/10.1038/nbt.3208
Li TG, Zhang DD, Zhou L, Kong ZQ, Hussaini AS, Wang D, Li JJ, Short DPG, Dhar N, Klosterman SJ, Wang BL, Yin CM, Subbarao KV, Chen JY, Dai XF (2018) Genome-wide identification and functional analyses of the CRK gene family in cotton reveals GbCRK18 confers Verticillium wilt resistance in Gossypium barbadense. Front Plant Sci 9. https://doi.org/10.3389/fpls.2018.01266
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lyons E, Pedersen B, Kane J, Alam M, Ming R, Tang H, Wang X, Bowers J, Paterson A, Lisch D, Freeling M (2008) Finding and comparing syntenic regions among Arabidopsis and the outgroups papaya, poplar, and grape: coge with rosids. Plant Physiol 148(4):1772–1781. https://doi.org/10.1104/pp.108.124867
Mitchell AL, Attwood TK, Babbitt PC et al (2019) InterPro in 2019: improving coverage, classification and access to protein sequence annotations. Nucleic Acids Res 47(D1):351–360. https://doi.org/10.1093/nar/gky1100
Msanne J, Lin J, Stone JM, Awada T (2011) Characterization of abiotic stress-responsive Arabidopsis thaliana RD29A and RD29B genes and evaluation of transgenes. Planta 234:97–107. https://doi.org/10.1007/s00425-011-1387-y
Ohmiya A, Kishimoto S, Aida R, Yoshika S, Sumitomo K (2006) Carotenoid Cleavage Dioxygenase (CmCCD4a) contributes to white color formation in chrysanthemum petals. Plant physiol 142:1193–1201. https://doi.org/10.1104/pp.106.087130
OlÍas R, Eljakaoui Z, Li J, De Morales P, MarÍn-Manzano MC, Pardo JM, Belver A (2009) The plasma membrane Na+/H+ antiporter SOS1 is essential for salt tolerance in tomato and affects the partitioning of Na+ between plant organs. Plant Cell Environ 32:904–916. https://doi.org/10.1111/j.1365-3040.2009.01971.x
Paterson AH, Wendel JF, Gundlach H et al (2012) Repeated polyploidization of Gossypium genomes and the evolution of spinnable cotton fibres. Nature 492(7429):423–427. https://doi.org/10.1038/nature11798
Pedrosa AM, Cidade LC, Martins CPS, Macedo AF, Neves DM, Gomes FP, Floh EIS, Costa MGC (2017) Effect of overexpression of citrus 9-cis-epoxycarotenoid dioxygenase 3 (CsNCED3) on the physiological response to drought stress in transgenic tobacco. Genet Mol Res 16(1):gmr16019292. https://doi.org/10.4238/gmr16019292
Roychoudhury A, Paul S, Basu S (2013) Cross-talk between abscisic acid-dependent and abscisic acid-independent pathways during abiotic stress. Plant Cell Rep 32:985–1006. https://doi.org/10.1007/s00299-013-1414-5
Rubio A, Rambla JL, Santaella M, Gómez MD, Orzaez D, Granell A, Gómez-Gómez L (2008) Cytosolic and plastoglobule-targeted carotenoid dioxygenases from Crocus sativus are both involved in β-ionone release. J Biol Chem 283:24816–24825. https://doi.org/10.1074/jbc.M804000200
Schultz J, Milpetz F, Bork P, Ponting CP (1998) SMART, a simple modular architecture research tool: identification of signaling domains. Proc Natl Acad Sci 95(11):5857–5864. https://doi.org/10.1186/gb-2000-1-1-reports234
Schwartz SH, Qin X, Loewen MC (2004) The biochemical characterization of two carotenoid cleavage enzymes from Arabidopsis indicates that a carotenoid-derived compound inhibits lateral branching. J Biol Chem 279(45):46940–46945. https://doi.org/10.1074/jbc.M409004200
Schwartz SH, Tan BC, Gage DA, Zeevart JAD, McCarty DR (1997) Specific oxidative cleavage of carotenoids by VP14 of maize. Science 276(5320):1872–1874. https://doi.org/10.1126/science.276.5320.1872
Seo M, Koshiba T (2002) Complex regulation of ABA biosynthesis in plants. Trends Plant Sci 7(1):41–48. https://doi.org/10.1016/s1360-1385(01)02187-2
Simkin AJ, Schwartz SH, Auldridge M, Taylor MG, Klee HJ (2004) The tomato carotenoid cleavage dioxygenase 1 genes contribute to the formation of the flavor volatiles β-ionone, pseudoionone, and geranylacetone. Plant J 40(6):882–892. https://doi.org/10.1111/j.1365-313X.2004.02263.x
Song MH, Lim SH, Kim JK, Jung ES, Jonh KMM, You MK, Ahn SN, Lee CH, Ha SH (2016) In planta cleavage of carotenoids by Arabidopsis carotenoid cleavage dioxygenase 4 in transgenic rice plants. Plant Biotechnol Rep 10:291–300. https://doi.org/10.1007/s11816-016-0405-8
Su W, Zhang C, Feng J, Feng A, You C, Ren Y, Wang D, Sun T, Su Y, Xu L, Chen N, Que Y (2021) Genome-wide identification, characterization and expression analysis of the carotenoid cleavage oxygenase (CCO) gene family in Saccharum. Plant Physiol Bioch 162:196–210. https://doi.org/10.1016/j.plaphy.2021.02.041
Sun Z, Hans J, Walter MH, Matusova R, Beekwilder J, Verstappen FWA, Ming Z, Echtelt EV, Strack D, Bisseling T, Bouwmeester HJ (2008) Cloning and characterization of a maize carotenoid cleavage dioxygenase (ZmCCD1) and its involvement in the biosynthesis of apocarotenoids with various roles in mutualistic and parasitic interactions. Planta 228:789–801. https://doi.org/10.2307/23390113
Tan BC, Joseph LM, Deng WT, Liu LJ, Li QB, Cline K, McCarty DR (2003) Molecular characterization of the Arabidopsis 9-cisepoxycarotenoid dioxygenase gene family. Plant J 35:44–56. https://doi.org/10.1046/j.1365-313X.2003.01786.x
Tong SM, Xi HX, Ai KJ, Hou HS (2017) Overexpression of wheat TaNCED gene in Arabidopsis enhances tolerance to drought stress and delays seed germination. Biol Plant 61(1):64–72. https://doi.org/10.1007/s10535-016-0692-5
Umehara M, Hanada A, Yoshida SA, Kiyama K, Arite T, Takeda-Kamiya N, Magome H, Kamiya Y, Shirasu K, Yoneyama K, Kyozuka J, Yamaguchi S (2008) Inhibition of shoot branching by new terpenoid plant hormones. Nature 455(7210):195–200. https://doi.org/10.1038/nature07272
Vallabhaneni R, Bradbury LMT, Wurtzel ET (2010) The carotenoid dioxygenase gene family in maize, sorghum, and rice. Arch Biochem Biophys 504(1):104–111. https://doi.org/10.1016/j.abb.2010.07.019
Vishwakarma K, Upadhyay N, Kumar N, Yadav G, Singh J, Mishra RK, Kumar V, Verma R, Upadhyay RG, Pandey M, Sharma S (2017) Abscisic acid signaling and abiotic stress tolerance in plants: a review on current knowledge and future prospects. Front Plant Sci 8:161. https://doi.org/10.3389/fpls.2017.00161
Vogel JT, Walter MH, Giavalisco P, Lytovchenko A, Kohlen W, Charnikhova T, Simkin AJ, Goulet C, Strack D, Bouwmeester HJ, Fernie AR, Klee HJ (2010) SlCCD7 controls strigolactone biosynthesis, shoot branching and mycorrhiza-induced apocarotenoid formation in tomato. Plant J 61:300–311. https://doi.org/10.1111/j.1365-313X.2009.04056.x
Walter MH, Strack D (2011) Carotenoids and their cleavage products: biosynthesis and functions. Nat Prod Rep 28:663. https://doi.org/10.1039/C0NP00036A
Walter MH, Floss DS, Hans J, Fester T, Strack D (2007) Apocarotenoid biosynthesis in arbuscular mycorrhizal roots: contributions from methylerythritol phosphate pathway isogenes and tools for its manipulation. Phytochemistry 68(1):130–138. https://doi.org/10.1016/j.phytochem.2006.09.032
Wang P, Lu S, Zhang X, Hyden B, Qin L, Liu L, Bai Y, Han Y, Wen Z, Xu J, Cao H, Chen H (2021) Double NCED isozymes control ABA biosynthesis for ripening and senescent regulation in peach fruits. Plant Sci 304:110739. https://doi.org/10.1016/j.plantsci.2020.110739
Wang RK, Wang CE, Fei YY, Gai JY, Zhao TJ (2013) Genome-wide identification and transcription analysis of soybean carotenoid oxygenase genes during abiotic stress treatments. Mol Biol Rep 40:4737–4745. https://doi.org/10.1007/s11033-013-2570-y
Wang Y, Ding GJ, Gu TT, Ding J, Li Y (2017) Bioinformatic and expression analyses on carotenoid dioxygenase genes in fruit development and abiotic stress responses in Fragaria vesca. Mol Genet Genomics 292:895–907. https://doi.org/10.1007/s00438-017-1321-5
Wei Y, Wan H, Wu Z, Wang R, Ruan M, Ye Q, Li M, Zhou G, Yao Z, Yang Y (2016) A comprehensive analysis of carotenoid cleavage dioxygenases genes in Solanum Lycopersicum. Plant Mol Biol Rep 34:512–523. https://doi.org/10.1007/s11105-015-0943-1
Xu G, Guo C, Shan H, Kong H (2012) Divergence of duplicate genes in exon-intron structure. Proc Natl Acad Sci U S A 109:1187–1192. https://doi.org/10.1073/pnas.1109047109
Yadav CB, Bonthala VS, Muthamilarasan M, Pandey G, Khan Y, Prasad M (2015) Genome-wide development of transposable elements-based markers in foxtail millet and construction of an integrated database. DNA Res 22:79–90. https://doi.org/10.1093/dnares/dsu039
Yamaguchi-Shinozaki K, Shinozaki K (1994) A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress. Plant Cell 6:251–264. https://doi.org/10.1105/tpc.6.2.251
Yang X, Xu Y, Yang F, Magwanga RO, Cai X, Wang X, Wang Y, Hou Y, Wang K, Liu F, Zhou Z (2019) Genome-wide identification of OSCA gene family and their potential function in the regulation of dehydration and salt stress in Gossypium hirsutum. J Cotton Res 2:11. https://doi.org/10.1186/s42397-019-0028-z
Yang Y, Zhou Q, Xu J, Li Q, Zhang S (2018) RNA interference of NtNCED3 reduces drought tolerance and impairs plant growth through feedback regulation of isoprenoids in Nicotiana tabacum. Environ Exp Bot 155:332–344. https://doi.org/10.1016/j.envexpbot.2018.07.016
Yuesen Y, Mingcai Z, Jiachang Z, Liusheng D, Li Z (2012) SOS1 gene overexpression increased salt tolerance in transgenic tobacco by maintaining a higher K+/Na+ ratio. J Plant Physiol 169:255–261. https://doi.org/10.1016/j.jplph.2011.10.007
Zhang JH, Jia WS, Yang JC, Ismail AM (2006) Role of ABA in integrating plant responses to drought and salt stresses. Field Crops Res 97(1):111–119. https://doi.org/10.1016/j.fcr.2005.08.018
Zhang T, Hu Y, Jiang W et al (2015) Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat Biotechnol 33(5):531–537. https://doi.org/10.1038/nbt.3207
Zhao J, Li J, Zhang J, Chen D, Zhang H, Liu C, Qin G (2021) Genome-wide identification and expression analysis of the carotenoid cleavage oxygenase gene family in five Rosaceae species. Plant Mol Biol Rep. https://doi.org/10.1007/s11105-021-01284-9
Zhou Q, Li Q, Li P, Zhang S, Liu C, Jin J, Cao P, Yang Y (2019) Carotenoid cleavage dioxygenases: identification, expression, and evolutionary analysis of this gene family in tobacco. Int J Mol Sci 20:5796. https://doi.org/10.3390/ijms20225796
Funding
This work was supported by the Science and Technology Plan Projects of Hebei Province of China (16226303D) and Modern Agricultural Science and Technology Innovation Project of Hebei Academy of Agriculture and Forestry Sciences (2019–3-7–4).
Author information
Authors and Affiliations
Contributions
XC, JZ, and XZ conceived and designed the experiments. LT, XL, and HW performed the experiments. SZ, CL, and JC analyzed the collected data. XC and ZJ drafted the manuscript. JZ and XZ revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Communicated by: Izabela Pawłowicz
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cai, X., Jiang, Z., Tang, L. et al. Genome-wide characterization of carotenoid oxygenase gene family in three cotton species and functional identification of GaNCED3 in drought and salt stress. J Appl Genetics 62, 527–543 (2021). https://doi.org/10.1007/s13353-021-00634-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-021-00634-3