Abstract
Endophytic fungi (EPF) are an important contributor to fungal diversity. It is surmised that EPF colonizing plant roots have high diversity. This study aimed to alleviate the scarcity of information regarding EPF in tropical forests, by isolationg and identifying EPF from a tropical forests in Indonesia. Soils were collected from five forests: (1) Tectona grandis monoculture; (2) Swietenia macrophylla monoculture; (3) Gmelina sp., Artocarpus champeden, Dipterocarp mixed; (4) Dipterocarp primary; (5) Macaranga sp. secondary. Four trees (Calliandra calothyrsus, Paraserianthes falcataria, Sesbania grandiflora, and Cassia siamea) and three crops (Sorghum bicolor, Allium fistulosum, and Trifolium repens) were grown in the forest soils to trap EPF. EPF were isolated from roots and isolation rates were calculated. Based on the isolation rates, P. falcataria and S. bicolor were chosen and grown again in forest soils. EPF were isolated and identified by their rDNA ITS1 region. Twelve and 21 EPF were isolated from 250 roots of P. falcataria and 300 roots of S. bicolor, respectively. Identified EPF were from genera Acrocalymma, Fusarium, Tolypocladium, Penicillium, Talaromyces, Exophiala, Dictyosporium, Pseudochaetosphaeronema, Mariannaea, Trichoderma, and Mycoleptodiscus. Acrocalymma, Tolypocladium, Penicillium, Exophiala, Pseudochaetosphaeronema, Mariannaea, and Mycoleptodiscus spp. were isolated from only one forest. Fusarium, Talaromyces, and Trichoderma spp. were isolated from more than one forest. The numbers of EPF isolated from Gmelina sp., Artocarpus champeden, Dipterocarp mixed forest, and Macaranga sp. secondary forest were higher than those from other forests, suggesting that different plant species in forests affect the root EPF community.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Endophytic fungi (EPF) colonize plant tissue without causing any visible disease symptoms at any particular moment (Schulz and Boyle 2005). Practically, EPF colonize almost any plant tissue, including leaf, stem, and root (Rodriguez et al. 2009). Since their discovery, EPF have been studied in many types of plants, including non-vascular, such as mosses (e.g., Schulz et al. 1993) and algae (e.g., Zuccaro et al. 2008), and vascular, such as shrubs (e.g., Schulz et al. 1993) and trees (e.g., Arnold and Lutzoni 2007). Most of the isolated EPF belong to the Ascomycota or Basidiomycota (Schulz and Boyle 2005).
A conservative estimate of worldwide fungal diversity has been proposed as 1.5 million species, which has been accepted as the working hypothesis and basis for the discovery of more fungal species (Hawksworth 2001). With only around 72,000 described species known so far, more than 1 million species are waiting to be found. EPF have been found in many plant species and are considered an important component of fungal diversity (Rodriguez et al. 2009). Despite the increasing number of studies of EPF in many countries, those based in the tropics are still lacking.
Arnold et al. (2000) isolated 418 EPF morphospecies colonizing the leaves of two understory tree species in a tropical forest in Panama, 59% of which were represented by single isolates. Cannon and Simmons (2002) isolated 64 EPF morphospecies colonizing the leaves of 12 tree species in a tropical forest in Guyana, 29 of which were from single leaf samples. These two studies reflect the high diversity of EPF colonizing tree leaves in tropical forests.
The different environmental factors in forests are expected to influence fungal diversity (Saikkonen 2007). However, most studies of the diversity of EPF colonizing leaves in tropical forests are limited to one forest site (e.g., Arnold et al. 2000, 2001; Cannon and Simmons 2002). Suryanarayanan et al. (2011) compared the EPF communities of 75 dicotyledonous trees belonging to 33 families from three tropical forest types in Southern India, namely, tropical dry thorn forest, dry deciduous forest, and montane evergreen forest. The type of forest appeared to have a larger effect on shaping the EPF community than the taxonomy of the host.
Existing studies of EPF in tropical forests are limited to EPF that colonize the above-ground part of plant, particularly leaf. EPF colonizing roots, further termed as root EPF, of tropical forest trees are rarely studied. Rodriguez et al. (2009) classified root EPF into two groups: class 2 EPF, which colonize the shoot, root, and rhizome, and class 4 EPF, also termed dark septate endophyte (DSE), which colonize the root only. In a review by Jumpponen and Trappe (1998), they noted that DSEs colonized approximately 600 plant species, representing 320 genera and 114 families, highlighting the abundance of DSE. However, DSE is not the only group of root EPF, indicating the possibility of an even higher abundance of root EPF in nature.
Root EPF are considered to play an important role in plant growth, similar to mycorrhizal fungi (Jumpponen and Trappe 1998). Researchers have recorded evidence about the growth promoting ability of several class 4 EPF. Exophiala pisciphila improved tolerance of maize (Zea mays L.) to cadmium toxicity (Wang et al. 2016). Leptodontidium orchidicola and Phialophora mustea improved growth of Tasmanian blue gum (Eucalyptus globulus) and birch (Betula pendula) (Berthelot et al. 2016). Veronaeopsis simplex suppressed Fusarium disease in Chinese cabbage (Brassica campestris) (Khastini et al. 2012). Heteroconium chaetospira promoted plant growth by increasing nitrogen uptake (Usuki and Narisawa 2007). Fusarium sp. improved plant resistance against fungal pathogen Gaeumannomyces graminis var. tritici (Maciá-Vicente et al. 2008). There have also been studies that recorded evidence about the growth promoting ability of class 2 EPF. Penicillium citrinum improved resistance of banana to a fungal pathogen, Fusarium oxysporum (Ting et al. 2012). Fusarium sp. improved resistance of eggplant to Verticillium wilt (Narisawa et al. 2002). Meta-analysis of data from temperate and boreal areas showed that root EPF colonization could have a negative, neutral, or positive effect on plant growth (Mandyam et al. 2013; Mayerhofer et al. 2013; Newsham 2011). Evidence of the positive effects of root EPF, along with the expectation of high EPF abundance in tropical forests, has underscored the necessity to conduct more studies of root EPF in tropical forests. However, studies of root EPF in tropical area remain a rarity (Mandyam and Jumpponen 2005).
The trap culture method is rarely used in the study of EPF, but is common in study of arbuscular mycorrhizal fungi (AMF). EPF and AMF have been found in the same plant community (Lingfei et al. 2005). Thus, utilization of the same plant species in trap culture for isolation of EPF is highly possible. Sorghum bicolor, Trifolium repens, and Allium fistulosum are crop species commonly used to isolate AMF (Del Val et al. 1999), thus can also be used to isolate EPF. Tree species are also a possible candidate to isolate EPF. Paraserianthes falcataria, Calliandra calothyrsus, Cassia siamea, and Sesbania grandiflora are common leguminous tree species in the tropics, particularly Indonesia. These tree species are fast-growing and are also candidate species for reforestation efforts (Otsamo et al. 1997, Wulandari et al. 2016). Considering the importance of the future utilization of EPF in reforestation, these tree species can also be used as host plants to isolate EPF. The objectives of present study were to (1) isolate root EPF from five forests soils, and identify them based on their rDNA ITS region, and (2) compare EPF community among different forests.
2 Materials and methods
2.1 Collection of forest soil
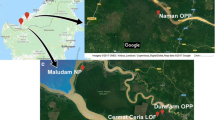
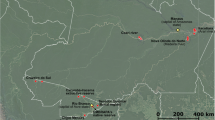
Forest soils were collected from five forests in Indonesia, with five replications in September 2012 (Table 1). Soils between 0 and 10 cm depth and within 0–5 cm distance from the root of representative seedlings, or around the main stem of representative tree species, were collected after removing the organic layer. Two kilograms of soil were collected for each replication. The distance between replications in one forest was ≥20 m. The latitude coordinates of each forest were recorded.
2.2 Chemical analysis of soil
Soil pH was measured at the soil:solution ratio of 1:2.5 in deionized water and 1 M KCl solution (Table 2). Soil available phosphate was determined following the Truog method (Truog 1930). Briefly, air-dried soil was suspended in 0.001 M sulfuric acid solution, stirred by a mechanical shaker, and filtered immediately using a filter paper (Advantec Toyo No. 6, Japan). Color development solution (Olsen and Sommers 1982) was added to the filtrate, and absorbance was measured with a spectrophotometer (U-2900, Hitachi, Japan) at 880 nm.
Cation exchange capacity (CEC) was determined by the semi-micro Schollenberger method, which is based on the displacement of soil cations by leaching soil with an excess of 1 M neutral (pH 7.0) NH4CH3COO solution, as described by the United States of Agriculture (USDA) (Soil survey staff 1992). Briefly, the soil was leached with NH4CH3COO solution, allowing NH4+ to replace soil cations. The soil was leached again with 1.35 M KCl solution to replace NH4+. By titration with formaldehyde and thymol blue, the amount of NH4+ was measured and CEC was calculated. Leached exchangeable cations (Ca, Mg, Na, and K) from the soil were measured by an atomic absorption spectrophotometer (Z-5000, Hitachi, Japan).
2.3 Experiment 1. Isolation of EPF from tree and crop species
Sand was acidified and sterilized by autoclaving at 80 °C for 45 min. Forty grams of sterilized sand was mixed with 40 g of forest soil, then used as growth medium. Seeds of Calliandra calothyrsus, Paraserianthes falcataria, Sesbania grandiflora, and Cassia siamea were sown on sterilized sand and incubated in a growth chamber (Biotron LPH-350S, NK System, Japan) at 27 °C with a 16-h photoperiod. One of two-leaf-stage seedlings of C. calothyrsus, P. falcataria, S. grandiflora, and C. siamea was transplanted onto the medium in a 50 ml syringe pot. Three, five, or 20 seeds of Sorghum bicolor, Allium fistulosum, or Trifolium repens, respectively, were sown onto the same medium. Ten grams of sterilized sand was further added into the syringe pot to cover the root system of seedlings or seeds. All plants were grown in the growth chamber for 90 days at 27 °C, with a 16-h photoperiod. Five to ten milliliters of 1 mg P L−1 nutrient solution (based on Wagatsuma et al. 1988) was applied once every two days. Twenty-five pots were prepared for each plant species.
C. calothyrsus, P. falcataria, S. grandiflora, C. siamea, S. bicolor, A. fistulosum, and T. repens were harvested 90 days after transplanting or sowing. Fresh roots were extracted from each soil and washed under running tap water. The roots were surface-sterilized following the method of Verma et al. (2012) by dipping in 90% EtOH (1 min), 5% NaClO (5 min), and 90% EtOH (10 s) and rinsing three times with sterilized deionized water. The roots were dried with sterilized Kimtowels™ and left to air dry. The air-dried roots were cut into 5 mm. Five pieces were plated on half-strength malt extract agar (MEA) containing 100 μg mL−1 Penicillin-Streptomycin (Lonza Biowhittaker, Penicillin-Streptomycin Mixture) (modified from Verma et al. 2012) and water agar. Five replication plates were made for each medium, for roots from one pot, for a total of 50 plates per pot. The plates were sealed with Parafilm™ and incubated in the dark at 25 °C. Emerging colonies from roots were subcultured on new half-strength MEA medium. EPF isolated from this experiment were used to calculate the isolation rate. Isolation rates were calculated by dividing the number of isolates by the number of initial plates.
2.4 Experiment 2. Isolation of EPF from P. falcataria and S. bicolor
EPF were isolated again from P. falcataria and S. bicolor. P. falcataria was chosen because this species would be used as the target species in future studies and the EPF isolation rates for the four tree species were not markedly different (Table 3). This species is not only a candidate for reforestation, but also economically profitable in a mixed plantation with crop species, or as single-species plantation (Siregar et al. 2007; Krisnawati et al. 2011), and a candidate for energy production (Amirta et al. 2016). S. bicolor was chosen because it had the highest EPF isolation rate among crop species, and also among all species.
EPF were isolated from P. falcataria and S. bicolor using the same method described above (Experiment 1). However, the nutrient solution for watering was changed to tap water to minimize nutrient input, mimicking the conditions in the forest. Seedling number per pot, plant height, leaf number, and symptoms of nutrient deficiency, on leaves of P. falcataria and S. bicolor were recorded before harvest (Tables S1 and S2). Plants showing good growth were harvested and EPF were isolated from the roots.
2.5 DNA extraction and amplification
EPF isolated in experiment 2 were subcultured on half-strength MEA for 1–3 weeks depending on the growth rate of each isolate. Hyphae of each isolate were collected using forceps and placed on the lid of a plastic tube containing 20 μL of InstaGene™ Matrix (Bio-Rad, USA). Hyphae were crushed with a pipet tip having a blunt end, further mixed with 180 μL of InstaGene™ Matrix, and vortexed. Next, rDNA was extracted following the manufacturer’s protocol for InstaGene™ Matrix. The extracted DNA was stored at −20 °C until use.
The internal transcribed spacer (ITS) region of the fungi was amplified using universal primers, ITS1F (5’-GTAACAAGGTTTCCGT-3′) and ITS1R (5’-CGTTCTTCATCGATG-3′) (Fujita et al. 2010), with an Expand High FidelityPLUS PCR system (Roche, Germany) at the following composition: 4 μL of 5× buffer with MgCl2, 0.2 μL of DNA polymerase, 2 μL of 2.0 mM dNTP, 0.4 μL of ITS1F, 0.4 μL of ITS1R, 11 μL of Milli-Q water, and 2 μL of DNA template. The reaction was performed in a Takara PCR Thermal Cycler Dice (Model TP600, Takara Bio, Japan) under the following conditions: initial denaturation at 94 °C for 120 s; 30 cycles of denaturation at 94 °C for 30 s, annealing at 55 °C for 60 s, and extension at 72 °C for 60 s; and final extension at 72 °C for 420 s. The PCR products were separated on 1.0% agarose gel (D1 Agarose Low EEO, Conda, Spain) in 1× Tris-borate-EDTA buffer, stained with SYBR® Safe DNA Gel Stain (Invitrogen, USA), and viewed under blue light (470 nm, MBP-LED, Bio-Pyramid, USA). PCR-amplified fragments were purified using a MonoFas DNA Purification Kit (GL Science, Japan) following the manufacturer’s protocol. Purified DNA was ligated into pT7Blue T-Vector (Novagen, USA) using a DNA Ligation Kit Ver 1 (Takara Bio, Japan). Twenty microliters of IPTG (Takara Bio, Japan) and 35 μL of X-Gal (Takara Bio, Japan) were applied to Luria Bertani (LB) medium containing 100 mg L−1 ampicillin. T-Vector containing DNA was transformed into Escherichia coli JM109 (Takara Bio, Japan) by plating onto this LB medium. Plates with E. coli were incubated at 37 °C for 16 h.
Single colonies of E. coli were collected and DNA was amplified using primers T7 (5’-TAATACGACTCACTATAG-3′) and U19 (5’-GTTTTCCCAGTCACGACT-3′) (Ikenaga et al. 2016) with GoTaq® DNA Polymerase (Promega, USA) at the following composition: 2 μL of 5 x reaction buffer, 0.05 μL of DNA polymerase, 0.8 μL of 2.0 mM dNTP, 0.2 μL of T7, 0.2 μL of U19, 6.75 μL of Milli-Q water, and a single colony of E. coli. The reaction was performed in a Takara PCR Thermal Cycler Dice (Model TP600, Takara Bio, Japan) under the following conditions: initial denaturation at 94 °C for 120 s; 35 cycles of denaturation at 94 °C for 15 s, annealing at 50 °C for 60 s, and extension at 72 °C for 80 s; and final extension at 72 °C for 600 s. The PCR products were separated on 1.0% agarose gel in 1× Tris-borate-EDTA buffer, stained with SYBR® Safe DNA Gel Stain, and viewed under blue light. The PCR products were used for sequencing.
Sequencing reactions were performed in a Bio-Rad DNA Engine Dyad PTC-220 Peltier Thermal Cycler using an ABI BigDye™ Terminator v3.1 Cycle Sequencing Kit with AmpliTaq DNA Polymerase (FS enzyme, Applied Biosystems, Japan) following the manufacturer’s protocol. Single pass sequencing was performed on each DNA template using a T7 promoter. Fluorescent-labeled fragments were purified from the unincorporated terminators by adopting an ethanol precipitation protocol. The samples were resuspended in distilled water and subjected to electrophoresis in an ABI 3730xl sequencer (Applied Biosystems, Japan).
2.6 Phylogenetic analyses
Sequences of EPF isolates were submitted for BLAST analysis (Altschul et al. 1990). The sequences and their corresponding BLAST top hits were aligned by Multiple Alignment using Fast Fourier Transform (MAFFT) (Katoh et al. 2002) through http://guidance.tau.ac.il.. Maximum parsimony method was performed by MEGA 7 (www.megasoftware.net) with 1000 replications of bootstrap analysis (Fig. 1).
3 Results
3.1 Soil chemical properties
Soil pH (H2O) ranged from 3.87 to 7.02 and soil pH (KCl), from 3.26 to 5.99 (Table 2). Based on USDA classification, soil pH (H2O) was neutral in both Tectona grandis and Swietenia macrophylla monoculture. Soil pH (H2O) was extremely acidic to very strongly acidic in Gmelina sp., Artocarpus champeden, and Dipterocarp mixed, Dipterocarp primary, and Macaranga sp. secondary forests. Available P concentration was 1.00 mg 100 g dry soil−1 in T. grandis monoculture and 0.99 mg 100 g dry soil−1 in S. macrophylla monoculture, and both values were higher than those in Gmelina sp., A. champeden, and Dipterocarp mixed, Dipterocarp primary and Macaranga sp. secondary forests. CEC ranged from 9.34 to 75.39 cmol kg−1; it was higher in T. grandis monoculture and S. macrophylla monoculture than in Gmelina sp., A. champeden, and Dipterocarp mixed, Dipterocarp primary or Macaranga sp. secondary forest. The same tendency as soil CEC was noted for exchangeable Ca, Mg, Na, and K.
3.2 Isolation of EPF
A total of 197 EPF were isolated from the roots of the seven host plant species grown in the five forest soils (Table 3). Among the 197 EPF, 13, 9, 17, 21, 75, 3, and 59 EPF were isolated from the roots of C. calothyrsus, P. falcataria, S. grandiflora, C. siamea, S. bicolor, A. fistulosum, and T. repens, respectively. S. bicolor and T. repens had higher EPF isolation rates than the other plant species. Moreover, among these, 197 EPF, 52, 55, 53, 13, and 24 EPF were isolated from T. grandis monoculture, S. macrophylla monoculture, Gmelina sp., A. champeden, and Dipterocarp mixed, Dipterocarp primary, and Macaranga sp. secondary forest, respectively. Soils of T. grandis monoculture, S. macrophylla monoculture, and Gmelina sp., A. champeden, and Dipterocarp mixed forest had higher EPF isolation rates than the other forest soils.
Leaf necrosis was detected in some seedlings of S. bicolor. Seedlings with higher plant height and leaf number, and less leaf necrosis, in 12 of a total 25 pots, were selected and used for isolation of EPF (Table S1). Twenty-one EPF were isolated from the roots of S. bicolor (Tables 4 and S1). Among the 21 EPF, 5, 2, 7, and 7 were isolated from soils of T. grandis monoculture, S. macrophylla monoculture, Gmelina sp., A. champeden, and Dipterocarp mixed, and Macaranga sp. secondary forest, respectively. EPF isolation rate for S. bicolor was 35%.
Leaf necrosis was not detected in all seedlings of P. falcataria. Seedlings with higher plant height and leaf number in 10 of a total 25 pots, were selected and used for isolation of EPF (Table S2). Twelve EPF were isolated from the roots of P. falcataria (Tables 4 and S2). Among the 12 EPF, 2, 7, and 3, were isolated from soils of T. grandis monoculture, Gmelina sp., A. champeden, and Dipterocarp mixed, and Macaranga sp. secondary forest, respectively. The EPF isolation rate for P. falcataria was 24%, which was higher than that of the first isolation of EPF.
3.3 Identification of EPF
The sequences of the ITS regions of all isolates were submitted to BLAST including uncultured or environmental samples. The similarity scores between isolated EPF and their closest relatives in GenBank were between 84 and 100% (Table 4). All sequences of isolated EPFs were deposited in the DNA Data Bank of Japan (DDBJ) under accession numbers LC334061–LC334093.
Seven genotypes of EPF isolated from the roots of P. falcataria closely matched fungi, identified to the species level (Table 4). One genotype closely matched a fungus identified to the phylum level. The remaining four genotypes closely matched uncultured fungi. Based on the National Center for Biotechnology Information (NCBI) (https://www.ncbi.nlm.nih.gov) database, the identified fungi were classified in orders Pleosporales, Hypocreales, Eurotiales, and Chaetothyriales.
Nine genotypes of EPF isolated from the roots of S. bicolor closely matched fungi identified to the species level (Table 4). One genotype closely matched a fungus identified to the genus level (Fusarium). One genotype closely matched a fungus identified to the family level (Clavicipitaceae). One genotype closely matched a fungus identified to the order level (Sordariales). One genotype closely matched a fungus identified to the class level (Dothideomycetes). There was one genotype that closely matched unidentified fungus (Fungal sp. voucher). The remaining seven genotypes closely matched uncultured fungi. Based on the NCBI database, the identified fungi were classified in the orders Pleosporales, Hypocreales, Eurotiales, and Magnaporthales.
4 Discussion
4.1 Specificity of EPF for host plants and forest sites
EPF from the orders Pleosporales, Hypocreales, Eurotiales, Magnaporthales, and Chaetothyriales were isolated in the present study (Table 4). EPF from orders Pleosporales, Hypocreales, and Eurotiales were isolated from both host plants P. falcataria and S. bicolor, whereas EPF from order Magnaporthales and Chaetothyriales were specifically isolated from P. falcataria and S. bicolor, respectively. EPF in these five orders were also isolated from the roots of subtropical Hordeum murinum in Ireland (Murphy et al. 2015). EPFs from two orders Pleosporales and Hypocreales were also isolated from the roots of plants growing in places with different environmental conditions, such as desert grass Bouteloua gracilis in New Mexico (Porras-Alfaro et al. 2008) and tropical shrub Sophora tonkinensis in China (Yao et al. 2017), showing the wide distribution of EPF from these two orders in nature.
EPF from the genera Acrocalymma, Fusarium, Tolypocladium, Penicillium, Talaromyces, Exophiala, Dictyosporium, Pseudochaetosphaeronema, Mariannaea, Trichoderma, and Mycoleptodiscus, have also been reported in other studies. Those EPFs were reported in the studies by Jin et al. (2017) (Acrocalymma vagum), Kwasna et al. (2016) Talaromyces verruculosus and Trichoderma spirale, Lin et al. 2007 (Dictyosporium sp.), Shubin et al. 2014 (Mycoleptodiscus sp. and Penicillium sp.), Yao et al. 2017 (Fusarium solani), Zhang et al. 2017a (Exophiala piscipila), Waipara et al. 1996 (Mariannaea sp.), Zhang et al. 2017b (Pseudochaetosphaeronema larense), and Sánchez Márquez et al. 2010 (Tolypocladium cylindrosporum). However, to our knowledge, the present study is the first to isolate EPF in these genera from the roots of P. falcataria. Amin (2013) isolated EPF belonging to a different genus, Nigrospora sp. (order Trichosphaeriales), from the roots of P. falcataria.
Sixteen of the 33 isolates had the closest match to fungi identified to the species level (Table 4). Ninety-seven percent similarity is widely used as the cut-off point (O’Brien et al. 2005) to determine whether the isolates are identical at the species level, or not. Thirteen of the 16 isolates were considered to be of the same species with the closest match. The remaining three isolates, 2354(1)-2, 2624(5), and 2655(2), were similar to Exophiala calicioides (84% similarity), Pseudochatoesphaeronema martinelli (95% similarity), and Mycoleptodiscus terrestris (94% similarity), respectively. These three isolates may be categorized in the same genera with the closest match.
Among the 16 isolates, 3 were specific to certain forest sites shared by P. falcataria and S. bicolor: Fusarium solani in T. grandis monoculture, Talaromyces verruculosus in Gmelina sp., A. champeden and Dipterocarp mixed, and Talaromyces aculeatus in Macaranga sp. secondary forest (Table 4). In addition, some of the isolates were specific to certain forest sites, but were not shared by the two host plants, examples of which are Dictyosporium heptasporum in T. grandis monoculture, Mariannaea camptospora in Gmelina sp., A. champeden, and Dipterocarp mixed, and Mycoleptodiscus sp. in Macaranga sp. secondary forest. These results indicated that EPF had low or high specificity for host plants, as well as forest sites. This phenomenon was also observed by Kernaghan and Patriquin (2011) in their study in a Boreal area. Kernaghan and Patriquin (2011) isolated root EPF from Betula papyrifera, Abies balsamea, and Picea glauca from two different sites and identified them by a molecular method. They revealed that the EPF communities were different among host trees in one site but not in another site. They also observed that some EPF were found only on certain host, showing the specificity of EPF to host tree species. Jumpponen and Trappe (1998) and Mandyam and Jumpponen (2005) clarified that root EPF, specifically DSE, colonized 587 plants, representing 320 genera and 114 families. Further inoculation experiments under natural and experimental conditions confirmed that DSE species had low host specificity.
The numbers of isolates in Gmelina sp., A. champeden, and Dipterocarp mixed forest and Macaranga sp. secondary forests were higher than those in the other forest sites (Table 4). The dominant species in Gmelina sp., A. champeden, and Dipterocarp mixed forest were Gmelina sp., A. champeden, and Dipterocarp sp. The dominant species in T. grandis monoculture forest and S. macrophylla monoculture forests were only one species each, T. grandis and S. macrophylla, respectively. The number of plant species in each forest might be the reason why the number of isolates was higher in Gmelina sp., A. champeden, and Dipterocarp mixed forest than the other forest sites. The dominant species in Macaranga sp. secondary forest was also one species: Macaranga sp. The reason why the number of isolates was larger in Macaranga sp. secondary forest than T. grandis monoculture forest and S. macrophylla monoculture forest is not known.
Our results demonstrate that the root EPF community differed among forest sites. Utilization of the trap culture method with the same host plant, P. falcataria or S. bicolor, still yielded different EPF among the five forests. Thus, differences in the root EPF community were mainly owing to differences in forest sites, involving the plant community and environmental factors, that resulted in the specific conditions in each forest.
4.2 Role of EPF from the same genera
Some of the root EPF isolated in the present study have also been isolated and studied for their importance, especially in relation with plant growth. Some were proven to promote plant growth under biotic and abiotic stress. Trichoderma sp. improved plant resistance against fungal pathogens (Vinale et al. 2008; Mukherjee et al. 2012), plant tolerance to salt stress (Brotman et al. 2013), and plant tolerance to drought (Bae et al. 2009). Exophiala sp. improved plant tolerance to heat stress (Khan et al. 2012) and cadmium toxicity (Wang et al. 2016). Talaromyces pinophilus promoted the growth of rice seedlings by gibberellin production (Khalmuratova et al. 2015). Penicillium citrinum improved plant resistance against fungal pathogens (Ting et al. 2012) and promoted growth by gibberellin production (Khan et al. 2008). Conversely, Mycoleptodiscus sp. was recorded to be a pathogen of Eurasian watermilfoil (Shearer et al. 2011). There is no information available regarding other remaining genera.
4.3 EPF isolation rate
The isolation of root colonizing microbes by trap culture is widely used in microbiological studies. However, the isolation from field-collected plants is a more common method in EPF studies, especially in studies that aim to isolate organic compounds for biotechnological applications (Strobel 2003). The trap culture method is common for studies that aim to clarify the role of EPF in protecting plants against soil pathogens and promoting plant growth (Amin 2013; Narisawa et al. 2002, 2007) or to clarify the existence of certain EPF in the field (Ahlich et al. 1998).
Studies of root EPF by some isolation methods might underestimate the number of species compared to studies that directly assess root samples by using molecular methods. However, the isolation method has one advantage, namely, the availability of culture for further studies. Brock et al. (2009) emphasized the importance of a herbarium or a fungal culture for the detailed assessment of unknown fungi.
In the present study, the trap culture method was adopted, and EPF were isolated from seven plant species grown on five forest soils. There was no difference in the EPF isolation rate among the four tree species (Table 3), but there was a difference among crop species, with S. bicolor showing the highest isolation rate. Among the seven plant species, S. bicolor showed the highest EPF isolation rate. Narisawa et al. (2002) isolated EPF by the trap culture method using eggplant, Chinese cabbage, tomato, melon, and strawberry as host plants. Among these five plant species, eggplant showed the highest EPF isolation rate. In our study, only representatives of EPF having the same morphology which emerged from root segments were isolated. In contrast, in the study of Narisawa et al. (2002), all emerging EPF were isolated. Thus, the EPF isolation rate in our study might be very low compared to that of Narisawa et al. (2002). In addition, there were also studies that isolated EPF from plant species that were taxonomically close to plant species of the present study: Sesbania bispinosa (Sreelalitha and Sridhar 2015), P. falcataria (Amin 2013), and Trifolium subterraneum (Mugerwa et al. 2013).
P. falcataria and S. bicolor were not originally part of the plant community in the five forest sites. Nevertheless, EPF were still isolated from their roots. Moreover, different EPF were isolated from different forests, indicating that the trap culture method can be used to evaluate differences in EPF community among forests. Considering that both host plants were not part of the plant community in the five forest sites, the effect of original plant species on the identity of isolated EPF was minimal. Therefore, differences in EPF isolates among the forest sites reflect the natural differences in the EPF community among the forest sites.
4.4 Relationship between soil chemical properties and EPF community
Soil chemical properties differ between forests in Java Island and Kalimantan Island. Different isolates were found in the two islands: Fusarium solani in Java Island and Talaromyces sp. in Kalimantan Island. There were specific isolates in each forest, that were not separated by the difference of soil chemical properties of two islands, such as F. solani and Talaromyces sp. Soil chemical properties may not be an important factor determining EPF community.
Root EPF were isolated from forest soil with pH (H2O) ranging from 3.87–4.82. The present study is not the only one to find EPF in soil with low pH. Hakim et al. (2015) isolated EPF from the roots of Shorea leprosula and Shorea selanica grown in reddish Latosol, a type of acid soil in the tropics. Göransson et al. (2008) observed EPF colonization in the roots of four woodland grasses (Elymus caninus, Poa nemoralis, Deschampsia cespitosa, and Deschampsia flexuosa) grown in subtropical forest soil with pH (H2O) ranging from 4.20 to 6.00. These findings emphasize the fact that EPF are commonly found in subtropical or tropical forest soils with low pH.
Root EPF were isolated from soil with available P lower than 1.00 mg 100 g soil−1. This value indicated low P availability for plant growth, according to USDA classification and Brady and Weil (2002). Della Monica et al. (2015) inoculated T. repens with four EPF isolates and cultivated the plant in soil:perlite (2:1 v:v) containing 3.42 mg 100 g−1 total P. One of the four EPF, Phialocephala glacialis, increased both P concentration in soil solution and shoot P content in T. repens. Hiruma et al. (2016) inoculated Arabidopsis thaliana with Colletotrichum tofieldiae and cultivated the plant in half-strength Murashige and Skoog medium, containing 0.68 mg 100 g−1 total P. C. tofieldiae increased the shoot fresh weight of A. thaliana. Root EPF isolated in the present study may have the potential to mineralize P and promote plant growth.
5 Conclusion
EPF were isolated from the roots C. calothyrsus, P. falcataria, S. grandiflora, C. siamea, S. bicolor, A. fistulosum, and T. repens, grown in five different forest soils in Indonesia. EPF isolation rates were not markedly different among the tree species. S. bicolor exhibited the highest EPF isolation rate among crop species, and also among all species used in the present study. Some EPF were isolated from more than one forest, and some specifically from only one forest. In addition, the number of isolated EPF from Gmelina sp., A. champeden, and Dipterocarp mixed were larger than that from other forests. These results indicated that EPF had specificity to the plant species. Future studies including EPF inoculation of plants is necessary to clarify the effect of EPF on plant growth.
References
Ahlich K, Rigling D, Holdenrieder O, Sieber TN (1998) Dark septate hyphomycetes in swiss conifer forest soils surveyed using Norway-spruce seedlings as bait. Soil Biol Biochem 30(8–9):1069–1075
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Amin N (2013) Investigation of culture filtrate of endophytic fungi Nigrospora sp. isolate Rs 10 in different concentrations towards root-knot nematode Meloidogyne spp. Indian J Sci Technol 6(9):5177–5181
Amirta R, Yuliansyah AEM, Ananto BR, Setiyono B, Haqiqi MT, Septiana HA, Lodong M, Oktavianto RN (2016) Plant diversity and energy potency of community forest in East Kalimantan, Indonesia: Searching for fast growing wood species for energy production. Nusantara Biosci 8(1):22–31
Arnold AE, Lutzoni F (2007) Diversity and host range of foliar fungal endophytes: are tropical leaves biodiversity hotspots? Ecology 88(3):541–549
Arnold AE, Maynard Z, Gilbert GS, Coley PD, Kursar TA (2000) Are tropical fungal endophytes hyperdiverse? Ecol Lett 3:267–274
Arnold AE, Maynard Z, Gilbert GS (2001) Fungal endophytes in dicotyledonous neotropical trees: patterns of abundance and diversity. Mycol Res 105(12):1502–1507
Bae H, Sicher RC, Kim MS, Kim SH, Strem MD, Melnick RL, Bailey BA (2009) The beneficial endophyte Trichoderma hamatum isolate DIS 219b promotes growth and delays the onset of the drought response in Theobroma cacao. J Exp Bot 60(11):3279–3295
Berthelot C, Leyval C, Foulon J, Chalot M, Blaudez D (2016) Plant growth promotion, metabolite production and metal tolerance of dark septate endophytes isolated from metal-polluted poplar phytomanagement sites. FEMS Microbiol Ecol 92(10):1–14. https://doi.org/10.1093/femsec/fiw144
Brady NC, Weil RR (2002) The nature and properties of soils, 13th edn. Pearson education Inc., United States of America, pp 39–40 100–109
Brock PM, Doring H, Bidartondo MI (2009) How to know unknown fungi: the role of a herbarium. New Phytol 181:719–724
Brotman Y, Landau U, Cuadros-Inostroza Á, Takayuki T, Fernie AR, Chet I, Viterbo A, Willmitzer L (2013) Trichoderma-plant root colonization: escaping early plant defense responses and activation of the actioxidant machinery for saline stress tolerance. PLoS Pathog 9(3):e1003221. https://doi.org/10.1371/journal.ppat.1003221
Cannon PF, Simmons CM (2002) Diversity and host preference of leaf endophytic fungi in the Iwokrama Forest Reserve, Guyana. Mycologia 94(2):210–220
Del Val C, Barea JM, Azcón-Aguilar C (1999) Diversity of arbuscular mycorrhizal fungus populations in heavy-metal-contaminated soils. Appl Environ Microbiol 65(2):718–723
Della Monica IF, Saparrat MCN, Godeas AM, Scervino JM (2015) The co-existence between DSE and AMF symbionts affects plant P pools through P mineralization and solubilization processes. Fungal Ecol 17:10–17
Fujita K, Furuya S, Kohno M, Suzuki S, Takayanagi T (2010) Analysis of microbial community in Japanese vineyard soils by culture-independent molecular approach. Int J Wine Res 2:75–104
Göransson P, Olsson PA, Postma J, Falkengren-Grerup U (2008) Colonisation by arbuscular mycorrhizal and fine endophytic fungi in four woodland grasses – variation in relation to pH and aluminium. Soil Biol Biochem 40:2260–2265
Hakim SS, Budi SW, Turjaman M (2015) Phosphate solubilizing and antifungal activity of root endophyte isolated from Shorea leprosula Miq. and Shorea selanica (DC) Blume. Journal Manajemen Hutan Tropika 21(3):138–146
Hawksworth DL (2001) The magnitude of fungal diversity: the 1.5 million species estimate revisited. Mycol Res 105(12):1422–1432
Hiruma K, Gerlach N, Sacristan S, Nakano RT, Hacquard S, Kracher B, Neumann U, Ramirez D, Bucher M, O’Connell RJ, Schulze-Lefert P (2016) Root endophyte Colletotrichum tofieldiae confers plant fitness benefits that are phosphate status dependent. Cell 165:464–474
Ikenaga M, Tabuchi M, Kawauchi T, Sakai M (2016) Application of locked nucleic acid (LNA) primer and PCR clamping by LNA oligonucleotide to enhance the amplification of internal transcribed spacer (ITS) regions in investigating the community structures of plant-associated fungi. Microbes Environ 31(3):339–348
Jin HQ, Liu HB, Xie YY, Zhang YG, Xu QQ, Mao LJ, Li XJ, Chen J, Lin FC, Zhang CL (2017) Effect of the dark septate endophytic fungus Acrocalymma vagum on heavy metal content in tobacco leaves. Symbiosis 74(2):89–95 https://doi.org/10.1007/s13199-017-0485-4
Jumpponen A, Trappe JM (1998) Dark septate endophytes: a review of facultative biotrophic root-colonizing fungi. New Phytol 140:295–310
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30(14):3059–3066
Kernaghan G, Patriquin G (2011) Host associations between fungal root endophytes and boreal trees. Microb Ecol 62(2):460–473
Khalmuratova I, Kim H, Nam YJ, Oh Y, Jeong MJ, Choi HR, You YH, Choo YS, Lee IJ, Shin JH, Yoon H, Kim JG (2015) Diversity and plant growth promoting capacity of endophytic fungi associated with halophytic plants from the west coast of Korea. Mycobiology 43(4):373–383
Khan SA, Hamayun M, Yoon H, Kim HY, Suh SJ, Hwang SK, Kim JM, Lee IJ, Choo YS, Yoon UH, Kong WS, Lee BM, Kim JG (2008) Plant growth promotion and Penicillium citrinum. BMC Microbiol 8:231. https://doi.org/10.1186/1471-2180-8-231
Khan AL, Hamayun M, Waqas M, Kang SM, Kim YH, Kim DH, Lee IJ (2012) Exophiala sp. LHL08 association gives heat stress tolerance by avoiding oxidative damage to cucumber plants. Biol Fertil Soils 48:519–529
Khastini RO, Ohta H, Narisawa K (2012) The role of a dark septate endophytic fungus, Veronaeopsis simplex Y34, in Fusarium disease suppression in Chinese cabbage. J Microbiol 50(4):618–624
Krisnawati H, Varis E, Kallio M, Kanninen M (2011) (L.) Nielsen: ecology, silviculture and productivity. Center for International Forestry Research, pp 2–4
Kwasna H, Szewczyk W, Behnke-Borowczyk J (2016) Fungal root endophytes of Quercus robur subjected to flooding. For Pathol 46:35–46
Lin X, Lu C, Huang Y, Zheng Z, Su W, Shen Y (2007) Endophytic fungi from a pharmaceutical plant, Camptotheca acuminata: isolation, identification and bioactivity. World J Microbiol Biotechnol 23:1037–1040
Lingfei L, Anna Y, Zhiwei Z (2005) Seasonality of arbuscular mycorrhizal symbiosis and dark septate endophytes in a grassland site in southwest China. FEMS Microbiol Ecol 54:367–373
Maciá-Vicente JG, Jansson HB, Mendgen K, Lopez-Llorca LV (2008) Colonization of barley roots by endophytic fungi and their reduction of take-all caused by Gaeumannomyces graminis var. tritici. Can J Microbiol 54:600–609
Mandyam K, Jumpponen A (2005) Seeking the elusive function of the root-colonising dark septate endophytic fungi. Stud Mycol 53:173–189
Mandyam KG, Roe J, Jumpponen A (2013) Arabidopsis thaliana model system reveals a continuum of responses to root endophyte colonization. Fungal Biol 117:250–260
Mayerhofer MS, Kernaghan G, Harper KA (2013) The effects of fungal root endophytes on plant growth: a meta-analysis. Mycorrhiza 23:119–128
Mugerwa TTM, Saleeba JA, McGee PA (2013) A variety of melanised root-associated fungi from the Sydney basin form endophytic associations with Trifolium subterraneum. Fungal Ecol 6:70–82
Mukherjee M, Mukherjee PK, Horwitz BA, Zachow C, Berg G, Zeilinger S (2012) Trichoderma–plant–pathogen interactions: advances in genetics of biological control. Indian J Microbiol 52(4):522–529
Murphy BR, Nieto LM, Doohan FM, Hodkinson TR (2015) Profundae diversitas: the uncharted genetic diversity in a newly studied group of fungal root endophytes. Mycology 6(3–4):139–150
Narisawa K, Kawamata H, Currah RS, Hashiba T (2002) Suppression of Verticillium wilt in eggplant by some fungal root endophytes. Eur J Plant Pathol 108:103–109
Narisawa K, Hambleton S, Currah RS (2007) Heteroconium chaetospira, a dark septate root endophyte allied to the Herpotrichiellaceae (Chaetothyriales) obtained from some forest soil samples in Canada using bait plants. Mycoscience 48:274–281
Newsham KK (2011) A meta-analysis of plant responses to dark septate root endophytes. New Phytol 190:783–793. https://doi.org/10.1111/j.1469-8137.2010.03611.x
O’Brien HE, Parrent JL, Jackson JA, Moncalvo JM, Vilgalys R (2005) Fungal community analysis by large-scale sequencing of environmental samples. Appl Environ Microbiol 71(9):5544–5550
Olsen SR, Sommers LE (1982) Phosphorus. In: Page AL (ed) Methods of soil analysis. Part 2. Chemical and microbiological properties. Soil science society of America, Madison, pp 403–430
Otsamo A, Adjers G, Hadi TS, Kuusipalo J, Vuokko R (1997) Evaluation of reforestation of 83 tree species planted on Imperata cylindrica dominated grassland. New For 14:127–143
Porras-Alfaro A, Herrera J, Sinsabaugh RL, Odenbach KJ, Lowrey T, Natvig DO (2008) Novel root fungal consortium associated with a dominant desert grass. Appl Environ Microbiol 74(9):2805–2813
Rodriguez RJ, White JF Jr, Arnold AE, Redman RS (2009) Fungal endophytes: diversity and functional roles. New Phytol 182(2):134–330
Saikkonen K (2007) Forest structure and fungal endophytes. Fungal Biol Rev 21:67–74
Sánchez Márquez S, Bills GF, Domínguez Acuña L, Zabalgogeazcoa I (2010) Endophytic mycobiota of leaves and roots of the grass Holcus lanatus. Fungal Divers 41:115–123
Schulz B, Boyle C (2005) The endophytic continuum. Mycol Res 109(6):661–686
Schulz B, Wanke U, Draeger S, Aust HJ (1993) Endophytes from herbaceous plants and shrubs: effectiveness of surface sterilization methods. Mycol Res 97:1447–1450
Shearer JF, Durham BD, Harms N (2011) Screening of biological control pathogens isolated from Eurasian watermilfoil. J Aquat Plant Manag 49:118–121
Shubin L, Juan H, RenChao Z, ShiRu X, YuanXiao J (2014) Fungal endophytes of Alpinia officinarum rhizomes: Insights on diversity and variation across growth years, growth sites, and the inner active chemical concentration. PLoS One 9(12):e115289. https://doi.org/10.1371/journal.pone.011528
Siregar UF, Rachmi A, Massijawa MY, Ishibashi N, Ando K (2007) Economic analysis of sengon (Paraserianthes falcataria) community forest plantation, a fast growing species in East Java, Indonesia. Forest Policy Econ 9:822–829
Soil survey staff (1992) Soil survey laboratory methods manual. Version No 2.0. USDA-NRCS. Soil Survey Investigations Report No. 42. U.S. Govt. Print. Office, Washington, DC
Sreelalitha SJ, Sridhar KR (2015) Endophytic fungi of wild legume Sesbania bispinosa in coastal sand dunes and mangroves of the Southwest coast of India. J For Res 26(4):1003–1011. https://doi.org/10.1007/s11676-015-0103-3
Strobel GA (2003) Endophytes as sources of bioactive products. Microbes Infect 5:535–544
Suryanarayanan TS, Murali TS, Thirunavukkarasu N, Govinda Rajulu MB, Venkatesan G, Sukumar R (2011) Endophytic fungal communities in woody perennials of three tropical forest types of the Western Ghats, Southern India. Biodivers Conserv 20:913–928
Ting ASY, Mah SW, Tee CS (2012) Evaluating the feasibility of induced host resistance by endophytic isolate Penicillium citrinum BTF08 as a control mechanism for Fusarium wilt in banana plantlets. Biol Control 61:155–159
Truog E (1930) The determination of the readily available phosphorus of soils. J Am Soc Agron 2:874–882
Usuki F, Narisawa K (2007) A mutualistic symbiosis between dark septate endophytic fungus, Heteroconium chaetospira, and a nonmycorrhizal plant, Chinese cabbage. Mycologia 99(2):175–184
Verma VC, Gond SK, Kumar A, Kharwar RN, Boulanger LA, Strobel GA (2012) Endophytic fungal flora from roots and fruits of an Indian Neem plant Azadirachta indica A. Juss. and impact of culture media on their isolation. Indian J Microbiol 51(4):469–476
Vinale F, Sivasithamparam K, Ghisalberti EL, Marra R, Woo SL, Lorito M (2008) Trichoderma–plant–pathogen interactions. Soil Biol Biochem 40:1–10
Wagatsuma T, Kawashima T, Tawaraya K (1988) Comparative stainability of plant-root cells with basic dye (methylene blue) in association with aluminum tolerance. Commun Soil Sci Plant Anal 19:1207–1215
Waipara NW, Di Menna ME, Cole ALJ, Skipp RA (1996) Potential pathogenicity of pasture plant root-colonising fungi to seedlings of legumes and grasses. Proc. 49th New Zealand Protection Conference, pp. 212–215
Wang JL, Li T, Liu GY, Smith JM, Zhao ZW (2016) Unraveling the role of dark septate endophyte (DSE) colonizing maize (Zea mays) under cadmium stress: physiological, cytological and genic aspects. Sci Rep 6:22028. https://doi.org/10.1038/srep22028
Wulandari D, Saridi, Cheng W, Tawaraya K (2016) Arbuscular mycorrhizal fungal inoculation improves Albizia saman and Paraserianthes falcataria growth in post-opencast coal mine field in East Kalimantan, Indonesia. For Ecol Manag 376:67–73
Yao YQ, Lan F, Qiao YM, Wei JG, Huang RS, Li LB (2017) Endophytic fungi harbored in the root of Sophora tonkinensis Gapnep: Diversity and biocontrol potential against phytopathogens. MicrobiologyOpen 6:e437. https://doi.org/10.1002/mbo3.437
Zhang Q, Gong M, Yuan J, Hou Y, Zhang H, Wang Y, Hou X (2017a) Dark septate endophyte improves drought tolerance in sorghum. Int J Agric Biol 19:53–60
Zhang Y, Lan TJ, Liao ST, Chen YL, Qin LP, Zhang WL, Nong Q, Xie L (2017b) Diversity of endophytic fungi in mangrove plants of Beibu Gulf, Guangxi. Microbiology China 44(4):783–794 Chinese
Zuccaro A, Schoch CL, Spatafora JW, Kohlmeyer J, Draeger S, Mitchell J (2008) Detection and identification of fungi associated with the brown seaweed Fucus serratus. Appl Environ Microbiol 74:931–941
Acknowledgements
The authors are grateful to Dr. Handojo Hadi Nurjanto and Dr. Widiyatno (Universitas Gadjah Mada) for help in gaining access to forest sites in Java Island. We are also grateful to Balai Penelitian Teknologi Konservasi Sumber Daya Alam (BPTKSDA) in Samboja for providing access to forest sites in Kalimantan Island and help in soil collection. This work was supported by JSPS KAKENHI Grant Number No. JP15H05246 from the Japan Society for the Promotion of Science (JSPS), INPEX, and Hashiya Scholarship Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Maulana, A.F., Turjaman, M., Sato, T. et al. Isolation of endophytic fungi from tropical forest in Indonesia. Symbiosis 76, 151–162 (2018). https://doi.org/10.1007/s13199-018-0542-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-018-0542-7