Abstract
Pathogenic changes to TAR DNA-binding protein 43 (TDP-43) leading to alteration of its homeostasis are a common feature shared by several progressive neurodegenerative diseases for which there is no effective therapy. Here, we developed Drosophila lines expressing either wild type TDP-43 (WT) or that carrying an Amyotrophic Lateral Sclerosis /Frontotemporal Lobar Degeneration-associating G384C mutation that recapitulate several aspects of the TDP-43 pathology. To identify potential therapeutics for TDP-43-related diseases, we implemented a drug repurposing strategy that involved three consecutive steps. Firstly, we evaluated the improvement of eclosion rate, followed by the assessment of locomotive functions at early and late developmental stages. Through this approach, we successfully identified fingolimod, as a promising candidate for modulating TDP-43 toxicity. Fingolimod exhibited several beneficial effects in both WT and mutant models of TDP-43 pathology, including post-transcriptional reduction of TDP-43 levels, rescue of pupal lethality, and improvement of locomotor dysfunctions. These findings provide compelling evidence for the therapeutic potential of fingolimod in addressing TDP-43 pathology, thereby strengthening the rationale for further investigation and consideration of clinical trials. Furthermore, our study demonstrates the utility of our Drosophila-based screening pipeline in identifying novel therapeutics for TDP-43-related diseases. These findings encourage further scale-up screening endeavors using this platform to discover additional compounds with therapeutic potential for TDP-43 pathology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
TAR DNA-binding protein 43 (TDP-43) is a 43 kDa highly conserved heterogeneous nuclear ribonuclear protein (hnRNP) composed of 414 amino acids and is encoded by the TARDBP gene located on chromosome 1 (1p36.22) [1]. TDP-43 primarily resides in the nucleus where it performs most of its physiological functions related to RNA processing, yet it can be transferred to the cytoplasm. The TDP-43 nucleo-cytoplasmic shuttling is a finely controlled process that requires both the nuclear localization sequence (NLS) and nuclear export signal (NES) of the TDP-43, and it occurs in a transcription-dependent manner [2,3,4,5,6,7,8].
Pathological changes to TDP-43 including mislocalization, misfolding, aberrant liquid–liquid phase separation, oligomerization, and post-translational modifications eventually lead to the formation of hyperphosphorylated and ubiquitylated cytoplasmic TDP-43 aggregates in combination with the loss of nuclear TDP-43 function [9,10,11,12,13,14,15,16,17]. These characteristic TDP-43-related pathological features are usually referred to as TDP-43 proteinopathy.
Intensive works have been carried out over the past several years to understand the consequences of the pathological TDP-43 changes. Numerous in vivo and in vitro studies demonstrated that a loss of TDP-43 nuclear RNA processing function globally undermines the transcriptome by either impairing pre-mRNA splicing or altering the regulation of both cryptic exons and retrotransposon activation [8, 18,19,20,21,22,23]. On the other hand, trapping TDP-43 into pathological cytoplasmic aggregates produces dramatic effects on RNA granule trafficking, and compartmentalized translation in neurons and mitochondria [24,25,26,27].
TDP-43 pathology is a progressive process that culminates in the formation of pathogenic TDP-43 deposits visible in the brain and spinal cord of patients that present across a spectrum of neurodegenerative diseases such as Amyotrophic Lateral Sclerosis (ALS), Frontotemporal Dementia (FTD), Alzheimer’s disease (AD), Parkinson's disease (PD), Huntington disease (HD) and Limbic-predominant age-associated TDP-43 encephalopathy (LATE) [28,29,30,31,32,33,34,35,36,37,38].
Although TDP-43 mislocalization and aggregation have been observed in several central nervous system (CNS) disorders, it is important to note that disease-causing mutations of TDP-43 are specific to amyotrophic lateral sclerosis (ALS) and frontotemporal degeneration (FTD). Particularly, A382T and G348C TARDBP are the most common missense mutations in familial ALS, both located in the glycine-rich domain which plays a critical role in TDP-43 aggregation [39]. In fact, either A382T or G348C TDP-43 show an increased propensity toward aggregation along with greater resistance to protein degradation. Therefore, these acquired abnormal properties extend the TDP-43 half-live and eventually accelerate the disease onset [40, 41].
Although a very small subsets of sporadic and familial ALS patients are associated with TDP-43 mutations, TDP-43 proteinopathy can be present in up to 97% of ALS patients [10, 11, 42] confirming that the alteration of TDP-43 homeostasis due to its genetic variations represents only a small portion of the disease mechanism underlying TDP-43 pathology.
Since TDP-43 proteinopathy was identified in ALS/FTD, various therapeutic approaches have been designed in cell and animal models including invertebrates and that literature have been recently reviewed [43, 44]. Notably, promoting the clearance of TDP-43 aggregates once already formed is the major therapeutic approach currently being tested in patients. The approach of repurposing existing FDA-approved drugs has proven to be an effective strategy in expediting the timeline for therapy development. Despite this, it is important to note that no TDP-43-directed therapies have yet demonstrated success in slowing down disease progression or reversing these neurodegenerative conditions in humans.
The invertebrate model Drosophila melanogaster has proven to be an excellent platform to underpin the pathological mechanisms underlying the TDP-43 pathology given the evolutionary conservation of TDP-43 [9, 45, 46]. Primarily, Drosophila has been employed to perform genetic modifier screening which has significantly broadened our understanding of TDP-43 pathology and facilitated the identification of novel genomic regulators [47,48,49,50,51,52,53]. Moreover, many studies have been also carried out to rapidly test the efficacy of novel small molecule therapeutics or to accelerate drug repurposing [54,55,56,57,58,59].
In this study, we generated novel Drosophila models of TDP-43 pathology by either conditional or constitutive expression of human TDP-43 wild type (WT) or TDP-43 G348C in different types of neurons. Subsequently, we employed these transgenic lines to perform a drug repurposing screening of twelve FDA-approved drugs that are recognized for their neuroprotective effects and currently under clinical trial and/or pre-clinical investigation for neurodegenerative diseases.
Methods
Plasmid Construction and Establishment of Transgenic Flies
The full coding sequence of human TDP-43 (Uniplot ID: Q13148) wild type (WT) and its mutant (G348C) were synthesized by Genescript and were inserted into pUC57 vector (Addgene). To construct the plasmid pUAST-TDP-43(WT) and pUAST-TDP-43(G348C), the full coding sequence of human TDP-43 was amplified by PCR with pUC57-TDP43(WT) and pUC57-TDP43(G348C), respectively. In the PCR, primers carrying EcoRI (5’-TCTGGgaattcCTAGATATCGGATCCCCAC-3’) and XhoI sites (5’-TAGAActcgagTTACGCCAAGCTTGCAT-3’) were used. The PCR products were digested with EcoRI and XhoI and inserted between the EcoRI and XhoI sites of the plasmid pUAST [60]. The constructed plasmids were verified by nucleotide sequencing. The obtained plasmids were then injected into embryos to obtain stable transformant lines carrying human TDP-43 wild type or G348C mutant (hereafter referred to as TDP-43 WT and TDP-43 G348C, respectively). P-element mediated germline transformation by injecting pUAST-TDP-43WT or pUAST-TDP-43 G348C into embryos was carried out as described earlier [61]. F1 transformants were selected on the basis of white eye color rescue as described in [62]. A total of 10 and 12 independent lines were selected for TDP-43 WT and G348C, respectively, and the viability of each was assessed by driving transgene expression with the motor neuron-specific D42-GAL4 driver.
Fly Stocks and Husbandry
Flies were reared on a standard food containing 10% glucose, 5% corn flour, 4% dry yeast, 3% rice bran and 0.65% agar in a 25 °C temperature-controlled condition. Fly lines carrying w[*]; P{w[+ mC] = GAL4 elav.L}CG16779[3]} (RRID:BDSC_8750) (elav-GAL4), w[*];P{w[+ mW.hs] = GawB}D42 (RRID:BDSC_8816) (D42-GAL4), y[1] w[*]; P{w[+ mC] = elav-Switch.O}GSG301 (RRID:BDSC_43642) (elavGS-GAL4), y[1] sc[*] v[1] sev[21]; P{y[+ t7.7] v{+ t1.8] = VALIUM22-EGFP.shRNA.4}attP40 (RRID:BDSC_41550) (UAS-EGFP), w[1118]; P{w[+ mC] = UAS-lacZ.B}Bg4-1–2 (RRID:BDSC_1776) (UAS-LacZ) were obtained from Bloomington Drosophila Stock Center. To minimize the effects of genetic background, the flies used in this study were backcrossed 6 times to w strain.
Drug Treatment
Drug screening method was performed as previously described in [57] and a few modifications were applied. To express TDP-43 in motor neurons, virgin females carrying D42-GAL4 were crossed with males carrying either UAS-TDP-43 WT or UAS-TDP-43 G348C. Twenty F1 embryos of each genotype were collected and transferred to vials containing either drugs or vehicle controls. A total of five replicates were performed for each condition. Subsequently, locomotive ability, survival rate, and visualization of neuromuscular junctions (NMJ) were assessed using the following procedures.
The drug concentration was selected using prior research to demonstrate that TDP-43 therapeutics have the maximum beneficial effect within a range of 1 to 25 µM [57]. To study the effects of fingolimod on adult flies we further employed the GeneSwitch (GS) inducible transgene expression system. In this approach, a fusion protein of GAL4 and the progesterone receptor (PR) is expressed only in neurons via the pan-neuronal elav promoter [63]. Upon an addition of the PR ligand mifepristone (RU-486), the fusion protein is translocated to the nucleus and GAL4 can bind to UAS elements to activate the transgene expression. To achieve this, virgin females carrying elavGS-GAL4 were crossed at 25 °C with males carrying either UAS-TDP-43 WT or UAS-TDP-43 G348C, to express the transgenes in all neuronal subtypes. The progenies at adult stage were then transferred to vials containing 80 µg/mL (1X) or 40 µg/mL (0.5X) of RU-486 or a combination of RU-486 and drugs respectively. The details of vehicles and drug concentration are shown in Table S4. The equivalent volume of each vehicle was used as control. Drug.
Pupal Lethality Assay
The parental crossing was conducted using instant food (Formula 4–24, Carolina Biological Supply Company) under controlled conditions of 25 °C temperature and 65% humidity. Subsequently, twenty embryos were transferred into vials containing various concentrations of either drugs or vehicles, as appropriate for the experiment. To ensure robustness and reliability, five replicates were performed in parallel for each condition. The eclosion rate, representing the number of adult flies that emerged per embryos analyzed, was used as the measurement for evaluating the effects of the treatments.
Larval Locomotive Assay
The parental crossing and drug administration were carried out as above described. Locomotive abilities of wandering third instar larvae were measured by crawling assay as previously described in [64]. Briefly, each larva was gently transferred onto a 15-cm petri dish containing 2% agarose. The videos were recorded when larvae started moving and finished at a 30-s recording. The crawling paths were analyzed by using ImageJ software with plugged-in wrMTrck (NIH). Larvae were collected from three independent parental crosses, and a total of 30 larvae were included in the assay.
Adult Locomotive Assay
The locomotion abilities of adult flies were evaluated using a negative geotaxis assay. The parental crossing and development of progeny were conducted on standard food. For the GeneSwitch (GS) inducible transgene expression system, adult progenies were transferred into vials containing either RU-486 or a combination of fingolimod and RU-486, depending on the experimental condition. Two concentrations of RU-486, 80 µg/mL (1X) or 40 µg/mL (0.5X), were used to induce transgene expression in combination with fingolimod or vehicle. Fingolimod was provided at different concentrations: 0.5, 1, 5, and 10 µM. Climbing assay was conducted at various time points, including 5, 10, 15, and 20 days post-eclosion, specifically on flies expressing the transgenes in motor neurons using the D42-GAL4 driver. Only two doses of fingolimod were tested in this assay, and no RU-486 was used. For flies expressing the inducible transgene expression system, climbing assay was performed solely 5 days post-eclosion. On the day of experiment, adult male flies from each group were collected and placed into a glass cylinder at a density of 10 flies per vial. They were given 15 min to recover from CO2 exposure. The vial containing flies was tapped on the bench at 30-s intervals for a total of 5 times, and the entire process was recorded. The climbing index for each group was calculated by summing the number of flies in each score category, multiplying it by the corresponding score, and then dividing by the total number of flies in the group. Offspring from three independent parental crosses were collected for the climbing assay of transgenes driven by elavGS-GAL4, with a total of 100 adult flies approximately used for each condition. However, due to a significant reduction in survival of flies expressing the transgene in motor neurons with D42-GAL4, the number of flies tested at 15- and 20-days post eclosion was less or equal to 70.
Visualization of NMJ Morphology
The morphology of the neuromuscular junction (NMJ) was inspected in twenty larvae for each experimental condition. Larvae were collected from at least two independent parental crosses, and the NMJ morphology was studied by adapting a previously described protocol [65]. Briefly, the third instar larvae were dissected in an ice-cold HL3 medium, and their body walls were fixed in 4% paraformaldehyde/PBS at 25 °C for 30 min. The fixed body walls were then blocked with 2% normal goat serum/0.1% Triton X-100 in PBS at 25 °C for 30 min before being stained with fluorescein isothiocyanate (FITC)-conjugated goat anti-horseradish peroxidase (HRP) IgG (1:1000, ThermoFischer). After that, the samples were probed with mouse monoclonal anti-Discs large (Dlg) (1:300, DHSB) at 4 °C for 16 h. After extensive washing with 0.1% Triton-X-100 in PBS, the samples were further incubated with the fluorescence-conjugated secondary antibody Alexa-594 anti-mouse IgG (1:400) at 25 °C for 2 h. The samples were mounted with Vectashield (Vector Laboratories) and inspected under a confocal laser-scanning microscope (OLYMPUS FluoView FV10i). Images were analyzed with the program MetaMoph Imaging System 7.7 (Molecular Devices Inc.).
Larval Brain Visualization
Whole larval brain visualization was performed as previously described [66]. Briefly, ten larvae per genotype were dissected in an ice-cold PBS medium and fixed in 4% paraformaldehyde/PBS at 25 °C for 30 min. The fixed brains were then blocked with 2% normal goat serum/0.1% Triton X-100 in PBS at 25 °C for 30 min before being incubated with the high-affinity F-actin probe (Phalloidin) conjugated to FITC (1:1000, ThemoFischer). Samples were further incubated with anti-TDP-43 rabbit IgG 1:1000 (Cell Signaling Technology, UAS) at 4 °C for 16 h. After extensive washing with 0.1% Triton-X-100 in PBS, the samples were further incubated with the fluorescence-conjugated secondary antibody Alexa-594 anti-rabbit IgG (1:400) at 25 °C for 2 h. DAPI was then used to stain nuclei of the samples before being mounted with Vectashield (Vector Laboratories) and inspected under a confocal laser-scanning microscope (OLYMPUS FluoView FV10i). Images were analyzed with the program MetaMoph Imaging System 7.7 (Molecular Devices Inc.).
Protein Extraction, Fractionation, and Western Blotting
For the protein extraction, crude extracted were obtained from 20 adult heads as follow. A 100 µl lysis buffer (30 mM Hepes–KOH pH 7.6, 20 mM KCl, 10 mM MgCl2, 0.2 mM EDTA pH 8.0, 1% NP-40 4 M urea 1X protease inhibitor) was added to 20 heads. Homogenization was performed on ice using TT-13 K Mini Handheld Homogenizer (Hercuvan) followed by a 20 min incubation at 4C. Total proteins were recovered with a Centrifuge 14,000 × g 15 min 4C. The supernatant, which contained the recovered total proteins, was carefully collected for further analysis.
For the TDP-43 solubility test, we adapted a previously described protocol [67]. In brief, 35 adult fly heads were dissected and homogenized in 250 µl of RIPA buffer. The RIPA buffer consisted of 50 mM Tris–HCl (pH 8), 150 mM NaCl, 2 mM EDTA, 1% Nonidet-P40 (v/v), 0.1% SDS, 1% Na-deoxycholate, and a cocktail of protease inhibitors (Roche cOmplete™). The homogenized samples were incubated under agitation at 4 °C for 1 h and then centrifuged at 1000 g for 10 min at 4 °C. At this point, a 50 µl aliquot was taken as the input sample. The remaining supernatant was further centrifuged at 100,000 × g for 30 min at 4 °C to collect the soluble fraction. The pellet obtained was re-extracted using 150 µl of urea buffer, which contained 8 M urea, 50 mM Tris–HCl (pH 8), 1% CHAPS, and a cocktail of protease inhibitors (Roche cOmplete™). The re-extracted sample was spun down to remove any precipitate, and the resulting supernatant containing the urea-soluble proteins was collected as the insoluble fraction. To analyze the protein samples, a 10% SDS-PAGE was performed. The input, RIPA fraction, and urea fraction were loaded onto the gel in equal proportions (1:1:1 ratio) for comparison.
For the study of TDP-43 cellular compartmentalization, a nuclear-cytoplasmic extraction protocol was adapted from [68]. In brief, 50 adult fly heads were homogenized in 200 μL of CytoEB1X buffer (or CytoEB2X buffer for subsequent steps), which consisted of 30 mM Hepes–KOH (pH 7.6), 20 mM KCl, 10 mM MgCl2, 0.2 mM EDTA (pH 8.0), and 20% Glycerol. To ensure complete cytoplasmic extraction, three washing steps were performed using 150 μL of WASH-100, WASH-150, and WASH-250 buffers, respectively. Each wash buffer was prepared by supplementing CytoEB 1X with 100 mM, 150 mM, or 250 mM Sucrose, respectively. After the washing steps, the pellet was re-suspended in 200 μL of RIPA buffer. The sample was vortexed and incubated at 4 °C for 30 min, followed by centrifugation at 14,000 g for 15 min. The resulting supernatant was collected as the nuclear fraction for further analysis.
For immunoblotting, protein extracts were run on 10% SDS–polyacrylamide gels and blotted to polyvinylidene difluoride (PVDF) membranes (Bio-Rad). Membranes were blocked with TBS 0.05% Tween-20 containing 5% BSA at 25 °C for 1 h and further incubated 16 h in 5% BSA blocking solution containing primary antibodies including polyclonal rabbit anti-TDP-43 IgG (1:1000, CST #89,789), monoclonal mouse anti-Actin IgG (1:1000, DSHB #JLA20), monoclonal mouse anti-Lamin C (1:1000, DSHB #LC28.26) and monoclonal mouse anti-β-tubulin (1:100, DSHB, #E7). Next, membranes were incubated with appropriate secondary antibodies including goat anti‐rabbit (1:3000, Biorad) anti-mouse (1:3000, Biorad) antibodies conjugated to horseradish peroxidase (HRP) for 2 h, at 25 °C. The Western blot signal was detected with Clarity Max ECL western blotting substrates (Biorad), and images were taken and analyzed with C-DiGit Blot Scanner (LI-COR). The intensity of each immunoreactive band was analyzed by Image J.
Semi-quantitative PCR
Total RNA was isolated from thirty adult heads with spin column QIAGEN and 500 ng were retrotranscribed with RevertAid First Strand cDNA Synthesis Kit and Oligo(dT)18 Primer (ThermoFisher) according to the manufacture instruction. Quantitative real-time PCR was performed with SensiFAST™ Probe Lo-ROX Kit and analyzed by 7500 Real-Time PCR Systems (Applied Biosystems™). Both TDP-43 and TBPH transcripts were normalized on Cyp1 and ACT5 reference gene, and folds were calculated versus untreated control. (four biological replicates; real-time reactions performed in triplicate for each biological replicate).
List of primers used: TDP43-FR: 5’-ATGGGTGGTGGGATGAACTTT-3’, TDP43-RV: 5’-CAGTGGGCCTGACTGGTTCT-3’, Cyp1-FR: 5’-TCGGCAGCGGCATTTCAGAT-3’, Cyp1-RV: 5’-TGCACGCTGACGAAGCTAGG-3’, ACT5-FR:5’-CAACTGGGACGATATGGAGAAG-3’, ACT5-RV:5’-GTCTCGAACATGATCTGGGTC- 3’.
Statistical Analyses
Data analyses were performed blind to the experimental conditions and all results are represented as mean ± SEM or mean ± SD. Multiple comparisons were performed by Kruskal–Wallis one-way ANOVA following post hoc Dunnet’s test. P ≤ 0.05 was considered a significant difference. GraphPad Prism 9.5.1 was used to perform all analyses.
Results
Development of Novel Drosophila Models of TDP-43 Pathology
The sequences of TDP-43 WT and the ALS/FTD-associating mutant form G348C (Fig. 1A) were obtained by PCR with pUC57-TDP43 WT and pUC57-TDP43 G348C, respectively, and further cloned into pUAST vectors for microinjections into Drosophila embryos. Insertion was designed downstream of the five tandemly arrayed optimized GAL4 binding sites followed by the hsp70 TATA box and upstream of the SV40 small T intron and polyadenylation site (Fig. 1B). The obtained plasmids were verified by sequencing and then injected into embryos to obtain stable transformant lines carrying TDP-43 WT or TDP-43 G348C. Results of Drosophila embryonic microinjection are summarized in Table S1. A total of ten and twelve Drosophila lines carrying TDP-43 WT and TDP-43 G348C were further tested, respectively. Pupal lethality of motor neuron-driven transgene expression was chosen as an initial readout for further drug screening (Tables S2 and S3). The TDP-43 WT transgenic line (line number 7) and G348C transgenic line (line number 12) carrying the P-element inserted in the same third chromosome whose transgene expression caused pupal lethality were selected for further studies (Tables S2 and S3). Both moderately toxic TDP-43 WT and TDP-43 G348C lines, specifically line number 1 (referred to as TDP-43 WT moderate) and line number 1 (referred to as TDP-43 G348C moderate), were chosen for further analysis in this study. These lines were selected because they do not result in lethality when expressed in motor neurons, allowing for long-term drug treatment and the assessment of drug effectiveness.
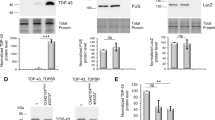
The newly established transgenic fly lines show common features of TDP-43 proteinopathy. A Schematic representation of major domains of TDP-43. The mutation used in this study (G348C) is also marked. The annotation of domains and motifs was retrieved from UniProt (https://www.uniprot.org) and listed as N-terminal (N-term) to C-terminal (C-term) orientation: nuclear localization signal (NLS), RNA-binding domain 1 (RRM1) and 2 (RRM2), nuclear export signal (NES), glycine-rich domain also known as disorder domain. B The simplified map of transgenic constructs cloned into pUAST vectors and used to microinject Drosophila embryos. The TDP-43 sequences, whether wild-type or carrying the G348C mutation, were cloned using the EcoRI-XbaI orientation. Transgenes were allocated downstream the five tandemly arrayed optimized GAL4 binding sites followed by the hsp70 TATA box. The SV40 small T intron and polyadenylation site is present downstream the transgene. Additionally, the P-element ends (P3' and P5') along with the white gene, which serves as markers indicating successful integration into the Drosophila genome, were also included in the construct. C Locomotive functions of wandering third instar larvae carrying w1118; UAS-LacZ/+; D42-GAL4/+ (CTRL, n= 24), w1118; +; D42-GAL4/UAS-TDP-43 WT (TDP-43 WT, n= 20) and w1118; +; D42-GAL4/UAS-TDP-43 G348C (TDP-43 G348C, n= 21) were tested by crawling assay. Distance (cm) covered in 30 sec was measured and plotted. (****) P<0.0001. (Median [cm/30 sec]: CTRL= 4.16 ± 0.70; TDP-43 WT= 2.13 ± 0.94; TDP-43 G348C =2.62 ± 0.68) D-E TDP-43 levels were assessed by performing western blotting on crude extracts obtained from the whole bodies of third instar larvae carrying w1118; +; D42-GAL4/UAS-TDP-43 WT (TDP-43 WT) and w1118; +; D42-GAL4/UAS-TDP-43 G348C (TDP-43 G348C). The relative abundance of the main TDP-43 band (43 kDa) was determined by quantifying the signal intensity using actin as an internal control. Statistical analysis was conducted on three independent extractions, and the results of two of these extractions are presented in the figure. F The solubility of TDP-43 was assessed by performing immunoblotting on RIPA and UREA fractions, obtained as described in the Materials and Methods section. G The distribution of TDP-43 in different cellular compartment was determined using nuclear and cytoplasmic protein fractionations. The effectiveness of the fractionation was confirmed by measuring the relative distribution of cytoplasmic β-tubulin and nuclear Lamin C in the fractions. H TDP-43 immunohistochemistry was conducted using dissected brains of third instar larvae carrying w1118; UAS-LacZ/+; elav-GAL4/+ (CTRL), w1118; +; elav-GAL4/UAS-TDP-43 WT (TDP-43 WT) and w1118; +; elav-GAL4/UAS-TDP-43 G348C (TDP-43 G348C). Dotted circles highlight the nuclei. Nuclei are stained with DAPI (blue); actin filaments are stained with fluorescein isothiocyanate (FITC)-conjugated Phalloidin (Pha). Anti-rabbit Alexa-Fluor 594 antibody was stained against anti-rabbit TDP-43 antibody (red). Scale bars: 100 μm (longer white line); 10 μm (shorter white line). A lower magnification showing the whole larval brain is shown in Fig. S1
The newly established transgenic fly lines exhibited characteristic features of TDP-43 proteinopathy (Fig. 1C-H). When the transgenes were expressed in motor neurons, locomotion defects, NMJ alterations, and predominant cytoplasmic localization of both wild-type and mutant TDP-43 were observed (Figs. 1C and G, S1A–C). Similarly, the expression of the transgenes using a pan-neuronal driver led to the cytoplasmic localization of both TDP-43 WT and G348C in the ventral cord of larval brains (Figs. 1H and S1D). Despite TDP-43 G348C being expressed at higher levels than TDP-43 WT (Fig. 1D-E), both proteins were predominantly enriched in the RIPA fractions (Fig. 1F). Importantly, the pupal lethality observed in these lines did not show any macroscopic alteration in body morphology, as the flies removed from the pupal pharate displayed all the typical attributes of insects (Fig. S1E).
Five FDA-approved Drugs Improved Pupal Lethality of TDP-43 Expressing Flies
We further developed a pilot drug screening platform utilizing the newly generated transgenic lines to assess the efficacy of FDA-approved drugs in addressing TDP-43 pathology. In this phase, we randomly selected ten drugs known to have potential neuroprotective effects in various neurological conditions, as confirmed by either pre-clinical or clinical evidence. Additionally, we included both riluzole and edaravone, as they are the two drugs currently prescribed for ALS patients. This important undertaking sets the stage for further investigations and the possibility of identifying promising candidates for the treatment of TDP-43-related conditions. Basic information of the drugs used in this study are summarized in Table S4. Therefore, we designed a time-saving and sustainable experimental workflow composed of three consecutive steps in which an improvement of eclosion rate, and locomotive functions at larval and adult stages, respectively were used to identify novel potential TDP-43 therapeutics (Fig. 2).
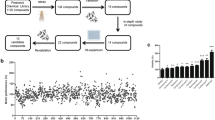
As an initial step in the drug screening process, we focused on assessing the potential of the screened drugs to rescue the pupal lethality associated with the expression of either wild-type TDP-43 or the TDP-43 G348C mutant specifically in motor neurons. The drugs were administered to the progeny at the embryonic stage and subsequently replenished every four days until the pupal stage. This continuous drug administration regimen ensured sustained exposure to the drugs throughout the developmental period. (Fig. 3A). A low dose of fingolimod (1 µM, Fng-L) and a high dose of pioglitazone (10 µM, Pgl-H) were the only drugs demonstrated significant improvement in the eclosion rate for both TDP-43 WT and TDP-43 G348C (Fig. 3B, C). (TDP-43 WT untreated vs Fng-L treated mean: 0.09 ± 0.03 vs 0.40 ± 0.10; TDP-43 G348C untreated vs Fng-L treated mean: 0.10 ± 0.04 vs 0.18 ± 0.08; TDP-43 WT untreated vs Pgl-H treated mean: 0.07 ± 0.07 vs 0.29 ± 0.06; TDP-43 G348C untreated vs Pgl-H treated mean: 0.02 ± 0.02 vs 0.32 ± 0.15)
Rescue of pupal lethality serves as a quick screening of drug repurposing for TDP-43 proteinopathy in Drosophila transgenic lines. A Embryos were collected and transferred to vials containing either the respective drugs or vehicle controls. The eclosion rate was then assayed to determine the number of flies that successfully emerged from the pupal stage B-C The eclosion rate was calculated as a ratio between the number of adult flies that successfully emerged from each vial and divided it by the initial number of embryos in that specific vial. A total of 20 larvae carrying B w1118; +; D42-GAL4/UAS-TDP-43 WT (TDP-43 WT) or C w1118; +; D42-GAL4/UAS-TDP-43 G348C (TDP-43 G348C) was allocated in a vial with different treatments (5 replicates per each treatment). Treatments are grouped according to the solvent used: distilled sterile water (H2O), (0.02-0.1%) ethanol (EtOH), (0.02-0.1%) dimethyl sulfoxide (DMSO). Drugs were tested in either at low (L, 1µM) or high (H, 10µM) dose and are listed in alphabetical order. Ceftriaxone (Cft), Carbamazepine (Crb), Dexpramipexole (Dxp), Edaravone (Edr), Fingolimod (Fng), Gabapentin (Gbp), Methylcobalamin (Mth), Olesoxime (Ols), Pioglitazone (Pgl), Riluzole (Rlz), Rapamyicin (Rpm), Tricostatin A (Trc). Yellow and orange boxplots represent the low and high doses of the selected drug used in the screening, respectively, while the white boxplots indicate the vehicle control. The median is shown as a horizontal line within each boxplot. The whiskers represent the 10th and 90th percentiles of the data. *P<0.05, **P<0.001
Moreover, a low dose of ceftriaxone (5 µM, Cft-L) or dexpramipexole (1 µM, Dxp-L) had a significative effect on TDP-43 WT flies (TDP-43 WT Cft-L treated mean: 0.38 ± 0.29; TDP-43 WT Dxp-L treated mean: 0.35 ± 0.24) with a trend to improve pupal lethality in TDP-43 G348C flies. Similarly, a low dose of edaravone (1 µM, Edr-L) significantly increased the eclosion rate of TDP-43 G348C expressing flies (TDP-43 G348C Edr-L treated mean: 0.20 ± 0.16) with a positive trend in TDP-43 WT expressing flies.
Based on these findings, we selected five positive candidates which are ceftriaxone, dexpramipexole, edaravone, fingolimod, and pioglitazone for examining the further readouts.
Administration of Fingolimod At the Early Developmental Stage Ameliorated Locomotion of Larvae Expressing TDP-43
We proceeded with the drug screening by evaluating the effects of five selected drugs on the locomotor functions of larvae expressing TDP-43 in motor neurons. The drugs were administered to the embryos as described earlier, and the crawling ability of third instar larvae was assessed (Fig. 4A).
Crawling assay of Drosophila larvae expressing TDP-43 in motor neurons. A Schematic diagram of experimental procedures (crawling assay). B-C Locomotive functions of wandering third instar larvae carrying B w1118; +; D42-GAL4/UAS-TDP-43 WT (TDP-43 WT, n= 30) and C w1118; +; D42-GAL4/UAS-TDP-43 G348C (TDP-43 G348C, n= 30) treated with different drugs or untreated (UNT) were tested by crawling assay. Distance (cm) covered in 30 sec was measured and plotted. *P<0.05, **P<0.001. Yellow and orange boxplots represent the low and high doses of the selected drug used in the screening, respectively, while the white boxplots indicate the vehicle control. The median is shown as a horizontal line within each boxplot. The whiskers represent the 10th and 90th percentiles of the data
Among the tested drugs, only fingolimod at a low dose (Fng-L) showed a significant improvement in locomotor deficits for both TDP-43 WT and TDP-43 G348C expressing larvae. The mean crawling scores for TDP-43 WT untreated vs Fng-L treated larvae were 2.23 ± 0.94 and 3.26 ± 0.99, respectively. For TDP-43 G348C untreated vs Fng-L treated larvae, the mean crawling scores were 2.72 ± 0.68 and 3.79 ± 0.44, respectively (Fig. 4B, C).
Additionally, a low dose of edaravone (Edr-L) and a high dose of pioglitazone (Pgl-H) were found to increase crawling length in TDP-43 WT larvae (TDP-43 WT Edr-L treated mean: 3.27 ± 0.77; TDP-43 WT Pgl-H treated mean: 3.29 ± 0.67). Furthermore, a low dose of ceftriaxone (Cft-L) exhibited a positive effect on the locomotion of TDP-43 G348C larvae (TDP-43 G348C Cft-L treated mean: 3.91 ± 0.52). Notably, there was also a trend towards improvement in locomotor function observed when administering Edr-L to TDP-43 G348C larvae and Cft-L to TDP-43 WT larvae.
Considering the substantial improvement in locomotor dysfunction observed in both TDP-43 WT and G348C expressing larvae, fingolimod emerged as the best candidate for subsequent experiments.
Administration of Fingolimod At the Early Developmental Stages Lacks Significant NMJ Morphology Improvement
According to previous reports, ectopically constitutive expression of human TDP-43 in motor neurons leads to a reduction in axonal branch length and bouton numbers [69], a feature that is also recapitulated by the transgenic lines in this study (Fig. S1). Therefore, we next examined whether fingolimod was capable to improve such deficits (Fig. 5A).
Inspection of the NMJ morphology upon fingolimod administration. A Schematic diagram of experimental procedures (NMJ inspection). B-E The NMJs innervating body wall muscle 4th in abdominal segment 3-4 (m4 in A3-4) were stained with antibodies targeting the postsynaptic density marker, Dlg (red) and the neuronal membrane marker, HRP (green) and were further analyzed. The representative HRP and Dlg in third instar larvae carrying w1118; +; D42-GAL4/UAS-TDP-43 WT (TDP-43 WT) and w1118; +; D42-GAL4/UAS-TDP-43 G348C (TDP-43 G348C) treated with fingolimod at 1µM (low dose, Fng-L) or 10 µM (high dose, Fng-H) and untreated (UNT) were shown in panel B and D, respectively. The number of boutons at the NMJ muscle 4th were quantified and compared with untreated condition (UNT) (C and E). n = 15, scale bar: 20 µm
A positive trend to increase the number of boutons of the NMJ4’s main branch was found upon administration of a low dose of fingolimod to larvae expressing TDP-43 WT (TDP-43 WT untreated vs Fng-L treated mean: 9.6 ± 2.6 vs 11.7 ± 2.4 n. boutons) (Fig. 5B and C). However, a similar effect was barely seen on larvae expressing TDP-43 G348C (11.2 ± 2.7 vs 12.4 ± 3.0 n. boutons) (Fig. 5D, E).
Fingolimod Post-transcriptionally Reduces the Level of TDP-43 WT to Improve Locomotive Functions in Adult Flies
To investigate the effects of fingolimod on adult flies, we utilized the GeneSwitch (GS) inducible transgene expression system. This approach was chosen because the expression of TDP-43 transgenes driven by D42-GAL4 in motor neurons resulted in pupal lethality, as indicated in Tables S2 and S3 of our supplementary materials. Additionally, we observed similar outcomes when using plain elav-Gal4-driven expression in all subtypes of neurons (data are not shown). Therefore, we conducted crosses between UAS-TDP-43 flies (both WT and G348C) and elav-GS flies, and the resulting offspring were subjected to a 5-day treatment with RU-486 (see method section). During this experiment, two different concentrations of RU-486, and the fingolimod dose–response assessment was performed by comparing the climbing scores of the flies (Fig. 6A). The protein levels of both transgenes were verified to be higher, corresponding to an increase in the concentration of RU-486 employed (Fig. S2).
Fingolimod improves locomotive functions in adult flies and post-transcriptionally reduces the level of TDP-43. A Schematic diagram of experimental procedures. Locomotive function was examined in adult flies expressing TDP-43 WT or TDP-43 G348C upon eclosion. Conditional expression of transgene was achieved by five consecutive days administration of mifepristone (RU-486) in flies carrying w1118; +; elavGS-GAL4/UAS-TDP-43 WT (TDP-43 WT) and w1118; +; elavGS-GAL4/UAS-TDP-43 G348C (TDP-43 G348C). Treatment with fingolimod was initiated upon fly eclosion and continued for 5 consecutive days. Fingolimod was administered at four different concentrations: 0.5 µM, 1 µM, 5 µM, and 10 µM. The climbing scores of the treated flies were compared to those of the untreated flies (0 µM). B-E The climbing abilities of five-day-old flies expressing TDP-43 WT or G348C, treated with 0.5X (B and D) or 1X (C and E) fingolimod, were assessed using a negative geotaxis assay. Statistical analysis was conducted on five independent replicates, Total flies assayed n = 95-110, *P<0.05. F-I After five consecutive days of inducing TDP-43 expression and treating with fingolimod, crude extracts were obtained from fly heads. TDP-43 immunoblots were performed to assess the levels of TDP-43 WT (F and H) or G348C (G and I), and Actin was used as a reference to quantify the relative abundance of the major TDP-43 band (43 kDa). Statistical analysis was performed on three independent Western blot experiments. *P<0.05; ***P<0.001; ns = not significant
Significantly, the locomotive functions of flies expressing TDP-43 WT were restored by treatment with 0.5 and 1 µM concentrations of fingolimod when a half-dose of RU-486 was used for induction (TDP-43 WT untreated vs 0.5-Fng or 1-Fng treated, respectively: mean 2.23 ± 0.17 vs 2.96 ± 0.21 or vs 3.17 ± 0.18 climbing score). However, the lowest concentration of fingolimod demonstrated ineffectiveness in rescuing locomotive functions when flies were induced with a full dose of RU-486. Notably, only the treatment with 1 µM fingolimod exhibited a statistically significant amelioration of climbing performance. (TDP-43 WT untreated vs 1-Fng treated: mean 2.49 ± 0.27 vs 3.06 ± 0.21 climbing score) (Fig. 6B, C). In contrast, flies expressing TDP-43 G348C exhibited an improvement in climbing scores when treated with 1 and 5 µM concentrations of fingolimod during induction with a half-dose (0.5X) of RU-486 (TDP-43 G348C untreated vs 1-Fng or 5-Fng treated, respectively: mean 2.41 ± 0.20 vs 2.86 ± 0.21 or vs 2.90 ± 0.10 climbing score). However, no benefit of fingolimod treatment was observed when the mutant TDP-43 was induced with a higher dose of RU-486 (Fig. 6D, E). Based on these observations, we discovered that the beneficial effect of fingolimod was more pronounced when we induced lower levels of TDP-43. These findings suggested a plausible link between the mechanism by which fingolimod rescues TDP-43 toxicity and the level of TDP-43 itself, indicating that the efficacy of fingolimod may be more limited in the presence of high TDP-43 levels.
To begin investigating the mechanism of how fingolimod exerts beneficial effects on TDP-43 pathology, post-climbing assay, total proteins were extracted from both fingolimod-treated and untreated flies. Immunoblotting analysis was performed to assess the levels of TDP-43. The results demonstrated a reduction in the TDP-43 protein level in flies expressing TDP-43 WT or G348C corresponding to the observed improvement in climbing scores. Specifically, a statistically significant reduction in TDP-43 WT levels was observed in flies treated with 1 µM fingolimod. Furthermore, a trend towards decreased TDP-43 levels was observed in flies treated with 0.5 and 1 µM fingolimod when a half dose (0.5X) of RU-486 was used for transgene induction. In contrast, a significant reduction in TDP-43 G348C levels was found only when flies were treated with 1 µM fingolimod, and transgene expression was induced using a half dose of RU-486. However, an intriguing trend towards decreased TDP-43 G348C levels was also observed in flies treated with 5 µM fingolimod, when a half dose of RU-486 was used for induction. (Fig. 6F–I; uncropped western blots are provided in Fig. S5).
To understand whether such a decline in TDP-43 amount was due to modulation of the elav-GS system at the transcriptional level, we further measured the level of TDP-43 mRNA by semi-quantitative PCR. No difference was observed in the comparison with untreated controls suggesting that fingolimod is capable to regulate the TDP-43 post-transcriptionally (Fig. S3A–D).
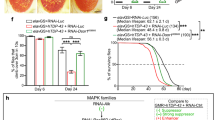
To evaluate the long-term impact of fingolimod on locomotive function, we used two moderately toxic TDP-43 lines that do not cause lethality when expressed constitutively in motor neurons (refer to Tables S1, S2, and S3). We measured the climbing score of flies that were raised with or without fingolimod (Fig. S4A). Without fingolimod treatment, both TDP-43 WT moderate and TDP-43 G348C moderate expressing flies exhibited a rapid decline in locomotive abilities. Interestingly, fingolimod treatment had slightly different effects on these lines: while TDP-43 WT moderate flies treated with 1 µM fingolimod maintained a higher climbing score for up to fifteen days post-eclosion, a significant beneficial effect of fingolimod was only observed in aged TDP-43 G348C moderate flies (Fig. S4B, C).
The underlying reasons for this apparent differential effect may require further investigation to be fully elucidated. However, it is worth noting that the amino acid substitution in G348C, which makes the protein more resistant to protein degradation as discussed earlier, could potentially play a role in this differential response.
Taken together, these data indicate that fingolimod is likely beneficial against TDP-43 pathology, at least in part, due to its ability to post-transcriptionally reduce TDP-43 levels.
Discussion
The objective of this study was to establish a Drosophila-based drug screening pipeline for the identification of US-FDA-approved drugs that could be repurposed for the treatment of TDP-43 pathology. A comprehensive screening was conducted using a three-step assay to evaluate pupal lethality rescue, as well as improvements in larval and adult locomotive function. For this pilot test, we randomly selected ten drugs currently undergoing clinical trials for their potential neuroprotective effects in different neurological conditions, along with two FDA-approved drugs for ALS patients. The primary goal was to gain initial insights and gather preliminary data to inform future investigations and refine the drug screening approach. The screening platform utilized in the study identified ceftriaxone, dexpramipexole, and pioglitazone as compounds that improved the eclosion rate in flies expressing human TDP-43 WT. Additionally, edaravone and pioglitazone were found to improve the eclosion rate in flies expressing the ALS/FTD-associated G348C mutant. However, fingolimod stood out as a particularly promising drug candidate since it demonstrated positive effects on all tested phenotypes in both TDP-43 WT and mutant flies, despite the reduction in beneficial effects at higher doses. Subsequently, in vitro assays supported the potential of fingolimod by showing its ability to reduce TDP-43 levels post-transcriptionally. These findings validate the effectiveness of this platform in identifying novel therapeutics that target TDP-43 pathology.
However, it is important to acknowledge a few limitations of this workflow that should be considered for future screening endeavors. One such limitation is the initial screening method, which focused on evaluating improvements in eclosion rate. This approach excludes the use of transgenic TDP-43 flies, such as those available at the Bloomington stock center, so far that exhibit vital phenotypes. Overcoming this limitation could involve transitioning to a life span readout assay as an alternative screening method. However, to implement this approach may significantly increase both the time and cost required for the screening process.
Another limitation of this platform is its reliance on two-dose testing, which may not fully capture the dose-dependent effects of the drugs, potentially missing out on optimal therapeutic concentrations. To address this limitation, an iterative testing approach can be considered. This involves retesting promising drugs identified in the initial screening using additional doses in subsequent experiments. As we have done with fingolimod based on promising results, this iterative process allows for a more refined assessment of dose–response relationships and increases the likelihood of identifying promising drugs.
We also acknowledge the limitations related to certain drugs that exert their neuroprotective function through specific receptors, which may not be conserved in Drosophila. In fact, drugs that rely on specific receptors not present in Drosophila may not be suitable for this screening platform. However, it is worth noting that all the drugs tested in this study either have conserved receptors or are known to exhibit neuroprotective effects through receptor-independent mechanisms. Therefore, the lack of significant results observed for some of the drugs tested in this study cannot solely be attributed to the absence of a specific receptor in Drosophila.
Fingolimod (also known as FTY720) was approved as the first oral drug for the treatment of multiple sclerosis by the US-FDA in 2010. Previous studies have demonstrated the protective effects of fingolimod in SOD1-ALS mice, where it modulated microglial activation and innate immunity [70]. Additionally, fingolimod has shown safety and tolerability in phase 2 clinical trials for ALS (ClinicalTrials.gov Identifier: NCT01786174) [71]. The well-established mechanism of action for fingolimod involves its binding to S1P receptors expressed on various cells, including neurons, astrocytes, microglia, and endothelial cells of blood vessels [72, 73]. However, an intriguing aspect of our study is the use of Drosophila melanogaster, which lacks the sphingosine-1-phosphate (S1P) receptor, allowing us to explore the receptor-independent functions of fingolimod. To the best of our knowledge, our study is the first to reveal a positive functional interaction between fingolimod and TDP-43 independent of S1P receptor.
Among the receptor-independent effects described thus far, there is growing evidence suggesting a role for fingolimod in epigenetic modulations [74,75,76]. Specifically, the phosphorylated form of fingolimod, generated by the activities of evolutionary conserved sphingosine kinases SK1 and SK2 [77], has been shown to act as a specific inhibitor of class I HDACs, including HDAC1 [75, 78, 79]. Intriguingly, recent findings have revealed a connection between HDAC1 and TDP-43, whereby HDAC1 becomes sequestered within cytoplasmic aggregates formed by TDP-43, resulting in the loss of its nuclear functions related to DNA repair and cell cycle regulation [59]. Consequently, genetic, and pharmacological manipulations targeting HDAC1 have demonstrated their potential to improve TDP-43-induced cell death both in vitro and in vivo [80]. Furthermore, HDAC1 inhibition has been recently shown to promote the degradation of TDP-43 [81].
Based on the emerging evidence suggesting a link between fingolimod and HDAC1, along with the previously reported genetic and physical interactions between HDAC1 and TDP-43, it is plausible to attribute the ability of fingolimod to reduce the level of TDP-43 to its inhibitory effect on HDAC1 activity. The evolutionary conservation of HDCA1 across mammals and Drosophila, with a similarity of 87% and an identity of 77% as retrieved from the Drosophila Integrative Ortholog Prediction Tool (DIOPT), strongly supports the possibility of functional conservation or similarity between the two species. As a result, the observed extension of survival and improvement in motor functions may be attributed at least in part to a potential regulatory role of fingolimod on TDP-43 via HDAC1. This hypothesis will require further studies to be validated, particularly those employing human cell data or patient-derived cells to translate these novel findings to TDP-43 pathology in ALS patients. Currently, fingolimod is undergoing clinical trials for ALS, but the precise mechanism of its beneficial effects on the disease remains unknown. Therefore, the discovery of fingolimod's receptor-independent benefits for TDP-43 pathology, as demonstrated in this study, encourages further investigation in this direction. Drosophila could play a critical role in these pioneering efforts, advancing our understanding of fingolimod's mechanisms and its potential application for TDP-43-related disorders.
The regulatory effect observed for fingolimod on TDP-43 is likely to be similar for both TDP-43 WT and G348C variants, as evidenced by the similar trend of decreased TDP-43 protein levels following fingolimod administration. However, due to the higher expression level of TDP-43 G348C in these fly lines compared to TDP-43 WT, the rescue of locomotive functions in the mutant flies is only achieved when a half dose of the RU-486 inducer is provided. The higher expression of TDP-43 G348C compared to TDP-43 WT may be attributed to multiple factors. One possible reason is the random insertion of the transgenes in the fly's genome, which by chance may result in the G348C variant being in a more active chromatin locus. Additionally, the biochemical properties of the G348C mutant form of TDP-43 may contribute to its higher expression. The substitution of glycine with cysteine at position 348 is known to increase the propensity of TDP-43 to aggregate, making it more prone to form protein clumps. This aggregation can render TDP-43 more resistant to proteasomal degradation, thereby extending its half-life and ultimately resulting in higher levels of the G348C variant compared to the WT form [40].
The idea that fingolimod could regulate TDP43 levels in both TDP-43 WT and G348C flies appears to be valid, despite a few observed differences. Specifically, when fingolimod treatment was provided at the embryo stage in flies expressing moderately toxic TDP-43 WT or G348C in motor neurons, there were differences in the time required for treatment to improve climbing ability. The treatment needed at least 10 days for TDP-43 WT flies and 15 days for G348C flies. Additionally, while the efficacy of the treatment diminished after 15 days in TDP-43 WT flies, it seemed to remain beneficial in TDP-43 G348C flies until 20 days post-treatment. In addition to the biochemical properties that make G348C more resistant to degradation, as mentioned earlier, to interpret these findings, it is important to consider the limitations related to the longevity of untreated flies. Flies expressing TDP-43 WT or G348C in motor neurons have poor survival beyond 20 days, and the number of flies that survive until that time point is very limited. Moreover, the climbing score of untreated TDP-43 G348C flies experienced a sudden drop at the 20-day point of analysis, making statistical comparison between untreated and treated flies challenging and prone to misinterpretation.
The discrepancy between the observed improvement in locomotion and the lack of improvement in the morphology of neuromuscular junctions (NMJs) in these experimental conditions raised an intriguing question. There could be several plausible explanations for this discrepancy including time frame and mechanistic differences. For instance, the fingolimod-induced improvement in locomotion may occur rapidly, while changes in NMJ morphology may require a longer time to manifest. It is possible that the drug's effects on locomotion are independent of NMJ morphology and involve other mechanisms or pathways. Further investigations may be necessary to clarify the underlying mechanisms responsible for the fingolimod-induced locomotor improvements independent of NMJ morphology changes.
Taken together, this study identified fingolimod as a novel promising therapeutic for TDP-43 pathology, thus encouraging further clinical investigations to repurpose its use on a wide spectrum of neurodegenerative diseases.
Data Availability
The authors confirm that the data supporting the findings of this study are available within the article and its supplementary material.
References
Stover CM, Lynch NJ, Hanson SJ, Windbichler M, Gregory SG, Schwaeble WJ. Organization of the MASP2 locus and its expression profile in mouse and rat. Mamm Genome. 2004;15(11):887–900. https://doi.org/10.1007/s00335-004-3006-8.
Ayala YM, Zago P, D’Ambrogio A, Xu YF, Petrucelli L, Buratti E, et al. Structural determinants of the cellular localization and shuttling of TDP-43. J Cell Sci. 2008;121(Pt 22):3778–85. https://doi.org/10.1242/jcs.038950.
Nishimura AL, Zupunski V, Troakes C, Kathe C, Fratta P, Howell M, et al. Nuclear import impairment causes cytoplasmic trans-activation response DNA-binding protein accumulation and is associated with frontotemporal lobar degeneration. Brain. 2010;133(Pt 6):1763–71. https://doi.org/10.1093/brain/awq111.
Winton MJ, Igaz LM, Wong MM, Kwong LK, Trojanowski JQ, Lee VM. Disturbance of nuclear and cytoplasmic TAR DNA-binding protein (TDP-43) induces disease-like redistribution, sequestration, and aggregate formation. J Biol Chem. 2008;283(19):13302–9. https://doi.org/10.1074/jbc.M800342200.
Ederle H, Funk C, Abou-Ajram C, Hutten S, Funk EBE, Kehlenbach RH, et al. Nuclear egress of TDP-43 and FUS occurs independently of Exportin-1/CRM1. Sci Rep. 2018;8(1):7084. https://doi.org/10.1038/s41598-018-25007-5.
Polymenidou M, Lagier-Tourenne C, Hutt KR, Huelga SC, Moran J, Liang TY, et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat Neurosci. 2011;14(4):459–68. https://doi.org/10.1038/nn.2779.
Tollervey JR, Curk T, Rogelj B, Briese M, Cereda M, Kayikci M, et al. Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat Neurosci. 2011;14(4):452–8. https://doi.org/10.1038/nn.2778.
Ling JP, Pletnikova O, Troncoso JC, Wong PC. TDP-43 repression of nonconserved cryptic exons is compromised in ALS-FTD. Science. 2015;349(6248):650–5. https://doi.org/10.1126/science.aab0983.
Li Y, Ray P, Rao EJ, Shi C, Guo W, Chen X, et al. A Drosophila model for TDP-43 proteinopathy. Proc Natl Acad Sci U S A. 2010;107(7):3169–74. https://doi.org/10.1073/pnas.0913602107.
Neumann M, Sampathu DM, Kwong LK, Truax AC, Micsenyi MC, Chou TT, et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science. 2006;314(5796):130–3. https://doi.org/10.1126/science.1134108.
Arai T, Hasegawa M, Akiyama H, Ikeda K, Nonaka T, Mori H, et al. TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem Biophys Res Commun. 2006;351(3):602–11. https://doi.org/10.1016/j.bbrc.2006.10.093.
Garcia Morato J, Hans F, von Zweydorf F, Feederle R, Elsasser SJ, Skodras AA, et al. Sirtuin-1 sensitive lysine-136 acetylation drives phase separation and pathological aggregation of TDP-43. Nat Commun. 2022;13(1):1223. https://doi.org/10.1038/s41467-022-28822-7.
Zhang YJ, Xu YF, Cook C, Gendron TF, Roettges P, Link CD, et al. Aberrant cleavage of TDP-43 enhances aggregation and cellular toxicity. Proc Natl Acad Sci U S A. 2009;106(18):7607–12. https://doi.org/10.1073/pnas.0900688106.
Wang YT, Kuo PH, Chiang CH, Liang JR, Chen YR, Wang S, et al. The truncated C-terminal RNA recognition motif of TDP-43 protein plays a key role in forming proteinaceous aggregates. J Biol Chem. 2013;288(13):9049–57. https://doi.org/10.1074/jbc.M112.438564.
Chen AK, Lin RY, Hsieh EZ, Tu PH, Chen RP, Liao TY, et al. Induction of amyloid fibrils by the C-terminal fragments of TDP-43 in amyotrophic lateral sclerosis. J Am Chem Soc. 2010;132(4):1186–7. https://doi.org/10.1021/ja9066207.
Chang Z, Deng J, Zhao W, Yang J. Amyloid-like aggregation and fibril core determination of TDP-43 C-terminal domain. Biochem Biophys Res Commun. 2020;532(4):655–61. https://doi.org/10.1016/j.bbrc.2020.08.096.
Babinchak WM, Haider R, Dumm BK, Sarkar P, Surewicz K, Choi JK, et al. The role of liquid-liquid phase separation in aggregation of the TDP-43 low-complexity domain. J Biol Chem. 2019;294(16):6306–17. https://doi.org/10.1074/jbc.RA118.007222.
Briese M, Saal-Bauernschubert L, Luningschror P, Moradi M, Dombert B, Surrey V, et al. Loss of Tdp-43 disrupts the axonal transcriptome of motoneurons accompanied by impaired axonal translation and mitochondria function. Acta Neuropathol Commun. 2020;8(1):116. https://doi.org/10.1186/s40478-020-00987-6.
Hasan R, Humphrey J, Bettencourt C, Newcombe J, Consortium NA, Lashley T, et al. Transcriptomic analysis of frontotemporal lobar degeneration with TDP-43 pathology reveals cellular alterations across multiple brain regions. Acta Neuropathol. 2022;143(3):383–401. https://doi.org/10.1007/s00401-021-02399-9.
Hunter M, Spiller KJ, Dominique MA, Xu H, Hunter FW, Fang TC, et al. Microglial transcriptome analysis in the rNLS8 mouse model of TDP-43 proteinopathy reveals discrete expression profiles associated with neurodegenerative progression and recovery. Acta Neuropathol Commun. 2021;9(1):140. https://doi.org/10.1186/s40478-021-01239-x.
Ma XR, Prudencio M, Koike Y, Vatsavayai SC, Kim G, Harbinski F, et al. TDP-43 represses cryptic exon inclusion in the FTD-ALS gene UNC13A. Nature. 2022;603(7899):124–30. https://doi.org/10.1038/s41586-022-04424-7.
Romano G, Klima R, Feiguin F. TDP-43 prevents retrotransposon activation in the Drosophila motor system through regulation of Dicer-2 activity. BMC Biol. 2020;18(1):82. https://doi.org/10.1186/s12915-020-00816-1.
Tam OH, Rozhkov NV, Shaw R, Kim D, Hubbard I, Fennessey S, et al. Postmortem Cortex Samples Identify Distinct Molecular Subtypes of ALS: Retrotransposon Activation, Oxidative Stress, and Activated Glia. Cell Rep. 2019;29(5):1164–77 e5. https://doi.org/10.1016/j.celrep.2019.09.066.
Altman T, Ionescu A, Ibraheem A, Priesmann D, Gradus-Pery T, Farberov L, et al. Axonal TDP-43 condensates drive neuromuscular junction disruption through inhibition of local synthesis of nuclear encoded mitochondrial proteins. Nat Commun. 2021;12(1):6914. https://doi.org/10.1038/s41467-021-27221-8.
Birsa N, Bentham MP, Fratta P. Cytoplasmic functions of TDP-43 and FUS and their role in ALS. Semin Cell Dev Biol. 2020;99:193–201. https://doi.org/10.1016/j.semcdb.2019.05.023.
Riemenschneider H, Guo Q, Bader J, Frottin F, Farny D, Kleinberger G, et al. Gel-like inclusions of C-terminal fragments of TDP-43 sequester stalled proteasomes in neurons. EMBO Rep. 2022;23(6):e53890. https://doi.org/10.15252/embr.202153890.
Suk TR, Rousseaux MWC. The role of TDP-43 mislocalization in amyotrophic lateral sclerosis. Mol Neurodegener. 2020;15(1):45. https://doi.org/10.1186/s13024-020-00397-1.
Amador-Ortiz C, Lin WL, Ahmed Z, Personett D, Davies P, Duara R, et al. TDP-43 immunoreactivity in hippocampal sclerosis and Alzheimer’s disease. Ann Neurol. 2007;61(5):435–45. https://doi.org/10.1002/ana.21154.
Borroni B, Bonvicini C, Alberici A, Buratti E, Agosti C, Archetti S, et al. Mutation within TARDBP leads to frontotemporal dementia without motor neuron disease. Hum Mutat. 2009;30(11):E974–83. https://doi.org/10.1002/humu.21100.
Davidson Y, Amin H, Kelley T, Shi J, Tian J, Kumaran R, et al. TDP-43 in ubiquitinated inclusions in the inferior olives in frontotemporal lobar degeneration and in other neurodegenerative diseases: a degenerative process distinct from normal ageing. Acta Neuropathol. 2009;118(3):359–69. https://doi.org/10.1007/s00401-009-0526-z.
Huang W, Zhou Y, Tu L, Ba Z, Huang J, Huang N, et al. TDP-43: From Alzheimer’s Disease to Limbic-Predominant Age-Related TDP-43 Encephalopathy. Front Mol Neurosci. 2020;13:26. https://doi.org/10.3389/fnmol.2020.00026.
Josephs KA, Murray ME, Whitwell JL, Parisi JE, Petrucelli L, Jack CR, et al. Staging TDP-43 pathology in Alzheimer’s disease. Acta Neuropathol. 2014;127(3):441–50. https://doi.org/10.1007/s00401-013-1211-9.
Kabashi E, Valdmanis PN, Dion P, Spiegelman D, McConkey BJ, Vande Velde C, et al. TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat Genet. 2008;40(5):572–4. https://doi.org/10.1038/ng.132.
Nelson PT, Dickson DW, Trojanowski JQ, Jack CR, Boyle PA, Arfanakis K, et al. Limbic-predominant age-related TDP-43 encephalopathy (LATE): consensus working group report. Brain. 2019;142(6):1503–27. https://doi.org/10.1093/brain/awz099.
Schwab C, Arai T, Hasegawa M, Yu S, McGeer PL. Colocalization of transactivation-responsive DNA-binding protein 43 and huntingtin in inclusions of Huntington disease. J Neuropathol Exp Neurol. 2008;67(12):1159–65. https://doi.org/10.1097/NEN.0b013e31818e8951.
Sreedharan J, Blair IP, Tripathi VB, Hu X, Vance C, Rogelj B, et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science. 2008;319(5870):1668–72. https://doi.org/10.1126/science.1154584.
Steinacker P, Barschke P, Otto M. Biomarkers for diseases with TDP-43 pathology. Mol Cell Neurosci. 2019;97:43–59. https://doi.org/10.1016/j.mcn.2018.10.003.
Van Deerlin VM, Leverenz JB, Bekris LM, Bird TD, Yuan W, Elman LB, et al. TARDBP mutations in amyotrophic lateral sclerosis with TDP-43 neuropathology: a genetic and histopathological analysis. Lancet Neurol. 2008;7(5):409–16. https://doi.org/10.1016/S1474-4422(08)70071-1.
Kuhnlein P, Sperfeld AD, Vanmassenhove B, Van Deerlin V, Lee VM, Trojanowski JQ, et al. Two German kindreds with familial amyotrophic lateral sclerosis due to TARDBP mutations. Arch Neurol. 2008;65(9):1185–9. https://doi.org/10.1001/archneur.65.9.1185.
Watanabe S, Kaneko K, Yamanaka K. Accelerated disease onset with stabilized familial amyotrophic lateral sclerosis (ALS)-linked mutant TDP-43 proteins. J Biol Chem. 2013;288(5):3641–54. https://doi.org/10.1074/jbc.M112.433615.
Yang C, Tan W, Whittle C, Qiu L, Cao L, Akbarian S, et al. The C-terminal TDP-43 fragments have a high aggregation propensity and harm neurons by a dominant-negative mechanism. PLoS One. 2010;5(12).
Mackenzie IR, Bigio EH, Ince PG, Geser F, Neumann M, Cairns NJ, et al. Pathological TDP-43 distinguishes sporadic amyotrophic lateral sclerosis from amyotrophic lateral sclerosis with SOD1 mutations. Ann Neurol. 2007;61(5):427–34. https://doi.org/10.1002/ana.21147.
Hayes LR, Kalab P. Emerging Therapies and Novel Targets for TDP-43 Proteinopathy in ALS/FTD. Neurotherapeutics. 2022. https://doi.org/10.1007/s13311-022-01260-5.
Buratti E. Targeting TDP-43 proteinopathy with drugs and drug-like small molecules. Br J Pharmacol. 2021;178(6):1298–315. https://doi.org/10.1111/bph.15148.
Liguori F, Amadio S, Volonte C. Fly for ALS: Drosophila modeling on the route to amyotrophic lateral sclerosis modifiers. Cell Mol Life Sci. 2021;78(17–18):6143–60. https://doi.org/10.1007/s00018-021-03905-8.
Yamaguchi M, Lee IS, Jantrapirom S, Suda K, Yoshida H. Drosophila models to study causative genes for human rare intractable neurological diseases. Exp Cell Res. 2021;403(1).
Appocher C, Mohagheghi F, Cappelli S, Stuani C, Romano M, Feiguin F, et al. Major hnRNP proteins act as general TDP-43 functional modifiers both in Drosophila and human neuronal cells. Nucleic Acids Res. 2017;45(13):8026–45. https://doi.org/10.1093/nar/gkx477.
Azpurua J, El-Karim EG, Tranquille M, Dubnau J. A behavioral screen for mediators of age-dependent TDP-43 neurodegeneration identifies SF2/SRSF1 among a group of potent suppressors in both neurons and glia. PLoS Genet. 2021;17(11).
Deng X, Sun X, Yue W, Duan Y, Hu R, Zhang K, et al. CHMP2B regulates TDP-43 phosphorylation and cytotoxicity independent of autophagy via CK1. J Cell Biol. 2022;221(1). https://doi.org/10.1083/jcb.202103033.
Kim HJ, Raphael AR, LaDow ES, McGurk L, Weber RA, Trojanowski JQ, et al. Therapeutic modulation of eIF2alpha phosphorylation rescues TDP-43 toxicity in amyotrophic lateral sclerosis disease models. Nat Genet. 2014;46(2):152–60. https://doi.org/10.1038/ng.2853.
Pons M, Prieto S, Miguel L, Frebourg T, Campion D, Sune C, et al. Identification of TCERG1 as a new genetic modulator of TDP-43 production in Drosophila. Acta Neuropathol Commun. 2018;6(1):138. https://doi.org/10.1186/s40478-018-0639-5.
Sreedharan J, Neukomm LJ, Brown RH Jr, Freeman MR. Age-Dependent TDP-43-Mediated Motor Neuron Degeneration Requires GSK3, hat-trick, and xmas-2. Curr Biol. 2015;25(16):2130–6. https://doi.org/10.1016/j.cub.2015.06.045.
Zhan L, Hanson KA, Kim SH, Tare A, Tibbetts RS. Identification of genetic modifiers of TDP-43 neurotoxicity in Drosophila. PLoS One. 2013;8(2).
Cheng CW, Lin MJ, Shen CK. Rapamycin alleviates pathogenesis of a new Drosophila model of ALS-TDP. J Neurogenet. 2015;29(2–3):59–68. https://doi.org/10.3109/01677063.2015.1077832.
Cragnaz L, Spinelli G, De Conti L, Bureau EA, Brownlees J, Feiguin F, et al. Thioridazine reverts the phenotype in cellular and Drosophila models of amyotrophic lateral sclerosis by enhancing TDP-43 aggregate clearance. Neurobiol Dis. 2021;160.
Francois-Moutal L, Felemban R, Scott DD, Sayegh MR, Miranda VG, Perez-Miller S, et al. Small Molecule Targeting TDP-43’s RNA Recognition Motifs Reduces Locomotor Defects in a Drosophila Model of Amyotrophic Lateral Sclerosis (ALS). ACS Chem Biol. 2019;14(9):2006–13. https://doi.org/10.1021/acschembio.9b00481.
Joardar A, Menzl J, Podolsky TC, Manzo E, Estes PS, Ashford S, et al. PPAR gamma activation is neuroprotective in a Drosophila model of ALS based on TDP-43. Hum Mol Genet. 2015;24(6):1741–54. https://doi.org/10.1093/hmg/ddu587.
Mollasalehi N, Francois-Moutal L, Scott DD, Tello JA, Williams H, Mahoney B, et al. An Allosteric Modulator of RNA Binding Targeting the N-Terminal Domain of TDP-43 Yields Neuroprotective Properties. ACS Chem Biol. 2020;15(11):2854–9. https://doi.org/10.1021/acschembio.0c00494.
Sanna S, Esposito S, Masala A, Sini P, Nieddu G, Galioto M, et al. HDAC1 inhibition ameliorates TDP-43-induced cell death in vitro and in vivo. Cell Death Dis. 2020;11(5):369. https://doi.org/10.1038/s41419-020-2580-3.
Brand AH, Perrimon N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development. 1993;118(2):401–15. https://doi.org/10.1242/dev.118.2.401.
Spradling AC, Rubin GM. Transposition of cloned P elements into Drosophila germ line chromosomes. Science. 1982;218(4570):341–7. https://doi.org/10.1126/science.6289435.
Robertson HM, Preston CR, Phillis RW, Johnson-Schlitz DM, Benz WK, Engels WR. A stable genomic source of P element transposase in Drosophila melanogaster. Genetics. 1988;118(3):461–70. https://doi.org/10.1093/genetics/118.3.461.
Osterwalder T, Yoon KS, White BH, Keshishian H. A conditional tissue-specific transgene expression system using inducible GAL4. Proc Natl Acad Sci U S A. 2001;98(22):12596–601. https://doi.org/10.1073/pnas.221303298.
Brooks DS, Vishal K, Kawakami J, Bouyain S, Geisbrecht ER. Optimization of wrMTrck to monitor Drosophila larval locomotor activity. J Insect Physiol. 2016;93–94:11–7. https://doi.org/10.1016/j.jinsphys.2016.07.007.
Jantrapirom S, Enomoto Y, Karinchai J, Yamaguchi M, Yoshida H, Fukusaki E, et al. The depletion of ubiquilin in Drosophila melanogaster disturbs neurochemical regulation to drive activity and behavioral deficits. Sci Rep. 2020;10(1):5689. https://doi.org/10.1038/s41598-020-62520-y.
Jantrapirom S, Lo Piccolo L, Yoshida H, Yamaguchi M. A new Drosophila model of Ubiquilin knockdown shows the effect of impaired proteostasis on locomotive and learning abilities. Exp Cell Res. 2018;362(2):461–71. https://doi.org/10.1016/j.yexcr.2017.12.010.
Cragnaz L, Klima R, Skoko N, Budini M, Feiguin F, Baralle FE. Aggregate formation prevents dTDP-43 neurotoxicity in the Drosophila melanogaster eye. Neurobiol Dis. 2014;71:74–80. https://doi.org/10.1016/j.nbd.2014.07.009.
Lo Piccolo L, Bonaccorso R, Onorati MC. Nuclear and Cytoplasmic Soluble Proteins Extraction from a Small Quantity of Drosophila’s Whole Larvae and Tissues. Int J Mol Sci. 2015;16(6):12360–7. https://doi.org/10.3390/ijms160612360.
Langellotti S, Romano V, Romano G, Klima R, Feiguin F, Cragnaz L, et al. A novel Drosophila model of TDP-43 proteinopathies: N-terminal sequences combined with the Q/N domain induce protein functional loss and locomotion defects. Dis Model Mech. 2016;9(6):659–69. https://doi.org/10.1242/dmm.023382.
Potenza RL, De Simone R, Armida M, Mazziotti V, Pezzola A, Popoli P, et al. Fingolimod: A Disease-Modifier Drug in a Mouse Model of Amyotrophic Lateral Sclerosis. Neurotherapeutics. 2016;13(4):918–27. https://doi.org/10.1007/s13311-016-0462-2.
Berry JD, Paganoni S, Atassi N, Macklin EA, Goyal N, Rivner M, et al. Phase IIa trial of fingolimod for amyotrophic lateral sclerosis demonstrates acceptable acute safety and tolerability. Muscle Nerve. 2017;56(6):1077–84. https://doi.org/10.1002/mus.25733.
Xu Z, Ikuta T, Kawakami K, Kise R, Qian Y, Xia R, et al. Structural basis of sphingosine-1-phosphate receptor 1 activation and biased agonism. Nat Chem Biol. 2022;18(3):281–8. https://doi.org/10.1038/s41589-021-00930-3.
Singh SK, Kordula T, Spiegel S. Neuronal contact upregulates astrocytic sphingosine-1-phosphate receptor 1 to coordinate astrocyte-neuron cross communication. Glia. 2022;70(4):712–27. https://doi.org/10.1002/glia.24135.
Saddoughi SA, Gencer S, Peterson YK, Ward KE, Mukhopadhyay A, Oaks J, et al. Sphingosine analogue drug FTY720 targets I2PP2A/SET and mediates lung tumour suppression via activation of PP2A-RIPK1-dependent necroptosis. EMBO Mol Med. 2013;5(1):105–21. https://doi.org/10.1002/emmm.201201283.
Hait NC, Wise LE, Allegood JC, O’Brien M, Avni D, Reeves TM, et al. Active, phosphorylated fingolimod inhibits histone deacetylases and facilitates fear extinction memory. Nat Neurosci. 2014;17(7):971–80. https://doi.org/10.1038/nn.3728.
Leo A, Citraro R, Amodio N, De Sarro C, Gallo Cantafio ME, Constanti A, et al. Fingolimod Exerts only Temporary Antiepileptogenic Effects but Longer-Lasting Positive Effects on Behavior in the WAG/Rij Rat Absence Epilepsy Model. Neurotherapeutics. 2017;14(4):1134–47. https://doi.org/10.1007/s13311-017-0550-y.
Herr DR, Fyrst H, Creason MB, Phan VH, Saba JD, Harris GL. Characterization of the Drosophila sphingosine kinases and requirement for Sk2 in normal reproductive function. J Biol Chem. 2004;279(13):12685–94. https://doi.org/10.1074/jbc.M310647200.
Hait NC, Avni D, Yamada A, Nagahashi M, Aoyagi T, Aoki H, et al. The phosphorylated prodrug FTY720 is a histone deacetylase inhibitor that reactivates ERalpha expression and enhances hormonal therapy for breast cancer. Oncogenesis. 2015;4.
Hashemian M, Ghasemi-Kasman M, Parsian H, Sadeghi F. Fingolimod (FTY720) improves the functional recovery and myelin preservation of the optic pathway in focal demyelination model of rat optic chiasm. Brain Res Bull. 2019;153:109–21. https://doi.org/10.1016/j.brainresbull.2019.08.014.
Wu CC, Jin LW, Wang IF, Wei WY, Ho PC, Liu YC, et al. HDAC1 dysregulation induces aberrant cell cycle and DNA damage in progress of TDP-43 proteinopathies. EMBO Mol Med. 2020;12(6):e10622. https://doi.org/10.15252/emmm.201910622.
Luo K, Wang Z, Zhuang K, Yuan S, Liu F, Liu A. Suberoylanilide hydroxamic acid suppresses axonal damage and neurological dysfunction after subarachnoid hemorrhage via the HDAC1/HSP70/TDP-43 axis. Exp Mol Med. 2022. https://doi.org/10.1038/s12276-022-00761-9.
Acknowledgements
We thank Ms Sureena Phosa, Ms Pawita Chaiya, Ms Titaree Yamsrii, Mr. Phatarawat Thaklaewphan, Dr. Phateep Hankittichai and the executive staffs at CMUTEAM for their support to facilitate laboratory procedures and administrative tasks at the Faculty of Medicine, Chiang Mai University. We also thank Dr. Ryo Tanaka and Dr. Itaru Yamamoto for their precious help and technical advice at the Kyoto Institute of Technology, Japan.
Funding
This study was supported by National Research Council of Thailand (NRCT5-TRG63004-02 to SJ), Genomics Thailand, Health Systems Research Institute (HSRI) (grant 64/122 and 65/036 to LLP), and the JSPS Core-to-Core Program, Asia-Africa Science Platform B (to MY and HY).
Author information
Authors and Affiliations
Contributions
Conceptualization and Organization: MY, YG, HY, SJ, LLP, Execution and analysis: SJ, TU, RY, LLP, Manuscript preparation: SJ, MY, LLP, Review and critique: LLP, TU, RY, SP, WN, VP, YG, HY, MY, SJ.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lo Piccolo, L., Umegawachi, T., Yeewa, R. et al. A Novel Drosophila-based Drug Repurposing Platform Identified Fingolimod As a Potential Therapeutic for TDP-43 Proteinopathy. Neurotherapeutics 20, 1330–1346 (2023). https://doi.org/10.1007/s13311-023-01406-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13311-023-01406-z