Abstract
Neuronal as well as glial cells contribute to higher order brain functions. Many observations show that neurons and glial cells are not only physically highly intermingled but are physiologically tightly connected and mutually depend at various levels on each other. Moreover, macroglia classes like astrocytes, NG2 cells and oligodendrocytes are not at all homogenous cell populations but do possess a markedly heterogeneity in various aspects similar to neurons. The diversity of differences in morphology, functionality and, cellular activity has been acknowledged recently and will be integrated into a concept of brain function that pictures a neural rather than a puristical neuronal world. With the recent progress in “omic” technologies, an unbiased and exploratory approach toward an enhanced understanding of glial heterogeneity has become possible. Here, we provide an overview on current technical transcriptomic and proteomic approaches used to dissect glial heterogeneity of the brain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Increasing evidence suggest that both neuronal and glial heterogeneity contribute to and are essential for higher order brain functions. A plethora of observations have been made showing that neurons and glial cells are not only physically highly intermingled but are physiologically tightly connected and mutually depend at various levels on each other. For example, astroglial cells are essential for maintaining the metabolic integrity of neurons and participate in synaptic transmission (“tripartite synapse”) [1, 4]. Therefore, the view emerges that disturbed neuron–glia interactions may contribute to the initiation and progression of neurodegenerative and psychiatric diseases. Intriguingly and similarly to neurons, macroglia classes like astrocytes, NG2 cells, and oligodendrocytes are not at all homogenous cell populations but do possess a markedly heterogeneity in morphology, functionality, and cellular activity as well, that only recently has been started to be acknowledged and integrated into a concept of brain function that pictures a neural world rather than a puristical neuronal world. In the light of these findings, it is not too surprising that glial cells are also increasingly considered as attractive targets for clinical intervention strategies aiming at re-balancing disturbed brain function.
So far, the application of conventional approaches to characterize neurons and more recently also glial cells at global levels by applying so-called -omic technologies has been limited for the brain [8]. This has several reasons: (1) the cellular complexity of the nervous system is extreme, with a high degree of regional variability of the different cell types and, hence, susceptibility in different diseases. (2) Under physiological and pathological conditions, the proper function of neurons cannot be dissociated from supporting glial cells—and vice versa. (3) Most neurodegenerative and psychiatric disorders are late-onset chronic “systems-diseases” that affect different neuron–glia networks, which complicates comparison between different studies (4). The analysis of brain tissue is still state of the art for molecular and biochemical studies, although cell-type relevant processes are likely to be masked due to the fact that all cells share a rather large pool of identical proteins ranging from housekeeping proteins, protein signaling cascades, adhesion molecules, and receptors and ion channels. Because of these facts, several methods needed to be developed and are still in progress of refined development that allow to specifically label and isolate either different glial and neuronal cells or cell-specific sub-cellular compartments to high purity and at conditions that allow for the application of state-of-the art -omic approaches. The appendix “omics” added to the class of biological molecules under investigation circumscribes respective globally scaled technologies, such as Genomics, Transcriptomics, Proteomics, Lipidomics, and Metabolomics. By definition, omics technologies claim to cover the majority of different molecules of a given subclass within one experiment providing a systems-level view onto a biological sample. Thus, unbiased and exploratory (“hypothesis-generating”) approaches toward an enhanced understanding of physiological or pathological cellular states are possible. Subsequently, we will provide an overview on current technical approaches used to perform transcriptomics and proteomics to dissect glial heterogeneity of the brain.
Strategies to isolate neuronal and glial cell types of the brain
Defined ex vivo culture conditions have been developed for all major cell types of the brain, such as astrocytes, oligodendrocyte precursor cells (OPCs or NG2 glia), mature oligodendrocytes, and different neuronal subtypes as well as microglia. Thus, it is possible to culture large quantities of defined cell types for extended periods in vitro. Although the basic electrophysiological and molecular characteristics of cultured neuronal and glial cells appear to be sustained in vitro, it is widely accepted that ex vivo cultures will only partially reflect the in vivo situation even when different culture conditions are employed and compared. The molecular identity of a given cell type depends not only on the availability of nutrients and soluble survival factors (“media and supplements”) but also critically on regional cues depending for example on the nature and activity of surrounding cells, composition of the local extracellular matrix as well as many other factors. Thus, the local microenvironment determines the molecular and structural features of cells in vivo, which cannot be exactly mirrored in vitro. For globally scaled proteomics and also the analysis of noncoding RNAs (ncRNAs), however, large homogenous pools of intact cells including all compartments and membranous extrusions (filopodia, neurites, etc.) may be the preferred substrate for analysis. Under these circumstances, ex vivo cultures for example of different glial cells—both primary and cell lines—at high purity are still the substrate of choice to date.
To overcome the limitations of ex vivo culturing and enrichment, several isolation techniques have been developed. A powerful method for isolating small defined brain regions and individual cell-types is “Laser-capture microdissection” (LCM), which combines high-resolution microscopy with precise microlaser cutting in one single device. Simple Nissl-staining can be applied to precisely guide regional selection and to identify a few neuronal cell types such as large-sized Purkinje cells of the cerebellum and motor neurons of the spinal cord [13, 19]. We have adapted this technique by using a nuclear targeted fluorescent protein to genetically label projection neurons of the adult cortex and could show that samples with cellular resolution revealed transcriptomic signatures, which were masked even in layer-specific cortical microregions [19]. Nonetheless, LCM is considered rather an enrichment than purification strategy given the fact that for example minor proportions of glial transcripts originating most likely from cellular processes will be co-isolated from semithin cryosections. Small-sized glial cells are at the limit of the resolution of the technique. Moreover, due to the low amount of isolated sample globally scaled proteomics has not been combined with LCM purification.
A more widely applied and less labor intensive isolation technique, particularly for small-sized glial cells, relies on tissue trituration followed by fluorescence-activated cell sorting (FACS). This strategy has been successfully used by us and others in combination with quantitative real-time polymerase chain reaction, microarray analysis, and more recently RNAseq to profile mRNA expression of glial cell populations [3, 12, 14, 15]. Both, transgenic labels introducing for example green fluorescent protein in astrocytes and antibodies directed against extracellular epitopes have been applied [14]. The caveats of FACS mediated approaches are ex vivo incubation steps that need to be strictly standardized and which could potentially cause artifacts. The induction of immediate-early or stress-associated genes has, however, not been observed [14]. The advantage of this method is that all RNA species, mRNAs as well as noncoding transcripts, can be harvested simultaneously (see below). Of particular importance for regulatory mechanisms in nuclear organization and transcriptional, posttranscriptional as well as epigenetic processes in the brain appear to be small and long ncRNAs [18]. Comprehensive characterizations of these molecules have not been described in different glia populations so far. Given the enormous progress in the sensitivity and coverage of mass spectometry based technologies, globally scaled proteomic analyses for example of different FACS enriched cell populations are already on the way to deliver the full complement of all expressed proteins in neuronal and glial cell types.
A partially complementary approach relies on ribosome pull-down protocols that allow, by using a transgenically tagged ribosome component, to study selectively the translated transcriptome of a pool of defined cell types from different brain regions, however, ncRNAs cannot be analyzed [6].
Transcriptomic approaches to unravel glial heterogeneity
Transcriptomic analyses with DNA microarrays were among the most widely applied globally scaled genomic technologies and have been successfully used to profile neural gene expression in the context of normal brain physiology and disease, however, mostly not at the cell-type-specific level [8]. More recently, second or next generation sequencing technologies (“Deep-Sequencing”) principally allow obtaining a much deeper insight into the “whole transcriptome” in an unbiased matter [22]. With RNA-Seq, the detection and quantification of all expressed mRNA splice variants and ncRNAs is possible without the a priori knowledge of their exact nature. It has been shown that RNAseq provides a more accurate, quantitative, and sensitive analysis of global transcriptomes as compared with hybridization based methods such as microarrays [22]. The maximally meaningful application of these globally scaled technologies most likely requires a coordinated approach since particularly ncRNAs are essential players in the control of mRNA abundance and initiation of protein translation.
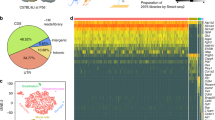
As a first step toward the characterization of region-specific molecular signatures of astrocytes, we have combined FACS with 3’ digital gene expression [20]. In this RNAseq approach, we selectively focused on mRNAs expression profiles of astrocytes FACS isolated from the cortex (Cx), hippocampus (Hi), and brainstem (Bs). We hypothesized that differential expression levels of genes encoding for transporter proteins correlating with the Cx- and Hi-specific uptake of sulforhodamine (SR101) into astrocytes would allow us to identify the SR101 transporter. From the identified candidates, Slco1c1 was validated with corresponding biochemical assays and mouse mutants as the Cx/Hi-specific astrocyte SR101 transporter (Fig. 1a–d) [20]. This study was purely hypothesis driven and utilized only a small fraction of the obtained data. Nonetheless, it clearly showed that transcriptomic datasets from different astrocytes population can help to better understand functional consequences of expression differences. By performing quality controls at all steps of the protocol (average r2 values of > 0.99 for technical replicates indicate a high level of reproducibility, see Fig. 1e) and by applying stringent statistical criteria, we are convinced that an unbiased bioinformatic analysis of the complete data set will reveal a deep insight into the regional differences of the transcriptomes of astrocytes isolated at P10 from Cx, Hi, and Bs. We can conclude from a preliminary analysis that astrocytes from these regions are highly heterogenous with respect to differential gene expression beyond transporter. The number of differentially expressed genes at a stringent cutoff (p value > 10−5) is high (Hi vs. Bs = 277, Hi vs. Cx = 1105, and Cx vs. Bs = 724), which most likely reflects several additional subregion-specific mechanisms operating in these astrocyte populations. A preliminary pathway analysis shows, for example, that—among many others—Ca2+ signaling-related gene sets are enriched in forebrain astrocytes (Kannayian and Rossner, unpublished).
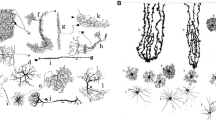
Identification of transporter genes enriched in forebrain astrocytes by transcriptome analysis. a Experimental strategy: Cortex (Cx), Hippocampus (Hi), and Brainstem (Bs) tissues from brains of mice expressing enhanced green fluorescent protein (EGFP) under the control of the glial-specific promoter glial fibrillary acidic protein (hGFAP-EGFP) were triturated and 50k EGFP positive cells were purified by fluorescence-activated cell sorting (FACS), followed by RNA isolation and transcriptome profiling with RNAseq. SR101 candidate uptake transporter genes (of the solute carrier, Slc family) were hypothesized to display a higher expression in Cx and Hi versus Bs astrocytes (Cx/Hi > Bs). b Scatter plots of FACS analysis with green fluorescent protein intensities plotted versus forward scatter (FSC) given as arbitrary units. Blue dots represent Hoechst 33342 positive and GFP negative cells, green dots represent Hoechst 33342 and GFP double positive cells representing different astrocyte populations. c Venn diagram of all forebrain-enriched transporter genes. Seven mRNAs were significantly elevated both in Cx and Hi. d Differentially expressed mRNAs coding for solute carriers (Slc and Slco gene families). Depicted are the gene symbols, accession, and average read numbers in Cx, Hi, and Bs samples. The seven detected forebrain-enriched mRNAs correspond to five genes (Slc1a2 and Slco1c1 annotated by two mRNAs each). Detection cut-off was determined with an adjusted p value cut-off < 0.05. Cx (n = 2), Hi (n = 3), Bs (n = 4) biological replicates. e Scatter plot analysis shows high degree of reproducibility of technical replicates, biological replicates, and astrocyte from different region as well as comparison of astrocytes and neurons display as expected increasing levels of divergence. f Inspection of ISH data (from Allen Brain Atlas) for top candidates being differentially expressed between Hi and Bs validates regional NGS profiles (a–d). (modified from (Schnell et al. 2013)
Proteomic approaches to unravel glial heterogeneity
With an estimated number of approximately 10,000 different proteins in each single mammalian cell [16], in-depth identification of a cell’s entire proteome is still a major challenge for modern proteomic technologies. In addition, the total number of different/unique proteins per synapse is estimated to be at approximately 2000–2500 [17], which are often under control of different activity-dependent turnover rates. While today’s state-of-the-art MS instruments routinely sequence single purified proteins with subfemtomolar sensitivity, the effective identification of low-abundance proteins is orders of magnitude lower in complex mixtures due to limited dynamic range and sequencing speed and due to the common, strong bias toward acquiring MS/MS data on higher abundance molecules. Hence, to tackle activity-dependent proteome alterations entire, functionally heterogeneous brain regions with different subtypes of neurons and macroglial cells are usually being sampled at the loss of cell-type specificity and spatial resolution. The characterization of a proteome is an even more difficult challenge if temporal and spatial aspects of a proteome or a subpopulation of the proteome have to be taken into consideration. Thus, the separation and enrichment of the subproteome in question is key for its successful characterization. To overcome the above mentioned limitations, several cellular and biochemical (subcellular) enrichment strategies combined with proteomics have been developed to increase sensitivity and selectivity for the analysis of neuronal and glial proteomes, and to finally dissolve cellular heterogeneity of neural cells in the brain (Fig. 2a-b).
Concerning cellular selectivity, intracellular proteomes and secretomes from cultured primary astrocytes and astroglial cell lines have been characterized in detail by several labs as proteomic approaches as indicated above require a much larger quantity than transcriptomic approaches. Analysis of primary cortical astrocytes revealed the astonishingly high number of proteins in conditioned media including 1247 putative secreted proteins [21]. More recently 6000 unique protein groups belonging to the secretome and 7265 unique intracellular protein groups were identified from cultured C8-D1A astrocytes [10]. Moreover, these experiments demonstrate activity-dependent changes in intracellular and secreted proteomes even under ex vitro conditions. Similar to in-depth analysis of postsynaptic and presynaptic protein fractions, biochemical fractionation approaches have been recently used to attain detailed pictures of defined subcellular glial proteomes, such as myelin membranes of oligodendrocytes or astroglial membrane fractions. Analysis of freshly purified human and murine myelin fractions revealed more than 1000 proteins and about 700 different lipids [9]. In a very recent study, Carney et al. characterized the proteome of gliosomes purified from murine brain tissue. Gliosomes are enriched for proteins of the astrocytic exocytosis machinery (marker protein VAMP3), perisynaptic astrocyte processes (marker protein Ezrin), and astrocyte plasma membrane proteins (e.g.astrocyte membrane glycoprotein Basigin44). While common synaptosomal proteins (such as the presynaptic SNARE proteins and postsynaptic proteins NR1 and PSD-95) were depleted in the gliosome fraction, the fraction contained glutamate receptors and a plethora of heteromeric G-protein subunits and small GTPases related to perisynaptic function. In combination with fluorescence-activated sorting, such as recently done for VGLUT1-Venus-labeled synaptosomes [2], specialized astrocytic gliosomes could be investigated in unprecedented detail in the future. Another approach to decipher glial proteomic heterogeneity might employ bioorthogonal metabolic protein labeling (Fig. 2c) as recently shown by us for different cell types including glial and neuronal cells in Drosophila melanogaster [7]. This technique relies on a mutated methionine tRNA synthetase (MetRSLtoG), which incorporates, instead of methionine, the noncanonical amino acid azidonorleucine (ANL) into nascent proteins. By transgenically expressing the MetRSLtoG cell-specificity can be achieved, whereas restricting feeding ANL to transgenic flies within a desired time enables temporal specificity. Thus, only in the cells expressing the mutated MetRSLtoG and only during the ANL feeding, the synthesized proteins incorporate ANL. Subsequently and depending of the type of analysis, ANL is clicked either to a fluorescent tag for visualization of ANL-harboring proteins (FUNCAT; [5]) or clicked to an affinity tag with subsequent analysis of tagged proteins using immunoblot or bulk identification via mass spectrometry (BONCAT [11]). Notably, this approach revealed for the first time the expression of the scaffolding protein Dlg in glia [5, 7] pointing once more to the importance of sub-type-specific proteome analysis for understanding brain function on the global and on the regional level.
Conclusions
Because of its apparent limitations, none of these methods qualifies as “the unique” and generally applicable strategy toward transcriptomic and proteomic approaches to unravel the molecular complexity for example of regional glial cell populations of the brain. Unresolved issues for proteomic approaches include the analysis of splice variants, alternative translation initiation sites, and point mutations of proteins due to both dynamic range and sequence coverage limitations. Finally, current proteomics is unable to simultaneously unlock all the critical determinants of cellular proteostasis because specific purification and separation procedures have to be employed for each single protein modifications making it necessary to address posttranslational modifications one by one. In parallel to the limitations mentioned for the proteomics, exploring the whole universe of RNA molecules is still challenging although sequencing methodologies have dramatically improved throughout the last decade. Nonetheless, kinetic and structural variations as well as for example allele-specific expression require sophisticated bioinformatics analyses, which are not yet available for all purposes.
Finally, to obtain a more complete picture and to have the opportunity to identify different technical biases, most likely several -omic and isolation strategies should be combined and analyzed in an integrated fashion to increase our understanding of glial heterogeneity of the brain.
Marcoglial proteomics approaches. Schematic strategy for the investigation of marcoglial proteomes. a In vitro cultivation of purified primary marcoglia cells or respective cell lines enables the analysis of the intracellular protein entity as well as secretomes, whereas fractionation and extraction procedures of brain homogenates b allows the identification of subcellular proteomes such as astroglial gliosomes, myelin, and others, however, at the expense of true cell-type specificity. Metabolic labeling of newly synthesized proteins using the noncanonical amino acid azidonorleucine (ANL) and a mutated methionine tRNA synthetase expressed in selected cell types may facility cell-type-specific proteome analysis in combination with “click chemistry” under in situ conditions prior biochemical and mass spectrometrical analyses
References
Allen NJ, Barres BA (2005) Signaling between glia and neurons: focus on synaptic plasticity. Curr Opin Neurobiol 15:542–548
Biesemann C, Grønborg M, Luquet E, Wichert SP, Bernard V, Bungers SR, Cooper B, Varoqueaux F, Li L, Byrne JA et al (2014) Proteomic screening of glutamatergic mouse brain synaptosomes isolated by fluorescence activated sorting. EMBO J 33:157–170
Cahoy JD, Emery B, Kaushal A, Foo LC, Zamanian JL, Christopherson KS, Xing Y, Lubischer JL, Krieg PA, Krupenko SA et al (2008) A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J Neurosci 28:264–278
Clarke LE, Barres BA (2013) Emerging roles of astrocytes in neural circuit development. Nat Rev Neurosci 14:311–321
Dieterich DC, Hodas JJL, Gouzer G, Shadrin IY, Ngo JT, Triller A, Tirrell DA, Schuman EM (2010) In situ visualization and dynamics of newly synthesized proteins in rat hippocampal neurons. Nat Neurosci 13:897–905
Doyle JP, Dougherty JD, Heiman M, Schmidt EF, Stevens TR, Ma G, Bupp S, Shrestha P, Shah RD, Doughty ML et al (2008) Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell 135:749–762
Erdmann I, Marter K, Kobler O, Niehues S, Abele J, Müller A, Bussmann J, Storkebaum E, Ziv T, Thomas U et al (2015) Cell-selective labeling of proteomes in Drosophila melanogaster. Nat Commun doi:10.1038/ncomms8521
Geschwind DH, Konopka G (2009) Neuroscience in the era of functional genomics and systems biology. Nature 461:908–915
Gopalakrishnan G, Awasthi A, Belkaid W, De Faria O, Liazoghli D, Colman DR, Dhaunchak AS (2013) Lipidome and proteome map of myelin membranes. J Neurosci Res 91:321–334
Han D, Jin J, Woo J, Min H, Kim Y (2014) Proteomic analysis of mouse astrocytes and their secretome by a combination of FASP and StageTip-based, high pH, reversed-phase fractionation. Proteomics 14:1604–1609
Hodas JJL, Nehring A, Höche N, Sweredoski MJ, Pielot R, Hess S, Tirrell DA, Dieterich DC, Schuman EM (2012) Dopaminergic modulation of the hippocampal neuropil proteome identified by bioorthogonal noncanonical amino acid tagging (BONCAT). Proteomics 12:2464–2476
Lalo U, Pankratov Y, Wichert SP, Rossner MJ, North RA, Kirchhoff F, Verkhratsky A (2008) P2 × 1 and P2 × 5 subunits form the functional P2X receptor in mouse cortical astrocytes. J Neurosci 28:5473–5480
Lobsiger CS, Boillée S, Cleveland DW (2007) Toxicity from different SOD1 mutants dysregulates the complement system and the neuronal regenerative response in ALS motor neurons. Proc Natl Acad Sci U S A 104:7319–7326
Lovatt D, Sonnewald U, Waagepetersen HS, Schousboe A, He W, Lin JH-C, Han X, Takano T, Wang S, Sim FJ et al (2007) The transcriptome and metabolic gene signature of protoplasmic astrocytes in the adult murine cortex. J Neurosci 27:12255–12266
Malatesta P, Hartfuss E, Götz M (2000) Isolation of radial glial cells by fluorescent-activated cell sorting reveals a neuronal lineage. Development 127:5253–5263
Pandey A, Mann M (2000) Proteomics to study genes and genomes. Nature 405:837–846
Pielot R, Smalla K-H, Müller A, Landgraf P, Lehmann A-C, Eisenschmidt E, Haus U-U, Weismantel R, Gundelfinger ED, Dieterich DC (2012) SynProt: a database for proteins of detergent-resistant synaptic protein preparations. Front Synaptic Neurosci 4:1
Qureshi IA, Mehler MF (2012) Emerging roles of non-coding RNAs in brain evolution, development, plasticity and disease. Nat Rev Neurosci 13:528–541
Rossner MJ, Hirrlinger J, Wichert SP, Boehm C, Newrzella D, Hiemisch H, Eisenhardt G, Stuenkel C, von Ahsen O, Nave K-A (2006) Global transcriptome analysis of genetically identified neurons in the adult cortex. J Neurosci 26:9956–9966
Schnell C, Shahmoradi A, Wichert SP, Mayerl S, Hagos Y, Heuer H, Rossner MJ, Hülsmann S (2015). The multispecific thyroid hormone transporter OATP1C1 mediates cell-specific sulforhodamine 101-labeling of hippocampal astrocytes. Brain Struct Funct 220:193–203. doi: 10.1007/s00429-013-0645-0
Skorupa A, Urbach S, Vigy O, King MA, Chaumont-Dubel S, Prehn JHM, Marin P (2013) Angiogenin induces modifications in the astrocyte secretome: relevance to amyotrophic lateral sclerosis. J Proteomics 91:274–285
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10:57–63
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Dieterich, D., Rossner, M. Dissecting the regional diversity of glial cells by applying -omic technologies. e-Neuroforum 6, 63–68 (2015). https://doi.org/10.1007/s13295-015-0009-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13295-015-0009-8