Abstract
Conversion of logged-over peatlands in Malaysia to oil palm plantations have led to changes in the microbial community in peat soil, mainly fungi. Therefore, this study was conducted to identify and compare the fungal species in four sites with different gradient of disturbances, namely a primary forest (Maludam NP), a logged-over forest (Cermat Ceria LOF), a 7.5-year-old oil palm plantation (Durafarm OPP), and an 11.5-year-old oil palm plantation (Naman OPP) by using a culture-dependent method. Fungi were isolated and identified based on morphological and molecular characterizations. The 18S rRNA gene was amplified, sequenced, and compared for the closest match in GenBank. Quantification of total fungal viable count, soil nutrients, and environmental aspects was also carried out. The total fungal viable count was not significantly different in all sites. Phylum Ascomycota was the dominant taxon isolated from all sites, which was about 90%. Basidiomycota and Zygomycota were also identified but found in low percentages. The most prevalent species for the different sites were Aspergillus fumigatus (20.6%) in Maludam NP, Penicillium chrysogenum in Cermat Ceria LOF (24.3%), Hypocrea muroiana in Durafarm OPP (27.4%), and Hypocrea atroviridis in Naman OPP (20%). Most of the species isolated were saprophytic fungi involved in decomposition of peat. In conclusion, fungal species composition in Sarawak peatlands with different gradient of disturbances was different in each site.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Tropical peatlands in Malaysia are located at low-elevation areas and characterized by a wet tropical climate with high annual rainfall and temperature (Page et al. 2006). Peatlands in Malaysia are unique, having a combination of rainforest and peatland ecosystems. In Malaysia, peatlands are located along the coastal areas of Peninsular Malaysia, Sabah, and Sarawak. Sarawak possesses about 70% (1.6 million hectares) of total peatlands in Malaysia (Melling 2016). These areas represent a pool of biodiversity for various flora and fauna, including microorganisms (Yule 2010; Adila et al. 2017; Shuhada et al. 2017).
Peat is formed from the accumulation of organic materials over the decomposition rate in soil due to the acidic environment and waterlogged conditions (Zinck 2011; Cobb et al. 2017). In tropical areas, peat is formed by the accumulation of partially decomposed organic matter from trees and dead fauna over thousands of years (Kanokratana et al. 2011). Peatlands in Sarawak have a high annual rainfall and are ombrogenous, acidic (pH 2.5–4.7), and dominated by dipterocarp trees (Melling 2016).
Despite their environmental conditions, peatlands provide a habitat for a wide range of animals and plants including soil microorganisms. Soil microorganisms such as fungi, bacteria, protists, and archaea have important ecological roles in regulating the ecosystem function via global carbon cycling (Winsborough and Basiliko 2010) and nutrient mineralization in peatlands (Andersen et al. 2013). Fungi are the primary decomposers (Thormann 2006; Myers et al. 2012) and dominant microbes found in peat; they are involved in biogeochemical processes such as the carbon cycle (Tveit et al. 2013) and emission of greenhouse gases (Toma et al. 2011). Fungi have the ability to sequester soil carbon and consequently provide ecosystem equilibrium (Treseder and Holden 2013). Therefore, the classification of fungi in peat is important in order to forecast carbon dynamics in the future.
Currently, the molecular approach is widely used to identify fungal species and resolve the taxonomic relationship among taxa (Hibbet et al. 2011; Asemaninejad et al. 2016; Sanders and Rodriguez 2016). Molecular identification provides more reliable results of identification compared to morphological characterization alone. Several markers of fungi such as protein-coding gene regions and nuclear ribosomal DNA (rDNA) are used in species identification. Ribosomal DNA that consists of a large subunit (28S), internal transcribed spacer (ITS), and small subunit (18S) are conserved within the fungi kingdom (Seifert 2009; Bellemain et al. 2010). The 18S rRNA gene is commonly used as a universal marker that can amplify across a wide range of fungi species with variations at the intraspecific level (Liu et al. 2015). The amplification of 18S rRNA gene has been used in many studies of diversity to assess fungal communities in a particular ecological niche such as forests and agricultural soils (Lim et al. 2010; Siles and Margesin 2016; Turrini et al. 2017).
Recently, peatlands in Sarawak are subjected to anthropogenic disturbances when logged-over degraded peatlands are converted to oil palm plantations. These disturbances may affect the diversity of soil microorganisms, mainly fungi, which later causes alteration of soil functions (Kerfahi et al. 2014; Mcguire et al. 2014; Nurulita et al. 2015). Therefore, the aim of this study was to identify and compare the fungal diversity among sites with different gradients of disturbances. The sites consisted of a primary forest, a logged-over forest, and two oil palm plantations.
2 Materials and methods
2.1 Sampling sites
Sampling sites were selected in four different forms of peat swamp ecosystems, which consisted of a primary forest (Maludam NP), a logged-over forest (Cermat Ceria LOF), and two oil palm plantations (Durafarm OPP and Naman OPP) (Fig. 1). Maludam NP (1°37′ 43.15″N, 111°02′ 22.12″E) represents the least disturbed ecosystem of peat swamp forest. Cermat Ceria LOF (1°23′ 58.85″N, 111°24′ 08.61″E) is a logged-over forest which has been recovered naturally after timber was harvested years ago. Durafarm OPP (1°23′ 50.63″N, 111°24′ 50.59″E) with 7.5 years old oil palm while Naman OPP (2°09′ 47.03″N, 111°55′ 22.32″E) with oil palm aged 11.5 years (Fig. 2). Durafarm OPP and Naman OPP have represented the highly disturbed areas. Maludam NP, Cermat Ceria LOF, and Durafarm OPP were located in Sri Aman, Sarawak while Naman OPP in Sibu, Sarawak. The distance between each site was ranged between 4 km up to 102 km.

Source of maps: Google maps Imagery©2017 https://www.google.com.my/maps
Maps showing the location of study sites. a Borneo Island where the sites are located in Sarawak, Malaysia. b Naman OPP is located at Sibu and c Maludam NP, Cermat Ceria LOF and Durafarm are located at Sri Aman, Sarawak. Cermat Ceria LOF is adjacent to the Durafarm OPP.
Sites with different gradient of disturbances. a Primary peat swamp forest, Maludam NP. b Cermat Ceria LOF is a logged-over forest which has been recovered naturally after logging. c Durafarm oil palm plantation with 7.5-year-old oil palm and d Naman oil palm plantation with oil palm aged 11.5 years
Sampling was carried out in October 2012 during the wet season (monthly precipitation > 200 mm). Peat samples were collected at ten sampling points for each site with depth between 0 and 60 cm, using peat auger (Eijkelkamp, The Netherlands). The sampling points were at least 10 m apart from each other. 50 ml tubes were used to keep the peat samples and transported in boxes containing ice packs to the laboratory for analysis. Based on soil survey conducted by Param Agricultural Soil Surveys (M) Sdn. Bhd. (Paramananthan 2012a), peat in Maludam NP is classified as decomposed sapric organic materials (sapric; fibre content less than 33%). Meanwhile, peat in Cermat Ceria LOF, Durafarm OPP and Naman OPP consisted of sapric with undecomposed wood (Paramananthan 2012b).
2.2 Fungal isolation
Fungi from peat samples were isolated on potato dextrose agar (PDA) and malt extract agar (MEA) using spread plate method. A volume of 100 µl of dilutions, 10−4 and 10−5 were spread onto PDA and MEA using hockey stick and incubated at 25 ± 2 °C for 3 days. Each different colony grown was subcultured onto new PDA to obtain pure isolates. The pure isolates were stored as the conidial disc made using cork borer in mineral oil for future use.
2.3 Enumeration of total fungal viable count
The soil dilutions of 10−4 and 10−5 were plated on PDA and MEA and incubated at 25 ± 2 °C. The viable colonies on the plate were counted and recorded after 3 days of incubation. The total fungal viable count was calculated as colonies forming unit per 1 g of dry soil (cfu/g).
2.4 Morphological characterization
The macroscopic characteristic such as colony appearance was observed on PDA after 5 days’ incubation at 25 ± 2 °C. For microscopic characteristics, the conidia, conidiophores and phialides shape were examined using slide culture method. Hypocrea sp. were identified based on taxonomy key by Gams and Bissett (1998). The characteristics of Aspergillus and Penicillium species were observed and identified according to Diba et al. (2007) and Pitt (1979), respectively. Besides, morphological characteristics of other species were identified according to Watanabe (2002).
2.5 Amplification of 18S rDNA
The species identification of all isolates was done using DNA sequencing of 18S rDNA region. The pure isolates were grown on PDA for 3 days (fast-growing fungi) and 7 days (slow-growing fungi) before proceeding with the DNA extraction. The mycelium was harvested and extracted for genomic DNA using Fungi/Yeast Genomic DNA Isolation Kit (Norgen Biotek Corp., Canada) according to manufacturer’s instruction. The 18S rDNA was amplified using a pair of primer: EF4-F (5′-GGAAGGG[G/A]TGTATTTATTAG-3′) and fung5-R (5′-GTAAAAGTCCTGGTTCCCC-3′) (Smit et al. 1999). The PCR amplification was conducted in 25 µl reaction master mix that consists of 1 × PCR buffer, 1.0 µM of each primer, 0.2 mM of each dNTPs, 0.3 U of GoTaq Flexi DNA polymerase (Promega, Madison, WI, USA), 2.5 mM MgCl2, 0.3% of bovine serum albumin (BSA) and 1.0 µl DNA template. The amplification was performed in a thermal cycler (Applied Biosystems Veriti™ Thermal Cycler, USA) with an initial denaturation at 94 °C for 3 min followed by 30 cycles of denaturation at 94 °C for 1 min, annealing at 48 °C for 1 min, extension at 72 °C for 1 min and 5 min final extension at 72 °C.
2.6 DNA sequencing
The PCR products were loaded on 1% agarose gel added with gel stain (FloroSafe DNA Stain, First BASE Laboratories, Malaysia). The gel electrophoresis was run at 80 V for 30 min and viewed under UV light. The amplicon sized 550 bp was excised from agarose gel and purified using EasyPure® Quick Gel Extraction Kit (TransGen Biotech, China). The purified PCR products were sent to First BASE Laboratories (Malaysia) for sequencing. DNA sequences obtained were analysed and compared using Basic Local Alignment Search Tool (BLAST) to determine the closest match sequences from a database (http://www.ncbi.nlm.nih.gov).
2.7 Phylogenetic tree
The sequences were aligned using CLUSTAL W in Molecular Evolutionary Genetics Analysis version 6.0 (MEGA 6.0). The aligned sequences then were used to construct a phylogenetic tree. The tree was generated based on neighbour-joining (NJ) method using Jukes–Cantor with 1000 bootstrap replications.
2.8 Soil physiochemical analysis
Soil pH was determined by adding 10 g of peat samples to 50 ml of distilled water and stirring for 30 min before measuring using a pH meter (Mettler Toledo, Switzerland) (Gheda and Ahmed 2015). The moisture content of soil was determined by drying 5 g of peat samples in the oven at 105 °C for 24 h (Kelly and Sivakumar 2014). After 24 h, the peat samples were weighed again and the moisture content was calculated following Latiffah et al. (2011). Rainfall data were recorded for the sampling month (October 2012). On field measurement such as peat depth was also recorded for all sites. For carbon (C) and nitrogen (N) determination, peat samples were dried at 25 °C for 2 weeks until constant weight achieved. The peat samples were combusted at 1040 °C using a model of Primacs SNC analyser (Skalar, UK). Total C was determined by using infrared detection and total N using a thermal conductivity detector. For micronutrients, peat samples were dried at 60 °C for 2–3 days until constant weight was reached. About 0.3 g of dry samples was extracted with 95% nitric acid, digested in Titan MPS Microwave Sample Preparation System (PerkinElmer SCIEX, USA) and diluted with MiliQ water before determination using Inductively Coupled Plasma Mass Spectrometer (ICP-MS) ELAN DRC (PerkinElmer SCIEX, USA). Analysis was conducted according to manufacturer’s protocol.
2.9 Statistical analysis
The mean value of soil physiochemical properties for each site was calculated. The data of soil physiochemical properties and total fungal viable count were analysed by one-way ANOVA using SPSS version 22.0 (SPSS Inc., Chicago, IL, USA) to determine variation between each site.
3 Results
3.1 Total fungal viable count
Based on total fungal viable count (Fig. 3), Cermat Ceria LOF recorded the highest culturable fungal population on MEA with 5.0 × 105 cfu/g compared to other sites. Meanwhile, fungal population was the highest on PDA for Naman OPP with 7.4 × 105 cfu/g. Statistical analysis revealed no significant difference in total fungal viable counts on PDA and MEA when compared between sites.
3.2 Species identification
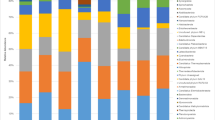
A total of 294 isolates were recovered from peat samples in all sites. A total of 72, 74, 73, and 75 isolates were obtained from Maludam NP, Cermat Ceria LOF, Durafarm OPP, and Naman OPP, respectively (Table 1). Based on a BLAST search, all isolates showed similarity between 73 and 100% with sequences in GenBank (Online Resource 1). Three phyla which comprised of Ascomycota, Basidiomycota, and Zygomycota were identified. Peat soils in Sarawak are dominated by filamentous ascomycetes. Basidiomycota and Zygomycota were found in Cermat Ceria LOF and Durafarm OPP.
The total number of fungal species varied among sites. Naman OPP had the highest species richness with 18 species. Durafarm OPP accounted for 17 species, followed by Maludam NP (16 species) and Cermat Ceria LOF (15 species).
Among the 36 species identified, Aspergillus fumigatus (40 isolates) was commonly found across the sites. The second most common species was P. chrysogenum (37 isolates) followed by Aspergillus sp. (30 isolates) and A. niger (21 isolates). Hypocrea sp. identified as H. atroviridis (34 isolates) and H. muroiana (31 isolates) were the most common species in plantation areas, Durafarm OPP and Naman OPP. Other than that, Penicillium roqueforti (19 isolates) was found only in oil palm plantations.
Three species of higher fungi (Antrodia variiformis, Byssomerulius albostramineus, and Ganoderma australe) were isolated from Cermat Ceria LOF, and an isolate of Pulcherricium caeruleum was found in Durafarm OPP. A species of Zygomycota, Mortierella chlamydospora was obtained from all sites except Naman OPP, while Umbelopsis isabellina was only recorded from Maludam NP.
In Maludam NP, about 20.6% of total fungal isolates was dominated by A. fumigatus followed by Penicillium corylophilum (17.8%). Penicillium corylophilum can also be found in Naman OPP at a low percentage. A teleomorphic species of Penicillium known as Talaromyces purpureogenus was isolated and found in Maludam NP. The other species of Penicillium identified were P. chrysogenum, P. citrinum, and P. namyslowskii.
The most prevalent fungus in Cermat Ceria LOF was P. chrysogenum (24.3%) followed by Aspergillus species including Aspergillus sp. (21.6%), A. fumigatus (20.3%), and A. niger (16.2%). Trichoderma sp. and T. harzianum, the anamorphic stage of Hypocrea, were also identified in Cermat Ceria LOF. Other fungi species found were Cladosporium cladosporioides, P. javanicum, Sarcladium sp., T. leycettanus, and M. chlamydospora.
The fungal species in the two oil palm plantations were relatively the same. Hypocrea muroiana and H. atroviridis were commonly isolated from Durafarm OPP (27.4 and 26%) and Naman OPP (15 and 20%). About 17.8% of the isolates in Durafarm OPP consisted of P. roqueforti, while only 8% of P. roqueforti was found in Naman OPP. Besides that, P. chrysogenum was isolated from both sites with 9.3% in Naman OPP and 2.7% in Durafarm OPP. The uncommon species found in Durafarm OPP were Daldinia sp., Hypoxylon haematostroma, Phomopsis sp., Pulcherricium caeruleum, and M. chlamydospora. On the other hand, the least isolated species in Naman OPP were Eurotiomycetes sp., Fusarium oxysporum, Neurospora crassa, P. namyslowskii, P. purpurogenum, and Purpureocillium lilacinum.
Based on observations of isolates on PDA, the colony morphology varied. Aspergillus fumigatus and A. niger produced white-to-yellowish mycelia which turned into dark green and black with the formation of spores (Fig. 4a, b). Meanwhile, Aspergillus sp. produced black colonies and abundant white-to-yellowish mycelia (Fig. 4c). Hypocrea muroiana and H. atroviridis formed whitish-to-green circular colonies on PDA (Fig. 4d, e). Penicillium corylophilum produced yellow and light green colonies towards the edge of the colonies (Fig. 4f). The colonies of Penicillium sp. were dark green in colour (Fig. 4g). On the other hand, P. javanicum showed flat colonies of grey to whitish on PDA (Fig. 4h). The morphology of P. chrysogenum was different from other Penicillium species. It produced flat mycelia with yellow-and-white colony on PDA (Fig. 4i). Umbelopsis isabellina produced white sparse aerial mycelia (Fig. 4j), while M. chlamydospora formed fluffy white mycelia after five days of incubation (Fig. 4k).
Morphological characteristics of fungi isolated from different sites in Sarawak based on macroscopic observation on PDA after 5 days. a A. fumigatus, b A. niger, c Aspergillus sp., d H. muroiana, e H. atroviridis, f P. corylophilum, g Penicillium sp., h P. javanicum, i P. chrysogenum, j U. isabellina, and k M. chlamydospora
For microscopic characterization, conidiophore of A. niger was long with dark spore masses at the tip (Fig. 5A). The conidia of A. niger were globose and found in clusters (Fig. 5a). Hypocrea muroiana produced branch conidiophores and phialides (Fig. 5B) while the conidia were round (Fig. 5b). The conidiophore of H. atroviridis grew from the main hyphae and stout phialides were formed from the branch (Fig. 5C). The conidia shape was ovoid to round (Fig. 5c). Penicillium chrysogenum produced biverticillate branching and bore conidia in chains (Fig. 5D). The conidia were found in clusters and chains with an ovoid shape (Fig. 5d). Penicillium corylophilum formed long monoverticillate conidiophores with chains of conidia (Fig. 5E). The conidia of P. corylophilum were phialosporous and globose (Fig. 5e). P. roqueforti formed long conidiophore that bore oval-to-round conidia in chains (Fig. 5F, f). Umbelopsis isabellina produced round and globose sporangiospores (Fig. 5g). The special thick wall of resting structure known as chlamydospore was observed in the M. chlamydospora culture. The chlamydospores were globose and granulate, occurred singly, and were found in the intercalary and terminal sections of the hyphae (Fig. 5h).
3.3 Phylogenetic tree analysis
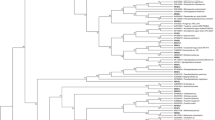
The tree was divided into four clades (Fig. 6). Clade A consisted of Ascomycota with a bootstrap value of 65%. Clades B and C consisted of isolates of Basidiomycota, while clade D consisted of Zygomycota. Clade A was divided into several subclades of Eurotioles (Aspergillus, Penicillium, and Talaromyces), Hypocreales (Fusarium, Hypocrea and its anamorphic stage, Trichoderma, and Purpureocillium), Xylariales (Hypoxylon haematostroma), Capnodiales (Cladosporium cladosporioides), and Diaporthales (Phomopsis sp.). Clade B consisted of an isolate of Byssomerulius albostramineus, a basidiomycete. Several species of basidiomycetes obtained from tropical peat soil were placed on clade C. On the other hand, M. chlamydospora, a zygomycete, was located on clade D with a high bootstrap value of 100%. In contrast, two isolates of U. isabellina (Zygomycota) were placed distinctly from the other clades.
A neighbour-joining tree showing the relationship between all isolates based on 18S rRNA gene. The tree was constructed using Jukes–Cantor method with 1000 replications of bootstrap value. The analysis involved 294 nucleotide sequences and the length of nucleotide for aligned sequences was 891 nucleotides. The tree was applied cut-off 50% with bootstrap values shown next to the branch. Symbol of ‘filled circle’ represented condensed clades which are consisted of the isolates in the same order
3.4 Soil physiochemical analysis
Total C content was the highest at the oil palm plantations, Naman OPP (55.77 ± 0.27%) and Durafarm OPP (53.42 ± 0.44%) compared to Maludam NP and Cermat Ceria LOF (Table 2). Meanwhile, total N content in peat soil was the highest at Durafarm OPP (1.88 ± 0.44%) and lowest at Cermat Ceria LOF (1.28 ± 0.48%). Concerning the C:N ratio, Cermat Ceria LOF had a higher value (38.88 ± 18.11) than other sites.
A comparison of the soil pH showed no significant difference among sites. The soil pH ranged between 3.37 and 3.41. The moisture content in all sites was significantly different between each other. The moisture content in Durafarm OPP and Naman OPP was the highest with 65.06 ± 1.14% and 65.09 ± 1.41%, respectively. On the contrary, Maludam NP had the lowest moisture content with 49.5 ± 1.4%. For peat depth, there was no significant difference and it ranged from 104.20 to 113.70 cm.
The concentrations of B, Mg, P, K, Ca, Mn, Fe, and Ni were significantly higher in Durafarm OPP compared to the other sites (Table 3). On the other hand, peat soil of Naman OPP showed a high content of trace elements such as Cu and Zn. In terms of heavy metal contents, Mo, Cd, and Pb were significantly greater in Cermat Ceria LOF compared to the other sites. In Maludam NP, Mg, P, K, Ca, Ni, Cu, Zn, Mo, and Cd showed the lowest value.
4 Discussion
Amongst sites, Naman OPP had the highest population of fungi on PDA, which was reflected by total fungal viable count. In this study, the oil palm plantations supported a large number of fungi compared to the forest. Fertilizer application in the plantations provides a favourable condition for fungal growth and enhances their population (Kerekes et al. 2013; Krashevska et al. 2013). Nurulita et al. (2015) also showed that fungal population was higher in oil palm and rubber plantation than in forest. Cermat Ceria LOF showed slightly higher fungal population on MEA as compared to the other sites. Nevertheless, both media used in this study were rich with carbon and nitrogen, allowing the growth of sporulating species from all sites.
In this study, Ascomycota was recorded as the common taxon isolated from peatlands in Sarawak. This phylum was reported as the dominant phylum in bog and fen peat (Lin et al. 2012), as well as in permafrost peatlands (Tveit et al. 2013). They are usually fast-growing fungi with a high spore production rate, which enables them to thrive in the peat soil.
Several species of Aspergillus are common in Sarawak peat soil and were found in all sites. Aspergillus species are ubiquitous soil fungi that globally play a role in carbon and nitrogen cycle (Dagenais and Keller 2009). Aspergillus fumigatus was the most abundant species in the primary forest as compared to the other sites. Conversely, A. niger, a soft rot fungus, was prevalent in Cermat Ceria LOF. A study by Hamed (2013) showed that A. niger degrades cellulose, rather than lignin, in wood decomposition. Thus, A. niger would have the same role to degrade simple structures during slow litter decomposition in logged-over forest. Aspergillus fumigatus and A. niger are tolerant to acid and temperature and are involved in phosphate solubilization process in mineral soil (Rinu and Pandey 2010).
The genus of Penicillium is a cosmopolitan fungus found in a wide range of ecosystems including peatlands. Penicillium chrysogenum is a soft rot fungus, which decomposes hardwood rather than softwood trees (Hamed 2013). This is supported by the result in this study where P. chrysogenum was highly prevalent in both primary and logged-over forests. This species is able to live in saline and acidic soil (Hujslová et al. 2010). Penicillium corylophilum has been reported to inhabit mangroves, where oxygen is depleted (Arfi et al. 2012). Thus, it can survive in peat soil. Penicillium roqueforti was present in peat soil of both oil palm plantations. However, P. roqueforti has only been widely used in cheese production before; other information on its association in peat soil remains unknown. Talaromyces purpurogenus is able to solubilize insoluble phosphorus in soil for plant utilization (Vieira and Nahas 2005). This species was isolated from tropical forests in India (Devi et al. 2012) and in acidic soils located in the Czech Republic (Hujslová et al. 2010).
The Hypocrea genus is ubiquitous, widely distributed, and associated with a wide range of substrates (Chaverri et al. 2015). Hypocrea spp. are also the predominant species of soil mycobiota (Jaklitsch and Voglmayr 2015). In the agricultural industry, Hypocrea species are economically important as biocontrol agents against diseases and pests, as well as agents to promote nutrient uptake in plants (Druzhinina et al. 2011; Harman et al. 2012). In the present study, H. atroviridis (= Trichoderma atroviride) and H. muroiana (= T. koningii) were abundantly distributed in oil palm plantations. Hypocrea atroviridis was commonly found in peat soil of oil palm plantations (Ramlah Ali et al. 2016). Besides that, H. atroviridis produces laccase enzymes that are used in lignin biodegradation (Chakroun et al. 2010). In the denitrification process, H. muroiana is involved in nitrous oxide (N2O) production that occurs in soil (Mothapo et al. 2015). Both species work as an extensive network to increase the nutrient uptake of oil palm. The anamorphic stage of Hypocrea, namely Trichoderma sp. and T. harzianum, were found in Cermat Ceria LOF and Naman OPP. Trichoderma harzianum is widely reported as having antagonistic properties against plant pathogens, such as Ganoderma boninense in oil palm plantations (Naher et al. 2012). Presence of T. harzianum in Naman OPP has a positive impact on oil palm health, reducing the basal stem rot incidence in this site.
In a matured oil palm plantation like Naman OPP, a few isolates of Fusarium oxysporum were isolated. This species causes vascular wilt disease (Rusli et al. 2015) and crown disease (Hafizi et al. 2013) on oil palm. Fusarium oxysporum was previously isolated from peat bog (Golovchenko et al. 2013) and tropical peat (Nurul Farah et al. 2016) as it is involved in the N2O-producing activity in soil (Mothapo et al. 2015).
Basidiomycetes were found in Cermat Ceria LOF and Durafarm OPP in low frequency. Basidiomycetes may take the role as polymer decomposers in logged-over forests and plantations for decomposition process. These fungi, widely known as complex polymer degraders, degrade lignin and plant biomass polysaccharide (Lundell et al. 2010). Antrodia variiformis, found in Cermat Ceria LOF, is a polypore; the same species was isolated from an ombrophilous mixed forest in Brazil (Westphalen and Silveira 2012).
Other species of basidiomycetes, such as Byssomerulius albostramineus and Ganoderma australe isolated in this study, are commonly reported to be associated with plants as wood-decay fungi (Stokland and Larsson 2011; Schwarze et al. 2012). The population of Basidiomycetes was underestimated since this study focused on soil samples. The culture-dependent method generally isolates sporulating fungi in the Ascomycota and Zygomycota divisions as compared to Basidiomycota. Besides that, the fruiting body production of basidiomycetes is erratic and influenced by moist weather (Rydin and Jeglum 2013).
Two species of Zygomycota were found in Sarawak peat. Members of Mortierella are common saprobic fungi associated with soil (Wagner et al. 2013). Based on micromorphological analysis of M. chlamydospora, it was observed to have a special thick wall structure and chlamydospores that can survive and remain dormant in peat soil. The percentage of occurrence of Mortierella species in peat is low, as reported by Dobrovol’skaya et al. (2012). Meanwhile, another species of zygomycetes, U. isabellina, was isolated from the root of conifers in peatlands and act as an endophytic fungus (Terhonen et al. 2014).
The molecular data resolved three divisions of fungi, indicating a variation of intraspecific level as shown in the neighbour-joining tree. In this study, isolates of T. purpurogenum formed a monophyletic group with P. purpurogenum (synonym of T. purpurogenum) and were placed on different clade from Penicillium genus. This is because the Talaromyces genus was introduced to accommodate the Penicillium subgenus Biverticillium, which is phylogenetically distinct from Penicillium (Yilmaz et al. 2014). Thus, isolates of T. purpurogenum were not grouped together with other Penicillium species. Besides, U. isabellina was placed apart from Mortierella group as they formed a new family, Umbelopsidaceae (Meyer and Gams 2003).
Molecular identification in this study also supported the morphological characterization based on macroscopic and microscopic characteristics. Most of the isolated fungi, such as Penicillium sp., Aspergillus sp., and Hypocrea sp., were prolific spore producers on media (Thormann 2006). The characteristics of the conidia with minute size enabled them to colonize and distribute well in peat soil.
Fungal abundance and richness increased with high abundance of plant diversity and richness, especially in forest soil (Costa et al. 2012; Dassen et al. 2017). However, the present study showed slightly higher species richness and abundance of fungi in oil palm plantations as compared to forest sites. This result did not reflect the previous report by Zak et al. (2003), where the abundance of plant diversity enhanced the fungal community in the soil. The difference in results was probably because of the characteristics of peat itself and the abundance of organic materials in oil palm plantations to provide a good niche to fungi. Moreover, there were limitations and biases of the culture-dependent method, since this method overestimated sporulating species as compared to slow-growing fungi and fungi which cannot grow on media. The nutrient contents in soil might contribute to high species richness in oil palm plantation ecosystems, which are scarce in the forests.
Abiotic factors such as pH, organic contents, and moisture influence the distribution and diversity of fungal species (Devi et al. 2012). Generally, fungi favour acidic conditions for growth and reproduction (Nurul Farah et al. 2016). Most of the species isolated in this study were previously reported to be tolerant to low pH and high temperature, such as Penicillium, Aspergillus, and Trichoderma (Rinu and Pandey 2010). Nutrients in peat soil influenced the fungal species composition. A direct relationship showed that oil palm plantations have higher nutrient contents in soil and a higher number of fungal species as compared to the forests.
The rate of decomposition is reflected by the C:N ratio in the soil (Hazelton and Murphy 2007). A high total C and low N indicate a slow decomposition rate in soil (Melling 2016). Maludam NP showed the highest decomposition rate, followed by Durafarm OPP, Naman OPP, and Cermat Ceria LOF. According to Banerjee et al. (2016), the decomposition rate in an ecosystem is determined by the abundance of the keystone taxa, which is associated with organic matter degradation. Even though Maludam NP exhibited a low number of fungal species (16 species, 72 isolates), the abundance of the same species, such as A. fumigatus and P. chrysogenum, was large enough to carry out a high decomposition rate as compared to oil palm plantations, Durafarm OPP and Naman OPP. Besides, the peat in Maludam NP (decomposed sapric materials) was associated with high decomposition rate in peat soil. On the other hand, Cermat Ceria LOF showed the slowest decomposition rate, which may be due to chemical recalcitrant in plant debris (Edmonds 1991). Other factors, such as climate may also affect the rate of soil decomposition (Lutzow et al. 2006).
In this study, peat soil in oil palm plantations has a high moisture content as compared to primary and logged-over forests. This result contradicts with a study by Nurulita et al. (2015), where forests showed higher moisture content than oil palm plantations. This was because the samples were taken from disturbed edges of Maludam NP (Online Resource 2). Microbes respond to moisture content changes to survive and carry out their functional activities in the soil (Manzoni et al. 2012).
5 Conclusions
In the peatlands of Sarawak, Malaysia, different species of fungi were identified from the phyla Ascomycota, Basidiomycota, and Zygomycota. Most of the species isolated were saprophytic fungi that are able to survive in peat soil. These fungi had prominent roles in maintaining peat ecosystem function. Fungi species’ composition in peat soil of Sarawak differed among sites and this might be driven by the nutrient contents and abundance of organic material in the sites. Species of Aspergillus and Penicillium were predominant in peat forests while Hypocrea species were commonly found in oil palm plantations. However, the culture-dependent method still showed no clear difference for fungal species richness and abundance in Sarawak peatlands. Therefore, a different approach such as metagenomics is needed to study the fungal diversity, richness, and composition in peat soil comprehensively.
References
Adila N, Sasidhran S, Kamarudin N, Puan CL, Azhar B, Lindenmayer DB (2017) Effects of peat swamp logging and agricultural expansion on species richness of native mammals in Peninsular Malaysia. Basic Appl Ecol 22:1–10
Andersen RRE, Chapman SJ, Artz RRE (2013) Microbial communities in natural and disturbed peatlands: a review. Soil Biol Biochem 57:979–994
Arfi Y, Marchand C, Wartel M, Record E (2012) Fungal diversity in anoxic-sulfidic sediments in a mangrove soil. Fungal Ecol 5:282–285
Asemaninejad A, Weerasuriya N, Gloor GB, Lindo Z, Thorn RG (2016) New primers for discovering fungal diversity using nuclear large ribosomal DNA. PLoS ONE 11:e0159043
Banerjee S, Kirkby CA, Schmutter D, Bissett A, Kirkegaard JA, Richardson AE (2016) Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol Biochem 97:188–198
Bellemain E, Carlsen T, Brochmann C, Coissac E, Taberlet P, Kauserud H (2010) ITS as an environmental DNA barcode for fungi: an in silico approach reveals potential PCR biases. BMC Microbiol 10:189
Chakroun H, Mechichi T, Martinez MJ, Dhouib A, Sayadi S (2010) Purification and characterization of a novel laccase from the ascomycete Trichoderma atroviride: application on bioremediation of phenolic compounds. Process Biochem 45:507–513
Chaverri P, Branco-Rocha F, Jaklitsch WM, Gazis RO, Degenkolb T, Samuels GJ (2015) Systematics of the Trichoderma harzianum species complex and the reidentification of commercial biocontrol strains. Mycologia 107:558–590
Cobb AR, Hoyt AM, Gandois L, Eri J, Dommain R, Abu Salim K, Kai FM, Haji Su’ut NS, Harvey CF (2017) How temporal patterns in rainfall determine the geomorphology and carbon fluxes of tropical peatlands. Proc Natl Acad Sci USA 114:E5187–E5196
Costa PMO, Souza-Motta CM, Malosso E (2012) Diversity of filamentous fungi in different systems of land use. Agrofor Syst 85:195–203
Dagenais TRT, Keller NP (2009) Pathogenesis of Aspergillus fumigatus in invasive Aspergillosis. Clin Microbiol Rev 22:447–465
Dassen S, Cortois R, Martens H, Hollander Md, Kowalchuk GA, Putten WHVD, Deyn GBD (2017) Differential responses of soil bacteria, fungi, archaea and protists to plant species richness and plant functional group identity. Mol Ecol. https://doi.org/10.1111/mec.14175
Devi LS, Khaund P, Nongkblaw FMW, Josbi SR (2012) Diversity of culturable soil micro-fungi along altitudinal gradients of Eastern Himalayas. Mycobiology 40:151–158
Diba K, Kordbacheh P, Rezaie S, Mahmoudi M (2007) Identification of Aspergillus species using morphological characteristics. Pakistan J Med Sci 23:867–872
Dobrovol’skaya TG, Golovchenko AV, Kukharenko OS, Yakushev AV, Semenova TA, Inisheva LA (2012) The structure of the microbial communities in low moor and high moor peat bogs of Tomsk Oblast. Eurasian Soil Sci 45:273–281
Druzhinina IS, Seidl-seiboth V, Herrera-estrella A, Zeilinger S, Grigoriev IV, Kubicek CP (2011) Trichoderma: the genomics of opportunistic success. Nat Rev Microbiol 9:749–759
Edmonds RL (1991) Organic matter decomposition in Western United States forests. In: Proceedings management and productivity of western-montane forest soils, pp 118–128
Gams W, Bissett J (1998) Morphology and identification of Trichoderma. In: Kubicek CP, Harman GE (eds) Trichoderma and gliocladium, vol 1. basic biology, taxonomy and genetics. Taylor and Francis Ltd, London, pp 3–34
Gheda SF, Ahmed DA (2015) Improved soil characteristics and wheat germination as influenced by inoculation of Nostoc kihlmani and Anabaena cylindrica. Rend Fis Acc Lincei 26:121–131
Golovchenko AV, Kurakov AV, Semenova TA, Zvyagintsev DG (2013) Abundance, diversity, viability, and factorial ecology of fungi in peatbogs. Eurasian Soil Sci 46:74–90
Hafizi R, Salleh B, Latiffah Z (2013) Morphological and molecular characterization of Fusarium solani and F. oxysporum associated with crown disease of oil palm. Braz J Microbiol 44:959–968
Hamed SAM (2013) In-vitro studies on wood degradation in soil by soft-rot fungi: Aspergillus niger and Penicillium chrysogenum. Int Biodeterior Biodegradation 78:98–102
Harman GE, Herrera-estrella AH, Horwitz BA, Lorito M (2012) Special issue: Trichoderma—from basic biology to biotechnology. Microbiology 158:1–2
Hazelton P, Murphy B (2007) Interpreting soil test results: what do all the number mean?. CSIRO Publishing, Collingwood
Hibbet DS, Ohman A, Glotzer D, Nuhn M, Krik P, Nilsson RH (2011) Progress in molecular and morphological taxon discovery in fungi and options for formal classification of environmental sequences. Fungal Biol 25:38–47
Hujslová M, Kubátová A, Chudicková M, Kolarik M (2010) Diversity of fungal communities in saline and acidic soils in the Soos National Natural Reserve, Czech Republic. Mycol Prog 9:1–15
Jaklitsch WM, Voglmayr H (2015) Biodiversity of Trichoderma (Hypocreaceae) in Southern Europe and Macaronesia. Stud Mycol 80:1–87
Kanokratana P, Uengwetwanit T, Rattanachomsri U, Bunterngsook B, Nimchua T, Tangphatsornruang S, Plengvidhya V, Champreda V, Eurwilaichitr L (2011) Insights into the phylogeny and metabolic potential of a primary tropical peat swamp forest microbial community by metagenomic analysis. Environ Microbiol 61:518–528
Kelly BCO, Sivakumar V (2014) Water content determinations for peat and other organic soils using the oven-drying method. Dry Technol 32:631–643
Kerekes J, Kaspari M, Stevenson B, Nilsson RH, Hartmann M, Amend A, Bruns TD (2013) Nutrient enrichment increased species richness of leaf litter fungal assemblages in a tropical forest. Mol Ecol 22:2827–2838
Kerfahi D, Tripathi BM, Lee J, Edwards DP, Adams JM (2014) The impact of selective-logging and forest clearance for oil palm on fungal communities in Borneo. PLoS ONE 9:e111525
Krashevska V, Sandmann D, Maraun M, Scheu S (2013) Moderate changes in nutrient input alter tropical microbial and protist communities and belowground linkages. ISME J 8:1126–1134
Latiffah Z, Yee TL, Maziah Z, Salleh B (2011) Diversity of microfungi in sandy beach soil of Teluk Aling, Pulau Pinang. Trop Life Sci Res 22:71–80
Lim YW, Kim BKB, Kim C, Jung HS, Kim BKB, Lee J, Chun J (2010) Assessment of soil fungal communities using pyrosequencing. J Microbiol 48:284–289
Lin X, Green S, Tfaily MM, Prakash O, Konstantinidis KT, Corbett JE, Chanton JP, Cooper WT, Kostka JE (2012) Microbial community structure and activity linked to contrasting biogeochemical gradients in bog and fen environments of the glacial. Appl Environ Microbiol 78:7023–7031
Liu J, Yu Y, Cai Z, Bartlam M, Wang Y (2015) Comparison of ITS and 18S rDNA for estimating fungal diversity using PCR–DGGE. World J Microbiol Biotechnol 31:1387–1395
Lundell TK, Mäkelä MR, Hildén K (2010) Lignin-modifying enzymes in filamentous basidiomycetes—ecological, functional and phylogenetic review. J Basic Microbiol 50:5–20
Lutzow MV, Kogel-Knabner I, Ekschmitt K, Matzner E, Guggenberger G, Marschner B, Flessa H (2006) Stabilization of organic matter in temperate soils: mechanisms and their relevance under different soil conditions-a review. Eur J Soil Sci 57:426–445
Manzoni S, Schimel JP, Porporato A (2012) Responses of soil microbial communities to water stress: results from a meta-analysis. Ecology 93:930–938
Mcguire KL, D’Angelo H, Brearley FQ et al (2014) Responses of soil fungi to logging and oil palm agriculture in Southeast Asian tropical forests. Microb Ecol 69:737–747
Melling L (2016) Peatland in Malaysia. In: Osaki M, Tsuji N (eds) Tropical peatland ecosystems. Springer Japan, Tokyo, pp 59–73
Meyer W, Gams W (2003) Delimitation of Umbelopsis (Mucorales, Umbelopsidaceae fam. nov.) based on ITS sequence and RFLP data. Mycol Res 107:339–350
Mothapo N, Chen H, Cubeta MA, Grossman JM, Fuller F, Shi W (2015) Phylogenetic, taxonomic and functional diversity of fungal denitrifiers and associated N2O production efficacy. Soil Biol Biochem 83:160–175
Myers B, Webster KL, Mclaughlin JW, Basiliko N (2012) Microbial activity across a boreal peatland nutrient gradient: the role of fungi and bacteria. Wetl Ecol Manag 20:77–88
Naher L, Tan SG, Yusuf UK, Ho CL, Abdullah F (2012) Biocontrol agent Trichoderma harzianum strain FA11326 as an enhancer of oil palm growth. Pertanika J Trop Agric Sci 35:173–182
Nurul Farah AK, Masratulhawa M, Nik Mohd Izham MN, Latiffah Z (2016) Saprophytic and potentially pathogenic Fusarium species from peat soil in Perak and Pahang. Trop Life Sci Res 27:1–20
Nurulita Y, Adetutu EM, Kadali KK, Zul D, Mansur AA, Ball AS (2015) The assessment of the impact of oil palm and rubber plantations on the biotic and abiotic properties of tropical peat swamp soil in Indonesia. Int J Agric Sustain 13:150–166
Page SE, Rieley JO, Wust R (2006) Lowland tropical peatlands of Southeast Asia. In: Martini IP, Martinez Cortizas A, Chesworth W (eds) Developments in earth surface processes. Elsevier B.V., Amsterdam, pp 145–172
Paramananthan S (2012a) Report of Soils of the MPOB’s biodiversity project sites. Maludam National Park & Tanjung Baru Forest, Selangor, Malaysia
Paramananthan S (2012b) Report of Soils of the MPOB’s biodiversity study area proposed Cermat Ceria and Durafarm plantations. Selangor, Malaysia
Pitt JI (1979) The genus Penicillium and teleomorphic states Eupenicillium and Talaromyces. Academic Press Inc., London
Ramlah Ali SA, Sakinah S, Mohd Shawal Thakib M, Shamsilawani AB, Nur Aziemah AG (2016) Soil fungal community associated with peat in Sarawak identified using 18 rDNA marker. J Oil Palm Res 28:161–171
Rinu K, Pandey A (2010) Temperature-dependent phosphate solubilization by cold- and pH-tolerant species of Aspergillus isolated from Himalayan soil. Mycoscience 51:263–271
Rusli MH, Idris AS, Cooper RM (2015) Evaluation of Malaysian oil palm progenies for susceptibility, resistance or tolerance to Fusarium oxysporum f. sp. elaeidis and defence-related gene expression in roots. Plant Pathol 64:638–647
Rydin H, Jeglum JK (2013) The biology of peatlands. Oxford University Press, UK
Sanders IR, Rodriguez A (2016) Aligning molecular studies of mycorrhizal fungal diversity with ecologically important levels of diversity in ecosystems. ISME J 10:2783–2786
Schwarze FWMR, Jauss F, Spencer C, Hallam C, Schubert M (2012) Evaluation of an antagonistic Trichoderma strain for reducing the rate of wood decomposition by the white rot fungus Phellinus noxius. Biol Control 61:160–168
Seifert KA (2009) Progress towards DNA barcoding of fungi. Mol Ecol Resour 9:83–89
Shuhada SN, Sabiha S, Zubaid A, Azhar B (2017) Logged peat swamp forest supports greater macrofungal biodiversity than large scale oil palm plantations and smallholdings. Ecol Evol 7:7187–7200
Siles JA, Margesin R (2016) Abundance and diversity of bacterial, archaeal, and fungal communities along an altitudinal gradient in Alpine forest soils: what are the driving factors? Microb Ecol 72:207–220
Smit E, Leeflang P, Glandorf B, Dirk JVE, Wernars K (1999) Analysis of fungal diversity in the wheat rhizosphere by sequencing of cloned PCR-amplified genes encoding 18S rRNA and temperature gradient gel electrophoresis. Appl Environ Microbiol 65:2614–2621
Stokland JN, Larsson K (2011) Legacies from natural forest dynamics: different effects of forest management on wood-inhabiting fungi in pine and spruce forests. For Ecol Manag 261:1707–1721
Terhonen E, Keriö S, Sun H, Asiegbu FO (2014) Endophytic fungi of Norway spruce roots in boreal pristine mire, drained peatland and mineral soil and their inhibitory effect on Heterobasidion parviporum in vitro. Fungal Ecol 9:17–26
Thormann MN (2006) Diversity and function of fungi in peatlands: a carbon cycling perspective. Can J Soil Sci 5:281–293
Toma Y, Takakai F, Darung U, Kuramochi K, Suwido HL, Dohong S, Hatano R (2011) Nitrous oxide emission derived from soil organic matter decomposition from tropical agricultural peat soil in central Kalimantan, Indonesia. Soil Sci Plant Nutr 57:436–451
Treseder KK, Holden SR (2013) Fungal carbon sequestration. Science 339:1528–1529
Turrini A, Agnolucci M, Palla M, Tomé E, Tagliavini M, Scandellari F, Giovannetti M (2017) Species diversity and community composition of native arbuscular mycorrhizal fungi in apple roots are affected by site and orchard management. Appl Soil Ecol 116:42–54
Tveit A, Schwacke R, Svenning MM, Urich T (2013) Organic carbon transformations in high-Arctic peat soils: key functions and microorganisms. ISME J 7:299–311
Vieira FCS, Nahas EÃ (2005) Comparison of microbial numbers in soils by using various culture media and temperatures. Microbiol Res 160:197–202
Wagner L, Stielow B, Hoffmann K, Petkovits T, Papp T, Hoog GSD, Verkley G, Voigt K (2013) A comprehensive molecular phylogeny of the Mortierellales (Mortierellomycotina) based on nuclear ribosomal DNA. Persoonia 30:77–93
Watanabe T (2002) Pictorial atlas of soil and seed fungi. CRC Press, Boca Raton
Westphalen MC, Silveira RMB (2012) Resupinate polypores from mixed ombrophilous forests in southern Brazil. Mycotaxon 122:111–122
Winsborough C, Basiliko N (2010) Fungal and bacterial activity in northern peatlands. Geomicrobiol J 27:315–320
Yilmaz N, Visagie CM, Houbraken J, Frisvad JC, Samson RA (2014) Polyphasic taxonomy of the genus Talaromyces. Stud Mycol 78:175–341
Yule CM (2010) Loss of biodiversity and ecosystem functioning in Indo–Malayan peat swamp forests. Biodivers Conserv 19:393–409
Zak DR, Holmes WE, White DC, Peacock AD, Tilman D (2003) Plant diversity, soil microbial communities, and ecosystem function: are there any links? Ecology 84:2042–2050
Zinck JA (2011) Tropical and subtropical peats: an overview. In: Zinck JA, Huber O (eds) Peatlands of the Western Guayana Highlands, Venezuela. Springer, Berlin, pp 5–28
Acknowledgements
The authors would like to thank Datuk Dr. Ahmad Kushairi Din, the Director-General of Malaysian Palm Oil Board (MPOB) and Dr. Norman Kamaruddin, the Director of Biological Research Division MPOB for the permission to publish this article. The authors also thank Dr. Mohd Haniff Harun, the Head of TROPI Unit, MPOB for constructive comments. This research was supported by the MPOB Research Fund (R009711000).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kusai, N.A., Ayob, Z., Maidin, M.S.T. et al. Characterization of fungi from different ecosystems of tropical peat in Sarawak, Malaysia. Rend. Fis. Acc. Lincei 29, 469–482 (2018). https://doi.org/10.1007/s12210-018-0685-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12210-018-0685-8