Abstract
Background
Centromeres are specialized chromosomal domains involved in kinetochore formation and faithful chromosome segregation. Despite a high level of functional conservation, centromeres are not identified by DNA sequences, but by epigenetic means. Universally, centromeres are typically formed on highly repetitive DNA, which were previously considered to be silent. However, recent studies have shown that transcription occurs in this region, known as centromeric-derived RNAs (cenRNAs). CenRNAs that contribute to fundamental aspects of centromere function have been recently investigated in detail. However, the distribution, behavior and contributions of centromeric transcripts are still poorly understood.
Objective
The aim of this article is to provide an overview of the roles of cenRNAs in centromere formation and function.
Methods
We describe the structure and DNA sequence of centromere from yeast to human. In addition, we briefly introduce the roles of cenRNAs in centromere formation and function, kinetochore structure, accurate chromosome segregation, and pericentromeric heterochromatin assembly. Centromeric circular RNAs (circRNAs) and R-loops are rising stars in centromere function. CircRNAs have been successfully identified in various species with the assistance of high-throughput sequencing and novel computational approaches for non-polyadenylated RNA transcripts. Centromeric R-loops can be identified by the single-strand DNA ligation-based library preparation technique. But the molecular features and function of these centromeric R-loops and circRNAs are still being investigated.
Conclusion
In this review, we summarize recent findings on the epigenetic regulation of cenRNAs across species, which would provide useful information about cenRNAs and interesting hints for further studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Centromeres play an essential role in kinetochore assembly and equal chromosome segregation, and are marked by a specific histone H3 variant (CENP-A in human and fission yeast; CENH3 in plants and Cse4 in budding yeast) (Henikoff et al. 2020; Dhatchinamoorthy et al. 2018). There are three major classes of centromeres, point centromere, regional centromere and holocentromere. Point centromeres are 120–200-bp, and are found in budding yeast (Kobayashi et al. 2015). Regional centromeres are most prevalent among human, mice, fission yeast, plants and other higher eukaryotes, which may reflect the ancestral centromere organization. Holocentromeres are found in Caenorhabditis elegans, for example, and encompass the length of a chromosome (Henikoff et al. 2020; Pluta 1995). Most plant and animal centromeres favor AT-rich DNA that comprise retrotransposons and tandemly repetitive DNA known as satellites (Fig. 1). Even though their functions are evolutionary highly conserved, the underlying centromeric DNAs are highly variable in sequence and evolve quickly, which are not essential for centromere identity (Cleveland et al. 2003; Stimpson and Sullivan 2010), suggesting that epigenetic marks are involved in establishing the centromeric state, like associated RNAs, proteins and other epigenetic modifications.
An optimal level of RNA transcription is required for maintaining the proper function of the centromere. Both sense and antisense cenRNAs including circRNA transcripted from centromeric repeats are detected. A low level of cenRNAs would lead to aberrant mitosis, micronuclei and autophosphorylation of Aurora B increase. Conversely, a high level of cenRNA would also cause mislocalization of centromere-associated proteins, centromere inactivation, and centromeric epigenetic changes
Centromeres are dynamic, rather than being inert. A common feature of centromeres is that they occur in gene-free regions but include genes that are transcribed at a very low level (Su et al. 2019; Henikoff and Talbert 2018). Although centromeric transcripts are a conserved epigenetic mechanism regulating centromeres across species, they vary dramatically in size. CenRNAs are associated with a broad range of functions, including participating in the regulation of chromosome behavior, gene transcription, and chromatin architecture (Arunkumar and Melters. 2020).
Various kinds of RNAs of eukaryotes and prokaryotes are attracting a lot of attention from researchers, including small RNAs, long noncoding RNAs (lncRNAs), circular RNAs (circRNAs), and RNA: DNA hybrids. Given the growing number of studies showing the role of cenRNAs, our current knowledge is largely derived from work in animals and yeasts; their global function are still enigmatic. We refer readers to a comprehensive summary that covers the possible biological roles of cenRNAs.
-
1.
A balanced level of cenRNA is essential for maintaining the proper function of centromeres
Centromeres comprise two domains: the central core and the flanking pericentric heterochromatin, serving for assembly of the kinetochore and centromere cohesion separately (Corless et al. 2020). Centromeres are the most condensed region of a chromosome. However, a low level transcription of the core and the flanking pericentric regions is detected in human cells (Wong et al. 2007), mouse cells (Ferri et al. 2009), fission yeast (Choi et al. 2011) and plants (Lv et al. 2020), suggesting that CENP-A chromatin contains some open chromatin. A human neocentromere contains 51 genes, of which about one-third are expressed (Saffery et al. 2003). Centromeric α-satellite transcripts are estimated to be about 0.5% that of a housekeeping gene (Chan et al. 2012). Genes transcription is also detected in de novo centromeric regions of maize (Su et al. 2016). In Arabidopsis, more than 47 expressed genes were found to be flanked by core centromeric repetitive sequence such as cen180 (Arabidopsis Genome 2000; May et al. 2005). In rice centromere 8, four of the genes present have normal transcripts (Nagaki et al. 2004). It has long been known that yeast centromere function can be switched off or on by controlling transcription induction or repression from the GAL1 promoter producing of a conditional centromere (Hill et al. 1987). There is also evidence that shows direct links between transcription and centromere activation from yeast to human. In fission yeast, centromeric histone H3 with CENP-ACnp1 tends to associate with a subset of RNA polymerase II (RNA PolII) promoters where RNA PolII binding is high. Similar findings were also found in S. cerevisiae CENP-ACse4 (Choi et al. 2011; Ólafsson et al. 2020). In Schizosaccharomyces pombe, Ams2 is a cell cycle-regulated GATA-like transcription factor, depletion of which results in the reduction of CENP-A binding to centromeres and thus chromosome missegration. Conversely, with the accumulation of Ams2, association of CENP-A mutant protein with a centromere is restored (Chen et al. 2003), implicating that transcription acts in centromere function.
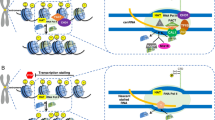
Transcription of centromeres is largely dependent on activation of RNA PolII and varies between development at stages and tissues (McNulty et al. 2017; Maison et al. 2010). Point centromere activity requires an optimal level of centromeric noncoding RNA. RNA PoII-mediated centromeric transcriptional level that is excessively high or low leads to centromere inactivation and failures in segregation (Ling et al. 2019; Ohkuni et al. 2011). CenRNA over-expression in cbf1 (centromere-binding protein 1) and htz1 (histone H2A variant) deletion increases budding yeast minichromosome loss. Minichromosome loss was also significantly increased when all the cenRNAs were knocked down (Ling et al. 2019). Nakano et al. (2008) developed a human artificial chromosome with an operable epigenetic state and also found that only moderate levels of transcription are compatible with correct centromere function. Functional centromere activity was deactivated by strong transcription from an artificial promoter, and was restored when centromeric transctipts decreased (Collins et al. 2005; Ohkuni et al. 2011). Thus, there is an optimal level of RNA transcription required for centromere and kinetochore assembly (Fig. 1).
Currently, several transcriptional regulators including RNA PolII are essential for keeping cenRNA balance, which were described in yeast, mouse and human cells. A nuclear protein ZFAT binds to centromeres to control centromeric non-coding RNA transcription through a specific 8-bp DNA sequence in human and mouse cells. (Ishikura et al. 2020). In budding yeast, centromeric transcription is suppressed to a low level by kinetochore protein Cbf1 and histone H2A variant H2A.ZHtz1 (Ling et al. 2019). In mice, MIWI regulates the post-transcription of mRNA, lncRNA and transposons. MIWI- and Dicer-mediated cleavage of the centromeric satellite RNAs prevents aneuploidy by preventing the over-expression of satellite RNAs (Hsieh et al. 2020). In human cells, alpha-satellite expression was repressed by centromere-nucleolar interactions (Bury et al. 2020). Other studies suggest cenRNAs remain at centromeres. However, Bury et al. (2020) found that alpha-satellite RNA transcripts were broadly distributed within the cytoplasm during mitosis, which provides a different perspective for cenRNA function. This may be explained by MuNulty’s (2017) opinion that each human alpha satellite array produces a unique set of non-coding transcripts to perform different functions (Fig. 2).
-
2.
CenRNAs are essential for CENP-A loading onto centromeres.
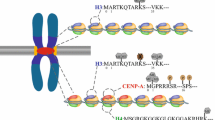
Loading of CENP-A at centromeres occurs in a cell cycle-specific manner. Synthesis of new CENP-A is deposited in metaphase in D. melanogaster S2 cells (Mellone et al. 2011), during telophase and G1 in human (Jansen et al. 2007), and prior to mitosis in G2 in plants such as maize, barley, rye and arabidopsis (Topp et al. 2004; Lermontova et al. 2007; Schubert et al. 2014; Lermontova et al. 2011). Consistently, cenRNAs levels are also cell cycle-regulated. In recent years, a direct RNA–protein interaction between centromeric RNAs and CENP-A has been found in many eukaryotes. In humans and Drosophila, centromeres are actively transcribed by RNA polymerase II from late mitosis to early G1 (Jansen et al. 2007; Dunleavy et al. 2012). A 1.3-kb RNA that originates from centromeres was associated with CENP-A. The long-term loss of centromeric transcripts led to the loss of CENP-A recruitment and its chaperone HJURP to centromeres, whereas its overexpression increases CENP-A and HJURP recruitment (Quénet et al. 2014). Yet, there is ample evidence from human and Xenopus oocytes that knock-down of centromeric transcripts results in reduced CENP-A levels at centromeres (Quénet et al. 2014; Saffery et al. 2003; Bergmann et al. 2011; Grenfell et al. 2016). McNulty et al. (2017) also found non-coding RNAs transcribed from human alpha satellite are complexed with CENP-A and CENP-C. Loss of CENP-A does not affect transcript abundance, but CENP-A and CENP-C at the targeted centromere are reduced when cenRNA is depleted. In mouse, minor repeats yield transcripts up to 4-kb long, and may impair centromeric architecture and function under stress (Bouzinba-Segard et al. 2006). In maize, nearly half of the centromeric retrotransposons (CRMs) and satellite repeats (CentC) RNA, which is larger than 40-nt in length were bound to CENH3. and siRNA-sized (22–30-nt) molecules were not detected (Topp et al. 2004).
There has been great progress in understanding centromere and kinetochore function over the last few years. A key question of how CENP-A recognizes DNA and targets the proper chromosomal location still remains. Henikoff and researchers have suggested that the replacement of histone H3 with CENH3 is often associated with active transcription, which can disrupt nucleosome and open chromatin, similar to the role of human Xist RNA in regulating X-inactivation by facilitating the replacement of histone H2 with macroH2 (Boeger et al. 2003; Plath et al. 2002; Jiang et al. 2003; Sullivan et al. 2001; Choo et al. 2001). However, Nechemia-Arbely (2019) found support for the idea that CENP-A was assembled into nucleosomes onto more than ten thousand transcriptionally active sites on the chromosome arms. DNA replication acts as an error correction mechanism to remove non-centromeric CENP-A.
RNA is also an essential structural and functional component of neocentromere chromatin. In humans, the L1 retrotransposon (~ 6-kb in size) belongs to the only active subfamily of LINEs. A significant enrichment of FL-L1b RNA (one of the elements of L1RNA) in the CENP-A bound fractions at the 10q25 neocentromeric chromatin was observed by anti-CENP-A RNA ChIP-seq, indicating that RNA transcribed from the L1 retrotransposon of a neocentromere could be incorporated into the core neocentromere chromatin and serve as a critical epigenetic determinant in chromosome remodeling, leading to neocentromere formation (Chueh et al. 2009). Taken together, ceRNAs appear to assist in cenH3 loading.
-
3.
CenRNAs are required for kinetochore structure and accurate chromosome segregation.
The kinetochore is a multiprotein complex that adheres to centromeric chromatin through the inner plate and binds to microtubules through the outer plate, which is essential to accurate chromosome segregation (Rošić et al. 2016; Yamagishi et al. 2014). Although RNA was first observed in kinetochores in the 1970s (Rieder, 1979; Braseton 1975), Topp et al. (2004) found that cenRNAs played a role in assembly and stabilization of kinetochore chromatin structure. Centromeric transcripts are bound by several kinetochore proteins that involve kinetochore assembly. Both sense and antisense cenRNA interact with the inner kinetochore protein CENP-C, as found in D. melanogaster (Rošić et al. 2014), plants (Du et al. 2010) and human cells (Wong et al. 2007). In human, CENP-C binds three different cenRNAs (Henikoff et al. 2018). Aurora B kinase interacts with ncRNA transcribed from centromeric satellite I, and knock down of satellite I RNA displays mitotic chromosomes segregation errors by inducing the defective attachment of microtubules to kinetochores (Wong et al. 2007; Indue et al. 2014). CenRNAs are also required for activation of Aurora B kinase in X. laevis eggs and mouse cells (Ferri et al. 2009; Blower 2016). CenRNAs processing contributes to proper spindle and kinetochore assembly in Xenopus egg extracts. Inhibition of transcription initiation or RNA splicing result in spindle defects (Grenfell et al. 2016). A-satellite RNA is also a key component in the assembly of other kinetochore proteins like Sgo1 (Talbert et al. 2018), CENP-A, and CENPC1 (Wong et al 2007). The over-accumulation of major and minor satellite transcripts alters meiotic kinetochore assembly and causes chromosome mis-segregation (Table 1; Fig. 1). Cell cycle-regulated cenRNAs may stabilize the binding of CENP-C to DNA, and help to recruit CENP-A loading and kinetochore assembly to regulate centromere function and facilitate accurate chromosome segregation.
-
4.
Cell-cycle-dependent cenRNAs act in pericentromeric heterochromatin assembly
During mitosis, heterochromatin formation is essential for gene regulation and maintaining centromere stability to ensure accurate chromosome segregation. As centomeres and pericentromeres are populated with enormous amounts of repeat sequence, the centromere is the most condensed and constricted region of a chromosome. However, a low level transcription of the core and the flanking pericentric regions is detected. Centromeric transcription mediated chromatin remodeling is favorable for transition of CENP-A to incorporate nucleosomes at the centromere (Georg et al. 2018). Transcription is actually required to inititiate heterochromatin formation (Reinhart and Bartel 2002). In mouse, major satellite RNAs stabilize pericentromeric heterochromatin retention of H3K9me3 methyltransferases by forming a RNA: DNA hybrid (Camacho et al. 2017). In Drosophila S2 cells, repeated RNAs are principally derived from active retrotransposons, especially gypsy elements, acting in both cis and trans on chromatin to help maintain pericentromeric hetetochromatin (Hao et al. 2020). Antisense transcripts can occur in the presence of heterochromatin, sense transcripts are repressed by Clr6 complexes (Volpe et al. 2002; Nicolas et al. 2007). Pathways to establish centromeric and pericentromeric heterochromatin have been better described in fission yeast and S. pombe. Formation of heterochromatin at centromeres relies on the RNA interference (RNAi) machinery, which involves processing of centromeric noncoding RNAs (Verdel et al. 2004; Chen et al. 2008). Similar findings also have been identified in plants (Lippman et al. 2004; Neumann et al. 2007). Transcripts from both strands of centromeric DNA are cell cycle regulated. The forward transcripts with preferential accumulation during S phase indicate the accessibility of heterochromatin structures in this phase (Chen et al. 2008).
-
5.
CenRNAs are associated with centromeres via RNA: DNA hybrids.
Topological organization of centromeric chromatin has recently gained increasing attention. R-loops are three strand nucleic acid structures consisting of an RNA: DNA hybrid and a displaced single-stranded DNA (Fig. 4). As to their functions, R-loops have been reported to be associated with DNA replication initiation (Yu et al. 2003), DNA-damage response (Hamperl et al. 2017), gene transcription (Fang et al. 2019), DNA repair (Lu et al. 2018), and genome instability (Frederic and Craig 2020). The formation of RNA–DNA hybrids is also an important mechanism of sequence-specific targeting of RNA to chromatin (Maldonado et al. 2019). Non-coding RNAs as a structural chromatin component is well documented for telomeric heterochromatin, and has been implicated to remain associated with telomeric chromatin by forming RNA: DNA hybrids that mediate telomere length and heterochromatin formation (Nakama et al. 2012; Schoeftner et al. 2009; Graf et al. 2017; Feretzaki et al. 2020). Centromeric ncRNA research falls behind that of telomeres. Apart from budding yeast, centromeric R-loops have been identified in human, rice, Arabidopsis, maize and other eukaryotic organisms (Kabeche et al. 2018; Fang et al. 2019; Xu et al. 2017) However, research on centromeric R-loops remains less explored. Centromeric R-loops are generated through RNAPII-mediated transcription during mitosis (Mishra et al. 2020). In maize, high levels of R-loops in centromeric retrotransposons led to a reduced localization of CENH3 (Liu et al. 2020). In mouse, major satellite RNAs stabilize pericentromeric heterochromatin retention of H3K9me3 methyltransferases by forming a RNA: DNA hybrid (Camacho et al. 2017). In human, R-loops are detected at centromeres in mitosis; and an R-loop-driven signaling pathway promotes faithful chromosome segregation and genome stability (Kabeche et al. 2018). Interestingly, the opposite effects of centromeric R-loops on chromosomal instability was also reported. Using hpr1∆ strains that accumulate R-loops. Mishra and other researchers find that R-loops at centromere chromatin contribute to defects in kinetochore integrity and chromosomal instability. They also found that R-loops at centromeres were not accumulated when centromeric non-coding RNA is increased (Mishra et al. 2020; Unoki et al. 2020). These findings indicate the negative and positive impact of R-loops on the function of kinetochores and centromeres. Importantly, R-loops are also observed in neocentromere regions in maize (Han et al. unpublished), which suggests a role of R-loops in neocentromere formation.
Circular RNAs (circRNAs) are a novel class of noncoding RNAs that are involved in gene expression regulation, and has been extensively explored in worm, metazoans, fruit fly, mouse, monkey, and human (Ivanov et al. 2015; Westholm et al. 2014; Fan et al. 2015; Memczak et al. 2013 and Salzman et al. 2012). Genomewide circRNAs also have been were identified in plants, including Arabidopsis thaliana, Oryza sativa, maize, wheat, barely, tomato, soybean (Ye et al. 2015; Chen et al. 2018; Wang et al. 2017a, b; Darbani et al. 2016; Zhao et al. 2017a, b; Zhou et al. 2016; Wang et al. 2017a, b; Zeng et al. 2018; Zuo et al. 2016). However, due to the limitation of bioinformatics tools identifying circRNAs and the repetitive nature of centromeric DNA, centromeric circRNAs can’t be identified easily. Liu et al (2020) first reported the role of centromeric circRNAs derived from retrotransposons in maize, which act by binding to the centromere through R-loops (Figs. 3, 4). The molecular features and function of these centromeric circRNAs are still being investigated. These clues shed new light on the function of centromeric circRNAs and R-loops, and it's an appealing line of research on the function and stabilization of centromeres.
Distribution of centromeric specific DNA repeats in plant chromosomes. a The distribution of centromeric retrotransposon of wheat (red) signals along the wheat chromosome. b The distribution of CRM1(green) and CentC(red) signals along the maize chromosome. DAPI-stained chromosomes are blue. Bar = 10 μm
Various types of cenRNAs play an important role in centromere function. The centromere-specific nucleosomes are distributed in a specific reigon of chromosome. Centromeric transcripts are processed, including small RNAs, lncRNAs, circRNAs and DNA-RNA hybrids, which are associated with CENP-A, CENP-C, Aurora B, pericentric heterochromatin and so on
Conclusion and perspective
How does CENH3 recognize and target centromeric DNA? Which factors are involved in centromere assembly and function? These questions remain subjects for further investigation. Besides, the repetitive nature of centromeric DNA and indefinite origin and length of RNA will continue to challenge centromere transcription and identity. Studying the function of centromeric circRNA and R-loops appears to offer a breakthrough. Centromeric DNA transcription and RNA localization is independent of CENP-A (McNulty 2017). Variant centromeric transcripts interact properly with different centromere proteins. Dissecting different centromeric RNA-binding proteins might bring some new clues for determining the mechanisms of centromere formation and function, centromere inactivation or other biological processes.
References
Arabidopsis Genome I (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408:796–815
Arunkumar G, Melters DP (2020) Centromeric transcription: a conserved swiss-army knife. Genes 11:911
Bergmann JH, Rodríguez MG, Martins NMC, Kimura H, Kelly DA, Masumoto H et al (2011) Epigenetic engineering shows H3K4me2 is required for HJURP targeting and CENP-A assembly on a synthetic human kinetochore. EMBO J 30:328–340
Blower MD (2016) Centromeric transcription regulates aurora-B localization and activation. Cell Rep 15:1624–1633
Boeger H, Griesenbeck J, Strattan JS, Kornberg RD (2003) Nucleosomes unfold completely at a transcriptionally active promoter. Mol Cell 11:1587–1598
Bouzinba-Segard H, Guais A, Francastel C (2006) Accumulation of small murine minor satellite transcripts leads to impaired centromeric architecture and function. Proc Natl Acad Sci U S A 103:8709–8714
Braselton JB (1975) Ribonucleoprotein staining of Allium cepa kinetochores. Cytobiologie 12:148–151
Bury L, Moodie B, Ly J, Mckay LS, Miga KH, Cheeseman IM (2020) Alpha-satellite RNA transcripts are repressed by centromere-nucleolus associations. Elife 9:e59770
Camacho OV, Galan C, Rosowska KS, Ching R, Gamalinda M, Karabiber F, Velazquez IDLR, Engist B, Koschorz B, Shukeir N et al (2017) Major satellite repeat RNA stabilize heterochromatin retention of Suv39h enzymes by RNA-nucleosome association and RNA:DNA hybrid formation. Elife 6:e25293
Chan FL, Wong LH (2012) Transcription in the maintenance of centromere chromatin identity. Nucleic Acids Res 40:11178–11188
Chen ES, Saitoh S, Yanagida M, Takahashi K (2003) A cell cycle-regulated GATA factor promotes centromeric localization of CENP-A in fission yeast. Mol Cell 11:175–187
Chen ES, Zhang K, Nicolas E, Hugh PC, Zofall M, Grewal SS (2008) Cell cycle control of centromeric repeat transcription and heterochromatin assembly. Nature 451:734–737
Chen L, Zhang P, Fan Y et al (2018) Circular RNAs mediated by transposons are associated with transcriptomic and phenotypic variation in maize. New Phytol 217:1292–1306
Choi ES, Stralfors A, Castillo AG, Durand-Dubief M, Ekwall K, Allshire RC (2011) Identification of noncoding transcripts from within CENP-A chromatin at fission yeast centromeres. J Biol Chem 286:23600–23607
Choi ES, Strålfors A, Catania S, Castillo AG, Svensson JP, Pidoux AL, Ekwall K, Allshire RC (2012) Factors that promote H3 chromatin integrity during transcription prevent promiscuous deposition of CENP-A (Cnp1) in fission yeast. PLoS Genet 8:e1002985
Choo KHA (2001) Domain organization at the centromere and neocentromere. Dev. Cell 1:165–177
Chueh AC, Northrop EL, Brettingham-Moore KH, Choo KH (2009) LINE retrotransposon RNA is an essential structural and functional epigenetic component of a core neocentromeric chromatin. PloS Genet 5:e1000354
Cleveland DW, Mao Y, Sullivan KF (2003) Centromeres and kinetochores: from epigenetics to mitotic checkpoint signaling. Cell 112:407–421
Collins KA, Castillo AR, Tatsutani SY, Biggins S (2005) De novo kinetochore assembly requires the centromeric histone H3 variant. Mol Biol Cell 16:5649–5660
Corless S, Höcker S, Erhardt S (2020) Centromeric RNA and its function at and beyond centromeric chromatin. J Mol Biol 432:4257–4269
Darbani B, Noeparvar S, Borg S (2016) Identification of circular RNAs from the parental genes involved in multiple aspects of cellular metabolism in Barley. Front Plant Sci 7:776
Dhatchinamoorthy K, Mattingly M, Gerton JL (2018) Regulation of kinetochore configuration during mitosis. Curr Genet 64:1197–1203
Du Y, Topp CN, Dawe RK (2010) DNA binding of centromere protein C (CENPC) is stabilized by single-stranded RNA. PLoS Genet 6:e1000835
Dunleavy EM, Beier NL, Gorgescu W, Tang J, Costes SV, Karpen GH (2012) The cell cycle timing of centromeric chromatin assembly in Drosophila meiosis is distinct from mitosis yet requires CAL1 and CENP-C. PLoS Biol 10:e1001460
Fan X, Zhang X, Wu X, Guo H, Hu Y, Tang F, Huang Y (2015) Single-cell RNA-seq transcriptome analysis of linear and circular RNAs in mouse preimplantation embryos. Genome Biol 16:148
Fang Y, Chen LF, Lin K, Feng YL, Zhang PY, Pan XC et al (2019) Characterization of functional relationships of R-loops with gene transcription and epigenetic modifications in rice. Genome Res 29:1287–1297
Feretzaki M, Pospisilova M, Fernandes RV, Lunardi T, Krejci L, Lingner J (2020) RAD51-dependent recruitment of TERRA ncRNA to telomeres through R-loops. Nature 587:303–308
Ferri F, Bouzinba-Segard H, Velasco G et al (2009) Non-coding murine centromeric transcripts associate with and potentiate Aurora B kinase. Nucleic Acids Res 37:5071–5080
Frederic C, Craig JB (2020) Emerging roles for R-loop structures in the management of topological stress. J Biol Chem 3:4684–4695
Georg OM, Bobkov NG, Patrick H (2018) Centromere transcription allows CENP-A to transit from chromatin association to stable incorporation. J Cell Biol 217:1957–1972
Graf M et al (2017) Telomere length determines TERRA and R-loop regulation through the cycle. Cell 170:2–85
Grenfell AW, Heald R, Strzelecka M (2016) Mitotic noncoding RNA processing promotes kinetochore and spindle assembly in Xenopus. J Cell Biol 214:133–141
Hamperl S, Bocek MJ, Saldivar JC, Swigut T, Cimprich KA (2017) Transcription-replication conflict orientation modulates r-loop levels and activates distinct DNA damage responses. Cell 70:774–786
Hao YJ, Wang DP, Wu SH, Li X, Shao CW, Zhang P, Chen JY, Lim DH, Fu XD et al (2020) Active retrotransposons help maintain pericentromeric heterochromatin required for faithful cell division. Genome Res 30:1570–1582
Heieh CL, Xia J, Lin HF (2020) MIWI prevents aneuploidy during meiosis by cleaving excess satellite RNA. EMBO J 39:e103614
Henikoff S, Talbert PB (2020) What makes a centromere? Exp. Cell Res. 389: 111895 Henikoff S, Ahmad K, Malik HS (2001) The Centromere Paradox: Stable Inheritance with Rapidly Evolving DNA. Science 293:1098–1110
Henikoff S, Buell CR, Jiang J (2004) Sequencing of a rice centromere uncovers active genes. Nat Genet 36:138–145
Hill A, Bloom K (1987) Genetic manipulation of centromere function. Mol Cell Biol 7:2397–2405
Ishikura S, Nakabayashi K, Nagai M, Tsunoda T, Shirasawa S (2020) ZFAT binds to centromeres to control noncoding RNA transcription through the KAT2B-H4K8ac-BRD4 axis. Nuclei Acids Res 48:10848–10866
Ivanov S, Memczak E, Wyler F, Torti HT, Porath MR, Orejuela M, Piechotta EY, Levanon M, Landthaler C, Dieterich N, Rajewsky, (2015) Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep 10:170–177
Jansen LE, Black BE, Foltz DR, Cleveland DW (2007) Propagation of centromeric chromatin requires exit from mitosis. J Cell Biol 176:795–805
Jiang J, Birchler JA, Parrott WA, Dawe RK (2003) A molecular view of plant centromeres. Trends Plant Sci 8:570–575
Kabeche L, Nguyen HD, Buisson R, Zou L (2018) A Mitosis-specific and R loop-driven ATR pathway promotes faithful chromosome segregation. Science 359:108–114
Kobayashi N, Suzuki Y, Schoenfeld LW, Müller CA et al (2015) Discovery of an unconventional centromere in budding yeast redefines evolution of point centromeres. Curr Biol 3:2026–2033
Lee HR, Neumann P, Macas J, Jiang J (2006) Transcription and evolutionary dynamics of the centromeric satellite repeat CentO in rice. Mol Biol Evol 23:2505–2520
Lefrançois P, Euskirchen GM, Auerbach RK, Rozowsky J, Gibson T, Yellman CM, Gerstein M, Snyder M (2009) Efficient yeast ChIP-Seq using multiplex short-read DNA sequencing. BMC Genomics 10:37
Lermontova I, Fuchs J, Schubert V, Schubert I (2007) Loading time of the centromeric histone H3 variant differs between plants and animals. Chromosoma 116:507–510
Lermontova I, Rutten T, Schubert I (2011) Deposition, turnover, and release of CENH3 at Arabidopsis centromeres. Chromosoma 120:633–640
Ling YH, Wing K, Yuen Y (2019) Centromeric non-coding RNA as a hidden epigenetic factor of the point centromere. Curr Genet 65:1165–1171
Lippman Z, Martienssen R (2004) The role of RNA interference in heterochromatic silencing. Nature 431:364–370
Liu Y, Su H, Zhang J, Liu Y, Feng C, Han FP (2020) Back-spliced RNA from retrotransposon binds to centromere and regulates centromeric chromatin loops in maize. PLoS Biol 18:e3000582
Lu WT, Hawley BR, Skalka GL, Baldock RA, Smith EM, Bader AS, Malewicz M, Watts FZ, Wilczynska A, Bushell M (2018) Drosha drives the formation of DNA: RNA hybrids around DNA break sites to facilitate DNA repair. Nat Commun 9:532
Lv J, Yu K, Wei J, Gui H, Liu CX, Liang D, Wang YL et al (2020) Generation of paternal haploids in wheat by genome editing of the centromeric histone CENH3. Nat Biotechnol. 38:1397–1401
Maison C, Quivy JP, Probst AV, Almouzni G (2010) Heterochromatin at mouse pericentromeres: a model for de novo heterochromatin formation and duplication during replication. Cold Spring Harb Symp Quant Biol 75:155–165
Maldonado R, Schwartz U, Silberhorn E, Längst G (2019) Nucleosomes Stabilize ssRNA-dsDNA Triple Helices in Human Cells. Mol Cell 73:1243–1254
May BP, Lippman ZB, Fang Y, Spector DL, Martienssen RA (2005) Differential regulation of strand-specific transcripts from arabidopsis centromeric satellite repeats. PLoS Genet 1:e79
Mellone BG, Grive KJ, Shteyn V, Bowers SR, Oderberg I, Karpen GH (2011) Assembly of drosophila centromeric chromatin proteins during mitosis. PLoS Genet 7:e1002068
Memczak S, Jens S, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M et al (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495:333–338
Miga KH, Shoshani O, Asron A, McMahon MA, Lee AY et al (2019) DNA replication acts as an error correction mechanism to maintain centromere identity by restricting CENP-A to centromeres. Nat Cell Biol 21:743–754
Mishra PK, Chakrabortyb K, Yehc E, Feng WY, Bloomc KS, Basraia M (2020) R-loops at centromeric chromatin contribute to defects in kinetochore integrity and chromosomal instability in budding yeast. Mol Biol Cell 1:mbcE20060379
MuNulty SM et al (2017) Human centromeres produce chromosome-specific and array-specific alpha satellite transcripts that are complexed with CENP-A and CENP-C. Dev Cell 42:226–240
Nagaki K, Cheng Z, Ouyang S, Talbert PB, Kim M, Jones KM, Nechemia-Arbely Y, Ideue T, Cho Y, Nishimura K, Tani T (2014) Involvement of satellite I noncoding RNA in regulation of chromosome segregation. Genes Cells 19:528–538
Nakano M, Cardinale S, Noskov VN, Gassmann R, Vagnarelli P, Kandels-lewis S, Larionov V, Earnshaw WC, Masumoto H (2008) Inactivation of a human kinetochore by specific targeting of chromatin modifiers. Dev Cell 14:507–522
Neumann P, Yan H, Jiang J (2007) The centromeric retrotransposons of rice are transcribed and differentially processed by RNA interference. Genetics 176:749761
Nicolas E, Yamada T, Cam HP, Fitzgerald PC, Kobayashi R, Grewal SIS (2007) Distinct roles of HDAC complexes in promoter silencing, antisense suppression and DNA damage protection. Nat Struct Mol Biol 14:372–380
Ohkuni K, Kitagawa K (2011) Endogenous Transcription at the Centromere Facilitates Centromere Activity in Budding Yeast. Curr. Biol. 21:1695–1703
Ólafsson G, Thorpe PH (2020) Polo kinase recruitment via the constitutive centromere-associated network at the kinetochore elevates centromeric RNA. PLoS Genet 18:e1008990
Path K, Mlynarcayk-Evans S, Nusinow D, Panning B (2002) Xist RNA and the mechanism of X chromosome inactivation. Annu Rev Genet 36:233–278
Pluta AF, Mackay AM, Ainsztein AM, Goldberg IG, Earnshaw WC (1995) The centromere: hub of chromosomal activities. Science 270:1591–1594
Quénet D, Dalal Y (2014) A long non-coding RNA is required for targeting centromeric protein a to the human centromere. Elife 3:e03254
Reinhart BJ, Bartel DP (2002) Small RNAs correspond to centromere heterochromatic repeats. Science 297:1831
Rieder CL (1979) Ribonucleoprotein staining of centrioles and kinetochores in newt lung cell spindles. J Cell Biol 80:1–9
Rošić S, Köhler F, Erhardt S (2014) Repetitive centromeric satellite RNA is essential for kinetochore formation and cell division. J Cell Biol 207:335–349
Saffery R, Sumer H, Hassan S, Wong LH, Craig JM, Todokoro K, Anderson M, Safford A, Choo KHA (2003) Transcription within a functional human centromere. Mol Cell 12:509–516
Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO (2012) Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 7:e30733
Schoeftner S, Blasco MA (2009) A “higher order” of telomere regulation: telomere heterochromatin and telomeric RNAs. EMBO J 28:2323–2336
Schubert V, Lermontova I, Shubert I (2014) Loading of the centromeric histone H3 variant during meiosis – how does it differ from mitosis? Chromosoma 123: 491–7 Shao JJ, Wang LQ, Liu XY, Yang M, Chen HM, Wu B, Liu C (2019) Identification and characterization of circular RNAs in Ganoderma lucidum. Sci Rep 9:16522
Shuhei I, Kazuhiko N, Masayoshi N, Toshiyuki T, Senji S (2020) ZFAT binds to centromeres to control noncoding RNA transcription through the KAT2B–H4K8ac–BRD4 axis. Nucleic Acids Res 4:10848–10966
Stimpson KM, Sullivan BA (2010) Epigenomics of centromere assembly and function. Curr Opin Cell Biol 22:772–780
Su H, Liu YL et al (2016) Dynamic chromatin changes associated with de novo centromere formation in maize euchromatin. Plant J 88:854–866
Su H, Liu YL et al (2019) Centromere Satellite Repeats Have Undergone Rapid Changes in Polyploid Wheat Subgenomes. Plant Cell 31:2015–2051
Sullivan KF (2001) A solid foundation: functional specialization of centromeric chromatin. Curr Opin Genet 11:182–188
Sullivan BA, Karpen GH (2004) Centromeric chromatin exhibits a histone modification pattern that is distinct from both euchromatin and heterochromatin. Nat Struct Mol Biol 11:1076–1083
Tallbert PB, Henikoff S (2018) Transcribing centromeres: noncoding RNAs and kinetochore assembly. Trends Genet 34:587–599
Topp CN, Zhong CX, Dawe RK (2004) Centromere-encoded RNAs are integral components of the maize kinetochore. Proc Natl Acad Sci USA 101:15986–15991
Unoki M, Sharif J, Saito YC et al (2020) CDCA7 and HELLS suppress DNA:RNA hybrid-associated DNA damage at pericentromeric repeats. Sci Rep 10:17865
Verdel A, Jia S, Gerber S, Sugiyama T, Gygi S, Grewal SI et al (2004) RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303:672–676
Volpe TA, Kider C, Hall IM, Teng G, Grewal SIS, Martienssen RA (2002) Regulation of heterochromatic silencing and histone H3 Lysine-9 Methylation by RNAi. Sci 297:1833–1837
Wang Z, Liu Y, Li D et al (2017a) Identification of circular RNAs in Kiwifruit and their species-specific response to bacterial canker pathogen invasion. Front Plant Sci 8:413
Wang Y, Yang M, Wei S et al (2017b) Identification of circular RNAs and their targets in leaves of Triticum aestivum L under dehydration stress. Front Plant Sci 7:224
Westholm JO, Miura P, Olson S, Shenker S, Joseph B, Sanfilippo P, Celniker SE, Graveley BR, Lai EC (2014) Genome-wide analysis of Drosophila circular RNAs reveals their structural and sequence properties and age-dependent neuralaccumulation. Cell Rep. 9:1966–1980
Wong LH, Brettingham-Moore KH, Chan L, Quach JM, Anderson MA, Northrop EL, Hannan R, Saffery R, Shaw ML, Williams E, Choo KH (2007) Centromere RNA is a key component for the assembly of nucleoproteins at the nucleolus and centromere. Genome Res 17:1146–1160
Xu W, Xu H, Li K, Fan YX, Liu Y, Yang XR, Sun QW (2017) The R-loop is a common chromatin feature of the Arabidopsis genome. Nat Plants 3:704–714
Yamagishi Y, Sakuno T, Goto Y, Watanabe Y (2014) Kinetochore composition and its function: lessons from yeasts. FEMS Microbiol Rev 38:185–200
Ye CY, Liu C, Liu C, Zhu QH, Fan LJ (2015) Widespread noncoding circular RNAs in plants. New Phytol 208:88–95
Yu K, Chedin F, Hsieh CL, Wilson TE, Lieber MR (2003) R-loops at immunoglobulin class switch regions in the chromosomes of stimulated B cells. Nat Immunol 4:442–451
Zeng RF, Zhou JJ, Hu CG et al (2018) Transcriptome-wide identification and functional prediction of novel and flowering-related circular RNAs from trifoliate orange (Poncirus trifoliata L. Raf.). Planta 247:1191–1202
Zhang X, Ma X, Ning L, Li Z, Zhao K, Li K, He J, Yin D (2019) Genome-wide identification of circular RNAs in peanut (Arachis hypogaea L). BMC Genomics 20:653
Zhao T, Wang L, Li S et al (2017a) Characterization of conserved circular RNA in polyploid Gossypium species and their ancestors. FEBS Lett 591:3660–3669
Zhao W, Cheng Y, Zhang C et al (2017b) Genome-wide identification and characterization of circular RNAs by high throughput sequencing in soybean. Sci Rep 7:5636
Zuo J, Wang Q, Zhu B et al (2016) Deciphering the roles of circRNAs on chilling injury in tomato. Biochem Biophys Res Commun 479:132–138
Acknowledgments
This work was supported by the National Natural Science Foundation of China (31920103006 and 31630049) and a National Science Foundation plant genome grant (IOS-1444514).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, Q., Liu, Y., Shi, Q. et al. Emerging roles of centromeric RNAs in centromere formation and function. Genes Genom 43, 217–226 (2021). https://doi.org/10.1007/s13258-021-01041-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-021-01041-y