Abstract
Introduction
In native heart tissue, functions of cardiac fibroblasts (CFs) include synthesis, remodeling, and degradation of the extracellular matrix (ECM) as well as secreting factors that regulate cardiomyocyte (CM) function. The influence of direct co-culture and CF-derived ECM on CM mechanical function are not fully understood.
Methods
Here we use an engineered culture platform that provides control over ECM geometry and substrate stiffness to evaluate the influence of iPSC-CFs, and the ECM they produce, on the mechanical function of iPSC-CMs. Mechanical analysis was performed using digital image correlation to quantify maximum contractile strain, spontaneous contraction rate, and full-field organization of the contractions.
Results
When cultured alone, iPSC-CFs produce and remodel the ECM into fibers following the underlying 15° chevron patterned ECM. The substrates were decellularized and confirmed to have highly aligned fibers that covered a large fraction of the pattern area before reseeding with iPSC-CMs, alone or in co-culture with iPSC-CFs. When seeded on decellularized ECM, larger maximum contractile strains were observed in the co-culture condition compared to the CM Only condition. No significant difference was found in contractile strain between the Matrigel and decellularized ECM conditions; however, the spontaneous contraction rate was lower in the decellularized ECM condition. A methodology for quantifying alignment of cell contraction across the entire field of view was developed based on trajectories approximating the cell displacements during contraction. Trajectory alignment was unaltered by changes in culture or ECM conditions.
Conclusions
These combined observations highlight the important role CFs play in vivo and the need for models that enable a quantitative approach to examine interactions between the CFs and CMs, as well as the interactions of these cells with the ECM.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
While cardiomyocytes (CMs) are the cell type primarily responsible for contraction in the heart, cardiac fibroblasts (CFs) are also prevalent and play an important role in maintaining cardiac function [1,2,3]. Important functions of CFs include maintenance of the extracellular matrix (ECM), cell-cell communication with CMs and other fibroblasts, and secretion of growth factors and cytokines [4]. The bidirectional communication between CFs and CMs occurs through formation of gap junctions or membrane nanotubes, via mechanical forces, and paracrine signaling [5, 6]. CFs are both sources and targets of environmental stimuli and can respond to various types of stimuli, including electrical, mechanical, and chemical. Their response may be cellular, including proliferation, migration, or differentiation, or molecular by producing ECM or small molecules such as cytokines, peptides, and enzymes [3, 7].
One of the primary functions of CFs is synthesis, remodeling, and degradation of the ECM, which surrounds and interconnects cells and allows for signaling and transmission of mechanical forces throughout the myocardium. The ECM helps to distribute mechanical forces throughout the myocardium and convey mechanical signals to cells via cell surface ECM receptors [4]. CFs produce collagen and fibronectin which are crucial for maintaining cardiac morphology and function [7]. The production and maintenance of ECM proteins are essential for maintaining structural, mechanical, and electrical functions of the myocardium [8].
CFs also influence the functional properties of CMs when in co-culture; for example, CFs increase the amplitude of force generated by CMs [9]. A higher spontaneous beat rate of induced pluripotent stem cell-derived CMs (iPSC-CMs) when in co-culture with CFs has also been reported [10, 11]. Our previous work has also shown CFs improve calcium kinetics and increase contractile strain when in co-culture with CMs [12]. CFs have been shown to promote the formation and contractile function of engineered cardiac tissues fabricated using polymer scaffolds [13, 14], hydrogels [15, 16] and detachable cell sheets [17]. These findings highlight the importance of developing in vitro models with multiple cell types that allow for controlled myocyte-fibroblast-ECM interactions to better understand myocyte-fibroblast communication and its effect on CM contractile function.
Varying the properties of the ECM can exert differential effects on CMs and CFs. For example, CF fate can be regulated by the stiffness of the ECM [18], with stiffer substrates promoting CF activation to myofibroblasts [5]. CFs respond to substrate surface cues as well; for example migration can be influenced by topography at both the nano and micro scales [19]. Modification of substrate properties can influence the deposition of ECM such as promoting aligned fibers by culturing CFs on a polydimethylsiloxane (PDMS) substrate with nanogrooves [20] or microcontact printed lanes [12]. These anisotropic scaffolds can be decellularized and used to culture other cell types [20]. CFs have been shown to promote the maturation of iPSC-CMs in coculture [10, 21].
2D culture studies are beneficial due to simplicity, convenience, amenability to a wide variety of analytical techniques, and higher throughput capability [22] . While 2D systems do have limitations, use of these models to precede and/or support more complex three-dimensional (3D) cultures, animal experiments, and clinical investigation may provide additional valuable information [22]. Platforms that include cues such as physiologically-relevant substrate stiffness [23, 24] , electrical and mechanical conditioning [25,26,27] and co-culture [28, 29] or combinations of multiple signaling factors [26, 30, 31] have demonstrated improvement in iPSC-CM function. Standard in vitro cardiac cell culture models do not reproduce the in vivo structural organization of cardiac fibroblasts or their interrelation with myocytes [1].
In the present study, we aimed to develop an engineered culture platform that better recapitulated the native myocardium and allowed for analysis of the interaction of iPSC-CMs with iPSC-CFs. We aimed to determine if iPSC-CF produced ECM alone was sufficient to enhance the mechanical function of iPSC-CMs and could mimic the advantages of co-culture systems. A previously developed 2D cell culture platform [32] was used to control intracellular features to produce organized and connected CMs. While standard in vitro cell culture models do not recapitulate in vivo structural organization, spatially restricted ECM patterns can be used to replicate important features of cardiac tissue-architecture [1]. Microcontact printing was used to pattern ECM proteins onto 10 kPa PDMS substrates, the stiffness of which agrees with the Young’s modulus of healthy heart muscle as determined by nanoindentation [22]. We have previously shown that the pattern influences CM shape and sarcomere organization by providing defined attachment regions, while the bridges increased connectivity to allow for synchronized contractions across large arrays of CMs [12, 33]. The pattern also influenced how CFs secreted and remodeled the ECM, which in turn modulated the stiffness, alignment, and topography of the secreted matrix on which CMs were to be seeded. CFs responded to the signal of narrow lane widths and remodeled and produced ECM aligned primarily in the longitudinal direction of the pattern [12].
CFs contribute to structural, biochemical, mechanical and electrical properties of the myocardium [1]. This work builds on a prior study using iPSC-CFs cultured on micropatterned PDMS. When co-cultured, iPSC-CMs maintained their alignment and ECM was produced by iPSC-CFs. Optical mapping analysis of electrical activity demonstrated that anisotropic impulse propagation was preserved in the co-culture conditions as seen previously within cultures containing only iPSC-CMs [33]. The contractile strains were higher in the co-culture models compared to iPSC-CMs alone [12]. Building on our prior work [12, 32, 33] we utilized the 2D micropatterned platform to allow iPSC-CFs to remodel and produce aligned ECM. The ECM organization was quantified in terms of alignment of fibers and a percent of area covered with ECM using a previously developed software package called Scanning Gradient Fourier Transform (SGFT) [34]. The substrates were then decellularized and reseeded with iPSC-CMs alone or in co-culture with iPSC-CFs. The focus of the study was mechanical function; mechanical analysis was performed using digital image correlation (DIC) followed by spatial and temporal quantification of contractile displacements and strains. An overview of the workflow is shown in Fig. 1.The maximum contractile strain, spontaneous contraction rate, and full-field organization of the contractions were quantified to evaluate the effect of ECM and co-culture on the contractile function of iPSC-CMs. Together, this platform and these analysis techniques enable a quantitative approach to examine interactions between the fibroblasts and myocytes, as well as the interactions of these cells with the ECM.
Schematic of workflow. A Cells are cultured on either micropatterned Matrigel, or substrates of micropatterned Matrigel that have been remodeled by cardiac fibroblasts and decellularized, to provide a more physiologic matrix environment. Not to scale. B Human iPSCs are differentiated into cardiomyocytes (CMs) and cardiac fibroblasts (CFs). These cells are cultured alone or in combination for the experimental conditions on Matrigel or decellularized ECM conditions in monolayers or patterns. Mechanical characterization is performed using digital image correlation to quantify function.
Methods
iPSC Cardiomyocyte Differentiation
A human induced pluripotent stem cell (iPSC) line iPS-DF19-9-11T (WiCell) was used for both CM and CF differentiation. iPSC-CMs were differentiated using a modified version of the small molecule GiWi method [35]. Briefly, hiPSCs were seeded on Matrigel (Corning) coated plates (8.7 μg/cm2) in mTesR1 medium supplemented with 10 μM ROCK inhibitor (Y-27632) (Tocris) until 100% confluent (5 days). On day 0 the medium was changed to RPMI 1640 (Thermo Fisher Scientific) plus B27 minus insulin supplement (Thermo Fisher Scientific) and 12 μM of the GSK3 inhibitor CHIR 99021 (Biogems). On day 1, 24 h after CHIR was added, the medium was changed to RMPI 1640 with B27 minus insulin supplement. On day 3, 72 h from the time CHIR was added, cells were treated with RPMI 1640 supplemented with B27-insulin and 5 μM of Wnt inhibitor IWP2 (StemGent). On day 5 and day 7, the medium was changed to RPMI 1640 with B27-insulin. On day 9, the medium was changed to RPMI 1640 plus B27 complete supplement, and the medium changed every other day until day 15 of differentiation. Contractions began on day 10–13.
iPSC-CMs were cryopreserved on day 15 of differentiation in 90% Fetal Bovine Serum (FBS, Invitrogen) plus 10% DMSO (Sigma). Cryopreserved day 15 iPSC-CMs were thawed onto Matrigel-coated 12 well plates at a density of 2.5 million cells/well in EB20 medium: DMEM/F12 (Life Technologies), 20% FBS (Life Technologies), NEAA (Life Technologies), GlutaMax (Life Technologies) and 2-Mercaptoethanol (Sigma). For cells to be used without freezing, cells were replated into 12 well plates with EB20 media on day 15. Cells were dissociated with 10x TrypLE™ (Thermo Fisher Scientific) and resuspended in EB20 medium. On day 17 of culture, cells were purified using lactate medium consisting of RPMI without glucose (Life Technologies), B27 complete supplement and 5 mM lactate (Sigma) [36]. Cells were fed every other day for 10 days, and then changed to EB2 medium: DMEM/F12 (Life Technologies), 2% FBS (Life Technologies), 1% NEAA (Thermo Fisher Scientific), 0.5% GlutaMax (Life Technologies) and 0.007% 2-Mercaptoethanol (Sigma), which was replaced every three days thereafter. iPSC-CM purities of greater than 95% after lactate purification were confirmed via flow cytometry [33].
iPSC Cardiac Fibroblast Differentiation
iPSC-CFs were differentiated as described previously [37]. Briefly, iPSCs were dissociated with 1 ml/well Versene solution (Invitrogen) at 37 °C for 5 min, and seeded on Matrigel (GFR; BD Biosciences) coated 6-well plates at the density of 2 × 106 cells/well in mTeSR1 medium supplemented with 10 μM ROCK inhibitor (Y-27632) (Tocris). Cells were cultured for 5 days in mTeSR1 medium with medium changed daily until 100% confluence was reached. On the first day of differentiation, day 0, the medium was changed to 2.5 ml RPMI supplemented with B27 minus insulin and 12 µM CHIR99021 (Tocris). At day 1, 24 h later, medium was changed to 2.5 ml RPMI plus B27 minus insulin (Life Technologies). At day 2, the medium was changed to 2.5 ml of the defined fibroblast culture medium (CFBM) [37] supplemented with 75 ng/ml basic Fibroblast Growth Factor (bFGF; WiCell). Cells were fed every other day with CFBM supplemented with 75 ng/ml bFGF and cultured until day 20 for flow cytometry analysis and subculture of a high purity population of cardiac fibroblasts [37]. iPSC-CFs were maintained in FibroGro plus 2% FBS medium (Millipore EDM) on tissue culture plastic. The cryopreserved iPSC-CFs were thawed in FibroGro plus 2% FBS and plated onto 6-well tissue culture plastic plates at a density of 55,100 cells/well in 2 ml of FibroGRO medium (Millipore EMD) plus 2% FBS (Gibco). iPSC-CFs were maintained in FibroGRO plus 2% FBS medium which was replaced every 2 days. iPSC-CFs were passaged at least once after thawing before use in experiments.
Patterning
Microcontact printing and soft lithography was used to generate compliant substrates with patterned extracellular matrix proteins in defined geometries [12, 33]. For experimental samples, PDMS substrates were made with a Young’s modulus of 10 kPa by blending Sylgard 184 and Sylgard 527 (Dow Corning) at a ratio of 52:1 Sylgard 527 to Sylgard 184 [38]. Sylgard 184 was made by mixing ten parts base to one part curing agent. Sylgard 527 was made by mixing equals part of components A and B. Both were mixed individually for 5 min with a glass stir rod and combined in the prescribed ratio to achieve the desired stiffness. Once the components were mixed, PDMS was poured into a petri dish, placed under vacuum for 20 min to remove air bubbles, and cured for 12 h at 60 °C. Mechanical characterization of 10 kPa PDMS prepared following the described protocol was previously performed to confirm the targeted stiffness was obtained [39]. After curing, the samples were cut to the desired size with a razor blade and attached to a 12-well plate using Sylgard 184 and allowed to cure for 12 h at 60 °C. Substrates were UV sterilized prior to use.
Patterns were fabricated on a master Si wafer (FlowJEM, Toronto, ON, Canada) and used to produce reusable PDMS (Sylgard 184) stamps for microcontact printing. PDMS was poured onto the patterned wafer, allowed to sit under vacuum for 20 min to remove air bubbles, cured at 60 °C for 6 h, then removed and cut into individual stamps. Stamps were coated with 83 μg/ml Matrigel at 37 °C and allowed to sit overnight. The following day the ECM was removed, and excess ECM and moisture removed with nitrogen airstream. A polyvinyl alcohol (PVA) film was made by dissolving 0.5 g of PVA (Sigma) in 10 ml of deionized water and dried overnight in a petri dish. The PVA film was removed from the petri dish and cut to size before the ECM-coated stamps were brought into contact. The stamps were placed in contact with the PVA film and kept in an incubator with a glass slide and 50 g weight placed on top to promote complete transfer of ECM. After one hour, the PVA film was removed from the PDMS stamp and then brought into conformal contact with the sterile 10 kPa PDMS culture substrate. After 20 min the substrate was washed 3 times with PBS to dissolve the PVA film, leaving behind the patterned proteins.
Seeding onto PDMS Substrates
Cardiac fibroblasts were all freshly passaged before use. No analysis was done to ensure that frozen versus fresh cardiomyocyte behavior was the same, however no differences were observed in results, and all conditions used both fresh and frozen cardiomyocytes. On day 30, purified iPSC-CMs were dissociated and singularized with TyrpLE 10x (Life Technologies) for 10–12 min and centrifuged for 5 min at 300 rpm. CMs were resuspended in EB20 medium in the absence of ROCK inhibitor and seeded onto patterned substrates, referred to as day 0 for the experimental time point. The patterned substrates were seeded at a density of 2528 CMs/mm2 (90,000CMs/35.605mm2). iPSC-CFs were treated with TrypLE Express (Life Technologies) for 3 min, centrifuged for 5 min at 300 rpm, and seeded onto patterned PDMS using EB20 medium. For both iPSC-CF alone and co-culture with CMs (CM-CF), a seeding density of 253 iPSC-CFs/mm2 (9000 CMs/35.605 mm2) was used. To control cell attachment, a 1 cm PVC tube was placed on top of the PDMS substrates and the cell suspension was added inside the tube. The PVC tube was removed the following day and additional EB20 medium was added. 48 h after seeding, the medium was changed to EB2 medium and exchanged every 2 days during the length of culture.
Decellularization
To remove iPSC-CFs for ECM visualization or reseeding, PDMS substrates were decellularized using a previously published protocol [20]. Briefly, samples were washed with PBS prior to the addition of the first decellularization solution (1 M NaCl, 10 mM Tris, and 5 mM EDTA; Sigma) and then placed on an orbital shaker for 1 h at room temperature. The samples were again washed with PBS and placed in a second decellularization solution (0.5% SDS, 10 mM Tris, and 25 mM EDTA; Sigma) and shaken for 15 min at room temperature on an orbital shaker. After a PBS wash, the samples were rinsed twice, for 24 h each, in DMEM medium with 20% FBS at room temperature. Samples were rinsed with PBS and dried with a nitrogen airstream prior to reseeding.
Immunofluorescence
Samples were washed once with PBS and fixed in 4% paraformaldehyde (PFA; Electron Microscopy Sciences) for 15 min at room temperature and then washed with PBS. Samples were then washed with PBS and blocking was performed with 5% nonfat dry milk (Bio-Rad) in 0.2% Triton X-100 solution and incubated for 1.5 h at room temperature on a rotator followed by two washes with PBS [37]. Primary antibodies were diluted in 1% BSA (Sigma) in PBS with 0.1% Triton X-100 and incubated overnight at 4 °C. The following day the samples were washed twice with 0.2% Tween 20 (Sigma) in PBS and then the secondary antibodies were applied in 1% BSA (Sigma) in PBS with 0.1% Triton X-100 for 1 hour. After application of the secondary antibodies, the samples were washed with 0.2% Tween 20 in PBS and then in PBS and transferred to coverslips, where they were mounted using ProLong Gold Antifade (Life Technologies). Primary antibodies used include Collagen 1a (1:250 dilution, Santa Cruz), Laminin (1:250, Sigma) and Fibronectin (1:250, Santa Cruz). Secondary antibodies include Alexa Fluor 488 (IgG1) goat anti-mouse, 488 (IgG1) goat anti-rabbit, Alexa Flour 640 (IgG1) goat anti-mouse and Alexa Fluor 647 (IgG2b) goat anti-mouse (Thermo Fisher Scientific).
FN Image Alignment/Percent Area Coverage
Decellularized samples labeled with fibronectin antibodies were imaged using a Nikon A1RSi Confocal Microscope with an attached Photometrics CoolSNAP HQ2 camera. To capture representative images of the entire pattern, images of areas containing only lanes as well as areas containing lane and bridge regions were collected for all samples. After image acquisition of fibronectin, the scanning gradient Fourier transform (SGFT) open source MATLAB software was used to provide unbiased quantification of ECM organization [34]. High-resolution images captured with a 60x oil-immersion lens were used for analysis. The following parameters were used as SGFT inputs on images with a resolution of 4096x4096 pixels: 0.05 µm/pix, a pattern size of 1.0 and a scan resolution of 16. The pattern size was used due to the relatively low spacing between fibronectin fibers and diameter of the fibers as compared to sarcomeres in higher magnification images, for which this software was primarily developed to analyze. To determine the area occupied by fibronectin fibers, the threshold for fiber identification within sft_main_v026.m (Line 687) was increased from 0.1 to 0.5. This value was determined by comparing the relative intensities of the fluorescent signals identified by the software in images containing fibronectin compared to previously analyzed images stained for the sarcomere protein α-actinin, for which the SGFT analysis was originally developed. Comparison between fibronectin and α-actinin images was performed only to determine an accurate threshold value, and no direct comparison occurs in the results of this work. Intensity was not used as a measure of fibronectin deposition in this context, as the relative intensity between images was similar. Rather, area quantification of fibronectin fibers was performed using automated analysis through the SGFT software to determine the area of the region occupied by fibronectin. The fluorescent threshold and automated area quantifications were further validated by counting the number of pixels within a patterned region for a manually binarized image compared to the number of pixels as measured by the SGFT software. The variability in the fibronectin quantification occurs due to intentional image capture of both bridge and lane regions of each decellularized substrate, which cannot be captured in a single image at the resolution required and have heterogeneity in their geometry at high magnification. These regions were sought to provide a representative analysis of the full pattern.
Contractile Strain Analysis
Phase contrast videos of cells spontaneously contracting were acquired on experiment days 6, 12, and 18 using a Nikon Eclipse Ti microscope with a Plan Flour 10x NA 0.3 objective and Nikon DS-QiMc camera with samples maintained at 37 °C. At least two spontaneous contractions or a maximum of 400 frames were acquired for each sample with an acquisition rate of 18.9 fps. Mechanical analysis was done using previously developed open source software, with custom add-ons and modifications [12, 33, 40]. The random high contrast pattern from the phase contrast of the cells is sufficient to allow tracking of displacements using the open-source digital image correlation software, Fast Iterative Digital Image Correlation (FIDIC) [41]. Subsequent frames were analyzed relative to the first frame which contained cardiomyocytes in a relaxed state. Full field 2D displacements, Ux and Uy, were computed for each frame using the input parameters of a target subset size of 48 pixels (31.2 μm) and subset spacing of 12 pixels (7.8 μm). From the displacements, x and y strains were calculated by taking the gradient of the displacement data, and ultimately principal strains, ε1 and ε2, were computed for each time point throughout the contraction event. A binary mask was created using edge detection techniques to eliminate data from areas not occupied by cells [42]. Second principal strain (i.e. contractile strain) was used as the metric of mechanical function. Full field contractile strain values were averaged for each frame of the video and the maximum of this averaged value (i.e. maximum contractile strain) was used as the metric for comparison between sample conditions.
Displacement Trajectory Analysis
The trajectories of displacements for videos of contracting cells were computed to quantify the organization of contractions. Each video analyzed contained at least two spontaneous contractions. Analysis was conducted using previously developed software with custom modifications for additional processing: https://github.com/jknotbohm/Cell-Velocity-Analysis/compute_cell_trajectories.m. The full field 2D displacements, Ux and Uy, computed using FIDIC were taken as inputs. A down sampling factor of 2 was used to create the initial positions of each trajectory, and the displacements in x and y directions added frame by frame across all locations. The total displacements for each location at each time point were then used to compute the angle and magnitude of the resulting vector. Trajectories with small magnitudes, likely occurring in regions of the pattern where no cells were present, were filtered by eliminating values below a threshold of 0.75 of the median trajectory magnitude for that sample. To quantify directionality, the four-quadrant inverse tangent was used to compute angles by using the start and end point of each trajectory. To evaluate overall organization from 0 to 180°, 180° was added to negative angles. The histogram of all trajectory angles was plotted using 20 bins, and the center of the bin with the highest count was taken as the primary angle. The angle of each trajectory was evaluated, and those within ± 20° of the primary angle were counted and the total percent of aligned trajectories calculated. Analysis was done for each video, and the percent alignment was used to assess multiple samples per condition.
Statistical Analysis
For brightfield microscopy, the reported sample size refers to biological replicates, rather than repeated acquisition at one timepoint within an individual micropatterned culture. Alternatively stated, a sample denotes one location on a micropattern at a given experimental timepoint. Only one location is used per micropattern, and there is one micropatterned sample per well. For fluorescent microscopy, the number of locations imaged per micropattern sample is noted in the respective results section. Sample sizes and p values are reported in each figure legend and statistical analyses were performed using MATLAB. For statistical analysis of two groups, an unpaired two sample t-test was used. For analysis with more than two conditions a one-way ANOVA test was performed followed by multiple comparison test to perform pairwise comparisons between pairs of data using Tukey’s honestly significant difference criterion. Distributions were normal prior to statistical testing as determined by a Kolmogorov–Smirnov test for each data set (p < .05). Statistical significance was defined as *p < .05, **p < .01, ***p < 0.001.
Results
Analysis of Decellularized ECM
We have previously demonstrated that iPSC-CFs remodel and deposit aligned ECM fibers, but these substrates were not used for culturing iPSC-CMs [12]. The optimal timepoint for decellularization of iPSC-CF modified ECM was determined by seeding iPSC-CFs on Matrigel in the chevron pattern and allowing them to produce and remodel ECM for 11, 14, or 18 days (Fig. 2A). The chevron pattern, consisting of 30 µm lanes intersected at a 15° angle, was used to provide connectivity across the pattern. This ECM geometry has been used to generate of iPSC-CMs that mimic the structure‐function relationship of native tissue in terms of morphology, electrical, and mechanical function [33]. The timepoints were selected based on prior studies where 2 days was not sufficient to produce continuous, aligned ECM fibers, whereas patterns appeared filled with aligned ECM fibers after 18 days [12].
A 10 kPa PDMS substrates patterned with Matrigel using the 15° chevron pattern and decellularized after 11, 14 and 18 days of culture with iPSC-CFs. Scale bar = 50 µm. B Quantification of the amount of patterned area covered by fibronectin fibers. C Quantification of the amount of fibronectin fibers aligned within 15° of the superior angle using SGFT software. No significant difference in area coverage or fiber alignment between timepoints. White = Fibronectin. N = 15 Day 11, N = 11 Day 14 and 18.
After decellularization, samples were fixed and stained for fibronectin (Fig. 2A), and a custom open source MATLAB SGFT software [34] was used to quantify the orientation and percent alignment of ECM fibers. Images of areas containing bridge regions and narrow lanes were evaluated for all timepoints. The percentage of area of pattern covered by fibers, taken as the area of image containing fibers over the area of the image containing patterned regions, was not significantly different between the 3 timepoints (Fig. 2B). The alignment of fibronectin fibers, measured as the percentage of fibers aligned within 15° of the primary angle, was also not significantly different between the 3 timepoints (Fig. 2C). Decellularized substrates were observed to have deposited Collagen Type I, as well as an increased thickness on days 11, 14, and 18 in comparison to day 0 Matrigel only (Supplemental Fig. 1). However, no statistical differences existed between the day 11, 14, and 18 timepoints. Given these results a decellularization timepoint of day 14 was chosen for convenience of experimental design.
Contractile Strain on Matrigel and Decellularized ECM
Following decellularization, iPSC-CMs were seeded alone or in co-culture (CM-CF) with iPSC-CFs on experimental day 0 (E0) and cultured for 18 days (E18). A CM:CF ratio of 10:1 was previously determined as an optimal ratio to allow for maintenance of iPSC-CM and iPSC-CF cellular attachment with some pattern augmentation [12]. Bright field videos of spontaneously contracting iPSC-CMs were captured at days 6, 12, and 18 and analyzed with DIC. Displacements in the x and y directions were first computed using DIC. The contraction primarily occurs in the y direction, which is the orientation of the major axis of the chevron pattern, and therefore the general alignment of the CMs. Although the major axis of the pattern aligns with the y axis, the bridge regions that are 15° offset from vertical contain cells whose contractions are not primarily in the y direction. To further quantify contraction and better capture the behavior of the cells in this orientation, the strain tensor was computed as described in the Methods section. All cardiomyocytes contracting in the field of view were analyzed and the second principal strain ε2, i.e., contractile strain, was used as the quantitative measure of mechanical output (Supplemental Fig. 2). Mechanical function was evaluated at days 6, 12, and 18 . The overall chevron pattern was maintained, however more regions appeared to be merged thus obscuring the original pattern in the CM-CF condition (Fig. 3A). While iPSC-CMs are not proliferating, iPSC-CFs continue to proliferate as well as migrate in culture to fill the spaces previously absent of ECM. As a result, progressive pattern augmentation and a transition from patterned to monolayer appearance occurs in CM-CF cultures with resident fibroblasts, but is not observed in CM Only cultures. On days 12 and 18, the maximum strain generated in the co-culture conditions was significantly higher compared to the CM Only condition (*p < .05, **p < .01; Fig. 3B).
A Bright field images of patterned iPSC-CMs and iPSC-CFs on Day 6 (left), Day 12 (middle), and Day 18 (right) of culture. Bright field locations are the same in the CM Only and CM-CF condition, respectively, so that the same area of the pattern can be tracked over time. Scale bar = 100 µm. B The maximum contractile strain for the two culture conditions on decellularized ECM on Day 6, 12 and 18 of culture. Maximum contractile strain is significantly higher in CM-CF condition on days 12 and 18. *p < .05, **p < .01, two-way ANOVA with post hoc Tukey tests. CM-CF: N = 22 Day 6, N = 19 Day 12 and Day 18. CM Only: N = 22 Day 6, N = 19 Day 12, N = 16 Day 18. C The maximum contractile strain for the two culture conditions on Matrigel on Day 6, 12 and 18 of culture. Maximum contractile strain is significantly higher in CM-CF condition at all timepoints. ***p < .001, two-way ANOVA with post hoc Tukey tests. N = 31 CM-CF day 6, N = 25 CM Day 6 and both conditions Day 12, N = 18 Day 18. D The maximum contractile strain for all timepoints in both ECM and culture conditions shown in B and C. Maximum contractile strain is significantly higher in CM-CF conditions, but there is no significant difference between ECM conditions. N = 68 CM Only Matrigel, N = 57 CM Only Decellularized, N = 74 CM-CF Matrigel, N = 60 CM-CF Decellularized.
In order to validate previously observed contractile behavior [12], iPSC-CMs were seeded alone or in co-culture with iPSC-CFs on PDMS substrates coated with patterned Matrigel. At each time point, there was a significant difference between the maximum strain generated in the co-culture conditions compared to the CM Only condition (***p < .001; Fig. 3C). Over the three time points tested there was no significant difference in strain within each of the two conditions. The contractile strain for both ECM conditions, Matrigel and decellularized iPSC-CF modified, and both culture conditions, CM Only and CM-CF, were compared. To compare ECM conditions, all timepoints (6, 12, and 18) are combined into one dataset per culture condition. The maximum contractile strain was significantly higher in the co-culture condition on both ECM conditions (***p < .001; Fig. 3D). There was no significant difference between the ECM conditions for either culture condition.
To understand other features of contractile behavior, the spontaneous contraction rate was evaluated. Only videos collected on Day 6 were evaluated as not all samples at later timepoints captured greater than one contraction. The contraction rate was determined for each sample by calculating the average time between contraction peaks and converting to beats per minute (BPM). The rate of contraction was significantly faster in the CM-CF condition compared to CM Only on both ECM, and in the CM-CF condition on Matrigel compared to decellularized ECM (Fig. 4, p < .001).
Spontaneous contraction rate for Day 6 samples in both Chevron patterned Matrigel and decellularized ECM and culture conditions. The contraction rate is significantly higher in CM-CF compared to CM Only condition. The contraction rate is significantly lower on decellularized ECM than Matrigel in CM-CF condition. ***p < .001, two-way ANOVA with post hoc Tukey tests. N = 24 CM Matrigel, N = 31 CM-CF Matrigel, N = 23 CM Decellularized, and N = 25 CM-CF Decellularized.
Displacement Trajectory Organization
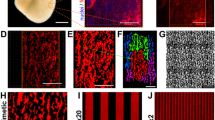
iPSC-CFs migrate and continue to proliferate in co-culture and fill the spaces absent of ECM, meaning the patterns appear more similar to monolayers under brightfield microscopy. We previously observed that the ECM deposited and remodeled by iPSC-CFs in co-culture was well aligned and generally followed the underlying chevron pattern [12], but the spatial organization of the contractions was not quantified in the prior research. To demonstrate the methodology for quantification, three initial ECM deposition conditions were used: 15° chevron, 30 μm lanes, and monolayer (illustrated in Fig. 5). It has been previously observed that the structural organization and electrophysiology of 15° chevron pattern is significantly improved compared to a monolayer [33]. In terms of mechanical function, iPSC-CMs in chevron and monolayers cultures produce similar maximum contractile strain, and both conditions produce significantly higher strains than the narrow 30 μm lane pattern (Fig. 5).
A Representative brightfield images of 15° chevron, 30 μm lane, and monolayer patterns. Scale bar = 100 um. B The maximum contractile strain for the 3 pattern conditions (chevron = red, lanes = blue, monolayer = green). Maximum contractile strain is significantly lower in lanes pattern than chevron and monolayer across timepoints (days 3−15) ***p < .001, two-way ANOVA with post hoc Tukey tests. N = 58 chevron, N = 59 lanes, N = 49 monolayer.
The lane and chevron patterns produce more organized iPSC-CMs with highly aligned sarcomeres, and visually the contractions appear spatially organized. To quantify this alignment, the displacement trajectories were computed across the entire field of view, and the predominant angles of the trajectories determined, as described in Methods. The percentage of trajectories within 20° of the predominant angle was computed to determine alignment (example workflow for 3 pattern conditions shown in Supplemental Fig. 3). Comparing the displacement trajectory alignment of the 3 pattern conditions, the alignment was statistically highest in the 15° chevron pattern, with alignment being lowest in the monolayer condition (Fig. 6). Retrospective analysis of samples having varying lane width between 20 μm and 140 μm showed higher contractile spatial alignment for lanes of 60 – 100 μm (Supplemental Fig. 5A).
Quantification of displacement trajectory alignment. Percent trajectory alignment for two different pattern types (Chevron and Lanes) and Monolayer conditions for combined days 3–15 in culture. Alignment is significantly higher in chevron pattern than lanes and monolayers. (Chevron = red, lanes = blue, monolayer = green). *p < .05, ***p < .001, two-way ANOVA with post hoc Tukey tests. N = 58 chevron, N = 59 lanes, N = 49 monolayer.
Once the trajectory alignment method was validated on the test patterns, the alignment was quantified for both CM Only and CM-CF culture conditions on both patterned ECM conditions. Despite the CM-CF samples appearing more like a monolayer (shown in Fig. 3A), no significant difference was found in the alignment between the culture or ECM conditions investigated across either all timepoints (Fig. 7) or individual days 6, 12, and 18 (Supplemental Fig. 4).
Percent of displacement trajectories with 20° alignment for all timepoints in both Matrigel and decellularized ECM and culture conditions on chevron patterned substrates (CM Only = black, CM-CF = red). Data from Day 6, 12 and 18 combined. No significant difference in overall alignment. N = 68 CM Matrigel, N = 57 CM Decellularized, N = 74 CM-CF Matrigel, N = 60 CM-CF Decellularized.
Discussion
While CMs are contractile cell type of the heart, CFs play an important role in maintaining proper function [5]. They contribute to structural, biochemical, mechanical and electrical properties of the myocardium [3]. One of the primary functions of CFs is the physiological turnover of the ECM, where the structural and biological condition of the ECM also impact cellular structure and function [3]. The goal of this research was to develop and characterize an engineered substrate with ECM produced by iPSC-CFs and evaluate the influence of this ECM on the mechanical function of iPSC-CMs. In this work we have analyzed the structural and mechanical influence of combined co-culture of iPSC-CMs with iPSC-CFs on a soft, micropatterned substrate with cell-derived ECM. The stiffness of healthy myocardium ranges from 10 to 30 kPa; in these experiments a compliant PDMS substrate with a Young’s modulus of 10 kPa was used. This addresses an issue with common tissue culture plastics that have a Young’s modulus of around 3 GPa. When cultured on substrates with a modulus on the order of GPa, CFs undergo activation to myofibroblasts [5]. With the use of microcontact printing, defined environmental cues can be used to control the alignment of ECM that is remodeled and produced by iPSC-CFs. The underlying 15° chevron pattern influenced iPSC-CF behavior such that ECM was produced and remodeled into fibers that primarily followed the lanes of the pattern.
iPSC-CFs also have been shown to increase the amount of force generated by CMs in comparison to when CMs are cultured alone [9, 43]. Our previous findings [12] of increased contractile strain in co-culture conditions compared to iPSC-CMs alone was replicated, confirming that co-culture with iPSC-CFs leads to an improvement in iPSC-CM contractility. Other groups have explored combinations of CMs and CFs [28, 29, 44]. We previously found a 10:1 iPSC-CM to iPSC-CF ratio to be optimal to allow for maintenance of iPSC-CM and iPSC-CF cellular attachment with some pattern augmentation [12]. This CM:CF ratio also matches ratios used in 3D collagen gel compaction studies [25]. Having two separate iPSC-CM and iPSC-CF cell populations allowed for combination in specific ratios, as well as consistently using newly differentiated iPSC-CMs. iPSC-CMs were seeded 30 days after they were differentiated, as reports suggest the responsiveness of iPSC-CMs to physical stimuli declines as differentiation progresses [26]. Other groups have also reported that low passage number after differentiation or fetal CFs enhance CM function and maturity better than adult CFs [6, 15, 45, 46].
We next sought to utilize the anisotropic ECM scaffolds modified by iPSC-CFs as a platform for iPSC-CM culture. First, the optimal timepoint for decellularization was identified, as defined by ECM having highly aligned fibers that covered a large fraction of the pattern area on the substrate. There was no statistical difference in these parameters between the timepoints explored, so the middle timepoint, day 14, was used. Variability in fiber alignment occurred in part due to some images capturing predominantly lanes while others captured bridge regions, or areas where lanes meet. This was done intentionally to obtain representative analyses of the entire pattern. When seeded on decellularized ECM, we report similar findings with the co-culture condition having larger contractile strains compared to the CM Only condition. No significant difference was found in contractile strain between the two ECM conditions; however, the spontaneous contraction rate on day 6 in culture was decreased on the decellularized ECM in the co-culture condition. Newly differentiated iPSC-CMs beat spontaneously in culture, but lose automaticity as they mature [47,48,49]. In future studies electrical pacing could be employed to study contractile kinetics, in addition to maximum contractile strain. Further analysis of structural makers including cell size, multinucleation, and myofibril density and alignment, as well as expression of key cardiac structural makers including β-myosin heavy chain and connexin-43 could be conducted to explore the degree of maturation [50]. Electrical pacing could also be used to control for the difference in spontaneous rate to understand the implications of electrophysiological changes driving the mechanics in the observed phenotype.
We previously observed that the ECM deposited and remodeled by iPSC-CFs in co-culture was well aligned and generally followed the underlying chevron pattern [12], but the spatial organization of the contractions had yet to be quantified. A methodology for quantifying contraction alignment was validated using three culture patterns: 15° chevron, 30 μm lanes, and monolayer. The myofibril structural organization and electrophysiology of the chevron pattern was previously observed to be significantly improved compared to a monolayer [33]. To quantify contraction alignment, the displacement trajectories were computed across the entire field of view, and the percentage of trajectories with angles within 20° of the predominant angle was computed. The displacement trajectory alignment was highest in the 15° chevron pattern, with alignment being lowest in the monolayer condition. Interestingly, the alignment was higher in the chevron pattern than the lane pattern, even though iPSC-CMs were spatially confined to be highly aligned in the lane pattern. We hypothesize this was due to the increased number of nearest neighbor cells in the chevron pattern relative to the narrow lane pattern and was supported by analysis of trajectory alignment within lanes of different width ( Supplemental Fig 5C). Whereas lanes wider than 100 μm have an increased number of neighboring cells, the disorganization can lead to diminishing returns in trajectory alignment (Supplemental Fig 5A). The 15° chevron pattern incorporates narrow feature widths and an increased number of nearest neighbors, allowing for the highest degree of contraction alignment observed. Trajectory alignment was not significantly different between culture or ECM conditions. This suggests that although co-culture and decellularized ECM samples superficially appear more like a monolayer over time (as observed with brightfield imaging), the underlying chevron pattern to which the iPSC-CMs initially adhere was sufficient to retain organization. This is further supported by the finding that sarcomere alignment was not statistically significantly different in the co-culture condition compared to CM Only on decellularized ECM (Supplemental Fig. 6). We previously demonstrated, however, that myocytes cultured on a non-patterned surface in a monolayer have decreased sarcomere alignment in comparison to patterned lanes, or to the values identified for CM Only and co-culture conditions in this study [34].
The influence of direct co-culture and cell-modified ECM on iPSC-CM mechanical function were explored, but iPSC-CFs also generate paracrine signals that may influence iPSC-CM behavior.
The influence of paracrine signaling of CFs on CMs has been explored by others through use of transwell plates and conditioned media [45, 51, 52]. Future work would explore the influence of paracrine signaling of iPSC-CMs on Matrigel and decellularized ECM cultured in standard or iPSC-CF conditioned media. The work presented here has focused on mechanical function, but CFs have other effects on CMs including electrophysiology, maturation, and protein expression [6, 10, 11, 33, 51, 53]. Additional analyses could be done to quantify changes in these parameters due to co-culture, decellularized ECM, and conditioned media.
A limitation of this study is that Matrigel, which is a heterogenous and animal-derived ECM, was used. We demonstrated that the decellularized substrates contain CF-deposited ECM fibers that are aligned with the underlying pattern, with an increase in matrix thickness between the Matrigel alone and decellularized substrates. Our collaborators have previously demonstrated that hiPSC-CFs differentiated through the described protocol deposit a dense matrix of fibronectin and collagen type I, while also upregulating an assortment of ECM-associated genes [37]. This matrix deposition is consistent with our findings, as well as with the fibrillar matrix structure and composition of the native human myocardium [54, 55].
To further understand what ECM proteins are being produced and remodeled, a single protein such as collagen, fibronectin, or laminin could be used for pattering. It has previously been demonstrated that iPSC-CFs remodel and produce new ECM when seeded on patterns containing only collagen I or fibronectin [12]. This methodology would allow for improved characterization of the ECM that is deposited by CFs rather than an artifact of the patterned Matrigel, including non-fibrillar components. While we showed qualitatively that collagen and fibronectin are produced, use of a defined ECM could have effects on iPSC-CF remodeling and iPSC-CM function. Fibronectin has been previously demonstrated as essential for both in vivo cardiac repair and in vitro cardiac differentiation. Following myocardial infarction, FN is necessary for the endogenous expansion of cardiac progenitor cells that repair and stabilize cardiac function [56]. Fibronectin knockdown by shRNA is reported to inhibit Brachyury+ mesodermal specification and cardiomyocyte generation of in vitro hiPSCs, further implicating the importance of defined ECM substrates on cardiac differentiation and maturation [57].
Given that this platform is based entirely on iPSCs, it enables the creation of patient specific disease models [58]. Other researchers have shown that adult CFs [6, 46] or CFs from murine models with diseases such as pressure-overloaded hearts [51] allows for study of the interaction of healthy and diseased cells. To generate a human model, this engineered culture platform could be used with iPSCs derived from patients harboring a specific mutation or genetically modified to create a specific disease model affecting CFs, CMs, or both.
Conclusion
While CMs are the cell type primarily responsible for contraction in the heart, CFs also play an important role in maintaining cardiac function including synthesis and remodeling of the ECM, as well as cell-cell communication with CMs. When cultured on compliant micropatterned substrates, iPSC-CFs remodel and produce aligned ECM as well as increasing contractile strain in iPSC-CMs when in co-culture compared to iPSC-CMs cultured alone. iPSC-CF derived ECM was decellularized and used to culture iPSC-CMs alone and in co-culture with iPSC-CFs. The contractile strain was found to be higher in co-culture as compared to the CM-only condition, and no significant difference was identified between the ECM conditions studied. In co-culture or alone, iPSC-CMs maintained spatial organization of their contractions across the sample region. While contractile function of iPSC-CMs was more significantly influenced by co-culture than the ECM condition, use of iPSC-CF derived ECM may be useful to study a variety of iPSC-CM disease models based on interaction with iPSC-CFs and their ECM. These findings highlight the importance of utilizing CFs to produce more physiologically relevant culture systems for studying CMs.
References
Camelliti, P., T. K. Borg, and P. Kohl. Structural and functional characterisation of cardiac fibroblasts. Cardiovasc. Res. 65(1):40–51, 2005. https://doi.org/10.1016/j.cardiores.2004.08.020.
MacKenna, D., S. R. Summerour, and F. J. Villarreal. Role of mechanical factors in modulating cardiac fibroblast function and extracellular matrix synthesis. Cardiovasc. Res. 46(2):257–263, 2000. https://doi.org/10.1016/S0008-6363(00)00030-4.
Fan, D., A. Takawale, J. Lee, and Z. Kassiri. Cardiac fibroblasts, fibrosis and extracellular matrix remodeling in heart disease. Fibrogenesis Tissue Repair. 5(1):15, 2012. https://doi.org/10.1186/1755-1536-5-15.
Souders, C. A., S. L. K. Bowers, and T. A. Baudino. Cardiac fibroblast: the renaissance cell. Circ. Res. 105(12):1164–1176, 2009. https://doi.org/10.1161/CIRCRESAHA.109.209809.
Hall, C., K. Gehmlich, C. Denning, and D. Pavlovic. Complex relationship between cardiac fibroblasts and cardiomyocytes in health and disease. J Am. Heart Assoc.10(5):e019338, 2021. https://doi.org/10.1161/JAHA.120.019338.
Li, Y., H. Asfour, and N. Bursac. Age-dependent functional crosstalk between cardiac fibroblasts and cardiomyocytes in a 3D engineered cardiac tissue. Acta Biomater. 55:120–130, 2017. https://doi.org/10.1016/j.actbio.2017.04.027.
Porter, K. E., and N. A. Turner. Cardiac fibroblasts: at the heart of myocardial remodeling. Pharmacol. Ther. 123(2):255–278, 2009. https://doi.org/10.1016/j.pharmthera.2009.05.002.
Doppler, S. A., et al. Cardiac fibroblasts: more than mechanical support. J Thorac Dis. 9(Suppl 1):S36–S51, 2017. https://doi.org/10.21037/jtd.2017.03.122.
Zhang, D., I. Shadrin, J. Lam, H.-Q. Xian, R. Snodgrass, and N. Bursac. Tissue-engineered Cardiac Patch for Advanced Functional Maturation of Human ESC-derived Cardiomyocytes. Biomaterials. 34(23):5813–5820, 2013. https://doi.org/10.1016/j.biomaterials.2013.04.026.
Giacomelli, E., et al. Human-iPSC-derived cardiac stromal cells enhance maturation in 3D cardiac microtissues and reveal non-cardiomyocyte contributions to heart disease. Cell Stem Cell. 26(6):862-879.e11, Jun.2020. https://doi.org/10.1016/j.stem.2020.05.004.
Beauchamp, P., et al. 3D co-culture of hiPSC-derived cardiomyocytes with cardiac fibroblasts improves tissue-like features of cardiac spheroids. Front Mol Biosci. 7:14, Feb.2020. https://doi.org/10.3389/fmolb.2020.00014.
Napiwocki, B. N., et al. Micropattern platform promotes extracellular matrix remodeling by human PSC-derived cardiac fibroblasts and enhances contractility of co-cultured cardiomyocytes. Physiol. Rep.9(19):e15045, 2021. https://doi.org/10.14814/phy2.15045.
Pfannkuche, K., et al. Fibroblasts facilitate the engraftment of embryonic stem cell-derived cardiomyocytes on three-dimensional collagen matrices and aggregation in hanging drops. Stem Cells Dev. 19(10):1589–1599, Oct.2010. https://doi.org/10.1089/scd.2009.0255.
Kensah, G., et al. Murine and human pluripotent stem cell-derived cardiac bodies form contractile myocardial tissue in vitro. Eur. Heart J. 34(15):1134–1146, Apr.2013. https://doi.org/10.1093/eurheartj/ehs349.
Liau, B., N. Christoforou, K. Leong, and N. Bursac. Pluripotent stem cell-derived cardiac tissue patch with advanced structure and function. Biomaterials. 32(35):9180–9187, 2011. https://doi.org/10.1016/j.biomaterials.2011.08.050.
Thavandiran, N., et al. Design and formulation of functional pluripotent stem cell-derived cardiac microtissues. Proc. Natl. Acad. Sci. USA. 2013. https://doi.org/10.1073/pnas.1311120110.
Matsuura, K., et al. Creation of mouse embryonic stem cell-derived cardiac cell sheets. Biomaterials. 32(30):7355–7362, 2011. https://doi.org/10.1016/j.biomaterials.2011.05.042.
Kurose, H. Cardiac fibrosis and fibroblasts. Cells. 10(7):1716, 2021. https://doi.org/10.3390/cells10071716.
Kim, D.-H., et al. Nanoscale cues regulate the structure and function of macroscopic cardiac tissue constructs. Proc. Natl. Acad. Sci. USA. 107(2):565–570, 2010. https://doi.org/10.1073/pnas.0906504107.
Xing, Q., C. Vogt, K. W. Leong, and F. Zhao. Highly aligned nanofibrous scaffold derived from decellularized human fibroblasts. Adv. Funct. Mater. 24(20):3027–3035, 2014. https://doi.org/10.1002/adfm.201303460.
Ogle, B. M., et al. Distilling complexity to advance cardiac tissue engineering. Sci. Transl. Med. 8(342):342, 2016. https://doi.org/10.1126/scitranslmed.aad2304.
van Putten, S., Y. Shafieyan, and B. Hinz. Mechanical control of cardiac myofibroblasts. J. Mol. Cell. Cardiol. 93:133–142, 2016. https://doi.org/10.1016/j.yjmcc.2015.11.025.
Hazeltine, L. B., et al. Effects of substrate mechanics on contractility of cardiomyocytes generated from human pluripotent stem cells. Int J Cell Biol. 2012. https://doi.org/10.1155/2012/508294.
Ribeiro, A. J. S., et al. Contractility of single cardiomyocytes differentiated from pluripotent stem cells depends on physiological shape and substrate stiffness. Proc. Natl. Acad. Sci. USA. 112(41):12705–12710, 2015. https://doi.org/10.1073/pnas.1508073112.
Zhao, Y., et al. A platform for generation of chamber-specific cardiac tissues and disease modeling. Cell. Jan.2019. https://doi.org/10.1016/j.cell.2018.11.042.
Ronaldson-Bouchard, K., et al. Advanced maturation of human cardiac tissue grown from pluripotent stem cells. Nature. 2018. https://doi.org/10.1038/s41586-018-0016-3.
Shadrin, I. Y., et al. Cardiopatch platform enables maturation and scale-up of human pluripotent stem cell-derived engineered heart tissues. Nat. Commun. 8(1):1, 2017. https://doi.org/10.1038/s41467-017-01946-x.
Kim, D.-H., K. Han, K. Gupta, K. W. Kwon, K.-Y. Suh, and A. Levchenko. Mechanosensitivity of fibroblast cell shape and movement to anisotropic substratum topography gradients. Biomaterials. 30(29):5433–5444, 2009. https://doi.org/10.1016/j.biomaterials.2009.06.042.
Malte, Tiburcy, et al. Defined engineered human myocardium with advanced maturation for applications in heart failure modeling and repair. Circulation. 135(19):1832–1847, 2017. https://doi.org/10.1161/CIRCULATIONAHA.116.024145.
Nunes, S. S., et al. Biowire: a new platform for maturation of human pluripotent stem cell derived cardiomyocytes. Nat. Methods. 10(8):781–787, 2013. https://doi.org/10.1038/nmeth.2524.
Ruan, J.-L., et al. Mechanical stress conditioning and electrical stimulation promote contractility and force maturation of induced pluripotent stem cell-derived human cardiac tissue. Circulation. 134(20):1557–1567, 2016. https://doi.org/10.1161/CIRCULATIONAHA.114.014998.
Salick, M. R., et al. Micropattern width dependent sarcomere development in human ESC-derived cardiomyocytes. Biomaterials. 35(15):4454–4464, 2014. https://doi.org/10.1016/j.biomaterials.2014.02.001.
Napiwocki, B. N., et al. Aligned human cardiac syncytium for in vitro analysis of electrical, structural, and mechanical readouts. Biotechnol. Bioeng. 118(1):442–452, 2021. https://doi.org/10.1002/bit.27582.
Salick, M. R., B. N. Napiwocki, R. A. Kruepke, G. T. Knight, R. S. Ashton, and W. C. Crone. The scanning gradient Fourier transform (SGFT) method for assessing sarcomere organization and alignment. J. Appl. Phys.127(19):194701, 2020. https://doi.org/10.1063/1.5129347.
Lian, X., et al. Robust cardiomyocyte differentiation from human pluripotent stem cells via temporal modulation of canonical Wnt signaling. Proc. Natl. Acad. Sci. USA. 109(27):E1848–E1857, 2012. https://doi.org/10.1073/pnas.1200250109.
Tohyama, S., et al. Distinct metabolic flow enables large-scale purification of mouse and human pluripotent stem cell-derived cardiomyocytes. Cell Stem Cell. 12(1):127–137, Jan.2013. https://doi.org/10.1016/j.stem.2012.09.013.
Zhang, J., et al. Functional cardiac fibroblasts derived from human pluripotent stem cells via second heart field progenitors. Nat. Commun. 10(1):1–15, May2019. https://doi.org/10.1038/s41467-019-09831-5.
Palchesko, R. N., L. Zhang, Y. Sun, and A. W. Feinberg. Development of polydimethylsiloxane substrates with tunable elastic modulus to study cell mechanobiology in muscle and nerve. PLoS ONE.7(12):e51499, Dec.2012. https://doi.org/10.1371/journal.pone.0051499.
B. N. Napiwocki, M. R. Salick, R. S. Ashton, and W. C. Crone, “Controlling hESC-CM Cell Morphology on Patterned Substrates Over a Range of Stiffness,” in Mechanics of Biological Systems and Materials, Volume 6, C. S. Korach, S. A. Tekalur, and P. Zavattieri, Eds., in Conference Proceedings of the Society for Experimental Mechanics Series. Cham: Springer International Publishing, 2017, pp. 161–168. doi: https://doi.org/10.1007/978-3-319-41351-8_23.
Notbohm, J., et al. Two-dimensional culture systems to enable mechanics-based assays for stem cell-derived cardiomyocytes. Exp Mech. 59(9):1235–1248, 2019. https://doi.org/10.1007/s11340-019-00473-8.
Bar-Kochba, E., J. Toyjanova, E. Andrews, K.-S. Kim, and C. Franck. A fast iterative digital volume correlation algorithm for large deformations. Exp Mech. 55(1):261–274, 2015. https://doi.org/10.1007/s11340-014-9874-2.
Treloar, K. K., and M. J. Simpson. Sensitivity of edge detection methods for quantifying cell migration assays. PLoS ONE.8(6):e67389, 2013. https://doi.org/10.1371/journal.pone.0067389.
Naito, H., et al. Optimizing engineered heart tissue for therapeutic applications as surrogate heart muscle. Circulation. 114(1):1–72, 2006. https://doi.org/10.1161/CIRCULATIONAHA.105.001560.
Miragoli, M., G. Gaudesius, and S. Rohr. Electrotonic modulation of cardiac impulse conduction by myofibroblasts. Circ. Res. 98(6):801–810, 2006. https://doi.org/10.1161/01.RES.0000214537.44195.a3.
LaFramboise, W. A., et al. Cardiac fibroblasts influence cardiomyocyte phenotype in vitro. Am. J. Physiol. Cell Physiol. 292(5):C1799–C1808, 2007. https://doi.org/10.1152/ajpcell.00166.2006.
Liau, B., C. P. Jackman, Y. Li, and N. Bursac. Developmental stage-dependent effects of cardiac fibroblasts on function of stem cell-derived engineered cardiac tissues. Sci. Rep. 7(1):1, 2017. https://doi.org/10.1038/srep42290.
Sartiani, L., E. Bettiol, F. Stillitano, A. Mugelli, E. Cerbai, and M. E. Jaconi. Developmental changes in cardiomyocytes differentiated from human embryonic stem cells: a molecular and electrophysiological approach. Stem. Cells. 25(5):1136–1144, 2007. https://doi.org/10.1634/stemcells.2006-0466.
Snir, M., et al. Assessment of the ultrastructural and proliferative properties of human embryonic stem cell-derived cardiomyocytes. Am. J. Physiol. Heart Circ. Physiol. 285(6):H2355–H2363, 2003. https://doi.org/10.1152/ajpheart.00020.2003.
Guo, Y., and W. T. Pu. Cardiomyocyte maturation. Circ. Res. 126(8):1086–1106, Apr.2020. https://doi.org/10.1161/CIRCRESAHA.119.315862.
Lundy, S. D., W.-Z. Zhu, M. Regnier, and M. A. Laflamme. Structural and functional maturation of cardiomyocytes derived from human pluripotent stem cells. Stem Cells Dev. 22(14):1991–2002, Jul.2013. https://doi.org/10.1089/scd.2012.0490.
Cartledge, J. E., et al. Functional crosstalk between cardiac fibroblasts and adult cardiomyocytes by soluble mediators. Cardiovasc. Res. 105(3):260–270, 2015. https://doi.org/10.1093/cvr/cvu264.
Zhang, X., et al. Cardiomyocyte differentiation induced in cardiac progenitor cells by cardiac fibroblast-conditioned medium. Exp. Biol. Med. (Maywood). 239(5):628–637, 2014. https://doi.org/10.1177/1535370214525323.
Trieschmann, J., et al. The interaction between adult cardiac fibroblasts and embryonic stem cell-derived cardiomyocytes leads to proarrhythmic changes in in vitro cocultures. Stem Cells Int. 2016:2936126, 2016. https://doi.org/10.1155/2016/2936126.
McCurdy, S., C. F. Baicu, S. Heymans, and A. D. Bradshaw. Cardiac extracellular matrix remodeling: fibrillar collagens and secreted protein acidic and rich in cysteine (SPARC). J. Mol. Cell Cardiol. 48(3):544–549, 2010. https://doi.org/10.1016/j.yjmcc.2009.06.018.
Lindsey, M. L., M. Jung, M. E. Hall, and K. Y. DeLeon-Pennell. Proteomic analysis of the cardiac extracellular matrix: clinical research applications. Expert Rev. Proteomics. 15(2):105–112, 2018. https://doi.org/10.1080/14789450.2018.1421947.
Konstandin, M. H., et al. Fibronectin is essential for reparative cardiac progenitor cell response following myocardial infarction. Circ. Res. 2013. https://doi.org/10.1161/CIRCRESAHA.113.301152.
Zhang, J., et al. “Cardiac differentiation of human pluripotent stem cells using defined extracellular matrix proteins reveals essential role of fibronectin. eLife. 11:69028, 2022. https://doi.org/10.7554/eLife.69028.
Stempien, A., et al. Identifying features of cardiac disease phenotypes based on mechanical function in a catecholaminergic polymorphic ventricular tachycardia model. Front. Bioeng. Biotechnol.10:873531, May2022. https://doi.org/10.3389/fbioe.2022.873531.
Acknowledgements
The authors would like to thank Jodi Lawson for her assistance in data processing and manuscript preparation. This research was funded by the University of Wisconsin-Madison, through the Karen Thompson Medhi Professorship, Graduate School, and Office of the Vice Chancellor for Research and Graduate Education (WCC). Support was also provided by the National Institutes of Health, under Ruth L. Kirschstein National Research Service Award T32 HL 007936 from the National Heart Lung and Blood Institute to the University of Wisconsin-Madison Cardiovascular Research Center (AS). TJK and and JZ were supported for this work by National Institutes of Health U01HL134764 and National Science Foundation 1648035. This material is based upon work supported by (while serving at) the National Science Foundation (WCC). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health nor the National Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Associate Editor Jane Grande-Allen, PhD oversaw the review of this article.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13239_2024_711_MOESM1_ESM.tif
(A) Representative 10x objective images of decellularized ECM used in this study on days 0 (prior to CF seeding on micropatterned substrates), 11. 14, and 18 immunolabeled for Collagen Type I (COL1A; teal) and Laminin (green). Laminin present on day 0 originates from Matrigel patterning. Scale bar = 250 µm. (B) Representative 100x objective images of Laminin remodeling at bridge regions from day 0 to 18. Scale bar = 25 µm. (C) Z – stack thickness of 100x images of decellularized scaffolds on days 0 (Matrigel only), 11, 14, and 18. N = 4 for each day. **p < .01, *** p < .001. Supplementary file9 (TIF 19348 kb)
13239_2024_711_MOESM2_ESM.tif
Heatmap images of the maximum state of contraction for CM Only on Matrigel (top left), CM-CF coculture on Matrigel (top right), CM Only on decellularized ECM (bottom left) and CM-CF coculture on decellularized ECM (bottom right). Maximum states of contraction were identified as the maximum average second principal strain calculated by DIC software. Regions of the imaging field not occupied by cells were excluded from calculations and are denoted by the white regions in the heatmap still. The second principal strain values reported in the legend are in percent. Full length heatmap videos, as well as the corresponding raw phase contrast video for each representative image are included as Supplemental Videos 1-8. Supplementary file10 (TIF 5081 kb)
13239_2024_711_MOESM3_ESM.tif
(Left) From a brightfield video of contracting iPSC-CMs, full-field displacements in x and y and computed for the duration of the video. (Middle) The resulting displacement trajectories are shown. The angle of each vector is computed and small trajectories, likely occurring in the regions where cells are not present, are filtered based on the trajectory magnitudes. (Right) A histogram of the trajectory angles is made, and the primary angle used to compute the percent of all trajectories within 20° of the peak angle. Representative histograms of the trajectory angles for (top) chevron, (middle) lane, and (bottom) monolayer pattern. Supplementary file11 (TIF 1231 kb)
13239_2024_711_MOESM4_ESM.tif
Percent of displacement trajectories with 20° alignment for days 6 (top), 12 (center), and 18 (bottom) in both Matrigel and decellularized ECM and culture conditions on chevron patterned substrates (CM Only = black, CM-CF = red). No significant difference exists in overall alignment on any day. Day 6: N= 25 CM Matrigel, N= 22 CM Decellularized, N= 31 CM-CF Matrigel, N = 22 CM-CF Decellularized. Day 12: N= 25 CM Matrigel, N= 19 CM Decellularized, N= 25 CM-CF Matrigel, N = 19 CM-CF Decellularized. Day 18: N= 18 CM Matrigel, N= 16 CM Decellularized, N= 18 CM-CF Matrigel, N = 19 CM-CF Decellularized. Supplementary file12 (TIF 2734 kb)
13239_2024_711_MOESM5_ESM.tif
Effects of micropatterned lane width on displacement trajectory alignment (A), maximum contractile strain (B), and number of neighboring CMs (C). The number of neighboring CMs was determined as the number of nuclei within 100 μm of each individual nucleus. The threshold of 100 μm was chosen as it is the physiological length of adult CMs. For displacement alignment and strain quantification: N = 26 (20 μm), 27 (40 μm), 31 (60 μm), 32 (80 μm), 32 (100 μm), 32 (120 μm), and 29 (140 μm). For the quantification of neighboring CMs, N = 4 biological replicates, with one image of each lane width of interest or chevron pattern taken from each sample. *p < .05, **p < .01 ***p < .001, two-way ANOVA with post hoc Tukey tests. All experiments were performed with CMs only, in the absence of CFs for the duration of culture. Supplementary file13 (TIF 1838 kb)
13239_2024_711_MOESM6_ESM.docx
(A) Quantification of the percent of myofibrils aligned within 10 degrees of the superior angle for the two culture conditions tested determined by SGFT. N = 8 (4 ROIs within a sample, 2 independent samples). (B) α-actinin expression after 18 days of culture for the CM Only (left) and CM-CF (right). α-actinin = green. Scale bars = 50 µm. Supplementary file14 (DOC 6002 kb)
Representative brightfield video recording of a day 12 CM Only sample on Matrigel. Captured on a Nikon Eclipse Ti microscope with a Plan Flour 10x NA 0.3 objective and Nikon DS-QiMc camera with samples maintained at 37°C. Scale bar = 250 µm. Acquisition rate = 18.92 fps (5.55 s). Supplementary file1 (AVI 26955 kb)
Representative brightfield video recording of a day 12 CM-CF sample on Matrigel. Captured on a Nikon Eclipse Ti microscope with a Plan Flour 10x NA 0.3 objective and Nikon DS-QiMc camera with samples maintained at 37°C. Scale bar = 250 µm. Acquisition rate = 18.92 fps (7.45 s). Supplementary file2 (AVI 36264 kb)
Representative brightfield video recording of a day 12 CM Only sample on decellularized ECM. Captured on a Nikon Eclipse Ti microscope with a Plan Flour 10x NA 0.3 objective and Nikon DS-QiMc camera with samples maintained at 37°C. Scale bar = 250 µm. Acquisition rate = 18.92 fps (8.62 s). Supplementary file3 (AVI 45398 kb)
Representative brightfield video recording of a day 12 CM-CF sample on decellularized ECM. Captured on a Nikon Eclipse Ti microscope with a Plan Flour 10x NA 0.3 objective and Nikon DS-QiMc camera with samples maintained at 37°C. Scale bar = 250 µm. Acquisition rate = 18.92 fps (7.93 s). Supplementary file4 (AVI 43510 kb)
Representative heatmap video of day 12 CM Only sample on Matrigel, corresponding to the Supplementary Video 1. Acquisition rate = 18.92 fps (5.55 s). Supplementary file5 (AVI 19269 kb)
Representative heatmap video of a day 12 CM-CF sample on Matrigel, corresponding to the Supplementary Video 2. Acquisition rate = 18.92 fps (7.45 s). Supplementary file6 (AVI 24307 kb)
Representative heatmap video of a day 12 CM Only sample on decellularized ECM, corresponding to the Supplementary Video 3. Acquisition rate = 18.92 fps (8.62 s). Supplementary file7 (AVI 27644 kb)
Representative heatmap video of a day 12 CM-CF sample on decellularized ECM, corresponding to the Supplementary Video 4. Acquisition rate = 18.92 fps (7.93 s). Supplementary file8 (AVI 24139 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Stempien, A., Josvai, M., Notbohm, J. et al. Influence of Remodeled ECM and Co-culture with iPSC-Derived Cardiac Fibroblasts on the Mechanical Function of Micropatterned iPSC-Derived Cardiomyocytes. Cardiovasc Eng Tech 15, 264–278 (2024). https://doi.org/10.1007/s13239-024-00711-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13239-024-00711-8