Abstract
The aim of this paper is to introduce a novel method using short-term EEG signals to separate depressed patients from healthy controls. Five common frequency bands (delta, theta, alpha, beta and gamma) were extracted from the signals as linear features, as well as, wavelet packet decomposition to break down signals into certain frequency bands. Afterwards, two entropy measures, namely sample entropy and approximate entropy were applied on the wavelet packet coefficients as nonlinear features, and significant features were selected via genetic algorithm (GA). Three machine-learning algorithms were used for classification; including support vector machine (SVM), multilayer perceptron (MLP) a novel enhanced K-nearest neighbors (E-KNN), which uses GA to optimize the feature-space distances and provides a feature importance index. The highest accuracy obtained by using frequency-based features was from gamma oscillations which resulted in 91.38%. Performance of nonlinear features were better compared to the frequency-based features and the results showed 94.28% accuracy. The combination of the features showed 98.44% accuracy with the new proposed E-KNN classifier.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The world health organization (WHO) has predicted that depression will become the second leading cause of disability by 2020 [1]. If it is diagnosed at an early stage, the patient’s condition will be significantly improved via treatment, medication or lifestyle changes.
Electroencephalogram (EEG) is a signal recorded non-invasively and represents the brain electrical activity. Due to its high temporal resolution and reliability, EEG seems to be an effective tool for diagnosis of depression. Hinrikus et al. [2] presented a novel spectral asymmetry index (SASI) which is based on the calculation of relative power differences of two specific frequency bands of the EEG signal. Their calculations exclude the central frequency band (alpha band) and their analysis is on 30-min recordings of 36 subjects (18 normal and 18 depressed). Puthankattil et al. [3] studied 30 subjects with 5-min EEG recording in resting state condition. Initially an eight-level discrete wavelet transform was used to decompose the signal into high and low frequency components (detail and approximate coefficients respectively), consequently, wavelet entropies (WE) based on the Shannon entropy and relative wavelet energy (RWE) were extracted from each level of the decomposition. Hosseinifard et al. [4] experimented with 45 subjects in each group (healthy and depressed). They extracted four non-linear features, namely, detrended fluctuation analysis (DFA) which displays long-term correlation of the signal, Higuchi’s fractal dimension (HFD), correlation dimension and Lyapunov exponent (LE) which measures the randomness of signal; and four band powers (delta, theta, alpha, beta) as linear features. Next, k-nearest neighbor, logistic regression and linear discriminant analysis (LDA) were used for classification. Their study indicates that non-linear features can discriminate between normal and depressed patients more efficiently than linear features. Alpha and theta band powers also showed higher discrimination rate specially in the left-hemisphere. However, they used long time EEG signal which might be undesirable in real world applications. Faust et al. [5] performed an experiment on two thousand samples of the EEG recordings from 30 subjects. The process starts with a two-level wavelet packet decomposition followed by five different entropy techniques namely, approximate entropy, sample entropy, Rényi entropy and Bispectral phase entropy applied on wavelet packet coefficients. In the mentioned study, Probabilistic neural network (PNN) achieved the highest discrimination rate compared to other models that were used. Acharya et al. [6] studied 30 subjects and proposed a depression diagnosis index to automatically discriminate between normal and depressed subjects. They extracted non-linear features from EEG signals consisting of fractal dimension, largest Lyapunov exponent (LLE), sample entropy, DFA, Hurst’s exponent, higher order spectra and recurrence quantification analysis. Mumtaz et al. [7] performed an experiment on 33 MDD patients and 30 normal participants and derived linear features (Band powers and alpha inter-hemispheric asymmetry) from EEG signals. They used receiver operating characteristic (ROC) to rank the most important features. Bachmann et al. [8] introduced a simple method to detect the depression depending on a single channel EEG based on spectral asymmetry index (SASI) and non-linear detrended fluctuation (DFA) analysis on 17 depressed and 17 control subjects. The first 5 min of each recording was selected in this analysis. Acharya et al. [9] analyzed 5-min EEG signals of 15 normal and 15 depressed patients to introduce an automatic depression detection method based on one dimensional convolutional neural network (CNN). The proposed 13-layered CNN was composed of five conventional layers, five pooling layers and three fully-connected layers. They also demonstrated that EEG signals recorded from the right hemisphere were more suitable for deep learning classification. Fitzgerald et al. [10] reviewed many studies suggesting that gamma band is able to discriminate more efficiently between depressed patients and healthy controls. Azizi et al. [11] studied recordings of 40 subjects (20 subjects in each group) and they presented a number of new differences based on geometrical methods: centroid to centroid distance, centroid to 45-degree line shortest distance and incenter radius. Significance of features were identified by t-test and their study concluded that right hemisphere features are more distinct among the two groups. A number of researchers also have used Entropy-based features from EEG signals in conjunction with deep learning schemes. Gao et al. [12] presented an innovative spatial–temporal convolution neural network (ESTCNN) based on EEG signals in order to identify driver fatigue. Chai et al. [13] analyzed 43 participants for classifying fatigue and alert states. They used autoregressive modeling and Bayesian neural network for feature extraction and classification respectively. Li et al. [14] experimented with 25 healthy controls and 17 PD (Parkinson Disease) patients and the preprocessing stage was done by applying three-level discrete Wavelet transform for splitting EEG signals into approximate and detail coefficients. Afterwards, Sample Entropy was used for extracting features from approximate coefficients. Finally, optimal center constructive covering algorithm (O_CCA) was used for discrimination between the groups. Al-ani [14] et al. introduced a novel method for recognizing the most relevant channels in each time segment of EEG signals to deal with the curse of high dimensionality. Also, for the preprocessing stage, many researchers [15,16,17,18,19,20,21] have proposed automated techniques for real-time removing of ocular, muscular and eye blinks artifacts from EEG signals.
The aim of this study is to provide a novel method on short-term EEG signals for classifying depressed and healthy control subjects according to the power and the complexity of frequency components of the signals and to propose a novel way to identify useful features and hence channels. Five frequency band powers (delta, theta, alpha, beta and gamma) and a combination of the wavelet packet decomposition and entropies are used as inputs to the classifiers. Genetic algorithm is exploited to reduce the dimensionality of the feature space by removing irrelevant features. We also investigate the efficiency of an enhanced KNN classification method alongside support vector machine (SVM) and multilayer perceptron (MLP).
Material and methods
Database
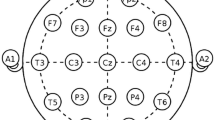
The data set used in this paper is provided by Mumtaz et al. [22], which is open to the public. Recordings include 34 depressed patients (17 females) with ages ranging from 27 to 53 and 30 normal subjects (9 females) with ages ranging from 22 to 53. EEG data was recorded in the eyes-closed state for 5 min. Recordings were obtained from 19 electro-cap electrodes that were mounted on the scalp according to 10–20 international standard electrode position classification system. The sampling frequency was also 250 Hz. A notch filter was applied to reject 50 Hz power line noise. All EEG signals were band-pass-filtered with cutoff frequencies at 0.5 Hz and 70 Hz.
Preprocessing
Wavelet-based de-noising
Discrete wavelet transform (DWT) can be defined as an implementation of wavelet transform (WT) using discrete set of wavelets that complies with certain regularities. After applying the wavelet transform, the signal is decomposed into detail and approximate coefficients. Afterwards, a threshold is to remove the coefficients corresponding to the noisy signal and artifacts [23]. Finally, the artifact-free signal will be reconstructed with all the detail and last level approximate coefficients. In this study, the db8 mother wavelet function [23] was selected to decompose noisy signals.
Feature extraction
Band power
Decomposing data into practically distinguishable frequency bands such as delta (0.5–4 Hz), theta (4–8 Hz), alpha (8–13 Hz), beta (13–30 Hz), and gamma (30–50 Hz) is one of the most commonly used methods to extract information from EEG signals. Initially, we determined an estimate of the power spectral density to compute average band powers in different frequencies. Welch’s periodogram method for estimating power spectrum includes averaging of Fourier transform on the successive windowed signal with overlapping segments [24]. For every channel, Welch’s method was performed to compute power spectrum and the parameters used were Hamming window and 50% overlap between segments.
Wavelet packet decomposition
Wavelet packet decomposition (WPD) is an extended version of classical wavelet decomposition. In discrete wavelet transform (DWT) signals are categorized into high and low frequency components known as detail and approximation coefficients respectively [25, 26]. The approximation coefficients then are split into next-level approximate and detail coefficients and the process continues based on the desired wavelet level. In comparison with DWT, WPD passes data to more filters and also decomposes detail coefficients in addition of approximation coefficients at each level (Fig. 1). Hence more frequency resolution can be obtained by implementing a wavelet packet transform and this method shows better performance in applications where higher frequency components represent more valuable information. Through trial and error and based on classification performance, we used four level wavelet decomposition and db4 as mother wavelet function.
Approximate entropy
Approximate entropy (ApEn) is a type of embedding entropies which is directly performed on the time-series signals. It is a criterion designed for measuring the amount of complexity or irregularity of the system [27, 28]. ApEn is less affected by the noise and can be used to process short-length data. Predictable and regular time-series result in low ApEn value while highly unpredictable time-series show greater ApEn. Briefly, ApEn is calculated by negative logarithm probability with series of length m to predict new series with length of m + 1 within a filter level r for N data points \(x\left(1\right), x\left(2\right),\dots .x\left(N\right)\). ApEn can be obtained using the following equation:
\({C}_{k}^{m}\left(r\right)\) is the correlation integral and parameters are: N refers to number of samples, m refers to embedding dimension or the number of lagged points used for making prediction and r refers to noise threshold.
Sample entropy
Sample entropy (SampEn) is an improvement of approximate entropy (ApEn) [29, 30]. Compared to ApEn, this technique is less affected by the time-series length. Lower values of SampEn specify more self-similarity or less noise in data. SampEn value for data of length N can be extracted by Eq. 2:
where \({U}^{m}\) is the number of vectors having the \(d[{X}_{m}\left(k\right),{X}_{m+1}\left(j\right)]<r\); N, m and r are number of samples, embedding dimension and tolerance respectively and \(d\) is a distance measure.
Feature selection
Feature selection is one of the main concepts in machine learning which extremely impacts the performance of the classification. In this paper, Genetic algorithm (GA) is used as main feature selection method. GA is an iterative process; where at each iteration, it produces new population called new generation. To create next generation, genetic algorithm selects certain individuals in the current population known as parents who contribute their genes to their individuals in next generation known as children [31, 32]. A fitness function decides which individuals will be selected for reproduction based on fitness scores for each individual. In this study, through trial and error, the size of population was set to 50, crossover rate and mutation rates were fixed to 80% and to 10% respectively.
Classification
The aim of classification is to accurately predict the labels for each category in the dataset. In this section, the selected features by GA were used as inputs to the classifiers; Based on these selected features, the signal is classified as normal and abnormal (depressed). In this study, three different classifiers were used which will be discussed shortly.
Enhanced KNN
In this paper, we propose Enhanced KNN, a generalization of the famous classification algorithm. K-nearest neighbor is a non-parametric machine-learning algorithm and it’s simple to execute and comprehend. KNN works by finding the distances between a test sample the specified number of samples nearest to the test sample, then votes for the most frequent label [33]. In general, the KNN classification implies that all features have the equal weights and equal impact on the performance of the classifier. Obviously, noisy and irrelevant features in the training set which have the same weights as the highly relevant features, reduce the classification accuracy from the optimal desired value. Feature weighting is an approach for determining the influence of each feature on the classification performance. In this work, we used a weighted KNN model based on the genetic algorithm in order to minimize the adverse effects of noisy training data. The modified method involves scaling more important features by associating greater weights to them, while applying the same process to less sensitive features with smaller weights. From the feature-space distance perspective, the total distance between two samples which consists of the summation of distances between all N features will be less affected by a redundant feature, and is more sensitive to the discriminative one. The representation of the distance function and weighted features can be written as,
where \(x and y\) denote two samples in the dataset. The distances between the unlabeled sample \(x\) and all of the \(M\) training examples \({y}_{m}\) will result in \({D}_{x,{y}_{m}}\). Then based on the k-value associated with the KNN, the label is decided based on the labels of those k samples that have the least distance compared to the unknown sample. Variable \(p\) shows the Minkowski distance parameter that results in Euclidean and Manhattan distance for \(p\) values of 1 and 2 respectively. In traditional KNN, the k nearest prototypes with distances calculated from the above equation determine a new samples class, whereas in our proposed method each of the \(N\) features have an optimum contribution to the summation called \(w\left(n\right)\) and to the result, which is found using the training samples:
where \(f\) refers to feature vector containing samples from both training and test sets denoted by \(x,y\) and \(w\) refers to weights computed for each feature respectively. Putting all that together the final proposition would be:
All of the weights in this technique were selected by genetic algorithm so that an optimum representation of the features can be achieved. The population size was set to 50. The crossover rate also set to 80%. The elite count number was two and the number of nearest neighbors used in this work was three (k = 3).
Linear SVM
Support vector machine (SVM) is a powerful learning model for classification problems which uses a decision boundary known as a hyperplane to separate data points in different categories. There are many possible hyperplanes that can be selected for classifying data although the aim of the support vector machine algorithm is to maximize this margin distance between hyperplane and closest data points in each category in an N-dimensional space [34].
MLP
Multilayer perceptron is one of the most common classification algorithms for the pattern recognition. A typical neural network consists of a number of processing units called neurons associated by learning weights. These weights are determined in a repetitive process using training samples [35]. Multilayer perceptron is composed of three main successive layers: an input layer where each neuron corresponds to a feature data, a hidden layer (20 neurons were used in this work) consists of neurons as training vectors and output layer that make predictions for unknown data samples. The neural network classification method used in this study was trained by back propagation algorithm. The most prevalent activation functions are the Rectified Linear Unit (ReLU) and Sigmoid, which are used in hidden and output layers respectively. These functions were also selected for this study due to better performance.
Evaluation
To prevent overfitting in the predictive model, tenfold cross validation (10-CV) was used to validate the outcomes of classification. The purpose of this technique is to assess the potential of the model to predict new data that were not used during evaluating of the model [36, 37]. The 10-CV was performed by dividing the initial dataset into tenfolds of complement subsets. First fold regarded as the validation set and the remaining folds regarded as the training set. Sensitivity, Specificity and Accuracy extracted from confusion matrix were provided by following equations. TP, TN, FP, FN stand for true positive, true negative, false positive and false negative respectively.
Results
In this work, student’s t-test was performed in order to verify the usefulness of the features. A t-test is the most common statistical procedure which uses variances for investigating the possibility of a significant difference in the mean of two different classes. For classification complications, low p-value is required which indicates that two classes are independent. An alpha of 0.05 is used as the limit for the validity in analysis. p-Value less than 0.05 denies the null hypothesis and deduces that there is a notable difference.
Frequency results
The p-value of different power bands are summarized in Table 1 using T-test. Based on the results, O1, O2, F7 and T5 channels of delta band power differ significantly between two groups (P < 0.05). Theta band power also demonstrates distinct p-values in channels O1, O2, T3, T5, T6, P4 and F7. For Beta band, all channels expect for O1 and O2 show notable discrimination rates. Gamma powers represent significant difference between depressed and control in all channels while no channel is significantly different in Alpha band.
Time–frequency results
The p-value of nonlinear features is shown in Tables 2 and 3 for the sub-bands with the lowest overall p-value. All non-linear features extracted from signals with wavelet packet decomposition and entropies applied on their coefficients were significant (P < 0.05).
Classification results
Frequency results
The results with power bands are summarized in Table 4. The number of input features was 19 according to the number of channels. Highest accuracy was achieved from gamma oscillations in all the three classifiers. Combination of the all band powers improved the accuracy of the classifiers. The highest accuracy was gained from combination of the band powers with E-KNN which resulted in 92.00%.
Time–frequency results
Table 5 shows the results of nonlinear features with the entropies applied on wavelet packet coefficients. Best accuracy was achieved from a combination of the features when E-KNN was used as classification algorithm.
Combinations of features and features selection results
Table 6 summarizes the classification characteristics of the proposed approach in this study. According to the table, using all features alongside the linear SVM classifier yields 93.75% accuracy. Furthermore, as Table 7 shows, the proposed weighted E-KNN classifier containing 14 features (Fig. 2) selected with GA yields 98.44% accuracy (specificity 100%, sensitivity = 96.80%). Also, Fig. 3 shows the confusion matrix for the aforementioned results with the E-KNN method. Figure 2 Demonstrates the 14 features selected and weighted by genetic algorithm in which \(x, w\) indicate features and weights respectively. According to the figure, highest weight calculated by genetic algorithm \({w}_{9}\)=1.92, is related to O1 channel and gamma band power features. Other weights are: \({w}_{10}\)=1.40, \({w}_{8}\)=1.05 and \({w}_{7}\)=0.91, representing O2, FP2 and P3 channels respectively, which are also gamma band power features. \({w}_{14}\)=0.83 is associated with T6 channel and sample entropy features from (1,1) sub-band. \({w}_{12}\)=0.72 connects to Pz electrode and sample entropy in (2,3) sub-band. The other weights, \({w}_{1}\)=0.44, \({w}_{2}\)=0.60, \({w}_{3}\)=0.61, \({w}_{4}\)=0.59, \({w}_{5}\)=0.67, \({w}_{7}\)=0.53 and \({w}_{13}\)=0.61 were in the same range and are related to approximate entropy. Lowest weight \({w}_{6}\)=0.16 was associated with C3 channel and gamma band power.
To further demonstrate how the E-KNN method works, Fig. 4 shows O1, O2 and P4 gamma power features (p < 0.05) before and after weighting by genetic algorithm in 3-dimensional space. According to the figures, an increase in the discrimination of the samples can be clearly observed, which improves the process of classification of healthy and depressed subjects by creating a better decision boundary.
Also, as depicted in Fig. 5, the GA optimization increased the total accuracy in a reasonable number of generations. The process was done on tenfold cross validation to prevent overfitting.
Discussion
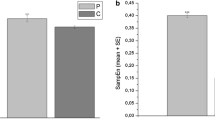
As it was seen in the results section, gamma powers were more distinct in all channels (Table 1). In all earlier discussed channels, the mean gamma powers were higher for depressed patients than normal group as depicted in Fig. 6. De Aguiar Neto et al. [38] reviewed EEG-based depression detection studies and concluded that gamma and theta oscillations as biomarkers can differentiate between depressed patients and healthy subjects. Strelets et al. [39] also reported that the power of the gamma rhythms in frontal and parietal cortex was considerably higher in depressed patients in comparison of normal subjects. Non-linear classification results indicate that SVM classifier yields the highest classification performance when entropies are applied on the higher frequency components of wavelet packet decomposition consisting of S(1,1), S(2,3), S(3,7) which corresponds to (35–70 Hz), (17.5–35 Hz) and (8.25–17.5 Hz) frequency ranges respectively. S(4,1) with (4.5–8 Hz) frequency range (theta) also showed better discrimination rate compared to other detail coefficients. In a similar study [5] the authors represented that approximate entropy applied on the S(2,3) coefficients demonstrates a significant p-value.
Ultimately, the linear and non-linear features were combined and applied as one feature vector to the classifier. In both feature groups, classifier accuracies were significantly improved. Combination of the band powers and E-KNN classifier reached an average accuracy of 92.00%. With non-linear methods the combination of the wavelet entropies features and E-KNN classifier resulted in 94.28% accuracy. Lastly, a combination of all features provided 93.75% accuracy with linear SVM. Genetic algorithm was used in order to reduce dimensionality of the feature-vector. 14 features were selected by genetic algorithm and were used as input to the proposed E-KNN obtained improved classification results (accuracy = 98.44, specificity 100%, sensitivity = 97.10%). The highest weighted features selected from genetic algorithm were related to O1, O2, Pz, T6, Fp2 channels.
Furthermore, as it can be seen in Fig. 7, the proposed E-KNN increasingly improves classification result with both significant and non-significant features (according to p-value). Also, in E-KNN, it is observed that for features of low significance, the results are more distinct compared to those of the most significant features as opposed to normal KNN, which implies the success of the E-KNN method to increase the separability of the classes.
Table 8 represents a comparison between the current study and a number of related studies. According to the table, Putankattil et al. [3] achieved an accuracy of 98.11% using ANNs with the model consisting of a single hidden layer with 20 neurons. Despite impressive results, the study suffers from the limited dataset size and also relatively long-time signals for each participant. Later on, Hosseinifard et al. [4] acquired an accuracy of 90.00% with Logistic regression as their best method. Their study had a reasonable dataset size although recording durations were as long as 5 min. Faust et al. [5] achieved a high accuracy of 99.50% performing the Probabilistic neural networks classification method, with respect to their high accuracy and novel method this experiment was highly limited by sample size. Acharya et al. [6] also used the SVM classifier and acquired an accuracy of 98.00% on a relatively small dataset of 30 participants. Mumtaz et al. [7] experimented with Naive Bayes, Logistic regression and SVM on long-term EEG signals with a duration of 5 min. The study yielded an accuracy of 98.40% using the Naïve Bayes algorithm. Acharya et al. [9] reported an accuracy of 94.99% using one dimensional convolutional neural network on the same dataset with the exact duration from their previous studies. In comparison, our study entails the following advantages and limitations: the developed E-KNN, being superior to KNN is also able to give information on feature importance with assigned weights, which can reveal important channels for further brain studies. The downside to this approach is that, on higher dimensions the performance of E-KNN and GA may be time-consuming. Taking that into consideration, the E-KNN won’t be a great option to those problems and instead using back propagation is recommended.
Conclusion
In this work, we analyzed two thousand samples (7.8 s) of 64 subjects (30 normal and 34 depressed) by frequency power of EEG signals and wavelet packet decomposition entropies. Three well-known classifiers, support vector machine (SVM), a novel enhanced K-nearest neighbors (E-KNN) and multilayer perceptron (MLP) were used to differentiate between two groups. Based on classification results with EEG linear features, the highest accuracy was obtained from gamma band powers, which demonstrated that gamma oscillations in comparison to other bands such as delta, theta, alpha and beta can more efficiently distinguish between depressed patients and healthy controls. Also, the proposed E-KNN demonstrated superior results with respect to famous classification algorithms. Thus, the E-KNN as a generalization to the KNN can always boost results and give away a feature importance index. This experiment demonstrates that the combination of power and complexity of high frequency components based on short-term EEG signals can highly discriminate between depressed patients and healthy controls. Our findings suggest that five specific electrodes considerably asses the ability to classify depressed patients and healthy controls. Also, our classification results were comparable to other experiments which several channels with various non-linear features performed on long-term EEG signals.
In conclusion, we suggest that gamma band should be addressed more seriously in depression-oriented studies.
References
WHO (2012) Depression: a global crisis. World mental health day.
Hinrikus H, Suhhova A, Bachmann M, Aadamsoo K, Võhma Ü, Lass J, Tuulik V (2009) Electroencephalographic spectral asymmetry index for detection of depression. Med Biol Eng Comput 47(12):1291
Puthankattil SD, Joseph PK (2012) Classification of EEG signals in normal and depression conditions by ANN using RWE and signal entropy. J Mech Med Biol 12:1240019
Hosseinifard B, Moradi MH, Rostami R (2013) Classifying depression patients and normal subjects using machine learning techniques and nonlinear features from EEG signal. Comput Methods Programs Biomed 109(3):339–345
Faust O, Ang PCA, Puthankattil SD, Joseph PK (2014) Depression diagnosis support system based on EEG signal entropies. J Mech Med Biol 14(3):1450035
Acharya UR, Sudarshan V, Adeli H, Santhosh J, Koh J, Adeli A (2015) Computer-Aided diagnosis of depression using EEG signals. Eur Neurol 73(2015):329–336
Mumtaz W, Xia L, Mhod Yasin MA, Azhar Ali SS, Malik AS (2017) Electroencephalogram (EEG)-based computer-aided technique to diagnose major depressive disorder (MDD). Biomed Signal Process Control 31:108–115
Bachmann M, Lass J, Hinrikus H (2017) Single channel EEG analysis for detection of depression. Biomed Signal Process Control 31:391–397
Acharya UR, Oh SL, Hagiwara Y, Tan JH, Adeli H, Subha DP (2018) Automated EEG-based screening of depression using deep convolutional neuralnetwork. Comput Methods Programs Biomed 161:103–113
Fitzgerald PJ, Watson BO (2018) Gamma oscillations as a biomarker for major depression anemergingtopic. Transl Psychiatry 8(1):1–7
Azizi A, Moridani MK, Saeedi A (2019) A novel geometrical method for depression diagnosis based on EEG signals. In: 2019 IEEE 4th conference on technology in electrical and computer engineering
Gao Z, Wang X, Yang Y, Mu C, Cai Q, Dang W, Zuo S (2019) EEG-based spatio–temporal convolutional neural network for driver fatigue evaluation. IEEE Trans Neural Netw Learn Syst 30(9):2755–2763
Chai R, Naik GR, Nguyen TN, Ling SH, Tran Y, Craig A, Nguyen HT (2016) Driver fatigue classification with independent component by entropy rate bound minimization analysis in an EEG-based system. IEEE J Biomed Health Inform 21(3):715–724
Liu G, Zhang Y, Hu Z, Du X, Wu W, Xu C, Li S (2017) Complexity analysis of electroencephalogram dynamics in patients with Parkinson’s disease. Parkinson’s Dis. https://doi.org/10.1155/2017/8701061
Al-Ani A, Koprinska I, Naik G (2017) Dynamically identifying relevant EEG channels by utilizing their classification behaviour. Expert Syst Appl 83:273–282
Acharyya A, Jadhav PN, Bono V, Maharatna K, Naik GR (2018) Low-complexity hardware design methodology for reliable and automated removal of ocular and muscular artifact from EEG. Comput Methods Programs Biomed 158:123–133
Butkevičiūtė E, Bikulčienė L, Sidekerskienė T, Blažauskas T, Maskeliūnas R, Damaševičius R, Wei W (2019) Removal of movement artefact for mobile EEG analysis in sports exercises. IEEE Access 7:7206–7217
Nejedly P, Cimbalnik J, Klimes P, Plesinger F, Halamek J, Kremen V, Worrell G (2018) Intracerebral EEG artifact identification using convolutional neural networks. Neuroinformatics. https://doi.org/10.1007/s12021-018-9397-6
Bhardwaj S, Jadhav P, Adapa B, Acharyya A, Naik GR (2015) Online and automated reliable system design to remove blink and muscle artefact in EEG. In: 2015 37th annual international conference of the IEEE engineering in medicine and biology society (EMBC). IEEE, pp 6784–6787
Jadhav PN, Shanamugan D, Chourasia A, Ghole AR, Acharyya AA, Naik G (2014) Automated detection and correction of eye blink and muscular artefacts in EEG signal for analysis of autism spectrum disorder. In: 2014 36th annual international conference of the IEEE engineering in medicine and biology society. IEEE, pp 1881–1884
Kilicarslan A, Contreras-Vidal JL (2019) Characterization and real-time removal of motion artifacts from EEG signals. J Neural Eng 16(5):056027
Mumtaz W, Xia L, Mhod Yasin MA, Azhar Ali SS, Malik AS (2017) A wavelet-based technique to predict treatment outcome for major depressive disorder. PLoS ONE 12(2):e0171409
Islam MK, Rastegarnia A, Yang Z (2017) Methods for artifact detection and removal from scalp EEG: a review. Neurophysiol Clin 46(4–5):287–305
Mamun M, Al-Kati M, Marufuzzaman M (2013) Effectiveness of wavelet denoising of electroencephalogram signals. J Appl Res Technol 11(1):156–160
Welch PD (1967) The use of Fast Fourier Transform, for the estimation of power spectra: a method based on the over short, modified periodograms. IEEE Trans Audio Electroacoust 15(2):70–73
Zhang Y, Dong Z (2015) Preclinical diagnosis of magnetic resonance (MR) brain images via discrete wavelet packet transform with Tsallis entropy and generalized eigenvalue proximal support vector machine (GEPSVM). Entropy 17(4):1795–1813
Coifman RR, Wickerhauser MV (1992) Entropy-based Algorithms for best basis selection. IEEE Trans Inf Theory 38(2):713–718
Pincus SM (1991) Approximate entropy as a measure of system complexity. Proc Natl Acad Sci 88(6):2297–2301
Richman JS, Moorman JR (2000) Physiological time-series analysis using approximate and sample entropy. Am J Physiol Heart Circ Physiol 278(6):H2039–H2049
Costa M, Goldberger A, Peng C-K (2005) Multiscale entropy analysis of biological signals. Phys Rev E 71(2):021906
Eiben AE, Raue PE, Ruttkay Z (1994) Genetic algorithms with multi-parent recombination. International conference on parallel problem solving from nature. Springer, Berlin, pp 78–87
Ting C-K (2005) On the mean convergence time of multi-parent genetic algorithms without selection. Advances in artificial life. Springer, Berlin, pp 403–412
Altman NS (1992) An introduction to kernel and nearest-neighbor nonparametric regression. Am Stat 46(3):175–185
Cortes C, Vapnik VN (1995) Support-vector networks. Mach Learn 20(3):273–229
Collobert R, Bengio S (2004) Links between Perceptrons, MLPs and SVMs. In: Proceedings of the international conference on machine learning (ICML)
Varma S, Simon R (2006) Bias in error estimation when using cross-validation for model selection. BMC Bioinform 7:91
Picard R, Cook D (1984) Cross-validation of regression models. J Am Stat Assoc 79(387):575–583
de Aguiar Neto F, Rosa J (2019) Depression biomarkers using non-invasive EEG: a review. Neurosci Biobehav Rev 105:83–93
Strelets VB, Garakh ZV, Novototskii-Vlasov VY (2007) Comparative study of the gamma rhythm in normal conditions, during examination stress, and in patients with first depressive episode. Neurosci Behav Physiol 37:387–394
Funding
The authors received no financial support for the research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
Approval was obtained by data owner from Human ethics committee of the Hospital university sains Malaysia(HUSM), Klentan, Malaysia.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Saeedi, M., Saeedi, A. & Maghsoudi, A. Major depressive disorder assessment via enhanced k-nearest neighbor method and EEG signals. Phys Eng Sci Med 43, 1007–1018 (2020). https://doi.org/10.1007/s13246-020-00897-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13246-020-00897-w