Abstract
Understanding the molecular mechanism underlying photoperiod sensitivity will play a crucial role in extending the cropping area of Cajanus cajan, a photoperiod sensitive major grain legume of India and Africa. In flowering plants, Flowering locus T (FT) gene is involved in the production of florigen molecule which is essential for induction of flowering, influenced largely by the duration of photoperiod. To understand the structural and regulatory nature of this gene, a genome-wide survey was carried out, revealing the presence of 13 PEBP (FT) family genes in C. cajan. Based on the gene expression profiling of 13 PEBP genes across the 30 tissues of C. cajan, CcFT6 and CcFT8 were found to be probable Flowering locus T genes responsible for the production of florigen as both of them showed expression in reproductive leaf. Expression analysis in photoperiod sensitive, MAL3 genotype revealed that CcFT6 is upregulated under SD. However, in photoperiod insensitive genotype (ICP20338) CcFT6 and CcFT8 were upregulated in SD and LD, respectively. Hence, in ICP20338 under SD, flowering induction occurs with the involvement of CcFT6 while under LD, flowering induction seems to be associated with the expression of CcFT8. CcFT6 was found to be expressed only under favourable photoperiodic condition (SD) in both MAL3 and ICP20338 and may be regulated through a photoperiod dependent pathway. The presence of additional florigen producing gene, CcFT8 in ICP20338 which has the ability to flower in a photoperiod independent manner under LD conditions might provide some clues on its photoperiod insensitive nature. This study will provide a detailed characterization of the genes involved in photoperiodic regulation of flowering in C. cajan.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The PEBP (phosphatidyl ethanolamine-binding protein) is an important family of proteins widely distributed across many organisms including bacteria, yeast, animals and plants (Danilevskaya et al. 2008). It is an evolutionarily conserved family of proteins, involved in numerous functions in animals like neural development, lipid binding, serine protease inhibitor and flower induction in plants (Banfield et al. 1998; Saunders et al. 1995; Hengst et al. 2001; Song et al. 2015). Along with flower induction, PEBP family of proteins are also involved in many biological processes such as embryogenesis, stress tolerance, (Karlgren et al. 2011) hormone signalling, tuberization and determination of plant architecture (Chen et al. 2018; Karlgren et al. 2011; Navarro et al. 2011). PEBP family proteins possess a common PEBP domain and can be classified into three subfamilies, which includes MOTHER OF FT AND TFL1 (MFT)-like proteins, TERMINAL FLOWER1 (TFL1)-like proteins and FLOWERING LOCUS T like proteins (Danilevskaya et al. 2008; Argiriou et al. 2008; Harig et al. 2012). Among the three subgroups of PEBP family, FLOWERING LOCUS T (FT) and TERMINAL FLOWER1 (TFL1) have been intensively studied in Arabidopsis due to the reason that, despite a 90% amino acid identity between these two, they showed antagonistic function with respect to flowering induction (Karlgren et al. 2011). In the presence of favourable photoperiod and temperature, FT promotes the flowering induction while TFL1 acts as a floral repressor (Higuchi 2018).
The molecular mechanism underlying the photoperiodic response in plants which leads to flower initiation is well characterized in Arabidopsis [long day plant (LD)] and rice [short day plant(SD)] (Jeong and Clark 2005; Izawa 2007). Flowering largely depends upon expression of flowering locus T (FT) in Arabidopsis and Hd3a and RFT1 in rice which encodes florigen, a systemic signalling molecule required for flower induction. It is transcribed and translated in the phloem companion cells of leaves and translocated through the phloem to the shoot apex (Corbesier et al. 2007; Tamaki et al. 2007; Cheng et al. 2018). At the shoot apex, it interacts with 14–3–3, a class of TPR proteins and FD proteins and forms a ‘florigen activation complex’ (FAC), which induces transcription of floral meristem identity genes such as APETALA1 (AP1) like and LFY gene leading to flower development (Taoka et al. 2011).
The amount of florigen inside the shoot tissues determines the timing of flowering in A. thaliana; under LD condition, expression of FT genes gets upregulated in leaf tissues leading to the synthesis of high amounts of florigen, which are further translocated to shoot apex through phloem resulting in rapid flowering; whereas under SD, expression of FT genes in the leaf tissue is suppressed and as a result, no florigen protein is synthesized in the leaf leading to delay in flowering (Kobayashi et al. 1999). It was reported that the CONSTANS (CO) gene, a member of the CCT gene family, is involved in the activation of FT expression under LD condition in A. thaliana (Samach et al. 2000). To induce FT expression, CO protein interacts with light signalling and circadian clock-associated proteins, forming FT activation complex (Samach et al. 2000). A. thaliana CO protein interacts with GIGANTEA (GI), CIRCADIAN CLOCK ASSOCIATED 1 (CCA1), FLAVIN-BINDING KELCH REPEAT F-BOX 1 (FKF-1), PRR, CYCLING DOF FACTORs (CDFs), LATE ELONGATED HYPOCOTYL (LHY), TOC1 etc., to sense external photoperiodic condition before induction of FT expression (David et al. 2006; Niwa et al. 2007; Sawa et al. 2007; Fornara et al. 2009; Nakamichi et al. 2010; Song et al. 2012).
In rice, Hd3a, an ortholog of Arabidopsis FT is responsible for the induction of photoperiod dependent flowering under SD photoperiodic condition (Komiya et al. 2008). Besides Hd3a, RICE FLOWERING LOCUS T1 (RFT1, a Hd3a paralog) was found to be responsible for the induction of flowering under LD condition. The flowering induction in rice at LD condition was regulated through histone modification at chromatin level with respect to gene Ehd1, RICE FLOWERING LOCUS T1 (RFT1, a Hd3a paralog) and their upstream regulators (Sun et al. 2012). In Sorghum bicolor, among the 19 PEBP genes, three genes viz. SbFT1, SbFT8 and SbFT10 were reported to be the potential candidates for florigen production (Wolabu et al. 2016). In Zea mays, among the seven CENTRORADIALIS (ZCN) genes encoding FT proteins, only ZCN8 was found to be responsible for florigenic activity (Meng et al. 2011). In Cicer arietinum, the cluster of FTa1, FTa2 and FTc genes were observed to be involved in the regulation of photoperiod dependent flowering (Ortega et al. 2019).
Pigeon pea (Cajanus cajan) is a prominent pulse crop of rainfed areas. It is a high protein (20–23%) containing food legume, providing for nutritional security of the Indian and parts of the African subcontinent. Globally, it ranks sixth in the grain legume production and is grown in 7.02 million hectares (mha) with a yearly production of 6.80 million tons and productivity of 969.1 kg/ha (FAOSTAT 2017). In India alone, C. cajan is cultivated in about 5.38 mha area (75% of the global area) with an average production of 4.87 million tonnes, accounting for 71% of the global production (FAOSTAT 2017). It occupies a prime niche in sustainable farming systems of small and marginal rainfed farmers. It is a quantitative short-day plant, requires 11–11.5 h of critical day length for induction of flowering. Warm weather combined with long photoperiod promotes vegetative growth, while short photoperiod and milder mean ambient temperatures (> 18 °C and < 25 °C) promote flowering. Traditionally, C. cajan is sown at the onset of the rainy season and harvested after 6–9 months. Due to its strong photoperiod requirement, C. cajan cultivation is restricted to the latitudes between 30° N and S, with a temperature range of 20–40 °C (Sinha 1977). The development of photoperiod insensitive cultivars in pigeon pea would be useful to ameliorate the situation. Hence, identification of photoperiod insensitive genotypes and understanding of the molecular mechanism underlying photoperiod insensitivity in this crop will aid in the expansion of its cultivation to diverse latitudes.
Material and methods
Screening of genotype for photoperiod insensitivity
Set of twelve genotypes of C. cajan (Table S1) were grown in three replications at 30 °C temperature and 60% relative humidity conditions under long day (LD) 16 hr/8 hr (light/dark) and short day (SD) 11.5 hr/12.5 hr (light/dark) photoperiod condition at National Phytotron Facility, ICAR-IARI, New Delhi. The same set of genotypes was screened consecutively for two years and the capability of each genotype to initiate flowering under two different photoperiod condition was recorded (Table S1).
Genome-wide identification of PEBP family genes
Genome-wide scanning of PEBP family of genes was carried out using C. cajan genome available at Legume Information System (LIS) database. In the LIS database, ‘gene family search’ option was used for identifying members of FT family. The PEBP family genes were searched using ‘FLOWERING LOCUS T PROTEIN’ as a keyword in the description field. Similarly, the PEBP family protein sequences of legume species viz. Vigna angularis, Glycine max, Vigna radiata, Cicer arietinum, Vigna unguiculata and Medicago truncatula were also downloaded from Legume Information System (LIS) database. The PEBP family protein sequences of O. sativa and A. thaliana were downloaded from the Rice Genome Annotation Project Database (http:// rice.plantbiology.msu.edu/index.shtml) and The Arabidopsis Information Resource (TAIR: https://www.arabidopsis.org) respectively. Chromosomal positions of each C. cajan PEBP family gene were obtained by using the BLAT search option available at the LIS database at default parameter. Chromosome wise distribution pattern of each gene was drawn using Mapchart software 2.3 (Voorrips 2002).
Phylogenetic analysis of PEBP family genes
The phylogenetic tree of C. cajan and Arabidopsis PEBP family genes was constructed using MEGA 10.0.5 software (Tamura et al. 2013) to establish the sub-grouping of C. cajan PEBP family genes into various classes. The phylogenetic analysis was performed using the maximum likelihood method with a bootstrap value of 1000, Jones Taylor-Thornton (JTT) model and complete deletion option available in the MEGA 10.0.5 software. To understand the phylogenetic relationship of C. cajan PEBP family proteins with other legumes, a combined phylogenetic tree was generated by adopting similar methodology using the PEBP protein sequences obtained from Vigna angularis, Glycine max, Vigna radiata, Cicer arietinum, Vigna unguiculata, Medicago truncatula, O. sativa and A. thaliana.
Nomenclature and domain prediction PEBP family genes
The C. cajan PEBP family genes were classified into three subgroups on the basis of phylogenetic relationship with A. thaliana PEPB genes and were renamed with increasing order of genomic position (Zhang et al. 2015). The protein sequences of PEBP genes were subjected to domain prediction using ‘batch web CD-search tool’ (https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi.) and the motif composition of each gene was identified using MEME motif prediction tools (https://meme-suite.org/).
Preparation of time tree
Phylogenetic tree was constructed through MEGA-10.0.5 using Realtime ML method which uses Tamura-Nei model for substitution at uniform rates. Time tree was developed using default parameters without adding any calibration point. While developing the time tree, AtMFT protein sequence was used as out-group. Relative time estimates using Realtime-ML method are useful for determining the relative ordering and spacing of speciation events, identifying lineages and estimating absolute times of divergence (Tamura et al. 2012).
Expression pattern analysis of PEBP family genes using gene expression atlas
Expression atlas of C. cajan developed by Pazhamala et al. (2017) from 30 tissues of a C. cajan cv. Asha was used to visualize the expression profile of all CcFT genes in different tissues. The gene expression data in the form of FPKM values of each selected genes were filtered from the gene expression atlas and used for the preparation of heatmap using gplots package of ‘R’.
Plant material and sample collection
The photoperiod sensitive (MAL3) and photoperiod insensitive (ICP20338) genotypes (Table 1) of C. cajan were grown in three replications at 30 °C temperature and 60% relative humidity conditions under long day (LD) 16 hr/8 hr (light/dark) and short day (SD) 12 hr/12 hr (light/dark) photoperiod condition at National Phytotron Facility, ICAR-IARI, New Delhi. It has been reported that after completing vegetative growth phase, flower induction signals are initiated in the leaf tissue of plant (Komiya et al. 2008); hence, the leaf tissue harvested during vegetative phase [vegetative leaf (VL)] and leaf tissue harvested at flower induction phase [reproductive leaf (RL)] seem to be most appropriate to validate our results through qRT-PCR. The VL and RL tissues from MAL3 and ICP20338 genotypes, grown under SD and LD photoperiod were harvested for RNA isolation. ICP20338 flowered in 55 DAS, hence VL tissue was harvested after 30 DAS and RL tissue was harvested at 50 DAS from both SD and LD grown plants. MAL3 flowered at 120 DAS, hence, VL tissue was harvested at 50 DAS and RL tissue was harvested 115 DAS from both SD and LD grown plants. In MAL3, RL tissue harvested from LD grown plant was not really reproductive leaf because the plant remains in the vegetative phase, hence we designated it as a pseudo reproductive leaf (PRL).
RNA isolation and cDNA synthesis
Total RNA was isolated using Spectrum Plant Total RNA Kit (Sigma Aldrich) from RL and VL tissues sampled from MAL3 and ICP20338 of C. cajan grown under SD and LD photoperiod condition. DNase (Thermo Fisher Scientific, Waltham, USA) treatment was given to remove DNA contamination. Quantification of total RNA was carried out using Nanodrop ND-1000 spectrophotometer (Thermo Fisher Scientific, USA). Integrity and quality of RNA were checked by electrophoresis on 1.2% denaturing agarose gel. The cDNA synthesis was performed using RevertAid first-strand cDNA synthesis kit (Thermo Fisher Scientific, Waltham, USA).
Expression analysis of candidate CcFT genes through qRT-PCR
The qRT-PCR primers were designed using primer quest software of Integrated DNA Technologies (IDT) (Table S2); and were custom synthesized using the facility of Bioengineering Services, India Private Ltd. The qRT-PCR analysis was performed using Light Cycler qRT-PCR (Roche) instrument using Brilliant III Ultra-Fast SYBR Green qPCR master mix kit (Agilent Technologies, USA). Each qRT-PCR reaction comprised of 2 µl of cDNA, 0.5 µl of each primer, 10 µl SYBR green master mix and 7.5 µl DEPC water. The Actin gene-specific primers (C. cajan) were taken as an internal control. The reaction condition was set at 94 °C for 3 min, followed by 40 cycles of 94 °C for 30 s, 60 °C for 15 s and 72 °C for 20 s, to carry out quantification of expressed mRNA. Similar reaction conditions were followed for melting curve analysis (56 °C to 95 °C) with increasing annealing temperature at 0.5 °C every 10 s.
Results
Screening of photoperiod insensitive genotypes of C. cajan
A set of twelve genotypes of C. cajan were grown in two plant growth chambers in three replications under controlled environmental conditions. The environmental conditions viz temperature and humidity were set uniformly for both the growth chambers except photoperiod. The photoperiod of growth chamber ‘A’ was set for 12 /12 h (light/dark) and chamber ‘B’ for16hr/8 hr (light/dark) and considered as SD and LD photoperiod respectively. Observations were recorded on days to flowering (DF) with the hypothesis that photoperiod sensitive genotypes will flower under SD conditions while photoperiod insensitive genotypes will flower under both SD and LD condition irrespective of photoperiod duration. The experiments revealed that out of the 12 accessions, three accessions namely ICP14923, ICP14952, and ICP20338 flowered after 96, 86, and 55 days, respectively, under both short-day as well as long-day conditions while the rest of the nine accessions flowered only under short-day conditions (Table 1).
Genome-wide identification and characterization of PEBP family genes
A total of 13 PEBP family genes were identified through genome-wide screening of C. cajan genome and they were arranged in ascending order and depicted in Fig. 1 (detail information is provided in Table S3). Phylogenetic analysis of C. cajan PEBP genes along with A. thaliana formed 3 clusters in which 8 genes clustered together with Arabidopsis AtFT, 4 genes clustered together with Arabidopsis AtTFL1 gene and the remaining 1 gene showed significant divergence and formed a separate clade and also exhibited maximum similarity with Arabidopsis AtMFT. The C. cajan PEBP genes were subgrouped into three classes viz. CcMFT, CcTFL1 and CcFT and renamed as CcMFT, CcTFL1-CcTFL4 and CcFT1-CcFT8 on the basis of their phylogenetic similarities with well characterized PEBP family of A. thaliana (Fig. 2).
The phylogenetic tree of C. cajan and A. thaliana PEBP proteins generated using the maximum likelihood method implemented in the MEGA 10.0.5 software with 1000 bootstrap iteration, JTT model and complete deletion option. The dotted line of clades depicts A. thaliana protein while solid line depicts C. cajan protein. Each sub group has been shown with different colour
Domain and Motif prediction in C. cajan PEBP genes
All the 13 C. cajan PEBP family proteins were subjected to domain prediction analysis using ‘batch CD search’ analysis tools. The obtained results confirmed the presence of a conserved PEBP domain in all the analysed proteins (Fig. 3a). Presence of conserved motifs was identified using the MEME (Multiple Em for Motif Elicitation) analysis tool. MEME motif analysis revealed the presence of around 1–6 different motifs present in each PEBP genes. The CcFT4 was observed to be the smallest (79 amino acid) protein in the PEBP family which possesses a single motif while the CcMFT was observed to be the longest (187 amino acid) protein possessing 6 motifs (Fig. 3b).
Phylogenetic analysis of PEBP family genes with other legumes
The phylogenetic relationship of the C. cajan PEBP family genes with other legume species was investigated a total of 109 PEPB protein sequences obtained from C. cajan (13), V. angularis (10), G. max (24), M. truncatula (11) V. radiata (6), C. arietinum (8), V. unguiculata (13), O. sativa (18) and A. thaliana (6) which were subjected to multiple sequence alignment followed by phylogenetic tree construction. In the phylogenetic tree, all the MFT (11 genes) were clustered together, similarly all the TFL1 (39 genes) and FT (58 genes) formed separate clusters while Arabidopsis BFT protein showed significant divergence from all the three clusters and aligned on thhhe separate clade of phylogenetic tree. Single copy of MFT was present in A. thaliana, M. truncatula, G. max, C. cajan and V. angularis, two copies of MFT in C. arietinum, V. unguiculata and O. sativa while no MFT gene was found to be present in V. radiata. The absence of MFT gene in V. radiata could be likely due to either insufficient coverage of the genome or due to its absence in the genome. In the TFL1 subgroup, AtTFL1 and AtACT/CEN were from Arabidopsis, four genes each were from C. cajan, M. truncatuala, C. arietinum, V. radiata, V. angularis, V. unguiculata and O. sativa, while nine genes from G. max. Presence of more numbers of TFL genes in Glycine max reflects its complex and larger genome. The FT subgroup of phylogenetic tree composed of largest number of genes including AtFT and AtTSF from Arabidopsis, 8 genes from C. cajan, 11 from O. sativa, 14 from G. max, 6 from M. truncatula, 7 from V. unguiculata, 5 from V. angularis, 3 from V. radiata and 2 from C. arietinum (Fig. 4).
Maximum likelihood phylogenetic tree of 109 PEBP genes of C. cajan and six legume species. The phylogenetic tree was constructed using MEGA-10.0.5 software with 1000 bootstrap iteration, JTT model and complete deletion option. Distribution of C. cajan PEBP genes among the various clades are highlighted with blue fonts
Divergence time estimation among the PEBP family members
The time tree of PEBP family genes showed an evolutionary relationship and divergence time in million years ago (mya) among the members. The time tree revealed that CcMFT was the evolutionary ancestral in PEBP family, and all the other members evolved subsequently with the course of time. The two lineages, CcTFL1 and CcFT diverged from the common ancestral CcMFT around 0.20 million years ago (mya). Among the TFL1 members divergence point of CcTFL1_2, CcTFL1_3 and CcTFL1_1 were 0.11, 0.07 and 0.04 mya, respectively. Similarly, among the CcFTs, CcFT3 and CcFT6 diverged 0.20 mya, CcFT4 and CcFT5 at 0.17 mya, CCFT8 at 0.16 mya, CcFT2 at 0.10 mya, CcFT7 and CcFT1 at 0.07 mya. The divergence time between CcFT3 and CcFT6 and between CcFT4 and CcFT5 was 0.03 mya and 0.02 mya, respectively (Fig. 5).
Expression analysis of CcFT family genes using Gene Expression Atlas of C. cajan
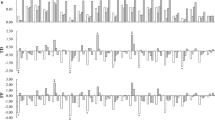
Gene expression Atlas-based analysis of PEBP family genes across the 30 tissues collected from various growth and development stages of C. cajan revealed that CcMFT showed higher expression during seed and pod development. Similarly, CcTFL1, CcTFL2 and CcTFL3 exhibited upregulated expression in radicle, root tissues, reproductive petioles and during senescence stage, whereas CcTFL4 was upregulated in immature seeds and pod tissues. The CcFT1 gene was upregulated in cotyledon, reproductive stem, petiole, root and radicle. The CcFT3 gene was expressed in the post-reproductive phase of the plant life cycle. The CcFT4 gene was observed to be expressed in 20 tissues but its elevated expression was observed in embryo, hypocotyl, shoot apical meristem, seedling shoot, hypocotyls and vegetative leaf. The CcFT5 gene showed upregulated expression in all the flower tissues, pods and reproductive leaf. The CcFT7 gene was expressed in cotyledon, nodule and senescence leaf. The CcFT6 gene was highly upregulated in reproductive sepal, senescence leaf and reproductive leaf. The CcFT8 gene was expressed in senescence and reproductive leaf. It was found that CcFT6 and CcFT8 were highly expressed in reproductive leaf whereas they remained unexpressed in vegetative leaf (Fig. 6).
Validation of expression pattern FT genes using qRT-PCR
The expression pattern of two probable flowering inducible CcFT genes viz., CcFT6 and CcFT8 were analysed using qRT-PCR analysis between RL and VL leaf tissues harvested from ICP20338 and MAL 3 grown under short day and long day photoperiod. The fold change expression in RL tissues was calculated with reference to the expression level present in VL tissue using delta-delta Ct method. The qRT-PCR based expression analysis revealed that CcFT6 was highly upregulated in RL tissues under SD and remained unexpressed under LD in both genotypes ICP20338 and MAL3 (Fig. 7a). Similarly, the expression pattern analysis of CcFT8 showed upregulated expression in RL tissue only under LD in ICP20338 (Fig. 7b).
Discussion
C. cajan is a quantitative short-day plant, requiring less than 12 h of photoperiod for induction of flowering (Carberry et al. 1998). The photoperiod requirement for the induction of flowering, limits its cultivation to specific seasons and in a particular geographical region. For the expansion of pigeon pea cultivation from a single season to multiple seasons and to diverse latitudes, the understanding of the molecular mechanism behind photoperiod sensitivity will provide the necessary molecular resources.
In previously reported studies, it has been observed that FT is an important component for the induction of photoperiod dependent flowering (Song et al. 2015). The FT genes are members of the PEBP family responsible for ‘florigen’ production (Higuchi 2018). Hence, to understand the molecular mechanism behind the photoperiod dependent flowering, we focussed on the PEBP family genes to identify probable CcFT genes responsible for the production of florigen in the C. cajan.
Genome-wide identification of PEBP family genes revealed that a total of thirteen members of PEBP family genes are present in the C. cajan genome. Upon protein domain analysis, it was found that all the thirteen members of C. cajan PEBP family genes possess phosphatidyl-ethanolamine-binding protein (PEBP) domain. It has been reported that PEBP family genes are involved in multiple functions like flower induction, seed dormancy, seed germination, embryogenesis, stress tolerance, hormone signalling, tuberization and controlling plant architecture (Chen et al. 2018; Karlgren et al. 2011; Navarro et al. 2011; Pin and Nilsson 2012; Li et al. 2014; Kwasniewski et al. 2016). To carry out such diverse roles, PEBP family genes diverged from one another during evolution (Wickland and Hanzawa 2015). Based on the evolutionary and functional relationship, PEBP family genes were classified into three subgroups, namely, FLOWERING LOCUS T (FT), mother of FT and TLF1 (MFT) and TERMINAL FLOWER 1 (TFL1) (Karlgren et al. 2011). Each class of proteins has a specialized function; for example, MFT has a role in seed embryogenesis and hormone signalling; FT acts as a positive regulator of flowering while TLF1 acts as a negative regulator (Kobayashi et al. 1999; Yamaguchi et al. 2005; Banfield and Brady 2000; Hedman et al. 2009).
Phylogenetic analysis of PEBP family of genes with other legumes revealed that most of the analyzed legume species carries low copy (either one or two) number of MFT genes except V. radiata which does not possess any MFT gene. MFT is considered as the least evolved and phylogenetically ancestral in the PEBP family and FT and TFL1 has been reported to be evolved from MFT with the course of time (Karlgren et al. 2011). It was found that almost all the analysed legume species except G. max possesses 4 TFL1 genes. Retention of an equal number of TFL1 gene in all the legume species despite variable rate of divergence from common ancestor indicates its essential role in legume growth and development. The FT clade of phylogenetic tree accommodates the largest number of genes; the FT genes from O. sativa formed three separate sub clusteres while genes from legumes were observed to be grouped together, thereby indicating their closeness to each other.
The results of gene expression analysis revealed that CcMFT exhibited higher expression during seed and pod development and might be involved in seed development and germination related activities. Similar kind of expression pattern of MFT was observed in Dimocarpus longan Lour and found to be actively involved in embryogenesis, stress tolerance and hormone signalling (Chen et al. 2018). Similarly, CcTFL1 showed upregulated expression in radicle, root tissues, reproductive petioles and during senescence stage. TFL1 is known to be a negative regulator of flower induction and its upregulation in the root, radicle and vegetative tissues indicate its role in suppression of flowering in these tissues. In Chenopodium rubrum, it is reported that darkness activates CrTFL1 expression in roots (Drabesova et al. 2016). Upregulated expression of CcTFL1, CcTFL2 and CcTFL3 in root tissues including radicle and root nodules showed their involvement in the suppression flowering in these underground tissues (Danilevskaya et al. 2008). Among the 8 CcFTs, CcFT4 was expressed irrespective of stage and tissues indicating their involvement in the basic metabolism required throughout the plant life cycle. CcFT7 was found to be expressed in nodule and may be involved in the modulating the growth of root and root nodules. In Solanum tuberosum, FT homolog StSPA6 synthesize ‘tuberigen’ for induction of tuberization process (Navarro et al. 2011). Further, it was observed that CcFT6 and CcFT8 were highly expressed in reproductive leaf whereas they remain unexpressed in vegetative leaf. In the earlier studies, it was reported that the FT gene is transcribed and translated in phloem companion cells of leaf and translocated through the phloem to the shoot apex (Chen et al. 2018). In our study, it was observed that CcFT6 and CcFT8 expression is upregulated in reproductive leaf tissues as compared to vegetative leaf tissues indicating their probable role in flower induction. Hence, these two CcFT genes were further selected for validation of their expression pattern in RL and VL tissues of MAL3 and ICP20338 using qRT-PCR.
The expression pattern of two probable flowering inducible CcFT genes viz., CcFT6 and CcFT8 revealed that CcFT6 was highly upregulated in RL tissue under SD and remained unexpressed under LD in both genotypes ICP20338 and MAL3, while CcFT8 showed upregulated expression in RL tissue only under LD grown ICP20338. The observed results suggest that in MAL3 genotype under SD photoperiod CcFT6 is actively involved in ‘florigen’ production probably leading to flower induction, while under LD photoperiod no ‘florigen’ production seems to occur, as a result plant remains in the vegetative phase. In the case of ICP20338, both CcFT6 and CcFT8 showed upregulated expression in SD and LD respectively, hence ‘florigen’ production could have occurred at both SD and LD photoperiod resulting in flowering in both conditions irrespective of day length. Rice is a SD plant having two florigen producer genes viz., Hd3a and RFT1; under SD photoperiod condition both Hd3a and RFT1 expressed and induced flowering, while under LD photoperiod only RFT1 was responsible for the induction of flowering (Komiya et al. 2008, 2009). No significant similarities were found between the protein sequences of OsRFT1 and CcFT8.
Conclusion
A total 13 PEBP family genes are present in the C. cajan genome. These genes were further sub-grouped into three categories CcMFT(1), CcTFL1(4) and CcFT(8). Among the 8 CcFT genes, CcFT6 and CcFT8 could be the major florigen (FT) producing genes responsible for induction of flowering in C. cajan based on their expression analysis. CcFT6 is expressed under SD photoperiod in both MAL3 and ICP20338 while CcFT8 expresses only under LD in ICP20338. Hence, in MAL3, ‘florigen’ is only synthesized under SD, whereas in ICP20338 ‘florigen’ was synthesized in both under SD and LD photoperiod. As a result, MAL3 flowered under SD while ICP20338 flowered in both LD and SD irrespective of day length. Accumulating more molecular evidence for the role of these genes in the initiation of flowering under SD and LD conditions would prove valuable identification of the candidate genes, their regulatory pathways and aid in the understanding of the photoperiod dependent flowering pathways in pigeonpea.
References
Argiriou A, Michailidis G, Tsaftaris AS (2008) Characterization and expression analysis of TERMINAL FLOWER1 homologs from cultivated alloteraploid cotton (Gossypium hirsutum) and its diploid progenitors. J Plant Physiol 165:1636–1646. https://doi.org/10.1016/j.jplph.2007.10.013
Banfield M, Brady R (2000) The structure of Antirrhinum centroradialis protein (CEN) suggests a role as a kinase regulator, Edited by I. A Wilson J Mol Biol 297:1159–1170. https://doi.org/10.1006/jmbi.2000.3619
Banfield MJ, Barker JJ, Perry ACF, Brady RL (1998) Function from structure? The crystal structure of human phosphatidylethanolamine-binding protein suggests a role in membrane signal transduction. Structure 6:1245–1254. https://doi.org/10.1016/S0969-2126(98)00125-7
Carberry PS, Ranganathan R, Reddy LJ et al (1998) Predicting growth and development of pigeonpea: Flowering response to photoperiod. F Crop Res 69:151–162. https://doi.org/10.1016/S0378-4290(00)00138-6
Chen Y, Xu X, Chen X et al (2018) Seed-specific gene MOTHER of FT and TFL1(MFT)involved in embryogenesis, hormones and stress responses in Dimocarpus longan Lour. Int J Mol Sci. https://doi.org/10.3390/ijms19082403
Cheng X, Li G, Tang Y, Wen J (2018) Dissection of genetic regulation of compound inflorescence development in Medicago truncatula. Development 145:158766. https://doi.org/10.1242/dev.158766
Corbesier L, Vincent C, Jang S et al (2007) FT protein movement contributes to long-distance signaling in floral induction of Arabidopsis. Science 316:1030–1033. https://doi.org/10.1126/science.1141752
Danilevskaya ON, Meng X, Hou Z et al (2008) A genomic and expression compendium of the expanded PEBP gene family from maize. Plant Physiol 146:250–264. https://doi.org/10.1104/pp.107.109538
David KM, Armbruster U, Tama N, Putterill J (2006) Arabidopsis GIGANTEA protein is post-transcriptionally regulated by light and dark. FEBS Lett 580:1193–1197. https://doi.org/10.1016/j.febslet.2006.01.016
Drabesova J, Cerna L, Masterova H et al (2016) The evolution of the FT/TFL1 Genes in Amaranthaceae and their expression patterns in the course of vegetative growth and flowering in Chenopodium rubrum. G3 6:3065–3076. https://doi.org/10.1534/g3.116.028639
FAOSTAT (2017) FAOSTAT. https://www.fao.org/faostat/en/#home. Accessed 19 Jan 2019
Fornara F, Panigrahi KCS, Gissot L et al (2009) Arabidopsis DOF transcription factors act redundantly to reduce CONSTANS expression and are essential for a photoperiodic flowering response. Dev Cell 17:75–86. https://doi.org/10.1016/j.devcel.2009.06.015
Harig L, Beinecke FA, Oltmanns J et al (2012) Proteins from the FLOWERING LOCUS T-like subclade of the PEBP family act antagonistically to regulate floral initiation in tobacco. Plant J 72:908–921. https://doi.org/10.1111/j.1365-313X.2012.05125.x
Hedman H, Källman T, Lagercrantz U (2009) Early evolution of the MFT-like gene family in plants. Plant Mol Biol 70:359–369. https://doi.org/10.1007/s11103-009-9478-x
Hengst U, Albrecht H, Hess D, Monard D (2001) The phosphatidylethanolamine-binding protein is the prototype of a novel family of serine protease inhibitors. J Biol Chem 276:535–540. https://doi.org/10.1074/jbc.M002524200
Higuchi Y (2018) Florigen and anti-florigen: Flowering regulation in horticultural crops. Breed Sci 68:109–118. https://doi.org/10.1270/jsbbs.17084
Izawa T (2007) Daylength measurements by rice plants in photoperiodic short‐day flowering. In: International review of cytology. pp 191–222
Jeong S, Clark SE (2005) Photoperiod regulates flower meristem development in Arabidopsis thaliana. Genetics 169:907–915. https://doi.org/10.1534/genetics.104.033357
Karlgren A, Gyllenstrand N, Källman T et al (2011) Evolution of the PEBP gene family in plants: functional diversification in seed plant evolution. Plant Physiol 156:1967–1977. https://doi.org/10.1104/pp.111.176206
Kobayashi Y, Kaya H, Goto K et al (1999) A pair of related genes with antagonistic roles in mediating flowering signals. Science 286:1960–1962
Komiya R, Ikegami A, Tamaki S et al (2008) Hd3a and RFT1 are essential for flowering in rice. Development 135:767–774. https://doi.org/10.1242/dev.008631
Komiya R, Yokoi S, Shimamoto K (2009) A gene network for long-day flowering activates RFT1 encoding a mobile flowering signal in rice. Development 136:3443–3450. https://doi.org/10.1242/dev.040170
Kwasniewski M, Daszkowska-Golec A, Janiak A et al (2016) Transcriptome analysis reveals the role of the root hairs as environmental sensors to maintain plant functions under water-deficiency conditions. J Exp Bot 67:1079–1094. https://doi.org/10.1093/jxb/erv498
Li Q, Fan C, Zhang X et al (2014) Identification of a Soybean MOTHER of FT and TFL1 homolog involved in regulation of seed germination. PLoS ONE 9:1–9. https://doi.org/10.1371/journal.pone.0099642
Meng X, Muszynski MG, Danilevskaya ON (2011) The FT like ZCN8 gene functions as a floral activator and is involved in photoperiod sensitivity in maize. Plant Cell 23:942–960. https://doi.org/10.1105/tpc.110.081406
Nakamichi N, Kiba T, Henriques R et al (2010) PSEUDO-RESPONSE REGULATORS 9, 7, and 5 are transcriptional repressors in the Arabidopsis Circadian Clock. Plant Cell 22:594–605. https://doi.org/10.1105/tpc.109.072892
Navarro C, Abelenda JA, Cruz-Oró E et al (2011) Control of flowering and storage organ formation in potato by FLOWERING LOCUS T. Nature 478:119–122. https://doi.org/10.1038/nature10431
Niwa Y, Ito S, Nakamichi N et al (2007) Genetic linkages of the circadian clock-associated genes, TOC1, CCA1 and LHY, in the photoperiodic control of flowering time in Arabidopsis thaliana. Plant Cell Physiol 48:925–937. https://doi.org/10.1093/pcp/pcm067
Ortega R, Hecht VFG, Freeman JS et al (2019) Altered Expression of an FT cluster underlies a major locus controlling domestication-related changes to chickpea phenology and growth habit. Front Plant Sci 10:824. https://doi.org/10.3389/fpls.2019.00824
Pazhamala LT, Purohit S, Saxena RK et al (2017) Gene expression atlas of pigeonpea and its application to gain insights into genes associated with pollen fertility implicated in seed formation. J Exp Bot 68:2037–2054. https://doi.org/10.1093/jxb/erx010
Pin PA, Nilsson O (2012) The multifaceted roles of FLOWERING LOCUS T in plant development. Plant, Cell Environ 35:1742–1755. https://doi.org/10.1111/j.1365-3040.2012.02558.x
Samach A, Onouchi H, Gold SE et al (2000) Distinct roles of CONSTANS target genes in reproductive development of Arabidopsis. Science 288:1613–1616
Saunders PTK, McKinnell C, Millar MR et al (1995) Phosphatidylethanolamine binding protein is an abundant secretory product of haploid testicular germ cells in the rat. Mol Cell Endocrinol 107:221–230. https://doi.org/10.1016/0303-7207(94)03447-2
Sawa M, Nusinow DA, Kay SA, Imaizumi T (2007) FKF1 and GIGANTEA complex formation is required for day-length measurement in Arabidopsis. Science 318:261–265. https://doi.org/10.1126/science.1146994
Sinha SK (1977) Food legumes: distribution, adaptability and biology of yield. Food Legum Distrib Adapt Biol yield
Song YH, Smith RW, To BJ et al (2012) FKF1 conveys timing information for CONSTANS stabilization in photoperiodic flowering. Science 336:1045–1049. https://doi.org/10.1126/science.1219644
Song YH, Shim JS, Kinmonth-Schultz HA, Imaizumi T (2015) Photoperiodic flowering: time measurement mechanisms in leaves. Korea Annu Rev Plant Biol 66:441–464. https://doi.org/10.1146/annurev-arplant-043014-115555
Sun C, Fang J, Zhao T et al (2012) The Histone methyltransferase SDG724 mediates H3K36me2/3 deposition at MADS50 and RFT1 and promotes flowering in rice. Plant Cell 24:3235–3247. https://doi.org/10.1105/tpc.112.101436
Tamaki S, Matsuo S, Wong HL et al (2007) Hd3a protein is a mobile flowering signal in rice. Science 80(316):1033–1036. https://doi.org/10.1126/science.1141753
Tamura K, Battistuzzi FU, Billing-Ross P et al (2012) Estimating divergence times in large molecular phylogenies. Proc Natl Acad Sci USA 109:19333–19338. https://doi.org/10.1073/pnas.1213199109
Tamura K, Stecher G, Peterson D et al (2013) MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Taoka K, Ohki I, Tsuji H et al (2011) 14–3-3 proteins act as intracellular receptors for rice Hd3a florigen. Nature 476:332–335. https://doi.org/10.1038/nature10272
Voorrips RE (2002) MapChart: Software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78. https://doi.org/10.1093/jhered/93.1.77
Wickland DP, Hanzawa Y (2015) The FLOWERING LOCUS T/TERMINAL FLOWER 1 Gene Family: functional evolution and molecular mechanisms. Mol Plant 8:983–997. https://doi.org/10.1016/j.molp.2015.01.007
Wolabu TW, Zhang F, Niu L et al (2016) Three FLOWERING LOCUS T -like genes function as potential florigens and mediate photoperiod response in sorghum. New Phytol 210:946–959. https://doi.org/10.1111/nph.13834
Yamaguchi A, Kobayashi Y, Goto K et al (2005) TWIN SISTER OF FT (TSF) acts as a floral pathway integrator redundantly with FT. Plant Cell Physiol 46:1175–1189. https://doi.org/10.1093/pcp/pci151
Zhang L, Li Q, Dong H et al (2015) Three CCT domain-containing genes were identified to regulate heading date by candidate gene-based association mapping and transformation in rice. Sci Rep 5:7663. https://doi.org/10.1038/srep07663
Acknowledgements
We acknowledge the support provided by Director, ICAR-NIPB, Post Graduate School, ICAR-IARI. We acknowledge the support extended by the National Phytotron Facility, ICAR- IARI, New Delhi for providing plant growth chambers with controlled environment condition.
Author information
Authors and Affiliations
Contributions
KG and KT conceived and designed the study, KT analysed all the expression data and conducted all the wet lab experiments, KD provided the samples for real-time analysis, AD and KK, HS performed qRT-PCR and data analysis, AMSV helped in data interpretation, KT, AD, HS, KK, KD, Sandhya, AMSV, PKJ and KG compiled and interpreted the data and wrote the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declares no competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tribhuvan, K.U., Das, A., Srivastava, H. et al. Identification and characterization of PEBP family genes reveal CcFT8 a probable candidate for photoperiod insensitivity in C. cajan. 3 Biotech 10, 194 (2020). https://doi.org/10.1007/s13205-020-02180-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-020-02180-x