Abstract
Background
Non-small cell lung cancer (NSCLC) is a major contributor of cancer-related mortality. Long non-coding RNAs (lncRNAs) are indicated to participate in the pathogenesis of NSCLC.
Objective
In this research, the effects of lncRNA HOXA transcript antisense RNA, myeloid-specific 1 (HOTAIRM1) on NSCLC progression and underlying mechanism were revealed.
Methods
The expression levels of HOTAIRM1 and microRNA-498 (miR-498) were detected by quantitative real time polymerase chain reaction (qRT-PCR) in NSCLC tissues, cells or exosomes. The protein expression of CD63, CD81, hexokinase 2 (HK2) and ATP binding cassette subfamily E member 1 (ABCE1) was determined by western blot. Cell viability, apoptosis, migration and invasion were investigated by cell counting kit-8 (CCK-8), flow cytometry, transwell migration and invasion assays, respectively. Cell glycolysis metabolism was revealed by glucose uptake and lactate production assays and western blot analysis. The binding relationship between miR-498 and HOTAIRM1 or ABCE1 was predicted by DIANA-LncBase v2 and starBase online database, and identified by dual-luciferase reporter assay. The effects of HOTAIRM1 on NSCLC growth in vivo were revealed by in vivo tumor formation assay.
Results
HOTAIRM1 expression was dramatically upregulated, whereas miR-498 expression was significantly downregulated in NSCLC tissues cells or exosomes as compared to control groups. Mechanistically, HOTAIRM1 knockdown repressed cell viability, migration, invasion and glycolysis metabolism, whereas induced cell apoptosis in NSCLC; however, miR-498 inhibitor hindered these effects. Functionally, HOTAIRM1 functioned as a sponge of miR-498 and miR-498 targeted ABCE1. In addition, HOTAIRM1 silencing inhibited NSCLC growth in vivo by downregulating ABCE1 and upregulating miR-498 expression.
Conclusions
HOTAIRM1 knockdown repressed cell glycolysis metabolism and tumor development by reducing ABCE1 expression through sponging miR-498 in NSCLC, which provided a theoretical basis for further studying NSCLC progression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancers have high morbidities and mortalities, accounting for more than 20 % of cancer-caused deaths (Sun et al. 2020). About 80 % of lung cancer patients are diagnosed with non-small-cell lung cancers (NSCLC) (Zhang et al. 2015). Clinically, patients are usually diagnosed with NSCLC at the advanced stage, and as a result, optimal treatment time was lost (Jun et al. 2020). At present, there are no efficacious treatments for NSCLC. Therefore, deeply understanding NSCLC genesis is necessary to seek effective biomarkers or develop therapeutic methods for NSCLC.
Long non-coding RNAs (lncRNAs) are a class of noncoding transcripts with more than 200 nucleotides in size (Wang et al. 2020a, b, c). Previous researches presented that lncRNAs were involved in cancer progression (Dhamija and Diederichs 2016; Wei and Wang 2017). For example, lncRNA DCST1-AS1 was shown to promote cell proliferation in glioblastoma (Hu et al. 2020). LINC00665 repressed cell proliferation and metastasis in NSCLC (Wang et al. 2020a, b, c). Yang et al.. indicated that lncRNA LINC01123 hindered cell apoptosis in endometrial cancer (Yang et al. 2020). HOTAIRM1, a lncRNA, was also reported to regulate cancer development (Li et al. 2019; Chao et al. 2020). However, there were few studies on the NSCLC progression regulated by HOTAIRM1.
MicroRNAs (miRNAs) are about 20 nucleotides in length with working by regulating the expression of their target genes (Treiber et al. 2019). In cancer development, a variety of miRNAs were implicated. For example, Zhou et al.. reported that miR-214-3p suppressed cell proliferation by targeting polymorphic adenoma-like protein 2 (PLAGL2) in colorectal cancer (Zhou et al. 2020). Fu et al.. exhibited that miR-126 inhibitor promoted cell metastasis in breast cancer (Fu and Tong 2020). MiR-498 was also demonstrated to modulate cell proliferation and apoptosis by binding to X-box binding protein 1 (XBP1) in esophageal squamous cell carcinoma (ESCC) (Jin et al. 2020). Furthermore, miR-498, a miRNA, has been found to take part in NSCLC development (Yan et al. 2019; Ji et al. 2020). ATP-binding cassette (ABC) sub-family E member 1 (ABCE1) belongs to ABC family, and mediates multiply biological behaviors owing to its repressive role in RNase L (Liang et al. 2019). Additionally, Li et al.. revealed that ABCE1 knockdown repressed cell proliferation and metastasis in esophageal cancer (Li et al. 2015). In the study of Tian et al.. ABCE1 was found to promote lung cancer cell clonogenicity and anchorage-independent growth (Tian et al. 2016). These evidences suggested that miR-498 and ABCE1 were related to lung cancer processes. Our previous data showed that miR-498 contained the binding sites between HOTAIRM1 and ABCE1.
Thus, the expression levels of HOTAIRM1, miR-498 and ABCE1 were detected in NSCLC tissues, cells or exosomes. The functional effects of HOTAIRM1, miR-498 and ABCE1 on NSCLC development were investigated. In addition, whether HOTAIRM1 regulated NSCLC development by miR-498/ABCE1 pathway was identified.
Materials and methods
Sample and cell culture
The Ethics Committee of The First People’s Hospital of Jingmen consented this research. NSCLC sufferers signed the written informed consent. NSCLC tissues and paracancerous normal lung tissues were collected from NSCLC patients in The First People’s Hospital of Jingmen. These tissues were kept in liquid nitrogen.
Otwobiotech (Shenzhen, China) provided NSCLC cell lines (NCI-H1299 and A549) and human normal lung epithelial cell BEAS-2B. Cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM, Thermo Fisher, Waltham, MA, USA) supplemented with 10 % fetal bovine serum (FBS) and 1 % streptomycin/penicillin (Thermo Fisher) at 37 °C in an incubator with 5 % CO2.
Cell transfection
Small interfering RNA targeting HOTAIRM1 (si-HOTAIRM1#1 and si-HOTAIRM1#2), miR-498 mimic, miR-498 inhibitor, the overexpression plasmid of ABCE1 (pc-ABCE1), short hairpin RNA against HOTAIRM1 (sh-HOTAIRM1) and their controls (si-NC, miRNA NC, inhibitor NC, pc-NC and sh-NC) were provided by GenePharma (Shanghai, China). Transfection was performed with Lipofectamine 2000 (Thermo Fisher) according to the manufacturer’s instructions. The sequences used in this part were listed as following. Si-HOTAIRM1#1 5′-GCAAAGGCCGATTTGGAGT-3′; si-HOTAIRM1#2 5′-CCGTTCAATGAAAGATGAA-3′; miR-498 inhibitor 5′-GAAAAACGCCCCCUGGCUUGAAA-3′; miR-498 mimic 5′-UUUCAAGCCAGGGGGCGUUUUUC-3′; si-NC 5′-GCACCGGTTAGGGTAAAGT-3′; inhibitor NC 5′-CAGUACUUUUGUGUAGUACAAA-3′ and miRNA NC 5′-UUUGUACUACACAAAAGUACUG-3′.
Exosome isolation
Exosome extraction kit (Ribobio, Guangzhou, China) was utilized to isolate exosomes. In brief, medium was collected and centrifuged at 2000 rpm for 30 min. Supernatant was incubated with exosome solution overnight. And exosomes were collected by centrifuging supernatant at 1500 rpm for 30 min. The obtained exosomes were identified by transmission electron microscopy (TEM; jeol, Tokyo, Japan), nanoparticle tracking analysis and by detecting protein markers CD63 and CD81.
Western blot analysis
NSCLC tissues, cells and exosomes were lysed using RIPA buffer (Beyotime, Jiangsu, China). The lysate was loaded by 12 % sodium dodecyl sulfonate-polyacrylamide gel electrophoresis (SDS-PAGE, Abcam, Cambridge, UK). Then protein bands were transferred onto polyvinylidene fluoride membranes (Membrane Solutions, Shanghai, China). After that, bands were blocked with 5 % skim milk. Primary antibodies, anti-CD63 (1:1000, Abcam), anti-CD81 (1:700, Abcam), anti-hexokinase 2 (anti-HK2) (1:1000; Abcam), anti-ABCE1 (1:250; Abcam) and anti-glyceraldehyde 3-phosphate dehydrogenase (anti-GAPDH; 1:10,000, Abcam), were utilized to incubate the membranes. Secondary antibody labeled with horse radish peroxidase (1:1000; Abcam) was then incubated with the membranes. Results were visualized by an ECL system (Pierce, Rockford, IL, USA). Protein expression was quantified by Image J software (NIH, Bethesda, MD, USA). GAPDH was chosen as a reference.
Quantitative real-time polymerase chain reaction (qRT-PCR)
NSCLC exosomes, tissues and cells were lysed and RNA was extracted using RNA isolation kit (Takara, Shiga, Japan). cDNA was amplified using prime Script™RT reagent kit (Takara). For detecting expression of HOTAIRM1, miR-498 and ABCE1, SYBR® Premix DimerEraser kit (Takara) was employed. Finally, results were analyzed with 2−ΔΔCt method. U6 and GAPDH were chosen as controls. The sequences of sense and anti-sense primers were HOTAIRM1 5′-AAAGGCCGATTTGGAGTGCT-3′ and 5′-GGGTTCAGGCAAAACAGACC-3′; miR-498 5′-CTCACGGAAAAACGCCCCCT-3′ and 5′-ACCTCAAGAACAGTATTTCCAGG-3′; ABCE1 5′-ACAGCGAGTAGCTTTAGCCC-3′ and 5′-GACGCGATCCGCTAGATAGG-3′; U6 5′-CTCGCTTCGGCAGCACA-3′ and 5′-CTCGCTTCGGCAGCACA-3′; GAPDH 5′-TTTGACGCTGGTGCTGGTAT-3′ and 5′-TGGGAGAATGGTCGCGTATC-3′.
Cell counting kit-8 (CCK-8) assay
NCI-H1299 and A549 cells were cultured in 96-well plate for 16 h. Cells were transfected with si-HOTAIRM1, si-NC, miR-498 inhibitor, inhibitor NC, miR-498 mimic, miRNA NC, pc-ABCE1 or pc-NC. After 24 h, 10 µL CCK-8 solution was added into plate. Following that, cells were cultured for another 4 h. The results were analyzed by detecting absorbance of wavelength 450 nm with Varioskan LUX Multimode microplate reader (Thermo Fisher).
Flow cytometry analysis
The apoptotic rate of NCI-H1299 and A549 cells was determined by an Annexin V-FITC/PI kit (Solarbio, Beijing, China). In short, NCI-H1299 and A549 cells at logarithmic growth phase were collected. Then, cells were rinsed with phosphate buffer solution (PBS, Thermo Fisher), followed by the suspension with binding buffer. Cells were incubated with Annexin V-FITC and propidium iodide, respectively. Finally, sample was determined by a FACSort flow cytometry (BD Biosciences, San Diego, CA, USA).
Transwell migration and invasion assays
The migratory and invasive abilities of NCI-H1299 and A549 cells were detected using transwell chamber without or with Matrigel (Corning, New York, Madison, USA). In brief, cells were cultivated in the upper chamber with FBS-free DMEM medium. DMEM medium with 10 % FBS was placed into the lower chamber and cells were cultured for 24 h. Supernatant was removed and cells were incubated with methanol and crystal violet (Solarbio). Results were analyzed by a microscope (100×) (Agilent, Chengdu, China).
Glucose uptake and lactate production assays
Glucose uptake kit (Sigma, St Louis, MO, USA) and lactate assay kit (Abcam) were chosen to detect the glucose uptake and lactate production. In short, NCI-H1299 and A549 cells were cultured in 6-well plate for 48 h after transfection of plasmids or oligonucleotides. Supernatant was collected by centrifuging and washed using PBS. Results were analyzed by a Varioskan LUX Multimode microplate reader (Thermo Fisher).
Dual‐luciferase reporter assay
The wild-type (WT) sequence of HOTAIRM1 and mutant (MUT) HOTAIRM1 sequence without the binding sites of miR-498 were inserted into psiCHECK2 vector (Promega, Madison, WI, USA), named as WT-HOTAIRM1 and MUT-HOTAIRM1. The 3’-untranslated regions (3’UTR) of ABCE1 containing the targeting sequence of miR-498 and MUT ABCE1 were cloned into psiCHECK2 vector, named as WT-ABCE1-3’UTR and MUT-ABCE1-3’UTR. The plasmids were co-transfected into NCI-H1299 and A549 cells with miR-498 or miRNA NC, respectively. Luciferase activities were detected by dual-luciferase reporter system (Promega) at 48 h after transfection. Ranilla luciferase activity was used as a control of firefly luciferase activity.
In vivo tumor formation assay
Charles River (Beijing, China) supplied BALB/c nude mice (5-week old). All nude mice were fed in pathogen-free environment. Nude mice were averagely divided into two groups (sh-NC group and sh-HOTAIRM1 group; N = 8 per group). 5 × 106 cells transfected with sh-HOTAIRM1 or sh-NC were injected into the flank of mice. Tumors volume was measured every 7 days. Nude mice were euthanized at the 28th after injection. Tumors weight was determined. The effects of HOTAIRM1 knockdown on the expression levels of miR-498 and ABCE1 in vivo were revealed by qRT-PCR. This study was permitted by the Animal Care and Use Committee of The First People’s Hospital of Jingmen.
Statistical analysis
Every experiment was repeated three times. Figures were made using GraphPad Prism 5.0 and image J software. The relationship between HOTAIRM1 and miR-498 was determined by Pearson correlation analysis. Data were shown as means ± standard deviations (SD). Significant differences were compared with two-tailed Student’s t-tests or one-way analysis of variance. P value < 0.05 was considered statistically significant.
Results
HOTAIRM1 expression is upregulated and miR-498 is downregulated in NSCLC tissues and exosomes secreted from NCI-H1299 and A549 cells
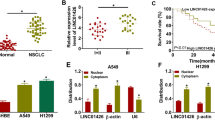
HOTAIRM1 and miR-498 expression were firstly detected in NCI-H1299 and A549 cell exosomes. TEM showed that the vesicles had a homogeneous structure, which was similar with the morphological characteristics of exosomes (Fig. 1a). Nanoparticle tracking analysis showed that the size of vesicles ranged from 30 to 150 nm (Fig. 1b). Meanwhile, western blot analysis detected the expression of exosome protein markers CD63 and CD81 in exosomes from NSCLC cells and normal cells (Fig. 1c). In addition, qRT-PCR results showed that HOTAIRM1 expression was obviously upregulated in NSCLC tissues and exosomes secreted from NSCLC cells as compared to control groups (Fig. 1d and g). MiR-498 was found to be downregulated in NSCLC tissues and exosomes secreted from NSCLC cells compared with control groups (Fig. 1e and h). Pearson correlation analysis explained that HOTAIRM1 was negatively correlated with miR-498 in expression (Fig. 1f). These data showed that HOTAIRM1 was upregulated, while miR-498 was downregulated in NSCLC tissues and exosomes from NCI-H1299 and A549 cells, and HOTAIRM1 expression was negatively related to miR-498 expression.
HOTAIRM1 is overexpressed and miR-498 is lowly expressed in NSCLC tissues and exosomes secreted from NCI-H1299 and A549 cells. a TEM was employed to visualize the size and morphology of the vesicles. b Nanoparticle tracking analysis was performed to determine the size and concentration of vesicles. c The protein expression of exosome markers (CD63 and CD81) were detected by western blot. d, e The expression levels of HOTAIRM1 and miR-498 were detected by qRT-PCR in NSCLC tissues and normal tissues. f Pearson correlation analysis was carried out to analyze the relationship between HOTAIRM1 and miR-498. g HOTAIRM1 expression was detected in exosomes secreted from BEAS-2B, NCI-H1299 and A549 cells by qRT-PCR. h QRT-PCR was employed to determine miR-498 expression in exosomes secreted from BEAS-2B, NCI-H1299 and A549 cells. *P < 0.05
HOTAIRM1 knockdown inhibits the progression of NSCLC
The functional effects of HOTAIRM1 on NSCLC progression were continued to be explored. Firstly, the expression level of HOTAIRM1 in NCI-H1299, A549 and BEAS-2B cells was detected. QRT-PCR results showed that HOTAIRM1 expression was dramatically upregulated in NCI-H1299 and A549 cells relative to BEAS-2B cells (Fig. 2a). QRT-PCR analysis showed that HOTAIRM1 expression was dramatically downregulated by si-HOTAIRM1#1 and si-HOTAIRM1#2 compared with control group (Fig. 2b). si-HOTAIRM1#1 was chosen for further study because of its higher interfering efficiency, and si-HOTAIRM1 substituted for si-HOTAIRM1#1 in subsequent trials. Subsequently, CCK-8 assay showed that HOTAIRM1 knockdown inhibited the viability of NCI-H1299 and A549 cells (Fig. 2c). Flow cytometry analysis presented that HOTAIRM1 silencing upregulated the apoptosis rate of NCI-H1299 and A549 cells (Fig. 2d). Transwell migration and invasion assays revealed that the migratory and invasive abilities of NCI-H1299 and A549 cells were repressed by HOTAIRM1 depletion (Fig. 2e and f). Warburg effect is a metabolic pathway that glucose metabolism is transformed from oxidative phosphorylation to glycolysis, consuming glucose and generating lactic acid (Zhao et al. 2018). Warburg effect indicates that cancer cells exhibit an increased dependence on glucolysis for energy needs. Thus, we further studied whether HOTAIRM1 knockdown affected Warburg effect. Glucose uptake and lactate production assays showed that HOTAIRM1 knockdown repressed the glucose uptake and lactate production (Fig. 2g and h). Western blot results explained that the protein level of HK2 was dramatically repressed by HOTAIRM1 depletion (Fig. 2i). Overall, all data suggested that HOTAIRM1 knockdown inhibited NSCLC progression.
HOTAIRM1 knockdown represses NSCLC progression in NSCLC. a The expression level of HOTAIRM1 was determined by qRT-PCR in NCI-H1299, A549 and BEAS-2B cells. b QRT-PCR was employed to determine the interfering efficiency of si-HOTAIRM1#1 and si-HOTAIRM1#2 in NCI-H1299 and A549 cells. c The effect of HOTAIRM1 knockdown on the viability of NCI-H1299 and A549 cells was revealed by CCK-8 assay. d Flow cytometry assay was employed to analyze the effect of HOTAIRM1 silencing on the apoptosis rate of NCI-H1299 and A549 cells. e, f The impacts of HOTAIRM1 depletion on migratory and invasive abilities of NCI-H1299 and A549 cells were detected by transwell migration and invasion assays. g, h Glucose uptake and lactate production assays were performed to investigate the influences of HOTAIRM1 knockdown on glucose uptake and lactate production. i Western blot was carried out to examine the impact of HOTAIRM1 depletion on protein expression of HK2. *P < 0.05
HOTAIRM1 functions as a sponge of miR-498
The miRNA associated with HOTAIRM1 was further sought in this part. The DIANA-LncBase v2 online database showed that HOTAIRM1 contained the binding sites of miR-498 (Fig. 3a). Dual-luciferase reporter assay revealed that the luciferase activity was dramatically repressed after WT-HOTAIRM1 and miR-498 mimic co-transfection in NCI-H1299 and A549 cells, whereas there was no obvious change in MUT-HOTAIRM1 + miR-498 mimic group (Fig. 3b and c). Subsequently, the effects between HOTAIRM1 knockdown and miR-498 inhibitor on miR-498 expression were revealed. The interfering efficiency of miR-498 inhibitor was firstly detected by qRT-PCR. Results showed that miR-498 expression was obviously repressed by miR-498 inhibitor (Fig. 3d). HOTAIRM1 knockdown evidently upregulated miR-498 expression in NCI-H1299 and A549 cells, whereas this effect was restrained by miR-498 inhibitor (Fig. 3e and f). These data illustrated that HOTAIRM1 was associated with miR-498.
HOTAIRM1 serves as a sponge of miR-498. a DIANA-LncBase v2 online database was performed to predict the binding sequence between HOTAIRM1 and miR-498. b, c Dual-luciferase reporter assay was employed to detect luciferase activities in NCI-H1299 and A549 cells. d The interfering efficiency of miR-498 inhibitor was determined by qRT-PCR. e, f The effects between HOTAIRM1 knockdown and miR-498 inhibitor on miR-498 expression were revealed by qRT-PCR. *P < 0.05
Inhibition of miR-498 compromises HOTAIRM1 knockdown-mediated effects in NSCLC
The effects between HOTAIRM1 and miR-498 on NSCLC progression were revealed by loss-of-function experiments. CCK-8 assay showed that HOTAIRM1 knockdown inhibited cell viability, but miR-498 inhibitor decreased this inhibition effect in NCI-H1299 and A549 cells (Fig. 4a and b). Flow cytometry analysis demonstrated that HOTAIRM1 silencing induced cell apoptosis in NCI-H1299 and A549 cells, whereas miR-498 inhibitor partially reversed this effect (Fig. 4c and d). Transwell migration assay revealed that the migratory ability of NCI-H1299 and A549 cells was suppressed by HOTAIRM1 depletion, which was partially attenuated by miR-498 inhibitor (Fig. 4e and f). Transwell invasion assay revealed that the invasive ability of NCI-H1299 and A549 cells was inhibited by HOTAIRM1 depletion, which was partially abolished by miR-498 inhibitor (Fig. 4g and h). In addition, glucose uptake and lactate production assays showed that HOTAIRM1 knockdown inhibited glucose consumption and lactate production; however, these effects were partially restored by miR-498 inhibitor (Fig. 4i and j). Western blot analysis demonstrated that the expression level of HK2 protein was inhibited by HOTAIRM1 silencing, whereas miR-498 inhibitor hindered this inhibition impact (Fig. 4k). All the above data revealed that HOTAIRM1 silencing repressed NSCLC development by sponging miR-498.
HOTAIRM1 knockdown inhibits NSCLC progression by sponging miR-498. a, b The effects between HOTAIRM1 knockdown and miR-498 inhibitor on the viability of NCI-H1299 and A549 cells were detected by CCK-8 assay. c, d Flow cytometry was performed to determine the impacts between HOTAIRM1 depletion and miR-498 inhibitor on the apoptosis of NCI-H1299 and A549 cells. e, f Transwell migration assay was carried out to explain the effects between HOTAIRM1 silencing and miR-498 inhibitor on cell migration in NCI-H1299 and A549 cells. g, h Transwell invasion assay was conducted to determine the impacts between HOTAIRM1 knockdown and miR-498 inhibitor on cell invasion in NCI-H1299 and A549 cells. i, j Glucose uptake and lactate production assays were carried out to reveal the effects between HOTAIRM1 depletion and miR-498 inhibitor on cell glucose consumption and lactate production. k The effects between HOTAIRM1 silencing and miR-498 inhibitor on HK2 expression were explored by western blot. *P < 0.05
MiR-498 targets ABCE1 in NCI-H1299 and A549 cells
The target gene of miR-498 was predicted by starBase online database. Result showed that ABCE1 contained the target sequence of miR-498 (Fig. 5a). Dual-luciferase reporter assay showed that the luciferase activity was dramatically repressed in WT-ABCE1-3’UTR + miR-498 mimic group in NCI-H1299 and A549 cells; however, there was no obvious change after MUT-ABCE1-3’UTR and miR-498 transfection (Fig. 5b and c). Subsequently, the impacts between miR-498 mimic and ABCE1 overexpression on ABCE1 expression were determined. The overexpression efficiency of miR-498 mimic and pc-ABCE1 was firstly detected. QRT-PCR results showed that the expression of miR-498 was greatly upregulated after miR-498 mimic transfection (Fig. 5d). Western blot analysis disclosed that ABCE1 expression level was significantly promoted by pc-ABCE1 (Fig. 5e). Then, western blot results displayed that ABCE1 expression was inhibited by miR-498 mimic in NCI-H1299 and A549 cells, whereas ABCE1 overexpression hindered this effect (Fig. 5f and g). These results revealed that miR-498 targeted ABCE1 in NCI-H1299 and A549 cells.
MiR-498 targets ABCE1 in NCI-H1299 and A549 cells. a The binding sequence between miR-498 and ABCE1 was predicted by starBase online database. b, c The luciferase activities were detected by dual-luciferase reporter assay in NCI-H1299 and A549 cells. d QRT-PCR was performed to investigate the overexpression efficiency of miR-498 mimic in NCI-H1299 and A549 cells. e The effect of pc-ABCE1 on ABCE1 protein expression was determined by western blot in NCI-H1299 and A549 cells. f, g Western blot assay was carried out to explain the impacts between miR-498 mimic and ABCE1 overexpression on ABCE1 expression in NCI-H1299 and A549 cells. *P < 0.05
MiR-498 inhibits cell glycolysis metabolism and NSCLC progression by targeting ABCE1
The effects between miR-498 mimic and ABCE1 overexpression on NSCLC development were further studied. CCK-8 assay displayed that miR-498 mimic repressed the viability of NCI-H1299 and A549 cells, whereas this effect was attenuated by ABCE1 overexpression ( Fig. 6a and b). Flow cytometry analysis revealed that miR-498 mimic promoted cell apoptosis, while this phenomenon was restrained by ABCE1 overexpression in NCI-H1299 and A549 cells (Fig. 6c and d). Transwell migration assay explained that cell migration was repressed by miR-498 mimic, whereas this inhibition effect was hindered by ABCE1 overexpression (Fig. 6e and f). Transwell invasion assay demonstrated that miR-498 mimic inhibited cell invasion; however, ABCE1 overexpression partially attenuated this effect (Fig. 6g and h). In addition, glucose uptake and lactate production assays revealed that miR-498 mimic inhibited glucose uptake and lactate production in NCI-H1299 and A549 cells, which was partially restored by ABCE1 overexpression (Fig. 6i and j). Meanwhile, western blot results showed that miR-498 mimic repressed HK2 protein expression, but ABCE1 overexpression retained this effect (Fig. 6k). Collectively, all evidences showed that miR-498 mimic repressed cell glycolysis metabolism, proliferation, migration and invasion, whereas induced cell apoptosis by binding to ABCE1 in NSCLC.
MiR-498 suppresses cell glycolysis metabolism and NSCLC progression by binding to ABCE1. a, b The effects between miR-498 mimic and ABCE1 overexpression on cell viability were detected by CCK-8 assay in NCI-H1299 and A549 cells. c, d The effects between miR-498 mimic and ABCE1 overexpression on cell apoptosis were determined by flow cytometry analysis in NCI-H1299 and A549 cells. e, f Transwell migration assay was performed to explain the effects between miR-498 mimic and ABCE1 overexpression on cell migration. g, h Transwell invasion assay was employed to explain the effects between miR-498 mimic and ABCE1 overexpression on cell invasion. i, j Glucose uptake and lactate production assays were carried out to reveal the effects between miR-498 mimic and ABCE1 overexpression on glucose uptake and lactate production in NCI-H1299 and A549 cells. k The effects between miR-498 mimic and ABCE1 overexpression on the protein expression of HK2 were investigated by western blot in NCI-H1299 and A549 cells. *P < 0.05
HOTAIRM1 knockdown represses the protein expression of ABCE1 by regulating miR-498
Given the binding relationship between miR-498 and HOTAIRM1 or ABCE1, whether HOTAIRM1 regulated ABCE1 by interacting with miR-498 was revealed in this part by western blot. Results showed that HOTAIRM1 knockdown repressed the protein expression of ABCE1 in NCI-H1299 and A549 cell, whereas this effect was hindered by ABCE1 overexpression (Fig. 7a and b). In addition, western blot also demonstrated that the protein expression of ABCE1 was inhibited by HOTAIRM1 silencing, whereas miR-498 inhibitor decreased this impact in NCI-H1299 and A549 cells (Fig. 7c and d). These results suggested that HOTAIRM1 inhibited ABCE1 expression by binding to miR-498.
HOTAIRM1 silencing represses ABCE1 expression by regulating miR-498. a, b The effects between HOTAIRM1 knockdown and ABCE1 overexpression on ABCE1 protein expression were investigated by western blot in NCI-H1299 and A549 cells. c, d The impacts between HOTAIRM1 silencing and miR-498 inhibitor on ABCE1 protein expression were revealed by western blot in NCI-H1299 and A549 cells. *P < 0.05
HOTAIRM1 knockdown represses NSCLC growth in vivo by regulating miR-498 and ABCE1 expression
The effects of HOTAIRM1 knockdown on NSCLC growth in vivo were studied by in vivo tumor formation assay. The effects of HOTAIRM1 silencing on tumor volume and weight were firstly revealed. Results showed that HOTAIRM1 depletion repressed tumor volume and weight (Fig. 8a and b). Subsequently, the impacts of HOTAIRM1 silencing on the expression levels of miR-498 and ABCE1 were investigated. The knockdown efficiency of sh-HOTAIRM1 was firstly detected, and qRT-PCR results showed that the expression of HOTAIRM1 was dramatically downregulated by sh-HOTAIRM1 when compared with control group (Fig. 8c). Then, qRT-PCR and western blot analysis showed that miR-498 expression was significantly upregulated and ABCE1 protein expression was downregulated after sh-HOTAIRM1 transfection as compared to control groups, respectively (Fig. 8d and e). These results showed that HOTAIRM1 depletion repressed NSCLC growth in vivo by upregulating miR-498 and downregulating ABCE1 expression.
HOTAIRM1 silencing inhibits NSCLC growth in vivo. a The effect of HOTAIRM1 knockdown on tumor volume was revealed. b The effect of HOTAIRM1 silencing on tumor weight was determined. c QRT-PCR was employed to reveal the knockdown efficiency of sh-HOTAIRM1. d The impact of HOTAIRM1 depletion on miR-498 expression was detected by qRT-PCR. e Western blot was employed to investigate the impact of HOTAIRM1 knockdown on the protein expression of ABCE1
Discussion
NSCLC is a common prevalent tumor and poses a heavy burden to human health. The 5-years survival rate of NSCLC patients is low because of the chemotherapeutic resistance and high metastasis of NSCLC (Ruysscher et al. 2018). Therefore, studying more about NSCLC tumorgenesis is necessary to seek reliable biomarkers for its treatment.
HOTAIRM1 was shown to regulate cancer progression. For instance, in glioma, HOTAIRM1 was exhibited to accelerate cell proliferation and suppress cell apoptosis (Xia et al. 2020). Hao et al. explained that HOTAIRM1 was overexpressed and its depletion repressed cell invasion in glioblastoma (Hao et al. 2020). Li et al.. also suggested that HOTAIRM1 promoted cell metastasis in endometrial cancer (Li et al. 2019). In addition, Chen et al.. reported that HOTAIRM1 repressed cell glycolysis metabolism by regulating glucose consumption and lactate production (Chen et al. 2020). Consistently, in our study, HOTAIRM1 was upregulated in NSCLC tissues and cells. And HOTAIRM1 promoted cell proliferation, metastasis and glycolysis metabolism, whereas repressed cell apoptosis in NSCLC. Additionally, HOTAIRM1 was shown to be upregulated in exosomes derived from NSCLC cells and targeted miR-498.
MiR-498 expression was reported to be downregulated and inhibit cell proliferation and invasion in ESCC cells (Wang et al. 2020a, b, c). You et al.. revealed that miR-498 suppressed cell metastasis in gastric cancer (You et al. 2020). MiR-498 was also explained to induce cell apoptosis in acute myeloid leukemia (Moghaddam et al. 2018). Similarly, we found that miR-498 inhibitor promoted cell proliferation, metastasis and hindered cell apoptosis in NSCLC. Furthermore, miR-498 was shown to be downregulated in exosomes from NSCLC cells. And miR-498 depletion contributed to cell glycolysis metabolism.
In our experiments, we also found miR-498 targeted ABCE1. ABCE1 was revealed to regulate cancer progression. Tian et al. . unclosed that ABCE1 overexpression promoted cell proliferation and metastasis in lung cancer (Tian et al. 2016). Wei et al. unveiled that miR-145 inhibitor restrained cell apoptosis by targeting ABCE1 (Wei et al. 2017), suggesting ABCE1 constrained cell apoptosis. Similarly, we explained that ABCE1 promoted cell proliferation and metastasis, whereas restrained cell apoptosis in NSCLC. In addition, ABCE1 was also discovered to accelerate cell glycolysis metabolism.
Conclusions
Collectively, HOTAIRM1 expression was upregulated and miR-498 was downregulated in NSCLC tissues, cells or exosomes compared with control groups. HOTAIRM1 knockdown repressed cell proliferation, metastasis and glycolysis metabolism, whereas accelerated cell apoptosis by downregulating ABCE1 expression through sponging miR-498 in NSCLC. Furthermore, HOTAIRM1 silencing repressed tumors growth in vivo. These data not only provide a theoretical basis for further studying NSCLC genesis and but also demonstrate that HOTAIRM1 may be employed as a diagnosis biomarker for NSCLC.
Abbreviations
- NSCLC:
-
Non-small cell lung cancer
- lncRNAs:
-
Long non-coding RNAs
- HOTAIRM1:
-
lncRNA HOXA transcript antisense RNA, myeloid-specific 1
- ABCE1:
-
ATP binding cassette subfamily E member 1
References
Chao H, Zhang M, Hou H, Zhang Z, Li N (2020) HOTAIRM1 suppresses cell proliferation and invasion in ovarian cancer through facilitating ARHGAP24 expression by sponging miR-106a-5p. Life Sci 243:117296
Chen L, Hu N, Wang C, Zhao H (2020) HOTAIRM1 knockdown enhances cytarabine-induced cytotoxicity by suppression of glycolysis through the Wnt/beta-catenin/PFKP pathway in acute myeloid leukemia cells. Arch Biochem Biophys 680:108244
De Ruysscher D, Wanders R, Hendriks LE, van Baardwijk A, Reymen B, Houben R, Bootsma G, Pitz C, van Eijsden L, Dingemans AC (2018) Progression-free survival and overall survival beyond 5 years of nsclc patients with synchronous oligometastases treated in a prospective phase II Trial (NCT 01282450). J Thorac Oncol 13(12):1958–1961
Dhamija S, Diederichs S (2016) From junk to master regulators of invasion: lncRNA functions in migration, EMT and metastasis. Int J Cancer 139(2):269–280
Fu R, Tong JS (2020) miR-126 reduces trastuzumab resistance by targeting PIK3R2 and regulating AKT/mTOR pathway in breast cancer cells. J Cell Mol Med 24(13):7600–7608
Hao Y, Li X, Chen H, Huo H, Liu Z, Chai E (2020) Over-expression of long noncoding RNA HOTAIRM1 promotes cell proliferation and invasion in human glioblastoma by up-regulating SP1 via sponging miR-137. Neuroreport 31(2):109–117
Hu S, Yao Y, Hu X, Zhu Y (2020) LncRNA DCST1-AS1 downregulates miR-29b through methylation in glioblastoma (GBM) to promote cancer cell proliferation. Clin Transl Oncol 22(12):2230–2235
Ji X, Tao R, Sun LY, Xu XL, Ling W (2020) Down-regulation of long non-coding RNA DUXAP8 suppresses proliferation, metastasis and EMT by modulating miR-498 through TRIM44-mediated AKT/mTOR pathway in non-small-cell lung cancer. Eur Rev Med Pharmacol Sci 24(6):3152–3165
Jin G, Yang Y, Tuo G, Wang W, Zhu Z (2020) LncRNA TUG1 promotes tumor growth and metastasis of esophageal squamous cell carcinoma by regulating XBP1 via competitively binding to miR-498. Neoplasma 67(4):751–761
Jun N, Hanping W, Xiaoyan S, Yan X, Mengzhao W, Xiaotong Z, Li Z (2020) Treatment with or without bevacizumab as a first-line and maintenance therapy for advanced non-squamous non-small cell lung cancer: a retrospective study. Thorac Cancer 11(7):1869–1875
Li XR, Yang LZ, Huo XQ, Wang Y, Yang QH, Zhang QQ (2015) Effects of silencing the ATP-binding cassette protein E1 gene by electroporation on the proliferation and migration of EC109 human esophageal cancer cells. Mol Med Rep 12(1):837–842
Li X, Pang L, Yang Z, Liu J, Li W, Wang D (2019) LncRNA HOTAIRM1/HOXA1 Axis promotes cell proliferation, migration and invasion in endometrial cancer. Onco Targets Ther 12:10997–11015
Liang Z, Yu Q, Ji H, Tian D (2019) Tip60-siRNA regulates ABCE1 acetylation to suppress lung cancer growth via activation of the apoptotic signaling pathway. Exp Ther Med 17(4):3195–3202
Moghaddam Y, Andalib A, Mohammad-Ganji M, Homayouni V, Sharifi M, Ganjalikhani-Hakemi M (2018) Evaluation of the effect of TIM-3 suppression by miR-498 and its effect on apoptosis and proliferation rate of HL-60 cell line. Pathol Res Pract 214(9):1482–1488
Sun L, Zhang Z, Yao Y, Li W-Y, Gu J (2020) Analysis of expression differences of immune genes in non-small cell lung cancer based on TCGA and ImmPort data sets and the application of a prognostic model. Ann Transl Med 8(8):550
Tian Y, Tian X, Han X, Chen Y, Song CY, Jiang WJ, Tian DL (2016) ABCE1 plays an essential role in lung cancer progression and metastasis. Tumour Biol 37(6):8375–8382
Treiber T, Treiber N, Meister G (2019) Regulation of microRNA biogenesis and its crosstalk with other cellular pathways. Nat Rev Mol Cell Biol 20(1):5–20
Wang Y, Wang H, Ruan J, Zheng W, Yang Z, Pan W (2020a) Long non-coding RNA OIP5-AS1 suppresses multiple myeloma progression by sponging miR-27a-3p to activate TSC1 expression. Cancer Cell Int 20:155
Wang H, Wang L, Zhang S, Xu Z, Zhang G (2020b) Downregulation of LINC00665 confers decreased cell proliferation and invasion via the miR-138-5p/E2F3 signaling pathway in NSCLC. Biomed Pharmacother 127:110214
Wang Z, Liu J, Wang R, Wang Q, Liang R, Tang J (2020c) Long non-coding RNA taurine upregulated gene 1 (TUG1) Downregulation constrains cell proliferation and invasion through regulating cell division cycle 42 (CDC42) expression via mir-498 in esophageal squamous cell carcinoma cells. Med Sci Monit 26:e919714
Wei GH, Wang X (2017) lncRNA MEG3 inhibit proliferation and metastasis of gastric cancer via p53 signaling pathway. Eur Rev Med Pharmacol Sci 21(17):3850–3856
Wei D, Yang L, Lv B, Chen L (2017) Genistein suppresses retinoblastoma cell viability and growth and induces apoptosis by upregulating miR-145 and inhibiting its target ABCE1. Mol Vis 23:385–394
Xia H, Liu Y, Wang Z, Zhang W, Qi M, Qi B, Jiang X (2020) Long noncoding RNA HOTAIRM1 maintains tumorigenicity of glioblastoma stem-like cells through regulation of hox gene expression. Neurotherapeutics 17(2):754–764
Yan R, Jiang Y, Lai B, Lin Y, Wen J (2019) The positive feedback loop FOXO3/CASC11/miR-498 promotes the tumorigenesis of non-small cell lung cancer. Biochem Biophys Res Commun 519(3):518–524
Yang Y, Wu J, Zhou H, Liu W, Wang J, Zhang Q (2020) STAT1-induced upregulation of lncRNA LINC01123 predicts poor prognosis and promotes the progression of endometrial cancer through miR-516b/KIF4A. Cell Cycle 19(12):1502–1516
You D, Wang D, Liu P, Chu Y, Zhang X, Ding X, Li X, Mao T, Jing X, Tian Z et al (2020) MicroRNA-498 inhibits the proliferation, migration and invasion of gastric cancer through targeting BMI-1 and suppressing AKT pathway. Hum Cell 33(2):366–376
Zhang H, Zhu X, Li N, Li D, Sha Z, Zheng X, Wang H (2015) miR-125a-3p targets MTA1 to suppress NSCLC cell proliferation, migration, and invasion. Acta Biochim Biophys Sin 47(7):496–503
Zhao W, Li W, Dai W, Huang N, Qiu J (2018) LINK-A promotes cell proliferation through the regulation of aerobic glycolysis in non-small-cell lung cancer. Onco Targets Ther 11:6071–6080
Zhou Z, Wu L, Liu Z, Zhang X, Han S, Zhao N, Bao H, Yuan W, Chen J, Ji J et al (2020) MicroRNA-214-3p targets the PLAGL2-MYH9 axis to suppress tumor proliferation and metastasis in human colorectal cancer. Aging 12(10):9633–9657
Funding
There is no funding to report.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Dongping Chen, Yashan Li, Yukang Wang and Jinlian Xu The authors declare that they have no financial conflicts of interest.
Ethical approval
This study had been approved by the The First People’s Hospital of Jingmen. Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, D., Li, Y., Wang, Y. et al. LncRNA HOTAIRM1 knockdown inhibits cell glycolysis metabolism and tumor progression by miR-498/ABCE1 axis in non‐small cell lung cancer. Genes Genom 43, 183–194 (2021). https://doi.org/10.1007/s13258-021-01052-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-021-01052-9