Abstract
Species of the Trichophyton benhamiae complex are predominantly zoophilic pathogens with a worldwide distribution. These pathogens have recently become important due to their epidemic spread in pets and pet owners. Considerable genetic and phenotypic variability has been revealed in these emerging pathogens, but the species limits and host spectra have not been clearly elucidated. In this study, we used an approach combining phylogenetic analysis based on four loci, population-genetic data, phenotypic and physiological analysis, mating type gene characterization and ecological data to resolve the taxonomy of these pathogens. This approach supported the inclusion of nine taxa in the complex, including three new species and one new variety. Trichophyton benhamiae var. luteum var. nov. (“yellow phenotype” strains) is currently a major cause of zoonotic tinea corporis and capitis in Europe (mostly transmitted from guinea pigs). The isolates of the “white phenotype” do not form a monophyletic group and are segregated into three taxa, T. benhamiae var. benhamiae (mostly North America; dogs), T. europaeum sp. nov. (mostly Europe; guinea pigs), and T. japonicum sp. nov. (predominant in East Asia but also found in Europe; rabbits and guinea pigs). The new species T. africanum sp. nov. is proposed for the “African” race of T. benhamiae. The introduction to new geographic areas and host jump followed by extinction of one mating type gene have played important roles in the evolution of these pathogens. Due to considerable phenotypic similarity of many dermatophytes and phenomena such as incomplete lineage sorting or occasional hybridization and introgression, we demonstrate the need to follow polyphasic approach in species delimitation. Neutrally evolving and noncoding DNA regions showed significantly higher discriminatory power compared to conventional protein-coding loci. Diagnostic options for species identification in practice based on molecular markers, phenotype and MALDI-TOF spectra are presented. A microsatellite typing scheme developed in this study is a powerful tool for the epidemiological surveillance of these emerging pathogens.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dermatophytes are a group of fungal pathogens that cause inflammatory and contagious skin diseases that are usually referred to as dermatophytoses, tinea or ringworm. These are among the most common diseases of warm-blooded animals, including humans, and their prevalence can reach dozens of percent in both human and animal populations (Agnetti et al. 2014; Ahdy et al. 2016; Cafarchia et al. 2010; Duarte et al. 2010; Havlickova et al. 2008; Kupsch et al. 2017; Seebacher et al. 2008). The treatment and prevention of these infections in humans, companion animals and pets require a considerable amount of funding every year (Benedict et al. 2018; Bond 2010; Chermette et al. 2008; Kane and Summerbell 1997; Shenoy and Jayaraman 2019).
The incidence of zoonotic dermatomycoses transmitted to humans from livestock decreased significantly in developed countries with the intensification of agriculture, introduction of preventive measures (e.g., vaccination in cattle) and advances in treatment options (Borman et al. 2007; Lund et al. 2014). In contrast, zoonotic infections transmitted from pets remain an important public health concern worldwide (Hubka et al. 2018d). Microsporum canis and Trichophyton mentagrophytes remain major agents of dermatophytosis in many domestic animals and cause a significant number of zoonotic dermatophytoses in humans (Hayette and Sacheli 2015). In addition to these well-known causal agents, several emerging zoonotic pathogens are increasingly reported in both humans and pets, and most of them belong to the Trichophyton benhamiae complex.
The Trichophyton benhamiae complex currently comprises six species: T. benhamiae, T. bullosum, T. concentricum, T. erinacei, T. eriotrephon and T. verrucosum (de Hoog et al. 2017; Lysková et al. 2015). These species are predominantly zoophilic, with the exception of anthropophilic T. concentricum, an agent of tinea imbricata in tropical regions (Bonifaz et al. 2004; Bonifaz and Vazquez-Gonzalez 2011; Pihet et al. 2008). Trichophyton verrucosum, a cause of dermatophytosis in cattle and other ruminants, is one of the best-known members of the complex. It has a worldwide distribution and causes economic losses in the food (negative impacts on milk and meat production), hide and skin industries (Bond 2010; Chermette et al. 2008). The incidence of infections in cattle has decreased in many regions in response to vaccination programmes or changes in agricultural systems, and the rate of infections in humans has decreased proportionally (Seebacher et al. 2008; Lund et al. 2014). By contrast, a lack of prophylaxis accounts for the high infection rates observed in countries such as Italy (Moretti et al. 2013). Trichophyton verrucosum grows slowly in culture and frequently produces only chlamydospores as its main microscopic characteristic. In this respect, it is superficially very similar to T. bullosum, which causes infections in donkeys and horses, but is much less common and is geographically restricted to the Middle East, Africa and Europe (Lysková et al. 2015; Sabou et al. 2018; Sitterle et al. 2012). Scant data are available on the distribution of T. eriotrephon, which is only known from several poorly documented cases of dermatophytosis in humans and dogs (Hubka et al. 2018d; Rezaei-Matehkolaei et al. 2013; Sabou et al. 2018). The remaining two zoophilic species, T. benhamiae and T. erinacei, are currently considered emerging pathogens, as their incidence as a cause of infections in pets and humans has increased significantly in the last decade (Hubka et al. 2018d).
A strikingly high incidence of zoonotic T. benhamiae (syn. Arthroderma benhamiae) infections, contracted mostly from guinea pigs, is currently reported in various European countries. Although this species was considered less clinically important in recent decades, it became one of the most common agents of zoonotic dermatophytoses after 2010 (Hubka et al. 2018b; Nenoff et al. 2014; Sabou et al. 2018; Symoens et al. 2013; Uhrlaß et al. 2015). It has been shown that the prevalence of the pathogen in guinea pig breeds and pet shops reaches up to 90% (Bartosch et al. 2019; Drouot et al. 2009; Guillot et al. 2018; Kupsch et al. 2017; Overgaauw et al. 2017). Infections occur more frequently in young guinea pigs and are usually asymptomatic. The presence of skin lesions with hair loss (mostly on the muzzle, forehead, ears and around eyes) is also reported in some individuals (Kraemer et al. 2013, 2012). When transmitted to the human host, the infections manifest most commonly as highly inflammatory tinea of glabrous skin and tinea capitis and less commonly as onychomycosis (Nenoff et al. 2014; Skořepová et al. 2014). The presence of asymptomatic infections in animal hosts contributes to the successful spread of the pathogen between animals kept in groups. Such asymptomatic infections also facilitate transmission to pet owners and the occurrence of small familial outbreaks or general infections among pet breeders, pet shop workers and others. In addition to guinea pigs, this pathogen has been reported in dogs, rabbits, cats, North American porcupines, various small rodents and foxes (Aho 1980; Fréalle et al. 2007; Hiruma et al. 2015; Needle et al. 2019; Sieklucki et al. 2014; Takeda et al. 2012; Ziółkowska et al. 2015).
Trichophyton benhamiae was originally described as Arthroderma benhamiae, a sexual and heterothallic species, from several canine and human infections in North America (Ajello and Cheng 1967). In subsequent years, Takashio (1974, 1977) recognized two races among strains of T. benhamiae based on biological compatibility experiments: an “Americano-European” race and an “African” race of Arthroderma benhamiae. Furthermore, two phenotypically different groups among strains of the Americano-European race have recently been recognized by different authors and designated the “yellow phenotype” and “white phenotype” strains (Brasch et al. 2016; Hiruma et al. 2015; Nenoff et al. 2014; Symoens et al. 2013). The characterization of mating type genes showed that the MAT1-1-1 idiomorph was significantly prevalent among strains of the yellow phenotype, while MAT1-2-1 prevailed among strains of the white phenotype (Symoens et al. 2013). Similar observations of a lack of one MAT gene or significant bias towards one MAT idiomorph have been made in several other primary pathogenic dermatophytes, while the prevalence of both mating types in a balanced ratio is common in geophilic species (Kosanke et al. 2018; Metin and Heitman 2017).
It was demonstrated that the vast majority of European infections are caused by yellow phenotype strains that emerged relatively recently (Hubka et al. 2014; Nenoff et al. 2014; Symoens et al. 2013; Uhrlaß et al. 2015). The first documented cases of infections due to yellow phenotype strains were recorded between 2002 and 2008 in France and Switzerland (Contet-Audonneau and Leyer 2010; Charlent 2011; Khettar and Contet-Audonneau 2012; Symoens et al. 2013). The first cases in Germany and the Czech Republic were described shortly before 2010, and the pathogen became rapidly epidemic during the following years. Currently, T. benhamiae is the most important agent of dermatophytoses transmitted from animals in the Czech Republic and Germany (Hubka et al. 2018b; Hubka et al. 2014; Kupsch et al. 2019; Nenoff et al. 2014; Uhrlaß et al. 2015). The origin of yellow phenotype strains of T. benhamiae and the reason for the sudden increase in the incidence of human and animal infections in Europe after 2010 are unknown. As the breeding of guinea pigs has been popular in Europe for decades, the epidemic cannot be explained by a change in pet owner behaviour. Therefore, the spread of a new virulent and highly transmissible genotype/lineage was hypothesized (Čmoková 2015; Hubka et al. 2018d). The occurrence of T. benhamiae infections in non-European countries is generally poorly known except for individual reported cases. This is mostly due to insufficient surveillance and a lack of long-term epidemiological studies supported by molecular-based identification of dermatophytes.
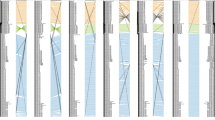
In contrast to yellow phenotype strains, white phenotype strains have probably existed worldwide for a long time. Sporadic human and animal infections due to white phenotype strains were described from various European countries, Japan and the USA before the widespread dispersal of yellow phenotype strains in Europe (Aho 1980; Ajello and Cheng 1967; Hejtmánek and Hejtmánková 1989; Kano et al. 1998; Takashio 1974). In Japan, white phenotype strains were first reported in 1996 from an infected rabbit (Kano et al. 1998); human cases were reported in the following years (Nakamura et al. 2002), and the infections were summarized by Kimura et al. (2015). The increasing number of people breeding pets, together with the increasing import of animals to Japan, is considered a cause of the increased incidence in Japan (Hiruma et al 2015; Kimura et al 2015; Takeda et al 2012). Chronology of reports of white and yellow phenotype strains in various countries is summarized in Fig. 1.
Chronology of reports of Trichophyton benhamiae phenotypes from various countries. Yellow-phenotype isolates correspond to T. benhamiae var. luteum proposed in this study. White-phenotype strains correspond to T. benhamiae var. benhamiae and two novel species proposed here: T. europaeum and T. japonicum. The reports are mostly sorted according to the phenotypic characters of cultures reported by the authors and, in more recent studies, by a combination of DNA sequencing and morphology. The icons of the hosts are explained in Fig. S1
The aim of this study was to elucidate the species boundaries, host spectrum, and population structure of emerging pathogens in the Trichophyton benhamiae complex. We examined a large set of clinical isolates associated with human and animal infections that were mostly collected in European countries but also in the USA and Japan. We conducted DNA sequencing of four genetic loci, phylogenetic analyses, and analyses of morphology and physiology to examine whether the previously detected level of phenotypic and genetic variability reflects undescribed species diversity or a high level of infraspecific variability. The levels of recombination/clonality within species and populations, respectively, were estimated by calculating the index of association and determining the ratios between MAT locus idiomorphs. MALDI-TOF MS spectra were compared between species of the T. benhamiae complex to test the possibility of their differentiation in the clinical setting. A set of highly variable microsatellite markers were developed to analyse the population structure and relationships between strains with differences in their geographic origin, host spectrum and phenotype. The new taxonomic classification and microsatellite typing scheme proposed in this study will enable the monitoring of changes in the frequencies of individual species and genotypes. This will help to evaluate the results of preventive measures and interventions and is a basic prerequisite for the development of epidemiological studies.
Materials and methods
Source of isolates
More than three hundred strains isolated from human and animal patients with dermatophytosis caused by pathogens from the T. benhamiae complex were obtained for this study from various clinical laboratories, hospitals and universities (Table S1): Laboratory for Medical Microbiology (Mölbis, Germany), College of Veterinary Medicine, University of Illinois at Urbana-Champaign (USA), College of Bioresource Sciences, Nihon University (Japan), Laboratory of Mycology, Department of Veterinary Sciences, University of Turin (Italy), and various institutions in the Czech Republic (Institute of Public Health in Ostrava and Usti nad Labem, General University Hospital in Prague, University Hospital in Pilsen, Hospital České Budějovice, Hospital in Pardubice and Labvet veterinary laboratory in Prague). This set of strains was further supplemented with isolates from culture collections, especially BCCM/IHEM Fungi Collection: Human and Animal Health (Brussels, Belgium) and CBS culture collection housed at the Westerdijk Institute (Utrecht, The Netherlands).
Selected isolates were deposited into the Culture Collection of Fungi (CCF), Department of Botany, Charles University, Prague, Czech Republic; herbarium specimens of newly described species were deposited into the herbarium of the Mycological Department, National Museum in Prague, Czech Republic (PRM).
Molecular studies
DNA was extracted from seven-day-old colonies using the ArchivePure DNA Yeast and Gram2 + Isolation Kit (5 PRIME Inc., Gaithersburg, Maryland) according to the manufacturer’s instructions as updated by Hubka et al. (2018c). The quality of the extracted DNA was evaluated by NanoDrop 1000 Spectrophotometer.
The ITS rDNA region (ITS1-5.8S-ITS2 cluster) was amplified using the primer set SR6R and LR1 (White et al. 1990) or ITS1F and ITS4 (Gardes and Bruns 1993; White et al. 1990), partial gapdh gene encoding glyceraldehyde-3-phosphate dehydrogenase was amplified with primers GPDF and GPDR (Kawasaki et al. 2011), partial tubb gene encoding β-tubulin with primers Bt2a and Bt2b (Glass and Donaldson 1995), and tef1α gene encoding translation elongation factor 1-α with primers EF-DermF and EF-DermR (Mirhendi et al. 2015). All primer combinations are listed in Table S2. Reaction volume of 20 µL contained 1 µL (50 ng mL−1) of DNA, 0.3 µL of both primers (25 pM mL−1), 0.2 µL of My Taq Polymerase and 4 μL of 5 × My Taq PCR buffer (Bioline, London, UK). PCR conditions followed protocol described by Hubka et al. (2018a). PCR product purification followed protocol of Réblová et al. (2016). Automated sequencing was performed at Macrogen Sequencing Service (Amsterdam, The Netherlands) using both terminal primers. The DNA sequences obtained in this study were deposited into the GenBank database (www.ncbi.nlm.nih.gov) under the accession numbers listed in Table 1.
Phylogenetic analysis
Alignments of the ITS, gapdh, tubb and tef1α regions were performed using the FFT-NS-i option implemented with the MAFFT online service (Katoh et al. 2017). The alignments were trimmed, concatenated and then analysed using maximum likelihood (ML) and Bayesian inference (BI) methods. Suitable partitioning schemes and substitution models (Bayesian information criterion) for the analyses were selected using a greedy strategy implemented in PartitionFinder 2 (Lanfear et al. 2017) with settings allowing introns, exons, codon positions and segments of the ITS region to be independent datasets. The optimal partitioning schemes for each analysed dataset along with basic alignment characteristics are listed in Table S3. The ML trees were constructed with IQ-TREE version 1.4.4 (Nguyen et al. 2015) with nodal support determined by nonparametric bootstrapping (BS) with 1000 replicates. The trees were rooted with Trichophyton rubrum. Bayesian posterior probabilities (PP) were calculated using MrBayes 3.2.6 (Ronquist et al. 2012). Optimal partitioning scheme and substitution models were selected as described above and are listed in Table S3. The analysis ran for 107 generations, two parallel runs with four chains each were used, every 1000th tree was retained, and the first 25% of trees were discarded as burn-in. The convergence of the runs and effective sample sizes were checked in Tracer v1.6 (https://tree.bio.ed.ac.uk/software/tracer).
The modified complex indel coding (MCIC) algorithm implemented in SeqState version 1.25 (Müller 2005) was used to code gaps. The TCS network method (Clement et al. 2000) was used to generate haplotype networks implemented in the program PopART (Leigh and Bryant 2015).
Development of microsatellite markers
Microsatellite motifs were identified in the available genomic sequence of T. europaeum CBS 112371 = IHEM 20161 = CCF 6479 (https://www.broadinstitute.org/) using WebSat online software (Martins et al. 2009). The same program suggested optimal primers for the amplification of target loci. We selected di-, tri-, and tetranucleotide repeats based on the loci with the highest repeat numbers. Interrupted repeats as well as loci containing two or more repeat motifs within the fragments delimited by particular primer pairs were excluded. A pilot set of eight strains was used to evaluate microsatellite polymorphism for all candidate loci following the method of Schuelke (2000). PCR conditions were as follows: one cycle at 95 °C for 1 min; 27 cycles at 95 °C for 30 s, 55 °C for 30 s, 72 °C for 45 s, followed by eight cycles at 95 °C for 30 s, 53 °C for 30 s, 72 °C for 45 s and a final extension at 72 °C for 10 min. A set of 24 loci exhibiting the highest level of polymorphism was selected from the 160 tested loci. The PCR products were screened for the presence of undesirable polymorphisms in the microsatellite flanking regions and the presence of polymorphisms in the microsatellite regions by sequencing. Emphasis was also placed on the selection of loci that were approximately uniformly distributed across the genome. Primer–primer interactions were checked before assembling multiplexes using Multiple Primer Analyzer (https://www.thermoscientificbio.com/webtools/multipleprimer/). The forward primers of ten selected loci were tagged with fluorescent dye and arranged into a single multiplex panel (Table 2). The reaction volume of 5 µL for PCR contained 50 ng DNA, 0.5 μL of the mixture of primers and 2.5 μL of Multiplex PCR Master Mix (QIAGEN, Germany). The PCR conditions were chosen according to the manufacturer's recommendations. The PCR products (diluted in water 1:50) were mixed with 10 µL of deionized formamide and 0.2 µL of the GeneScan™ 600 LIZ size standard and denatured for 5 min at 95 °C, followed by analysis on an ABI 3100 Avant Genetic Analyzer.
Statistical analysis of microsatellite data
The discriminatory power of these newly designed loci was calculated using Simpson’s index of diversity as described previously (Hunter and Gaston 1988). A binary and allele data matrix was created using GeneMarker 1.51 software (SoftGenetics, LLC, State College, PA, USA) and used to estimate the similarities between individuals using Jaccard’s similarity coefficient calculation in the program FAMD (Schlueter and Harris 2006). A neighbour-joining tree based on Jaccard’s similarity coefficient matrix was constructed using the same software. Genetic distances were calculated from the same matrix and used for the construction of the NeighborNet network in the SplitsTree 4 program (Huson 1998).
A Bayesian model-based clustering algorithm with a clustering number (K) = 1–10 was applied to the clone-corrected allele data matrix using the software STRUCTURE (Pritchard et al. 2000). Ten simulations were calculated at the www.bioportal.uio.no server (Lifeportal, University of Oslo) using the admixture model and 106 MCMC replicates; 5 × 108 replicates were discarded as burn-in. The no-admixture model and uncorrelated allele frequencies were chosen for the analysis. The optimal clustering number (K) was estimated using ΔK and similarity coefficients (Evanno et al. 2005), and both values were calculated using the script structure-sum (Ehrich 2006) in the R version 3.3.4 program (R Core Team 2016).
The genetic variability within and between clusters was analysed for ten variable loci on the clone-corrected dataset via analysis of molecular variance (AMOVA) (Excoffier et al. 1992) in the Arlequin program (Schneider et al. 2000). The degree of gene flow among clusters was estimated using a pairwise fixation index (FST) and a coefficient of genetic differentiation (GST) calculated in Arlequin (Schneider et al. 2000) and POPGENE (Yeh et al. 1999), respectively.
The degree of clonality or recombination within particular clusters was estimated by calculating the index of association (IA) in the program MultiLocus 1.3 (Agapow and Burt 2001), which is used for measuring the linkage disequilibrium between alleles and is useful in inferring the occurrence of cryptic recombination in putatively asexual populations (Burt et al. 1996). Random mating is suggested if no linkage is detected between the alleles of different loci (randomly distributed alleles); in that case IA is expected to be nearly zero or zero. We tested for significant deviation from 10,000 random multilocus permutations of genotypes under a random mating model.
To measure within-population diversity, Nei’s genotype diversity (Dg) was calculated based on frequencies of genetically distinct individuals, and Nei’s gene diversity (D) was calculated based on the frequencies of alleles at individual loci (Kosman 2003; Nei 1987). The effective number of genotypes (Geff) (Parker 1979) was calculated based on the number of equally abundant genotypes required to reflect the value of a diversity measure. It was calculated to obtain diversity values comparable between the clusters. The degree of genetic divergence was investigated by rarity index of (DW index; frequency down-weighted marker values) (Schönswetter and Tribsch 2005). All mentioned population indexes (Dg, D, DW, Geff) were calculated from the clone-corrected binary data matrix using script AFLPdat (Ehrich 2006) in R 3.0.2. Frequency histograms of pairwise differences between individuals were generated using the same program.
MAT locus determination
A partial sequence of the MAT1-1-1 gene encoding the alpha box domain was amplified with the primers MF1 and MF5, and a partial sequence of the MAT1-2-1 gene encoding the high mobility group (HMG) domain was amplified with the primers Ab_HMG_F and Ab_HMG_R or TmHMG3S and TmHMG3R (Kano et al. 2012; Kosanke et al. 2018; Symoens et al. 2013). The PCR volume of 10 µL contained 25 ng of DNA, 0.15 µL of both primers (25 pM mL−1), 0.15 µL of My Taq Polymerase and 2 μL of buffer. The PCR conditions are described above. The PCR products were visualized in an electrophoretogram (1% agarose gel with 0.5 μg mL−1 ethidium bromide). Several PCR products of each MAT idiomorph were subjected to sequencing for the confirmation of product specificity.
Phenotypic studies
The morphology of the colonies on malt extract agar (MEA, HiMedia, Mumbai, India) at 25 °C was documented in all strains. At least five strains from each species (if available) were subjected to a detailed analysis that involved macromorphology on MEA, potato dextrose agar (PDA, Himedia, Mumbai, India) and Sabouraud dextrose agar [SAB, Atlas (2010)] at 25, 30 and 37 °C. The macromorphology of the colonies was documented using an Olympus SZ61 or Canon EOS 500D binocular loupe (with Olympus Camedia C-5050 Zoom camera). Colony colour determinations were made using the ISCC-NBS Centroid Colour Charts (Kelly 1964); https://tx4.us/nbs/nbs-1.htm.
Micromorphology was documented using an Olympus BX-51 microscope. Particular micromorphological characteristics were recorded at least 35 times for each isolate (at least five strains selected per species). The variance inflation factor (VIF) was assessed before performing the analysis of variance to test the correlation between variables. Statistical differences in particular phenotypic characteristics were tested with one-way analysis of variance (ANOVA) followed by Tukey’s honestly significant difference (HSD) test in program R version 3.3.4 (R Core Team 2016).
MALDI-TOF mass spectrometry
The cultivation of strains from the T. benhamiae clade (up to five strains from each species, if available) was performed in liquid cultivation medium for 22–24 h. The strains were prepared according to Schrödl et al. (2012) and analysed by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS). In brief, for MALDI-TOF MS analysis, all samples were prepared using the liquid cultivation method and ethanol / formic acid extraction method. One milliliter of each overnight culture was centrifuged for 2 min at about 10,000 × g. The supernatant was carefully removed and the fungal pellet was resuspended in 1 ml water, mixed thoroughly, and centrifuged for further 5 min at 10,000 × g. After removing the supernatant the pellet was resuspended in a mixture of 300 µL bidistilled water and 900 µL absolute ethanol. After centrifugation, the fungal cells were dried shortly and mixed thoroughly with 50 µL of 70% formic acid and 50 µL pure acetonitrile, followed by centrifugation for 2 min at 10,000×g. A volume of 1 µL supernatant was placed onto a MALDI target plate (Bruker Daltonik GmbH, Germany) and allowed to dry at room temperature. Eight MALDI target positions per strain were prepared in parallel. Each sample position (including one Bruker Bacterial Test Standard position) was overlaid with 1 µL of matrix (HCCA portioned; Bruker Daltonik GmbH, Germany) and air dried at room temperature. MALDI-TOF MS measurement was conducted on a Microflex LT benchtop instrument operated by FlexControl software (Bruker Daltonik GmbH, Leipzig, Germany). Spectra were acquired in linear positive mode at a laser frequency of 200 Hz within a mass range from 2000 to 20,000 Da by using the standard flexControl and AutoX methods. For each sampled spot up to three sum spectra were accumulated resulting in 24 MALDI spectra per strain. Finally, five spectra were selected for better spectra handling and visualization.
Results
Phylogeny of the Trichophyton benhamiae complex
We assessed 340 combined ITS, gapdh, tubb and tef1-α sequences from members of the T. benhamiae species complex (TBSC) in the phylogenetic analysis. The final alignment included 2371 characters, with 247 variable and 152 parsimony informative sites, and Trichophyton rubrum CBS 202.88 was used as the outgroup. The detailed alignment characteristics together with the partitioning schemes and substitution models are listed in Table S3. The isolation source and accession numbers for the DNA sequences are available in Table 1 and Table S1. The alignments are available in the online supplementary material.
Members of the TBSC were resolved into three major monophyletic clades in the best scoring multiple-gene ML tree shown in Fig. 2, (single-gene trees are shown in Figs. S2–S5).
Multilocus phylogeny of the Trichophyton benhamiae complex inferred with the maximum likelihood method based on the gapdh, tubb, ITS rDNA and tef1-α loci (alignment characteristics, partitioning scheme and substitution models are listed in Table S3). Maximum likelihood bootstrap values and Bayesian posterior probabilities are appended to the nodes; only support values higher than 70% and 0.90, respectively, are shown. The ex-type strains are designated with a superscripted T. Trichophyton rubrum CBS 202.88 was used as the outgroup
The T. benhamiae clade contains anthropophilic T. concentricum (n = 3) and the Americano-European race of T. benhamiae (n = 318). The isolates of the Americano-European race do not form a monophyletic lineage and are paraphyletic with respect to T. concentricum. These strains are segregated into three major subclades: T. benhamiae s. str. and two newly proposed species, T. japonicum sp. nov. and T. europaeum sp. nov. Isolates of T. benhamiae s. str. originating mostly from Europe and North America, and they comprise both white and yellow phenotype strains. They form a monophyletic and fully supported (100% bootstrap supports, bs/1.00 posterior probability, pp) subclade together with T. concentricum, which can be differentiated by only two unique substitutions in the ITS region and three in the tef1-α gene (the tubb and gapdh genes are identical).
Species from the T. benhamiae clade show a low level of intraspecific genetic variability. In total, there are only seven unique multilocus genotypes (MLST) among 318 isolates belonging to the T. benhamiae clade (Fig. 3). Two MLST genotypes are present among T. benhamiae strains, represented by a single substitution in the tef1-α gene (Fig. S4). Two MLST genotypes are present in T. japonicum, caused by a single substitution in the ITS1 region. Trichophyton japonicum can be differentiated from the closely related T. europaeum by a single substitution in the ITS region and four conserved substitutions in the gapdh gene (Figs. S2-S5). No intraspecific variability is detectable among the isolates of T. europaeum. The only exception is the isolate of “T. europaeum” IHEM 25139, which presents an abnormal ITS1 region sequence that contains 6 additional substitutions compared to the T. europaeum isolates. Some of these positions are critical for the differentiation of the T. europaeum/T. japonicum lineage from T. benhamiae s. str., suggesting that this strain could be a hybrid between T. benhamiae clade species. The gapdh gene sequence of IHEM 25139 is typical of T. europaeum.
Haplotype network of the Trichophyton benhamiae clade based on multilocus data (gapdh, tubb, ITS rDNA and tef1-α loci). Haplotypes are indicated by circles whose sizes correspond to the number of analysed strains, and dashes on the connecting lines indicate substitutions (indels are excluded). The upper figure shows the species identity and genotypic diversity, the middle figure shows the distribution of MAT gene idiomorphs, and the lower figure shows the geographic distribution of particular genotypes
Both MAT gene idiomorphs were only detected among strains of T. benhamiae. Trichophyton japonicum and T. concentricum strains exhibited only the MAT1-1-1 idiomorph, while T. europaeum comprised strains characterized by the presence of the MAT1-2-1 idiomorph. Only “T. europaeum” strain IHEM 25139 showed MAT1-1-1 idiomorph.
The T. erinacei clade comprises three species: T. erinacei, an agent of mycoses in hedgehogs (genera Erinaceus, Aterelix); T. verrucosum, an agent of cattle ringworm; and T. eriotrephon, with poorly known ecological characteristics (Fig. 2). All analyzed isolates of T. erinacei and T. verrucosum presented the MAT1-2-1 idiomorph, while T. eriotrephon exhibited only the MAT1-1-1 idiomorph.The T. bullosum clade contains three isolates of the African race of Arthroderma benhamiae from humans, and T. bullosum which is a causal agent of dermatomycoses in horses and donkeys. Isolates of the African race apparently represent an independent taxonomic entity, and we propose the name T. africanum for this species (Fig. 2). Both MAT gene idiomorphs were detected in T. africanum, while T. bullosum isolates exhibited only the MAT1-1-1 idiomorph.
Analysis of the T. benhamiae clade with newly designed microsatellite markers
A total of 160 microsatellite markers with di- or trinucleotide repeats and motifs longer than eleven repetitions were extracted from the available genome of T. europaeum CBS 112371 using WebSat software (Martins et al. 2009). The number of repeats was inferred by subtracting the known length of the flanking sequence from the total amplicon length. Only 24 regions contained the required repeat and showed length polymorphism in the microsatellite region and an absence of polymorphism in the flanking region. A total of ten markers with an even distribution in the genome and different lengths (for the purpose of multiplexing) were selected for the final analysis (Table 2). The Simpson’s diversity index calculated for particular loci yielded values ranging from 0.34 (TC20 locus) to 0.59 (TAG16 locus). The whole panel consisting of ten markers yielded a diversity index of 0.77 (Table S4).
This newly developed microsatellite typing scheme was applied to a total number of 318 isolates belonging to the T. benhamiae clade. Forward primers of all loci were marked with fluorescent dye and arranged in a multiplex panel (Table 2). The highest number of alleles was found at the TAG16 (n = 12) locus, followed by the CT21 (n = 10) locus. In contrast, the fewest alleles were found in the AG21 (n = 5) and TC20 (n = 5) loci. The remaining loci included 6–9 alleles (Table S4). All loci were successfully amplified in all examined strains (null alleles were not found). The dependence of genotypic diversity on the number of loci showed that a sufficient number of markers was used to resolve the population structure of the T. benhamiae clade. It was apparent from the curves (Fig. 4) that genetic diversity would not increase significantly with the addition of additional markers.
A Bayesian model-based clustering algorithm was used to determine how many groups were included in the dataset. The highest ΔK value was observed at K = 6, and a much lower peak was present at K = 4 (Fig. 5). The estimated population structure inferred from this analysis is shown in Fig. 5. The analysis revealed a total of 41 genotypes among T. benhamiae clade isolates clustering into six clusters (C1–C6).
The population structure of the Trichophyton benhamiae clade (ten microsatellite loci, 318 isolates). The neighbour-joining tree was calculated from the multilocus microsatellite profiles using the Jaccard distance matrix measure in FAMD 1.3 (Schlueter and Harris 2006) and is used solely for the comprehensive presentation of the results. Genetic structure was revealed with STRUCTURE software by Bayesian clustering (the peak of ΔK was observed at K = 6); clones were discarded from the analysis; the number of isolates representing each haplotype is indicated in parentheses following the isolate number; the geographic origin of the isolates representing particular haplotypes is indicated using abbreviations: Europe (Eu), Japan (Jpn), United States of America (USA), Indonesia (Indon), Polynesia (Poly). Individual haplotypes are represented by horizontal bars; the colours were attributed according to the clusters delimited by STRUCTURE
The distribution of the isolates into clusters was correlated with their geographic distribution and main primary hosts (Fig. 6). The cluster C1 was found most abundantly in Europe and was associated with guinea pigs. These isolates are responsible for the current outbreak of infections in Central Europe and consist exclusively of yellow phenotype strains. We propose the name T. benhamiae var. luteum for this cluster. Clusters C2 and C3 comprised white phenotype strains from North America isolated mostly from dogs and characterized by highly variable microsatellite data (T. benhamiae var. benhamiae). Cluster C4 (T. japonicum) comprised the majority of strains from Japan analysed in this study and some European strains (rabbits, guinea pigs and human infections contracted from them). Cluster C5 (T. europaeum) comprised strains from Europe (infections mostly contracted from guinea pigs). The isolate IHEM 25139 was assigned to T. europaeum but its haplotype was intermediate between T. europaeum and T. japonicum (alleles CT21 and CT21b were characteristic of T. japonicum, while the remaining 8 alleles were from T. europaeum). Cluster C6 was represented by three human isolates of T. concentricum from tropical regions.
Population structure of the Trichophyton benhamiae clade revealed by the analysis of ten microsatellite loci in 318 strains. The NeighborNet network was built with FAMD 1.3 software and visualized in SplitsTree 4.13 using the Jaccard index-based distance matrix (Delta score: 0.1778, Q-residual score: 0.01222). The assignment of strains to main clusters and species is indicated by different colours. The labels of each cluster show the geographic origin of strains with the number of isolates and main host(s). The icons of the hosts are explained in Fig. S1
The clustering based on the microsatellite data was correlated with MAT gene distribution and single-gene DNA data (tubb gene was excluded due lack of variability in T. benhamiae clade) (Fig. 7, Fig. S6). It is evident from the visualisation that clustering of isolates according to the single-gene genotype and MAT idiomorphs was in general agreement with microsatellite data and proposed species hypothesis. However, the clusters C1–C3 are not supported by any DNA locus sequences in study and are only distinguishable by microsatellites. Trichophyton benhamiae var. luteum (C1) was characterized by low variability of microsatellite data and exclusively consisted of isolates with MAT1-1-1 idiomorph. The isolates of T. benhamiae var. benhamiae cluster C2 were exclusively of the MAT1-2-1 idiomorph, while those of cluster C3 were exclusively of the MAT1-1-1 idiomorph. Despite obvious phenotypic and population genetic differences between T. benhamiae var. benhamiae and T. benhamiae var. luteum, these two varieties are not distinguishable by any of the DNA sequence markers used in this study. The only detected DNA sequence variant, represented by a single substitution in the tef1-α gene, did not fully correspond to the two varieties delimited by microsatellite markers (Fig. 7).
Phylogenetic tree of the Trichophyton benhamiae clade revealed by the analysis of ten microsatellite loci in 318 strains constructed in FAMD software using a Jaccard index-based distance matrix. Coloured circles display the genotype diversity of the ITS, gapdh and tef1-α loci and the distribution of MAT gene idiomorphs (blue: MAT1-1-1; pink: MAT1-2-1) across Trichophyton benhamiae clade species. Isolate numbers are displayed in Fig. S6
Genetic diversity and population structure analysis of T. benhamiae clade
Population characteristics were calculated from microsatellite data to test significance of clonal expansion versus recombination, and genetic diversity within clusters. Besides the inability to reproduce sexually due to missing opposite mating type in most of species, the clonality is indicated by the skewed distribution of pairwise differences between individuals (Fig. 8). Consequently, all populations are genetically uniform which is evident from low value of Nei’s gene diversity (D) (Table S5) that ranged from 0.02 in T. benhamiae var. benhamiae cluster 3 to 0.156 in T. benhamiae var. benhamiae cluster 2. The low Nei’s genotype diversity (Dg = 0.35) of T. benhamiae var. luteum compared to other taxa reflects the fact that the population consisted of several abundant clones (Table S5). Asexual reproduction prevails in all populations for long time which is supported by the low effective number of genotype (Geff) values that were significantly lower than observed number of genotypes (Table S5). The exception was T. benhamiae var. benhamie cluster C2 (Table S5). However, recombination in cluster C2 was not confirmed by calculation of index of association (IA) (Table S5), possibly due to low number of samples available. The recombination was not rejected only in T. europaeum population according to IA on significance level p < 0.05 (IA = 0.24, p < 0.0042) (Fig. 9, Table S5).
To test cluster-specific differences, AMOVA was performed on the microsatellite data. The diversity between six clusters contributed to a total variability of 68.1%, while the diversity within clusters contributed to only 31.9% (p < 0.0001). Thus, there is a low level of genetic information exchange between clusters, reflected in a high number of fixed alleles (FST = 0.89, GST = 0.75, p < 0.0001).
Trichophyton concentricum and T. benhamiae var. benhamiae cluster C2 shared the greatest number of alleles in common (FST = 0.451, GST = 0.46). The lowest number of shared alleles was found between T. benhamiae var. luteum and all other clusters (FST = 0.90–0.95; Table S6). The strongly fixed set of alleles in T. benhamiae var. luteum indicates low or no gene flow between this cluster and the remaining clusters. Relatively low DW index value (DW = 0.06; Table S5) indicate recent origin of T. benhamiae var. luteum. On the other hand, high DW values in other taxa indicate long-term isolation due to accumulation of unique alleles (Table S5).
Phenotypic studies
Initially, the phenotype of all isolates was recorded on malt extract agar (MEA). It was observed that the morphotypes within the T. benhamiae clade generally corresponded to the clusters delimited by microsatellite analysis. Notable exceptions were the strains showing signs of degeneration (poorly sporulating, white, cottony colonies usually producing no pigments). Such a phenotype is commonly described in dermatophytes and indicates degeneration, usually caused by long-term strain passaging and preservation (de Hoog et al. 2017). These strains were excluded from further phenotype analyses. At least five strains (if available) from each group were selected, and their phenotypes were analysed on three cultivation media (Fig. 10). Growth rates were recorded at three temperatures (Fig. 11), and micromorphology was measured on MEA (Fig. 12). Cultivation on MEA and potato dextrose agar (PDA) promoted sporulation and pigment production most effectively.
Among the taxa from the T. benhamiae clade, the strains of T. concentricum and T. benhamiae var. luteum were characterized by the slowest growth on all media and at all tested temperatures (Fig. 11). No sporulation was observed in the T. concentricum strains examined in this study. Overall, poor sporulation, the production of intense yellow pigmentation as the colony reverse colour and the absence of macroconidia and spiral hyphae were characteristic of T. benhamiae var. luteum (yellow phenotype strains of T. benhamiae). All three remaining species from the T. benhamiae clade produced both micro- and macroconidia and whitish colonies, usually with a brownish, red-brown or red colony reverse colour (white phenotype strains of T. benhamiae). Trichophyton benhamiae var. benhamiae grew more rapidly at 25 °C than the other species from this clade (Fig. 11) and exhibited larger microconidia on average (Fig. 12). The obverse colony colour was whitish or showed a brownish tint, and red-brown pigmentation on the reverse side was commonly arranged into sectors (Fig. 10). The growth parameters and micromorphology of T. japonicum and T. europaeum were very similar (Fig. 11–12), and all strains extensively sporulated.
The phylogenetically distant T. africanum (formerly called “African race”) is characterized by relatively long microconidia (comparable to those of T. benhamiae var. benhamiae) growing on unbranched or loosely branched conidiophores. Compared to T. africanum, the conidiophores of T. benhamiae clade members were either poorly differentiated from vegetative hyphae (conidia sessile on the hyphae) or short with many lateral branches under the top (branched in a pyramidal pattern, grape-like). A more detailed differential diagnosis of particular species with their relatives is included in the Notes in the Taxonomy section.
To compare phenotypic characteristics, the ANOVA was performed on microconidia width, length and growth rates (MEA, SAB, PDA at 25, 30 and 37 °C), followed by a post hoc analysis using Tukey's HSD pairwise comparisons based on the mean values for each strain and a confidence interval of 0.95. All growth rate variables and conidium size variables were strongly correlated. Growth rate and conidium size variables can therefore be used interchangeably (Fig. 13). The analysis showed that there were statistically significant differences between T. benhamiae clade species according to any combination of characteristics, including conidia size and growth rates (p < 0.001). Furthermore, growth rates measured at 25 °C on MEA or PDA can be used independently to distinguish the majority of species (p < 0.001) (Fig. 13, Fig. S7, Table S7). Variables such as microconidium length (Table S8) and width (Table S9) can also be used independently to distinguish particular species, except for T. japonicum and T. benhamiae var. luteum, which cannot be differentiated at the specified significance level.
Principal component analysis (PCA) of morphological characteristics. The two major axes of the plot show all variables, including the growth rates (cultivation on MEA, SAB, and PDA at 25, 30 and 37 °C) and microconidium sizes (mean values of length and width) (a). The correlation matrix shows the Pearson correlation coefficients between variables such as growth rates (three different media and temperatures) and microconidia sizes (length and width). A darker colour indicates stronger correlations, which means that all variables within a growth rate or microconidium size group were strongly correlated (b), indicating the possibility of reducing the number of variables
MALDI-TOF mass spectrometry
Representative isolates of each species from the T. benhamiae clade were analysed using MALDI-TOF mass spectrometry; T. africanum isolates were also included for comparison (Fig. 14). All samples could be measured very well and delivered high quality (peak rich) MALDI spectra. In the mass range between approximately 5900–6200 m/z (as a representative example), the MALDI-TOF mass spectra were very similar both between and within all groups, and differentiation of the groups was not possible within this range. In contrast to this high similarity, several specific peaks could be found for all analysed taxa in the entire mass range of approximately 4000 to 12,000 m/z. The most variable mass range of approximately 4000 to 8000 m/z is shown on Fig. 14. Trichophyton africanum significantly differed from all of the samples in many peaks in its spectrum (Fig. 14a). Trichophyton benhamiae var. luteum and T. benhamiae var. benhamiae shared peaks at 7150 and 7745 m/z in their mass spectra but different peaks at 4112 and 4680 m/z, which are typical of var. luteum, and 6515 and 6530 m/z, which are typical of var. benhamiae (Fig. 14c, d). Both mentioned species differ from T. europaeum and T. japonicum in the absence of a peak at 7150 m/z (data not shown). Trichophyton europaeum differed from T. japonicum in the presence of a peak at 7745 m/z and the absence of a peak at 7715 m/z (Fig. 14b). Trichophyton concentricum differed from both T. benhamiae varieties in its peaks at 4770, 6435 and 7145 m/z (Fig. 14d) and also differed from the rest of the samples in several peaks. To prove the general applicability of the here presented MALDI peaks more strains of the mentioned species / varieties should be analyzed in the future and incorporated into the presented MALDI-based differentiation model.
MALDI-TOF mass spectra in the Trichophyton benhamiae clade members; only variable regions are shown. Comparison of spectra in the species of the former Americano-European race (T. benhamiae var. benhamiae and T. benhamiae var. luteum, T. europaeum and T. japonicum), the African race (T. africanum) and T. concentricum (a). Comparison of T. europaeum, T. japonicum and T. concentricum (b). Comparison of spectra of T. concentricum and two varieties of T. benhamiae (c, d)
Taxonomy
Trichophyton benhamiae clade
Trichophyton benhamiae (Ajello & S.L. Cheng) Y. Gräser & de Hoog [Index Fungorum 356: 2. 2018] var. benhamiae var. nov. (automatically generated; Art. 26.3 [Turland et al. (2018)])—Fig. 15.
Macromorphology and micromorphology of Trichophyton benhamiae var. benhamiae. Colonies after two weeks of cultivation at 25 °C on Sabouraud’s dextrose agar (a, b), malt extract agar (c, d) and potato dextrose agar (e, f). Conidiophores bearing microconidia (g–i) and macroconidia (j); macroconidia (k–p), frequently with mycelial fragments at one or both ends (k–n, p); microconidia (r); spiral hyphae (s). Scale bars = 20 μm
Typus: USA, Missouri, human, L. Ajello, NCDC B765d (holotype), PRM 944659 (epitype, designated here, MBT 394322) a dried culture derived from strain IHEM 4710; ex-epitype culture IHEM 4710 (= CBS 623.66 = ATCC 16781 = CABIM 124768 = CCF 6484 = CDC X-797 = CECT 2892 = IMI 124768 = IP 1064.74 = NCPF 0410 = RV 23303 = UAMH 2822).
Vegetative hyphae smooth, septate, hyaline, 1.5–4 µm diam (mean ± sd; 2.5 ± 0.7). Conidiophores poorly differentiated from vegetative hyphae, mostly unbranched, conidia sessile or born on short lateral branches; pyramidally branches conidiophores less common and with sparse branching. Microconidia abundant, pyriform to clavate, truncate, 2.5–6 (3.8 ± 0.5) × 1.6–3.5 (2.6 ± 0.4) μm. Macroconidia sparse to abundant, cylindrical or elongated fusiform, with pointed or rounded ends, easily disintegrate into fragments with truncate ends, developing intercalary or terminally on vegetative hyphae, frequently released with short to long mycelial fragments at one or both ends, predominantly 3–10-septate (median 8), 23–82 (59.2 ± 15.5) × 4.5–7.5 (6.1 ± 0.8) μm. Chlamydospores present. Spiral hyphae absent or rare. Heterothallic. Sexual state fide Ajello & Cheng (1967) and Čmoková (2015): cleistothecia white to yellowish-white, covered with dichotomously branched peridial hyphae and spiral appendages. Peridial hyphae composed of asymmetrical peridial cells, dumb-bell shaped, echinulate, 8.5–10.5 (9.1 ± 1.8) µm in length, 2.5–4.5 (2.8 ± 0.7) µm in width at enlarged ends, internode width 2–4 µm (2.4 ± 1.2); intercalary conidia sparse, cylindrical or barrel-shaped. Asci globose, eight-spored, ascospores ovate, hyaline to pale yellow, longer dimension up to 3 μm, shorter dimension up to 2 μm.
Culture characteristics (7 days at 25 °C): Colonies on SAB 28–34 mm diam (⌀ = 32 mm), white (#F2F3F4), velvety to powdery, centrally raised, radially furrowed in some strains, edge diffuse, reverse pale orange yellow (#FAD6A5) to light orange yellow (#FBC97F) in the marginal part, vivid orange (#F38400) to deep brown (#593319) in the center. Colonies on MEA 30–35 mm diam (⌀ = 34 mm), velvety to granular, pale yellow-gray (#C7ADA3) to light yellow (#FAD6A5), umbonate, edge diffuse, reverse pale orange yellow (#FAD6A5) to brilliant orange yellow (#FFC14F), red pigment produced in sectors by some strains—deep reddish orange (#AA381E). Colonies on PDA 27–32 mm diam (⌀ = 30 mm), white (#F2F3F4) to light yellow (#FAD6A5), velvety to granular, centrally raised, occasionally with filamentous sectors, reverse pale orange yellow (#FAD6A5) to brilliant orange yellow (#FFC14F), red pigment produced in sectors by some strains—deep reddish orange pigment (#AA381E). Colonies in 7 days at 30 °C grow faster than at 25 °C: SAB 37–45 mm diam (⌀ = 39 mm); PDA 35–43 mm diam (⌀ = 37 mm); MEA 8–43 mm diam (⌀ = 40 mm). Colonies at 37 °C in 7 d: SAB 27–39 mm diam (⌀ = 30 mm); PDA 30–35 mm diam (⌀ = 34 mm); MEA 30–35 mm diam (⌀ = 33 mm).
Material examined: USA, Missouri, human, L. Ajello (PRM 944659, epitype); ex-epitype culture IHEM 4710 (= CBS 623.66 = ATCC 16781 = CABIM 124768 = CCF 6484 = CDC X-797 = CECT 2892 = IMI 124768 = IP 1064.74 = NCPF 0410 = RV 23303 = UAMH 2822). USA, Urbana, dog, 2009 (USA 3208). USA, Urbana, dog, 2006 (USA 3209); ibid., USA 3216. USA, Urbana, cat, 2006 (USA 3220). USA, Urbana, dog, 2007 (USA 3329). USA, Urbana, dog, 2010 (USA 3350 = CCF 6485); ibid., USA 3355; ibid., USA 3356. USA, Urbana, chinchilla, 2011 (USA 3360 = CCF 6486). USA, Urbana, dog, 2011 (USA 3361). USA, Urbana, unknown source, 1991 (USA 3368). USA, Urbana, unknown source, 1989 (USA 3369). USA, Urbana, unknown source, 1997 (USA 3370). USA, Urbana, unknown source, 2001 (USA 3371). USA, Urbana, unknown source, 1996 (USA 3376). USA, Urbana, unknown source, 1995 (USA 3378). In-vitro, monoascospore isolate, 1970, M. Takashio [IHEM 3287 = RV 26678 = CCF 6483; isolate from cross between IHEM 24908 (ex dog, USA) × IHEM 4710 (ex human, USA)]. In-vitro, monoascospore isolate, 1970, M. Takashio [IHEM 3288 = BER 1464 = DSM 6916 = JS 83-006 = RV 26680 = SM 0104 = VUT 77012 = CCRC 31780 = IAM 12705 = JCM 1886; isolate from cross between IHEM 24908 (ex dog, USA) × IHEM 4710 (ex human, USA)].
Typification Ajello & Cheng (1967) designated the specimen NCDC B765d as a holotype of Arthroderma benhamiae, and a dried culture with ascomata was generated by crossing the isolates TM-20 (= ATCC 16781 = IHEM 4710 = CBS 623.66 = CABIM 124768 = CDC X-797 = CECT 2892 = IMI 124768 = IP 1064.74 = NCPF 0410 = RV 23303 = UAMH 2822 = CCF 6484; ex human; MAT1-2-1) × TM-17 (= ATCC 16782 = CBS 624.66 = IHEM 24908 = RV 23302 = CDC X-798 = CECT 2893 = IMI 124769 = NCPF 411 = UAMH 2823; ex dog; MAT1-1-1). Although this specimen exhibits both sexual and asexual morphs in its life cycle, it is not suitable for the purposes of the recent taxonomy for several reasons. First, it is not clear which of the two cultures contained within the type should be considered the ex-holotype culture. Additionally, interspecific hybrids can be induced by crossing opposite mating type strains of unrelated species in vitro as shown in previous studies on dermatophytes (Anzawa et al. 2010; Kawasaki et al. 2009, 2011, 2010), and the deposition of a resultant ‘hybrid’ type could lead to ambiguities. Because it is not possible to recognize which portion of the holotype belongs to a particular isolate, we designated an epitype PRM 944659 (dried culture) derived from the strain IHEM 4710.
Distribution and ecology: Trichophyton benhamiae var. benhamiae is a zoophilic dermatophyte, and isolates examined in this study originated from dogs (n = 8), cats (isolate USA 3220), chinchillas (isolate USA 3360) and unknown hosts (n = 6). Previously reported cases of human infections were probably transmitted from animals (Ajello and Cheng 1967). Another important host of this pathogen is probably the North American porcupine (Erethizon dorsatum) (Needle et al. 2019; Takahashi et al. 2008), a close relative of the guinea pig (Cavia porcellus). Isolates from the North American porcupine exhibited ITS rDNA identical to that of T. benhamiae, and their morphology showed characteristics typical of T. benhamiae var. benhamiae (Needle et al. 2019; Takahashi et al. 2008). All strains examined here were collected in North America (the in vitro-derived isolates IHEM 3287, IHEM 3288, IHEM 4710 were also based on strains of American origin). A recently reported Chinese case of tinea faciei, likely contracted from fox, was probably also caused by T. benhamiae var. benhamiae based on the ITS sequence and morphology of the isolate (Tan et al. 2020).
Notes: The macromorphology of T. benhamiae var. benhamiae most closely resembles those of T. europaeum, T. japonicum and T. mentagrophytes in the production of a red-brown pigment on the reverse side of colonies and abundant microconidia. It differs from T. europaeum and T. japonicum in its host spectrum and higher growth rates, especially on MEA and PDA at 25 °C (Fig. 11). Macroconidia of T. benhamiae var. benhamiae are usually more abundantly produced compared to T. europaeum and T. japonicum, and they are most frequently cylindrical or elongated fusiform with terminal fragments of vegetative hyphae. Closely related T. concentricum differs significantly in its ecology. It is an anthropophilic species occurring in tropical regions, grows very slowly, produces cerebriform colonies without red-brown pigment on the colony reverse and usually does not sporulate. Trichophyton behamiae var. luteum is also strikingly different in its host spectrum (mostly guinea pigs), distribution (mainly Europe) and morphology (slow growth, yellow pigmentation, relatively poor sporulation, absence of macroconidia). Trichophyton benhamiae var. benhamiae does not produce intense yellow pigment on SAB supplemented with chloramphenicol and cycloheximide and MEA, in contrast to T. benhamiae var. luteum. The ratio of MAT1-1-1 and MAT1-2-1 strains was 14:5.
Trichophyton benhamiae (Ajello & S.L. Cheng) Y. Gräser & de Hoog [Index Fungorum 356: 2. 2018] var. luteum Cmokova & Hubka, var. nov.—MycoBank MB835887; Fig. 16
Etymology: Refers to the bright yellow colony reverse produced on all examined media.
Typus: SWITZERLAND, Lausanne, University Hospital Vaudois, dermatophytosis in human, arm skin (tinea corporis), 2009, M. Monod, PRM 944414 (holotype); ex-holotype culture IHEM 25068 (= CCF 6500).
Vegetative hyphae smooth, septate, hyaline, 1–3.5 µm diam (mean ± sd: 1.9 ± 0.5). Conidiophores branched in a pyramidal (grape-like) pattern, sometimes poorly differentiated from vegetative hyphae, unbranched or poorly branched, conidia sessile or born on short lateral branches. Microconidia sparse to abundant, pyriform, less commonly clavate, 2.5–4.9 (3.2 ± 0.4) × 1.5–3.4 (2.1 ± 0.3) μm. Macroconidia not observed in any of the isolates examined. Chlamydospores were not observed. Spiral hyphae not observed. Sexual morph unknown.
Culture characteristics (7 days at 25 °C): colonies on SAB 10–20 mm diam (⌀ = 13 mm), white (#F2F3F4) to yellowish white (#F0EAD6), velvety, flat with radially furrowed center, edge filliform, reverse vivid yellow (#F3C300). Colonies on MEA 6–17 mm diam (⌀ = 12 mm), pale yellow (#F3E5AB), filamentous, in large extent submerged, flat, edge filliform, reverse light yellow (#F8DE7E) to vivid yellow (#F3C300). Colonies on PDA 9–17 mm diam (⌀ = 13 mm), light yellow (#F8DE7E) to pale yellow (#F3E5AB), velvety, flat, radially furrowed, edge filliform, reverse brilliant orange yellow (#FFC14F) to vivid yellow (#F3C300). Colonies at 30 °C in 7 d: SAB 15–26 mm diam (⌀ = 21 mm); PDA 18–22 mm diam (⌀ = 21 mm); MEA 21–22 mm diam (⌀ = 22 mm). Colonies at 37 °C in 7 d: SAB 15–20 mm diam (⌀ = 18 mm); PDA 10–17 mm diam (⌀ = 12 mm); MEA 11–13 mm diam (⌀ = 11 mm).
Material examined: Switzerland, Lausanne, University Hospital Vaudois, dermatophytosis in human, arm skin (tinea corporis), 2009, M. Monod (PRM 944414, holotype, dried culture; PRM 944415, isotype); ex-holotype culture IHEM 25068 = CCF 6500. Japan, common degu, 2012 (NUBS 13001). Switzerland, Lausanne, University Hospital Vaudois, human skin, 2009, M. Monod (IHEM 25066). Czechia, Prague, guinea pigs (Cavia porcellus), 2014, J. Koubová (KOUB 23); ibid., KOUB 51; ibid., KOUB 77. Germany, Berlin, dermatophytosis in human, 2010 (BER 24); ibid., BER 211; ibid., BER 212; ibid., BER 213. Czechia, České Budějovice, dermatophytosis in human, 2012 (D126); ibid., D295; ibid., D375; ibid., D417; ibid., D521. Germany, Mölbis, dermatophytosis in human, 2015 (DE 200156); ibid., DE 200351; ibid., DE 200465. Belgium, Brussels, dermatophytosis in human, 2012 (IHEM 25744 = CCF 6476); ibid., IHEM 25743; ibid., IHEM 25742 = CCF 6474; ibid., IHEM 25466; ibid., IHEM 25745. Czechia, Prague, dermatophytosis in human, 2012 (CCF 4849); ibid., CCF 4850; ibid., CCF 4851; ibid., CCF 4852. All 236 strains examined in this study are listed in Table S1.
Distribution and ecology: Trichophyton benhamiae var. luteum is a zoophilic species with the guinea pig as the main host (Hubka et al. 2018d). It is widely distributed in Europe, but it has also been detected in common degu (Octodon degus) in Japan (Hiruma et al. 2015) and was recently isolated from human dermatophytosis in Brazil (de Freitas et al. 2019; Grisólia 2019) and Iraq (S. Uhrlaß, unpublished data) (Table S10).
The European strains of T. benhamiae var. luteum (n = 236) examined here were predominantly obtained from humans (~ 72% from females and ~ 28% from males, median age 12 years) who mostly reported contact with guinea pigs; the remaining strains were recovered from animals (guinea pigs and common degu) (Table S1). The human infections mostly manifested as highly inflammatory tinea corporis, tinea faciei and tinea capitis (Fig. 17). By contrast, infected animals were mostly symptomless. Symptomatic guinea pigs usually showed localized lesions with scaling and crusting or alopecia located predominantly on the head, less frequently on the other body parts (Fig. 17). Green fluorescence of infected tissues may be observed under Wood's light in some strains, similar to M. canis (Skořepová et al. 2014). Only the MAT1-1-1 idiomorph was detected in the T. benhamiae var. luteum isolates examined here.
Clinical presentation of infections caused by Trichophyton benhamiae clade species in guinea pigs and humans. Guinea pigs: area of alopecia with scaling located in the temporal area (a); areas with scaling on the ear (b); area of alopecia with scaling located on the back (c); itchy area of alopecia behind the ear (d) and on the guinea pig's abdomen (e); weeping lesion under the eye (f). Zoonotic infections in humans: tinea corporis located on the thigh (g) and chest (h), tinea faciei (i), tinea barbae (j), tinea capitis profunda (k, l)
Notes: The macromorphology of T. benhamiae var. luteum resembles that of Microsporum canis in the production of intense yellow pigments. However, M. canis usually produces abundant spindle-shaped macroconidia, which are absent in T. benhamiae var. luteum. The differentiation of sterile M. canis isolates may be more difficult but is possible according to its higher growth parameter values. In addition, these species differ in their main hosts, which are cats and dogs in M. canis and guinea pigs in T. benhamiae var. luteum. The closely related anthropophilic species T. concentricum differs in its ecology, colony characteristics (no yellow pigment produced) and microscopic characteristics (usually no sporulation). Other taxa from the T. benhamiae clade differ in showing higher growth rates (Fig. 11), the production of red/brown pigments and the production of macroconidia, which are absent in T. benhamiae var. luteum. In addition to these differences, T. benhamiae var. luteum can be clearly distinguished from T. benhamiae var. benhamiae and other species in the T. benhamiae clade by microsatellite data (Figs. 5–6) and MALDI-TOF MS spectra (Fig. 14).
Trichophyton concentricum R. Blanch., Traité de Pathologie Générale 2: 916. 1896—Fig. 18
Macromorphology and micromorphology of Trichophyton concentricum. Colonies after three weeks of cultivation at 25 °C on Sabouraud’s dextrose agar (a, b), malt extract agar (c, d) and potato dextrose agar (e, f). Vegetative hyphae (g–l), frequently consisting of inflated cells and containing intercalary or terminal chlamydospores (h, i), occasionally proliferating in a zigzag pattern (k, l). Scale bars = 20 μm
Vegetative hyphae smooth, septate, frequently inflated, occasionally with knob-like terminations, often proliferating in a zigzag pattern, hyaline, 1.5–4 µm diam (mean ± sd; 2.7 ± 0.7). Chlamydospores common, usually globose or ovate, intercalar, terminal or in short chains. Conidiophores, conidia, pectinate hyphae and favic chandeliers were not observed among the examined strains. Sexual morph unknown.
Culture characteristics (7 days at 25 °C): Colonies on SAB 6–16 mm diam (⌀ = 11 mm), pale orange yellow (#FAD6A5) to pale yellowish pink (#ECD5C5), membranous to slightly velvety, raised, umbonate or cerebriform, deeply furrowed, edge filiform or lobate, reverse light orange yellow (#FBC97F). Colonies on MEA 9–16 mm diam (⌀ = 15 mm), pale orange yellow (#FAD6A5) to pale yellowish pink (#ECD5C5), membranous to slightly velvety, umbonate, edge filiform, reverse light orange yellow (#FBC97F) to brilliant orange yellow (#FFC14F), vivid yellow (#F3C300) in narrow centre. Colonies on PDA 5–12 mm diam (⌀ = 11 mm), pale orange yellow (#FAD6A5) to pale yellowish pink (#ECD5C5), membranous, raised, deeply furrowed to cerebriform, edge irregular to lobate, reverse light orange yellow (#FBC97F) to brilliant orange yellow (#FFC14F), vivid yellow (#F3C300) in narrow centre. Colonies at 30 °C in 7 d: SAB 8–20 mm diam (⌀ = 16 mm); MEA 8–15 mm diam (⌀ = 13 mm); PDA 10–14 mm diam (⌀ = 11 mm). Colonies at 37 °C in 7 d: SAB 5–14 mm diam (⌀ = 10 mm); MEA 5–13 mm diam (⌀ = 10 mm); PDA 5–13 mm diam (⌀ = 9 mm).
Material examined: Polynesia, human, 1926, A. Castellani (ex-neotype strain CBS 196.26 = IFO 5972). Fiji, human skin, 1963 (CCF 5303 = IHEM 13435 = RV 30442). Indonesia, Manado, human, arm and trunk skin, 1990, W. Warow (CCF 5302 = IHEM 5470).
Distribution and ecology: Trichophyton concentricum is an anthropophilic species distributed in Oceania, Southeast Asia, and Central and South America. It is a cause of tinea imbricata (tokelau) usually affecting rural indigenous populations. The clinical manifestation is very characteristic and gives human skin ornate appearance due to the presence of concentric squamous plaques (Bonifaz et al. 2004; Bonifaz and Vazquez-Gonzalez 2011; Pihet et al. 2008).
Notes: The morphology of T. concentricum resembles those of the slow-growing species T. verrucosum, T. bullosum (for differentiation see T. bullosum description) and T. schoenleinii. Closely related species from the T. benhamiae clade are easily distinguished from T. concentricum by higher growth rates (Fig. 11) and relatively abundant sporulation. Differentiation from these species is usually not problematic in practice due to the different host spectra and geographic distributions of these species. Only the MAT1-1-1 idiomorph was detected in the T. concentricum isolates examined here; in contrast, isolates of T. verrucosum and T. schoenleinii exclusively show the MAT1-2-1 idiomorph (Kano et al. 2014; Kosanke et al. 2018).
Trichophyton concentricum usually grows as a sterile mycelium in culture; however, the production of clavate microconidia and smooth-walled macoconidia has been observed by some authors (Pihet et al. 2008; Rippon 1988), while favic chandeliers and pectinate hyphae (“antler” tips) are more frequently reported (Bonifaz et al. 2004; Dvořák and Otčenášek 1969). We did not observe these structures in any of the isolates examined.
Trichophyton europaeum Cmokova & Hubka, sp. nov.—MycoBank MB835888; Fig. 19
Macromorphology and micromorphology of Trichophyton europaeum. Colonies after two weeks of cultivation at 25 °C on Sabouraud’s dextrose agar (a, b), malt extract agar (c, d) and potato dextrose agar (e, f). Conidiophores bearing microconidia (g–i); macroconidia (j–m); microconidia (n); spiral hyphae (o, p). Scale bars = 20 μm
Etymolog: Refers to the origin of the examined strains.
Typus: SWITZERLAND, Lausanne, guinea pig (Cavia porcellus), 2008, M. Monod, PRM 944419 (holotype); ex-holotype culture IHEM 22725 (= CCF 6499).
Vegetative hyphae smooth, septate, hyaline, 1–3 µm diam (mean ± sd: 1.9 ± 0.3). Conidiophores branched in a pyramidal (grape-like) pattern or poorly differentiated from the hyphae and represented by conidiogenous hyphae with sparse to numerous short lateral branches. Microconidia abundant, sessile on lateral or terminal branches, pyriform to clavate, 2.5–3.9 (3 ± 0.3) × 1.5–2.8 (2.1 ± 0.2) μm. Macroconidia rare to sparse, born terminally on hyphae, usually consisting of 2–7 cells (median = 4) with an unequal diameter, 45–76 (51.2 ± 7.3) × 3–10.5 (5 ± 1.3) μm, elongated, clavate, less frequently fusiform, with a tapering rounded apex and truncate end, cylindrical fragments of macroconidia common, macroconidia consisting of irregular and bloated cells common. Chlamydospores present. Spiral hyphae absent to rare in 14-days-old cultures, usually consisting of one to several coils. Sexual morph unknown, pseudo-ascomata are formed by some isolates after prolonged incubation.
Culture characteristics (7 days at 25 °C): Colonies on SAB 24–29 mm diam (⌀ = 25 mm), White (#F2F3F4), velvety to floccose, flat, in some strains with radially wrinkled or elevated center, edge filiform, diffuse or entire, reverse brilliant yellow (#FADA5E), to deep orange yellow (#C98500). Colonies on MEA 20–30 mm diam (⌀ = 26 mm), white (#F2F3F4) to light yellow (#F8DE7E), velvety, floccose to coarsely granular, flat with an umbonate center, edge entire to diffuse, reverse in shades of brown [strong orange yellow (#EAA221) to deep orange (#BE6516)] or red [vivid reddish orange (#E25822) to vivid red (#BE0032)]. Colonies on PDA 19–23 mm diam (⌀ = 21 mm), white (#F2F3F4) to light yellow (#F8DE7E), velvety, floccose to coarsely granular, flat with an umbonate center, edge irregular, lobate dendritic, reverse yellow (#F3C300) in the marginal part, strong orange yellow (#EAA221) to deep orange (#F38400) in the center. Colonies at 30 °C in 7 d: SAB 32–37 mm diam (⌀ = 35 mm); PDA 29–31 mm diam (⌀ = 30 mm); MEA 32–39 mm diam (⌀ = 36 mm). Colonies at 37 °C in 7 d: SAB 23–31 mm diam (⌀ = 28 mm); PDA 24–31 mm diam (⌀ = 29 mm); MEA 20–30 mm diam (⌀ = 27 mm).
Material examined: Switzerland, Lausanne, guinea pig (Cavia porcellus), 2008, M. Monod (PRM 944419, holotype, dried culture); ex-holotype culture IHEM 22725 (= CCF 6499). France, Lyon, guinea pig, 1963 (IHEM 25139 = CBS 806.72 = RV 14387 = ATCC 28061 = CCF 6480). Switzerland, Lausanne, human dermatophytosis (contact with guinea pig), 2002, M. Monod (IHEM 20159 = CBS 112370); ibid., IHEM 25062 = CCF 6477. Switzerland, Lausanne, human dermatophytosis (contact with guinea pig), 2007, M. Monod (IHEM 25064 = CCF 6478). Switzerland, Lausanne, tinea corporis (contact with guinea pig), 2010, M. Monod (IHEM 25075). Switzerland, Lausanne, tinea faciei, 2011, Monod (HEM 25076). Switzerland, Lausanne, guinea pig, 2002, M. Monod (IHEM 22723). Czechia, Malhotice, toenail (onychomycosis), 2012, S. Dobiášová (CCF 4917). Czechia, Prague, human dermatophytosis (tinea faciei), 2012, M. Skořepová (CCF 4848). Czechia, Bylany, dermatophytosis in human (tinea corporis), M. Skořepová (CCF 4853). All 40 strains of T. europaeum examined in this study are listed in Table S1.
Distribution and ecology: Trichophyton europaeum is a zoophilic species that is widely distributed in guinea pigs in Europe (Fréalle et al. 2007; Fumeaux et al. 2004; Sabou et al. 2018; Symoens et al. 2013) but is less prevalent than T. benhamiae var. luteum. The species has also been reported from fox in Poland (Ziółkowska et al. 2015), guinea pigs in Japan (Takeda et al. 2012) and human dermatophytosis in Iran (Rezaei-Matehkolaei et al. 2016). Dermatophytosis in horses reported in Egypt is an unusual finding (Tartor et al. 2016).
The European strains of T. europaeum (n = 41) examined here were predominantly obtained from humans (~ 80% from females and ~ 20% from males, median age 12 years) who mostly reported contact with guinea pigs (66%), and the remaining strains were recovered from guinea pigs or dogs (Table S1). The infections mostly manifested as tinea corporis (79%) and tinea faciei (21%). Only the MAT1-2-1 idiomorph was detected in the T. europaeum isolates examined here, with the exception of the IHEM 25139 strain.
Notes: The morphology of T. europaeum most closely resembles those of T. benhamiae var. benhamiae, T. japonicum and T. mentagrophytes. Trichophyton europaeum shares many morphological characteristics with T. japonicum, including the red/brown pigmentation of the colony reverse colour on MEA in some strains, the production of conidiophores branched in a pyramidal pattern and abundant sporulation. The ratio of MAT1-1-1 and MAT1-2-1 strains in the T. europaeum strains examined here was 1:39; by contrast, all T. japonicum strains exhibited only the MAT1-1-1 idiomorph (Figs. 3, 7). These two species can be reliably differentiated only by means of molecular methods (ITS and gapdh gene sequences, microsatellite markers, MALDI-TOF MS). Trichophyton benhamiae var. benhamiae differs from T. europaeum and T. japonicum in its host spectrum, higher growth rates, especially on MEA and PDA at 25 °C (Fig. 11) and macroconidia characteristics. The differentiation of T. mentagrophytes from T. europaeum and T. japonicum is sometimes difficult by morphological methods. In general, the obverse of T. mentagrophytes colonies is more intensively coloured in shades of yellow–brown to brown, and the colony reverse colour is usually dark brown. Trichophyton mentagrophytes isolates usually produce abundant spiral hyphae, which are rather rare in T. europaeum and T. japonicum after 2 weeks. To differentiate T. europaeum from other species, see the descriptions of T. benhamiae var. benhamiae and T. benhamiae var. luteum.
Trichophyton japonicum Cmokova & Hubka, sp. nov.—MycoBank MB835889; Fig. 20
Macromorphology and micromorphology of Trichophyton japonicum. Colonies after two weeks of cultivation at 25 °C on Sabouraud’s dextrose agar (a, b), malt extract agar (c, d) and potato dextrose agar (e, f). Conidiophores bearing microconidia (g–k); macroconidia (l–p); microconidia (r); spiral hyphae (s, t). Scale bars = 20 μm
Etymology: Refers to the origin of the majority of the examined strains.
Typus: SPAIN, human, 1963, P. Miguens, PRM 944416 (holotype); ex-holotype culture IHEM 17701 = ATCC 28063 = CBS 807.72 = CECT 2894 = RV 14988 = CCF 6481.
Vegetative hyphae smooth, septate, hyaline, 1.5–4 µm diam (mean ± sd: 2.5 ± 0.6). Conidiophores usually poorly differentiated from hyphae and represented by conidiogenous hyphae with sparse to numerous short lateral branches; conidiophores branched in a pyramidal (grape-like) pattern relatively rare. Microconidia abundant, born terminally on hyphae, pyriform to clavate, 2.5–5 (3.2 ± 0.4) × 1.5–3.6 (2.3 ± 0.3) µm. Macroconidia rare to abundant, born terminally on hyphae, sparse to abundant depending on the isolate, consisting of 3–8(–12) cells (median = 5), 11–79 (55.2 ± 12.4) × 5–11 (6.8 ± 1.5) μm, elongated, cigar-shaped, clavate, with a tapering rounded apex and truncate end, macroconidia consisting of irregular and bloated cells common, long macroconidia easily disintegrate into cylindrical fragments. Chlamydospores present. Spiral hyphae absent to sparse in 14-days-old colonies. Sexual morph unknown.
Culture characteristics (7 days at 25 °C): Colonies on SAB 16–36 mm diam (⌀ = 23 mm), white (#F2F3F4) to pale yellowish pink (#ECD5C5), velvety to floccose, flat with sligtly elevated and furrowed center, edge entire to diffuse, reverse light orange (#FAB57F) to vivid orange yellow (#F6A600) in the marginal part, in some strains deep orange yellow (#C98500) center. Colonies on MEA 18–30 mm diam (⌀ = 26 mm), white (#F2F3F4), light yellow (#F8DE7E) to pale yellowish pink (#ECD5C5), floccose to granular, flat, sometimes with an umbonate center, frequently with concentric ring pattern, margin entire to diffuse, reverse deep orange (#BE6516), strong reddish brown (#882D17) to vivid red (#BE0032). Colonies on PDA 16–27 mm diam (⌀ = 23 mm), white (#F2F3F4) to pale yellowish pink (#ECD5C5), floccose to granular, occasionally with cottony sectors, flat or umbonate, margin entire, reverse deep orange (#BE6516), strong reddish brown (#882D17) to vivid red (#BE0032). Colonies at 30 °C in 7 d: SAB 32–45 mm diam (⌀ = 38 mm); MEA 28–37 mm diam (⌀ = 33 mm); PDA 26–35 mm diam (⌀ = 30 mm). Colonies at 37 °C in 7 d: SAB 21–38 mm diam (⌀ = 26 mm); MEA 32–37 mm diam (⌀ = 35 mm); PDA 30–35 mm diam (⌀ = 33 mm).
Material examined: Spain, human, 1963, P. Miguens (PRM 944416, holotype, dried culture; PRM 944417, isotype); ex-holotype culture IHEM 17701 = ATCC 28063 = CBS 807.72 = CECT 2894 = RV 14988 = CCF 6481). Belgium, dog, 1971, De Vroey (IHEM 4030 = ATCC 28067 = CBS 809.72 = RV 28105 = CCF 6498). Japan, rabbit, 2009 (NUBS 09011). Japan, Saitama, human, 2000 (VUT 00003–2). Japan, Saitama, rabbit, 1999 (VUT 00002). Japan, Saitama, rabbit, 2000 (VUT 00003). Japan, human, 2013 (NUBS 12001). Japan, Hyogo, rabbit, 1997 (VUT 97010 = CCF 6489). Japan, unknown source (JPN3 = CCF 6487). Japan, unknown source (JPN6). Japan, human, unknown (NUBS 13002 = CCF 6488). Czechia, human, tinea corporis, 2013, N. Mallátová (D 35). Czechia, human, tinea corporis, 2011, S. Dobiášová (DMF 3061). Czechia, human, tinea corporis, 2012, S. Dobiášová (DMF 2446); ibid., DMF 3031. Czechia, human, tinea corporis, 2013, S. Dobiášová (DMF 1658). Czechia, guinea pig (Cavia porcellus), 2014, J. Koubková (KOUB 63). Czechia, Pardubice, human, tinea corporis, 2011, K. Mencl (ME 961). Czechia, Prague, human, tinea corporis, 2012, P. Lysková (PL 1773).
Distribution and ecology: Trichophyton japonicum is a zoophilic species occurring mostly in rabbits and guinea pigs. The species is widely distributed in Japan (mostly in rabbits) (Kimura et al. 2015; Takeda et al. 2012). In Europe it occurs mostly in guinea pigs and less frequently in rabbits and other hosts (Table S10). In guinea pigs it is less common than T. benhamiae var. luteum and T. europaeum (see discussion). It has also been detected in Iran (GenBank JX413540), Oman (Al-Hatmi 2010), Taiwan (Wang and Sun 2018), Thailand (Vu et al. 2019) and South Korea (P.-L. Sun, pers. comm.) (Table S10).
The European and Japan strains of T. japonicum (n = 19) examined here were predominantly obtained from humans (~ 63% from females and ~ 37% from males, median age 15 years) who mostly reported contact with rabbits, guinea pigs and dogs. The remaining strains were recovered from the mentioned animals (Table S1). The infections mostly manifested as tinea corporis (trunk skin 38%, extremities 63%). Only the MAT1-1-1 idiomorph was detected in the T. japonicum isolates examined here.
Notes: For the differentiation of T. japonicum from similar species, see the description of T. europaeum. Only the MAT1-1-1 idiomorph was detected in all examined strains, by contrast all T. europaeum strains exhibited the MAT1-2-1 idiomorph except for strain IHEM 25139.
Trichophyton erinacei clade
Trichophyton erinacei (J.M.B. Sm. & Marples) Quaife, J. Clin. Pathol. 19: 178. 1966—Fig. 21
Macromorphology and micromorphology of Trichophyton erinacei. Colonies after two weeks of cultivation at 25 °C on Sabouraud’s dextrose agar (a, b), malt extract agar (c, d) and potato dextrose agar (e, f). Conidiophores bearing microconidia (g–j) and macroconidia (intermediate forms) (k); macroconidia (l–n); free microconidia and macroconidia (two-celled intermediate forms) (o); intercalary conidia—marked with arrows (p). Scale bars = 20 μm
Vegetative hyphae smooth, septate, hyaline, 1–3 µm diam (mean ± sd: 1.9 ± 0.8). Conidiophores usually poorly differentiated from vegetative hyphae, conidiophores branched in a pyramidal (grape-like) pattern present only in some strains, conidia sessile on hyphae or short lateral and terminal branches. Microconidia abundant, mostly clavate or pyriform, 2.9–6.5 (4.3 ± 2.79) × 1.5–3.5 (2.7 ± 0.28) μm diam. Macroconidia rare to abundant, predominantly consisting of only two or few cells (intermediate forms between micro- and macroconidia), max. 5-celled (median = 2), clavate, cigar-shaped, 6–35 (11 ± 4.52) × 2.5–4.5 (3.4 ± 0.17) μm; intercalary conidia sparse to abundant, cylindrical, barrel-shaped or irregular. Chlamydospores present. Spiral hyphae not observed. Sexual morph unknown.
Culture characteristics (7 days at 25 °C): Colonies on SAB 19–32 mm diam (⌀ = 29 mm), white (#F2F3F4) to light orange yellow (#F3E5AB) in the centre, flat, finely to coarsely granular, edge diffuse, reverse light orange yellow (#F3E5AB) to vivid yellow (#FADA5E), deep reddish brown (#882D17) in the centre. Colonies on MEA 19–30 mm diam (⌀ = 25 mm), white (#F2F3F4), flat, finely to coarsely granular, edge diffuse, reverse light orange yellow (#F3E5AB) to vivid reddish orange (#F38400). Colonies on PDA 11–32 mm diam (⌀ = 24 mm) white (#F2F3F4) to pale orange yellow (#F3E5AB), flat to slightly raised in the center, finely to coarsely granular (velvety to cottony in some strains), edge diffuse (irregular or submerged in some strains), light orange yellow (#F3E5AB) to vivid yellow (#F3C300), frequently deep reddish brown (#882D17) in the centre. Colonies at 30 °C in 7 d: SAB 25–35 mm diam (⌀ = 31 mm); MEA 39–48 mm diam (⌀ = 39 mm); PDA 35–42 mm diam (⌀ = 36 mm). Colonies at 37 °C in 7 d: SAB 25–40 mm diam (⌀ = 34 mm); MEA 30–37 mm diam (⌀ = 32 mm); PDA 32–34 mm diam (⌀ = 33 mm).
Material examined: New Zealand, hedgehog (Erinaceus europaeus), M.J. Marples (ex-holotype culture CBS 511.73 = ATCC 28443 = IMI 101051 = NCPF 375). The Netherlands, Delft (Diagnostic Center SSDZ), arm skin, human, 1979 (CBS 344.79). United Kingdom, Bristol (General Hospital Bristol), human, 1972 (IHEM 19619 = RV 28925); ibid., culture IHEM 19621 = RV 28927.
Distribution and ecology: Trichophyton erinacei (Quaife 1966) is a zoophilic species that is common in wild-living and pet hedgehogs worldwide. The pathogen was originally described in the European hedgehog (Erinaceus europaeus), occurring naturally in the UK and Northern and Western Europe; it has also been imported to New Zealand and Japan (Morris and English 1969; Smith and Marples 1964; Takahashi et al. 2003). The African wild-living four-toed hedgehog (Atelerix albiventris) is another host of T. erinacei. The prevalence of the pathogen is high in both wild-living and pet hedgehogs, resulting in a significant increase in human infections due to T. erinacei, especially those contracted from pet hedgehogs, in recent years (Abarca et al. 2017; Hubka et al. 2018d; Kargl et al. 2018). The presentation in hedgehog range from asymptomatic infection (Fig. 22) to extensive involvement of the body surface. The infection is predominantly located on the head and usually spread slowly (Morris and English 1973; Schauder et al. 2007; Takahashi et al. 2002). In human, extremities are affected in cca 70‒80% of reported cases (Fig. 22), although tinea corporis, barbae, faciei (Fig. 22), capitis and onychomycosis have been also reported (Concha et al. 2012; English et al. 1962; Piérard-Franchimont et al. 2008).
Clinical presentation of infections caused by Trichophyton erinacei clade species in animals and humans. Trichophyton erinacei: four-toed hedgehog (Atelerix albiventris) without apparent clinical signs of infection (a), a source of tinea corporis infection in a pet breeder (Lysková et al. , 2018); wild European hedgehog (Erinaceus europaeus) with facial skin lesion (b), image courtesy of Veronica Risco-Castillo; tinea corporis on the left forearm caused by T. erinacei (c). Trichophyton verrucosum: discrete, scaling patches of hair loss located on the head and neck of cattle (d, e) and goat (f); tinea corporis on the forearm (g), infection that affected scalp skin after previous injury—the situation after surgical removal of necrotic parts (h)
Notes: The morphology of T. erinacei resembles that of T. africanum and T. mentagrophytes. Compared to T. mentagrophytes, with a dark colony reverse colour, the colony reverse colour of T. erinacei is pale. These species also differ in the production of spiral hyphae, which are absent in T. erinacei, and by the general shape of microconidia, which are mostly globose or subglobose in T. mentagrophytes. The species is strongly associated with hedgehogs, and identification is thus only difficult when isolated from infected humans with incomplete anamnestic data. The closely related taxa T. eriotrephon and T. verrucosum are easily distinguishable from T. erinacei by their slower growth rates (Fig. 11) and relatively poor sporulation (sporulation usually absent in T. verrucosum). Additionally, T. eriotrephon can be differentiated from T. erinacei by the production of an intense reddish-brown pigment and microconidia with variable shapes. Only the MAT1-1–2 idiomorph was detected among the T. erinacei isolates examined here.
Trichophyton eriotrephon Papegaay, Ned. Tijdschr. Geneesk. 69: 885. 1925—Fig. 23
Macromorphology and micromorphology of Trichophyton eriotrephon. Colonies after two weeks of cultivation at 25 °C on Sabouraud’s dextrose agar (a, b), malt extract agar (c, d) and potato dextrose agar (e, f). Conidiophores bearing microconidia and intercalary conidia (g–k); microconidia and intercalary conidia with variable shape (l). Scale bars = 20 μm
Vegetative hyphae smooth, septate, hyaline, 1.4–3.2 µm diam (mean ± sd: 2.2 ± 1.0). Well-differentiated conidiophores rare, usually only poorly differentiated from vegetative hyphae, lateral branches arise in a right-angle to the fertile hyphae, fertile hyphae frequently disintegrate into propagules (intercalary conidia and microconidia). Microconidia abundant, sessile, formed terminally or laterally on fertile hyphae, or on lateral branches, occasionally in short chains, variable in shape, mostly ovoid or pyriform, occasionally barrel-shaped, limoniform or irregular, 3.3–6.6 (4.6 ± 0.85) × 2.1–3.7 (3.4 ± 0.41) μm diam; intercalary conidia common, occasionally arranged in chains, barrel-shaped or irregular. Macroconidia absent. Spiral hyphae absent. Chlamydospores common. Sexual morph unknown.
Culture characteristics (7 days at 25 °C): Colonies on SAB 17–25 mm diam (⌀ = 21 mm), white (#F5F5F0) or light yellowish brown (#E3D6A1), flat with radially wrinkled centre, velvety to delicately granular, edge entire, reverse deep reddish brown (#882D17), diffuse pigment strong reddish brown (#6E2615) produced into the medium (less intense in IHEM 24340). Colonies on MEA 17–25 mm diam (⌀ = 21 mm), white (#F5F5F0) to pale yellow (#C2B280) in the centre, flat with or without radially wrinkled centre, velvety to delicately granular, edge submerged and filliform, reverse vivid red (#841B2D) to deep reddish brown (#882D17) (yellow reverse in IHEM 24340). Colonies on PDA 17–24 mm diam (⌀ = 22 mm), white (#F5F5F0) to light yellowish brown (#E3D6A1) in the centre, flat or umbonate, with raised centre (radially wrinkled in CBS 220.25), velvety or downy, edge submerged to filliform, reverse vivid orange (#F38400) to strong yellowish brown (#80461B) in the centre. Colonies at 30 °C in 7 d: SAB 22–32 mm diam (⌀ = 27 mm); MEA 28–31 mm diam (⌀ = 31 mm); PDA 22–25 mm diam (⌀ = 24 mm). Colonies at 37 °C in 7 d: SAB 0–3 mm diam (⌀ = 1 mm); no growth on MEA and PDA.
Material examined: The Netherlands, human dermatophytosis, 1925, J. Papegaay (ex-type culture CBS 220.25). Belgium, Marke, dog skin and hair (Jack Russell terrier), 2010 (IHEM 24340).
Distribution and ecology: Insufficient data are available regarding the distribution of T. eriotrephon, which is known from four cases of dermatophytosis in humans (tinea corporis, Netherlands; tinea manuum and tinea faciei, Iran; tinea barbae, France) (Papegaay 1925; Rezaei-Matehkolaei et al. 2013; Sabou et al. 2018) and a dog (isolate IHEM 24340 from Belgium). It is assumed that T. eriotrephon is a zoophilic species based on its phylogenetic relationships with other zoophilic species and the clinical manifestations of known infections in humans.
Notes: The morphology of T. eriotrephon only slightly resembles species from the T. benhamiae clade in its red-brown colony reverse colour. The conidiophores of T. eriotrephon are mostly loose and poorly branched compared to those of zoophilic species from the T. benhamiae clade, with grape-like conidiophores. Other typical characteristics include the production of a diffuse red-brown pigment on SAB, microconidia with variable shapes and the absence of macroconidia. These characteristics, together with the absence of or restricted growth at 37 °C, differentiate T. eriotrephon from all other species of the T. benhamiae complex. The MAT1-1-1 idiomorph of the mating type gene was detected in both T. eriotrephon isolates examined here.
Trichophyton verrucosum E. Bodin, Les champignons parasites de l'homme: 121. 1902—Fig. 24
Macromorphology and micromorphology of Trichophyton verrucosum. Colonies after three weeks of cultivation at 25 °C on Sabouraud’s dextrose agar (a, b), malt extract agar (c, d) and potato dextrose agar (e, f). Clumps of vegetative hyphae (g); chlamydospores in chains (h, i); macroconidia (j–m); conidiophores (fertile hyphae) with sessile microconidia (n, o); microconidia (p). Scale bars = 20 μm
Vegetative hyphae smooth, septate, frequently inflated, hyaline, 1–2.5 µm diam (mean ± sd: 1.7 ± 1.16). Conidiophores rare, poorly differentiated from vegetative hyphae, unbranched or sparsely branched, conidia sessile on hyphae or born on short lateral branches. Microconidia absent or rare, clavate, 3–6 (4.5 ± 0.7) × 1.9–3.5 (2.9 ± 0.45) μm. Macroconidia absent or rare, smooth-walled, clavate or fusiform with rounded apex and truncate end, usually consisting of 1–4 cells (median = 2), 16–50 × 4–8 μm. Chlamydospores abundant and frequently in the form of chains. Spiral hyphae absent. Sexual morph unknown.
Culture characteristics (7 days at 25 °C): Colonies on SAB 18–22 mm diam (⌀ = 20 mm), white (#F5F5F0) to pale orange yellow (#FFF587) or light orange yellow (#FAD6A5), flat, raised and furrowed, or cerebriform, velvety to slightly powdered, edge entire, lobate, or submerse, reverse light orange yellow (#F8DE7E) to deep orange yellow (#C9AE5D), dark brown in some strains. Colonies on MEA 5–18 mm diam (⌀ = 13 mm), white (#F5F5F0) to pale orange yellow (#F3E5AB), raised in the centre, frequently wrinkled, velvety or waxy, edge entire, lobate, or submerse, reverse light orange yellow (#F8DE7E) to vivid orange yellow (#F6A600), dark brown in some strains. Colonies on PDA 8–18 mm diam (⌀ = 14 mm), white (#F5F5F0) to pale orange yellow (#F3E5AB), flat or with raised centre, velvety or waxy, edge entire, lobate, or submerse, reverse light orange yellow (#F8DE7E), dark brown in some strains. Colonies at 30 °C in 7 d: SAB 10–23 mm diam (⌀ = 18 mm); MEA 8–10 mm diam (⌀ = 9 mm); PDA 9–10 mm diam (⌀ = 9 mm). Colonies at 37 °C in 7 d: SAB 9–15 mm diam (⌀ = 10 mm); MEA 11–12 mm diam (⌀ = 11 mm); PDA 12–14 mm diam (⌀ = 13 mm).
Material examined: Unknown locality, cow, 1953, F. Blank (ex-neotype culture CBS 365.53). Czechia, Pardubice, dermatophytosis in 21-year-old woman (contact with cattle), 2011, K. Mencl (CCF 4612). Czechia, Hlinsko, dermatophytosis in 58-year-old woman (contact with cattle), 2011, K. Mencl (CCF 4613). Czechia, Tábor, dermatophytosis in 38-year-old woman (contact with cattle), 2014, N. Mallátová (CCF 4889).
Distribution and ecology: Trichopyhton verrusocum is a zoophilic species typically found in cattle and other ruminants (Fig. 22), but it can easily spread to humans and animals, including horses, donkeys, camels, rabbits, dogs, cats, pigs, and even birds (Ali-Shtayeh et al. 1988; Dvořák et al. 1965; Georg 1960; Chermette et al. 2008; Khosravi and Mahmoudi 2003). The species is distributed worldwide, but the incidence of infections in cattle and man has been decreased in many regions by specific preventive measures, especially by vaccination programmes or changes in agricultural systems, such as reduction of the number of cattle in breeding units, and infections in humans have decreased proportionally (Lund et al. 2014; Seebacher et al. 2008). Human patients usually develop aggressive inflammatory skin lesions usually located on extremities and head (Fig. 22), which may be accompanied by constitutional symptoms, such as fever and lymphadenopathy (Courtellemont et al. 2017; Silver et al. 2008). Tinea barbae and capitis are relatively common clinical forms which can result in irreversible scarring and alopecia.
Notes: The morphology of T. verrucosum resembles that of T. bullosum and T. concentricum. For distinguishing characters see T. bullosum description. Only MAT1-2-1 idiomorph was detected in all strains examined here and in all strains analyzed by other reserchers (Kano et al. 2014; Kosanke et al. 2018).
Trichophyton bullosum clade
Trichophyton africanum Cmokova & Hubka, sp. nov.—MycoBank MB835890; Fig. 25
Macromorphology and micromorphology of Trichophyton africanum. Colonies after two weeks of cultivation at 25 °C on Sabouraud’s dextrose agar (a, b), malt extract agar (c, d) and potato dextrose agar (e, f). Conidiophores bearing microconidia (g–k); macroconidia (l–n); microconidia (o). Scale bars = 20 μm
Etymology: Refers to the origin of the ex-type strain.
Typus: MOZAMBIQUE, human, 1969, M.J. Campos-Magalhaes, PRM 944418 (holotype); ex-holotype culture IHEM 4032 (= ATCC 28064 = RV 25293 = CM 3440 = CCF 6493).
Vegetative hyphae smooth, septate, hyaline, 1–4 µm diam (mean ± sd: 2.2 ± 0.5). Conidiophores poorly differentiated from vegetative hyphae, unbranched or sparsely branched, conidia sessile on lateral or terminal branches. Microconidia abundant, pyriform to clavate, 2.5–5 (4 ± 0.5) × 1.9–2.9 (2.4 ± 0.3) µm. Macroconidia rare to sparse, cigar-shaped, 14–80.5 (64.2 ± 14.4) × 6–11 (8.2 ± 1.2) μm, consisting of 3–9(–13) cells (median = 6). Chlamydospores present. Spiral hyphae rare or absent. Sexual morph unknown.
Culture characteristics (7 days at 25 °C): Colonies on SAB 30–32 mm diam (⌀ = 31 mm), white (#F2F3F4) to pale yellow green (#F2F3E5), granular, slightly raised in the center, margin diffuse, reverse light orange yellow (#FBC97F) in the marginal part, strong orange (#ED872D) in the center. Colonies on MEA 28–35 mm diam (⌀ = 30 mm), white (#F2F3F4), granular, flat, margin entire, reverse uncoloured to light orange yellow (#FBC97F). Colonies on PDA 27–28 mm diam (⌀ = 28 mm), white (#F2F3F4) to pale yellow green (#F2F3E5), granular to floccose, slightly raised in the center, margin entire, reverse pale yellow (#F3E5AB) to pale orange yellow (#FAD6A5) in the marginal part, dark orange yellow (#BE8A3D) in the center. Colonies at 30 °C in 7 d: SAB 40–45 mm diam (⌀ = 43 mm); MEA 35–45 mm diam (⌀ = 39 mm); PDA 35–40 mm diam (⌀ = 36 mm). Colonies at 37 °C in 7 d: SAB 21–24 mm diam (⌀ = 24 mm); MEA 20–29 mm diam (⌀ = 23 mm); PDA 20–22 mm diam (⌀ = 21 mm).
Material examined: Mozambique, human, 1969, M.J. Campos-Magalhaes (PRM 944418, holotype, dried culture); ex-holotype culture IHEM 4032 = ATCC 28064 = RV 25293 = CM 3440 = CCF 6493). Belgium, Bruges, human fingernail, 1978 (IHEM 19628 = RV 40614). South Africa, human skin, 1971, K. Scott (IHEM 4033 = ATCC 28065 = CBS 808.72 = CECT 2895 = NCPF 456 = RV 27926).
Distribution and ecology: All three currently known strains are of human origin, but the low number of isolates does not allow us to draw conclusions about their ecology. The species probably occurs mainly in Africa.
Notes: The morphology of T. africanum may resemble zoophilic T. benhamiae clade species, T. erinacei or T. mentagrophytes sensu de Hoog et al. (2017). Trichophyton africanum shows a uncoloured or pale reverse on MEA, differing from the intense yellow or red/brown pigments typical of T. benhamiae clade species and T. mentagrophytes. The conidiophores of T. africanum are unbranched or sparsely branched; when branched, the resulting conidiophores have usually only few and relatively long lateral branches and are less compact than those of T. benhamiae var. luteum, T. europaeum and T. japonicum (pyramidal/grape-like with many short lateral branches). Trichophyton africanum has conidia of similar lengths to those of T. benhamiae var. benhamiae and in average longer than those of the remaining species from the T. benhamiae clade. The differentiation of this species from T. erinacei on the basis of morphology may be difficult, but T. erinacei is very strongly associated with hedgehogs. The most closely related species, T. bullosum, can be easily distinguished by its very slow growth, poor or absent sporulation, and abundant production of chlamydospores. The ratio of MAT1-1-1 and MAT1-2-1 strains in T. africanum was 2:1.
Trichophyton bullosum Lebasque, Les Champignons des Teignes du Cheval et des Bovidés: 53. 1933—Fig. 26
Macromorphology and micromorphology of Trichophyton bullosum. Colonies after three weeks of cultivation at 25 °C on Sabouraud’s dextrose agar (a, b), malt extract agar (c, d) and potato dextrose agar (e, f). Detail of colony with submerged, dendritic growth on Sabouraud’s dextrose agar supplemented with cycloheximide and chloramphenicol after 3 months of cultivation at 25 °C (g). Thick-walled vegetative hyphae with numerous intercalary or terminal chlamydospores (h, i); chlamydospores in chains and free chlamydospores (j), vegetative hyphae with terminal chlamydospores (k, l). Scale bars = 20 μm
Vegetative hyphae smooth, septate, inflated, often branched and with knob-like terminations, hyaline 1.5–4 µm diam (mean ± sd; 2.7 ± 0.7). Chlamydospores abundant, spherical, oval or irregular, occasionally in chains, 4–9(–20) µm in diam. Microconidia and macroconidia not observed in the isolates examined in this study, but they were observed by Lebasque (1933) under specific conditions. Spiral hyphae absent.
Culture characteristics (7 days at 25 °C): Colonies on SAB 11–12 mm diam (⌀ = 12 mm), white (#F2F3F4) to pale yellowish pink (#ECD5C5) or pale orange yellow (#FAD6A5), umbonate, radially furrowed, membranous or slightly velvety, edge submerged or filiform, reverse light yellow (#F8DE7E). Colonies on MEA 8–12 mm diam (⌀ = 10 mm), White (#F2F3F4), pale yellow (#F3E5AB) or vivid orange yellow (#F6A600), flat with raised and cerebriform center, membranous, edge entire or submerged with dendritic growth, reverse light yellow (#F8DE7E). Colonies on PDA 7–9 mm v diam (⌀ = 8 mm), white (#F2F3F4), pale yellowish pink (#ECD5C5) or pale orange yellow (#FAD6A5), circular, flat, umbonate, membranous, edge entire, reverse light yellow (#F8DE7E). Colonies at 30 °C in 7 d: SAB 11–14 mm diam (⌀ = 13 mm); MEA 11–12 mm diam (⌀ = 11 mm); PDA 12–13 mm diam (⌀ = 12 mm). Colonies at 37 °C in 7 d: SAB 8–10 mm diam (⌀ = 9 mm); MEA 9–10 mm diam (⌀ = 10 mm); PDA 9–10 mm diam (⌀ = 9 mm).
Material examined: France, horse, J. Lebasque (ex-type culture, CBS 363.35 = LP 770). Czechia, skin lesions in horse, 2013, P. Lysková (CCF 4831). Egypt, near Cairo, skin lesion in donkey (Equus asinus), 2015, A. Peano (CCF 5730).
Distribution and ecology: Trichophyton bullosum is a zoophilic species known from infections in donkeys and horses (Fig. 27). It is distributed in Europe, North Africa and the Middle East (Lebasque 1933; Lysková et al. 2015; Sabou et al. 2018; Sitterle et al. 2012).
Clinical presentation of infections caused by Trichophyton bullosum: patches of hair loss in the saddle area, shoulders, hip bones, withers and upper chest of a horse (a), isolate CCF 4831 (Lysková et al. 2015); scaling patches of hair loss located on the head, chest and legs of a donkey (b–e), isolate CCF 5730
Notes: Due to its slow grow rate, T. bullosum strongly resembles T. verrucosum and T. concentricum. These species either do not sporulate or sporulate poorly (especially on sugar-rich media such as SAB) but produce abundant chlamydospores, frequently in the form of chains. All mentioned species are relatively strongly associated with their hosts and/or with a typical clinical manifestation (cattle ringworm caused by T. verrucosum; dermatophytosis caused by T. bullosum in horses and donkeys; tinea imbricata caused by T. concentricum in humans). Therefore, detailed anamnestic data can facilitate their identification. Molecular genetic methods may be necessary to verify the identification of some isolates. For the differentiation of T. bullosum from the most closely related species, T. africanum, see the description of T. africanum. Only the MAT1-1-1 idiomorph was detected among the T. bullosum isolates examined here.
Discussion
Species delimitation issues in Trichophyton
Species delimitation in dermatophytes is based on a polyphasic approach (Gräser et al. 2008) combining ecological (distribution, host range) and clinical data, the analysis of DNA sequence data, the macro- and micromorphological examination of cultures, physiological and biochemical tests and mating tests. However, the application of the individual components of this concept is limited in many species complexes due to specific problems. As a result, the “polyphasic” approach is commonly applied in a restricted form in practice.
Phenotypic criteria are usually relatively effective in routine diagnostics for major dermatophyte species or species complexes. However, as in other fungi, we have found similarities between species or morphotypes across unrelated dermatophytes, resulting in misdiagnosis in practice (Lysková et al. 2015; Summerbell 2011; Uhrlaß et al. 2018). There is also considerable intraspecific phenotypic variability in other species or species complexes that is not correlated with molecular taxonomy (Heidemann et al. 2010; Kandemir et al. 2020; Su et al. 2019). Moreover, the success rate of phenotypic identification frequently depends on the age of isolates because of the rapid degeneration of important portions of cultures (de Hoog et al. 2017). Consequently, it can be difficult to maintain and reproduce phenotypic characters over decades for the purposes of taxonomic studies.
The high level of clonality in many primary pathogenic dermatophytes with a presumed recent origin is also associated with an extremely low level of genetic intraspecific variability. Consequently, there is a lack of sufficiently variable DNA sequence markers for the differentiation of some species and, therefore, ambiguities in the definition of their boundaries (de Hoog et al. 2017). Phenomena such as incomplete lineage sorting or occasional hybridization and introgression may further complicate the species delimitation of evolutionarily recently diverged species with semi-permeable reproductive barriers (Matute and Sepúlveda 2019; Steenkamp et al. 2018; Taylor et al. 2015). The divergence between these young species may be hidden when using some classical protein-coding phylogenetic markers. Neutrally evolving or noncoding DNA regions, such as microsatellites, introns and intergenic spacers, which accumulate mutations more rapidly, were shown to reveal the evolutionary trajectories of primary pathogenic dermatophytes with higher success (Gräser et al. 2008; Hubka et al. 2018d; Mochizuki et al. 2017).
The specific problems in species delimitation in Trichophyton can be demonstrated by the example of the T. mentagrophytes and Trichophyton rubrum complexes. It was generally assumed that the differentiation of zoophilic T. equinum (main host = horse) from closely related anthropophilic T. tonsurans would be possible based on the ecological preferences, nutritional requirements, and MAT gene idiomorphs (Summerbell et al. 2007; Woodgyer 2004). Kandemir et al. (2020) examined 67 isolates and found that none of the five selected phylogenetic markers were able to unambiguously separate these species (probably due to incomplete lineage sorting) according to differences in their MAT genes, ecology and nicotinic acid requirements. It is postulated that these species evolved very recently and that the speciation process might not yet be complete (Kandemir et al. 2020). Another taxonomically problematic species pair is T. mentagrophytes/T. interdigitale. According to the traditional concept promoted by de Hoog et al. (2017), T. mentagrophytes is a zoophilic species in which both MAT idiomorphs are present in the population, resulting in relatively high intraspecific genetic variability. By contrast, anthropophilic T. interdigitale is a clonal lineage (consisting only of the MAT1-1-1 idiomorph) that is almost exclusively associated with onychomycosis and tinea pedis. Although the correlation between the genotype and the clinical manifestation or source of isolates has been repeatedly demonstrated, the correlation between ITS genotype and phenotype is relatively poor (Dhib et al. 2017; Heidemann et al. 2010; Pchelin et al. 2016). Currently, the molecular diagnosis of these species is mostly based on several unique sites in the ITS region, and phylogenies usually resolve T. mentagrophytes as para- or polyphyletic with T. interdigitale (Hainsworth et al. 2020; Heidemann et al. 2010; Nenoff et al. 2019; Pchelin et al. 2019; Singh et al. 2019; Taghipour et al. 2019). Both species names remain in use, due to the epidemiological consequences associated with different sources of infections in particular. The laboratory diagnosis of T. mentagrophytes and T. interdigitale and that of T. equinum and T. tonsurans are further complicated by inaccurate or even impossible species differentiation using MALDI-TOF MS (da Cunha et al. 2018; Dukik et al. 2018; Hedayati et al. 2019; Nenoff et al. 2013; Suh et al. 2018).
Very similar species delimitation issues complicate the taxonomy of the anthropophilic T. rubrum complex, encompassing clonal lineages showing differences in their distribution and the clinical manifestation of associated infections (de Hoog et al. 2017; Gräser et al. 2000). The majority of molecular studies relying on the variability in the ITS region and microsatellite markers have revealed some support for 2–4 lineages (i.e., T. rubrum, T. violaceum and/or T. soudanense and/or T. yaoundei), but the number of species and their boundaries are still under debate (Gräser et al. 2007; Packeu et al. 2020; Su et al. 2019). MALDI-TOF MS showed promising results in the differentiation of these species/lineages (Packeu et al. 2020).
Detailed genomic, epigenetic and multigene phylogenetic studies on a large number of samples can resolve delimitation issues between these species in the future (Pchelin et al. 2019; Singh et al. 2019; Zhan et al. 2018).
SNP detection by whole-genome sequence typing can be used to infer the genetic relatedness of Trichophyton isolates. This approach will ultimately become one of the methods of choice in the future with decreasing costs (Hadrich and Ranque 2015). Currently, the sequencing of ITS rDNA and population genetic markers such as microsatellites (Gräser et al. 2007; Kaszubiak et al. 2004; Pasquetti et al. 2013) or mixed-marker approaches (Abdel-Rahman et al. 2010) offers higher discriminatory power in the species differentiation of primary pathogenic dermatophytes compared to MLST approaches based on the currently available loci.
Disentangling the taxonomy of the T. benhamiae complex based on a polyphasic approach
In this study, we encountered similar problems to those mentioned in the previous section in the T. mentagrophytes and T. rubrum complexes. However, a polyphasic approach combining independent molecular genetic markers (four DNA loci and 10 microsatellite loci) with phenotypic features and ecological data helped to overcome the majority of obstacles to species delimitation. Selected characteristics are schematically summarized in Fig. 28. Various forms of polyphasic approaches integrating different types of data, including morphological, physiological, exometabolite, ecological and molecular are increasingly used in many fungal order, e.g. Eurotiales, Pleosporales and Xylariales (Bhunjun et al. 2020; Houbraken et al. 2020; Kuhnert et al. 2017; Lambert et al. 2019; Samarakoon et al. 2020). These complex species delimitation approaches are basis for stable taxonomy that is less prone to errors compared to less robust approaches. Molecular data from gradually increasing number of non-linked loci become a regular part of these approaches which prevents the impact of paralogous genes, incomplete lineage sorting, non-reciprocal monophyly and other phenomena on taxonomic conclusions (Hubka and Kolařík 2012; Matute and Sepúlveda 2019; Stadler et al. 2020; Steenkamp et al. 2018).
Overview of selected data on ecology, phenotype and population genetics plotted on the simplified four-gene phylogeny of the Trichophyton benhamiae species complex. The icons of the hosts are explained in Fig. S1. Explanation of superscript numbers: 1macroconidia observed by some authors (Rippon 1988; Pihet et al. 2008), 2macroconidia were observed by Lebasque (1933) under specific conditions
In this study, we showed that isolates that were designated in the past as the Americano-European race of T. benhamiae harbour five taxa (three species and two varieties). The strains with the so-called white phenotype do not represent monophyletic entities and correspond to T. benhamiae var. benhamiae, T. japonicum and T. europaeum, while the yellow phenotype strains correspond to T. benhamiae var. luteum. Isolates of the African race of T. benhamiae referred to as T. africanum herein are phylogenetically distant and are most closely related to T. bullosum.
None of the four sequence markers alone was able to unequivocally differentiate all species within the T. benhamiae complex and provide accurate identification in 100% of cases. The ITS region contained a diagnostic position for all nine species, but the differentiation of T. europaeum and T. japonicum relied on a single substitution (Table S11). In addition, the identification of isolate IHEM 25139, with a probable hybrid origin, failed as described above. The gapdh gene was useful for differentiation between T. europaeum and T. japonicum, but some pairs of sister species shared identical sequences (i.e., T. benhamiae and T. concentricum, T. verrucosum and T. eriotrephon) (Table S11). The tef1-α gene differentiated all species except for T. europaeum and T. japonicum (Table S11). The tubb gene presented the least discriminatory power and failed to differentiate species within the T. benhamiae clade but could be used for species identification in the T. erinacei and T. bullosum clades. Insufficient discriminatory power of the tubb gene has been reported in many other Trichophyton species (Kandemir et al. 2020; Packeu et al. 2020; Suh et al. 2018). The unique substitutions observed within the DNA loci of the T. benhamiae clade species will be the basis for reliable species identification in practice (Table S11). The taxonomic significance of these unique sites is unambiguous, as they correspond to independent microsatellite markers and phenotypic and ecological data, indicating the reproductive isolation of recognized taxa.
While sequence markers were shown to be useful for the diagnosis of T. benhamiae complex species, they were not able to distinguish the two varieties of T. benhamiae. The only intraspecific variation was a single substitution in the tef1-α gene. This substitution was able to differentiate all T. benhamiae var. luteum isolates from the majority of T. benhamiae var. benhamiae strains, with the exception of two isolates from cluster C2, probably due to incomplete lineage sorting between these recently diverged lineages. The results of other analyses clearly indicated that T. benhamiae var. luteum is an emerging entity that is distinct both qualitatively (at the population genetic level and according to phenotypic differences) and ecologically (showing different hosts and distributions). The differentiation of this taxon has clinical relevance, due to which we decided to reassign it as variety of the nearest recombining ancestor, T. benhamiae var. benhamiae. We chose this conservative approach rather than the proposals of a new species because of the impossibility of distinguishing this entity using available DNA sequence markers, as is the current standard in fungal taxonomy.
In contrast to DNA sequence data, population genetic analysis based on the newly developed microsatellite typing scheme clearly separated all species in the T. benhamiae clade (Figs. 5, 6), including T. benhamiae var. benhamiae and T. benhamiae var. luteum. Similarly, pilot MALDI-TOF MS analysis was able to identify specific peaks for all species and varieties in the T. benhamiae clade, suggesting that this increasingly popular method can be used for species identification in clinical practice, but the analysis of additional isolates will be needed to generate a more robust database and confirm our preliminary observations.
Phenotypic and ecological data added another important piece to the taxonomic puzzle. Trichophyton benhamiae var. luteum can be identified by its slow growth on all media at all temperatures and its uniform phenotype (yellow reverse side of colonies and absence of macroconidia; all strains exhibit only mating type MAT1-1-1). The closely related T. benhamiae var. benhamiae is only found in the USA (mostly dogs) and exhibits strikingly different colonies with a brown to red-brown reverse side, macroconidium production and larger microconidia than T. benhamiae var. luteum. This variety shows the most rapid growth among the species from the T. benhamiae clade; isolates with both MAT gene idiomorphs were detected among the examined strains. Trichophyton europaeum is the second most common species from the T. benhamiae complex occurring in Europe and is responsible for human and guinea pig infections. While T. japonicum is currently responsible for the majority of human and animal (rabbits and guinea pigs) infections in Japan, it also occurs in Europe at low frequencies. Reliable differentiation of these species is only possible by molecular methods. Trichophyton japonicum and T. europaeum differ strikingly in the distribution of mating type genes in their populations. Detailed distinguishing characteristics of particular species are listed in the Taxonomy section, and proposed identification procedure in clinical practice is illustrated in Fig. 29.
Speciation through the geographic expansion, host jump and extinction of opposite mating type partners
The assessment of species boundaries via mating experiments (revealing biological compatibility) played an important role in the delimitation of many early species and the discovery of their sexual states. This approach based in principle on the biological species concept (BSC) is generally highly applicable in geophilic dermatophytes (Dawson and Gentles 1962; Hubka et al. 2015; Choi et al. 2012; Padhye and Carmichael 1972; Stockdale 1964). By contrast, the results of biological compatibility assessment can considerably disagree with the concept of classical species of anthropophilic and zoophilic dermatophytes. These species are evolutionarily young, and their phylogenetic divergence preceded the development of reproductive barriers, as demonstrated by interspecific hybrid induction between various primary pathogenic Trichophyton species in vitro (Anzawa et al. 2010; Kawasaki 2011; Kawasaki et al. 2009, 2010). However, it is highly unlikely that this kind of hybridization occurs naturally due to the different ecological niches of species, and the results of in vitro mating assays therefore cannot be extrapolated to a natural scenario. Additionally, the ratio of mating-type gene idiomorph is usually extremely imbalanced or one idiomorph is missing in the majority of anthropo- and zoophilic dermatophytes (Kosanke et al. 2018; Metin and Heitman 2017). This fact further limits or even prevents the possibility of using BSCs in the delimitation of these species. A similar phenomenon was observed by our group in almost all species from the T. benhamiae complex (Fig. 28), suggesting that the loss of opposite mating-type partners was an important driver of their evolution. The ancestors of many currently recognized pathogenic dermatophytes were likely sexually reproducing geophilic species and zoophilic species on free-living mammals (sexually reproducing, e.g., in soil surrounding burrows) with balanced ratios of opposite mating type individuals (Gräser et al. 2006; Summerbell 2011). Introduction to new areas and/or adaptation to a new host is probably a unique event in the evolution of many anthropo- and zoophilic dermatophytes, resulting in the extinction of one mating partner in the whole population of these species. Only some “clonal” offshoots of ancestral sexual dermatophytes probably maintain ongoing populations and follow independent evolutionary trajectories towards speciation (Gräser et al. 2006). Recent outbreak of T. benhamiae var. luteum follows the scenario reported in vertebrate pathogens Batrachochytrium dendrobatidis (James et al. 2009) and Pseudogymnoascus destructans (Trivedi et al. 2017). In that fungi, a single clone of one mating type migrated to new areas, meeting a naive host, what resulted in the high virulence, epidemic spread and formation of a clonal population. Alternatively, the extinction of one MAT gene in a population of dermatophytes may be caused by the preferential spread of strain(s) exhibiting an advantageous combination of alleles associated with higher virulence/transmission potential. Such a successful genotype may be significantly dominant in conditions with almost exclusive asexual transmission and may displace other genotypes. Such a situation is very likely to lead to an imbalance in the MAT gene ratio or even the loss of one MAT gene in the population. The extinction of strains belonging to one mating type is, for instance, observed in some populations of M. canis (Sharma et al. 2007), and different levels of virulence linked with mating-type idiomorphs have been repeatedly documented in fungal pathogens (Chang et al. 2000; Cheema and Christians 2011; Yue et al. 1999).
In the T. benhamiae clade, clonal reproduction is the dominant mode of dissemination (Dg, H, DW indices), and recombination is rare or absent in almost all populations according to the IA. Despite the fact that only MAT1-2-1 idiomorph strains were present within T. europaeum strains, the null hypothesis of random mating was not rejected (Table S6, Fig. 9), suggesting the existence of recent recombination events in this species. As T. japonicum and T. europaeum consist of a single mating type and no recent recombination or gene flow has occurred between them, they should be conceptualized as separate, albeit clonal species, despite potential in vitro interbreeding (Gräser et al. 2006; Summerbell 2002). The disruption of gene flow between T. benhamiae clade species was reflected in high number of fixed alleles (FST or GST indexes) (Table S5, Table S6) indicating reproductive isolation despite overlapping hosts (e.g. guinea pigs) and geographic distributions. This could be caused by pre- or postzygotic reproductive barriers, or absence of terrestrial reservoir for sexual reproduction.
In the T. benhamiae complex, there are at least two possible sexual ancestors of “clonal” species: T. benhamiae var. benhamiae and T. africanum, based on the presence of both MAT gene idiomorphs. While the ecology of T. africanum is poorly known, reservoirs of T. benhamiae var. benhamiae exist in free-living animals. It has been detected in the North American porcupine (Needle et al. 2019; Takahashi et al. 2008), but its host spectrum can be broader and may include members of family Canidae, as evidenced by repeated isolation from dogs (Ajello and Cheng 1967; Sieklucki et al. 2014) and patients who have come into contact with foxes (Tan et al. 2020). Due to close phylogenetic proximity, Trichophyton benhamiae var. benhamiae was very likely a common ancestor of at least some taxa in the T. benhamiae clade, especially anthropophilic T. concentricum (only MAT1-1-1) and zoophilic T. benhamiae var. luteum (only MAT1-1-1). The low genetic diversity within T. benhamiae var. luteum together with its recent origin (according to the DW index) may indicate a founder effect in the recent past. This may suggest that the origin of T. benhamiae var. luteum lies in North America and that one or a few strains were recently introduced to Europe.
The only exception among the examined isolates was strain IHEM 25139, isolated in 1963 by M. Takashio from guinea pig in France. This strain, identified here as T. europaeum based on the gapdh gene, shared some microsatellite alleles with T. japonicum. It also presented the MAT1-1-1 idiomorph of the MAT gene, typical of T. japonicum or T. benhamiae var. benhamiae cluster C3, and an atypical ITS1 region sequence with six substitutions compared to other T. europaeum strains, some of which are at positions crucial for the differentiation of T. benhamiae clade species (Fig. S2). It is possible that this strain originated from hybridization between T. europaeum and T. japonicum. The ecological niches of these species partially overlap, as they both occur in guinea pigs and some other animals that are frequently maintained together. In addition, the coinfection of guinea pigs with two species or morphotypes has been repeatedly documented (Bartosch et al. 2019; Kupsch et al. 2017). In such cases, the exchange of genetic information may likely occur not only through hybridization during saprophytic growth outside the host (possibly followed by introgressive hybridization) but also during coinfection of the same host through a parasexual cycle (anastomosis of hyphae, mitotic crossing-over and haploidization). Another strain with an ITS sequence identical to IHEM 25139 is IHEM 19622 (= RV 14389), which was not examined by our group (GenBank MK298816). These two strains with identical provenance were noted by Takashio to be atypical compared to other examined A. benhamiae isolates because of the less compact texture of their colonies (Takashio 1974). These strains represent unique material for studying natural hybridization in dermatophytes. Their origin and genomic arrangement remain to be elucidated by genomic studies. The absence of these genotypes among the more recently isolated strains examined here and by others (no additional occurrence in GenBank) suggests that they were replaced by more successful genotypes.
Geographical distribution of T. benhamiae clade species
To understand the global distribution of the newly reassigned species in the T. benhamiae clade, we analysed 255 ITS rDNA sequences deposited in GenBank. The analysis enabled the identification of these records to the species level based on the species-specific substitutions in the ITS region (Table S10). This fact further supported the feasibility of the novel taxonomic classification proposed here. The ecological data resulting from the analysis were used as a basis for mapping the distribution of T. benhamiae clade species (Fig. 30; Table S10). The main limitations are the unavailability of epidemiological and DNA data from America, many Asian countries and Africa. As a result, the majority of analysed ITS sequences are from European countries and Japan, where dermatophyte research has a long tradition, and DNA-based identification is more commonly used. Additionally, it is not possible to distinguish two varieties of T. benhamiae based on the ITS region, but macro- and micromorphological characters described in some studies enable clear distinction of the varieties; the variety characteristics described below refer to such cases.
In Europe, guinea pigs are hosts of all three pathogens, among which T. benhamiae var. luteum is the most prevalent, followed by T. europaeum and T. japonicum. The ITS-based identification of 30 T. benhamiae strains from guinea pigs from a single veterinary institution in Prague (Czech Republic) between 2014 and 2019 revealed a 24:4:2 ratio of these pathogens (Hubka and Prausová, unpubl. data). The corresponding ratio of these pathogens in human Czech patients is very similar, ~ 27:5:1 (Hubka et al. 2018b, Hubka et al. unpubl. data; Hubka et al. 2014). In addition to guinea pigs, another important reservoir of T. japonicum are rabbits, while other animal hosts of T. benhamiae clade members seem to be much less important.
Based on current knowledge, white-phenotype strains (T. europaeum and T. japonicum) occurred in Europe before epidemic spread of yellow phenotype strains (T. benhamiae var. luteum). The oldest European strains representing T. japonicum are IHEM 4030 (collected before 1988 in Belgium) and IHEM 17701 (collected before 1997 in Spain). The oldest strains representing T. europaeum were collected before 1988 in Finland (Aho 1980) (Table S10) and more recently from Switzerland, in 2002 (IHEM 20159, IHEM 20161, IHEM 20162, IHEM 20163). The identity of other old white-phenotype strains reported in various European countries from the 1960s to 2000 (Fig. 1) is unclear due to the unavailability of isolates and/or sequence data. Both T. japonicum and T. europaeum were subsequently detected in Japan and some other countries (Fig. 30, Table S10). Outbreaks of infections caused by T. benhamiae var. luteum now seem to be limited to Europe, but an increasing number of infections can be expected in non-European countries due to its recent introduction to other continents (de Freitas et al. 2019; Hiruma et al. 2015).
Zoophilic T. benhamiae clade members have probably been brought into Japan with imported animals on several occasions and spread in Japan by the transportation of animals by breeders or pet shops, as suggested in a series of publications (Hiruma et al. 2015; Kano et al. 1998; Mochizuki et al. 2001; Takeda et al. 2012). The most prevalent species in Japan and South Korea (Jun et al. 2004; Lee et al. 2018, and pers. comm. with PL Sun) is T. japonicum. Other species are probably much less common: T. europaeum has been detected in guinea pig (unknown year of isolation) (Takeda et al. 2012), T. benhamiae var. benhamiae was imported to a Japanese zoo from Canada and the USA (in 2000 and 2002) with North American Porcupines (Takahashi et al. 2008), and T. benhamiae var. luteum was detected in 2012 in common degu (Hiruma et al. 2015).
In addition to Europe and Japan, T. benhamiae var. luteum was recently reported in Brazil (de Freitas et al. 2019; Santana et al. 2020). Trichophyton benhamiae var. benhamiae was confirmed only in North America in our study but was also recently reported in China (Tan et al. 2020).
Animal trade certainly plays an important role in the spread of zoonotic dermatophytes to new geographic areas. It also erases original geographic areas of a species distribution. Consequently, it is difficult to trace the origin of particular species. The current worldwide distribution and prevalence of infections caused by T. benhamiae clade members are poorly known due to insufficient overall surveillance of dermatophytosis supported by molecular-based identification. This problem pertains to both human and veterinary medicine. In addition, our knowledge of the ecology of these pathogens is mostly limited to domestic animals and pets, and little is known about potential wild-living hosts. Therefore, any hypothesis about the species origin is based on very incomplete data and needs to be refined by future research.
Genotyping and surveillance of emerging pathogens in the T. benhamiae complex
The emergence and rapid spread of T. benhamiae in Europe in the last decade and the recent detection of this species in many other countries has been one of the major public health events in the field of zoonotic superficial mycoses in recent years. This fact underscores the need for the One Health integrative approach and closer collaboration between the veterinary profession, dermatologists, epidemiologists and public health personnel (Bontems et al. 2020; Hubka et al. 2018d; Nenoff et al. 2014). Infected and frequently asymptomatic animals may act as a recurrent source of infections in other animals and humans. Interdisciplinary cooperation is needed to establish effective preventive measures for the control of infections.
Genotyping techniques are often employed to gain insight into the dynamics of disease transmission, determine the source and routes of infections, confirm or rule out outbreaks, recognize virulent strains and regional and global changes in genotype patterns and evaluate the effectiveness of control measures (Ranjbar et al. 2014). Other common issues in dermatophytes concern the differentiation of relapse versus reinfection and the determination of whether the infection is caused by one or more strains and if genotypes differ in their clinical manifestation. Many methods have been developed for the subtyping of dermatophytes, but a significant number of them are now obsolete, and their utility is frequently limited due to poor reproducibility or unsatisfactory strain differentiation (Abdel-Rahman 2008; Hubka et al. 2018d; Mochizuki et al. 2017). MLST typing approaches have been widely applied to many fungal pathogens (Bernhardt et al. 2013; Debourgogne et al. 2012; Maitte et al. 2013; Meyer et al. 2009; Prakash et al. 2016), but no such typing scheme has been evaluated and developed for dermatophytes, and the currently available loci usually lack sufficient discriminatory power to study the population structure of Trichophyton and Microsporum species in detail. Microsatellite markers are still among the most effective tools available for the subtyping of dermatophytes. Typing schemes have been developed for a limited number of species, including only T. rubrum (Gräser et al. 2007), Nannizzia persicolor (Sharma et al. 2008) and M. canis (Pasquetti et al. 2013; Sharma et al. 2007).
Polymorphisms in T. benhamiae (Americano-European race) were previously investigated by the RFLP analysis of the NTS region, which produced 11 different patterns in 46 isolates; this method successfully confirmed laboratory-acquired infections as well as familial outbreaks transmitted from pets (Mochizuki et al. 2002; Takeda et al. 2012). In this study, we developed a microsatellite typing scheme consisting of ten variable markers. This new typing scheme is currently the most powerful tool for the subtyping of T. benhamiae clade species. It is easy to use and cost-effective due to its multiplex design. It is possible that the modified scheme can be used in other species in the T. benhamiae complex. Our preliminary data showed that at least 6 of 10 markers (CT21b, TAG16, TC20, TCA16, TC19, TC17a) are useful for the subtyping of another emerging pathogen, T. erinacei.
The establishment of global databases based on largely comparable data, such as that from microsatellites, SNPs and DNA sequences, is desirable. Such databases would enable us to understand the global epidemiology of dermatophytes and monitor changes in genotype spectra on a global scale. Although high-throughput sequencing facilities are now widely available and increasingly used even in the epidemiology of fungal infections, this option has not yet been exploited in dermatophytes.
The prevalence and spread of emerging pathogens from the T. benhamiae complex require close monitoring, particularly because infection rates in the principal hosts (guinea pigs, hedgehogs, porcupines, mammal pets and farm animals) are high. The new taxonomic classification and microsatellite typing scheme proposed in this study will enable the monitoring of changes in the frequencies of individual species and genotypes. It will help to evaluate the results of preventive measures and interventions and is a basic prerequisite for the preparation of epidemiological studies.
Data availability
The important fungal isolate used for experiments are publically available in the internationally recognized culture collections; newly generated DNA sequences are available in European Nucleotide Archive (ENA) database; alignments are available in the Supplementary material.
Change history
07 January 2021
The original version of this article has been revised: Missing supplementary material has been added and the captions to supplementary files 5-7 have been corrected.
References
Abarca M, Castellá G, Martorell J, Cabañes F (2017) Trichophyton erinacei in pet hedgehogs in Spain: occurrence and revision of its taxonomic status. Med Mycol 55:164–172
Abdel-Rahman SM (2008) Strain differentiation of dermatophytes. Mycopathologia 166:319–333
Abdel-Rahman SM et al (2010) Divergence among an international population of Trichophyton tonsurans isolates. Mycopathologia 169:1–13
Agapow PM, Burt A (2001) Indices of multilocus linkage disequilibrium. Mol Ecol Notes 1:101–102
Agnetti F et al (2014) Trichophyton verrucosum infection in cattle farms of Umbria (Central Italy) and transmission to humans. Mycoses 57:400–405
Ahdy AM, Sayed-Ahmed MZ, Younis EE, Baraka HN, El-khodery SA (2016) Prevalence and potential risk factors of dermatophytosis in Arabian horses in Egypt. J Equine Vet Sci 37:71–76
Aho R (1980) Pathogenic dermatophytes recovered from the hair of domestic animals in Finland between 1977 and 1980. Suomen Elainlaakarilehti 86:487–506
Ajello L, Cheng S-L (1967) The perfect state of Trichophyton mentagrophytes. Sabouraudia 5:230–234
Al-Hatmi AMS (2010) Pathogenic fungi isolated from clinical samples in Oman. Master Thesis, Sultan Qaboos University
Ali-Shtayeh M, Arda H, Hassouna M, Shaheen S (1988) Keratinophilic fungi on the hair of cows, donkeys, rabbits, cats, and dogs from the West Bank of Jordan. Mycopathologia 104:109–121
Anzawa K, Kawasaki M, Mochizuki T, Ishizaki H (2010) Successful mating of Trichophyton rubrum with Arthroderma simii. Med Mycol 48:629–634
Atlas RM (2010) Handbook of microbiological media, 4th edn. CRC Press, Boca Raton
Bartosch T et al (2019) Trichophyton benhamiae and T. mentagrophytes target guinea pigs in a mixed small animal stock. Med Mycol Case Rep 23:37–42
Benedict K, Jackson BR, Chiller T, Beer KD (2018) Estimation of direct healthcare costs of fungal diseases in the United States. Clin Infect Dis 68:1791–1797
Bernhardt A, Sedlacek L, Wagner S, Schwarz C, Würstl B, Tintelnot K (2013) Multilocus sequence typing of Scedosporium apiospermum and Pseudallescheria boydii isolates from cystic fibrosis patients. J Cyst Fibros 12:592–598
Bhunjun CS et al (2020) A polyphasic approach to delineate species in Bipolaris. Fungal Divers 102:225–256
Bond R (2010) Superficial veterinary mycoses. Clin Dermatol 28:226–236
Bonifaz A, Archer-Dubon C, Saúl A (2004) Tinea imbricata or Tokelau. Int J Dermatol 43:506–510
Bonifaz A, Vazquez-Gonzalez D (2011) Tinea imbricata in the Americas. Curr Opin Infect Dis 24:106–111
Bontems O, Fratti M, Salamin K, Guenova E, Monod M (2020) Epidemiology of dermatophytoses in Switzerland according to a survey of dermatophytes isolated in Lausanne between 2001 and 2018. J Fungi 6:95
Borman AM, Campbell CK, Fraser M, Johnson EM (2007) Analysis of the dermatophyte species isolated in the British Isles between 1980 and 2005 and review of worldwide dermatophyte trends over the last three decades. Med Mycol 45:131–141
Brasch J, Beck-Jendroschek V, Voss K, Uhrlaß S, Nenoff P (2016) Arthroderma benhamiae strains in Germany. Morphological and physiological characteristics of the anamorphs. Hautarzt 67:700–705
Burt A, Carter DA, Koenig GL, White TJ, Taylor JW (1996) Molecular markers reveal cryptic sex in the human pathogen Coccidioides immitis. Proc Natl Acad Sci 93:770–773
Cafarchia C et al (2010) Epidemiology and risk factors for dermatophytoses in rabbit farms. Med Mycol 48:975–980
Chang Y, Wickes BL, Miller G, Penoyer L, Kwon-Chung K (2000) Cryptococcus neoformans STE12α regulates virulence but is not essential for mating. J Exp Med 191:871–882
Charlent A-L (2011) Le complexe Trichophyton mentagrophytes, caractérisation mycologique et moléculaire d'un nouveau variant: Trichophyton mentagrophytes var. porcellae. Dissertation, Université Henri Poincaré
Cheema MS, Christians JK (2011) Virulence in an insect model differs between mating types in Aspergillus fumigatus. Med Mycol 49:202–207
Chermette R, Ferreiro L, Guillot J (2008) Dermatophytoses in animals. Mycopathologia 166:385–405
Choi JS, Gräser Y, Walther G, Peano A, Symoens F, de Hoog S (2012) Microsporum mirabile and its teleomorph Arthroderma mirabile, a new dermatophyte species in the M. cookei clade. Med Mycol 50:161–169
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659
Concha M, Nicklas C, Balcells E, Guzmán AM, Poggi H, León E, Fich F (2012) The first case of tinea faciei caused by Trichophyton mentagrophytes var. erinacei isolated in Chile. Int J Dermatol 51:283–285
Contet-Audonneau N, Leyer C (2010) Émergence d’un dermatophyte transmis par le cochon d’Inde et proche de Trichophyton mentagrophytes var. erinacei: T. mentagrophytes var. porcellae. J Mycol Med 20:321–325
Courtellemont L, Chevrier S, Degeilh B, Belaz S, Gangneux J-P, Robert-Gangneux F (2017) Epidemiology of Trichophyton verrucosum infection in Rennes University Hospital, France: A 12-year retrospective study. Med Mycol 55:720–724
Čmoková A (2015) Molecular typization of isolates from Arthroderma benhamiae complex, a zoonotic agent of epidemic dermatophytosis in Europe. Master Thesis, Charles University
da Cunha KC et al (2018) Fast identification of dermatophytes by MALDI-TOF/MS using direct transfer of fungal cells on ground steel target plates. Mycoses 61:691–697
Dawson CO, Gentles J (1962) The perfect states of Keratinomyces ajelloi van-Breuseghem, Trichophyton terrestre Durie & Frey and Microsporum nanum Fuentes. Sabouraudia 1:49–57
de Freitas RS, de Freitas THP, Siqueira LPM, Gimenes VMF, Benard G (2019) First report of tinea corporis caused by Arthroderma benhamiae in Brazil. Braz J Microbiol 50:985–987
de Hoog GS et al (2017) Toward a novel multilocus phylogenetic taxonomy for the dermatophytes. Mycopathologia 182:5–31
Debourgogne A, Gueidan C, de Hoog S, Lozniewski A, Machouart M (2012) Comparison of two DNA sequence-based typing schemes for the Fusarium solani species complex and proposal of a new consensus method. J Microbiol Methods 91:65–72
Dhib I, Khammari I, Yaacoub A, Slama FH, Saïd MB, Zemni R, Fathallah A (2017) Relationship between phenotypic and genotypic characteristics of Trichophyton mentagrophytes strains isolated from patients with dermatophytosis. Mycopathologia 182:487–493
Drouot S, Mignon B, Fratti M, Roosje P, Monod M (2009) Pets as the main source of two zoonotic species of the Trichophyton mentagrophytes complex in Switzerland, Arthroderma vanbreuseghemii and Arthroderma benhamiae. Vet Dermatol 20:13–18
Duarte A et al (2010) Survey of infectious and parasitic diseases in stray cats at the Lisbon Metropolitan Area, Portugal. J Feline Med Surg 12:441–446
Dukik K et al (2018) Ultra-high-resolution mass spectrometry for identification of closely related dermatophytes with different clinical predilections. J Clin Microbiol 56:e00102-00118
Dvořák J, Otčenášek M (1969) Mycological diagnosis of animal dermatophytoses. Academia, Prague, Czech Republic
Dvořák J, Otčenášek M, Komárek J (1965) Das Spektrum der aus Tierläsionen in Ostböhmen in den Jahren 1962–1964 isolierten Dermatophyten. Mycoses 8:126–127
Ehrich D (2006) AFLPdat: a collection of R functions for convenient handling of AFLP data. Mol Ecol Notes 6:603–604
English MP, Evans CD, Hewitt M, Warin RP (1962) Hedgehog ringworm. Br Med J 1:149–151
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Fréalle E et al (2007) Phylogenetic analysis of Trichophyton mentagrophytes human and animal isolates based on MnSOD and ITS sequence comparison. Microbiology 153:3466–3477
Fumeaux J et al (2004) First report of Arthroderma benhamiae in Switzerland. Dermatology 208:244–250
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes-application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118
Georg LK (1960) Animal ringworm in public health: diagnosis and nature. US Government Printing Office, Washington
Glass NL, Donaldson GC (1995) Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 61:1323–1330
Gräser Y, De Hoog S, Summerbell R (2006) Dermatophytes: recognizing species of clonal fungi. Med Mycol 44:199–209
Gräser Y, Fröhlich J, Presber W, de Hoog S (2007) Microsatellite markers reveal geographic population differentiation in Trichophyton rubrum. J Med Microbiol 56:1058–1065
Gräser Y, Kuijpers AFA, Presber W, De Hoog GS (2000) Molecular taxonomy of the Trichophyton rubrum complex. J Clin Microbiol 38:3329–3336
Gräser Y, Scott J, Summerbell R (2008) The new species concept in dermatophytes—a polyphasic approach. Mycopathologia 166:239–256
Grisólia ME (2019) Perfil de sensibilidade aos antifúngicos e de variabilidade genética de espécies de Trichophyton isolados de pacientes com infecção cutânea atendidos em um Serviço Público de Micologia em Manaus/AM. Fundação Oswaldo Cruz
Guillot J et al (2018) Emergence of Trichophyton benhamiae in guinea pigs: a retrospective study from the mycology laboratory of the veterinary college of Alfort. Med Mycol 56:S55–S55
Hadrich I, Ranque S (2015) Typing of fungi in an outbreak setting: lessons learned. Curr Fungal Infect Rep 9:314–323
Hainsworth S, Hubka V, Lawrie AC, Carter D, Vanniasinkam T, Grando D (2020) Predominance of Trichophyton interdigitale revealed in podiatric nail dust collections in Eastern Australia. Mycopathologia 185:175–185
Havlickova B, Czaika V, Friedrich M (2008) Epidemiological trends in skin mycoses worldwide. Mycoses 51(Suppl. 4):2–15
Hayette M-P, Sacheli R (2015) Dermatophytosis, trends in epidemiology and diagnostic approach. Curr Fungal Infect Rep 9:164–179
Hedayati MT et al (2019) Identification of clinical dermatophyte isolates obtained from Iran by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Current Medical Mycology 5:22–26
Heidemann S, Monod M, Gräser Y (2010) Signature polymorphisms in the internal transcribed spacer region relevant for the differentiation of zoophilic and anthropophilic strains of Trichophyton interdigitale and other species of T. mentagrophytes sensu lato. Brit J Dermatol 162:282–295
Hejtmánek M, Hejtmánková N (1989) Teleomorphs and mating types in Trichophyton mentagrophytes complex. Acta Univ Palacki Olomuc Fac Med 123:11–33
Hiruma J, Kano R, Harada K, Monod M, Hiruma M, Hasegawa A, Tsuboi R (2015) Occurrence of Arthroderma benhamiae genotype in Japan. Mycopathologia 179:219–223
Houbraken J et al (2020) Classification of Aspergillus, Penicillium, Talaromyces and related genera (Eurotiales): an overview of families, genera, subgenera, sections, series and species. Stud Mycol 95:5–169
Hubka V et al (2018a) Unravelling species boundaries in the Aspergillus viridinutans complex (section Fumigati): opportunistic human and animal pathogens capable of interspecific hybridization. Persoonia 41:142–174
Hubka V et al (2018b) Zoonotic dermatophytoses: clinical manifestation, diagnosis, etiology, treatment, epidemiological situation in the Czech Republic. Čes-slov Derm 93:208–235
Hubka V, Kolařík M (2012) β-tubulin paralogue tubC is frequently misidentified as the benA gene in Aspergillus section Nigri taxonomy: primer specificity testing and taxonomic consequences. Persoonia 29:1–10
Hubka V, Nissen C, Jensen R, Arendrup M, Cmokova A, Kubatova A, Skorepova M (2015) Discovery of a sexual stage in Trichophyton onychocola, a presumed geophilic dermatophyte isolated from toenails of patients with a history of T. rubrum onychomycosis. Med Mycol 53:798–809
Hubka V et al (2018c) Polyphasic data support the splitting of Aspergillus candidus into two species; proposal of Aspergillus dobrogensis sp. nov. Int J Syst Evol Microbiol 68:995–1011
Hubka V, Peano A, Cmokova A, Guillot J (2018) Common and emerging dermatophytoses in animals: well-known and new threats. In: Seyedmousavi S, de Hoog GS, Guillot J, Verweij PE (eds) Emerging and epizootic fungal infections in animals. Springer, Cham, pp 31–79
Hubka V et al (2014) Molecular epidemiology of dermatophytoses in the Czech Republic—two-year-study results. Čes-slov Derm 89:167–174
Hunter PR, Gaston MA (1988) Numerical index of the discriminatory ability of typing systems: an application of Simpson’s index of diversity. J Clin Microbiol 26:2465–2466
Huson DH (1998) SplitsTree: analyzing and visualizing evolutionary data. Bioinformatics 14:68–73
James TY et al (2009) Rapid global expansion of the fungal disease chytridiomycosis into declining and healthy amphibian populations. PLoS Pathog 5:e1000458
Jun JB, Sang YH, Chung SL, Choi JS, Suh SB (2004) The mycological and molecular biological studies on Arthroderma benhamiae isolated for the first time in Korea. Korean J Med Mycol 9:12–27
Kandemir H, Dukik K, Hagen F, Ilkit M, Gräser Y, de Hoog GS (2020) Polyphasic discrimination of Trichophyton tonsurans and T. equinum from humans and horses. Mycopathologia 185:113–122
Kane M, Summerbell R (1997) Laboratory handbook of dermatophytes. A clinical guide and laboratory manual of dermatophytes and other filamentous fungi from skin, hair and nails. Star Publishing Company, Belmont
Kano R, Kawasaki M, Mochizuki T, Hiruma M, Hasegawa A (2012) Mating genes of the Trichophyton mentagrophytes complex. Mycopathologia 173:103–112
Kano R et al (1998) The first isolation of Arthroderma benhamiae in Japan. Microbiol Immunol 42:575–578
Kano R et al (2014) Mating type gene (MAT1-2) of Trichophyton verrucosum. Mycopathologia 177:103–112
Kargl A, Kosse B, Uhrlaß S, Koch D, Krüger C, Eckert K, Nenoff P (2018) Hedgehog fungi in a dermatological office in Munich: case reports and review. Hautarzt 69:576–585
Kaszubiak A, Klein S, De Hoog G, Gräser Y (2004) Population structure and evolutionary origins of Microsporum canis, M. ferrugineum and M. audouinii. Infect Genet Evol 4:179–186
Katoh K, Rozewicki J, Yamada KD (2017) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20:1160–1166
Kawasaki M (2011) Verification of a taxonomy of dermatophytes based on mating results and phylogenetic analyses. Med Mycol J 52:291–295
Kawasaki M, Anzawa K, Mochizuki T, Ishizaki H, M. Hemashettar B, (2009) Successful mating of a human isolate of Arthroderma simii with a tester strain of A. vanbreuseghemii. Med Mycol J 50:15–18
Kawasaki M, Anzawa K, Ushigami T, Kawanishi J, Mochizuki T (2011) Multiple gene analyses are necessary to understand accurate phylogenetic relationships among Trichophyton species. Med Mycol J 52:245–254
Kawasaki M, Anzawa K, Wakasa A, Takeda K, Mochizuki T, Ishizaki H, Hemashettar B (2010) Matings among three teleomorphs of Trichophyton mentagrophytes. Jap J Med Mycol 51:143–152
Kelly KL (1964) Inter-society color council—National bureau of standards color name charts illustrated with centroid colors. US Government Printing Office, Washington
Khettar L, Contet-Audonneau N (2012) Cochon d’Inde et dermatophytose. Ann Dermatol Venereol 139:631–635
Khosravi A, Mahmoudi M (2003) Dermatophytes isolated from domestic animals in Iran. Mycoses 46:222–225
Kimura U, Yokoyama K, Hiruma M, Kano R, Takamori K, Suga Y (2015) Tinea faciei caused by Trichophyton mentagrophytes (molecular type Arthroderma benhamiae) mimics impetigo: a case report and literature review of cases in Japan. Med Mycol J 56:E1–E5
Kosanke S, Hamann L, Kupsch C, Garcia SM, Chopra A, Gräser Y (2018) Unequal distribution of the mating type (MAT) locus idiomorphs in dermatophyte species. Fungal Genet Biol 118:45–53
Kosman E (2003) Nei’s gene diversity and the index of average differences are identical measures of diversity within populations. Plant Pathol 52:533–535
Kraemer A, Hein J, Heusinger A, Mueller R (2013) Clinical signs, therapy and zoonotic risk of pet guinea pigs with dermatophytosis. Mycoses 56:168–172
Kraemer A, Mueller R, Werckenthin C, Straubinger R, Hein J (2012) Dermatophytes in pet guinea pigs and rabbits. Vet Microbiol 157:208–213
Kuhnert E et al (2017) Phylogenetic and chemotaxonomic resolution of the genus Annulohypoxylon (Xylariaceae) including four new species. Fungal Divers 85:1–43
Kupsch C, Berlin M, Gräser Y (2017) Dermophytes and guinea pigs: An underestimated danger? Hautarzt 68:827–830
Kupsch C, Berlin M, Ritter L, Heusinger A, Stoelker B, Graeser Y (2019) The guinea pig fungus Trichophyton benhamiae—Germany-wide distribution analysis of the zoonotic agent. In: Groschup MH, Ludwig S, Drosten C (eds) Zoonoses 2019 – International Symposium on Zoonoses Research, Berlin, Germany, 2020. Journal der Deutschen Dermatologischen Gesellschaft, p 12
Lambert C, Wendt L, Hladki AI, Stadler M, Sir EB (2019) Hypomontagnella (Hypoxylaceae): a new genus segregated from Hypoxylon by a polyphasic taxonomic approach. Mycol Prog 18:187–201
Lanfear R, Frandsen PB, Wright AM, Senfeld T, Calcott B (2017) PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol Biol Evol 34:772–773
Lebasque J (1933) Les champignons des teignes du cheval et des bovidés. Dissertation, Faculté des Sciences de Paris
Lee WJ, Eun DH, Jang YH, Lee S-J, Bang YJ, Jun JB (2018) Tinea faciei in a mother and daughter caused by Arthroderma benhamiae. Ann Dermatol 30:241–242
Leigh JW, Bryant D (2015) POPART: full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116
Lund A, Bratberg AM, Næss B, Gudding R (2014) Control of bovine ringworm by vaccination in Norway. Vet Immunol Immunopathol 158:37–45
Lysková P et al (2018) Five cases of dermatophytosis in man caused by zoophilic species Trichophyton erinacei transmitted from hedgehogs. Čes-slov Derm 93:237–243
Lysková P, Hubka V, Petřičáková A, Dobiáš R, Čmoková A, Kolařík M (2015) Equine dermatophytosis due to Trichophyton bullosum, a poorly known zoophilic dermatophyte masquerading as T. verrucosum. Mycopathologia 180:407–419
Maitte C, Leterrier M, Le Pape P, Miegeville M, Morio F (2013) Multilocus sequence typing of Pneumocystis jirovecii from clinical samples: how many and which loci should be used? J Clin Microbiol 51:2843–2849
Martins WS, Lucas DCS, de Souza Neves KF, Bertioli DJ (2009) WebSat-A web software for microsatellite marker development. Bioinformation 3:282–283
Matute DR, Sepúlveda VE (2019) Fungal species boundaries in the genomics era. Fungal Genet Biol 131:103249
Metin B, Heitman J (2017) Sexual reproduction in dermatophytes. Mycopathologia 182:45–55
Meyer W et al (2009) Consensus multi-locus sequence typing scheme for Cryptococcus neoformans and Cryptococcus gattii. Med Mycol 47:561–570
Mirhendi H, Makimura K, de Hoog GS, Rezaei-Matehkolaei A, Najafzadeh MJ, Umeda Y, Ahmadi B (2015) Translation elongation factor 1-α gene as a potential taxonomic and identification marker in dermatophytes. Med Mycol 53:215–224
Mochizuki T, Kawasaki M, Ishizaki H, Kano R, Hasegawa A, Tosaki H, Fujihiro M (2001) Molecular epidemiology of Arthroderma benhamiae, an emerging pathogen of dermatophytoses in Japan, by polymorphisms of the non-transcribed spacer region of the ribosomal DNA. J Dermatol Sci 27:14–20
Mochizuki T, Takeda K, Anzawa K (2017) Molecular markers useful for intraspecies subtyping and strain differentiation of dermatophytes. Mycopathologia 182:57–65
Mochizuki T, Watanabe S, Kawasaki M, Tanabe H, Ishizaki H (2002) A Japanese case of tinea corporis caused by Arthroderma benhamiae. J Dermatol 29:221–225
Moretti A et al (2013) Dermatophytosis in animals: epidemiological, clinical and zoonotic aspects. G Ital Dermatol Venereol 148:563–572
Morris P, English MP (1969) Trichophyton mentagrophytes var. erinacei in British hedgehogs. Sabouraudia 7:122–128
Morris P, English MP (1973) Transmission and course of Trichophyton erinacei infections in British hedgehogs. Sabouraudia 11:42–47
Müller K (2005) SeqState. Appl Bioinformatics 4:65–69
Nakamura Y, Kano R, Nakamura E, Saito K, Watanabe S, Hasegawa A (2002) Case report. First report on human ringworm caused by Arthroderma benhamiae in Japan transmitted from a rabbit. Mycoses 45:129–131
Needle DB et al (2019) Atypical Dermatophytosis in 12 North American Porcupines (Erethizon dorsatum) from the Northeastern United States 2010–2017. Pathogens 8:171
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nenoff P, Erhard M, Simon JC, Muylowa GK, Herrmann J, Rataj W, Gräser Y (2013) MALDI-TOF mass spectrometry-a rapid method for the identification of dermatophyte species. Med Mycol 51:17–24
Nenoff P et al (2014) Trichophyton species von Arthroderma benhamiae – a new infectious agent in dermatology. J Dtsch Dermatol Ges 12:571–582
Nenoff P, Verma SB, Uhrlaß S, Burmester A, Gräser Y (2019) A clarion call for preventing taxonomical errors of dermatophytes using the example of the novel Trichophyton mentagrophytes genotype VIII uniformly isolated in the Indian epidemic of superficial dermatophytosis. Mycoses 62:6–10
Nguyen L-T, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274
Overgaauw P, van Avermaete K, Mertens C, Meijer M, Schoemaker N (2017) Prevalence and zoonotic risks of Trichophyton mentagrophytes and Cheyletiella spp. in guinea pigs and rabbits in Dutch pet shops. Vet Microbiol 205:106–109
Packeu A, Stubbe D, Roesems S, Goens K, Van Rooij P, de Hoog S, Hendrickx M (2020) Lineages within the Trichophyton rubrum complex. Mycopathologia 185:123–136
Padhye A, Carmichael J (1972) Arthroderma insingulare sp. nov., another gymnoascaceous state of the Trichophyton terrestre complex. Sabouraudia 10:47–51
Papegaay J (1925) Over pathogene huidschimmels in Amsterdam voorkomend bij den mensch. Ned Tijdschr Geneeskd 69:879–890
Parker ED Jr (1979) Ecological implications of clonal diversity in parthenogenetic morphospecies. Am Zool 19:753–762
Pasquetti M, Peano A, Soglia D, Min ARM, Pankewitz F, Ohst T, Gräser Y (2013) Development and validation of a microsatellite marker-based method for tracing infections by Microsporum canis. J Dermatol Sci 70:123–129
Pchelin IM, Azarov DV, Churina MA, Scherbak SG, Apalko SV, Vasilyeva NV, Taraskina AE (2019) Species boundaries in the Trichophyton mentagrophytes/T. interdigitale species complex. Med Mycol 57:781–789
Pchelin IM et al (2016) Reconstruction of phylogenetic relationships in dermatomycete genus Trichophyton Malmsten 1848 based on ribosomal internal transcribed spacer region, partial 28S rRNA and beta-tubulin genes sequences. Mycoses 59:566–575
Piérard-Franchimont C, Hermanns J-F, Collette C, Pierard G, Quatresooz P (2008) Hedgehog ringworm in humans and a dog. Acta Clin Belg 63:322–324
Pihet M, Bourgeois H, Mazière J-Y, Berlioz-Arthaud A, Bouchara J-P, Chabasse D (2008) Isolation of Trichophyton concentricum from chronic cutaneous lesions in patients from the Solomon Islands. Trans R Soc Trop Med Hyg 102:389–393
Prakash A et al (2016) Evidence of genotypic diversity among Candida auris isolates by multilocus sequence typing, matrix-assisted laser desorption ionization time-of-flight mass spectrometry and amplified fragment length polymorphism. Clin Microbiol Infect 22:277.e271-277.e279
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Quaife R (1966) Human infection due to the hedgehog fungus, Trichophyton mentagrophytes var. erinacei. J Clin Pathol 19:177–178
R Core Team (2016) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Ranjbar R, Karami A, Farshad S, Giammanco GM, Mammina C (2014) Typing methods used in the molecular epidemiology of microbial pathogens: a how-to guide. New Microbiol 37:1–15
Réblová M, Hubka V, Thureborn O, Lundberg J, Sallstedt T, Wedin M, Ivarsson M (2016) From the tunnels into the treetops: new lineages of black yeasts from biofilm in the Stockholm metro system and their relatives among ant-associated fungi in the Chaetothyriales. PLoS ONE 11:e0163396
Rezaei-Matehkolaei A et al (2013) Molecular epidemiology of dermatophytosis in Tehran, Iran, a clinical and microbial survey. Med Mycol 51:203–207
Rezaei-Matehkolaei A, Rafiei A, Makimura K, Gräser Y, Gharghani M, Sadeghi-Nejad B (2016) Epidemiological aspects of dermatophytosis in Khuzestan, southwestern Iran, an update. Mycopathologia 181:547–553
Rippon JW (1988) Medical mycology. The pathogenic fungi and the pathogenic actinomycetes, 3rd edn. Saunders, Philadelphia
Ronquist F et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Sabou M et al (2018) Molecular identification of Trichophyton benhamiae in Strasbourg, France: a 9-year retrospective study. Med Mycol 56:723–734
Samarakoon MC et al (2020) Elucidation of the life cycle of the endophytic genus Muscodor and its transfer to Induratia in Induratiaceae fam. nov., based on a polyphasic taxonomic approach. Fungal Divers 101:177–210
Santana AE, Reche-Junior A, Sellera FP, Taborda CP (2020) A comment on “First report of tinea corporis caused by Arthroderma benhamiae in Brazil”. Braz J Microbiol 51:1463–1464
Schauder S, Kirsch-Nietzki M, Wegener S, Switzer E, Qadripur S (2007) Von Igeln auf Menschen: Zoophile Dermatomykose durch Trichophyton erinacei bei 8 Patienten. Hautarzt 58:62–67
Schlueter PM, Harris SA (2006) Analysis of multilocus fingerprinting data sets containing missing data. Mol Ecol Notes 6:569–572
Schneider S, Roessli D, Excoffier L (2000) ARLEQUIN: a software for population genetics data analysis, Version 2.000 vol 2. University of Geneva, Geneva
Schönswetter P, Tribsch A (2005) Vicariance and dispersal in the alpine perennial Bupleurum stellatum L. (Apiaceae). Taxon 54:725–732
Schrödl W et al (2012) Direct analysis and identification of pathogenic Lichtheimia species by matrix-assisted laser desorption ionization–time of flight analyzer-mediated mass spectrometry. J Clin Microbiol 50:419–427
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol 18:233–234
Seebacher C, Bouchara J-P, Mignon B (2008) Updates on the epidemiology of dermatophyte infections. Mycopathologia 166:335–352
Sharma R, De Hoog S, Presber W, Gräser Y (2007) A virulent genotype of Microsporum canis is responsible for the majority of human infections. J Med Microbiol 56:1377–1385
Sharma R, Presber W, Rajak RC, Gräser Y (2008) Molecular detection of Microsporum persicolor in soil suggesting widespread dispersal in central India. Med Mycol 46:67–73
Shenoy MM, Jayaraman J (2019) Epidemic of difficult-to-treat tinea in India: current scenario, culprits, and curbing strategies. Arch Med Health Sci 7:112–117
Sieklucki U, Oh SH, Hoyer LL (2014) Frequent isolation of Arthroderma benhamiae from dogs with dermatophytosis. Vet Dermatol 25:39–41
Silver S, Vinh DC, Embil JM (2008) The man who got too close to his cows. Diagn Microbiol Infect Dis 60:419–420
Singh A et al (2019) A unique multidrug-resistant clonal Trichophyton population distinct from Trichophyton mentagrophytes/Trichophyton interdigitale complex causing an ongoing alarming dermatophytosis outbreak in India: Genomic insights and resistance profile. Fungal Genet Biol 133:103266
Sitterle E et al (2012) Trichophyton bullosum: a new zoonotic dermatophyte species. Med Mycol 50:305–309
Skořepová M, Hubka V, Polášková S, Stará J, Čmoková A (2014) Our first experiences with Infections caused by Arthroderma benhamiae (Trichophyton sp.). Čes-slov Derm 89:192–198
Smith J, Marples MJ (1964) Trichophyton mentagrophytes var. erinacei. Sabouraudia 3:1–10
Stadler M, Lambert C, Wibberg D, Kalinowski J, Cox RJ, Kolařík M, Kuhnert E (2020) Intragenomic polymorphisms in the ITS region of high-quality genomes of the Hypoxylaceae (Xylariales, Ascomycota). Mycol Prog 19:235–245
Steenkamp ET, Wingfield MJ, McTaggart AR, Wingfield BD (2018) Fungal species and their boundaries matter – Definitions, mechanisms and practical implications. Fungal Biol Rev 32:104–116
Stockdale PM (1964) The Microsporum gypseum complex (Nannizzia incurvata Stockd., N. gypsea (Nann.) comb. nov., N. fulva sp. nov.). Sabouraudia 3:114–126
Su H et al (2019) Species distinction in the Trichophyton rubrum complex. J Clin Microbiol 57:e00352-e1319
Suh S-O, Grosso KM, Carrion ME (2018) Multilocus phylogeny of the Trichophyton mentagrophytes species complex and the application of matrix-assisted laser desorption/ionization–time-of-flight (MALDI-TOF) mass spectrometry for the rapid identification of dermatophytes. Mycologia 110:118–130
Summerbell R (2002) What is the evolutionary and taxonomic status of asexual lineages in the dermatophytes? Stud Mycol 47:97–101
Summerbell RC (2011) Trichophyton, Microsporum, Epidermophyton, and agents of superficial mycoses. In: Versalovic J, Carroll K, Funke G, Jorgensen J, Landry M, Warnock D (eds) Manual of clinical microbiology, 10th edn. American Society of Microbiology, Washington, pp 1919–1942
Summerbell RC, Moore MK, Starink-Willemse M, Van Iperen A (2007) ITS barcodes for Trichophyton tonsurans and T. equinum. Med Mycol 45:193–200
Symoens F, Jousson O, Packeu A, Fratti M, Staib P, Mignon B, Monod M (2013) The dermatophyte species Arthroderma benhamiae: intraspecies variability and mating behaviour. J Med Microbiol 62:377–385
Taghipour S et al (2019) Trichophyton mentagrophytes and T. interdigitale genotypes are associated with particular geographic areas and clinical manifestations. Mycoses 62:1084–1091
Takahashi H et al (2008) An intrafamilial transmission of Arthroderma benhamiae in Canadian porcupines (Erethizon dorsatum) in a Japanese zoo. Med Mycol 46:465–473
Takahashi Y, Haritani K, Sano A, Takizawa K, Fukushima K, Miyaji M, Nishimura K (2002) An isolate of Arthroderma benhamiae with Trichophyton mentagrophytes var. erinacei anamorph isolated from a four-toed hedgehog (Atelerix albiventris) in Japan. Jap J Med Mycol 43:249–255
Takahashi Y, Sano A, Takizawa K, Fukushima K, Miyaji M, Nishimura K (2003) The epidemiology and mating behavior of Arthroderma benhamiae var. erinacei in household four-toed hedgehogs (Atelerix albiventris) in Japan. Jap J Med Mycol 44:31–38
Takashio M (1974) Observations on African and European strains of Arthroderma benhamiae. Int J Dermatol 13:94–101
Takashio M (1977) The Trichophyton mentagrophytes complex. In: Iwata K (ed) Recent advances in medical and veterinary mycology. University of Tokyo Press, Tokyo, pp 271–276
Takeda K, Nishibu A, Anzawa K, Mochizuki T (2012) Molecular epidemiology of a major subgroup of Arthroderma benhamiae isolated in Japan by restriction fragment length polymorphism analysis of the non-transcribed spacer region of ribosomal RNA gene. Jpn J Infect Dis 65:233–239
Tan J, Liu X, Gao Z, Yang H, Yang L, Wen H (2020) A case of Tinea Faciei caused by Trichophyton benhamiae: first report in China. BMC Infect Dis 20:1–5
Tartor YH, El Damaty HM, Mahmmod YS (2016) Diagnostic performance of molecular and conventional methods for identification of dermatophyte species from clinically infected Arabian horses in Egypt. Vet Dermatol 27:401-e102
Taylor JW, Hann-Soden C, Branco S, Sylvain I, Ellison CE (2015) Clonal reproduction in fungi. Proc Natl Acad Sci USA 112:8901–8908
Trivedi J et al (2017) Fungus causing white-nose syndrome in bats accumulates genetic variability in North America with no sign of recombination. Msphere 2:e00271-e1217
Turland NJ et al (2018) International Code of Nomenclature for algae, fungi, and plants (Shenzhen Code) adopted by the Nineteenth International Botanical Congress Shenzhen, China, July 2017. Koeltz Botanical Books, Glashütten
Uhrlaß S, Krüger C, Nenoff P (2015) Microsporum canis: Aktuelle Daten zur Prävalenz des zoophilen Dermatophyten im mitteldeutschen Raum. Hautarzt 66:855–862
Uhrlaß S et al (2018) Molecular epidemiology of Trichophyton quinckeanum–a zoophilic dermatophyte on the rise. J Dtsch Dermatol Ges 16:21–32
Vu D et al (2019) Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Stud Mycol 92:135–154
Wang F-Y, Sun P-L (2018) Tinea blepharo-ciliaris in a 13-year-old girl caused by Trichophyton benhamiae. J Mycol Med 28:542–546
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Woodgyer A (2004) The curious adventures of Trichophyton equinum in the realm of molecular biology: a modern fairy tale. Med Mycol 42:397–403
Yeh FC et al. (1999) POPGENE version 1.31. A Microsoft window based freeware for population genetic analysis. University of Alberta, Canada
Yue C, Cavallo LM, Alspaugh JA, Wang P, Cox GM, Perfect JR, Heitman J (1999) The STE12α homolog is required for haploid filamentation but largely dispensable for mating and virulence in Cryptococcus neoformans. Genetics 153:1601–1615
Zhan P et al (2018) Phylogeny of dermatophytes with genomic character evaluation of clinically distinct Trichophyton rubrum and T. violaceum. Stud Mycol 89:153–175
Ziółkowska G, Nowakiewicz A, Gnat S, Trościańczyk A, Zięba P, Majer Dziedzic B (2015) Molecular identification and classification of Trichophyton mentagrophytes complex strains isolated from humans and selected animal species. Mycoses 58:119–126
Acknowledgements
We are very grateful to Jan Karhan and Lukáš Vít Rýdl for the concept of data visualization and help with graphical adjustments of analysis outputs. We thank Milada Chudíčkova, Petra Seifertová and Adéla Kovaříčková for their invaluable assistance in the laboratory and Peter Mikula for research support. We thank Jiřina Stará, Magdalena Skořepová, Stanislava Dobiášová and Jana Hanzlíčková for providing some of the strains used in this study. The research reported in this publication was part of the long-term goals of the ISHAM working group Onygenales.
Funding
Charles University Grant Agency (GAUK 600217): A. Čmoková; Czech Ministry of Health (AZV 17-31269A): M. Kolařík, R. Dobiáš, H. Janouškovcová, I. Kuklová, N. Mallátová, K. Mencl, T. Větrovský, V. Hubka; BIOCEV (CZ.1.05/1.1.00/02.0109) provided by the Ministry of Education, Youth and Sports of the Czech Republic and ERDF: V. Hubka; Charles University Research Centre program no. 204069: V. Hubka; Czech Academy of Sciences (Project RVO 67985939): M. Man.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflicts of interest. The authors alone are responsible for the content and the writing of the paper.
Electronic supplementary material
Below is the link to the electronic supplementary material.
13225_2020_465_MOESM2_ESM.pdf
Supplementary file2 Fig. S2 Maximum likelihood tree based on ITS region sequences. Maximum likelihood bootstrap values are appended to the nodes; only support values higher than 70% are shown; the ex-type strains are designated with a superscripted T; Trichophyton rubrum CBS 202.88 was used as the outgroup. Clades with >5 identical sequences are collapsed; *positions refer to the alignment available in the Supplementary material (PDF 76 KB)
13225_2020_465_MOESM3_ESM.pdf
Supplementary file3 Fig. S3 Maximum likelihood tree based on gapdh gene sequences. Maximum likelihood bootstrap values are appended to the nodes; only support values higher than 70% are shown; the ex-type strains are designated with a superscripted T; Trichophyton rubrum CBS 202.88 was used as the outgroup. Clades with >5 identical sequences are collapsed; *positions refer to the alignment available in the Supplementary material (PDF 63 KB)
13225_2020_465_MOESM4_ESM.pdf
Supplementary file4 Fig. S4 Maximum likelihood tree based on tef1-α gene sequences. Maximum likelihood bootstrap values are appended to the nodes; only support values higher than 70% are shown; the ex-type strains are designated with a superscripted T; Trichophyton rubrumTrichophyton rubrum CBS 202.88 was used as the outgroup. Clades with >5 identical sequences are collapsed; *positions refer to the alignment available in the Supplementary material (PDF 80 KB)
13225_2020_465_MOESM5_ESM.pdf
Supplementary file5 Fig. S5 Maximum likelihood tree based on tubb gene sequences. Maximum likelihood bootstrap values are appended to the nodes; only support values higher than 70% are shown; the ex-type strains are designated with a superscripted T; Trichophyton rubrum CBS 202.88 was used as the outgroup. Clades with >5 identical sequences are collapsed (PDF 54 KB)
13225_2020_465_MOESM6_ESM.png
Supplementary file6 Fig. S6 Phylogenetic tree of the Trichophyton benhamiae clade revealed by the analysis of ten microsatellite loci in 318 strains constructed in FAMD software using a Jaccard index-based distance matrix. Coloured circles display the genotype diversity of the ITS, gapdh and tef1-α loci and the distribution of MAT gene idiomorphs across Trichophyton benhamiae clade species (PNG 915 KB)
Rights and permissions
About this article
Cite this article
Čmoková, A., Kolařík, M., Dobiáš, R. et al. Resolving the taxonomy of emerging zoonotic pathogens in the Trichophyton benhamiae complex. Fungal Diversity 104, 333–387 (2020). https://doi.org/10.1007/s13225-020-00465-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-020-00465-3