Abstract
Orchidaceae is a highly evolved and largest angiosperm family, which includes enormous number of species and their hybrids. Recent molecular cytogenetic studies of orchid hybrids have successfully started to reveal their origin and chromosome evolution. Here, we constructed BAC libraries of the two orchid plants, Neofinetia falcata and Rhynchostylis coelestis, as molecular cytogenetic tools, which can be used for chromosome-based comparisons of specific regions between different species and their hybrids chromosomes. A total of 21,000 and 10,600 BAC clones with average insert sizes of 74.6 and 50.8 kb were obtained for the N. falcata and R. coelestis, respectively. Random BAC FISH analyses of the two orchid species revealed distribution of some repetitive sequences in these orchid chromosomes. Thus, these BAC clones are useful resources for understanding the genomic organization of the orchid plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant chromosomes contain a number of complex repetitive sequences, including retroelements, which influence their genome sizes and chromosome evolution [3, 11, 12, 14, 15]. Fluorescence in situ hybridization (FISH) is frequently used to visualize chromosomal distribution of these repetitive elements in plants [2, 7, 9, 13, 24] as a powerful tool for comparative genomics at the microscopic level. In addition, FISH using bacterial artificial chromosome (BAC) probes (BAC FISH) is a useful cytogenetic technique for physical mapping and chromosome marker screening. Because each plant BAC clone usually carries repetitive sequences, random BAC FISH without competitive DNA can visualize complex chromosome organization [18, 20].
The family Orchidaceae is a highly evolved and largest angiosperm family having enormous number of species. Their hybrids often retain its enchanting and splendid flowers. The economically important orchid hybrids have a big share in cut-flower industries worldwide. The genome and transcriptome data of some Orchidaceae plants have been reported recently [1, 4, 16, 22, 23, 25]. Therefore, molecular breeding of orchids has begun vigorously in Asia. Many Asian researchers are interested in the orchid molecular cytogenetics, because their chromosome number is not so variable and inter-species hybrids are easily obtained. Molecular cytogenetic techniques are undoubtedly valuable tools to elucidate genome organization in orchids; BAC-FISH analysis has important contribution in chromosome-based comparisons.
We are interested in the intergeneric hybrid of orchid, eg. Neostylis Lou Sneary, which is an economically valuable orchid hybrid of Neofinetia falcata (2n = 38) and Rhynchostylis coelestis (2n = 38). These two parental species are both aerophyte native to East Asia, which possess beautiful flowers (Fig. 1). Especially, N. falcata is historically famous in Japan and called as “Furan.” In this study, to elucidate the genomic and chromosomal organization of the highly crossable N. falcata and R. coelestis, we constructed BAC libraries of these plants, and performed BAC-FISH analysis. These BAC libraries can be further used as important tools for molecular cytogenetics in the orchid hybrids.
Materials and methods
Plant materials
Neofinetia falcata (2n = 38) and Rhynchostylis coelestis (2n = 38) were grown at room temperature. Young leaves and roots were used to isolate high-molecular-weight (HMW) DNA and chromosomal preparation, respectively.
Chromosomal preparation
Root tips were pretreated with 2 mM 8-hydroxyquinoline for 5 h at 14 °C. The pretreated root tips were fixed in ethanol/glacial acetic acid (3:1) for several days, and then squashed in 45 % acetic acid on glass slides as described by Mukai et al. [8].
Construction of BAC libraries
BAC libraries were constructed according to the conventional method as described before [5, 10, 18, 20, 21]. HMW DNA embedded in agarose plugs was prepared by the typical rapid procedure [17, 20], and was partially digested with HindIII (Takara) at 37 °C for 12 or 20 min. Partially-digested DNA was then subjected to pulsed field gel electrophoresis (PFGE) using a CHEF mapper (Bio-Rad) with a 1 % low melting point agarose gel (SeaPlaque GTG, FMC) in 0.5× TBE buffer with the conditions of 6 V/cm, constant linearly ramped pulse time of 90 s for 4 h at 14 °C, followed by 6 V/cm, constant linearly ramped pulse time of 6 s for 12 h at 14 °C. The target sizes of DNA fragments were released from the selected fractions (around 150 kb) of the gel with ß-Agarase I (New England Biolabs), and ligated into the 8.2-kb HindIII-digested CopyControl™ pCC1BAC™ vector (Epicentre) with T4 ligase (Promega). Subsequently, the ligated DNA was electroporated into Escherichia coli strain ElectroMAX DH10B (Invitrogen) using a Gene Pulser II (Bio-Rad) under the conditions of 1.25 or 1.5 kV, 25 μF, and 100 Ω. Recombinants were clearly distinguished from nonrecombinants on an LB agar plate containing 12.5 mg/l chloramphenicol, X-gal and IPTG after 20–24 h of incubation at 37 °C. Insert size of BAC DNA was determined by NotI digestion followed by PFGE.
Probe labeling and BAC-FISH analysis
Randomly selected BAC colonies were inoculated into 3 ml of LB with 12.5 mg/l chloramphenicol, and shaken at 37 °C overnight. BAC DNA purified by the standard alkali method was dissolved in 20 μl of TE, and the 4-μl aliquot was labelled with digoxigenin-11-dUTP or biotin-16-dUTP using a nick translation kit (Roche Diagnostics). For the FISH analysis, chromosomal DNA was denatured in 70 % formamide-2×SSC for 2 min at 69 °C and dehydrated in an ethanol series at -20 °C. The 10 μl of hybridization mixture (50 % formamide, 10 % dextran sulfate, 50 μg of blocking DNA, and 2×SSC, with 2.5 μl of the labelled BAC DNA), which had been denatured for 10 min at 100 °C and cool on ice for 10 min, was applied to each slide. Overnight hybridization was done in a moist chamber at 37 °C. After hybridization, the slides were washed in 2×SSC at room temperature for 5 min, 50 % formamide-2×SSC at 37 °C for 15 min, 2×SSC at room temperature for 15 min, 1×SSC at room temperature for 15 min, and 4×SSC at room temperature for 5 min. Biotin and digoxigenin were simultaneously detected with fluorescein isothiocyanate (FITC)-conjugated avidin (Roche Diagnostics) and rhodamine-conjugated sheep anti-digoxigenin Fab fragment (Roche Diagnostics), respectively. Slides were incubated in 2 μg/ml FITC-conjugated avidin and 2 μg/ml rhodamine-conjugated antidigoxigenin in detection buffer consisting of 4×SSC-1 % BSA for 1 h at 37 °C. After incubation, the slides were washed in 4×SSC for 10 min, 0.1 % Triton X-100 in 4×SSC for 10 min, 4×SSC for 10 min, and 2×SSC for 5 min, all at room temperature. Slides were mounted in a fluorescence antifade solution (1.25 % DABCO, 90 % glycerol). DAPI was used as chromosome DNA counterstaining in the antifade solution at 200 ng/ml. Each fluorescent signal on the slide was captured with an Axioskop fluorescence microscope (Zeiss) coupled to a cooled CCD camera (Hamamatsu Photonics, model 4880). Images were pseudo-colored and merged using Photoshop software (Adobe).
Results and discussion
The BAC libraries of N. falcata and R. coelestis were constructed from HindIII partially digested DNA by using the 8.2-kb pCC1BAC vector. In total, 21,000 and 10,600 clones were selected as glycerol stocks for N. falcata and R. coelestis BAC libraries, respectively. For the glycerol stocks, we used the 100-clone pooled procedure [19], which can be used for polymerase chain reaction (PCR) screening. Average inset sizes of N. falcata and R. coelestis BAC clones were 74.6 and 50.8 kb, respectively. Because the most abundant range of insert sizes was from 40 to 60 kb (Fig. 2), we concluded that the BAC clones were sufficient for FISH probes. Total insert sizes were calculated as 1.57 and 0.54 Gb in N. falcata and R. coelestis, respectively. Leitch et al. [6] estimated the DNA amounts corresponding to the 1C value of Neofinetia and Rhynchostylis as 2.4 and 2.6 pg, which can be converted to 2.35 and 2.54 Gb of genome sizes (using 1 pg = 978 Mb), respectively. Accordingly, our BAC libraries of N. falcata and R. coelestis estimate 0.67 and 0.21 genome coverage, respectively.
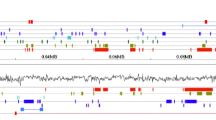
The insert size of the two orchid BAC libraries. a Distribution of the insert size in 125 BAC clones in Neofinetia falcata. b Representative results of PFGE analysis of the NotI-digested BAC clones in Rhynchostylis coelestis. Each insert DNA fragment migrated more slowly than the common pCC1BAC vector band (Vector)
Since we have previously reported that localization of repetitive DNA can be detected by BAC-FISH analysis without adding competitive DNA [18, 20], the same strategy was applied to the two orchid species. In N. falcata, 106 BAC clones were applied to the BAC-FISH analysis (Table 1). Almost half of the BAC clones showed dispersed repetitive signals on the chromosomes; this result was more similar to asparagus-type than onion-type of the BAC-FISH patterns [20]. It might be possible to speculate that 32 clones showing no signal might contain unique sequences, because high-quality chromosomal preparation in orchid was quite difficult to obtain higher resolution signals by the BAC-FISH analysis. We found comparatively high proportion (ca. 20 %) of clones showing proximal and distal BAC FISH signals, suggesting that chromosomal region specific localization of repetitive sequences is rather common in N. falcata.
Figure 3 showed the representative results of the BAC FISH analysis. Two different BAC signals of N. falcata were simultaneously detected on the N. falcata chromosomes (Fig. 3a), showing that BACs contain respective repetitive sequence(s) preferentially distributed to proximal and distal regions of all chromosomes. Similarly, signals from one BAC clone of R. coelestis were appeared on the distal regions of the most chromosomes (Fig. 3b). The distribution patterns of repetitive sequences are also known in other plants (eg. asparagus, onion, and wheat) [18, 20].
Representative BAC FISH results in Neofinetia falcata (a) and Rhynchostylis coelestis (b). a FISH signals from two BAC clones, C-2 (66 kb) and A-53 (65 kb), were simultaneously detected in proximal (red) and distal (green) regions of N. falcata chromosomes, respectively. b Distal repetitive signals (red) from a B-15 BAC clone (125 kb) were detected on R. coelestis chromosomes (blue). The Bars represent 10 μm
In the present study, we constructed BAC libraries of N. falcata and R. coelestis, and BAC-FISH revealed the chromosomal distribution of the orchid repetitive sequences. A previous study reported that centromeric BAC clones were obtained from a tobacco BAC library whose cloning site was HindIII [10]. Because our orchid BAC libraries were also used the HindIII digestion for cloning, we will try to isolate the centromeric DNAs in Neofinetia and Rhynchostylis, which are useful for comparative cytogenetic studies in their hybrids.
Abbreviations
- BAC:
-
Bacterial artificial chromosome
- FISH:
-
Fluorescence in situ hybridization
- FITC:
-
Fluorescein isothiocyanate
- HMW:
-
High-molecular-weight
- PCR:
-
Polymerase chain reaction
- PFGE:
-
Pulsed field gel electrophoresis
References
Cai J, Liu X, Vanneste K, Proost S, Tsai WC, Liu KW, Chen LJ, He Y, Xu Q, Bian C, Zheng Z, Sun F, Liu W, Hsiao YY, Pan ZJ, Hsu CC, Yang YP, Hsu YC, Chuang YC, Dievart A, Dufayard JF, Xu X, Wang JY, Wang J, Xiao XJ, Zhao XM, Du R, Zhang GQ, Wang M, Su YY, Xie GC, Liu GH, Li LQ, Huang LQ, Luo YB, Chen HH, Van de Peer Y, Liu ZJ. The genome sequence of the orchid Phalaenopsis equestris. Nat Genet. 2015;47:65–72.
Fukui KN, Suzuki G, Lagudah ES, Rahman S, Appels R, Yamamoto M, Mukai Y. Physical arrangement of retrotransposon-related repeats in centromeric regions of wheat. Plant Cell Physiol. 2001;42:189–96.
Hawkins JS, Kim H, Nason JD, Wing RA, Wendel JF. Differential lineage-specific amplification of transposable elements is responsible for genome size variation in Gossypium. Genome Res. 2006;16:1252–61.
Huang JZ, Lin CP, Cheng TC, Chang BC, Cheng SY, Chen YW, Lee CY, Chin SW, Chen FC. A de novo floral transcriptome reveals clues into Phalaenopsis orchid flower development. PLoS ONE. 2015;10:e0123474.
Ito T, Suzuki G, Ochiai T, Nakada M, Kameya T, Kanno A. Genomic organization of the AODEF gene in Asparagus officinalis L. Genes Genet Syst. 2005;80:95–103.
Leitch IJ, Kahandawala I, Suda J, Hanson L, Ingrouille MJ, Chase MW, Fay MF. Genome size diversity in orchids: consequences and evolution. Ann Bot. 2009;104:469–81.
Miller JT, Dong F, Jackson SA, Song J, Jiang J. Retrotransposon-related DNA sequences in the centromeres of grass chromosomes. Genetics. 1998;150:1615–23.
Mukai Y, Endo TR, Gill BS. Physical mapping of the 5S rDNA multigene family in common wheat. J Hered. 1990;81:290–5.
Mukai Y, Nakahara Y, Yamamoto M. Simultaneous discrimination of the three genomes in hexaploid wheat by multicolor fluorescence in situ hybridization using total genomic and highly repeated DNA probes. Genome. 1993;36:489–94.
Nagaki K, Shibata F, Suzuki G, Kanatani A, Ozaki S, Hironaka A, Kashihara K, Murata M. Coexistence of NtCENH3 and two retrotransposons in tobacco centromeres. Chromosom Res. 2011;19:591–605.
Neumann P, Koblízková A, Navrátilová A, Macas J. Significant expansion of Vicia pannonica genome size mediated by amplification of a single type of giant retroelement. Genetics. 2006;173:1047–56.
Piegu B, Guyot R, Picault N, Roulin A, Saniyal A, Kim H, Collura K, Brar DS, Jackson S, Wing RA, Panaud O. Doubling genome size without polyploidization: dynamics of retrotransposition-driven genomic expansions in Oryza australiensis, a wild relative of rice. Genome Res. 2006;16:1262–9.
Presting GG, Malysheva L, Fuchs J, Schubert I. A Ty3/gypsy retrotransposon-like sequence localizes to the centromeric regions of cereal chromosomes. Plant J. 1998;16:721–8.
SanMiguel P, Bennetzen JL. Evidence that a recent increase in maize genome size was caused by the massive amplification of intergene retrotransposons. Ann Bot. 1998;82:37–44.
Shirasu K, Schulman AH, Lahaye T, Schulze-Lefert P. A contiguous 66-kb barley DNA sequence provides evidence for reversible genome expansion. Genome Res. 2000;10:908–15.
Su C, Chao YT, Chang YCA, Chen WC, Chen CY, Lee AY, Hwa KT, Shih MC. De novo assembly of expressed transcripts and global analysis of the Phalaenopsis aphrodite transcriptome. Plant Cell Physiol. 2011;52:1501–14.
Suzuki G, Watanabe M, Toriyama K, Isogai A, Hinata K. Direct cloning of the Brassica S locus by using a P1-derived artificial chromosome (PAC) vector. Gene. 1997;199:133–7.
Suzuki G, Ura A, Saito N, Do GS, Seo BB, Yamamoto M, Mukai Y. BAC FISH analysis in Allium cepa. Genes Genet Syst. 2001;76:251–5.
Suzuki G, Do GS, Mukai Y. Efficient storage and screening system for onion BAC clones. Breeding Sci. 2002;52:157–9.
Suzuki G, Ogaki Y, Hokimoto N, Xiao L, Kikuchi-Taura A, Harada C, Okayama R, Tsuru A, Onishi M, Saito N, Do GS, Lee SH, Ito T, Kanno A, Yamamoto M, Mukai Y. Random BAC FISH of monocot plants reveals differential distribution of repetitive DNA elements in small and large chromosome species. Plant Cell Rep. 2012;31:621–8.
Tomita RN, Suzuki G, Yoshida K, Yano Y, Tsuchiya T, Kakeda K, Mukai Y, Kowyama Y. Molecular characterization of a 313-kb genomic region containing the Self incompatibility locus of Ipomoea trifida, a diploid relative of sweet potato. Breeding Sci. 2004;54:165–75.
Tsai WC, Fu CH, Hsiao YY, Huang YM, Chen LJ, Wang M, Liu ZJ, Chen HH. OrchidBase 2.0: comprehensive collection of orchidaceae floral transcriptomes. Plant Cell Physiol. 2013;54:e7.
Yan L, Wang X, Liu H, Tian Y, Lian J, Yang R, Hao S, Wang X, Yang S, Li Q, Qi S, Kui L, Okpekum M, Ma X, Zhang J, Ding Z, Zhang G, Wang W, Dong Y, Sheng J. The genome of Dendrobium officinale illuminates the biology of the important traditional Chinese orchid herb. Mol Plant. 2015;8:922–34.
Zhang P, Li W, Fellers J, Friebe B, Gill BS. BAC-FISH in wheat identifies chromosome landmarks consisting of different types of transposable elements. Chromosoma. 2004;112:288–99.
Zhang J, Wu K, Zeng S, Teixeira da Silva JA, Zhao X, Tian CE, Xia H, Duan J. Transcriptome analysis of Cymbidium sinense and its application to the identification of genes associated with floral development. BMC Genomics. 2013;14:279.
Acknowledgments
This work was supported in part by Grants-in-Aid for Scientific Research (C) (No. 25450006 to Y.M.; No. 25450515 to G.S.), and Scientific Research on Innovative Areas (Nos. 23113006, 23113001 to G.S.) from the Japan Society for Promotion of Science (JSPS).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Matsuba, A., Fujii, M., Lee, S.S. et al. Molecular cytogenetic use of BAC clones in Neofinetia falcata and Rhynchostylis coelestis . Nucleus 58, 207–210 (2015). https://doi.org/10.1007/s13237-015-0147-y
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13237-015-0147-y